Abstract
Ageing is defined as the progressive attrition of tissue/organ function resulting in an increased susceptibility to disease and death. The DNA mutation and damage theory of ageing posits that the accrual of genetic damage over time is the underlying cause of ageing. Evidence for this theory stems from the fact that numerous human progeroid syndromes are caused by inherited defects in genome maintenance mechanisms, linking excess genetic damage with accelerated ageing. These diseases have been modelled in mice and other organisms. However, the molecular mechanism by which genomic instability drives ageing is currently not known. The nematode, C. elegans, is a genetically tractable, well-studied model organism for investigating mechanisms of ageing and DNA repair pathways identified in mammalian systems are well conserved in the worm. Furthermore, proliferating and post-mitotic cells, which have distinct responses to genomic instability, are clearly delineated in the worm. Thus worms provide an opportunity to study the importance of genomic stability in each of these compartments in the context of a whole organism. Genomic instability can interfere with transcription, trigger apoptosis, attenuate proliferative capacity, and cause metabolic changes. Here, we first examine the DNA repair pathways that are conserved between worms and mammals, with an overview of the spatial and temporal activity of each of these repair pathways. This chapter then explores evidence from studies in the nematode that genomic instability, healthspan and organismal ageing are linked.
Access provided by CONRICYT-eBooks. Download chapter PDF
Similar content being viewed by others
Keywords
11.1 DNA Damage and Repair in C. elegans
The nuclear and mitochondrial genomes are constantly exposed to damaging agents from endogenous (i.e., spontaneous, e.g., reactive oxygen species ) and exogenous (i.e., environmental, e.g., ultraviolet radiation) sources. These agents cause chemical modification of DNA, impacting chromosomal replication and transcription . DNA replication is critical during worm development and in the gonads of adult worms, while transcription is crucial in all cells throughout life. The advent of whole genome sequencing revealed that the mutation frequency in C. elegans is ~6.7 × 10−10 per nucleotide per cell division [1] whereas, in humans the mutation rate is 5.0 × 10−11 per nucleotide per cell division [2]. This suggests that even with a short lifespan , worms accumulate a significant number of mutations, analogous to humans. Additionally, single-strand DNA breaks [3] and deletions in the mitochondrial genome [4] increase with age in the worm, which is also comparable to humans (reviewed in [5–7])
The DNA repair pathways that are conserved between mammals and C. elegans include the following [8]:
-
Base excision repair (BER): BER identifies and excises subtle lesions that don’t distort the helical structure of DNA. The kinds of lesions routinely repaired by BER are abasic (AP) sites, oxidized bases, alkylated bases, deaminated bases and single-strand breaks.
-
Nucleotide excision repair (NER): NER detects and repairs numerous different types of lesions that cause helical distortion, including UV-induced (6-4) photoproducts (6-4PPs) and cyclobutane pyrimidine dimers (CPDs), bulky adducts formed by environmental agents such as a by-product of tobacco smoke BP-7,8-diol-9,10-epoxide (BPDE). NER consists of two sub-pathways: Global Genome NER (GG-NER): lesions repaired anywhere in the nuclear genome and Transcription Coupled NER (TC-NER): repair of lesions occurring in the template strand of an actively transcribed gene.
-
Interstrand cross-link repair (ICLR): ICLR or the Fanconi pathway repairs lesions that covalently link both strands of DNA together.
-
Mismatch repair (MMR): MMR is a DNA repair mechanism that is responsible for correcting base-base mismatches, insertion/deletion mismatches and small hairpin structures resulting from misalignment that occurs during DNA replication and recombination.
-
Homologous recombination (HR): HR is used to repair DNA double-strand breaks (DSBs) using a sister chromatid or homologous chromosome as a template to acquire lost sequence information. In addition to double-strand breaks, HR is needed for the repair of interstrand crosslinks (ICL) and for the recovery of stalled replication forks.
-
Non-homologous end-joining (NHEJ): DNA double-strand breaks with two broken ends are repaired by NHEJ via a mechanism by which the two ends are ligated together.
Conventionally, each of these DNA repair mechanisms is thought to tackle a specific type of DNA damage. However, overlap between these pathways is becoming increasingly evident. DNA repair is tightly regulated by DNA damage sensors (proteins that detect damaged DNA) and signal transducers (protein cascades that transmit the damage signal to numerous effector proteins that regulate repair, cell cycle and cell fate) [9]. Translesion synthesis (TLS) is a mechanism by which DNA lesions are tolerated by replicating cells, but not repaired. When the replication machinery is stalled by a DNA lesion, a specialized DNA polymerase can be recruited to enable bypass of the lesion, enabling resumption of replication [10]. TLS is also well conserved between worms and mammals [11] (see Table 11.1).
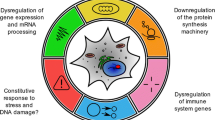
An important consideration when studying DNA damage and its role in ageing is that repair mechanisms are differentially utilized in tissues (reviewed in [12]). For instance, germ cells respond more strongly to DNA damage than somatic cells. Therefore, post-mitotic adult worms are relatively resistant to ionizing radiation, whereas germ cells are extremely sensitive. Proliferating and meiotic germ cells repair DSBs by HR, whereas, post-mitotic somatic cells utilize NHEJ. Similarly, GG-NER, BER and ICLR maintain DNA stability in the mitotic germ cell compartment. TLS is highly active during early embryonic growth, contributing to resistance to genotoxic stress during this phase of development. During development, somatic cell genome maintenance requires HR and NHEJ, whereas the post-mitotic adult is mostly dependent on TC-NER . These observations reveal complex spatial and temporal regulation of DNA repair mechanisms (Fig. 11.1).
Notably, there are a few key proteins involved in genome maintenance that appear to be lacking in the worm. γH2Ax, MDC1 and RNF8 DNA damage signalling proteins have not been identified in C. elegans. Similarly, some regulators of NHEJ and ICL repair pathway do not appear to be present in the worm (see Table 11.1). Another point to consider is that in higher organisms DNA damage can induce cellular senescence [13], which has been shown to drive ageing [14]. However, it is currently unclear if C. elegans have a cellular senescence programme [15]. Nonetheless, the importance of DNA repair mechanisms to genome stability and organismal lifespan is well documented in the worm. We focus here on each of the genome maintenance pathways and their relationship to ageing .
11.2 Changes in DNA Damage Levels and Mutation Frequency with Age
One of the earliest studies measuring DNA damage over the lifespan of worms was reported by Klass et al. in the 1980s. The authors observed a 34-fold increase in single-strand DNA breaks in day 15 adults compared with young, day 5 animals, using an Escherichia coli DNA polymerase I assay. Additionally, 5-methylcytosine (an epigenetic marker regulated to some extent by DNA repair ) was also exponentially increased in older worms compared to larvae and young adults. These changes were accompanied by reduced transcription [3]. These data are consistent with the notion that incomplete repair of DNA damage leads to damage accumulation with age.
Similarly, the oxidative DNA lesion 8-oxo-7,8-dihydro-2′-deoxyguanosine (8-oxo-dG) was measured in the short-lived mutant, mev-1 (a gene that encodes a subunit of complex II in the mitochondrial electron transport chain) [16] causing accumulation of dysfunctional mitochondria , reduced mitochondrial membrane potential [17], increased ROS, and hypersensitivity to oxidative stress [18]. A significant increase in adducts is detected in mev-1 mutants compared to wild-type worms by high-performance liquid chromatography coupled with electrochemical detection (HPLC-EC). mev-1 mutants also have a five to ten-fold higher mutation frequency than WT worms, based on a fem-3 mutation assay that detects loss-of-function mutations by measuring reversal of temperature sensitive sterility [19]. These studies are consistent with the idea that mitochondrial ROS contributes to nuclear genomic instability and mutagenesis, and promotes ageing. However, longitudinal studies in worms measuring DNA damage accumulation over the organism’s lifespan have yet to be reported.
11.3 DNA Repair Capacity with Age
Radiation-sensitive mutant strains (rad) were first isolated in the 1980s, based on their sensitivity to ultraviolet light (UV). These mutants are also sensitive to other DNA damaging agents such as methyl methane sulphonate (MMS) and ionizing radiation (IR) [20]. In this study, no significant differences in lifespan were observed in the rad mutants compared to wild-type worms. Five of the seven rad mutants (except rad-4 and rad-7) have slightly shortened lifespans after exposure to IR, but this is dependent on the dose of IR used. An intermediate dose of radiation (10–30 krad) leads to increased mean lifespans in all strains (significant in WT (N2) and rad-4), while a higher dose (>100 krad) shortens lifespan (significant in WT (N2) and rad-3). This can be attributed to hormesis , where low levels of damage induce stress responses that are beneficial [21]. However, higher levels of stress lead to irreversible, persistent damage that has detrimental effects.
Hartman et. al. reported that there is no correlation between lifespan and sensitivity to genotoxic stress in several inbred strains of worms that have lifespans ranging from 13 to 30.9 days [22]. In addition, there is no correlation between excision of UV induced lesions (a direct measure of NER) and lifespan, based on measurement of (6–4) photoproducts and CPDs by radioimmunoassay in UV irradiated worms. It is important to note that these experiments were performed 24–48 h after egg lay (small differences in NER between strains were observed at 24 h but not at 48 h), which may not accurately portray DNA repair capacity of adult worms.
In contrast, several studies show that most long-lived mutant strains are resistant to multiple stressors including UV irradiation. In mammals, there is evidence, although not overwhelmingly convincing, that long-lived animals have higher DNA repair capacity compared to short-lived species [23, 24]. To perform similar studies in C. elegans, Hyun et al. examined DNA repair capacity of WT and long-lived strains by measuring the number of pyrimidine dimers in a target gene, vps-45 of UV irradiated worms, using T4 endonuclease V (T4 endo V). T4 endo V incises DNA specifically at sites of pyrimidine dimers, which can be quantified by Southern blot. Using this technique in WT worms, the majority of repair is completed within 8 h and repair plateaus by 12 h (~67 % of lesions are repaired). However, in long-lived mutants, 85 % pyrimidine dimer repair is complete within 4 h and plateaus at 90 % by 12 h. This suggests that DNA repair capacity (at least NER) is faster in several long-lived mutants including age-1 fer-15, age-1 and daf-2 . Similarly, UV irradiation of adults (post-reproduction stage) significantly decreases lifespan in both WT and daf-16 worms (37.2 %) and in long-lived (~16.4 %) mutants, suggesting that failure to repair UV-induced adducts, does shorten lifespan [25]. Although these experiments were done with an exogenous source of DNA damage that may cause damage to other macromolecules , they do suggest that DNA repair capacity impacts lifespan.
11.4 DNA Repair Pathways and Ageing
One line of evidence that supports the theory that decreased repair capacity drives ageing comes from humans with genetically inherited defects in DNA repair pathways (reviewed in [26, 27]). These defects lead to hypersensitivity to DNA damaging agents, accumulation of DNA damage and accelerated ageing of one or more tissues. The identification of such progeroid syndromes in humans led to the design of mutant C. elegans strains that have defects in DNA repair mechanisms. Below, we examine each DNA repair pathway and evidence that links it to ageing.
11.4.1 Base Excision Repair (BER)
C. elegans possess two AP endonucleases, EXO-3 (exo III family) and APN-1 (endo IV family) [28]. EXO-3 (R09B3.1a) is an endonuclease required for BER that nicks DNA 5′ to an AP site. exo-3 mRNA levels decline 45 % by day 5 of adulthood and are maintained at low levels as worms age beyond that point [29]. RNAi depletion of exo-3 increases ROS and mitochondrial genome deletions, which are characteristics of aged worms. Knockdown of exo-3 also leads to other common ageing features such as neuronal damage and reduced motility.
Pharmacological suppression of ROS in exo-3 deficient worms inhibits neuronal damage and increases motility, suggesting that ROS is a key cause of morbidity in the mutant worms [29]. In accordance, exo-3 RNAi leads to a reduction in both mean (20 %) and maximum (10 %) lifespan of C. elegans. Interestingly, suppression of cep-1, ortholog of the tumour suppressor p53, rescues ageing phenotypes of exo-3 RNAi mutants. One possible explanation is that cep-1 is known to increase oxidative stress by inducing expression of pro-oxidant genes and repressing antioxidant genes, in response to cellular stress including genotoxic stress [29]. Thus deleting cep-1 should reduce ROS and oxidative DNA damage in the exo-3 mutant worms . Interestingly, in WT nematodes, suppression of cep-1 leads to upregulation of exo-3 and preserves healthspan (neuronal integrity and motility). This suggests that cep-1 and exo-3 coordinately respond to oxidative or genotoxic stress and this influences age-related decline .
Kato et al. further characterized the exo-3 deletion mutant (tm4374) and confirmed a reduced lifespan [30]. The short lifespan of exo-3 mutant is also suppressed by deletion of ung-1, a monofunctional uracil DNA glycosylase. UNG-1 acts upstream of EXO-3 in BER to remove uracil from DNA (caused by spontaneous hydrolysis of cytosine) creating an AP site. This reveals that AP sites are more deliterious than uracil lesions. Additionally, the authors reported a surprising difference between somatic versus germline cells (post-mitotic vs proliferating). In the germline, exo-3 is highly expressed and loss of EXO-3 leads to a reduced brood size (reflecting the proliferative capacity of germ cells). Interestingly, the impact of exo-3 on brood size requires the presence of nth-1, a second DNA glycosylase that removes oxidized pyrimidines [31]. This suggests that oxidative DNA lesions are a major substrate of BER in germ cells, whereas deamination products are more important in somatic cells.
Although one might predict that increased levels of oxidative DNA lesions would promote ageing, nth-1 null mutants show a normal mean and maximum lifespan [32]. Surprisingly, QPCR studies reveal that the rate of removal of damage caused by oxidative and alkylating agents in the WT and nth-1 adult worms is similar [33]. This could imply that there are redundant mechanisms for removing oxidized purines in nth-1 deficient somatic cells and that it is unlikely that nth-1 depletion induces ROS as occurs in exo-3 mutants. It is also possible that oxidative DNA lesions may not be a major determinant of lifespan in somatic cells. Collectively, these genetic studies help reveal what endogenous DNA lesions are apt to contribute to ageing and lifespan [30].
Suppression of the other AP endonuclease, apn-1, causes classic phenotypes associated with a DNA repair defect, including increased mutation frequency and sensitivity to DNA damaging agents [34]. Additionally, knock down of apn-1 causes a delay in the division of the P1 blastomere, typical of worms with increased DNA damage. However, unlike exo-3 mutants that have a shortened lifespan, apn-1 RNAi does not reduce the lifespan of worms unless they are treated with tert-butyl hydroperoxide (tert-BH) or MMS [34]. This suggests that EXO-3 may play a redundant role for APN-1 and is the major AP endonuclease in somatic maintenance. Collectively, these studies show that BER is required for normal lifespan of worms, implicating endogenous DNA damage as a driver of ageing .
11.4.2 Nucleotide Excision Repair (NER)
Inherited mutations affecting NER are responsible for several progeroid syndromes in humans including Xeroderma pigmentosum (XP), Cockayne syndrome (CS), trichothiodystrophy (TTD) and XFE progeroid syndrome [26]. These syndromes are all characterized by accelerated age-related decline of several tissues and the premature onset of diseases associated with old age. Many of these progeroid syndromes have been recapitulated in mice, often by single DNA repair gene mutations. These human syndromes fuelled several studies to interrogate whether NER promotes healths and longevity in worms.
In C. elegans , the mechanism of repair of UV-induced DNA lesions (i.e., NER) is very similar to humans [35]. There are several lines of evidence suggesting that DNA lesions that are substrates for NER promote ageing in worms. Expression of NER proteins is significantly lower in non-gravid adults (older adults) compared to gravid adults [35], indicating that NER is important for replicative longevity. glp-1 mutants have an arrested germline and therefore enable measurement of DNA repair exclusively in post-mitotic animals. Repair of UV lesions is slower in ageing glp-1adults compared to young worms. This diminished repair in somatic cells however, is not because of decreased expression of DNA repair proteins (at least at the mRNA level). This could mean that protein translation, subcellular localization or post-translational modification of DNA repair proteins is affected with age [35]. These studies contribute evidence that DNA repair capacity decreases with age.
XP Complementation Group A (XPA) is required for GG-NER and TC-NER and plays a key role before damage excision. As in humans and mice [36], rad-3/xpa-1 worms are hypersensitive to UV irradiation and have an increased mutation frequency in response to UV. Steady state levels of the oxidative lesions formamidopyrimidines (FapyGua and FapyAde) and 8-hydroxyadenine are significantly increased in xpa-1 (ok698) mutants [37]. Human XP-A lymphoblasts also show an accumulation of these oxidative lesions, suggesting the importance of NER in repairing these endogenous lesions [38]. These adducts block both replication and transcription, and are increased in several age-related diseases such as Alzheimer’s and cancer [39].
Reports on lifespan of xpa-1 (ok698) mutants vary. Hyun et al. report a ~20 % reduction in mean lifespan [25]. Lans et al. find no lifespan shortening when looking only at a population of healthy adults (but observe a shortened lifespan in an unbiased population consisting of developmentally delayed mutants) [40, 41]. Fensgård et al. see a reduction in mean lifespan but not in maximum lifespan, when strains are grown on standard E. coli OP50 bacteria [32]. Interestingly, nth-1 deletion (BER-see above) restores lifespan of xpa-1 mutants. Furthermore, transcription of several DNA damage response genes is attenuated in the double mutant (nth-1;xpa-1 compared to xpa-1 alone) [32]. Taken together, these data suggest that upon loss of xpa-1, nth-1 tries to process the lesions usually repaired by NER. However, NTH-1 and BER is apparently unable to resolve this damage through BER and instead causes an increased genome stress signal that culminates in a shortened lifespan. One possible interpretation of these data is that it is not the accumulation of DNA lesions itself that affects healthspan and lifespan but the damage-associated stress signal that is detrimental.
Many long-lived mutants , such as daf-2 and age-1, require the FOXO transcription factor , daf-16 for their extended lifespan [42, 43] (see Chap. 4). In the absence of cellular stress (or presence of insulin and IGF-1 ) DAF-16 is hyperphosphorylated by AKT and maintained in the cytoplasm under basal conditions. Upon stress, such as starvation, DAF-16 phosphorylation is attenuated and this allows for nuclear translocation and induction of several downstream target genes, including ROS scavengers and detoxifying enzymes. DAF-16 is predominantly in the nucleus in response to DNA damage (UV and in the xpa-1 mutant), and is required for growth and development in the presence of genotoxic stress. As worms age, DAF-16 nuclear translocation in response to UV radiation diminishes [44]. This would suggest that with age the responsiveness of DNA damage-associated stress-protective genes is attenuated. The ability to respond to stress and longevity has long been proposed to go hand-in-hand. Thus the loss of stress responses upon genomic instability may explain the shortened lifespan in some DNA repair mutants.
To determine if DNA repair and genomic stability is necessary for the increased lifespan of the longevity mutants, xpa-1 was knocked-down in age-1 mutants. Although age-1 mutants live ~1.6-fold times longer than N2, the lifespan of xpa-1 (RNAi);age-1 is similar to that of WT worms [25], suggesting that NER is critical for longevity. However, in stark contrast, knock-down of ERCC-1/XPF-1 expression further extends the life span of daf-2 mutants. This is puzzling since ERCC-1/XPF-1 functions downstream of XPA-1. However, interpretation of these results is complicated by the fact that ERCC-1/XPF-1 plays a role in several DNA repair pathways including DSB repair and ICL. Further studies, with suppression of different NER proteins in long-lived mutants, is required to resolve this conundrum.
11.4.3 Homologous Recombination (HR)
In humans, Bloom syndrome, caused by mutations in BLM, is characterized by genomic instability , chromosomal breaks and gross chromosomal rearrangements, growth retardation, facial erythema, impaired fertility and an elevated risk of cancer. HIM-6 is the C. elegans ortholog of human BLM RecQ helicase, a class of enzymes that play an integral role in HR. him-6 mutants exhibit several phenotypes that are characteristic of genomic instability [45]. For instance, him-6 worms have increased apoptosis and heightened sensitivity to ionizing radiation, a known inducer of double-strand breaks, as well as an increased frequency of small insertions and deletions . This is in accordance with human cells that lack BLM and have short deletions and duplications and an elevated number of sister-chromatid exchanges [46]. Although, Bloom patients do not display signs of classical premature ageing they do succumb to early onset of cancer. him-6 (ok412) mutant worms display a slight but significant decrease in lifespan [47]. Other healthspan measurements, such as motility or neuron maintenance have not been carefully examined in him-6 mutants.
DNA-2 helicase/endonuclease is involved in DNA replication and repair. It is recruited by BLM to cleave 5′ ssDNA during double-strand break repair. The worm ortholog CeDNA-2, protein is highly expressed in proliferative germ cells and during the early stages of embryo development, consistent with a role in a DNA replication [48]. dna-2 mutants display embryonic lethality (consistent with a role in replication and repair) and shortening of lifespan, which is more pronounced with each successive generation [49]. This result is similar to other DNA repair mutants such as mre-11, required for both homologous and non-homologous repair of double-strand breaks, RecQ5 in humans promotes DNA double-strand break repair by strand annealing [50, 51] and its loss is mainly associated with promoting cancer. In C. elegans , RCQ-5 is highly expressed in gonads, embryos and the intestine of adult worms. rcq-5 RNAi leads to increased sensitivity to ionizing radiation, consistent with a role in HR. Inhibition of rcq-5 leads to a 13 % decrease in lifespan at 20 °C and 37 % decrease at 25 °C [52]. This warrants further studies to investigate how temperature affects genomic instability in worms. An interesting analogy in humans is the exacerbation of the disease phenotypes in trichothiodystrophy patients with a defect in TC-NER when they have a fever [53].
11.4.4 Non-homologous End Joining (NHEJ)
Careful studies using a transgenic knock-in GFP-based NHEJ reporter mouse observed a significant decline in repair capacity in several tissues with age [54]. NHEJ deficiency leads to gross chromosomal rearrangements. Also, mice lacking Ku70, Ku80 or both, display signs of premature ageing including kyphosis, alopecia, osteoporosis, skin atrophy, and early onset of cancer [55]. Consistent with a role in NHEJ, RNAi of cku-70 or cku-80 (C. elegans orthologs of Ku70 and Ku80 respectively) show sensitivity to radiomimetics and MMS, but not to UV [56]. Interestingly, suppression of cku-70 both in WT and in long-lived daf-2 worms significantly increases thermotolerance (resistance to heat stress), in a daf-16 dependent manner. In WT worms, there is no concomitant increase in lifespan. In contrast, knockdown of cku-70 in an RNAi sensitive strain rrf-3 (pkl426) and in daf-2 mutants increases mean lifespan by 14 % and 35 %, respectively. One explanation is that the lifespan extension observed in daf-2 mutants maybe due to an HSF-1 dependent increase in multiple stress resistance. Interestingly, these longevity phenotypes are independent of germline signals, since glp-4;daf-2 mutants that lack germ cells also show an increase in lifespan (~9 %) when cku-70 expression is knocked-down.
In contrast to cku-70, knockdown of cku-80 does not confer thermotolerance and lifespan extension, suggesting possible divergent functions of these proteins that are thought to exist primarily as a complex [56]. Ku86 knock-out mice (cku-80 ortholog in mammals) display progeroid symptoms, premature senescence and a shortened lifespan [57]. Future studies to resolve this discrepancy between mice and worms are important.
Werner syndrome is a rare autosomal recessive disorder in humans, characterized by accelerated ageing. WRN belongs to the RecQ family of proteins and possesses an unusual exonuclease domain with 3–5′ activity. WRN plays a role in telomere maintenance, replication and interacts with Ku80/70 to facilitate NHEJ [58–60]. In the nematode, WRN-1 displays 43 % identity in protein sequence with human WRN. However, it lacks the exonuclease domain [61]. WRN-1 is expressed in larval stages, as well as in the hypodermis , intestine and germ cells of the adult, and protein expression decreases with age [61]. Loss of wrn-1 significantly reduces brood size and leads to increased growth arrest at larval stages. Additionally, wrn-1 mutants display increased lipofuscin accumulation, tissue deterioration in the head, and have shortened lifespans [61, 62]. Whether the role of WRN-1 in NHEJ is required for its role in lifespan maintenance needs to be further examined. Genetic studies to place the NHEJ proteins in pathways associated with ageing are key to further understanding its role in healthspan and longevity .
11.5 DNA Damage Response (DDR) and Ageing
The DNA damage response (DDR) is an integral part of damage recognition, recruitment of DNA repair proteins and maintenance of genomic stability (reviewed in [9]). The most upstream DDR kinases are: (1) ataxia telangiectasia mutated (ATM/ATM-1) and (2) ataxia telangiectasia and RAD3-related protein (ATR/ATL-1). ATM-1 usually recognizes double-strand breaks, such as those caused by IR and crosslinking drugs like mitomycin C (MMC), whereas ATL-1 responds to single-strand breaks and bulky adducts caused by agents such as UV [63]. These serine-threonine protein kinases, ATM-1 and ATL-1, phosphorylate a number of targets including the transcription factor p53/CEP-1, either directly or by first phosphorylating and activating checkpoint kinase 2 (CHK2). Additionally, ATR phosphorylates checkpoint kinase 1 (CHK1) effector protein, which in turn phosphorylates the dual-specificity phosphatase CDC25C, thus arresting cells prior to mitosis and affording time for repair. These signalling pathways are well conserved in the nematode. Loss of these checkpoint proteins causes severe disease in humans including ataxia-telangiectasia and related syndromes and predisposition to cancer [64]. Since C. elegans is not prone to cancer, it is easier to examine the role of DNA damage response proteins in maintaining healthspan and lifespan.
Poly ADP-ribose polymerase (PARP), is a family of proteins that detects single-strand breaks, binds to DNA, and begins the synthesis of a poly-ADP-ribose chain (PAR) that acts as a recruitment signal for other repair proteins [65]. PARP also directly binds and activates ATL-1, to recruit other repair proteins [66]. PARylation increases markedly in mice and nematodes with ageing, suggesting that DNA damage increases with age. Accordingly, xpa-1 (ok698) mutants have significantly higher levels of PAR than WT worms [67]. PARP activity requires NAD+. In a C. elegans pme-1 mutant, the worm PARP-1 homologue, both PARylation and NAD+ consumption is attenuated, implying that pme-1 is a major consumer of NAD+. Notably, NAD+ levels decline with age across species [68], indirectly supporting a rise in DNA damage with age.
Ageing-associated lipid peroxidation and lipofuscin accumulation is substantially reduced in pme-1 mutant worms, and suppression of pme-1 increases levels of NAD+ and leads to an extension of mean lifespan. NAD+ consumption by pme-1 leads to suppression of sir2.1 activity, which also requires NAD+ as a coactivator. sir2.1 in turn regulates mitochondrial unfolded protein response (UPRmt) and the daf-16-dependent antioxidant response needed to maintain mitochondrial homeostasis. This ties together nuclear DNA damage and repair mechanisms with mitochondrial function. Importantly, these signalling pathways are conserved in mammals [68].
Sirtuins also play a role in mitochondrial biogenesis by regulating PGC1α [69, 70] in mammals. Increased DNA damage (e.g., in xpa-1 mutants), leads to hyperactivation of PARP-1 and attenuation of the NAD+-SIRT1-PGC1α axis. In turn, mitophagy is compromised and defective mitochondria accumulate. Thus treatment of short-lived xpa-1 (ok698) mutants with a PARP inhibitor AZD2281 or NAD+ precursors (nicotinamide riboside (NR) and nicotinamide mononucleotide (NMN)) rescues lifespan of these DNA repair deficient worms [67]. This reveals a complex relationship between DNA damage response pathways and metabolic homeostasis, necessary for maintaining healthspan and lifespan.
CID-1 (caffeine induced death-1) shares homology with poly(A)+ polymerase domain proteins, which play a role in the S phase to mitosis checkpoint [71, 72]. Hydroxyurea (HU) works as a ribonucleotide reductase (RNR) inhibitor and depletes dNTP causing developmental arrest [73]. In contrast, cid-1 suppression (RNAi and mutant) permits normal development upon exposure to HU, suggesting a failure to induce checkpoint signalling that leads to cell cycle arrest or apoptosis. Loss of cid-1 leads to hsp-4 induction, increases thermotolerance and significantly increases lifespan. Although, hsp-4 is induced in cid-1 mutant worms, this is not accompanied by hsp-16 activation. This suggests a role for hsp-4 in stress resistance that is independent from the hsp-16-dependent unfolded protein response in the endoplasmic reticulum (UPRER) [73].
Other checkpoint proteins, such as cdc-25.1 (orthologue of CDC25C) and chk-1 (orthologue of CHK-1), also influence longevity [73]. Inactivation of cdc-25.1, cdc25.2, cdc-25.3 (but not cdc-25.4), results in stress resistance and an increase in lifespan. cdc-25.1 gain-of function mutants are thermosensitive and short-lived. Suppression of chk-1 leads to resistance to thermal stress (confirmed using the chk-1 inhibitor UCN-01), extends lifespan (~15 to 25 %) and induces hsp-4. Surprisingly, stress resistance upon chk-1 inhibition is not dependent on DAF-16 and DAF-12, as no change in nuclear localization of these proteins is observed. Consistently, loss of chk-1 increases lifespan in daf-16, daf-12 and in the long-lived daf-2 mutants [73]. This data suggests that these checkpoint proteins promote somatic maintenance in post-mitotic cells, clearly through processes that are different from their well-established role germline and IIS pathways.
C. elegans p53 ortholog, cep-1, shares homology with mammalian p53, both in form and in function. Upon DNA damage, CEP-1, a transcription factor translocates to the nucleus and regulates the DNA damage response [74]. Suppression of CEP-1 leads to an increase in chromosomal nondisjunction events [75]. In humans, p53 is a well-established tumour suppressor, mutation of which causes the cancer predisposition syndrome Li-Fraumeni. The importance of tight regulation of p53 is documented by the fact that mutations causing chronic activation of p53 leads to segmental progeria in mice, and decreases both median (23 %) and maximum (21 %) lifespan [76]. Deletion or loss-of-function mutations of p53 leads to increased cancer incidence. Although, C. elegans is a largely post-mitotic organism, cep-1 still seems to play a role in the DNA damage response and repair in the worm.
Suppression of cep-1, using RNAi or the deletion mutant (gk138), does not confer resistance to heat, high oxygen levels or UV, but does lead to increased mean lifespan. cep-1 requires daf-16 to exert its effects on lifespan, but this is not through differential nuclear localization of daf-16, suggesting that CEP-1 is not biochemically upstream of DAF-16. Of other proteins known to be involved in DNA repair clk-2 (qm37), rad-5 (mn159), him-7 (e1480), ced-4 (n1162), egl-1 (n487), msh-2 (ev679::Tc1) and hus-1), only suppression of hus-1, a checkpoint protein required for genomic stability, exhibits an increase in lifespan (~11 %). cep-1 RNAi in the hus-1 mutant does not further alter longevity, suggesting they both influence lifespan by the same pathway(s) [77].
11.6 Conclusions and Future Perspectives
Despite strong evidence in humans that DNA damage increases with age and is associated with several age-related diseases, whether it plays a causal role in driving ageing remains contentious. C. elegans as a model system has provided critical insights on the DNA damage theory of ageing. It is undeniable that several DNA repair pathways examined in the worm have an effect on lifespan. However, the mechanism remains unclear. Does chronic activation of the DNA damage response or multi-stress response mechanisms play a role? Or is it mutagenesis caused largely by transcription drive ageing?
Mutation accumulation has been measured in 24 different regions of the genome to compare the relative importance of BER vs. NER vs. MMR in protecting genomic stability [78]. This revealed that loss of MMR led to 48-fold increase in mutations, while NER mutants caused a 28-fold increase and BER deficient worms had a 17-fold increase compared to WT worms. In contrast, whole genome next generation sequencing (NGS) in WT and 17 different DNA repair -deficient mutants revealed no significant increase in mutation rate in the absence of various DNA repair mechanisms [1]. These differences could stem from the number of generations examined. However, both of these studies do not reveal any significant accumulation of mutations during one lifespan, suggesting minimal role of mutation accumulation on lifespan.
Likewise, another question that can be answered in the nematode, is the relative importance of DNA repair in proliferating versus post-mitotic cells and its effect on lifespan. Additionally, C. elegans does not seem to have the traditional cellular senescence and senescence associated secretory phenotype (SASP) [13]. This could be an advantage in understanding primary mechanisms that respond to DNA damage and impact cellular programming finally impacting ageing. Last, but not the least, the one question that plagues the field is whether improving DNA repair efficiency leads to longevity? This poses a challenging problem, since DNA damage recognition and repair processes are extremely complex. However, with the advent of newer techniques such CRISPR, generating transgenic overexpression lines in the worm is feasible, timely and cost-effective.
In conclusion, using the strengths of C. elegans to elucidate the role of DNA damage in maintaining healthspan and lifespan is key to understanding how evolutionary adaptations has led to lifespan differences in species .
References
Meier B et al (2014) C. elegans whole-genome sequencing reveals mutational signatures related to carcinogens and DNA repair deficiency. Genome Res 24(10):1624–1636
Drake JW et al (1998) Rates of spontaneous mutation. Genetics 148(4):1667–1686
Klass M, Nguyen PN, Dechavigny A (1983) Age-correlated changes in the DNA template in the nematode C. elegans. Mech Ageing Dev 22(3–4):253–263
Melov S et al (1995) Increased frequency of deletions in the mitochondrial genome with age of C. elegans. Nucleic Acids Res 23(8):1419–1425
Jacob KD et al (2013) Markers of oxidant stress that are clinically relevant in aging and age-related disease. Mech Ageing Dev 134(3–4):139–157
Yen TC et al (1992) Age-dependent 6 kb deletion in human liver mitochondrial DNA. Biochem Int 26(3):457–468
Bratic A, Larsson NG (2013) The role of mitochondria in aging. J Clin Invest 123(3):951–957
Friedberg EG, Wood RD (1996) DNA excision repair path ways. In: Pamphilis MLD (ed) DNA replication in eukaiyotic cells. Cold Spring Harbor Laboratory Press: Gold Spring Harbor, NY, pp 249–269
Stergiou L, Hengartner MO (2004) Death and more: DNA damage response pathways in the nematode C. elegans. Cell Death Differ 11(1):21–28
Ho TV, Scharer OD (2010) Translesion DNA synthesis polymerases in DNA interstrand crosslink repair. Environ Mol Mutagen 51(6):552–566
Roerink SF et al (2012) A broad requirement for TLS polymerases eta and kappa, and interacting sumoylation and nuclear pore proteins, in lesion bypass during C. elegans embryogenesis. PLoS Genet 8(6):e1002800
Lans H, Vermeulen W (2015) Tissue specific response to DNA damage: C. elegans as role model. DNA Repair (Amst) 32:141–148
d’Adda di Fagagna F (2008) Living on a break: cellular senescence as a DNA-damage response. Nat Rev Cancer 8(7):512–522
Baker DJ et al (2016) Naturally occurring p16(Ink4a)-positive cells shorten healthy lifespan. Nature 530(7589):184–189
Dmitrieva NI, Burg MB (2007) High NaCl promotes cellular senescence. Cell Cycle 6(24):3108–3113
Ishii N et al (1998) A mutation in succinate dehydrogenase cytochrome b causes oxidative stress and ageing in nematodes. Nature 394(6694):694–697
Senoo-Matsuda N et al (2003) A complex II defect affects mitochondrial structure, leading to ced-3- and ced-4-dependent apoptosis and aging. J Biol Chem 278(24):22031–22036
Senoo-Matsuda N et al (2001) A defect in the cytochrome b large subunit in complex II causes both superoxide anion overproduction and abnormal energy metabolism in C. elegans. J Biol Chem 276(45):41553–41558
Hartman P et al (2004) Mitochondrial oxidative stress can lead to nuclear hypermutability. Mech Ageing Dev 125(6):417–420
Hartman PS, Herman RK (1982) Radiation-sensitive mutants of C. elegans. Genetics 102(2):159–178
Johnson TE, Hartman PS (1988) Radiation effects on life span in C. elegans. J Gerontol 43(5):B137–B141
Hartman PS et al (1988) Radiation sensitivity and DNA repair in C. elegans strains with different mean life spans. Mutat Res 208(2):77–82
MacRae SL et al (2015) DNA repair in species with extreme lifespan differences. Aging (Albany NY) 7(12):1171–1184
Salmon AB, Ljungman M, Miller RA (2008) Cells from long-lived mutant mice exhibit enhanced repair of ultraviolet lesions. J Gerontol A Biol Sci Med Sci 63(3):219–231
Hyun M et al (2008) Longevity and resistance to stress correlate with DNA repair capacity in C. elegans. Nucleic Acids Res 36(4):1380–1389
Gurkar AU, Niedernhofer LJ (2015) Comparison of mice with accelerated aging caused by distinct mechanisms. Exp Gerontol 68:43–50
Navarro CL, Cau P, Levy N (2006) Molecular bases of progeroid syndromes. Hum Mol Genet 15(Spec No 2):R151–R161
Shatilla A et al (2005) Identification of two apurinic/apyrimidinic endonucleases from C. elegans by cross-species complementation. DNA Repair (Amst) 4(6):655–670
Schlotterer A et al (2010) Apurinic/apyrimidinic endonuclease 1, p53, and thioredoxin are linked in control of aging in C. elegans. Aging Cell 9(3):420–432
Kato Y et al (2015) C. elegans EXO-3 contributes to longevity and reproduction: differential roles in somatic cells and germ cells. Mutat Res 772:46–54
Morinaga H et al (2009) Purification and characterization of C. elegans NTH, a homolog of human endonuclease III: essential role of N-terminal region. DNA Repair (Amst) 8(7):844–851
Fensgard O et al (2010) A two-tiered compensatory response to loss of DNA repair modulates aging and stress response pathways. Aging (Albany NY) 2(3):133–159
Hunter SE et al (2012) In vivo repair of alkylating and oxidative DNA damage in the mitochondrial and nuclear genomes of wild-type and glycosylase-deficient C. elegans. DNA Repair (Amst) 11(11):857–863
Zakaria C et al (2010) C. elegans APN-1 plays a vital role in maintaining genome stability. DNA Repair (Amst) 9(2):169–176
Meyer JN et al (2007) Decline of nucleotide excision repair capacity in aging C. elegans. Genome Biol 8(5):R70
de Vries A et al (1995) Increased susceptibility to ultraviolet-B and carcinogens of mice lacking the DNA excision repair gene XPA. Nature 377(6545):169–173
Arczewska KD et al (2013) Active transcriptomic and proteomic reprogramming in the C. elegans nucleotide excision repair mutant xpa-1. Nucleic Acids Res 41(10):5368–5381
Lipinski LJ et al (1999) Repair of oxidative DNA base lesions induced by fluorescent light is defective in xeroderma pigmentosum group A cells. Nucleic Acids Res 27(15):3153–3158
Cooke MS et al (2003) Oxidative DNA damage: mechanisms, mutation, and disease. FASEB J 17(10):1195–1214
Lans H et al (2013) DNA damage leads to progressive replicative decline but extends the life span of long-lived mutant animals. Cell Death Differ 20(12):1709–1718
Lans H et al (2010) Involvement of global genome repair, transcription coupled repair, and chromatin remodeling in UV DNA damage response changes during development. PLoS Genet 6(5):e1000941
Kenyon C et al (1993) A C. elegans mutant that lives twice as long as wild type. Nature 366(6454):461–464
Johnson TE (1990) Increased life-span of age-1 mutants in C. elegans and lower Gompertz rate of aging. Science 249(4971):908–912
Mueller MM et al (2014) DAF-16/FOXO and EGL-27/GATA promote developmental growth in response to persistent somatic DNA damage. Nat Cell Biol 16(12):1168–1179
Wicky C et al (2004) Multiple genetic pathways involving the C. elegans Bloom’s syndrome genes him-6, rad-51, and top-3 are needed to maintain genome stability in the germ line. Mol Cell Biol 24(11):5016–5027
German J (1993) Bloom syndrome: a mendelian prototype of somatic mutational disease. Medicine (Baltimore) 72(6):393–406
Grabowski MM, Svrzikapa N, Tissenbaum HA (2005) Bloom syndrome ortholog HIM-6 maintains genomic stability in C. elegans. Mech Ageing Dev 126(12):1314–1321
Lee KH et al (2003) Dna2 requirement for normal reproduction of C. elegans is temperature-dependent. Mol Cells 15(1):81–86
Lee MH et al (2003) C. elegans dna-2 is involved in DNA repair and is essential for germ-line development. FEBS Lett 555(2):250–256
Hu Y et al (2007) RECQL5/Recql5 helicase regulates homologous recombination and suppresses tumor formation via disruption of Rad51 presynaptic filaments. Genes Dev 21(23):3073–3084
Paliwal S et al (2014) Human RECQ5 helicase promotes repair of DNA double-strand breaks by synthesis-dependent strand annealing. Nucleic Acids Res 42(4):2380–2390
Jeong YS et al (2003) Deficiency of C. elegans RecQ5 homologue reduces life span and increases sensitivity to ionizing radiation. DNA Repair (Amst) 2(12):1309–1319
Vermeulen W et al (2001) A temperature-sensitive disorder in basal transcription and DNA repair in humans. Nat Genet 27(3):299–303
Vaidya A et al (2014) Knock-in reporter mice demonstrate that DNA repair by non-homologous end joining declines with age. PLoS Genet 10(7):e1004511
Li H et al (2007) Deletion of Ku70, Ku80, or both causes early aging without substantially increased cancer. Mol Cell Biol 27(23):8205–8214
McColl G, Vantipalli MC, Lithgow GJ (2005) The C. elegans ortholog of mammalian Ku70, interacts with insulin-like signaling to modulate stress resistance and life span. FASEB J 19(12):1716–1718
Vogel H et al (1999) Deletion of Ku86 causes early onset of senescence in mice. Proc Natl Acad Sci U S A 96(19):10770–10775
Cooper MP et al (2000) Ku complex interacts with and stimulates the Werner protein. Genes Dev 14(8):907–912
Shen JC, Loeb LA (2000) The Werner syndrome gene: the molecular basis of RecQ helicase-deficiency diseases. Trends Genet 16(5):213–220
Crabbe L et al (2004) Defective telomere lagging strand synthesis in cells lacking WRN helicase activity. Science 306(5703):1951–1953
Lee SJ et al (2004) A Werner syndrome protein homolog affects C. elegans development, growth rate, life span and sensitivity to DNA damage by acting at a DNA damage checkpoint. Development 131(11):2565–2575
Dallaire A et al (2012) Down regulation of miR-124 in both Werner syndrome DNA helicase mutant mice and mutant C. elegans wrn-1 reveals the importance of this microRNA in accelerated aging. Aging (Albany NY) 4(9):636–647
Vermezovic J et al (2012) Differential regulation of DNA damage response activation between somatic and germline cells in C. elegans. Cell Death Differ 19(11):1847–1855
Sperka T, Wang J, Rudolph KL (2012) DNA damage checkpoints in stem cells, ageing and cancer. Nat Rev Mol Cell Biol 13(9):579–590
Lindahl T et al (1995) Post-translational modification of poly(ADP-ribose) polymerase induced by DNA strand breaks. Trends Biochem Sci 20(10):405–411
Kedar PS et al (2008) Interaction between PARP-1 and ATR in mouse fibroblasts is blocked by PARP inhibition. DNA Repair (Amst) 7(11):1787–1798
Fang EF et al (2014) Defective mitophagy in XPA via PARP-1 hyperactivation and NAD(+)/SIRT1 reduction. Cell 157(4):882–896
Mouchiroud L et al (2013) The NAD(+)/Sirtuin pathway modulates longevity through activation of mitochondrial UPR and FOXO signaling. Cell 154(2):430–441
Canto C et al (2009) AMPK regulates energy expenditure by modulating NAD+ metabolism and SIRT1 activity. Nature 458(7241):1056–1060
Rodgers JT et al (2005) Nutrient control of glucose homeostasis through a complex of PGC-1alpha and SIRT1. Nature 434(7029):113–118
Wang SW et al (2000) Cid1, a fission yeast protein required for S-M checkpoint control when DNA polymerase delta or epsilon is inactivated. Mol Cell Biol 20(9):3234–3244
Saitoh S et al (2002) Cid13 is a cytoplasmic poly(A) polymerase that regulates ribonucleotide reductase mRNA. Cell 109(5):563–573
Olsen A, Vantipalli MC, Lithgow GJ (2006) Checkpoint proteins control survival of the postmitotic cells in C. elegans. Science 312(5778):1381–1385
Bensaad K, Vousden KH (2007) p53: new roles in metabolism. Trends Cell Biol 17(6):286–291
Derry WB, Putzke AP, Rothman JH (2001) C. elegans p53: role in apoptosis, meiosis, and stress resistance. Science 294(5542):591–595
Tyner SD et al (2002) p53 mutant mice that display early ageing-associated phenotypes. Nature 415(6867):45–53
Arum O, Johnson TE (2007) Reduced expression of the C. elegans p53 ortholog cep-1 results in increased longevity. J Gerontol A Biol Sci Med Sci 62(9):951–959
Denver DR et al (2006) The relative roles of three DNA repair pathways in preventing C. elegans mutation accumulation. Genetics 174(1):57–65
DNA repair database. Available from: https://dnapittcrew.upmc.com/db/index.php
Acknowledgments
A.U.G. is supported by NIH/NIAK99 AG049126. M.S.G. is supported by NIH/NIA R01 AG036992. L.J.N. is supported by NIH/NIAP01 AG043376. In addition, the Niedernhofer lab is supported in part by a sponsored research agreement between the Scripps Research Institute and Aldabra Biosciences LLC, of which she is a co-founder. The authors apologize to those whose work could not be cited due to lack of space.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Gurkar, A.U., Gill, M.S., Niedernhofer, L.J. (2017). Genome Stability and Ageing. In: Olsen, A., Gill, M. (eds) Ageing: Lessons from C. elegans. Healthy Ageing and Longevity. Springer, Cham. https://doi.org/10.1007/978-3-319-44703-2_11
Download citation
DOI: https://doi.org/10.1007/978-3-319-44703-2_11
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-44701-8
Online ISBN: 978-3-319-44703-2
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)