Abstract
Salmonella species are causative organisms of salmonellosis, and the prevalence of multidrug-resistant Salmonella has increased dramatically. These multidrug-resistant isolates have been found in both humans and animals and thus pose a major public health concern. Drug resistance in Salmonella has been shown to be largely attributable to multiple target gene mutations and to active efflux by pumps. At least ten drug efflux system genes in the genome of this organism have been experimentally identified to date, and some efflux pump genes encoded in plasmids have been also identified. This chapter describes the drug resistance and virulence roles of efflux pumps and their regulation in Salmonella.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
Salmonella species exist all over the world and are responsible for causing acute gastroenteritis and typhoid/paratyphoid [1]. Salmonella enterica serovar Typhimurium is contagious in rodents, including mice, causing a systemic infectious disease, closely resembling human typhoid [2, 3]. In humans, it produces acute gastroenteritis and is a cause of food poisoning. Fluoroquinolones represent the drug of choice for the treatment of a wide range of human infectious diseases, and they were also introduced into veterinary medicine in Europe in the late 1980s through the early 1990s and the USA in 1995. Following their introduction, fluoroquinolone-resistant strains of Salmonella started to emerge [4]. Fluoroquinolone resistance in S. enterica serovar Typhimurium has been shown to be largely attributable to multiple target gene mutations and to active efflux by multidrug transporters [5, 6]. Also, the increasing prevalence of multidrug resistance has been found in Salmonella isolates from both humans and animals and thus poses an important public health concern [7, 8].
The genome sequences of Salmonella spp. indicate the presence of numerous efflux pump genes that encode transporters of various superfamilies and families [9, 10]. At least ten drug efflux pump genes in the genome of S. enterica serovar Typhimurium have been experimentally identified to date [11–15]. Some efflux pump genes encoded on plasmids have been also identified [16–18]. In addition to their roles in drug resistance, it was shown that the efflux pumps contribute to Salmonella virulence [13, 15, 19, 20]. Physiological functions of efflux pumps in Salmonella have been also reported with roles in metal resistance [21, 22], biofilm formation [23], colonization [11], adhesion, and cell invasion [19]. In this chapter, the roles of Salmonella efflux pumps in drug resistance and their physiological functions and regulation are described.
2 The AcrAB Efflux Pump in Salmonella
S. enterica serovar Typhimurium TnphoA mutants with increased susceptibility to biological and chemical detergents were reported [24], and it was found that one mutant LX1054 had a defect in a multidrug resistance pump AcrB [11]. Nikaido et al. [12] found that the previously reported drug-susceptible S. enterica serovar Typhimurium [25] carried a mutation in the acrAB operon. The mutant of acrAB exhibited increased susceptibility to a wide range of antimicrobial agents including antibiotics, bile salts, dyes, detergents, and disinfectants as shown in Table 10.1 [12, 13]. AcrA and AcrB in S. enterica serovar Typhimurium strain LT2 exhibit the amino acid identities of 92 and 95 % with those in Escherichia coli [13]. High-level fluoroquinolone resistance in S. enterica serovar Typhimurium phage type DT204 has been previously shown to be essentially due to both multiple target gene mutations and active efflux by the AcrAB–TolC efflux system [5, 6]. In other drug-resistant isolates of Salmonella, overexpression of acrB is also reported [29], and antimicrobial treatment of Salmonella results in the increased expression of acrB [30, 31]. A post-therapy isolate of S. enterica serovar Typhimurium (after treatment with fluoroquinolones and β-lactams) was found to carry a Gly288Asp substitution in AcrB [32]. This residue substitution is located in AcrB drug-binding pocket and significantly affects the structural and dynamic properties of AcrB, resulting in alternated substrate specificity (i.e., reduced susceptibility to fluoroquinolone but increased susceptibility to doxorubicin and minocycline) [32]. Low-level exposure of S. enterica serovar Typhimurium to a biocide, either a quaternary ammonium compound, an oxidative compound, or a halogenated tertiary amine compound, in the laboratory selected mutants that were cross-resistant to nalidixic acid, ciprofloxacin, chloramphenicol, tetracycline, and/or triclosan [33]. Among multiple mutations carried by these mutants, derepression of AcrAB–TolC expression was observed [33].
3 The Salmonella Drug Efflux Pumps Identified by Genomic Information
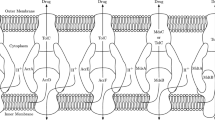
Genomic analyses revealed that Salmonella strains possess five putative RND efflux systems (http://www.membranetransport.org). Four of them, AcrAB (AcrA, membrane fusion protein; AcrB, RND transporter), AcrD, AcrEF (AcrE, membrane fusion protein; AcrF, RND transporter), and MdtABC (MdtA, membrane fusion protein; MdtB and MdtC, RND transporters), have homologs in E. coli with approximately ~90 % amino acid identity (Table 10.1) [13]. MdtB and MdtC are each an RND pump and usually function as one drug efflux system [34]. The last putative RND system is the Salmonella-specific MdsABC (MdsA, membrane fusion protein; MdsB, RND transporter; MdsC, outer membrane protein). In addition to the RND pumps, efflux systems belonging to the major facilitator superfamily (MFS) (EmrAB, MdfA, and SmvA), multidrug and toxic compound extrusion (MATE) family (MdtK), and the ATP-binding cassette (ABC) superfamily (MacAB) transporter families were also experimentally identified (Fig. 10.1) [13, 14, 35].
Diagrammatic representation of the structure of multidrug efflux pumps in Salmonella and their location on the membranes (Modified from Horiyama et al. [27])
The genes of acrAB, acrD, acrEF, mdtABC, mdsABC, emrAB, mdfA, mdtK, and macAB were cloned into the multicopy number plasmid, and their ability to confer drug resistance upon the Salmonella acrB mutant was investigated (Table 10.1) [13]. The plasmids carrying efflux operons or genes that confer multidrug resistance phenotypes against various antimicrobial compounds are shown in Table 10.1. It was also reported that the deletion mutant of the smvA gene showed increased susceptibility to a range of cytotoxic agents (Table 10.1) [14]. Overproduction of SmvA provided acriflavine resistance in the Salmonella acrB mutant (unpublished data). A recent study also showed that Salmonella EmrAB and AcrEF pumps may have additive effects with the major efflux system AcrAB in decreased susceptibility to triclosan [26]. Deletion of the tolC, acrB, or acrAB genes resulted in strains with increased susceptibility to various compounds, and the acrB, acrAB, and tolC mutant strains have overlapping substrate susceptibility profiles, which is in agreement with the notion that the encoded proteins interact as a tripartite efflux complex system. The tolC mutant was more susceptible to certain compounds including novobiocin, deoxycholate, and sodium dodecyl sulfate than the acrAB mutant [13, 36] – suggesting a functional role in other efflux systems. And a strain with nine drug exporters (acrAB, acrD, acrEF, mdtABC, mdsABC, emrAB, mdfA, mdtK, and macAB) deleted was shown to be more susceptible to novobiocin, deoxycholate, and sodium dodecyl sulfate, compared to the ∆acrAB mutant. On the other hand, strains deleted for the acrD, acrEF, mdtABC, mdsABC, emrAB, mdfA, mdtK, or macAB genes exhibited the same drug susceptibility as the wild-type strain [13]. These two lines of data suggest, similar to E. coli, a predominant if not overwhelming role of the AcrAB in the drug resistance phenotype. Furthermore, that other pump expression is minimal and/or their functions are masked by overlapping substrate repertoires with AcrAB. The expression levels of drug transporter genes under laboratory conditions were investigated by streaking out onto X-Gal (5-bromo-4-chloro-3-indolyl-β-D-galactopyranoside) LB agar plate strains in which the E. coli lacZY genes replaced the chromosomal copy of the drug efflux genes in Salmonella [13]. The tolC-lacZY and acrA-lacZY strains were blue on plates, whereas the acrD-lacZY, emrA-lacZY, mdfA-lacZY, mdtK-lacZY, and macA-lacZY strains were only faint blue. Thus, the AcrAB–TolC efflux system is expressed in the complex laboratory media, whereas the other efflux systems appear to require additional cues for expression [13].
TolC is required for the function of seven drug efflux systems AcrAB, AcrD, AcrEF, MdsAB, MdtABC, EmrAB, and MacAB in S. enterica serovar Typhimurium [27]. Therefore, plasmids carrying the acrAB, acrD, acrEF, mdsAB, mdtABC, emrAB, or macAB genes do not confer resistance to the tolC mutant, whereas they conferred drug resistance in the acrB mutant. Plasmids carrying mdsABC, mdfA, or mdtK provide resistance to the tolC mutant, indicating that these three efflux systems function without TolC. The crystal structure of TolC (i.e., ST50) from Salmonella Typhi was recently reported, showing the structural basis for TolC role in multidrug efflux pumps across the outer membrane [37]. The Salmonella-specific drug efflux system mdsABC operon codes for a putative outer membrane protein – MdsC – which is in contrast to the other operons coding for RND-type drug transporter genes. In E. coli, most operons coding for RND-type drug transporter homologs lack genes for outer membrane proteins [38] because they rely on TolC as their outer membrane component [39–42]. Overexpression of both the mdsABC and mdsAB genes produced drug resistance in the ∆acrB mdsABC strain. On the other hand, overexpression of mdsABC, but not mdsAB, resulted in drug resistance to the ∆acrB tolC mdsABC strain. These findings indicate that the drug resistance phenotype conferred by the MdsAB system is dependent on the presence of either the MdsC or TolC proteins and that the MdsAB system can function with both TolC and MdsC outer membrane components [13, 27].
Except for the acrD gene, all RND efflux system genes also code for a membrane fusion protein in the same operon. The overproduction of AcrD yielded multidrug resistance in the ΔacrB mutant against β-lactam antibiotics and other agents (Table 10.1). It was revealed that AcrD requires AcrA and TolC to function (Fig. 10.1) [27, 43]. One possibility for AcrD utilizing AcrA, coded in a different operon, is that AcrD may form a complex with AcrA and TolC when mutations occur in AcrB and compensate for the lost function of AcrAB–TolC multidrug efflux system. Another possibility is that AcrA contributes to different biological functions by forming complexes with two different RND pumps, AcrB and AcrD. Such a functional network of multidrug efflux pumps may contribute to bacterial adaptation to various environmental conditions [43].
4 Plasmid-Mediated Fluoroquinolone Efflux Pumps
In addition to the efflux systems encoded in the Salmonella genome, plasmid-mediated fluoroquinolone efflux pumps have been identified. The MFS efflux pump QepA was originally identified in Escherichia coli clinical isolate [44]. Resistance levels against ciprofloxacin, enrofloxacin, and norfloxacin were significantly elevated in E. coli transformants harboring qepA under AcrB–TolC-deficient conditions. The intracellular accumulation of norfloxacin was decreased in a qepA-expressing E. coli transformant [44]. In Salmonella, qepA was first detected in the clinical isolates obtained in the hospital clinic in Spain [45]. Subsequently, qepA was detected in several quinolone-resistant Salmonella spp. clinical isolates [46, 47].
Plasmid-encoded multidrug efflux genes oqxAB were also identified in Salmonella [18, 48–51]. The quinoxaline-di-N-oxide olaquindox has been a growth enhancer in pigs. Its antimicrobial activity is due to inhibition of DNA synthesis [52]. The oqxAB genes were originally identified from a conjugative plasmid isolated from E. coli [53]. OqxA, a membrane fusion protein, and OqxB, an inner membrane protein, are homologous to several RND family efflux systems from different species. Plasmids containing the oqxAB genes yielded high resistance to olaquindox in E. coli. The oqxAB-encoded pump also conferred high resistance to chloramphenicol [53]. H+-dependent ethidium efflux abilities of OqxAB were also confirmed in E. coli [53]. A derivative of the plasmid encoding OqxAB was readily transferred to enterobacterial pathogens and transconjugants showed reduced susceptibility to chloramphenicol, ciprofloxacin, and olaquindox [54]. OqxAB were found in human clinical isolates on a plasmid in E. coli and on the chromosome of Klebsiella pneumoniae. IS26-like sequences flanked the plasmid-mediated oqxAB genes, suggesting that they had been mobilized as part of a composite transposon [55]. After the first detection of oqxAB in Salmonella spp. isolated from food [47], the genes were identified in many Salmonella isolates which exhibited resistance to fluoroquinolones [48–51, 56, 57].
5 Virulence Roles of Salmonella Drug Efflux Pumps
Drug efflux systems are evolutionarily ancient and are found throughout the three domains of life [58, 59]. These systems are fundamental to the bacterial physiology and some have roles other than conferring resistance to antimicrobials. Recognizing that the AcrAB–TolC system serves as an important antimicrobial resistance determinant [11, 12], it was also reported that this efflux system is required for Salmonella resistance to bile salts [11, 60] which are found exclusively associated with higher vertebrates. It was shown that the acrB mutant of S. enterica serovar Typhimurium exhibited a reduced capacity to colonize the intestinal tract, and this suggests that AcrAB–TolC efflux system play an important role in mouse intestinal colonization [11]. It was also reported that the deletion of the macAB genes attenuated Salmonella virulence, and a strain lacking all drug efflux systems was avirulent when mice were inoculated by the oral route [13]. These results indicate that drug efflux genes are required for Salmonella’s ability to cause a lethal infection in mice. Utilizing similar approaches, Buckley et al. [19] studied the role of efflux systems on virulence of S. enterica serovar Typhimurium using efflux-defective mutants in a chicken model and found that mutants deficient in either acrB or tolC genes colonized poorly and did not persist in the avian gut, indicating that AcrAB–TolC system is essential for the colonization of S. enterica serovar Typhimurium in chickens. Experiments using BALB/c mice by the oral route with isogenic strains harboring deletions in efflux genes showed that the mutation in tolC of S. enterica serovar Typhimurium attenuated virulence [13], as reported for an S. enterica serovar Enteritidis tolC mutant [61]. Inactivation of the MarA or RamA activator (which upregulates AcrAB–TolC expression; see Sect. 10.7) reduced both the invasion and survival ability of Salmonella choleraesuis in the host cells and virulence in mice [62].
Salmonella MacAB pump plays a role in the detoxification of reactive oxygen species, compounds that salmonellae are exposed to at various stages of infection [63]. The macAB operon is induced upon exposure to hydrogen peroxide and is critical for survival of S. enterica serovar Typhimurium in the presence of oxidative stress. Furthermore, macAB is required for intracellular replication inside murine macrophages but is not required for survival in reactive oxygen species-deficient macrophages [63]. Bogomolnaya et al. [63] suggested the presence of a soluble anti-peroxide compound secreted by Salmonella cells through a MacAB-dependent mechanism. In E. coli, MacAB is involved in the secretion of heat-stable enterotoxin II [64], and MacA binds lipopolysaccharide core specifically with high affinity [65]. Also, it was recently reported that protoporphyrin is exported by MacAB–TolC in E. coli [66]. Because high protoporphyrin levels result in production of reactive oxygen species [67], Turlin et al. [66] proposed that MacAB is involved in the efflux of intracellular protoporphyrin which decreases reactive oxygen species formation in the bacterial cytoplasm, providing a possible explanation for the role of MacAB in Salmonella pathogenicity.
6 Physiological Functions of Salmonella Drug Efflux Pumps
There are several reports about the physiological functions of Salmonella drug efflux systems. The BaeSR two-component signal transduction system activates the acrD and mdtABC expression in response to indole, copper, and zinc. BaeSR, AcrD, and MdtABC contribute to copper and zinc resistance in Salmonella [21]; andiron and sodium tungstate are inducers of the BaeR regulon suggesting MdtA, AcrD, and AcrB exist for the waste disposal of tungstate from the cell [68]. Additionally, the MdsABC pump (also called GesABC) is required for gold resistance and the mdsABC operon is controlled by GolS which is a MerR-like sensor and highly selective for Au ions [22]. In contrast to heavy metal-specific CusCBA RND pump of E. coli, MdsABC, accommodates a large number of substrates including many antibiotics (Table 10.1) [28].
Recent studies have showed that defects in efflux activity impair biofilm formation. In S. enterica serovar Typhimurium, deletion of any efflux pump or chemical inhibition of the efflux activity results in compromised ability of Salmonella to form biofilm [23]. The defect of biofilm formation in efflux mutants resulted from transcriptional repression of curli biosynthesis genes and consequently inhibition of its production, but was not associated with altered aggregative ability or export of any biofilm-promoting factor [69] (also see Chap. 25).
7 Regulation of Salmonella Drug Efflux Pumps
The key to understanding how bacteria utilize multidrug efflux pumps lies in the regulation of pump expression. The data currently available show that multidrug efflux pumps are often expressed under precise and elaborate transcriptional control. For example, expression of macAB is controlled by the PhoPQ system, the master regulator for the virulence of Salmonella (Table 10.2) [13]. A sequence resembling the PhoP binding box exists in the upstream of the macAB operon [78]. DNase I footprinting analysis with the purified PhoP protein showed protection of the region upstream of the macA open reading frame [13], indicating that the PhoPQ two-component signal transduction system controls macAB directly. Analysis of mRNA levels of drug efflux genes revealed that the expression of macAB is induced when the organism infects macrophages [15]. A recent study also showed that hydrogen peroxide induces expression of macAB [63], supporting the induction of macAB inside macrophages and the existence of additional regulator to control the macAB genes responsive to hydrogen peroxide.
Moreover, positive regulation of the multidrug efflux pump mdtABC and acrD genes by the BaeSR two-component signal transduction system was found (Table 10.2) [21]. In addition to the roles of MdtABC, AcrD, and BaeSR in multidrug resistance, they contribute to copper and zinc resistance in Salmonella as described above. Both copper and zinc are essential for organisms but can be toxic at high levels, and microorganisms express diverse resistance mechanisms. The expression of mdtABC and acrD is induced by copper or zinc, and BaeSR is involved in this induction (Table 10.2). This finding indicates that the MdtABC and AcrD efflux systems have physiological roles in metal homeostasis beyond multidrug resistance [21]. It was also reported that GolS controls MdsABC in response to Au ions [22].
Mutations in acrR contribute to overexpression of acrAB and increases resistance to multiple drugs in Salmonella [73]. The histone-like protein (H-NS) modulates multidrug resistance through repression of the acrEF genes [77]. Eaves et al. [74] suggested that acrB, acrF, and acrD are coordinately regulated and that their expression is also influenced by the expression of the transcriptional activators marA and soxS. Nikaido et al. [75] found that acrAB induction in response to methyl viologen is dependent on SoxS. Indole, bile salts, and an E. coli-conditioned medium were also able to induce the expression of acrAB in Salmonella. The acrAB induction by these three signal sources is completely dependent on the Salmonella-specific regulator RamA, indicating that RamA plays a major role in inducing acrAB (Table 10.2) [70]. RamA belongs to the AraC transcriptional activator family, and this gene appears to be specific for Salmonella serovars and is absent in many other Gram-negative microorganisms; notable exceptions are Klebsiella pneumoniae and Enterobacter species [79–81]. The AcrAB induction pathway in Salmonella is different from that in E. coli. Bile induces AcrAB in both Salmonella and E. coli. In E. coli, the transcriptional factor Rob plays a major role in inducing acrAB expression in response to bile [82]. However, bile induction of acrAB in Salmonella is dependent on RamA, not Rob. Other regulators, including MarA, SoxS, SdiA, and AcrR, are not involved in AcrAB induction by indole and bile [70]. These facts suggest that RamA is the major regulator of Salmonella acrAB and may mask the contributions of any other acrAB regulators.
Abouzeed et al. [83] demonstrated that the inactivation of the ramR gene upstream of ramA resulted in an increased expression of ramA and the AcrAB efflux pump, indicating that RamR is a local repressor of ramA. Inactivation of marR, marA, soxR, and soxS did not affect the susceptibilities of the S. enterica serovar Typhimurium strain LT2, whereas the disruption of ramR resulted in a multidrug resistance phenotype with this strain. In E. coli, multiple regulators, including MarA, Rob, SoxS, and SdiA, work together in controlling acrAB expression in response to acrAB inducers. This may be related to the lack of RamA in E. coli. Indeed, overproduction of RamA has induced the drug resistance level of E. coli [84, 85]. There may also be different induction mechanisms for acrAB via the RamA regulator. Indole was shown to induce ramA expression, and such increased expression of ramA can induce acrAB, whereas bile binds to RamA. This is reminiscent of the binding of bile to the Rob protein involved in regulation of acrAB in E. coli [82]. It seems that RamA can be converted from a low-activity state to a high-activity state in response to bile. More recently, Baucheron et al. [71] also identified a different induction mechanism of acrAB in response to bile whereby the bile-mediated activation of the acrAB and tolC multidrug efflux genes occurs via transcriptional derepression of the ramA activator gene, likely via the RamR repressor protein controlling expression of ramA. Indole and bile salts are found in various internal human environments, especially in the intestine [86, 87]. Indole is produced by many enteric bacterial species [87], and bile is often present in high concentrations in the intestinal tract [86]. Therefore, RamA may be required for Salmonella to detect environmental signals and for subsequent induction of the AcrAB–TolC system, resulting in excretion of toxic compounds into the surrounding environment in the above examples, the intestine. A recent study showed heterogeneity in ramRA mutations and its differential impact on expression of regulator genes ramA, marA, soxS, and acrR and efflux component genes acrB, acrF, emrB, and tolC, revealing deletions that affected RamR-binding site exhibiting a major impact on the ramA transcript level and the multidrug resistance phenotype [88].
8 Structure of Multidrug Efflux Pump Regulator RamR with Multiple Drugs
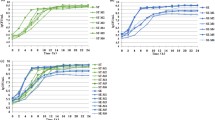
As described above, RamR and RamA are important regulators for AcrAB–TolC in Salmonella. From the structural and biochemical analysis of RamR, a multidrug recognition mechanism of RamR occurs, whereby the DNA-binding activity is controlled by multiple drugs in order to induce ramA expression [72]. Yamasaki et al. [72] identified five substrates of the RamR protein, including berberine, crystal violet, dequalinium, ethidium bromide, and rhodamine 6G (Fig. 10.2). Similar approaches in crystallizing the TetR family regulators with multiple drugs have been also reported in QacR [89], TtgR [90], and CmeR [91]. The molecular weight of RamR in solution was calculated to be 36 kDa using gel filtration chromatography, which was conducted during the purification of the RamR protein. Dissolved RamR was found to exist in the dimer form in solution, and the molecular weight of the RamR monomer was 21 kDa [72]. The structure of RamR was initially determined at a resolution of 2.6 Å by multiple wavelength anomalous dispersion using selenomethionine modification. Subsequently, the RamR structure was determined at 2.1 Å by molecular replacement. Approximate overall dimensions of the RamR dimer were 58 × 47 × 44 Å3. RamR is composed of nine α-helices, and the three-helix bundle structures formed at the N-terminus maintain a helix-turn-helix motif conserved in DNA-binding sites. The structure of the RamR DNA-binding site is similar to that of other TetR family regulators. By the surface plasmon resonance analysis, it was found that five compounds, berberine, crystal violet, dequalinium, ethidium bromide, and rhodamine 6G, bind to the RamR protein. In contrast, tetracycline did not show any indication of binding to RamR. Using a ramA reporter plasmid, a ß-galactosidase assay showed the enhanced promoter activity of ramA when bacterial cells were treated with berberine, crystal violet, dequalinium, ethidium bromide, or rhodamine 6G. The crystal structures of RamR in complex with berberine, crystal violet, dequalinium, ethidium bromide, and rhodamine 6G were determined at a resolution of 2.4, 2.2, 2.6, 1.6, and 2.5 Å, respectively [72]. The structure reveals that RamR binds two molecules of berberine, ethidium bromide, or rhodamine 6G per dimer. And RamR binds one crystal violet or dequalinium molecule per dimer. It was originally reported that all the ligands bind to QacR with a 1:2 stoichiometry (one ligand per QacR dimer) [89], while either 1:2 or 1:1 stoichiometry has been observed for RamR. Similar observations were reported in TtgR [90]. The orientation of all agents is parallel with the Phe155 of RamR, suggesting that all these drugs bind with RamR through π–π stacking interactions. In contrast to the common interaction of all of these drugs with Phe155, each individual drug was also found to interact with a different set of amino acid residues other than Phe155. The interaction of different sets of amino acid residues with each drug indicates that multiple drugs are recognized by the multisite binding of RamR [72]. Comparison of the liganded structures with an unliganded RamR structure reveals that drug binding triggers an expansion of the distance between the N-termini of the helix-turn-helix motifs in the RamR dimer. This expansion occurred as a result of the binding of all of the drugs examined. By the electrophoretic mobility shift assays and surface plasmon resonance experiments, RamR substrates interact with their recognition sites to reduce the DNA-binding affinity of RamR, resulting in the induction of ramA [72]. Because RamA has also been reported to negatively influence virulence in S. enterica serovar Typhimurium by downregulating expression of the Salmonella pathogenicity island 1 [92], determining the crystal structure of RamR is the first step in understanding the structural basis for the function of the regulatory proteins that control both drug resistance and virulence in pathogens. This effort extended our knowledge of transcriptional regulation mediated by RamR, a regulator of multidrug resistance in several enterobacterial pathogens.
Regulatory cascade and structure of RamR. (a) Model for gene regulation by RamR. RamR represses expression of the ramA gene, which encodes the activator protein for the acrAB efflux pump genes. RamR binds to the intergenic region between the ramR and ramA genes, and RamA binds to the upstream region of acrAB. (b) Crystal structure of the RamR dimer. Each monomer is colored as follows: the α-helices are represented in blue (α1), marine (α2), sky blue (α3), cyan (α4), green (α5), limon (α6), yellow (α7a), deep olive (α7b), orange (α8a), brown (α8b), and red (α9). (c) Multidrug recognition by RamR. Substrate binding site of RamR with a bound molecule berberine, crystal violet, dequalinium, ethidium bromide, or rhodamine 6G. Key residues are shown, including residue Phe155, which is involved in π–π stacking interactions with drugs. Carbon atoms of drugs and RamR are shown in magenta and green, respectively. Nitrogen, oxygen, and sulfur atoms are shown in blue, red, and yellow, respectively (Figure is modified from Yamasaki et al. [72])
9 Concluding Remarks
Post-genomic research has demonstrated that bacteria possess a large number of drug efflux system genes. As described in this chapter, at least ten drug efflux systems in the genome of S. enterica have been experimentally identified to date. Under normal growth conditions, most of drug efflux pumps are thought to be weakly expressed [13]. Increased expression of such efflux systems is possible when mutations occur in their regulatory factors. In fact, various types of mutations in ramR and the ramR–ramA intergenic region were identified in multidrug-resistant strains of S. Typhimurium, other S. enterica serovars, and K. pneumoniae, which result in increased expression of ramA and an increase in efflux-mediated multidrug resistance [83, 93, 94]. Also, it was reported that overexpression of the multidrug efflux operon acrEF occurs by insertional activation with IS1 or IS10 elements in S. enterica serovar Typhimurium DT204 acrB mutants selected with fluoroquinolones [76]. A mutation in acrR, the local repressor of acrAB, was found for two ciprofloxacin-resistant selected mutants of S. enterica serovar Typhimurium [73]. In addition to these mutations, the structural and biochemical analysis showed that toxic compounds bind to RamR resulting in the increased efflux activity of Salmonella to protect this organism against the compounds [72].
Association of resistance mechanism with two-component signal transduction systems, which control the expression of drug efflux pumps, has also been identified in Salmonella. These findings suggest that the expression of efflux systems is transiently induced through some types of stimulation. In fact, this induction occurs as a result of various environmental stressors, such as low pH, osmotic changes, metals, and oxidative stress. The mechanism by which efflux pumps are expressed in response to the environment suggests that they might be expressed in the growth environments of bacteria such as at infection sites. It is reasonable to assume that efflux systems are induced inside hosts because these contribute not only to drug resistance but also to bacterial virulence. Therefore, it is necessary to identify the regulatory network of multidrug transporters in order to understand their physiological functions. Moreover, determining the physiological substrate of efflux systems is an important area of study, which will contribute to the understanding of the role of drug efflux systems in virulence.
The mechanism by which drug efflux pumps contribute to bacterial virulence has three features. Firstly, the efflux system has the capacity to transport substrates necessary to establish virulence, for example, toxins. Secondly, the efflux system is able to export antibacterial substances present in the host (such as bile acid and antimicrobial peptides) in order to protect the bacteria from the host environment. Thirdly, it can transport factors contributing to bacterial homeostasis or promoting bacterial regulatory functions within the host (such as autoinducers). Currently, several research groups and pharmaceutical companies are conducting research to develop drug efflux pump inhibitors. As efflux systems contribute to multidrug resistance and bacterial virulence, efflux systems are an attractive target for the development of new drugs. If an effective inhibitor is found, it could play a role in the development of new therapies that could conquer bacterial multidrug resistance and virulence.
References
Scherer CA, Miller SI (2001) Molecular pathogenesis of Salmonella. In: Groisman EA (ed) Principles of bacterial pathogenesis. Academic Press, New York, pp 266–333
Finlay BB, Falkow S (1988) Virulence factors associated with Salmonella species. Microbiol Sci 5:324–328
Finlay BB, Falkow S (1989) Salmonella as an intracellular parasite. Mol Microbiol 3:1833–1841. doi:10.1111/j.1365-2958.1989.tb00170.x
Bager F, Helmuth R (2001) Epidemiology of resistance to quinolones in Salmonella. Vet Res 32:285–290. doi:10.1051/vetres:2001125
Baucheron S, Imberechts H, Chaslus-Dancla E, Cloeckaert A (2002) The AcrB multidrug transporter plays a major role in high-level fluoroquinolone resistance in Salmonella enterica serovar Typhimurium phage type DT204. Microb Drug Resist 8:281–289. doi:10.1089/10766290260469543
Baucheron S, Chaslus-Dancla E, Cloeckaert A (2004) Role of TolC and parC mutation in high-level fluoroquinolone resistance in Salmonella enterica serotype Typhimurium DT204. J Antimicrob Chemother 53:657–659. doi:10.1093/jac/dkh122
Cloeckaert A, Chaslus-Dancla E (2001) Mechanisms of quinolone resistance in Salmonella. Vet Res 32:291–300. doi:10.1051/vetres:2001105
Piddock LJ (2002) Fluoroquinolone resistance in Salmonella serovars isolated from humans and food animals. FEMS Microbiol Rev 26:3–16. doi:10.1111/j.1574-6976.2002.tb00596.x
Nishino K (2005) Bacterial multidrug exporters: insights into acquisition of multidrug resistance. Science. Online publication http://www.sciencemag.org/site/feature/data/prizes/ge/2004/nishino.xhtml
Ren Q, Paulsen IT (2007) Large-scale comparative genomic analyses of cytoplasmic membrane transport systems in prokaryotes. J Mol Microbiol Biotechnol 12:165–179. doi:10.1159/000099639
Lacroix FJ, Cloeckaert A, Grepinet O, Pinault C, Popoff MY, Waxin H, Pardon P (1996) Salmonella typhimurium acrB-like gene: identification and role in resistance to biliary salts and detergents and in murine infection. FEMS Microbiol Lett 135:161–167. doi:10.1111/j.1574-6968.1996.tb07983.x
Nikaido H, Basina M, Nguyen V, Rosenberg EY (1998) Multidrug efflux pump AcrAB of Salmonella typhimurium excretes only those β-lactam antibiotics containing lipophilic side chains. J Bacteriol 180:4686–4692
Nishino K, Latifi T, Groisman EA (2006) Virulence and drug resistance roles of multidrug efflux systems of Salmonella enterica serovar Typhimurium. Mol Microbiol 59:126–141. doi:10.1111/j.1365-2958.2005.04940.x
Villagra NA, Hidalgo AA, Santiviago CA, Saavedra CP, Mora GC (2008) SmvA, and not AcrB, is the major efflux pump for acriflavine and related compounds in Salmonella enterica serovar Typhimurium. J Antimicrob Chemother 62:1273–1276. doi:10.1093/jac/dkn407
Nishino K, Nikaido E, Yamaguchi A (2009) Regulation and physiological function of multidrug efflux pumps in Escherichia coli and Salmonella. Biochim Biophys Acta 1794:834–843. doi:10.1016/j.bbapap.2009.02.002
Nordmann P, Poirel L (2005) Emergence of plasmid-mediated resistance to quinolones in Enterobacteriaceae. J Antimicrob Chemother 56:463–469. doi:10.1093/jac/dki245
Robicsek A, Jacoby GA, Hooper DC (2006) The worldwide emergence of plasmid-mediated quinolone resistance. Lancet Infect Dis 6:629. doi:10.1016/S1473-3099(06)70599-0
Wong MH, Chen S (2013) First detection of oqxAB in Salmonella spp. isolated from food. Antimicrob Agents Chemother 57:658–660. doi:10.1128/AAC.01144-12
Buckley AM, Webber MA, Cooles S, Randall LP, La Ragione RM, Woodward MJ, Piddock LJ (2006) The AcrAB-TolC efflux system of Salmonella enterica serovar Typhimurium plays a role in pathogenesis. Cell Microbiol 8:847–856. doi:10.1111/j.1462-5822.2005.00671.x
Piddock LJ (2006) Multidrug-resistance efflux pumps – not just for resistance. Nat Rev Microbiol 4:629–636. doi:10.1038/nrmicro1464
Nishino K, Nikaido E, Yamaguchi A (2007) Regulation of multidrug efflux systems involved in multidrug and metal resistance of Salmonella enterica serovar Typhimurium. J Bacteriol 189:9066–9075. doi:10.1128/JB.01045-07
Pontel LB, Audero ME, Espariz M, Checa SK, Soncini FC (2007) GolS controls the response to gold by the hierarchical induction of Salmonella-specific genes that include a CBA efflux-coding operon. Mol Microbiol 66:814–825. doi:10.1111/j.1365-2958.2007.05963.x
Baugh S, Ekanayaka AS, Piddock LJ, Webber MA (2012) Loss of or inhibition of all multidrug resistance efflux pumps of Salmonella enterica serovar Typhimurium results in impaired ability to form a biofilm. J Antimicrob Chemother 67:2409–2417. doi:10.1093/jac/dks228
Lacroix FJ, Avoyne C, Pinault C, Popoff MY, Pardon P (1995) Salmonella typhimurium TnphoA mutants with increased sensitivity to biological and chemical detergents. Res Microbiol 146:659–670. doi:10.1016/0923-2508(96)81063-1
Sukupolvi S, Vaara M, Helander IM, Viljanen P, Makela PH (1984) New Salmonella typhimurium mutants with altered outer membrane permeability. J Bacteriol 159:704–712
Rensch U, Nishino K, Klein G, Kehrenberg C (2014) Salmonella enterica serovar Typhimurium multidrug efflux pumps EmrAB and AcrEF support the major efflux system AcrAB in decreased susceptibility to triclosan. Int J Antimicrob Agents 44:179–180. doi:10.1016/j.ijantimicag.2014.04.015
Horiyama T, Yamaguchi A, Nishino K (2010) TolC dependency of multidrug efflux systems in Salmonella enterica serovar Typhimurium. J Antimicrob Chemother 65:1372–1376. doi:10.1093/jac/dkq160
Conroy O, Kim EH, McEvoy MM, Rensing C (2010) Differing ability to transport nonmetal substrates by two RND-type metal exporters. FEMS Microbiol Lett 308:115–122. doi:10.1111/j.1574-6968.2010.02006.x
Webber M, Buckley AM, Randall LP, Woodward MJ, Piddock LJ (2006) Overexpression of marA, soxS and acrB in veterinary isolates of Salmonella enterica rarely correlates with cyclohexane tolerance. J Antimicrob Chemother 57:673–679. doi:10.1093/jac/dkl025
Usui M, Nagai H, Hiki M, Tamura Y, Asai T (2013) Effect of antimicrobial exposure on AcrAB expression in Salmonella enterica subspecies enterica serovar Choleraesuis. Front Microbiol 4:53. doi:10.3389/fmicb.2013.00053
Ferrari RG, Galiana A, Cremades R, Rodriguez JC, Magnani M, Tognim MC, Oliveira TC, Royo G (2013) Expression of the marA, soxS, acrB and ramA genes related to the AcrAB/TolC efflux pump in Salmonella enterica strains with and without quinolone resistance-determining regions gyrA gene mutations. Braz J Infect Dis 17:125–130. doi:10.1016/j.bjid.2012.09.011
Blair JM, Bavro VN, Ricci V, Modi N, Cacciotto P, Kleinekathfer U, Ruggerone P, Vargiu AV et al (2015) AcrB drug-binding pocket substitution confers clinically relevant resistance and altered substrate specificity. Proc Natl Acad Sci U S A 112:3511–3516. doi:10.1073/pnas.1419939112
Webber MA, Whitehead RN, Mount M, Loman NJ, Pallen MJ, Piddock LJ (2015) Parallel evolutionary pathways to antibiotic resistance selected by biocide exposure. J Antimicrob Chemother 70:2241–2248. doi:10.1093/jac/dkv109
Nagakubo S, Nishino K, Hirata T, Yamaguchi A (2002) The putative response regulator BaeR stimulates multidrug resistance of Escherichia coli via a novel multidrug exporter system, MdtABC. J Bacteriol 184:4161–4167. doi:10.1128/JB.184.15.4161-4167.2002
Santiviago CA, Fuentes JA, Bueno SM, Trombert AN, Hildago AA, Socias LT, Youderian P, Mora GC (2002) The Salmonella enterica sv. Typhimurium smvA, yddG and ompD (porin) genes are required for the efficient efflux of methyl viologen. Mol Microbiol 46:687–698. doi:10.1046/j.1365-2958.2002.03204.x
Baucheron S, Mouline C, Praud K, Chaslus-Dancla E, Cloeckaert A (2005) TolC but not AcrB is essential for multidrug-resistant Salmonella enterica serotype Typhimurium colonization of chicks. J Antimicrob Chemother 55:707–712. doi:10.1093/jac/dki091
Guan HH, Yoshimura M, Chuankhayan P, Lin CC, Chen NC, Yang MC, Ismail A, Fun HK et al (2015) Crystal structure of an antigenic outer-membrane protein from Salmonella Typhi suggests a potential antigenic loop and an efflux mechanism. Sci Rep 5:16441. doi:10.1038/srep16441
Nishino K, Yamaguchi A (2001) Analysis of a complete library of putative drug transporter genes in Escherichia coli. J Bacteriol 183:5803–5812. doi:10.1128/JB.183.20.5803-5812.2001
Fralick JA (1996) Evidence that TolC is required for functioning of the Mar/AcrAB efflux pump of Escherichia coli. J Bacteriol 178:5803–5805
Nishino K, Yamaguchi A (2002) EvgA of the two-component signal transduction system modulates production of the yhiUV multidrug transporter in Escherichia coli. J Bacteriol 184:2319–2323. doi:10.1128/JB.184.8.2319-2323.2002
Nishino K, Yamada J, Hirakawa H, Hirata T, Yamaguchi A (2003) Roles of TolC-dependent multidrug transporters of Escherichia coli in resistance to β-lactams. Antimicrob Agents Chemother 47:3030–3033. doi:10.1128/AAC.47.9.3030-3033.2003
Nishino K, Yamaguchi A (2004) Role of histone-like protein H-NS in multidrug resistance of Escherichia coli. J Bacteriol 186:1423–1429. doi:10.1128/JB.186.5.1423-1429.2004
Yamasaki S, Nagasawa S, Hayashi-Nishino M, Yamaguchi A, Nishino K (2011) AcrA dependency of the AcrD efflux pump in Salmonella enterica serovar Typhimurium. J Antibiot (Tokyo) 64:433–437. doi:10.1038/ja.2011.28
Yamane K, Wachino J, Suzuki S, Kimura K, Shibata N, Kato H, Shibayama K, Konda T et al (2007) New plasmid-mediated fluoroquinolone efflux pump, QepA, found in an Escherichia coli clinical isolate. Antimicrob Agents Chemother 51:3354–3360. doi:10.1128/AAC.00339-07
Lunn AD, Fabrega A, Sanchez-Cespedes J, Vila J (2010) Prevalence of mechanisms decreasing quinolone-susceptibility among Salmonella spp. clinical isolates. Int Microbiol 13:15–20. doi:10.2436/20.1501.01.107
Al-Gallas N, Abbassi MS, Gharbi B, Manai M, Ben Fayala MN, Bichihi R, Al-Gallas A, Ben Aissa R (2013) Occurrence of plasmid-mediated quinolone resistance determinants and rmtB gene in Salmonella enterica serovar Enteritidis and Typhimurium isolated from food-animal products in Tunisia. Foodborne Pathog Dis 10:813–819. doi:10.1089/fpd.2012.1466
Colobatiu L, Tabaran A, Flonta M, Oniga O, Mirel S, Mihaiu M (2015) First description of plasmid-mediated quinolone resistance determinants and β-lactamase encoding genes in non-typhoidal Salmonella isolated from humans, one companion animal and food in Romania. Gut Pathog 7:16. doi:10.1186/s13099-015-0063-3
Li L, Liao X, Yang Y, Sun J, Li L, Liu B, Yang S, Ma J et al (2013) Spread of oqxAB in Salmonella enterica serotype Typhimurium predominantly by IncHI2 plasmids. J Antimicrob Chemother 68:2263–2268. doi:10.1093/jac/dkt209
Li L, Liao XP, Liu ZZ, Huang T, Li X, Sun J, Liu BT, Zhang Q et al (2014) Co-spread of oqxAB and bla CTX-M-9G in non-Typhi Salmonella enterica isolates mediated by ST2-IncHI2 plasmids. Int J Antimicrob Agents 44:263–268. doi:10.1016/j.ijantimicag.2014.05.014
Wong MH, Yan M, Chan EW, Biao K, Chen S (2014) Emergence of clinical Salmonella enterica serovar Typhimurium isolates with concurrent resistance to ciprofloxacin, ceftriaxone, and azithromycin. Antimicrob Agents Chemother 58:3752–3756. doi:10.1128/AAC.02770-13
Kao CY, Chen CA, Liu YF, Wu HM, Chiou CS, Yan JJ, Wu JJ (2015) Molecular characterization of antimicrobial susceptibility of Salmonella isolates: first identification of a plasmid carrying qnrD or oqxAB in Taiwan. J Microbiol Immunol Infect. doi:10.1016/j.jmii.2015.03.004
Suter W, Rosselet A, Knusel F (1978) Mode of action of quindoxin and substituted quinoxaline-di-N-oxides on Escherichia coli. Antimicrob Agents Chemother 13:770–783. doi:10.1128/AAC.13.5.770
Hansen LH, Johannesen E, Burmolle M, Sørensen AH, Sørensen SJ (2004) Plasmid-encoded multidrug efflux pump conferring resistance to olaquindox in Escherichia coli. Antimicrob Agents Chemother 48:3332–3337. doi:10.1128/AAC.48.9.3332-3337.2004
Hansen LH, Jensen LB, Sørensen HI, Sørensen SJ (2007) Substrate specificity of the OqxAB multidrug resistance pump in Escherichia coli and selected enteric bacteria. J Antimicrob Chemother 60:145–147. doi:10.1093/jac/dkm167
Kim HB, Wang M, Park CH, Kim EC, Jacoby GA, Hooper DC (2009) oqxAB encoding a multidrug efflux pump in human clinical isolates of Enterobacteriaceae. Antimicrob Agents Chemother 53:3582–3584. doi:10.1128/AAC.01574-08
Wong MH, Chan EW, Liu LZ, Chen S (2014) PMQR genes oqxAB and aac(6′)Ib-cr accelerate the development of fluoroquinolone resistance in Salmonella typhimurium. Front Microbiol 5:521. doi:10.3389/fmicb.2014.00521
Lin D, Chen K, Wai-Chi Chan E, Chen S (2015) Increasing prevalence of ciprofloxacin-resistant food-borne Salmonella strains harboring multiple PMQR elements but not target gene mutations. Sci Rep 5:14754. doi:10.1038/srep14754
Saier MH Jr, Paulsen IT, Sliwinski MK, Pao SS, Skurray RA, Nikaido H (1998) Evolutionary origins of multidrug and drug-specific efflux pumps in bacteria. FASEB J 12:265–274
Blair JM, Richmond GE, Piddock LJ (2014) Multidrug efflux pumps in Gram-negative bacteria and their role in antibiotic resistance. Future Microbiol 9:1165–1177. doi:10.2217/fmb.14.66
Prouty AM, Brodsky IE, Falkow S, Gunn JS (2004) Bile-salt-mediated induction of antimicrobial and bile resistance in Salmonella Typhimurium. Microbiology 150:775–783. doi:10.1099/mic.0.26769-0
Stone BJ, Miller VL (1995) Salmonella enteritidis has a homologue of tolC that is required for virulence in BALB/c mice. Mol Microbiol 17:701–712. doi:10.1111/j.1365-2958.1995.mmi_17040701.x
Lee JJ, Hsuan SL, Kuo CJ, Wu YC, Chen TH (2015) MarA and ramA regulate virulence in Salmonella enterica serovar Choleraesuis. Vet Microbiol 181:323–327. doi:10.1016/j.vetmic.2015.09.006
Bogomolnaya LM, Andrews KD, Talamantes M, Maple A, Ragoza Y, Vazquez-Torres A, Andrews-Polymenis H (2013) The ABC-type efflux pump MacAB protects Salmonella enterica serovar Typhimurium from oxidative stress. mBio 4:e00630–13. doi:10.1128/mBio.00630-13
Yamanaka H, Kobayashi H, Takahashi E, Okamoto K (2008) MacAB is involved in the secretion of Escherichia coli heat-stable enterotoxin II. J Bacteriol 190:7693–7698. doi:10.1128/JB.00853-08
Lu S, Zgurskaya HI (2013) MacA, a periplasmic membrane fusion protein of the macrolide transporter MacAB-TolC, binds lipopolysaccharide core specifically and with high affinity. J Bacteriol 195:4865–4872. doi:10.1128/JB.00756-13
Turlin E, Heuck G, Simoes Brandao MI, Szili N, Mellin JR, Lange N, Wandersman C (2014) Protoporphyrin (PPIX) efflux by the MacAB-TolC pump in Escherichia coli. Microbiol Open 3:849–859. doi:10.1002/mbo3.203
Kochevar IE (1987) Mechanisms of drug photosensitization. Photochem Photobiol 45:891–895. doi:10.1111/j.1751-1097.1987.tb07899.x
Appia-Ayme C, Patrick E, Sullivan MJ, Alston MJ, Field SJ, AbuOun M, Anjum MF, Rowley G (2011) Novel inducers of the envelope stress response BaeSR in Salmonella Typhimurium: BaeR is critically required for tungstate waste disposal. PLoS One 6:e23713. doi:10.1371/journal.pone.0023713
Baugh S, Phillips CR, Ekanayaka AS, Piddock LJ, Webber MA (2014) Inhibition of multidrug efflux as a strategy to prevent biofilm formation. J Antimicrob Chemother 69:673–681. doi:10.1093/jac/dkt420
Nikaido E, Yamaguchi A, Nishino K (2008) AcrAB multidrug efflux pump regulation in Salmonella enterica serovar Typhimurium by RamA in response to environmental signals. J Biol Chem 283:24245–24253. doi:10.1074/jbc.M804544200
Baucheron S, Nishino K, Monchaux I, Canepa S, Maurel MC, Coste F, Roussel A, Cloeckaert A et al (2014) Bile-mediated activation of the acrAB and tolC multidrug efflux genes occurs mainly through transcriptional derepression of ramA in Salmonella enterica serovar Typhimurium. J Antimicrob Chemother 69:2400–2406. doi:10.1093/jac/dku140
Yamasaki S, Nikaido E, Nakashima R, Sakurai K, Fujiwara D, Fujii I, Nishino K (2013) The crystal structure of multidrug-resistance regulator RamR with multiple drugs. Nat Commun 4:2078. doi:10.1038/ncomms3078
Olliver A, Valle M, Chaslus-Dancla E, Cloeckaert A (2004) Role of an acrR mutation in multidrug resistance of in vitro-selected fluoroquinolone-resistant mutants of Salmonella enterica serovar Typhimurium. FEMS Microbiol Lett 238:267–272. doi:10.1111/j.1574-6968.2004.tb09766.x
Eaves DJ, Ricci V, Piddock LJ (2004) Expression of acrB, acrF, acrD, marA, and soxS in Salmonella enterica serovar Typhimurium: role in multiple antibiotic resistance. Antimicrob Agents Chemother 48:1145–1150. doi:10.1128/AAC.48.4.1145-1150.2004
Nikaido E, Shirosaka I, Yamaguchi A, Nishino K (2011) Regulation of the AcrAB multidrug efflux pump in Salmonella enterica serovar Typhimurium in response to indole and paraquat. Microbiology 157:648–655. doi:10.1099/mic.0.045757-0
Olliver A, Valle M, Chaslus-Dancla E, Cloeckaert A (2005) Overexpression of the multidrug efflux operon acrEF by insertional activation with IS1 or IS10 elements in Salmonella enterica serovar Typhimurium DT204 acrB mutants selected with fluoroquinolones. Antimicrob Agents Chemother 49:289–301. doi:10.1128/AAC.49.1.289-301.2005
Nishino K, Hayashi-Nishino M, Yamaguchi A (2009) H-NS modulates multidrug resistance of Salmonella enterica serovar Typhimurium by repressing multidrug efflux genes acrEF. Antimicrob Agents Chemother 53:3541–3543. doi:10.1128/AAC.00371-09
Zwir I, Shin D, Kato A, Nishino K, Latifi T, Solomon F, Hare JM, Huang H et al (2005) Dissecting the PhoP regulatory network of Escherichia coli and Salmonella enterica. Proc Natl Acad Sci U S A 102:2862–2867. doi:10.1073/pnas.0408238102
Komatsu T, Ohta M, Kido N, Arakawa Y, Ito H, Mizuno T, Kato N (1990) Molecular characterization of an Enterobacter cloacae gene (romA) which pleiotropically inhibits the expression of Escherichia coli outer membrane proteins. J Bacteriol 172:4082–4089
van der Straaten T, Zulianello L, van Diepen A, Granger DL, Janssen R, van Dissel JT (2004) Salmonella enterica serovar Typhimurium RamA, intracellular oxidative stress response, and bacterial virulence. Infect Immun 72:996–1003. doi:10.1128/IAI.72.2.996-1003.2004
Chollet R, Chevalier J, Bollet C, Pagès JM, Davin-Regli A (2004) RamA is an alternate activator of the multidrug resistance cascade in Enterobacter aerogenes. Antimicrob Agents Chemother 48:2518–2523. doi:10.1128/AAC.48.7.2518-2523.2004
Rosenberg EY, Bertenthal D, Nilles ML, Bertrand KP, Nikaido H (2003) Bile salts and fatty acids induce the expression of Escherichia coli AcrAB multidrug efflux pump through their interaction with Rob regulatory protein. Mol Microbiol 48:1609–1619. doi:10.1046/j.1365-2958.2003.03531.x
Abouzeed YM, Baucheron S, Cloeckaert A (2008) ramR mutations involved in efflux-mediated multidrug resistance in Salmonella enterica serovar Typhimurium. Antimicrob Agents Chemother 52:2428–2434. doi:10.1128/AAC.00084-08
van der Straaten T, Janssen R, Mevius DJ, van Dissel JT (2004) Salmonella gene rma (ramA) and multiple-drug-resistant Salmonella enterica serovar typhimurium. Antimicrob Agents Chemother 48:2292–2294. doi:10.1128/AAC.48.6.2292-2294.2004
Yassien MA, Ewis HE, Lu CD, Abdelal AT (2002) Molecular cloning and characterization of the Salmonella enterica serovar Paratyphi B rma gene, which confers multiple drug resistance in Escherichia coli. Antimicrob Agents Chemother 46:360–366. doi:10.1128/AAC.46.2.360-366.2002
Batta AK, Salen G, Batta P, Tint GS, Alberts DS, Earnest DL (2002) Simultaneous quantitation of fatty acids, sterols and bile acids in human stool by capillary gas-liquid chromatography. J Chromatogr B Anal Technol Biomed Life Sci 775:153–161. doi:10.1016/S1570-0232(02)00289-1
Sonnenwirth AC (1980) The enteric bacteria and bacteroides. In: Davis BD, Dulbecco R, Eisen HN, Ginsberg HS (eds) Microbiology, 3rd edn. Harper & Row, Publishers, Philadephia, pp 645–672
Fàbrega A, Balleste-Delpierre C, Vila J (2016) Differential impact of ramRA mutations on both ramA transcription and decreased antimicrobial susceptibility in Salmonella Typhimurium. J Antimicrob Chemother 71:617–624. doi:10.1093/jac/dkv410
Schumacher MA, Miller MC, Grkovic S, Brown MH, Skurray RA, Brennan RG (2001) Structural mechanisms of QacR induction and multidrug recognition. Science 294:2158–2163. doi:10.1126/science.1066020
Alguel Y, Meng C, Teran W, Krell T, Ramos JL, Gallegos MT, Zhang X (2007) Crystal structures of multidrug binding protein TtgR in complex with antibiotics and plant antimicrobials. J Mol Biol 369:829–840. doi:10.1016/j.jmb.2007.03.062
Lei HT, Shen Z, Surana P, Routh MD, Su CC, Zhang Q, Yu EW (2011) Crystal structures of CmeR-bile acid complexes from Campylobacter jejuni. Protein Sci 20:712–723. doi:10.1002/pro.602
Bailey AM, Ivens A, Kingsley R, Cottell JL, Wain J, Piddock LJ (2010) RamA, a member of the AraC/XylS family, influences both virulence and efflux in Salmonella enterica serovar Typhimurium. J Bacteriol 192:1607–1616. doi:10.1128/JB.01517-09
Kehrenberg C, Cloeckaert A, Klein G, Schwarz S (2009) Decreased fluoroquinolone susceptibility in mutants of Salmonella serovars other than Typhimurium: detection of novel mutations involved in modulated expression of ramA and soxS. J Antimicrob Chemother 64:1175–1180. doi:10.1093/jac/dkp347
Rosenblum R, Khan E, Gonzalez G, Hasan R, Schneiders T (2011) Genetic regulation of the ramA locus and its expression in clinical isolates of Klebsiella pneumoniae. Int J Antimicrob Agents 38:39–45. doi:10.1016/j.ijantimicag.2011.02.012
Acknowledgments
The author acknowledges funding from the Japan Agency for Medical Research and Development; the Japan Science and Technology Agency; the Japan Society for the Promotion of Science; the Ministry of Education, Culture, Sports, Science and Technology, Japan; and the Cabinet Office, Government of Japan.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Nishino, K. (2016). Antimicrobial Drug Efflux Pumps in Salmonella . In: Li, XZ., Elkins, C., Zgurskaya, H. (eds) Efflux-Mediated Antimicrobial Resistance in Bacteria. Adis, Cham. https://doi.org/10.1007/978-3-319-39658-3_10
Download citation
DOI: https://doi.org/10.1007/978-3-319-39658-3_10
Published:
Publisher Name: Adis, Cham
Print ISBN: 978-3-319-39656-9
Online ISBN: 978-3-319-39658-3
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)