Abstract
Increasing evidences have emphasized the importance of gut microbiota and integrity of the intestinal epithelium to avoid the occurrence of many diseases. Recently, microRNAs have emerged as important gene expression regulators in many conditions. A dysregulated microRNA expression is a common feature of various human diseases, such as cancer , developmental abnormalities, muscular and cardiovascular disorders, and inflammatory diseases. Moreover, exosomal microRNAs have been recently reported to have a crucial role in modulating the bacterial gene expression. So far, the interplays between microRNAs expression and gut microbiota modulation have not been explored in details. To provide further insights into this interesting relationship, in this chapter we discussed some papers appeared in the literature in the last few years.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
1 The Intestinal Epithelium and the Gut Microbiota
The human body contains a great variety of bacteria, collectively referred to as the human microbiota . The human intestinal tract harbors a diverse and complex microbial community, the gut microbiota, which plays a central role in human health. It has been estimated that our gut contains up to 100 trillion microbes, 1000 bacterial species and 100-fold more genes than those codified by the human genome (Ley et al. 2006b; Qin et al. 2010).
Humans have their first contact with bacteria during birth, when the baby passes through the mother’s birth canal (Dethlefsen et al. 2007; Ley et al. 2006a). In the postnatal period, the human intestine is colonized rapidly by an array of microbes. The conditions known to influence the colonization process include the gestational age, the mode of delivery (vaginal birth vs. assisted delivery), diet (breast milk vs. formula), sanitation, and antibiotic treatment (Adlerberth and Wold 2009; Marques et al. 2010). By the end of the first year of life, infants possess an individually distinct microbial profile, converging toward the characteristic microbiota of an adult. By 2–5 years of age, the microbiota fully resembles that of an adult in terms of composition and diversity (Koenig et al. 2011; Yatsunenko et al. 2012). In the adult, the abundance and the composition of the gut microbial population is different between individuals and this variability is influenced by life style, weight, and overall metabolic state of the host (Tagliabue and Elli 2013; Tehrani et al. 2012). This life-long process of gut colonization led to the formation of a complex ecosystem where the host and its microbiome form an equilibrium that represents a remarkable example of reciprocal adaptation .
Disruptions to the normal balance between the gut microbiota and the host, that can occurs either by changes of the gut microbiota composition or by alterations of the host response, is associated with many pathological conditions such as obesity (Ley et al. 2006b; Turnbaugh et al. 2008), malnutrition (Kau et al. 2011), inflammatory bowel disease (IBD) (Dicksved et al. 2008; Frank et al. 2007), neurological disorders (Gonzalez et al. 2011) and cancer (Lupton 2004).
A coordinated interplay between commensal microbiota and mucosal immune responses occurs to maintain the host intestinal immune homeostasis . In fact, the immune system is the principal regulator of the gut microbiota homeostasis and acts mainly by maintaining the equilibrium between a correct defense against pathogens and tolerance to commensals. Environmental stimuli elicit continuously the intestinal epithelium and many gut cells are necessary to form a barrier against them. In fact, several intestinal diseases are caused by deregulation of the intestinal barrier function (Krogius-Kurikka et al. 2009). The intestinal epithelium is the largest mucosal surface of the body, covering ~400 m2. Its main function is to prevent infections and protect by invading pathogens (Johansson et al. 2011). The intestinal epithelium is organized into crypts and villi, and contains different cells: (i) pluripotent intestinal epithelial stem cells (pluripotent IESCs), that reside at the base of crypts and continuously renew the surface, (ii) enterocytes (for metabolic and digestive functions) and (iii) secretory IECs, including enteroendocrine cells, goblet cells and Paneth cells specialized for maintaining the digestive or barrier function of the epithelium. Enteroendocrine cells represent a link between the central and enteric neuroendocrine systems through the secretion of numerous hormones that regulate the digestive function. The luminal secretion of mucins and antimicrobial proteins (AMPs) by goblet cells and Paneth cells, respectively, establishes a physical and biochemical barrier to microbial contact with the epithelial surface and underlying immune cells (Gallo and Hooper 2012; Kim and Ho 2010). Many regulatory mechanisms control the equilibrium between microbiota and the host intestinal cell response (Coombes and Powrie 2008; Sartor 2008; Strober 2009). In fact, pathogens in commensal bacteria, abnormal microbial composition (i.e., decreased concentrations of protective bacteria) or defective host containment of commensal bacteria (i.e., reduced secretion of antimicrobial peptides to reduce mucosal bacterial overgrowth) may determine an imbalance of this delicate interplay (Sartor 2008). Moreover, this equilibrium is mainly determined by mucosal dendritic cells, that have an important role in the regulation of intestinal immunity processes (Coombes and Powrie 2008; Strober 2009).
However, we cannot exclude that microRNAs as well may represent complementary molecular determinants potentially involved in these processes.
2 microRNAs Biogenesis and Processing
MicroRNAs (miRNAs) have emerged as major regulators of various biological processes and important mediators of immune development and virulence (Choi et al. 2014; O’Connell et al. 2010; Slaby et al. 2009). microRNAs (miRNAs) are short, highly conserved small noncoding RNA molecules naturally occurring in the genomes of plants and animals. miRNAs are 17–27 nucleotides long and regulate post-transcriptionally the mRNA expression, typically by binding to the 3′ untranslated region (3′UTR ) of the complementary mRNA sequence, resulting in translational repression and gene silencing (Bartel 2004). microRNAs are transcribed by RNA polymerase II (Pol II) (Cai et al. 2004) and RNA polymerase III (Pol III) (Borchert et al. 2006) in primitive transcripts , named pri-miRNA. Pri-miRNAs are processed into fragments of ~70-bp, the precursors (pre-miRNAs), in a two-step process catalyzed by the proteins Drosha and Dicer (Lee et al. 2003). The exportin-5 (Exp-5) recognizes the double-stranded pre-miR and transports it from the nucleus to the cytoplasm, irrespective of miRNA nucleotide sequence and the presence of diverse structural motifs (Lund et al. 2004; Okada et al. 2009). Once in the cytoplasm, the RNA III ribonuclease Dicer complex converts the pre-miRNA in a mature miRNA, producing a miRNA–miRNA* duplex (Cullen 2004), which displays a 2-nt 3′overhang at both ends. Only one miRNA strand (the guide strand, or -5p form) of the duplex is loaded into Argonaute protein (AGO) (O’Toole et al. 2006) to form the RISC complex (referred to as the miRISC) that is the effector of the reaction by recognizing the miRNA target in a sequence-specific manner and can mediate various type of gene silencing (Tijsterman and Plasterk 2004), mRNA degradation or translation inhibition (Djuranovic et al. 2012), whereas the inactive strand (the -3p form) is degraded (Kim 2005).
3 Interplays Between miRNAs and Microbiota
miRNAs have been also found to be implicated in gut microbiota-host interactions (Kaser et al. 2011). To investigate the mechanisms by which the host cell reprogram their transcription during colonization, germ-free mice were colonized with the microbiota from pathogen-free mice (Dalmasso et al. 2011). RNA extracted from ileum and colon of germ free and colonized mice, showed down- and up-regulated miRNAs: eight microRNAs were expressed in the ileum, whereas seven in the colon. The expression of host miRNAs is modulated in response to microbiota colonization and this indicates that microbiota modulates host miRNAs expression suggesting an implication of miRNAs in microbiota-mediated host gene regulation. In particular, by intersecting the microarray-detected dysregulated genes with the potential targets of dysregulated miRNAs (predicted by at least two algorithms), the authors identified only one gene, Abcc3, potentially targeted by mmu-miR-665 in the colon, whereas no overlapping genes were found in the ileum (Dalmasso et al. 2011). Abcc3 belongs to the multidrug resistance-associated protein family, which mediates the metabolism of xenobiotics and endogenous toxins (Hooper et al. 2001). Therefore, mmu-miR-665 was identified as a microRNA potentially implicated in the colonization of microbiota through the direct targeting and inhibition of Abcc3.
Many authors found that different intestinal tracts have distinct miRNAs expression patterns. By using germ-free and conventionally raised mice, the impact of the endogenous microbiota on the global expression of caecal miRNAs in vivo has been investigated by Singh et al. (2012). The murine miRNA signature in the caecum is affected by the presence of the microbiota. Moreover, authors found that 34 putative miRNA target genes encode for proteins involved in the regulation of the intestinal barrier function (i.e., glycosylation enzymes, junctional proteins, proteins found in the mucus layers) and in the immune regulation (i.e., MHCI and II proteins). They found that the expression of miRNAs depends on the endogenous microbiota and that 16 unique miRNAs were deregulated between germ-free and conventional raised mice. By cross-matching the list of intestinal barrier genes predicted to be modulated by differentially expressed miRNAs, with genes already demonstrated to be deregulated in the mucosa of intestinal-specific Dicer knock-out mice (McKenna et al. 2010) the authors supported the hypothesis that gut commensals impact the intestinal barrier via miRNAs expression modulation.
4 Inflammatory Diseases
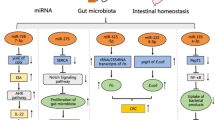
It is now apparent that a dysregulated miRNA expression is a common feature of various human diseases, such as cancer , developmental abnormalities, muscular and cardiovascular disorders, and inflammatory diseases such as inflammatory bowel diseases (IBD) (Takagi et al. 2010). In fact, a study by Xue et al. focused on the microbiota regulation of miRNAs expression and on the maintenance of intestinal homeostasis , and reported a connection between the expression of miR-10a and of its target IL-12/IL-23p40, a key molecule for innate immune responses to commensal bacteria (Xue et al. 2011). The authors found that commensal bacteria down-regulated dendritic cell miR-10a expression via TLR–TLR ligand interactions through a MyD88-dependent pathway and that mice with colitis expressed higher levels of IL-12/IL-23p40 and lower level of gut miR-10a, compared to control mice, opening new perspectives for the study of miRNAs regulation in intestinal diseases.
Intestinal inflammation is characterized by epithelial disruption, loss of barrier function, recruitment of immune cells, and host immune responses to gut microbiota . Recently, it has been observed that PepT1, a di/tripeptide transporter that uptakes bacterial products, is upregulated in inflamed colon tissue (Dai et al. 2015). This peptide has a role in bacterium-associated intestinal inflammation. The amount of this peptide is inversely correlated with the level of miR-193a-3p in inflamed colon tissues with active ulcerative colitis. Moreover, miR-193a-3p reduced PepT1 expression and activity as a target gene and subsequently suppressed the NF-κB pathway, suggesting that miR-193a-3p may have a crucial role to regulate the colonic inflammation process (through PepT1) and to maintain intestinal homeostasis .
Another example of microRNAs that regulate gut mucosal immunity has been reported by Biton et al. who studied miR-375 in mice with an inducible intestinal epithelial cell-specific deficiency in Dicer1 (Dicer1 Δgut) (Biton et al. 2011) (Fig. 3.1).
Biton et al. reported that Dicer1 depletion in the mice gut leads to goblet-cell depletion and that the regulation of goblet-cell differentiation is dependent on the expression of miR-375 (Biton et al. 2011). The expression of this miRNA is able to inhibit the translation of KLF5, an antagonist of the goblet cell–differentiation factor KLF4, supporting the differentiation of goblet cells. Moreover, they observed a lower expression of IL-4, IL-5 and IL-13 in Dicer1 Δgut mice and an enhanced susceptibility to infection by the helminth parasite Trichuris muris. IL-13, presumably supplied by TH2 cells, induces miR-375 in intestinal epithelial cells in vitro and a downstream production of the TH2-facilitating epithelial cytokine TSLP, indicating an appropriately balanced TH2 feed-forward loop regulated by miR-375. Based on their results, the authors suggested that miR-375 directs the differentiation of goblet cells and the promotion of antiparasitic TH2 immune responses. As the miR-375 expression is very high in the human intestine (Wu et al. 2010), mucosal expression of this particular miRNA might also be important in the regulation of intestinal homeostasis and protection against parasite infection in humans (Wu et al. 2010).
Previously (Masotti 2012) we reported a study by Chassin et al. who found that the TLR-4-mediated transcriptional activation of intestinal epithelial cells observed in mice immediately after birth, was induced by an oral ingestion of endotoxins from the environment and induced a post-transcriptional down-regulation of epithelial IRAK1 protein expression, which protected from secondary bacteria-induced epithelial damages (Chassin et al. 2010).
In a very recent paper, Runtsch et al. investigated the role of miR-146a in regulating intestinal immunity and barrier function and verified the miRNA expression in a variety of gut tissues in adult mice (Runtsch et al. 2015). By comparing intestinal gene expression in wild type (WT) and in miR-146a-/- mice, the authors demonstrated that miR-146a repressed a subset of immune-related signaling genes related to an increase of gut barrier and inflammation. Consistent with an enhanced intestinal barrier, Runtsch et al. found that miR-146a-/- mice, a model of Ulcerative Colitis (UC), are more resistant to the dextran sulphate-induced colitis compared to WT. The elevated expression of colonic miR-146a has been observed also in UC patients, therefore suggesting a crucial role for miR-146a in modulating the intestinal barrier function, which is a process that alters gut homeostasis and enhances some intestinal diseases. These results will constitute the basis of further research and will open new perspectives for therapeutic applications.
The same authors reviewed the literature and discussed the influence that miRNAs have on both immune and epithelial cell biology in the mammalian intestines and its impact on the microbiota . However, the authors emphasized the lack of studies aimed at deciphering the functions of specific miRNAs within the gut finalized to the understanding of the cellular mechanisms that promote intestinal homeostasis and the identification of potential molecular targets underlying intestinal diseases such as inflammatory bowel disease and colorectal cancer (Runtsch et al. 2014).
5 Symbiosis of Host and Guest
All of the papers discussed in the previous paragraphs described the interplay between the microbiota and the host. In particular, we discussed how microbiota modulates the gene expression of the host through miRNAs. So far, nothing has been know on how the host regulates the microbiota. This is a crucial point, because it represents the missing part in the big picture describing the symbiosis of the host and the guest (Fig. 3.2).
The human intestinal lumen is populated by microorganisms (gut microbiota ) that regulate the host gene expression through microRNAs. Similarly, the host produces extracellular vesicles containing microRNAs that regulate the expression of microbial genes. This ‘symbiotic loop’ is emerging as a powerful inter-kingdom communication system, although the precise molecular mechanisms underlying it are still not know. We have no doubt that this loop will be extensively explored in the next few years
To close this gap, a very recent work by Liu et al. described how the host selectively shapes the microbiota through miRNAs contained in extracellular vesicles (EVs) produced by the host itself (Liu et al. 2016). miRNAs, when contained in vesicles, are relatively stable compared to other RNAs (Jung et al. 2010). Fecal miRNAs can exist in EV-free forms, associated with high-density lipoproteins or argonaute protein (Creemers et al. 2012), or in a completely free form. Liu et al. reported that the miRNAs they have identified and characterized, can target specific bacterial genes after entering the bacteria, modulating their gene expression. In their work, Liu et al. used Escherichia coli and Fusobacterium nucleatum, two bacterial species that have been reported to promote colorectal cancer (Rubinstein et al. 2013). The authors demonstrated that different miRNAs have different ability to enter into bacteria and that miRNAs shapes bacteria with a temporal and spatial organization (Liu et al. 2016).
6 Conclusions
In this chapter, we discussed the papers that appeared in the literature in the last 5 years (Table 3.1), that studied the interplays between gut microbiota and gene expression modulation mediated by microRNAs.
We wondered whether microRNAs could be exploited therapeutically to modulate an altered gut microbiota composition and ultimately restore a healthy condition. For example, it has been reported recently that the incidence of type 1 diabetes cannot be explained only by genetics, epigenetics and environmental factors only (Gulden et al. 2015). Lifestyle, diet and the use of antibiotics also should be taken into account. The diet supplementation with pre-/pro-biotics has emerged as a potential mean to improve gut integrity and avoid the occurrence of diseases. However, we think that the recent work by Liu and colleagues (Liu et al. 2016) is a clear demonstration that other bacterial modulatory mechanisms can be elicited, as for example, the use of microRNAs for the dysregulation of bacterial gene expression. We still do not know if it will be possible to modulate gut bacterial composition by simply employing microRNAs (i.e., by modulating gut bacteria gene expression to activate cell death processes that could lead to a progressive enrichment or depletion of a given bacterial population). In any case, if validated, this kind of innovative ‘therapeutic’ intervention could be exploited also for other pathologies, and not be limited only to diabetes .
In the near future, many other works will surely prompt further research aimed at deciphering the existence of other types of interactions between the host microbiota and the guest itself. The interpretation of the complex ‘inter-kingdom communication’ system and all the ways and pathways by which these systems interact each other will be the next challenge that we are going to face
References
Adlerberth I, Wold AE (2009) Establishment of the gut microbiota in Western infants. Acta Paediatr 98(2):229–238. doi:10.1111/j.1651-2227.2008.01060.x
Bartel DP (2004) microRNAs: genomics, biogenesis, mechanism, and function. Cell 116(2):281–297
Biton M, Levin A, Slyper M, Alkalay I, Horwitz E, Mor H, Kredo-Russo S, Avnit-Sagi T, Cojocaru G, Zreik F, Bentwich Z, Poy MN, Artis D, Walker MD, Hornstein E, Pikarsky E, Ben-Neriah Y (2011) Epithelial microRNAs regulate gut mucosal immunity via epithelium-T cell crosstalk. Nat Immunol 12(3):239–246. doi:10.1038/ni.1994
Borchert GM, Lanier W, Davidson BL (2006) RNA polymerase III transcribes human microRNAs. Nat Struct Mol Biol 13(12):1097–1101. doi:10.1038/nsmb1167
Cai X, Hagedorn CH, Cullen BR (2004) Human microRNAs are processed from capped, polyadenylated transcripts that can also function as mRNAs. RNA 10(12):1957–1966. doi:10.1261/rna.7135204
Chassin C, Kocur M, Pott J, Duerr CU, Gutle D, Lotz M, Hornef MW (2010) miR-146a mediates protective innate immune tolerance in the neonate intestine. Cell Host Microbe 8(4):358–368. doi:10.1016/j.chom.2010.09.005
Choi EJ, Kim HB, Baek YH, Kim EH, Pascua PN, Park SJ, Kwon HI, Lim GJ, Kim S, Kim YI, Choi YK (2014) Differential microRNA expression following infection with a mouse-adapted, highly virulent avian H5N2 virus. BMC Microbiol 14:252. doi:10.1186/s12866-014-0252-0
Coombes JL, Powrie F (2008) Dendritic cells in intestinal immune regulation. Nat Rev Immunol 8(6):435–446. doi:10.1038/nri2335
Creemers EE, Tijsen AJ, Pinto YM (2012) Circulating microRNAs: novel biomarkers and extracellular communicators in cardiovascular disease? Circ Res 110(3):483–495. doi:10.1161/CIRCRESAHA.111.247452
Cullen BR (2004) Transcription and processing of human microRNA precursors. Mol Cell 16(6):861–865. doi:10.1016/j.molcel.2004.12.002
Dai X, Chen X, Chen Q, Shi L, Liang H, Zhou Z, Liu Q, Pang W, Hou D, Wang C, Zen K, Yuan Y, Zhang CY, Xia L (2015) MicroRNA-193a-3p Reduces Intestinal Inflammation in Response to Microbiota via Down-regulation of Colonic PepT1. J Biol Chem 290(26):16099–16115. doi:10.1074/jbc.M115.659318
Dalmasso G, Nguyen HT, Yan Y, Laroui H, Charania MA, Ayyadurai S, Sitaraman SV, Merlin D (2011) Microbiota modulate host gene expression via microRNAs. PLoS ONE 6(4):e19293. doi:10.1371/journal.pone.0019293
Dethlefsen L, McFall-Ngai M, Relman DA (2007) An ecological and evolutionary perspective on human-microbe mutualism and disease. Nature 449(7164):811–818. doi:10.1038/nature06245
Dicksved J, Halfvarson J, Rosenquist M, Jarnerot G, Tysk C, Apajalahti J, Engstrand L, Jansson JK (2008) Molecular analysis of the gut microbiota of identical twins with Crohn’s disease. ISME J 2(7):716–727. doi:10.1038/ismej.2008.37
Djuranovic S, Nahvi A, Green R (2012) miRNA-mediated gene silencing by translational repression followed by mRNA deadenylation and decay. Science 336(6078):237–240. doi:10.1126/science.1215691
Frank DN, St Amand AL, Feldman RA, Boedeker EC, Harpaz N, Pace NR (2007) Molecular-phylogenetic characterization of microbial community imbalances in human inflammatory bowel diseases. Proc Natl Acad Sci USA 104(34):13780–13785. doi:10.1073/pnas.0706625104
Gallo RL, Hooper LV (2012) Epithelial antimicrobial defence of the skin and intestine. Nat Rev Immunol 12(7):503–516. doi:10.1038/nri3228
Gonzalez A, Stombaugh J, Lozupone C, Turnbaugh PJ, Gordon JI, Knight R (2011) The mind-body-microbial continuum. Dialogues Clin Neurosci 13(1):55–62
Gulden E, Wong FS, Wen L (2015) The gut microbiota and Type 1 Diabetes. Clin Immunol 159(2):143–153. doi:10.1016/j.clim.2015.05.013
Hooper LV, Wong MH, Thelin A, Hansson L, Falk PG, Gordon JI (2001) Molecular analysis of commensal host-microbial relationships in the intestine. Science 291(5505):881–884. doi:10.1126/science.291.5505.881
Johansson ME, Larsson JM, Hansson GC (2011) The two mucus layers of colon are organized by the MUC2 mucin, whereas the outer layer is a legislator of host-microbial interactions. Proc Natl Acad Sci USA 108(Suppl 1):4659–4665. doi:10.1073/pnas.1006451107
Jung M, Schaefer A, Steiner I, Kempkensteffen C, Stephan C, Erbersdobler A, Jung K (2010) Robust microRNA stability in degraded RNA preparations from human tissue and cell samples. Clin Chem 56(6):998–1006. doi:10.1373/clinchem.2009.141580
Kaser A, Niederreiter L, Blumberg RS (2011) Genetically determined epithelial dysfunction and its consequences for microflora-host interactions. Cell Mol Life Sci 68(22):3643–3649. doi:10.1007/s00018-011-0827-y
Kau AL, Ahern PP, Griffin NW, Goodman AL, Gordon JI (2011) Human nutrition, the gut microbiome and the immune system. Nature 474(7351):327–336. doi:10.1038/nature10213
Kim VN (2005) MicroRNA biogenesis: coordinated cropping and dicing. Nat Rev Mol Cell Biol 6(5):376–385. doi:10.1038/nrm1644
Kim YS, Ho SB (2010) Intestinal goblet cells and mucins in health and disease: recent insights and progress. Curr Gastroenterol Rep 12(5):319–330. doi:10.1007/s11894-010-0131-2
Koenig JE, Spor A, Scalfone N, Fricker AD, Stombaugh J, Knight R, Angenent LT, Ley RE (2011) Succession of microbial consortia in the developing infant gut microbiome. Proc Natl Acad Sci USA 108(Suppl 1):4578–4585. doi:10.1073/pnas.1000081107
Krogius-Kurikka L, Lyra A, Malinen E, Aarnikunnas J, Tuimala J, Paulin L, Makivuokko H, Kajander K, Palva A (2009) Microbial community analysis reveals high level phylogenetic alterations in the overall gastrointestinal microbiota of diarrhoea-predominant irritable bowel syndrome sufferers. BMC Gastroenterol 9:95. doi:10.1186/1471-230X-9-95
Lee Y, Ahn C, Han J, Choi H, Kim J, Yim J, Lee J, Provost P, Radmark O, Kim S, Kim VN (2003) The nuclear RNase III Drosha initiates microRNA processing. Nature 425(6956):415–419. doi:10.1038/nature01957
Ley RE, Peterson DA, Gordon JI (2006a) Ecological and evolutionary forces shaping microbial diversity in the human intestine. Cell 124(4):837–848. doi:10.1016/j.cell.2006.02.017
Ley RE, Turnbaugh PJ, Klein S, Gordon JI (2006b) Microbial ecology: human gut microbes associated with obesity. Nature 444(7122):1022–1023. doi:10.1038/4441022a
Liu S, da Cunha AP, Rezende RM, Cialic R, Wei Z, Bry L, Comstock LE, Gandhi R, Weiner HL (2016) The Host Shapes the Gut Microbiota via Fecal MicroRNA. Cell Host Microbe 19(1):32–43. doi:10.1016/j.chom.2015.12.005
Lund E, Guttinger S, Calado A, Dahlberg JE, Kutay U (2004) Nuclear export of microRNA precursors. Science 303(5654):95–98. doi:10.1126/science.1090599
Lupton JR (2004) Microbial degradation products influence colon cancer risk: the butyrate controversy. J Nutr 134(2):479–482
Marques TM, Wall R, Ross RP, Fitzgerald GF, Ryan CA, Stanton C (2010) Programming infant gut microbiota: influence of dietary and environmental factors. Curr Opin Biotechnol 21(2):149–156. doi:10.1016/j.copbio.2010.03.020
Masotti A (2012) Interplays between gut microbiota and gene expression regulation by miRNAs. Front Cell Infect Microbiol 2:137. doi:10.3389/fcimb.2012.00137
McKenna LB, Schug J, Vourekas A, McKenna JB, Bramswig NC, Friedman JR, Kaestner KH (2010) microRNAs control intestinal epithelial differentiation, architecture, and barrier function. Gastroenterology 139(5):1654–1664, 1664 e1651. doi:10.1053/j.gastro.2010.07.040
O’Connell RM, Rao DS, Chaudhuri AA, Baltimore D (2010) Physiological and pathological roles for microRNAs in the immune system. Nat Rev Immunol 10(2):111–122. doi:10.1038/nri2708
O’Toole AS, Miller S, Haines N, Zink MC, Serra MJ (2006) Comprehensive thermodynamic analysis of 3′ double-nucleotide overhangs neighboring Watson-Crick terminal base pairs. Nucleic Acids Res 34(11):3338–3344. doi:10.1093/nar/gkl428
Okada C, Yamashita E, Lee SJ, Shibata S, Katahira J, Nakagawa A, Yoneda Y, Tsukihara T (2009) A high-resolution structure of the pre-microRNA nuclear export machinery. Science 326(5957):1275–1279. doi:10.1126/science.1178705
Qin J, Li R, Raes J, Arumugam M, Burgdorf KS, Manichanh C, Nielsen T, Pons N, Levenez F, Yamada T, Mende DR, Li J, Xu J, Li S, Li D, Cao J, Wang B, Liang H, Zheng H, Xie Y, Tap J, Lepage P, Bertalan M, Batto JM, Hansen T, Le Paslier D, Linneberg A, Nielsen HB, Pelletier E, Renault P, Sicheritz-Ponten T, Turner K, Zhu H, Yu C, Jian M, Zhou Y, Li Y, Zhang X, Qin N, Yang H, Wang J, Brunak S, Dore J, Guarner F, Kristiansen K, Pedersen O, Parkhill J, Weissenbach J, Bork P, Ehrlich SD (2010) A human gut microbial gene catalogue established by metagenomic sequencing. Nature 464(7285):59–65. doi:10.1038/nature08821
Rubinstein MR, Wang X, Liu W, Hao Y, Cai G, Han YW (2013) Fusobacterium nucleatum promotes colorectal carcinogenesis by modulating E-cadherin/beta-catenin signaling via its FadA adhesin. Cell Host Microbe 14(2):195–206. doi:10.1016/j.chom.2013.07.012
Runtsch MC, Round JL, O’Connell RM (2014) microRNAs and the regulation of intestinal homeostasis. Front Genet 5:347. doi:10.3389/fgene.2014.00347
Runtsch MC, Hu R, Alexander M, Wallace J, Kagele D, Petersen C, Valentine JF, Welker NC, Bronner MP, Chen X, Smith DP, Ajami NJ, Petrosino JF, Round JL, O’Connell RM (2015) MicroRNA-146a constrains multiple parameters of intestinal immunity and increases susceptibility to DSS colitis. Oncotarget 6(30):28556–28572. doi:10.18632/oncotarget.5597
Sartor RB (2008) Microbial influences in inflammatory bowel diseases. Gastroenterology 134(2):577–594. doi:10.1053/j.gastro.2007.11.059
Singh N, Shirdel EA, Waldron L, Zhang RH, Jurisica I, Comelli EM (2012) The murine caecal microRNA signature depends on the presence of the endogenous microbiota. Int J Biol Sci 8(2):171–186
Slaby O, Svoboda M, Michalek J, Vyzula R (2009) microRNAs in colorectal cancer: translation of molecular biology into clinical application. Mol Cancer 8:102. doi:10.1186/1476-4598-8-102
Strober W (2009) The multifaceted influence of the mucosal microflora on mucosal dendritic cell responses. Immunity 31(3):377–388. doi:10.1016/j.immuni.2009.09.001
Tagliabue A, Elli M (2013) The role of gut microbiota in human obesity: recent findings and future perspectives. Nutr Metab Cardiovasc Dis 23(3):160–168. doi:10.1016/j.numecd.2012.09.002
Takagi T, Naito Y, Mizushima K, Hirata I, Yagi N, Tomatsuri N, Ando T, Oyamada Y, Isozaki Y, Hongo H, Uchiyama K, Handa O, Kokura S, Ichikawa H, Yoshikawa T (2010) Increased expression of microRNA in the inflamed colonic mucosa of patients with active ulcerative colitis. J Gastroenterol Hepatol 25(Suppl 1):S129–S133. doi:10.1111/j.1440-1746.2009.06216.x
Tehrani AB, Nezami BG, Gewirtz A, Srinivasan S (2012) Obesity and its associated disease: a role for microbiota? Neurogastroenterol Motil 24(4):305–311. doi:10.1111/j.1365-2982.2012.01895.x
Tijsterman M, Plasterk RH (2004) Dicers at RISC; the mechanism of RNAi. Cell 117(1):1–3
Turnbaugh PJ, Backhed F, Fulton L, Gordon JI (2008) Diet-induced obesity is linked to marked but reversible alterations in the mouse distal gut microbiome. Cell Host Microbe 3(4):213–223. doi:10.1016/j.chom.2008.02.015
Wu W, He C, Liu C, Cao AT, Xue X, Evans-Marin HL, Sun M, Fang L, Yao S, Pinchuk IV, Powell DW, Liu Z, Cong Y (2015) miR-10a inhibits dendritic cell activation and Th1/Th17 cell immune responses in IBD. Gut 64(11):1755–1764. doi:10.1136/gutjnl-2014-307980
Wu F, Zhang S, Dassopoulos T, Harris ML, Bayless TM, Meltzer SJ, Brant SR, Kwon JH (2010) Identification of microRNAs associated with ileal and colonic Crohn’s disease. Inflamm Bowel Dis 16(10):1729–1738. doi:10.1002/ibd.21267
Xue X, Feng T, Yao S, Wolf KJ, Liu CG, Liu X, Elson CO, Cong Y (2011) Microbiota downregulates dendritic cell expression of miR-10a, which targets IL-12/IL-23p40. J Immunol 187(11):5879–5886. doi:10.4049/jimmunol.1100535
Yatsunenko T, Rey FE, Manary MJ, Trehan I, Dominguez-Bello MG, Contreras M, Magris M, Hidalgo G, Baldassano RN, Anokhin AP, Heath AC, Warner B, Reeder J, Kuczynski J, Caporaso JG, Lozupone CA, Lauber C, Clemente JC, Knights D, Knight R, Gordon JI (2012) Human gut microbiome viewed across age and geography. Nature 486(7402):222–227. doi:10.1038/nature11053
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Celluzzi, A., Masotti, A. (2016). Interplays Between Gut Microbiota and Gene Expression Regulation by miRNAs: Towards a Symbiotic Vision of Host and Guest. In: Leitão, A., Enguita, F. (eds) Non-coding RNAs and Inter-kingdom Communication. Springer, Cham. https://doi.org/10.1007/978-3-319-39496-1_3
Download citation
DOI: https://doi.org/10.1007/978-3-319-39496-1_3
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-39494-7
Online ISBN: 978-3-319-39496-1
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)