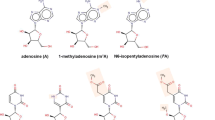
Abstract
A distinct feature of selenocysteine (Sec) biosynthesis is that this amino acid is synthesized on its tRNA, designated tRNA[Ser]Sec. Sec is then inserted into protein in response to the codon, UGA, as the 21st proteinogenic amino acid. In eukaryotes and archaea, Sec biosynthesis involves several steps. Transfer RNA[Ser]Sec is first aminoacylated by seryl-tRNA synthetase with serine. O-Phosphoseryl-tRNA[Ser]Sec kinase phosphorylates seryl-tRNA[Ser]Sec forming O-phosphoseryl-tRNA[Ser]Sec that in turn reacts with Sec synthase (SEPSECS) in the presence of selenophosphate yielding Sec-tRNA[Ser]Sec. Selenophosphate is generated by selenophosphate synthetase 2 (SPS2) from selenide and/or other selenium metabolites and ATP. Interestingly, sulfide can replace selenide in the reaction involving SPS2 yielding thiophosphate which can then form cysteine- (Cys)-tRNA[Ser]Sec in the presence of SEPSECS. The Cys moiety on Cys-tRNA[Ser]Sec can donate Cys to protein in response to UGA codons at internal positions of mammalian selenoprotein mRNAs. Cys/Sec replacement occurs naturally in vivo and the amount of replacement is dependent on the level of selenium in the diet.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Codon UGA
- Cysteine
- De novo biosynthesis
- Selenocysteine
- Selenocysteine tRNA
- Selenoprotein biosynthesis
- Selenoproteins
1 Introduction
The selenium-containing amino acid , selenocysteine (Sec), is biosynthesized unlike any other known amino acid in eukaryotes in that it is synthesized on its tRNA, designated tRNA[Ser] Sec . Transfer RNA[Ser]Sec has many unique features that distinguish it from all canonical tRNAs as detailed in Chap. 1. Other than seryl-tRNA synthetase (SERS), which initiates the biosynthesis of Sec by aminoacylating tRNA[Ser]Sec with serine, all enzymes involved in the synthesis of this amino acid are exclusive to the Sec pathway (see [1, 2] and references therein).
Interestingly, cysteine (Cys) may replace Sec on tRNA[Ser] Sec , and Cys may be incorporated into protein in place of Sec as discussed below. Replacement of Cys with Sec or Sec with Cys in cellular metabolism is not too surprising, since the structures of these two amino acids are so similar, differing only in the presence of a selenium versus a sulfur atom. Furthermore, these two amino acids have similar chemical properties and, when present in active sites of enzymes, may catalyze some of the same reactions. An example of the replacement of sulfur with selenium is also known that involves selenomethionine that may be incorporated into protein in place of methionine (reviewed in [3, 4]). In addition, generation of selenized yeast that largely contains selenium in the form of selenomethionine is widely used in the dietary supplement industry for producing selenium supplements [5–7]. Although sulfur replacing selenium in protein is less common, it has been shown to occur in vitro in mammalian cells in culture and in vivo in livers of mice (see [2] and references therein). The biosynthesis of Sec and the molecular mechanism of how Cys replaces Sec in specific selenoproteins are described below.
2 Sec Biosynthesis
The Sec codon, which occurs at internal positions of selenoprotein mRNAs, is UGA. The location of the UGA Sec codon within selenoprotein mRNAs can be anywhere from near the N-terminus to the penultimate codon at the C-terminus. Some selenoprotein mRNAs also employ UGA as a stop codon [8]. The distance between the UGA Sec codon and the Sec Insertion Sequence (SECIS ) element, which plays a major role in dictating a UGA codon as Sec, is an important factor in determining the efficiency of Sec insertion into protein (see [9, 10] and references therein). Interestingly, in the ciliate Euplotes, UGA can code for either Sec or Cys, even within the same mRNA, and the location of the SECIS element relative to the UGA Sec codon is the major governing factor dictating whether this codon designates Sec or Cys [11].
The biosynthesis of Sec on tRNA[Ser]Sec was initially established in Escherichia coli (E. coli) by Bӧck and coworkers (reviewed in [12]). Its synthesis in E. coli is detailed in Chap. 5 and will not be further discussed herein. The biosynthesis of Sec on tRNA[Ser] Sec in eukaryotes and archaea is more complex than in eubacteria [1, 13] and involves five major steps: (1) tRNA[Ser] Sec is initially aminoacylated with serine in the presence of SERS to form seryl-tRNA[Ser]Sec ; (2) a kinase, phosphoseryl-tRNA[Ser]Sec kinase (PSTK), phosphorylates the serine moiety on Ser-tRNA[Ser]Sec to form the intermediate, O-phosphoseryl-tRNA[Ser]Sec (pSer-tRNA[Ser] Sec ); (3) synthesis of the active selenium donor, selenophosphate (H2SePO3 −), is catalyzed by selenophosphate synthetase (SPS2); and (4) and (5) pSer-tRNA[Ser]Sec is converted to another intermediate, most likely dehydroalanyl-tRNA[Ser]Sec , and donation of H2SePO3 − to this intermediate forms Sec-tRNA[Ser]Sec , wherein both these steps are carried out by SEPSECS (see Fig. 4.1).
2.1 Discoveries That Provided the Foundation for Sec Biosynthesis
Sec tRNA[Ser] Sec was discovered in 1970 in mammalian and avian livers and initially characterized as a minor seryl-tRNA that either formed pSer-tRNA[Ser] Sec [14] or decoded specifically the termination codon, UGA [15]. Due to the fact that this tRNA recognized only UGA and decoded UGA in protein synthesis [16], it was proposed to be a nonsense suppressor tRNA. It was subsequently shown to have numerous novel features unique to this tRNA (Chap. 1). The important findings in these early studies were the observations that tRNA[Ser]Sec was aminoacylated with serine, decoded specifically UGA and formed pSer-tRNA[Ser] Sec .
2.2 Step 1: Aminoacylation of tRNA[Ser]Sec
As noted above, the first step in the biosynthesis of Sec is the attachment of serine to tRNA[Ser] Sec , which is carried out by SERS and requires ATP and Mg2+, as shown in Fig. 4.1.
2.3 Step 2: Phosphorylation of the Serine Moiety
PSTK remained elusive for more than 30 years until it was discovered using the methods of comparative genomics. The following rationale was used in identifying the PSTK gene (Pstk) [17]. Since SEPSECS, which synthesizes Sec in bacteria, is absent in archaea and eukaryotes, it was assumed that eukaryotes and archaea synthesize Sec by the same pathway that involves pSer-tRNA[Ser]Sec as an intermediate. Bioinformatics analyses for kinase genes present exclusively in those archaea that also encoded selenoprotein genes revealed four possible kinase genes that might be Pstk. Analysis of the sequences of these four kinase genes for orthologs in those eukaryotes, which possess the Sec biosynthetic and insertion machinery, revealed a single kinase gene that might encode PSTK. Subsequent biochemical characterization of the protein product demonstrated that it was indeed PSTK [17].
2.4 Step 3: Generation of Selenophosphate , the Active Selenium Donor
Two selenophosphate synthetase genes, designated Sps1 [18, 19] and Sps2 [20], were discovered in mammals that had homology to bacterial selD, and were proposed to also be responsible for synthesizing H2SePO3 −. Interestingly, Sps2 had a UGA codon in the position corresponding to a Cys codeword in selD suggesting that the expressed protein product synthesized from Sps2 self-regulates selenoprotein synthesis [20]. Biochemical analyses revealed that SelD and the SPS2 mutant in which Sec is replaced with Cys, synthesized H2SePO3 −, but SPS1 did not, demonstrating that SPS2 is the functional selenophosphate synthetase in mammals and that SPS1 must have another role in cellular metabolism [1, 21].
2.5 Sec Synthesis
Since no ortholog of bacterial selA, which synthesizes Sec on tRNA[Ser] Sec , was found in mammals, a comparative genomics approach was also used to identify this protein. Eukaryotic and archaea genomes were scanned for co-occurrence of candidate genes involved in Sec biosynthesis, selenoprotein genes and known components of Sec machinery [1]. This search revealed a gene that matched a protein identified previously in patients with autoimmune chronic hepatitis and named soluble liver antigen (SLA) [22]. SLA had been shown to co-precipitate with tRNA[Ser] Sec and to exist in a complex with other proteins involved in Sec metabolism, providing further evidence of its possible role in Sec biosynthesis .
The corresponding gene of SLA is now called Sepsecs [1]. PSer-tRNA[Ser]Sec was found to bind strongly to the putative SEPSECS, while tRNA[Ser] Sec bound less well, seryl-tRNA[Ser] Sec bound poorly, and tRNASer and seryl-tRNASer did not bind. The facts that (1) O-phosphoserine was efficiently hydrolyzed from this substrate by SEPSECS, and (2) the resulting intermediate readily accepted the active selenium donor to form Sec-tRNA[Ser] Sec further supported the hypothetical function of SECSEPS. Overall, the data demonstrated that SEPSECS functioned as the Sec synthase in eukaryotic and archaeal Sec biosynthesis [1].
3 De Novo Synthesis of Cys and Cys/Sec Replacement In Vitro and In Vivo
As noted above, Cys and Sec are structurally similar. Their genetic language is different, however, in that Cys is decoded by the codewords, UGU/UGC, while Sec is decoded by UGA. Replacement of Sec with Cys in thioredoxin reductase 1 (TXNRD1 ) has been observed in the livers of selenium deficient rats [23]; however, the mechanism was not determined, and thus, we assessed the mechanism of insertion of Cys into TXNRD1 in lieu of Sec as discussed below.
3.1 In vivo Studies
When thiophosphate (H2SPO3 −) was added to the media of NIH 3T3 cells and the intracellular selenoproteins analyzed, Cys was found to have replaced Sec in TXNRD1 virtually completely [24]. On the other hand, when mice were maintained on selenium deficient (0 ppm selenium), selenium adequate (0.1 ppm selenium), or selenium enriched (2.0 ppm selenium) diets, and TXNRD1 and thioredoxin reductase 3 (TXNRD3) in liver subsequently isolated, purified and analyzed by mass spectrometry [24], about 50 % of liver TXNRD1 and 3 were found to contain Cys in place of Sec in selenium deficient animals, about 10 % in selenium-adequate animals and no replacement in selenium-enriched animals. These studies provided the background for determining the precise mechanism of how such replacement occurs.
3.2 In Vitro Studies
To establish the replacement of Sec with Cys in vitro, we initially rationalized that sulfide could replace selenide in synthesizing Cys catalyzed by SPS2 [24]. Thus, the enzymes and other components required for synthesizing Sec-tRNA[Ser] Sec from pSer-tRNA[Ser]Sec were prepared and Sec biosynthesis carried out. Addition of H2SPO3 − to the reaction with pSer-tRNA[Ser] Sec and SEPSECS yielded Cys-tRNA[Ser]Sec as did incubation of pSer-tRNA[Ser]Sec , SEPSECS, sodium sulfide, ATP and SPS2(Cys). These two reactions demonstrated that H2SPO3 − could replace H2SePO3 − yielding Cys-tRNA[Ser]Sec [24]. De novo synthesis of Cys on tRNA[Ser] Sec in mammals is shown in Fig. 4.1 (lower panel).
4 Concluding Remarks
The biosynthetic pathway by which the essential element selenium is incorporated into Sec, the 21st amino acid in the genetic code, has been established in eukaryotes and archaea (Fig. 4.1). As shown in the figure, Sec biosynthesis occurs on its tRNA, tRNA[Ser] Sec , which represents the only known amino acid in eukaryotes whose synthesis takes place on its tRNA, and was the last proteinogenic amino acid in mammals whose biosynthesis was resolved. Sec-tRNA[Ser] Sec donates its Sec moiety to the nascent polypeptide chain in response to the UGA codon in mRNA generating a selenoprotein product (Chap. 2). Selenium and selenoproteins play major roles in the many health benefits attributed to selenium and some of these roles are: (1) serving as a cancer chemopreventive agent (Chap. 27); (2) delaying the onset of AIDS in HIV positive patients (Chap. 28); (3) reducing the incidence of heart disease, (4) boosting immune function (Chap. 42); (5) and regulating the aging process. Thus, assessing the pathway of how selenium makes its way into protein provides an important step in understanding the overall function of this element in health.
Two of the genes involved in Sec biosynthesis are Pstk and Sepsecs (Fig. 4.1). They were both identified in mammals using computational, comparative genomics approaches, followed by biochemical analyses. These data suggested that the selenoprotein biosynthetic pathway shown in Fig. 4.1 is the same used in all archaea and eukaryotes that synthesize selenoproteins .
The issue whether SPS1 or SPS2 or both enzymes were used to synthesize the active selenium donor, H2SePO3 −, was resolved largely by biochemical studies. SPS2 was found to be the enzyme that synthesizes H2SePO3 −, while the exact role of SPS1 is still elusive [1, 21]. However, the latter has recently been shown to be an essential protein, with a role in regulating redox homeostasis in mammals [25]. Cys in place of Sec in TXNRD1 and TXNRD3 was found to occur in vivo in both cells in culture and in mice, and the mechanism of how this replacement occurred was established using in vitro studies [24]. Thus, Cys can be synthesized de novo and the replacement of Sec with Cys suggests novel roles of Cys in mammalian metabolism.
References
XM Xu et al 2007 PLoS Biol 5:e4
AA Turanov et al 2011 Adv Nutr 2:122
GN Schrauzer 2000 J Nutr 130:1653
TG Sors et al 2005 Photosyn Res 86:373
M Kieliszek, S Blazejak 2013 Nutrition 29:713
MP Rayman 2004 Br J Nutr 92:557
CM Weekley, HH Harris 2013 Chem Soc Rev 42:8870
VM Labunskyy et al 2014 Physiol Rev 94:739
Z Stoytcheva et al 2006 Mol Cell Biol 26:9177
AA Turanov et al 2013 Nucleic Acids Res 41:6952
AA Turanov et al 2009 Science 323:259
S Yoshizawa, A Bock 2009 Biochim Biophys Acta 1790:1404
J Yuan et al 2006 Proc Natl Acad Sci U S A 103:18923
PH Maenpaa, MR Bernfield 1970 Proc Natl Acad Sci U S A 67:688
D Hatfield, FH Portugal 1970 Proc Natl Acad Sci U S A 67:1200
A Diamond et al 1981 Cell 25:497
BA Carlson et al 2004 Proc Natl Acad Sci U S A 101:12848
SC Low et al 1995 J Biol Chem 270:21659
IY Kim, TC Stadtman 1995 Proc Natl Acad Sci U S A 92:7710
MJ Guimaraes et al 1996 Proc Natl Acad Sci U S A 93:15086
XM Xu et al 2007 Biochem J 404:115
C Gelpi et al 1992 Proc Natl Acad Sci U S A 89:9739
J Lu et al 2009 FASEB 8:2394
XM Xu et al 2010 Proc Natl Acad Sci U S A 107:21430
R Tobe et al 2016 Biochem J doi:10.1042/BCJ20160393
Acknowledgements
This work was supported by the Intramural Research Program of the National Institutes of Health, NCI, Center for Cancer Research to D.L.H. and NIH grants CA080946, GM061603 and GM065204 to V.N.G.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer Science+Business Media, LLC
About this chapter
Cite this chapter
Gladyshev, V.N., Carlson, B.A., Hatfield, D.L. (2016). Pathways in De Novo Biosynthesis of Selenocysteine and Cysteine in Eukaryotes. In: Hatfield, D., Schweizer, U., Tsuji, P., Gladyshev, V. (eds) Selenium. Springer, Cham. https://doi.org/10.1007/978-3-319-41283-2_4
Download citation
DOI: https://doi.org/10.1007/978-3-319-41283-2_4
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-41281-8
Online ISBN: 978-3-319-41283-2
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)