Abstract
Nutrigenomics approaches have contributed to our understanding of how selenium impacts metabolic pathways, homeostatic control and disease risk. In this chapter, we discuss the known genetic polymorphisms in genes encoding selenoproteins and components of the selenoprotein synthesis machinery. Furthermore, the consequences of these genetic variants on the synthesis and activity of individual selenoproteins within the context of the metabolic pathway in which they are involved, including the impact of these variants on the overall selenoproteome, as a result of the shared synthesis machinery between selenoproteins are discussed. The evidence for the association of these genetic variants with several chronic diseases is presented, with a specific emphasis on functional variants.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Association study
- Cancer
- Nutrigenomics
- Selenoprotein
- Selenoprotein hierarchy
- Single nucleotide polymorphisms
1 Introduction
Selenoproteins play pivotal roles in many biochemical pathways and in particular in the response to oxidative stress and endoplasmic reticulum (ER) stress . In a nutritional context, it is important to consider the effect of selenium (Se) intake on the overall balance of selenoproteins and related pathways; similarly, from a genetic perspective, the effects of inter-individual variations need to be assessed in the overall Se pathway. Here, we discuss how genetic studies are providing novel perspectives about the mechanisms by which Se intake and metabolism are related to disease risk.
Single Nucleotide Polymorphisms (SNPs) represent 90 % of the genetic variations among individuals, and some genetic variants alter gene or protein regulation or protein activity and are thus functionally relevant. During evolution, the ability of our ancestors to survive relied on their capacity to combat metabolic stresses and infections and ensure their reproduction; these functions are strongly supported by various selenoproteins and therefore highly dependent on Se intake . It is possible that genetic variants which modulate selenoprotein synthesis or activity in different conditions of Se supply could have been selected as a result of the disparate geological distribution of Se and exposure to various pathogens and stresses. Thus, the pressure of selection imposed by environmental forces may partially explain the variations in allelic frequencies currently observed among different populations. Supporting this hypothesis, several studies have identified SNPs in selenoprotein genes corresponding to signatures of recent genetic adaptation , suggesting a local adaptation to low Se content in the soil. In Asian populations living in Se-deficient regions, it was reported that a selective sweep occurred recently at the GPX1 locus [1]. Similarly, allele frequency shifts for SNPs in selenoprotein genes were observed in populations living in the severe Se-deficient regions of China [2]. Moreover, a recent study identified a strong positive natural selection in individuals of European decent, for known functional variants in GPX1, SELENBP1, GPX3, GPX2 and SELO genes suggesting local adaptation tolow soil Se [3].
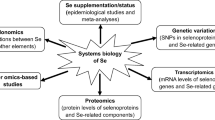
Schematic diagram illustrating interactions of selenoprotein SNPs and Se supply on selenoprotein function and disease risk. The diagram illustrates how SNP-Se interactions could affect selenoprotein synthesis and activity and consequently downstream biochemical pathways known to be crucial in the response to oxidative and ER (endoplasmic reticulum) stress and in the maintenance of mitochondrial redox status. As disruption of these biochemical pathways is commonly observed in many chronic disorders, the existence of genetic associations linking these SNP with diseases indicate the importance of selenoproteins’ function in these pathways. Genes (dark grey, italic) in which functional SNPs have been associated with disease risk are indicated at subcellular location in which the corresponding selenoproteins function is required
With the increase in human lifespan, genetic variations in Semetabolism have been proposed to be associated with risks for several age-related diseases, including cancer, diabetes, immunological or neurological disorders and cardiovascular diseases. Many of these disorders share a common basis for their development such as impaired cellular maintenance mechanisms and responses to stress. These metabolic pathways involve many selenoproteins. Thus, genetic factors affecting selenoprotein activity or synthesis, have the potential to modify individual risk to chronic disease (Fig. 13.1).
Three types of studies have shown links between genetic variants in selenoprotein genes and disease susceptibility (Tables 13.1, 13.2, 13.3, and 13.4): (1) the study of a small number of candidate SNPs of known functional relevance in which it is assumed that the functional consequences of the SNP on the gene/protein regulations could contribute to the disease risk; (2) the broader study of SNPs in several selenoprotein genes or screening of genes across the Se metabolic pathways, usually using a combination of tagSNPs and functional variants; and (3) large scale genome-wide association studies (GWAS) in which selenoprotein genes were associated with a disease trait. Focussed studies of well-characterized SNPs provide mechanistic backing to epidemiological studies and also often allow measurements of biomarkers of Se status and stress, particularly important in the case of Se since intake varies across different populations [4]. Screening studies involving a wider range of SNPs regardless of known functionality and large-scale GWAS have the advantage of wide genetic coverage but are less likely to have environmental and dietary data available.
2 Selenium Metabolism and Transport
Selenoprotein synthesis depends upon Se intake, Se transport from the liver to other organs, Se conversion to selenocysteine (Sec) and its incorporation into selenoproteins (Chaps. 2 and 4). Genetic variants in two genes involved in Sec synthesis (Table 13.4), Sec tRNA synthase (SEPSECS) and selenophosphate synthetase 1 (SEPHS1), have been linked to Crohn’s Disease in a Caucasian population from New Zealand with sub-optimal Se intake [5, 6]. Moreover, SNPs in SEPP1, the gene coding for selenoprotein P (SePP, SEPP1), which transports hepatic Se to other tissues, have been shown to affect Se delivery, Sec incorporation throughout the body and disease risk [7]. In particular, two genetic variants, rs3877899 and rs7579 (SEPP1), affect the plasma SePP isoform pattern, response to Se supplementation in healthy individuals with sub-optimal Se intake and selenoprotein synthesis [8, 9]. In support of this, expression of SePP and GPx1 was affected by both variants in patients with mild cognitive impairment supplemented with Se [10]. Moreover, rs3877899 and rs7579 were associated with risk for several cancers (Table 13.3) in different European populations [7, 11–13] and various SEPP1 variants were linked to colorectal (CRC) [14] or breast (BC) cancer in Native Americans [15]. In Europeans, risk of prostate cancer (PCA) or advanced PCA was linked to rs7579 (SEPP1) [12, 13] and modulated by an interaction between rs3877899 (SEPP1) and rs4880, a known functional SNP in SOD2 (manganese superoxide dismutase) [16]. In a US population, rs13168440 (SEPP1) was also shown to affect PCA risk [17], but other studies have failed to link SNPs in SEPP1 to PCA [13, 18, 19] or disease recurrence [20]. In addition, both rs3877899 and rs7579 (SEPP1) were also shown to influence body mass index, blood pressure, peripheral arterial disease occurrence and abdominal aortic aneurysm development in overweight and obese subjects [21].
Moreover, considerable mechanistic evidence suggest a role for SePP in glucose and insulin metabolism [22–24]. In support of this, four SNPs in SEPP1 (rs28919926, rs146125471, rs168727790 and rs7579) were shown to be associated with an altered fasting or acute insulin responses in two different Hispanic cohorts [25].
It is interesting to note that the effects of genetic variants in factors involved in Sec conversion and transport on disease risk are often influenced by Se status and/or ethnicity (hence genetic background), consistent with evolutionary adaptation of populations to geographical differences in Se distribution.
3 The Effects of SNPs on the Sec Incorporation Machinery
Sec incorporation requires the recoding of a UGA codon for Sec and the binding of factors from the selenoprotein synthesis machinery to the Sec insertion sequence (SECIS) stem-loop structure within the 3′untranslated region (3′UTR) of selenoprotein mRNAs (reviewed in Chap. 2). In addition, SECIS-binding protein 2 (SBP2) plays a major role in this process.
The 3′UTR regions of selenoprotein mRNAs play a key regulatory role in the so-called selenoprotein hierarchy [26], a mechanism by which the synthesis of the various selenoproteins is affected differently by limiting conditions of Se supply [27, 28]. The hierarchy reflects the fact that all selenoproteins share the same incorporation machinery and the same tRNA carrying Sec for their synthesis and therefore there is in essence a competition between the 25 selenoprotein mRNAs for the available synthesis machinery and Sec. As a result, a genetic variant in the 3′UTR of one selenoprotein has the potential to affect synthesis, not only of the selenoprotein coded by that mRNA, but also synthesis of other selenoproteins. Similarly, a SNP in a gene coding for factors involved in the selenoprotein biosynthesis machinery have the potential to affect the synthesis of all selenoproteins.
In particular, mutations in the selenoprotein N gene (SELN) within the gene region corresponding to the SECIS region of the 3′UTR were associated with reduced binding affinity of SBP2 for SECIS, lower expression of SelN and congenital muscular dystrophy [29]. Moreover,missense mutations in SBP2 led to poor Sec incorporation, poor thyroid function or muscular dystrophy, low expression of all selenoproteins and increased sensitivity to oxidative stress [30, 31]. However, mutations in selenoprotein genes that directly cause genetic disease are rare. On the contrary, common SNPs may have more subtle effects on selenoprotein metabolism, but in conjunction with other factors, such as dietary Se intake, can lead to altered risk for many diseases. Three functional genetic variants, one in GPX4 (rs713041) [32] and two in SEP15 (rs5859 and rs5845) [33], which induce base changes in a region nearby or within the SECIS element in corresponding transcripts, have been shown to reduce the efficiency of Sec incorporation in the corresponding protein and to affect the selenoprotein hierarchy and disease risk [27, 33–35].
Originally identified as a C/T variant in the gene region corresponding to the 3′UTR region of GPX4 mRNA [32], rs713041 illustrates how SNPs in selenoprotein genes can be studied from a functional perspective and how a SNP affecting the 3′UTR region impacts selenoprotein hierarchy . The C variant was shown to induce reporter gene expression to a greater extent than the T counterpart [36], to have a stronger binding affinity for the selenoprotein synthesis machinery inRNA-protein binding assays [35] and to alter the pattern of selenoprotein synthesis, especially during Se-depletion [36, 37]. Moreover, in healthy individuals, rs713041 was shown to affect expression of blood selenoproteins in response to Se supplementation, consistent with an effect on the selenoprotein hierarchy in vivo [35]. Finally, human umbilical vein endothelial cells from individual donors and monocytes [38] expressing the T-variant showed an increased expression of vascular cell adhesion protein 1 and adhesion to monocytes compared with cells from the CC individuals.
4 Genetic Variants Affecting Redox-Active Selenoproteins
Many selenoproteins are involved in the control of cellular redox balance and antioxidant defense and can be divided into two main classes of redox-active selenoenzymes, the GPX and thioredoxin reductases (TXNRD). Functional SNPs in GPX1-4 and TXNRD1-2 have been shown to affect anti-oxidant defense and disease risk (Tables 13.1, 13.2, 13.3, and 13.4).
4.1 Genetic Variants in GPX4
As mentioned above, rs713041 (GPX4) was shown to affect the selenoprotein hierarchy [35], as well as the sensitivity to oxidative challenge [37, 38]. Interestingly, in a Se-deficient Chinese population (Table 13.2), rs713041 together with rs4807542, another GPX4 SNP in high linkage disequilibrium with rs713041, were found to affect Kashin-Beck disease (KBD) risk, with GPX4 mRNA levels being reduced in KBD patients compared with controls [39]. Moreover, rs713041 has been linked to risk of CRC in Czech and Scottish populations [7, 36], to lung and laryngeal cancers in a Polish population [40] and risk of BC mortality in a British population [41]. In the Czech population, CRC risk was affected by polymorphisms in both SEPP1 (rs7579) and GPX4 (rs713041) [7] and was further modulated by significant genetic interactions between SNPs in SEPP1 (rs7579 and rs3877899), known to affect Se bioavailability, and variants in SEP15 (rs5859) or GPX4 (rs713041), known to affect Sec incorporation [7]. These interactions suggest that in carriers of the combined genotypes, the altered pattern of synthesis of selenoproteins affects the individual’s ability to respond to stress. Similarly, genetic interactions between rs713041 in GPX4, rs960531 in TXNRD2 and rs4880 in SOD2 mirror the interactions of these enzymes in mitochondrial redox function and suggest that the genetic interactions could affect an individual’s ability to counteract oxidative stress in the mitochondria [7]. Additionally, two independent GWAS linked the GPX4 locus to Crohn’s disease [42, 43], consistent with involvement ofGPx4 in inflammatory responses and NF-kB regulation [44, 45].
4.2 Genetic Variants in GPX1
Cellular GPx1 is an important antioxidant enzyme in mammals. In humans, the enzyme activity is affected by rs1050450, a coding C/T SNP in the GPX1 gene, inducing a Pro (CC) to Leu (TT) amino acid change at position 198 of the amino acid sequence. The Leu variant exhibits lower activity compared with the Pro counterpart [46, 47] and, during Se-supplementation, GPx1 activity is stimulated less in TT carriers compared with CC carriers [48, 49]. Moreover, significantly higher levels of DNA oxidation were observed in Leu carriers, probably as a result of reduced GPx1 activity [48, 49]. Surprisingly, however, an increased susceptibility to DNA strand breaks was observed in CC subjects, but not TT, during Se withdrawal [48].
Many studies have linked rs1050450 to risk for several disorders (Table 13.1), including various cancers [12, 13, 40, 46, 50–56], Alzheimer’s disease [57], metabolic syndrome and obesity [58, 59], and KBD [60]. In particular, rs1050450 was linked to BC risk in some [46], but not other US populations [61–63], although a meta-analysis found an increased BC risk among women of African descent [64]. However, the association was replicated in several European populations [11, 47, 65]. The consistent differences observed between studies of European and North American populations suggest that the effect of rs1050450 on BC risk may be partially influenced by Se status. In a Danish population, the Leu variant was associated with reduced GPx1 activity, increased risk of BC and a higher grade of ductal tumors [11, 47]. Additionally, pre-diagnostic erythrocyte GPx1 activity was lower in Leu females, following hormone replacement therapy and who develop BC later in life, compared with controls [11]. Furthermore, a GCG repeat polymorphism in GPX1, resulting in variant protein sequences containing between 5 and 7 alanines, was linked to BC risk [63], supporting a role for GPx1 activity in protecting breast tissue from carcinogenesis.
Variants in the GPX1 gene have also been linked to PCA, another cancer influenced by sex-hormones [13]. Moreover, rs1050450 together with rs18006688 (a GPX1 SNP in high linkage disequilibrium) modified the association between lead exposure and glioblastoma [66], suggesting that the reduced GPx1 activity of the Leu variant could impair the protection against oxidative damage generated by lead [66]. Like GPX4, GWAs have linked the GPX1 locus to inflammatory conditions such as Crohn’s disease [42], inflammatory bowel disease [43] and ulcerative colitis [67].
4.3 Genetic Variants in TXNRDs
Mammalian cells have three isozymes of thioredoxin reductases, including cytoplasmic and nuclear TXNRD1 and mitochondrial TXNRD2. The relationship between diseases and genetic variants in TXNRD1 and TXNRD2 has been investigated in studies using tagSNPs including a GWAs [68, 69], in a study investigating association of SNPs in several selenoprotein genes on PCA risk, grade and recurrence [20], and in a study investigating the association of polymorphic variants in Se metabolism with PCA risk [19]. In the latter study, carried out in a German population with low Se intake, tagSNPs in SELK, TXNRD1 and TXNRD2 were found to interact with plasma Se or SePP status to modulate risk of advanced disease [19] (Table 13.4).
In addition, SNPs in other selenoproteins involved in the protection against oxidative damage were linked to disease. In a US population, three SNPs in GPX3 and one in GPX2 were significantly associated with risk of rectal cancer, but not with either colon cancer or adenoma [70] (Tables 13.3, and 13.4). In the MSRA gene, coding for methionine sulfoxide reductase A, rs10903323, was associated with coronary artery disease risk in a Chinese population [71].
5 Genetics of Endoplasmic Reticulum Selenoproteins
In eukaryotic cells , the ER is not only responsible for the synthesis, post-translational modification and correct folding of membrane and secreted proteins, but also for intracellular Ca2+ homeostasis and lipid biosynthesis [72]. Alteration of protein folding caused by changes in intracellular Ca2+ levels, redox state, nutrient status, protein synthesis rate or inflammatory stimuli, can result in ER stress and activation of the unfolded protein response to remove misfolded proteins [72]. ER dysfunction and prolonged ER stress have been implicated in diseases such as cancer, diabetes and Alzheimer’s disease [72]. Seven human selenoproteins have been shown to be associated with the ER, which are the 15-kDa selenoprotein (Sep15), type 2 iodothyronine deiodinase and selenoproteins K, M, N, S, and T [73]. Selenoprotein S is a component of the ER-associated protein degradation pathway and is involved in the removal of misfolded proteins from the ER lumen [73]. Sep15 has been implicated in the formation of disulfide bonds and the quality control of protein folding [34], while SelK plays a key role in calcium signalling in immune cells [74].
Genetic polymorphisms in SELS, SELK and SEP15 have now been linked to various cancers and inflammatory conditions, consistent with these SNPs affecting correct ER function (Table 13.4). In SELS gene, rs34713741 affects levels of pro-inflammatory cytokines, IL-6, IL-1β and TNF-α [75], and risk of CRC in Korean [76] and Czech [7] populations. In the Czech population, rs34713741 was associated with greater CRC risk, and in the Korean population a second variant in close proximity led to increased risk (in females only). The replication of the association in these two diverse populations strongly indicates that, independently of lifestyle and dietary factors, SNPs in SELS influence CRC risk. Moreover, supporting a role of SelS in gastrointestinal function, rs34713741, was also linked to gastric cancer risk [77, 78]. No association was identified between six genetic variants in SELS and type 1 diabetes [79]. In contrast, rs225014 in (type 2 deiodinase gene), results in a Thr to Ala amino-acid change at position 92 in the protein sequence, with the Ala variant being less active and associated with type 2 diabetes [80–82].
In the SEP15 gene, rs5859 and rs5845, known to reduce Sec incorporation efficiency and Sep15 synthesis, affect BC risk in a Se-dependent manner in African American women, but not in Caucasians [33]. In breast tumors , a loss of heterozygosity at the SEP15 locus was observed [33, 83], indicating a potential tumor suppressor role of Sep15 in breast tissue. In addition, rs5859 and rs5845 were associated with increased risk of PCA in German [12] and New Zealand [51] males with low Se intake, and other SNPs in SEP15 were also linked to PCA mortality (Table 13.4) [20, 84]. CRC risk was also affected by rs5859 in a South Korean population [76] and by genetic interactions between SNPs in SEPP1 and rs5859 in SEP15 [7]. Moreover, rs5859 was linked to lung cancer in Polish individuals with low Se status [85]. The influence of these SNPs on disease risk and progression provides insight into the potential role of ER stress and protein folding control in disease etiology as well as on the importance of key selenoproteins in maintaining a healthy tissue.
6 Perspectives
Two main lessons can be learned from the studies described above. First, there is now considerable evidence for a number of genetic variants affecting selenoprotein synthesis and, in a limited number of cases, this has been linked to effects on response to dietary Se intake. Genetic epidemiology studies suggest that a number of these variants, notably those in SEPP1, SELS, GPX1, GPX4 and SEP15, modulate risk of various chronic diseases. In particular, these results suggest that variants in SEPP1, SEP15 and GPX1 affect PCA risk and progression, SNPs in SELS influence CRC risk and variants in GPX1 BC risk. However, the evidence linking these variants with disease risk comes largely from relatively small studies that often lack accompanying measures of Se status and often require replication. In addition, observed effects are often inconsistent between study populations, a phenomenon that likely reflects differences in the characteristics of the study populations, notably Se status. Further work is necessary to be confident of the clinical relevance of selenoprotein SNPs in different populations, both in terms of Se intake and ethnicity. Larger studies, combining genetics and biomarkers of Se status should provide a clearer picture of the links between Se, selenoprotein genetics and disease risk.
Second, identifying that SNPs in selenoprotein genes affect risk for several chronic diseases is compatible with the observation that most of these multifactorial diseases share a common basis, with the disruption of biochemical pathways involved in the response to oxidative and ER stress . As a result, these genetic associations help elucidate potential pathways affected in the etiology of disease for an individual. Selenoprotein synthesis depends on the distribution of Se between selenoproteins and the use of common synthesis machinery. Thus, the observation of genetic associations of a SNP with a disease may reflect an effect of this particular SNP on the whole selenoproteome, inviting us to combine genetic association studies with expression of selenoproteins in a tissue.
References
LH Foster, S Sumar 1997 Crit Rev Food Sci Nutr 37:211
L White et al 2015 Mol Biol Evol 32:1507
J Engelken et al 2016 Mol Biol Evol 33:738
GF Combs, Jr. 2001 Br J Nutr 85:517
L Gentschew et al 2012 Nutrients 4:1247
CD Thomson et al 2007 Br J Nutr 97:357
C Meplan et al 2010 Carcinogenesis 31:1074
C Meplan et al 2009 Antioxid Redox Signal 11:2631
C Meplan et al 2007 FASEB J 21:3063
BR Cardoso et al 2016 Food Funct 7:825
C Meplan et al 2013 PLoS ONE 8:e73316
A Steinbrecher et al 2010 Cancer Epidemiol Biomarkers Prev 19:2958
MS Geybels et al 2014 J Natl Cancer Inst 106:dju003
U Peters et al 2008 Cancer Epidemiol Biomarkers Prev 17:1144
AJ Pellatt et al 2013 PLoS ONE 8
ML Cooper et al 2008 Cancer Res 68:10171
KL Penney et al 2013 Prostate 73:700
MS Geybels et al 2015 Cancer Epidemiol Biomarkers Prev 24:178
C Meplan et al 2012 PLoS ONE 7:e48709
JP Gerstenberger et al 2015 Prostate 75:60
E Strauss et al 2014 Sci Rep 4:7061
H Misu et al 2010 Cell Metab 12:483
SJ Yang et al 2011 J Clin Endocrinol Metab 96:E1325
H Steinbrenner 2013 Free Radic Biol Med 65:1538
JN Hellwege et al 2014 Gene 534:33
FP Bellinger et al 2009 Biochem J 422:11
G Bermano et al 1996 Biochem J 320 ( Pt 3):891
V Pagmantidis et al 2005 FEBS Lett 579:792
V Allamand et al 2006 EMBO Rep 7:450
E Schoenmakers et al 2010 J Clin Invest 120:4220
E Schoenmakers et al 2016 in Diversity of Selenium Funcion in Health and Disease, R Brigelius-Flohé, H Sies Eds (CRC Press, Boca Raton, FL) p 343
S Villette et al 2002 Blood Cells Mol Dis 29:174
YJ Hu et al 2001 Cancer Res 61:2307
KV Korotkov et al 2001 J Biol Chem 276:15330
C Meplan et al 2008 Am J Clin Nutr 87:1019
G Bermano et al 2007 Genes Nutr 2:225
H Gautrey et al 2011 Biochim Biophys Acta 1810:284
LK Crosley et al 2013 Mol Nutr Food Res 57:2185
XH Du et al 2012 Br J Nutr 107:164
K Jaworska et al 2013 PLoS ONE 8:e59051
M Udler et al 2007 J Clin Oncol 25:3015
A Franke et al 2010 Nature Genetics 42:1118
L Jostins et al 2012 Nature 491:119
H Imai, Y Nakagawa 2003 Free Radic Biol Med 34:145
G Gong et al 2012 Genes Nutr 7:167
YJ Hu, AM Diamond 2003 Cancer Res 63:3347
G Ravn-Haren et al 2006 Carcinogenesis 27:820
E Jablonska et al 2015 Eur J Nutr doi:10.1007/s00394-015-1118-4
JC Miller et al 2012 American Journal of Clinical Nutrition 96:923
Z Arsova-Sarafinovska et al 2009 Int Urol Nephrol 41:63
N Karunasinghe et al 2012 J Nutrigenet Nutrigenomics 5:339
P Yang et al 2004 Carcinogenesis 25:1935
D Ratnasinghe et al 2000 Cancer Res 60:6381
O Raaschou-Nielsen et al 2007 Cancer Lett 247:293
Y Ichimura et al 2004 J Urol 172:728
H Zhao et al 2005 Urology 66:769
C Paz-y-Mino et al 2010 Am J Med Sci 340:373
M Kuzuya et al 2008 Am J Clin Nutr 87:1939
C Cominetti et al 2011 Nutrition 27:891
YM Xiong et al 2010 Osteoarthritis Cartilage 18:817
DG Cox et al 2004 Cancer Epidemiol Biomarkers Prev 13:1821
J Ahn et al 2005 Cancer Epidemiol Biomarkers Prev 14:2459
JA Knight et al 2004 Cancer Epidemiol Biomarkers Prev 13:146
J Hu et al 2010 Breast Cancer Res Treat 124:425
E Jablonska et al 2015 BMC Cancer 15:657
P Bhatti et al 2009 Cancer Epidemiol Biomarkers Prev 18:1841
A Julia et al 2014 Hum Mol Genet 23:6927
Z Ye et al 2014 Front Genet 5:125
EN Vithana et al 2012 Nat Genet 44:1142
U Haug et al 2012 Genes Chromosomes Cancer 51:598
H Gu et al 2013 Clin Biochem 46:1668
M Wang, RJ Kaufman 2014 Nat Rev Cancer 14:581
VA Shchedrina et al 2010 Antioxid Redox Signal 12:839
Z Huang et al 2011 J Biol Chem 286:34830
JE Curran et al 2005 Nat Genet 37:1234
A Sutherland et al 2010 Genes Nutr 5:215
T Shibata et al 2009 BMC Gastroenterol 9:2
H Mao et al 2015 Int J Clin Exp Med 8:10993
A Martinez et al 2008 BMC Genomics 9:329
LH Canani et al 2005 J Clin Endocrinol Metab 90:3472
JM Dora et al 2010 Eur J Endocrinol 163:427
AA Estivalet et al 2011 Obesity (Silver Spring) 19:825
MA Nasr et al 2003 Cancer Therapy 1:293
KL Penney et al 2010 Cancer Prev Res (Phila) 3:604
E Jablonska et al 2008 Eur J Nutr 47:47
A Cebrian et al 2006 Cancer Res 66:1225
M Abe et al 2011 BJU Int 107:126
JY Choi et al 2007 Cancer Epidemiol Biomarkers Prev 16:1115
R Hansen et al 2005 Cancer Lett 229:85
RD Hansen et al 2009 Mutat Res 664:13
A Rosenberger et al 2008 BMC Cancer 8:60
T Hamanishi et al 2004 Diabetes 53:2455
L Huang et al 2013 PLoS ONE 8:e71411
OH Al-Taie et al 2004 Nutr Cancer 48:6
AJ Cox et al 2013 Acta Diabetol 50:391
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer Science+Business Media, LLC
About this chapter
Cite this chapter
Méplan, C., Hesketh, J. (2016). Functional Genomics of Selenoproteins and Se-responsive Pathways. In: Hatfield, D., Schweizer, U., Tsuji, P., Gladyshev, V. (eds) Selenium. Springer, Cham. https://doi.org/10.1007/978-3-319-41283-2_13
Download citation
DOI: https://doi.org/10.1007/978-3-319-41283-2_13
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-41281-8
Online ISBN: 978-3-319-41283-2
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)