Abstract
Animal DNA barcoding has been advanced as a promising tool to aid species identification and discovery through the use of short, standardized mitochondrial gene COI targets in spite of extensive morphological taxonomic studies, for a variety of reasons the identification of fishes can be problematic, time consuming, even for experts. To overcome in this content though DNA barcoding is proving to be a useful tool. The present study use DNA barcoding to genetically identify specimen were collected from Nallavadu Lagoon ecosystem in Puducherry and morphologically identified. The COI was amplified and bi-directionally sequenced from 45 specimens (5 specimens/species) belonging to 9 species, five families. All studied specimens sequence were interrogated using analytical tools developed by Barcode of Life Data System (BOLD). All specimen exhibited discrete clusters of closely related haplogroups in NJ tree, which permitted the discrimination of 96–100 % genetic similarity of all the studied species. These studies represent a significant contribution to the worldwide barcode database from Indian continental regional and demonstrated the utility of COI gene sequences for specific regional species identification.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
Coastal lagoon is a shallow coastal water body separated from the sea by a barrier, connected at least sporadically to the ocean by one or more inlets and usually oriented parallel to shore. Coastal lagoons constitute about 13 % of global coastline (Farmworth and Golley 1974; Poyyamoli et al. 2011) and considered as a different coastal environment from estuaries (Kjerfve 1994). In India, there are 17 noteworthy lagoons are present on the east and west coast of India. Coastal lagoons notably act on the control of freshwater inflow and connections with the ocean (Crivelli and Ximenes 1992; Peja et al. 1996; Quinn et al. 1999), they are found to be highly dynamic, productive and extremely unpredictable systems (Barnes 1980). The fauna that occur in lagoons are of 3 groups, marine species, freshwater species and lagoon species. The presence of this specialist fauna make the conservation of lagoons important in maintaining the biodiversity. These ecosystem areas are important for the marine fisheries, serving as they do as nurseries for many species of fishes. Bouchet (2006) reported that 22, 9602 marine species had been described but the total species living on Earth may exceed more than 10 million as estimated by Grassle and Maciolek (1992). Traditionally taxonomy sciences are described based on the morphological characters, which develop the taxonomy keys. The conventional taxonomic sciences are very tedious to delineate species, in the last 250 years only 25 % of species were identified (Coyne and Orr 2004; Packer et al. 2009). The taxonomic scenario identification keys were initiated by Aristotle and later organized by Linnaeus. The fin fishes have been catalogued to about thirty-two thousand species from the aquatic environments, and it is divided into six classes, 62 orders, and 540 families (Eschmeyer 2010), which occupy >50 % of the all vertebrate species in world wide. Indian waters fishes were listed and identified by Day (1878) and Whitehead and Talwar (1976). Later, Talwar (1990) has reported 2546 fish species belongings to the 969 genera, 254 families and 40 order, among them 57 % of marine fish genera are common to the Indo-Pacific and Atlantic and Mediterranean regions. In Indian marine biodiversity studies are still untouched in minor phyla, non-commercially important fishes, and in remote parts of the islands and minor estuaries (Venkataraman and Wafar 2005). In current taxonomy scenario, the shortage of trained personnel limits development of taxonomic knowledge. The above factors have created a taxonomic impediment to biodiversity studies (Hebert et al. 2003). Despite ongoing scientific debate concerning the role of molecular methods in taxonomy DNA barcoding has emerged as a widely accepted tool for species identification because of its enhanced focus on standardization and data validation.
DNA barcoding is a method, based on short standardized gene sequences of DNA (Hebert et al. 2003), used for species identification. It has been used inter alia for identification of fish species (Hebert et al. 2004; Ward et al. 2005; Hubert et al. 2008; Persis et al. 2009; Steinke et al. 2009; Lakra et al. 2010; Sachithanandam et al. 2011, 2012). The effectiveness of barcoding has been demonstrated in diverse taxa, including butterflies (Hebert et al. 2003), birds (Hebert et al. 2004), fishes (Ward et al. 2005; Persis et al. 2009; Steinke et al. 2009; Lakra et al. 2010; Sachithanandam et al. 2011, 2012), sponges, bivalves (Jarnegren et al. 2007) and mammals (Clare et al. 2007). DNA barcoding is being applied in the field of fish conservation (Holmes et al. 2009), fisheries managements aspects (Rasmussen et al. 2009) and food safety analysis, with mislabeling being revealed through COI sequences (Wong and Hanner 2008).
In 2005 FISH-BOL (Fish Barcode of Life Initiative) was set up to establish a library of COI gene sequences of all fish species, to enable global taxonomic identification (Ward et al. 2009; Eschmeyer 2010). FISH-BOL’s primary work is led by a research team with responsibility for collection of samples, traditional identification using morphological keys, and COI gene sequencing of species in their geographic regions. Current progress of DNA barcoding in the Arctic and Antarctic regions is 74 and 50 %, respectively. Other regions from tropical to subtropical show good progress, with 20 % coverage from Australia, Mesoamerica continental and Oceania. However, species-rich regions of Asia, South America and Africa show low progress (Becker et al. 2011). In India, 11,023 fish species have been morphologically reported (Nelson 2006; Mecklenburg et al. 2010). Reports suggest that only 1918 of these (17.4 %) have been barcoded (Becker et al. 2011).
The aim of this study is to extend barcode coverage to Nallavadu brackish water fishes from Puducherry coastal area. We examined the patterns of barcode sequences genetic divergences among the 45 specimens identified as belonging to 9 species consists of five families representing highly exploitation for commercially values fishes from in this region.
2 Methods
2.1 Study Area
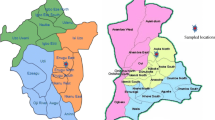
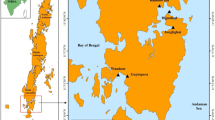
Puducherry located on the Coramandal coast 11° 52′ N, 79° 45′ E and 11° 59′ N and between 79° 52′ E covers an area of 480 km2. Nallavadu village located at a distance of about 14 km South from Puducherry main town. Nallavadu, Poornankupam, Andiarpalayam, Pillaiarthittu and Panithittu villages are the boundaries of Nallavadu lagoon (Fig. 1). The aerial length of the lagoon is 3.44 km and maximum depth is 10 m in its broadest part (Padmavathy et al. 2010; Poyyamoli et al. 2011). The climate is tropical dissymmetric with the bulk of the rainfall during northeast monsoon from October to December and some amount of rainfall during southwest monsoon from June–September. The lagoon is connected to the sea in its south part by an artificial channel and receives freshwater primarily from Penniar River in the north and east, respectively.
2.2 Sampling and Taxonomic Coverage
A total of 45 specimens of fishes collected in Nallavadu lagoon near Pooranankuppam village, representing 9 species, six genera, and 5 families were analyzed. Traditional taxonomic analyses were carried out used available literature to identify the specimen (FAO 1974; Rao 2003).
2.3 DNA Analysis
Total DNA was extracted from 0.25 g of tissue by using lysis buffer and followed by standard proteinase-K/phenol-chloroform-isoamyl alcohol-ethanol precipitation method (Sambrook et al. 1987) and fish DNA modified isolation protocol followed as described by Sachithanandam et al. (2012). The concentration of DNA was estimated using UV spectrophotometer method at 260/280 nm. Subsequently the DNA was diluted to final concentration of 100 ng/µL for further use. The 650–655 bp section of the mitochondrial (mt) DNA genome from the COI gene was amplified using a published universal degenerated primer set (Ward et al. 2005) synthesized by Sigma Aldrich Chemicals India Pvt. Ltd.
The polymerase chain reaction (PCR) was carried out in 25 µL consisting of 100 ng/µL of DNA and PCR master mix details described in Table 1. PCR was carried out in an Applied Bio systems AB-2720 Thermal cycler, and PCR thermal condition was followed (Sachithanandam et al. 2014, 2012). PCR products were resolved in 1 % agarose containing 0.5 mg/µL of ethidium bromide and viewed under UV Transilluminator and documented. Nucleotide sequencing was performed using the Sanger et al. (1977) method. Sequencing was performed using a BigDye Terminator Cycle Sequencing kit, following manufacturer’s instructions (Applied Biosystems, Foster City, CA, USA). The sequencing was done both in the forward and reverse directions.
2.4 Sequence Analysis
The DNA sequences were aligned both in forward and reverse directions for each individual fish and assembled using the SeqMan II version 5.03 (DNASTAR) and ChromaxSeq Version 3.1. The obtained studied species COI sequences analysis was done along with high similarity genetic identity reference sequences retrieved from the National Centre for Biotechnology Information (NCBI) GenBank. Studied species COI sequences used multiple and pairwise alignment was done using the ClustalW tool, and phylogenetic molecular evolutionary analyses were conducted through MEGA (Molecular Evolutionary Genetics Analysis) version 4.1. Sequence divergences were calculated using the Kimura two parameter (K2P) distance model and unrooted NJ phenograms based on K2P distances were created using MEGA 4.1 and were bootstrapped 1000 times to provide percentage bootstrap values for branch points (Tamura et al. 2007; Saitou and Nei 1987).
3 Results and Discussion
A total of 45 fish specimens belonging to 9 species, 6 genera, and 5 families were barcoded from Pooranankuppam brackish water in Nallavadu lagoon at Puducherry. The present study generated a total of 45 COI sequences for 9 species (No stop codon, insertions or deletions were found in any of the amplified sequences). All the amplified COI sequences were mean length 600 bp. Average nucleotide frequency were C (26.50 %), T (30.25 %), A (25.04 %) and G (18.21 %). The studied species were exhibited unique barcode haplotypes cluster of very closely related reference sequences, which permitted the discrimination of 96–98 % identity of species. All sequences were submitted to the NCBI GenBank.
3.1 COI Sequence Divergence Analysis
Out of the 9 species, 8 species belonging to perciformes and remaining one is belonging to Anguilla species were well discriminated though COI barcoding with average intraspecific genetic variability in the range of 0.005–0.020 compared with 0.203–0.354 for species within genera. The K2P genetic distances averaged 0.10 % within species, averaged 6.38 % within genera, 18.25 % within families, 21.55 % within orders and 23.18 % within classes, respectively were also obtained. Hence, overall taxonomic level increasing values, there was a 15 fold more pronounced difference among congeneric species than among conspecific individuals, and this value also produces a high level of resolution between clusters in the NJ tree to group the species to their corresponding genera and families with bootstrap values (Fig. 2).
3.2 Comments on Individual Genera
3.2.1 Ambassis spp.
The studied six genera, 9 species were obtained unrooted NJ tree shows that Ambassis spp. had separate cluster with three species. Among the three species Ambassis ambassis close genetic distance with Ambassis dussummieri. These species have clear external morphological features that allow differentiating each other; however A. commersoni are synonyms of A. ambassis through morphological taxonomy described but COI gene sequences had distinct genetic divergences obtained within A. commersoni and A. ambassis (K2P = 0.204). Moreover, genetic distance between this genus is 6.38 %, well above typical within genera values. Therefore, the lack of discrimination feature at morphological taxonomy level between these two species is likely misidentification. On the other hand, a reference sequence which was retrieved from NCBI data from Indian water and South Africa region also supports our data and shown in NJ tree (Fig. 3).
3.2.2 Anguilla sp.
Tropical eels are widely distributed about four species or subspecies in the Indian Ocean. In India two species of Anguilla (A. bicolor bicolor and A. nebulosa) reported. Based on the morphological characters study suggested that A. bicolor historically has been described as two subspecies, A. bicolor bicolor in the Indian Ocean and A. bicolor pacifica in the western Pacific Ocean and the seas around the northern parts of Indonesia (Ege 1939), but is apparently genetically homogeneous in the Indian Ocean (Minegishi et al. 2011). However, A. bicolor bicolor and A. bicolor pacifica COI genetic divergences study have been never investigated in details. The present study was to reveal the COI genetic divergency between A. bicolor bicolor collected from study site with close genetic identity reference sequences retrieved from GenBank and compared for further confirmation.
Collected specimen was identified as A. bicolor bicolor based on the morphological identification keys. Studied species COI gene sequence with nine reference sequence from GenBank used to constructed NJ phylogram formed 4 distinct clades (Fig. 4). The clade 1 in the top of the NJ phylogram contained Anguilla marmorata reference sequences JQ665825, HM345929, DQ521000 from different part had a cluster with 100 % identity. Clade 2 consists of (KF182302, KF182302) Anguilla bengalensis from Indonesian water, which showed 100 % similarity with same species from same region. Clade 3 contains studied species A. bicolor bicolor along with reference sequence of same species KF182304 (A. bicolor bicolor) from Malaysia with 100 % genetic similarity observed in NJ tree (Fig. 4). The studied species sequences with reference sequences retrieved from GenBank database to form clustered in NJ tree. From this analysis observed that COI gene sequences clearly proving the reliability of in identification of Anguilla spp.
4 Genetic Distance
A. bicolor bicolor Four specimen COI gene sequences Intraspecific mean genetic pair-wise distance was calculated and in the range of 0.03 from 0.02 to 0.050 and same species reference sequence (KF182304) from Malaysia also same values. Morphologically subspecies of A. bicolor pacifica reference sequence from Philippines water with studied species (A. bicolor bicolour) genetic diversity had 0.025. Interspecies genetic diversity analysis between studied species and A. marmorata (HM345929; JQ665825; JQ431413) showed 0.050 K2P values. Based on the A. bicolor bicolour species COI gene sequences are new reports from India for NCBI GenBank and BOLD system.
DNA barcoding is a well accepted taxonomic method which uses a short genetic marker to facilitate identification of a particular species even by non-specialist. CO1 has been accepted as universal barcode to delineate animal life. Hebert et al. (2004) proposed DNA barcoding works under the principle that interspecies variations are greater than the intra-species variations allowing one to distinguish the species using nucleotide sequences. Marine fish interspecific genetic distance well documented in different family in Indian water (Persis et al. 2009; Lakra et al. 2010; Ajmal Khan et al. 2011; Sachithanandam et al. 2012, 2014). Based on multiple studies Hebert et al. (2003) suggested an approximately 650 bp of the CO1 gene which is relatively easy to amplify with standard primers and is sufficient enough to obtain resolution on all levels between species and phylum for majority of the groups albeit with some exceptions (Ward et al. 2009). In the present study, earlier works in different animals are compared with the results of the pairwise comparisons within the genera 0.30 for perciformes spp., and Angullia sp. 0.022, the following results have been revealed: intraspecific variation in marine fishes has the value of 0.30 (Ward et al. 2005), Indian carangids has 0.24 (Persis et al. 2009), Indian Ocean marine fishes has 0.30 (Lakra et al. 2010), North American birds have 0.24, 0.23 and 0.27 (Hebert et al. 2004), Guyanese bats has 0.60 (Clare et al. 2011) and 0.46 for Lepidoptera (Hajibabaei et al. 2006). Atlantic and Brazilian cuzk eels species were discriminated 5.8 % genetic divergence observed (Mabragana et al. 2011). It is evident that congeneric distances are higher than the conspecific distances. In addition to the species identification, DNA barcoding has been used for identification of processed food products (Smith et al. 2008).
5 Conclusion
DNA barcoding is taking great endeavours of biological research genomics, phylogenetics and providing a comprehensive view into the biology, used to screen the large-scale genes, assign unknown individuals and discovery of new species. The present work concluded that studied species DNA barcode tool clearly discriminated in inter and intraspecific level without any ambiguity in genetic divergence of fish; these results support the utility of DNA barcodes for regional species identification of fishes. In future, Indian water need more groups of aquatic species for biodiversity assessment though COI gene sequence approach, will contribute increase biodiversity database. Moreover, COI gene based identification species enabled differentiation of species beyond doubt.
References
Ajmal Khan S, Lyla PS, Akbar John B, Prasanna Kumar C, Murugan S (2011) Identifying marine fin fishes using DNA barcodes. Curr Sci 101:9–12
Barnes RSK (1980) Coastal Lagoons. The natural history of a neglected habitat. Camb Stud Mod Biol 1:1–160
Becker S, Hanner R, Steinke D (2011) Five years of FISH-BOL: brief status report. Mitochondrial DNA 22(S.1):3–9
Bouchet P (2006) The magnitude of marine biodiversity. In: Duarte CM (ed) The exploration of marine biodiversity: scientific and technological challenges. Fundación BBVA, Bilbao, pp 31–64
Coyne JA, Orr HA (2004) Speciation. Sinauer Associates, Sunderland, p 545
Clare EL, Lim BK, Engstrom MD, Eger JL, Hebert PDN (2007) DNA barcoding of Neotropical bats: species identification and discovery within Guyana. Mol Ecol Notes 7:184–190
Clare EL, Lim BK, Fenton MB, Hebert PDN (2011) Neotropical Bats: estimating Species Diversity with DNA Barcodes. PLoS ONE 6(7):e22648. http://doi.org/10.1371/journal.pone.0022648
Crivelli AJ, Ximenes MC (1992) Alterations to the functioning of Mediterranean lagoons and their effects on fisheries and aquaculture. In: Finlayson M, Hollis T, Davis T (eds) IWRB. Special Publication, Grado. Managing Mediterranean Wetlands and Their Birds 20:134–140
Day F (1878) The fishes of India; being a natural history of the fishes known to inhabit the seas and fresh waters of India, Burma, and Ceylon. Parts 1–4. William Dawson and Sons Ltd., London, pp 778
Eschmeyer WN (2010) Catalog of fishes. http://research.calacademy.org/research/ichthyology/catalog/speciesByFamily.asp
Ege V (1939) A revision of the genus Anguilla Shaw: a systematic, phylogenetic and geographical study. Dana Rep 16:1–256
FAO (1974) Species identification sheet for fishery purposes. In: Fischer W, Whitehead PJP (eds) Eastern Indian Ocean (fishing area 57) and Western Central Pacific (fishing area 71), vol 1, pp C1–22
Farmworth EG, Golley FB (1974) Fragile ecosytems. Springer-Verlag, New York, pp 258
Grassle JF, Maciolek N (1992) Deep Sea species richness: regional and local diversity estimates from quantitative bottom samples. Am Nat Soc 39:313–334
Hebert PDN, Stoeckle MY, Zemlak TS, Francis CM (2004) Identification of birds through DNA barcodes. PLoS Biol 2:1657–1663
Hajibabaei M, Janzen DH, Burns JM, Hallwachs W, Hebert PDN (2006) DNA barcodes distinguish species of tropical Lepidoptera. Proc Nat Acad Sci USA 103:968–971
Hebert PDN, Cywinska A, Ball SL, DeWaard JR (2003) Biological identifications through DNA barcodes. Proc R Soc Lond B 270:313–321
Hubert N, Hanner R, Holm E, Mandrak NE, Taylor E (2008) Identifying Canadian freshwater fishes through DNA barcodes. PLoS ONE 3(6):e2490
Holmes BH, Steinke D, Ward RD (2009) Identification of shark and ray fins using DNA barcoding. Fish Res 95:280–288
Jarnegren J, Schander C, Sneli JA, Ronningen V, Young CM (2007) Four genes, morphology and ecology: distinguishing a new species of Acesta (Mollusca; Bivalvia) from the Gulf of Mexico. Mar Biol 152:43–55
Kjerfve B (1994) Coastal lagoon. In: Kjerfve B (ed) Coastal lagoon processes. Elsevier Oceanography Series, Amsterdam, pp 1–8
Lakra WS, Verma MS, Goswami M, Lal KK, Mohindra V, Punia P, Gopalakrishnan A, Singh KV, Ward R, Hebert PDN (2010) DNA barcoding Indian marine fishes. Mol Ecol Res 11:60–71
Mabragana E, Astarloa JMD, Hanner R, Zhang J, Castro MG (2011) DNA barcoding identifies Argentine fishes from marine and brackish waters. PLoS ONE 6(12):e28655
Mecklenburg CW, Moller PR, Steinke D (2010) Biodiversity of arctic marine fishes: taxonomy and zoogeography. Mar Biodiv 41(1):109–140
Minegishi Y, Gagnaire PA, Aoyama J, Bosc P, Feunteun E, Tsukamoto K, Berrebi P (2011) Present and past genetic connectivity of the Indo-Pacific tropical eel Anguilla bicolor. J Biogeogr. doi:10.1111/j.1365-2699.2011.02603.x
Nelson JS (2006) Fishes of the world, 4th edn. Wiley, New York, pp 601
Packer L, Gibbs J, Sheffield C, Hanner R (2009) DNA barcoding and the mediocrity of morphology. Mol Ecol Res 9:42–50
Padmavathy K, Poyyamoli G, Balachandran N (2010) Coastal dune flora, Nallavadu Village, Puducherry, India. Check List 6(2):198–200
Poyyamoli G, Padmavathy A, Balachandran N (2011) Coastal sand dunes—vegetation structure, diversity and disturbance in Nallavadu Village, Puducherry, India. Asian J Air Water Environ 8:115–122
Persis M, Reddy ACS, Rao LM, Khedkar GD, Ravinder K, Nasruddin K (2009) COI (Cytochrome oxidase—I) sequence based studies of Carangid fishes from Kakinada coast. India Mol Biol Rep 36:1733–1740
Peja N, Vaso A, Miho A, Rakaj N, Crivelli AJ (1996) Characteristics of Albanian lagoons and their fisheries. Fish Res 27:215–225
Quinn NW, Breen CN, Whitfield AK, Hearne JW (1999) An index for the management of South African estuaries for juvenile fish recruitment from marine environment. Fish Manag Ecol 6:421–436
Rao DV (2003) Guide to reef fishes of Andaman and Nicobar Islands. Zoological Survey of India, Kolkata, pp 555
Rasmussen RS, Morrissey MT, Hebert PDN (2009) DNA barcoding of commercially important salmon and trout species in North America. In: 3rd international barcode of life conference, Mexico City, p 84
Sachithanandam V, Mohan PM, Muruganandam N (2014) DNA barcoding of marine venomous and poisonous fish of family Scorpaenidae and Tetraodontidae from Andaman water. In: Venkataraman K, Sivaperuman C (eds) Marine faunal diversity in India taxonomy, ecology and conservation. Academic Press, Elsevier, London
Sachithanandam V, Mohan PM, Dhivya P, Muruganandam N, Baskaran R, Chaaithanya IK, Vijayachari P (2011) DNA barcoding, phylogenetic relationships and speciation of genus: Plectropomus in Andaman coast. J Res Biol 3:179–183
Sachithanandam V, Mohan PM, Muruganandam N, Chaaithanya IK, Dhivya P, Baskaran R (2012) DNA barcoding, phylogenetic study of Epinephelus spp. from Andaman coastal region. India Indian J Geo-Mar Sci 41:203–211
Saitou N, Nei M (1987) The Neighbour-joining method—a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sambrook J, Fritsch EF, Maniatus T (1987) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory Press, New York, pp 1659
Sanger F, Nicklen S, Coulson AR (1977) DNA sequencing with chain terminating inhibitors. Proc Nat Acad Sci USA 74(12):5463–5467
Smith PJ, Mc Veagh SM, Steinke D (2008) DNA barcoding for the identification of smoked fish products. J Fish Biol 72:464
Steinke D, Zemlak TS, Hebert PDN (2009) Barcoding Nemo: DNA-based identifications for the ornamental fish trade. PLoS ONE 4:e6300
Talwar PK (1990) Fishes of the Andaman and Nicobar Islands: a synoptic analysis. J Andaman Sci Assoc 6(2):71–102
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4. 0. Mol Biol Evol 24(8):1596–1599
Venkataraman K, Wafar M (2005) Coastal and marine biodiversity of India. Ind J Mar Sci 34(1):57–75
Ward RD, Zemlak TS, Innes BH, Last PR, Hebert PDN (2005) DNA barcoding Australia’s fish species. Philos Trans R Soc Ser Biol 360:847–1857
Ward RD, Hanner R, Hebert PDN (2009) The campaign to DNA barcode all fishes. FISHBOL J Fish Biol 74(2):329–356
Wong EHK, Hanner R (2008) DNA barcoding detects market substitution in North American seafood. Food Res Int 41:828–837
Whitehead D, Talwar PK (1976) Clupeoid fishes of the world (Suborder: clupeoidei): Engraulididae. American museum of natural history, New York, Part 4. Fishes 121:118–119
Acknowledgments
The Authors express their sincere acknowledgement to, Vice-Chancellor, Pondicherry University and the constant help and encouragement of Dr. P. Vijayachari, Director, Regional Medical Research Centre (ICMR) Port Blair during this study. First Author thanks the UGC-RFSMS fellowship and Ministry of Earth Sciences (MoES) for funding this work. The authors thank the Director, NCSCM, MoEF&CC, Chennai, for providing scientific and administrative support.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Sachithanandam, V. et al. (2016). DNA Barcoding Identifies Brackish Water Fishes from Nallavadu Lagoon, Puducherry, India. In: Trivedi, S., Ansari, A., Ghosh, S., Rehman, H. (eds) DNA Barcoding in Marine Perspectives. Springer, Cham. https://doi.org/10.1007/978-3-319-41840-7_16
Download citation
DOI: https://doi.org/10.1007/978-3-319-41840-7_16
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-41838-4
Online ISBN: 978-3-319-41840-7
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)