Abstract
The Cancer Genome Atlas effort has generated significant interest in a new paradigm shift in tumor tissue analysis, patient diagnosis and subsequent treatment decision. Findings have highlighted the limitation of sole reliance on histology, which can be confounded by inter-observer variability. Such studies demonstrate that histologically similar grade IV brain tumors can be divided into four molecular subtypes based on gene expression, with each subtype demonstrating unique genomic aberrations and clinical outcome. These advances indicate that curative therapeutic strategies must now take into account the molecular information in tumor tissue, with the goal of identifying molecularly stratified patients that will most likely to receive treatment benefit from targeted therapy. This in turn spares non-responders from chemotherapeutic side effects and financial costs. In advancing clinical stage drug candidates, the banking of brain tumor tissue necessitates the acquisition of not just tumor tissue with clinical history and robust follow-up, but also high quality molecular information such as somatic mutation, transcriptomic and DNA methylation profiles which have been shown to predict patient survival independent of current clinical indicators. Additionally, the derivation of cell lines from such tumor tissue facilitates the development of clinically relevant patient-derived xenograft mouse models that can prospectively reform the tumor for further studies, yet have retrospective clinical history to associate bench and in vivo findings with clinical data. This represents a core capability of Precision Medicine where the focus is on understanding inter- and intra-tumor heterogeneity so as to best tailor therapies that will result in improved treatment outcomes.
*Carol Tang and Beng Ti Christopher Ang are Co-corresponding authors for this chapter.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Glioblastoma
- Histology
- Glioma-propagating cells
- TCGA
- Patient-derived xenografts
- Tumor resource
- Precision medicine
- Connectivity map
- Patient stratification
- Bioinformatics
4.1 Introduction
Glioblastoma multiforme (GBM) remains a cancer with the worst prognosis. Patients often show a median survival period of 15 months, even with advanced surgical intervention, chemotherapy and radiation treatment [1]. Temozolomide, the standard of care drug in the clinic, has annual global sales of US$ 1 billion, yet it merely extends survival by 3 months. Among the reasons for the highly invasive and recurrent nature of the disease lies in the cellular and molecular heterogeneity of GBM. Recent deep molecular profiling efforts such as The Cancer Genome Atlas (TCGA) demonstrated that histologically identical GBM tumors are molecularly heterogeneous, further suggesting that their regulatory pathway networks determine each tumor’s sensitivity to targeted therapeutic approaches [2, 3]. While these computational analyses reveal the putative mechanisms underlying tumor resistance and recurrence, biological or functional validation in preclinical animal models is lacking. In the past decade, the use of mouse xenograft models created from commercially procured serum-grown glioma cells has been challenged by studies demonstrating that such xenografted tumors fail to recapitulate the patient’s original tumor morphology and transcriptomic profiles [4]. In addition, such in vitro serially passaged serum-grown cells often contain genomic aberrations not found in the original primary tumor [5, 6]. Indeed, the widely used US National Cancer Institute NCI-60 panel of human cancer cell lines commonly passaged in serum-containing media will soon be decommissioned, with an aim to launch a rejuvenated repository of cancer models that are derived from fresh patient samples and tagged with details about their clinical past [7].
Glioma-propagating cells (GPCs) have been isolated from malignant brain tumors, and cultivated as spheroid structures in serum-free media supplemented with growth factors [8]. This media composition is similar to that used to passage neural stem cells , and helps promote self-renewal and tumorigenicity of GPCs [9]. In contrast, the addition of serum encourages differentiation of GPCs, resulting in cessation of cell proliferation with subsequent involution of tumor growth. We and others previously demonstrated the preservation of karyotypic hallmarks in these cells, similar to the original tumors [6, 10]. We developed a method to passage these spheroid structures by mechanical trituration, thus avoiding constant use of harsh enzymatic solutions that have been shown to alter karyotypic patterns in human embryonic stem cells grown as embryoid bodies [11]. We discuss vitrification , a technique adapted from in vitro fertilization procedures, and emphasize the importance of preserving these spheroid structures with reduced water content to prevent damage from ice crystals during the thawing process. Collectively, these methodologies preserve the integrity of GPCs and their ability to establish patient-derived xenograft (PDX) tumors that faithfully phenocopy the original tumor pathophysiology, cytogenetic and transcriptomic profiles. This capability reflects the importance of establishing a cell line and tumor tissue bank that presents the most clinically relevant resource to test and validate computational predictions generated from large patient glioma databases.
Crucial to establishing clinical relevance of our brain tumor resource , we discuss computational platforms such as the Connectivity Map to link molecular data acquired from in vitro and animal studies with multi-dimensional clinical information such as the patient’s age, tumor grade, molecular information, Karnofsky score and magnetic resonance imaging (MRI) scans available in large clinical databases such as TCGA [12]. These computational advances have expanded the scope of studies made possible with our brain tumor resource [13–17]. Importantly, we now have a brain tumor biobank that facilitates studies in Precision Medicine where biological validation of patient-centric predictions is achievable.
4.2 Brain Tumor Resource
With the advance of TCGA efforts, several brain tumor banks containing low-passage cell lines have been established with a focus on acquisition of clinical history with deep content molecular information (genotype-phenotype databases). The goal of such tumor banks is to facilitate studies requiring capability to remodel the disease accurately so as to prospectively test computational predictions based on patient information [18]. Central to this biobank creation, we previously demonstrated three important criteria: (1) Growth of patient-derived glioma cells in serum-free media supplemented with growth factors, (2) Vitrification as a cryopreservation method, and (3) Ability to establish orthotopic PDX models that recapitulate the patient’s original tumor pathophysiology. As such, genotype–phenotype databases previously needed only simple computing technologies, including very basic data fields relating to pathogenicity, but did not capture the process of pathogenicity interpretation. Going forward, this approach will have to change, especially if we wish to deliver truly Precision Medicine-based findings, which will require mechanistic in addition to probabilistic modeling, and hence even more sophisticated sources of input information and tools for the recording of results.
GPCs can be maintained and propagated as tumor neurosphere cultures in defined serum-free condition supplemented with epidermal growth factor (EGF) and basic fibroblast growth factor (bFGF), a paradigm that is adopted from the traditional neurosphere culture [19, 20]. Furthermore, Lee and colleagues have shown that tumor stem-like cells grown in serum-free condition closely mimic the genotype, transcriptomic profile and morphological features of their parental tumors [6]. Thus, the establishment of a tumor neurosphere repository with preservation of essential features of tumor heterogeneity would provide a clinically relevant resource to investigate the disease. Such a method would also allow us to return to the same experimental cell line passages to reduce variability in experimental replication. Recent efforts by the Roadmap Epigenomics Project spearheaded by the National Institutes of Health (NIH) highlighted epigenetic silencing differences of in vitro cultured cells when compared to similar cell lineages in primary tissue, thus underscoring the importance of cryopreserving low-passage GPCs with retention of its molecular fingerprint [21]. In many studies involving the prospective isolation of tumor-propagating cells, only small amounts of clinical material are available, and this limitation is compounded by lack of appropriate methods to preserve such cells at convenient time points. Although in vivo serial passage of GPC-derived tumors has been described as the most reliable method to preserve the cells, practically, it is not always feasible to have access to suitably-aged immunocompromised mice [8]. We developed mechanical trituration as a method to passage three-dimensional spheroid cultures for the reason that acute dissociation of such structures into single cells promotes cellular senescence over an extended period [9]. In addition, induction of cellular differentiation, such as by the presence of serum, results in loss of tumorigenicity and consequent involution of tumor growth [22].
We adapted a method used in cryopreservation of embryos from in vitro fertilization procedures. Vitrification is a process of glass-like solidification in which an aqueous solution is prevented from crystallization by rapid cooling [23]. This method has been commonly used for the cryopreservation of embryos at different developmental stages from various species such as murine, rabbit, sheep and bovine [24–29]. Furthermore, human and mouse multi-cell embryos have been successfully cryopreserved using this strategy [30]. This highlights the feasibility of cryopreserving cellular aggregates. In addition, it has been demonstrated that vitrified embryonic stem cells retained their pluripotency, cytogenetic profile and viability upon thawing [31]. Taken together, vitrification could provide an effective means of storage of brain tumor-propagating cells cultured as spherical structures. Although adherent GPC cultures using laminin have been proposed, these growth conditions resulted in transcriptomic shift of the cells [32]. In support, Dirks and colleagues showed that a chemical genetics screen utilizing GPC spheroid cultures identified small molecules affecting neurotransmission in the adult central nervous system (CNS), thus suggesting that clinically approved neuromodulators may remodel the mature CNS and find application in the treatment of brain cancer [33].
To facilitate testing of patient stratification methods and identification of molecular mechanisms contributing to disease progression, a genomic roadmap is created to characterize the cells, tumor xenografts and primary tumors. The pivotal goal of molecular profiling of patient-derived cells aims to stratify patient cohorts and identify amenable therapeutic strategies. The content-rich patient tumor molecular profiles need to be systematically archived for efficient data mining to evaluate proof-of-concept studies. Initial steps at identifying potential cancer-specific biomarkers require patient-centric bioinformatics interrogation, information from which can then be used for the analysis of samples stored in tissue banks. In TCGA , the quality of samples acquired was assessed from several participating tissue banks that surprisingly showed only one percent of the samples being reliable for downstream molecular data acquisition [2, 34, 35]. Sample quality and the associated clinical information are important factors in tissue banking. The importance in having expertise at tissue sampling and culturing of patient-derived cells, with preservation of cytogenetic and transcriptomic hallmarks found in the original primary tumor has previously been reported by our colleagues [6, 10]. Precision Medicine-driven studies, dependent on this molecular heterogeneity, warrant a preclinical mouse model that recapitulates the patient’s original tumor pathophysiology and aids at advancing chemotherapeutic candidates into clinical trials .
4.3 Animal Model Established from the Biobank : An Informative Preclinical Mouse Model
Modeling brain tumors in animals reveals genetic events and molecular mechanisms that contribute to oncogenesis. The mouse shares extensive molecular and physiological similarities to human beings and is a powerful tool for studying cancer [36]. Unlike invertebrate model systems, tumor development in mice is accompanied by other complex processes such as angiogenesis and metastasis, similar to those in human cancer [37]. More importantly, mouse tumor models provide a temporal perspective and genetically-controlled system for studying the tumorigenic process, as well as response to specific therapies.
Genetically engineered mouse models (GEMMs) are a popular model to study tumor biology [37]. There are several limitations to using GEMMs as more than one driver mutation is required to initiate tumorigenesis [38, 39]. The expression of the transgene is often elevated to levels that exceed those in patients. Tumors that arise in this model are often sporadic, resulting in difficulty of study designs that require significant animal numbers for reproducibility. TCGA efforts have also demonstrated that the spectrum of driver mutations differs significantly among patients, thus the relevance of a particular GEMM may be limited [2, 40]. Nevertheless, GEMMs will likely provide useful insight into the tumor cell-of-origin and initiating events.
Xenograft models established from commercially procured serum-grown cell lines date back to the late 1960s. However, several studies demonstrated that such xenografted tumors exhibit significant morphological and molecular features not found in the original primary tissue [4, 6]. Such issues have subsequently been overcome through the development of PDX models. Several studies revealed that the orthotopic xenograft model established from patient-derived glioma cell lines or tumor explants bear more clinical relevance [41–43]. This is due to the presence of the microenvironment provided by the normal brain parenchyma, where better measurement of drug delivery and clearance kinetics can be evaluated. The orthotopic PDX model is useful as tumor formation with high incidence and the ability to generate large cohorts of animals in preclinical studies are attainable. Several investigators have provided evidence that PDX tumors phenocopy the pathophysiology and molecular features of their parental tumor [6, 44]. Importantly, this model allows assessment of therapeutic responses using stratified clinical material, a core capability of Precision Medicine. We and others previously integrated the use of such an in-house brain tumor resource to interrogate lab findings in clinical glioma databases [7, 10, 14–17]. We showed that transcriptomic patterns derived from in vitro drug-treated or genetically manipulated cells mapped to patient clinical databases, and dictated primary tumor phenotype.
Despite the frequent use of the PDX model in studying drug therapy response, we recognize that immunogenic and microenvironmental factors may not be fully represented. Additionally, engraftment inefficiency can be as high as 90 %, depending on the type of cancer [45]. Such limitations suggest that the use of mouse models should be carefully considered to provide maximum information about the study question .
4.4 Enabling Precision Medicine-Based Studies: Highlighting the Importance of a Biobank
A brain tumor biobank that merges multi-platform data from patient material (cells, xenograft and primary tumors) and seeks to address clinical and imaging phenotypes with molecular data will advance studies in stratified medicine. The development and preclinical validation of novel anti-cancer drugs require low-passage cell lines that are representative, scalable and reproducible in experimental models. In the absence of an integrated human brain biobank, research findings from in vitro and in vivo models of neurological disorders cannot be functionally validated in the actual disease context. TCGA efforts revealed that gene expression drives GBM disease progression and survival outcome [3, 46–48]. Although an important prognostic factor of glioma progression relies on the World Health Organization (WHO) grading scheme, the wide differences in treatment response and survival suggest that the aggressiveness of treatment cannot be decided just by histology alone. These findings underscore the limitation of relying solely on morphological criteria to diagnose patients. Future brain tumor classification must now include molecular information which in turn guides diagnosis and subsequent treatment decision [49]. Development and maintenance of biobanks as an international resource for the study of human diseases provides the scientific community with well-characterized cells and rich phenotypic data. Such resources facilitate prospective remodeling of the disease in mouse models, with retrospective clinical information to evaluate correlation patterns and directly validate the mechanism.
Targeted therapy is an attractive approach to overcome the highly infiltrative and recurrent nature of GBM. Recent characterization of the epigenome, somatic mutation profile and transcriptome of tumor tissue has now provided a deeper understanding of the alterations underlying the disease phenotype [2, 3, 50, 51]. Tumor cells are assessed for the underlying pattern using unsupervised computational approaches to discern their molecular heterogeneity. An important evaluation is to computationally identify regulatory pathways that can be targeted with small molecule drugs. These predictions are then functionally validated in PDX models. Transcriptomic resources of xenograft and primary tumors are scrutinized for highly variable genes which can reflect a common bias present among primary tumors and xenografts established from GPCs . This common and systematic bias can be controlled and nullified by statistical algorithms such as Anova-based batch effect removal and principal component based analysis [52, 53]. A scatter plot accounting for major principal components across matched primary and orthotopic tumors demonstrates close proximity of matched samples, indicating that GPCs can recreate the original tumor molecular profile in vivo (Fig. 4.1).
Recapturing molecular portrait of primary tumors in orthotopic xenograft tumors derived from GPCs . Principal Component Analysis (PCA) map demonstrates similar transcriptomic profiles between matched xenograft and primary tumors. Each color denotes similar patient material with corresponding xenograft tumor. Triangle, xenograft tumor; square, patient tumor
Computational evaluation of matched molecular data from biobanked patient tumors and GPCs requires systematic validation of molecular profiles using bioinformatics approaches. A computational pipeline is required to interrogate gene signatures (transcriptomic classifiers) derived from GPCs and a large set of independent predictive database collection from patients’ molecular data [12, 54, 55]. Molecular perturbation experiments on GPCs have demonstrated promising evidence in conferring the patient’s prognosis in predictive databases and in vivo experiments [13, 15]. The most comprehensive glioma patient’s database was established by TCGA , where key components of clinical phenotype and genomics information were catalogued into a multi-tier organizational structure for 33 different tumor types [56]. Each tier is confined with a data structure from genomic platforms including somatic mutation, copy number, methylation, transcriptomic and proteomic technologies. Importantly, we have merged our cell and xenograft tumor molecular patterns with international collections by adapting the statistical framework to control for systematic batch effects across different collections [52, 53]. This integrated evaluation also confirms the consistency across tumor cell types and provides greater statistical power [57]. We have successfully adapted a qualitative enrichment pipeline, the Connectivity Map (CMAP) to interrogate active pathway programs coded as gene signatures in our biobanked cells with our patients’ transcriptomic patterns [12, 13, 17]. These patients’ prognoses can then be retrospectively predicted by mapping survival and clinical response parameters with enrichment scores from the CMAP pipeline. Thus, molecular perturbation experiments tapping into our brain tumor biobank serve as effective tools to biologically validate these patient-centric computational predictions .
4.5 Conclusion
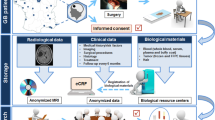
Biobanking coupled with deep molecular characterization is a core capability for Precision Medicine-based studies (Fig. 4.2). The ability to remodel brain tumors in mice facilitates biological and functional validation of computationally predicted pathway networks that should be therapeutically targeted.
Abbreviations
- bFGF:
-
Basic fibroblast growth factor
- CMAP:
-
Connectivity map
- CNS:
-
Central nervous system
- EGF:
-
Epidermal growth factor
- GBM:
-
Glioblastoma multiforme
- GEMM:
-
Genetically engineered mouse model
- GPCs:
-
Glioma-propagating cells
- MRI:
-
Magnetic resonance imaging
- NIH:
-
National Institutes of Health
- PDX:
-
Patient-derived xenograft
- TCGA:
-
The Cancer Genome Atlas
References
Louis DN, Ohgaki H, Wiestler OD, Cavenee WK, Burger PC, Jouvet A, Scheithauer BW, Kleihues P (2007) The 2007 WHO classification of tumours of the central nervous system. Acta Neuropathol 114(2):97–109
Cancer Genome Atlas Research N (2008) Comprehensive genomic characterization defines human glioblastoma genes and core pathways. Nature 455(7216):1061–1068
Verhaak RG, Hoadley KA, Purdom E, Wang V, Qi Y, Wilkerson MD, Miller CR, Ding L, Golub T, Mesirov JP, Alexe G, Lawrence M, O’Kelly M, Tamayo P, Weir BA, Gabriel S, Winckler W, Gupta S, Jakkula L, Feiler HS, Hodgson JG, James CD, Sarkaria JN, Brennan C, Kahn A, Spellman PT, Wilson RK, Speed TP, Gray JW, Meyerson M, Getz G, Perou CM, Hayes DN, Cancer Genome Atlas Research N (2010) Integrated genomic analysis identifies clinically relevant subtypes of glioblastoma characterized by abnormalities in PDGFRA, IDH1, EGFR, and NF1. Cancer Cell 17(1):98–110
Hodgson JG, Yeh RF, Ray A, Wang NJ, Smirnov I, Yu M, Hariono S, Silber J, Feiler HS, Gray JW, Spellman PT, Vandenberg SR, Berger MS, James CD (2009) Comparative analyses of gene copy number and mRNA expression in glioblastoma multiforme tumors and xenografts. Neuro Oncol 11(5):477–487
Clark MJ, Homer N, O’Connor BD, Chen Z, Eskin A, Lee H, Merriman B, Nelson SF (2010) U87MG decoded: the genomic sequence of a cytogenetically aberrant human cancer cell line. PLoS Genet 6(1):e1000832
Lee J, Kotliarova S, Kotliarov Y, Li A, Su Q, Donin NM, Pastorino S, Purow BW, Christopher N, Zhang W, Park JK, Fine HA (2006) Tumor stem cells derived from glioblastomas cultured in bFGF and EGF more closely mirror the phenotype and genotype of primary tumors than do serum-cultured cell lines. Cancer Cell 9(5):391–403
Ledford H (2016) US cancer institute to overhaul tumour cell lines. Nature 530(7591):391
Galli R, Binda E, Orfanelli U, Cipelletti B, Gritti A, De Vitis S, Fiocco R, Foroni C, Dimeco F, Vescovi A (2004) Isolation and characterization of tumorigenic, stem-like neural precursors from human glioblastoma. Cancer Res 64(19):7011–7021
Svendsen CN, ter Borg MG, Armstrong RJ, Rosser AE, Chandran S, Ostenfeld T, Caldwell MA (1998) A new method for the rapid and long term growth of human neural precursor cells. J Neurosci Methods 85(2):141–152
Chong YK, Toh TB, Zaiden N, Poonepalli A, Leong SH, Ong CE, Yu Y, Tan PB, See SJ, Ng WH, Ng I, Hande MP, Kon OL, Ang BT, Tang C (2009) Cryopreservation of neurospheres derived from human glioblastoma multiforme. Stem Cells 27(1):29–39
Pera MF (2004) Unnatural selection of cultured human ES cells? Nat Biotechnol 22(1):42–43
Lamb J, Crawford ED, Peck D, Modell JW, Blat IC, Wrobel MJ, Lerner J, Brunet JP, Subramanian A, Ross KN, Reich M, Hieronymus H, Wei G, Armstrong SA, Haggarty SJ, Clemons PA, Wei R, Carr SA, Lander ES, Golub TR (2006) The connectivity map: using gene-expression signatures to connect small molecules, genes, and disease. Science 313(5795):1929–1935
Chong YK, Sandanaraj E, Koh LW, Thangaveloo M, Tan MS, Koh GR, Toh TB, Lim GG, Holbrook JD, Kon OL, Nadarajah M, Ng I, Ng WH, Tan NS, Lim KL, Tang C, Ang BT (2016) ST3GAL1-associated transcriptomic program in glioblastoma tumor growth, invasion, and prognosis. J Natl Cancer Inst 108(2)
Choudhury Y, Tay FC, Lam DH, Sandanaraj E, Tang C, Ang B-T, Wang S (2012) Attenuated adenosine-to-inosine editing of microRNA-376a* promotes invasiveness of glioblastoma cells. J Clin Invest 122(11):4059–4076
Koh LW, Koh GR, Ng FS, Toh TB, Sandanaraj E, Chong YK, Phong M, Tucker-Kellogg G, Kon OL, Ng WH, Ng IH, Clement MV, Pervaiz S, Ang BT, Tang CS (2013) A distinct reactive oxygen species profile confers chemoresistance in glioma-propagating cells and associates with patient survival outcome. Antioxid Redox Signal 19(18):2261–2279
Ng FS, Toh TB, Ting EH, Koh GR, Sandanaraj E, Phong M, Wong SS, Leong SH, Kon OL, Tucker-Kellogg G, Ng WH, Ng I, Tang C, Ang BT (2012) Progenitor-like traits contribute to patient survival and prognosis in oligodendroglial tumors. Clin Cancer Res 18(15):4122–4135
Yeo CWS, Ng FSL, Chai C, Tan JMM, Koh GRH, Chong YK, Koh LWH, Foong CSF, Sandanaraj E, Holbrook JD, Ang B-T, Takahashi R, Tang C, Lim K-L (2012) Parkin pathway activation mitigates glioma cell proliferation and predicts patient survival. Cancer Res 72(10):2543–2553
Laks DR, Crisman TJ, Shih MYS, Mottahedeh J, Gao F, Sperry J, Garrett MC, Yong WH, Cloughesy TF, Liau LM, Lai A, Coppola G, Kornblum HI (2016) Large-scale assessment of the gliomasphere model system. Neuro-Oncology
Reynolds B, Weiss S (1992) Generation of neurons and astrocytes from isolated cells of the adult mammalian central nervous system. Science 255(5052):1707–1710
Reynolds BA, Weiss S (1996) Clonal and population analyses demonstrate that an EGF-responsive mammalian embryonic CNS precursor is a stem cell. Dev Biol 175(1):1–13
Roadmap Epigenomics C, Kundaje A, Meuleman W, Ernst J, Bilenky M, Yen A, Heravi-Moussavi A, Kheradpour P, Zhang Z, Wang J, Ziller MJ, Amin V, Whitaker JW, Schultz MD, Ward LD, Sarkar A, Quon G, Sandstrom RS, Eaton ML, Wu YC, Pfenning AR, Wang X, Claussnitzer M, Liu Y, Coarfa C, Harris RA, Shoresh N, Epstein CB, Gjoneska E, Leung D, Xie W, Hawkins RD, Lister R, Hong C, Gascard P, Mungall AJ, Moore R, Chuah E, Tam A, Canfield TK, Hansen RS, Kaul R, Sabo PJ, Bansal MS, Carles A, Dixon JR, Farh KH, Feizi S, Karlic R, Kim AR, Kulkarni A, Li D, Lowdon R, Elliott G, Mercer TR, Neph SJ, Onuchic V, Polak P, Rajagopal N, Ray P, Sallari RC, Siebenthall KT, Sinnott-Armstrong NA, Stevens M, Thurman RE, Wu J, Zhang B, Zhou X, Beaudet AE, Boyer LA, De Jager PL, Farnham PJ, Fisher SJ, Haussler D, Jones SJ, Li W, Marra MA, McManus MT, Sunyaev S, Thomson JA, Tlsty TD, Tsai LH, Wang W, Waterland RA, Zhang MQ, Chadwick LH, Bernstein BE, Costello JF, Ecker JR, Hirst M, Meissner A, Milosavljevic A, Ren B, Stamatoyannopoulos JA, Wang T, Kellis M (2015) Integrative analysis of 111 reference human epigenomes. Nature 518(7539):317–330
Piccirillo SGM, Reynolds BA, Zanetti N, Lamorte G, Binda E, Broggi G, Brem H, Olivi A, Dimeco F, Vescovi AL (2006) Bone morphogenetic proteins inhibit the tumorigenic potential of human brain tumour-initiating cells. Nature 444(7120):761–765
Rall WF, Wood MJ, Kirby C, Whittingham DG (1987) Development of mouse embryos cryopreserved by vitrification. J Reprod Fertil 80(2):499–504
Ali J, Shelton JN (1993) Design of vitrification solutions for the cryopreservation of embryos. J Reprod Fertil 99(2):471–477
Kasai M, Komi JH, Takakamo A, Tsudera H, Sakurai T, Machida T (1990) A simple method for mouse embryo cryopreservation in a low toxicity vitrification solution, without appreciable loss of viability. J Reprod Fertil 89(1):91–97
Kasai M, Nishimori M, Zhu SE, Sakurai T, Machida T (1992) Survival of mouse morulae vitrified in an ethylene glycol-based solution after exposure to the solution at various temperatures. Biol Reprod 47(6):1134–1139
Saha S, Otoi T, Takagi M, Boediono A, Sumantri C, Suzuki T (1996) Normal calves obtained after direct transfer of vitrified bovine embryos using ethylene glycol, trehalose, and polyvinylpyrrolidone. Cryobiology 33(3):291–299
Ali J, Shelton JN (1993) Successful vitrification of day-6 sheep embryos. J Reprod Fertil 99(1):65–70
Kasai M, Hamaguchi Y, Zhu SE, Miyake T, Sakurai T, Machida T (1992) High survival of rabbit morulae after vitrification in an ethylene glycol-based solution by a simple method. Biol Reprod 46(6):1042–1046
Mukaida T, Wada S, Takahashi K, Pedro PB, An TZ, Kasai M (1998) Vitrification of human embryos based on the assessment of suitable conditions for 8-cell mouse embryos. Hum Reprod 13(10):2874–2879
Reubinoff BE, Pera MF, Vajta G, Trounson AO (2001) Effective cryopreservation of human embryonic stem cells by the open pulled straw vitrification method. Hum Reprod 16(10):2187–2194
Pollard SM, Yoshikawa K, Clarke ID, Danovi D, Stricker S, Russell R, Bayani J, Head R, Lee M, Bernstein M, Squire JA, Smith A, Dirks P (2009) Glioma stem cell lines expanded in adherent culture have tumor-specific phenotypes and are suitable for chemical and genetic screens. Cell Stem Cell 4(6):568–580
Diamandis P, Wildenhain J, Clarke ID, Sacher AG, Graham J, Bellows DS, Ling EKM, Ward RJ, Jamieson LG, Tyers M, Dirks PB (2007) Chemical genetics reveals a complex functional ground state of neural stem cells. Nat Chem Biol 3(5):268–273
Scott CT, Caulfield T, Borgelt E, Illes J (2012) Personal medicine -the new banking crisis. Nat Biotech 30(2):141–147
Suh KS, Sarojini S, Youssif M, Nalley K, Milinovikj N, Elloumi F, Russell S, Pecora A, Schecter E, Goy A (2013) Tissue banking, bioinformatics, and electronic medical records: the front-end requirements for personalized medicine. J Oncol 2013:368751
Rosenthal N, Brown S (2007) The mouse ascending: perspectives for human-disease models. Nat Cell Biol 9(9):993–999
Wee B, Charles N, Holland EC (2011) Animal models to study cancer-initiating cells from glioblastoma. Front Biosci (Landmark Ed) 16:2243–2258
Alcantara Llaguno S, Chen J, Kwon C-H, Jackson EL, Li Y, Burns DK, Alvarez-Buylla A, Parada LF (2009) Malignant astrocytomas originate from neural stem/progenitor cells in a somatic tumor suppressor mouse model. Cancer Cell 15(1):45–56
Zheng H, Ying H, Yan H, Kimmelman AC, Hiller DJ, Chen AJ, Perry SR, Tonon G, Chu GC, Ding Z, Stommel JM, Dunn KL, Wiedemeyer R, You MJ, Brennan C, Wang YA, Ligon KL, Wong WH, Chin L, DePinho RA (2008) p53 and Pten control neural and glioma stem/progenitor cell renewal and differentiation. Nature 455(7216):1129–1133
Network TC (2013) Corrigendum: comprehensive genomic characterization defines human glioblastoma genes and core pathways. Nature 494(7438):506
Shapiro WR, Basler GA, Chernik NL, Posner JB (1979) Human brain tumor transplantation into nude mice. J Natl Cancer Inst 62(3):447–453
Kaye AH, Morstyn G, Gardner I, Pyke K (1986) Development of a xenograft glioma model in mouse brain. Cancer Res 46(3):1367–1373
Horten BC, Basler GA, Shapiro WR (1981) Xenograft of human malignant glial tumors into brains of nude mice. A histopatholgical study. J Neuropathol Exp Neurol 40(5):493–511
Joo KM, Kim J, Jin J, Kim M, Seol HJ, Muradov J, Yang H, Choi YL, Park WY, Kong DS, Lee JI, Ko YH, Woo HG, Lee J, Kim S, Nam DH (2013) Patient-specific orthotopic glioblastoma xenograft models recapitulate the histopathology and biology of human glioblastomas in situ. Cell Rep 3(1):260–273
Herter-Sprie GS, Kung AL, Wong KK (2013) New cast for a new era: preclinical cancer drug development revisited. J Clin Invest 123(9):3639–3645
Gravendeel LA, Kouwenhoven MC, Gevaert O, de Rooi JJ, Stubbs AP, Duijm JE, Daemen A, Bleeker FE, Bralten LB, Kloosterhof NK, De Moor B, Eilers PH, van der Spek PJ, Kros JM, Sillevis Smitt PA, van den Bent MJ, French PJ (2009) Intrinsic gene expression profiles of gliomas are a better predictor of survival than histology. Cancer Res 69(23):9065–9072
Li A, Walling J, Ahn S, Kotliarov Y, Su Q, Quezado M, Oberholtzer JC, Park J, Zenklusen JC, Fine HA (2009) Unsupervised analysis of transcriptomic profiles reveals six glioma subtypes. Cancer Res 69(5):2091–2099
Phillips HS, Kharbanda S, Chen R, Forrest WF, Soriano RH, Wu TD, Misra A, Nigro JM, Colman H, Soroceanu L, Williams PM, Modrusan Z, Feuerstein BG, Aldape K (2006) Molecular subclasses of high-grade glioma predict prognosis, delineate a pattern of disease progression, and resemble stages in neurogenesis. Cancer Cell 9(3):157–173
Louis DN, Perry A, Burger P, Ellison DW, Reifenberger G, von Deimling A, Aldape K, Brat D, Collins VP, Eberhart C, Figarella-Branger D, Fuller GN, Giangaspero F, Giannini C, Hawkins C, Kleihues P, Korshunov A, Kros JM, Beatriz Lopes M, Ng HK, Ohgaki H, Paulus W, Pietsch T, Rosenblum M, Rushing E, Soylemezoglu F, Wiestler O, Wesseling P, International Society of N-H (2014) International Society of Neuropathology – Haarlem consensus guidelines for nervous system tumor classification and grading. Brain Pathol 24(5):429–435
Brennan CW, Verhaak RG, McKenna A, Campos B, Noushmehr H, Salama SR, Zheng S, Chakravarty D, Sanborn JZ, Berman SH, Beroukhim R, Bernard B, Wu CJ, Genovese G, Shmulevich I, Barnholtz-Sloan J, Zou L, Vegesna R, Shukla SA, Ciriello G, Yung WK, Zhang W, Sougnez C, Mikkelsen T, Aldape K, Bigner DD, Van Meir EG, Prados M, Sloan A, Black KL, Eschbacher J, Finocchiaro G, Friedman W, Andrews DW, Guha A, Iacocca M, O’Neill BP, Foltz G, Myers J, Weisenberger DJ, Penny R, Kucherlapati R, Perou CM, Hayes DN, Gibbs R, Marra M, Mills GB, Lander E, Spellman P, Wilson R, Sander C, Weinstein J, Meyerson M, Gabriel S, Laird PW, Haussler D, Getz G, Chin L, Network TR (2013) The somatic genomic landscape of glioblastoma. Cell 155(2):462–477
Nevo I, Woolard K, Cam M, Li A, Webster JD, Kotliarov Y, Kim HS, Ahn S, Walling J, Kotliarova S, Belova G, Song H, Bailey R, Zhang W, Fine HA (2014) Identification of molecular pathways facilitating glioma cell invasion in situ. PLoS One 9(11):e111783
Johnson WE, Li C, Rabinovic A (2007) Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics 8(1):118–127
Reese SE, Archer KJ, Therneau TM, Atkinson EJ, Vachon CM, de Andrade M, Kocher JP, Eckel-Passow JE (2013) A new statistic for identifying batch effects in high-throughput genomic data that uses guided principal component analysis. Bioinformatics 29(22):2877–2883
Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, Smyth GK (2015) Limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res 43(7):e47
Tibshirani RJ, Efron B (2002) Pre-validation and inference in microarrays. Stat Appl Genet Mol Biol 1:Article 1. doi:10.2202/1544-6115.1000
Marx V (2013) Drilling into big cancer-genome data. Nat Meth 10(4):293–297
Lottaz C, Beier D, Meyer K, Kumar P, Hermann A, Schwarz J, Junker M, Oefner PJ, Bogdahn U, Wischhusen J, Spang R, Storch A, Beier CP (2010) Transcriptional profiles of CD133+ and CD133- glioblastoma-derived cancer stem cell lines suggest different cells of origin. Cancer Res 70(5):2030–2040
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Tan, S.Y.M., Sandanaraj, E., Tang, C., Ang, B.T.C. (2016). Biobanking: An Important Resource for Precision Medicine in Glioblastoma. In: Karimi-Busheri, F., Weinfeld, M. (eds) Biobanking and Cryopreservation of Stem Cells. Advances in Experimental Medicine and Biology, vol 951. Springer, Cham. https://doi.org/10.1007/978-3-319-45457-3_4
Download citation
DOI: https://doi.org/10.1007/978-3-319-45457-3_4
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-45455-9
Online ISBN: 978-3-319-45457-3
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)