Abstract
Lipids are important signaling compounds in plants. They can range from small lipophilic molecules like the dicarboxylic acid Azelaic acid to complex phosphoglycerolipids and regulate plant development as well as the response to biotic and abiotic stress. While their intracellular function is well described, several lipophilic signals are known to be found in the plant phloem and can, thus, also play a role in long-distance signaling. Mostly, they play a role in the pathogen response and systemic acquired resistance. This is particularly true for oxylipins, dehydroabietinal, and azelaic acid. However, several phospholipids have now been described in phloem exudates. Their intracellular function as well as implications and a model for long-distance signaling are discussed in this chapter.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
Introduction
The world population is projected to increase by 2.4 billion, from 7.2 billion in 2013 to 9.6 billion in 2050, with most of the increase occurring within less developed regions (www.un.com). Combined with the encroachment of cities onto arable land this necessitates an increase in agricultural yields. When faced with similar issues in the 1940s, the “green revolution,” led by Norman Borlaug, provided solutions through the development of high-yielding varieties of cereal grains, the modernization of management techniques and irrigation systems, as well as distribution of hybrid seeds, synthetic fertilizers, and pesticides to farmers. Today, the amount of arable land is limited and often, there is now a competition between food and fuel crops. In addition, changes in the global climate may impact future yields. Unlike animals, plants cannot move to escape adverse conditions. Plants had to develop mechanisms to detect changes in their environment, communicate these changes to different organs, and adjust development accordingly (Thomas and Vince-Prue 1997). These responses can occur within one cell or tissue; however, often we see a systemic response at a location distal from the detection of those environmental changes. One path to move these long-distance signals is the plant phloem. Its main conductive system is composed of large tubular cells, the sieve elements, which connect to form sieve tubes. The length of the phloem can reach up to 100 m in tall trees. To allow for an unobstructed flow, the sieve elements, cells through which transport occurs, have lost their nuclei, ribosomes, and most organelles during development. Cell walls at the interphase between two sieve elements contain large sieve pores. Sieve elements are left with the plasma membrane and a thin cytoplasm containing ER, phloem-specific plastids, and a few dilated mitochondria (VanBel and Knoblauch 2000; Turgeon and Wolfe 2009). The residual ER is found near the plasmodesmata which connect the sieve elements with the companion cells (Lucas et al. 2013). Molecules found within the sieve element are largely synthesized in the companion cells. For long-distance movement, these molecules need to be transported into the sieve element via plasmodesmata, moved along the sieve tube, and are either perceived at their target location or transported out of the phloem. Transport of photoassimilates as well as signaling molecules is thought to occur from source (photosynthetically active mature leaves) to sink (immature leaves, roots, fruits, flowers, etc.) in a mechanism driven by the osmotic gradient (“Pressure flow hypothesis ”; Münch 1930; for a review see Froehlich et al. 2011; Lucas et al. 2013). The view of the phloem function has evolved from that of simple assimilate transport to a complex trafficking system for stress signals and developmental regulators (Zeevaart 1976; Wu et al. 2003; Ding et al. 2003; Ayre and Turgeon 2004; Haywood et al. 2005) in the form of small molecules (Corbesier et al. 2003; Chen et al. 2001), peptides/proteins (Fisher et al. 1992; Kühn et al. 1997; Marentes and Grusak 1998; Kehr et al. 1999; Xoconostle-Cazares et al. 1999; Haebel and Kehr 2001; Hoffmann-Benning et al. 2002; Giavalisco et al. 2006; Lin et al. 2009; Benning et al. 2012), nucleic acids (Citovsky and Zambryski 2000; Haywood et al. 2005; Ding et al. 2003; Ruiz-Medrano et al. 1999; Yoo et al. 2004), and more recently, lipids (Madey et al. 2002; Behmer et al. 2010; Guelette et al. 2012; Benning et al. 2012; Tetyuk et al. 2013). Thus, the study of signaling compounds within the phloem is essential for our understanding of the transmission of environmental cues throughout the plant. While there is a large focus on the function of proteins, mRNAs and microRNAs in long-distance signaling, less attention has been paid to small lipophilic metabolites, and even less to more complex lipids. Yet, both, small lipophilic molecules as well as several phospholipids, di- and triacylglycerols (DAG and TAG, respectively) have been found in phloem exudates (Table 14.1). The former have already been shown to play an important role in long-distance signaling . Most play a role in the response to biotic stress and systemic-acquired resistance (SAR). Others are plant hormones with functions ranging from pathogen response to plant development. Their known function as well as possible roles of phospholipids will be discussed below.
Lipophilic Compounds in the Phloem
Plant Hormones
The phloem has been shown to contain several “lipid” plant hormones including Auxin, Gibberellins, Cytokinins, Jasmonate, and Abscisic acid (ABA; Hoad 1995), all of which signal different aspects of plant development and, in the case of ABA, certain abiotic stresses. Auxins, Cytokinins and ABA are found in the xylem as well. Auxins have been detected in the phloem of over 14 species and shown to be mobile. Whether auxin is synthesized in the sieve elements or imported from other cells is, as of yet, unknown (Hoad 1995). Auxin and Cytokinin play an important role in signaling/regulating the development of both the xylem and the phloem (see Lucas et al. 2013). Auxin is thought to integrate the phosphate status with the plant growth response by changing root architecture (López-Bucio et al. 2002; Chiou and Lin 2011). It is also implied in systemic acquired resistance (Truman et al. 2010). ABA is transported in the phloem of many woody and herbaceous species in response to many stresses including drought. A function in phloem loading and unloading has also been proposed (Hoad 1995). It will be discussed further below. Plant hormones such as systemin, salicylic acid (SA) and jasmonic acid (JA) , which provoke a pathogen response, have been identified and transported within the phloem in response to pathogen infection (Schilmiller and Howe 2005; Truman et al. 2007). In addition, the phloem contains methyl salicylate (2-hydroxy benzoic acid), which plays a role in thermogenesis (Raskin 1992) and plant defense/systemic acquired resistance:
Small Lipophilic Metabolites and Systemic Acquired Resistance (SAR)
Systemic acquired resistance is a response to pathogen infection that puts distal, pathogen-free parts of a plant into a state of increased preparedness against further infections. As a result, defense molecules are induced faster and stronger during subsequent infections. This response can be passed on through several generations in an epigenetic mechanism (Luna and Ton 2012). However, its origin lies within one plant and is mediated by proteins and small metabolites. An essential component of SAR is the movement of a SAR -signal from the infection site to distal organs of the plant, most likely through the phloem (Tuzun and Kuc 1985; Maldonado et al. 2002; Chaturvedi et al. 2008; Jung et al. 2009; Chanda et al. 2011). Candidates for such a SAR-signal include several small lipophilic molecules: methyl salicylate (Chaturvedi and Shah 2007), dehydroabietinal (Chaturvedi et al. 2012), a Glycerol-3-phosphate-dependent factor (Chanda et al. 2011; Mandal et al. 2011), and azelaic acid (Jung et al. 2009). All four are increased in leaves infected with pathogens.
Salicylic acid (SA) is considered one of the key factors in SAR. Both, SA as well as its glucoside are increased in tissues infected by a variety of pathogens (Shah et al. 2014). There they lead to the subsequent expression of SA-responsive genes like PR1 (pathogenesis-related 1) and a local defense response. In addition to this localized response, SA is converted to methyl salicylate (Me-SA) and elicits a systemic response as well. While SA has been found in phloem exudates, its movement is still debated and not sufficient to explain the increase of SA in distal leaves (Malamy et al. 1990; Métraux et al. 1990). One possibility is an increased SA biosynthesis in the distal leaves in response to additional mobile signals. This includes Me-SA, which is produced from SA in Tobacco-Mosaic-Virus (TMW)-infected leaves, and moves through the phloem to elicit a SAR response in the distal (non-infected) tobacco leaves (Park et al. 2007). A second important factor in SA accumulation and SAR is the abietane diterpenoid dehydroabietinal (DA ; Chaturvedi et al. 2012): DA was purified and identified as a small hydrophobic factor in SAR. Picomolar applications of DA to leaves led to its movement to distal leaves as well as SA accumulation and a SAR response. While the DA level does not increase in petiole exudates from infected plants, 2H-DA can rapidly translocate from the point of application to the rest of the plant, suggesting a mobile DA-derivative (Chaturvedi et al. 2012). Interestingly, free dehydroabietic acid was found in phloem exudates from non-infected plants (Guelette et al. 2012). DA, on the other hand, is found in a pathogen-induced high-molecular-weight complex (HMW), which also includes DIR1 (see Shah et al. 2014). While this complex is likely not phloem mobile, it is possible that DIR 1 facilitates the long-distance movement of DA. An additional factor in systemic signaling is a Glycerol-3-phosphate-derivative of unknown structure. Glycerol-3-phosphate (G3P) in itself is not a lipid; yet, it forms the backbone of glycerolipids, which include membrane-, storage-, and signaling lipids. During SAR, G3P-levels increase in the infected leaf, petiole exudates, and the distal, non-infected leaves, however, when radiolabeled G3P was applied to leaves, it did not move. Instead, the radioactivity was recovered in an unknown compound within the distal leaf (Chanda et al. 2011). This suggests that rather than G3P, an as of yet, unidentified G3P derivative (G3P*) may be the mobile signal. Azelaic acid (AzA) , a small dicarboxylic acid, was found in phloem exudates of infected plants (Jung et al. 2009). It is derived through the oxidation of C18 unsaturated fatty acids in a ROS-mediated process. Only a small fraction of the labeled AzA that had been applied to a leaf could be recovered in a distal leaf making long-distance movement questionable. It may, however, act in SAR by increasing the production of G3P (Yu et al. 2013).
In addition to these four lipophilic molecules, the SAR response is mediated by two proteins: DIR1 (Defective in induced resistance 1 ) and AZI1 (Azelaic acid induced 1 ). DIR1 is a lipid transfer protein that has been found within the phloem (Maldonado et al. 2002; Mitton et al. 2009) and was shown to move systemically throughout the plant (Champigny et al. 2013; Yu et al. 2013); DIR1 can interact with itself as well as with AZI1. The proposed networking between these proteins and G3P, DA, and AzA has been summarized comprehensively in Shah et al. 2014.
Oxylipins/Jasmonic Acid
Additional, more complex lipophilic signals moving through the phloem are the oxylipins. Oxylipins are oxygenated polyunsaturated fatty acids derived from the polyunsaturated acyl groups in chloroplast membrane galactolipids (Howe and Schilmiller 2002; Feussner and Wasternack 2002; Yan et al. 2013). Jasmonates are formed though a LOX-catalyzed peroxidation of the trienoic fatty acids to form 13-hydroperoxide. They are further modified through the allene oxide synthase pathway (AOS) or the hydrodyperoxide lyase pathway (HPL), which partition oxylipin biosynthesis into two pools of molecules with varying functions (Chehab et al. 2008). In the AOS pathway, 12-OPDA (cis12-oxo-phytodienoic acid) and JA are produced, while the HPL pathway produces aldehydes and their volatile relatives (green leaf volatiles; GLV). Jasmonates include Jasmonic acid (JA), methyl-jasmonate (Me-JA), jasmonate-isoleucine (JA-Ile), and OPDA. They play a role in the regulation of plant growth and development as well as in the response to biotic and abiotic stress. 12-OPDA is not just a JA precursor but a signaling molecule itself (Dave and Graham 2012). In response to wounding or herbivory, JA is synthesized and moves throughout the plant to elicit a (systemic) defense response (Truman et al. 2007; Thorpe et al. 2007). This response is not limited to herbivores but extends to bacterial and fungal pathogens as well (Yan et al. 2012). In many cases jasmonates display a cooperative response with other plant hormones – it can act synergistically with ethylene, salicylic acid, ABA, and gibberellins (Yan et al. 2013). Different environmental stresses produce different oxylipins profiles: Savchenko et al. (2014) found that while wounding induces JA and its precursor 12-OPDA, drought only induces 12-OPDA. This induction is correlated with a differential increase in plant ABA levels under the same conditions. More importantly, external application of ABA and/or 12-OPDA could overcome the defect in stomatal closure in ABA- and 12-OPDA biosynthesis mutants, suggesting ABA/OPDA crosstalk during drought signaling (Savchenko et al. 2014). Interestingly, this mirrors findings that OPDA plays a role in seed development and germination, which are cellular processes that are also mediated by ABA (Dave et al. 2011).
(Phospho-)Glycerolipids in the Phloem: A Possible Model of their Transport
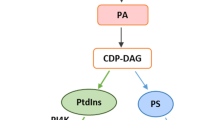
In analogy to animal systems, lipids and lipid-binding proteins may play an important role in stress and developmental signaling in plants. Lipids, specifically phospholipids, are mainly found within cell membranes to which they provide both structure and act as mediators that regulate various aspects of plant development and environmental interactions (Wang and Chapman 2013; Wang 2004; Xue et al. 2009). Signaling lipids include glycerolipids, sphingolipids, fatty acids, oxylipins, and sterols. Phospholipid signaling cascades are normally grouped according to the phospholipase that catalyzes the formation of other messengers such as PtdOH, DAG, DAG pyrophosphate (DGPP), lysophospholipids, free fatty acids, oxylipins, phosphoinositides, and inositol phosphates (Zhu 2002; Wang 2004; Wang et al. 2007).
Unlike structural lipids, signaling lipids occur only in minute amounts, but increase rapidly and transiently in response to certain stimuli. The resulting signal is quickly downregulated. Consequently, signaling lipids typically have a higher turnover than other structural lipids (Testerink and Munnik 2005). Lipids such as phosphatidic acid (PtdOH), phosphatidyl inositol (PtdIns), and its phosphates (PtdInsPs) are well-known secondary messengers within plant cells (Wang 2004; Munnik and Testerink 2009), but essentially nothing is known about their involvement in long-distance signaling. A recent analysis of phloem exudates of Arabidopsis thaliana, led to the identification of several glycerolipids within the phloem including phosphatidic acid (PtdOH), phosphatidyl choline (PtdCho), phosphatidyl inositol (PtdIns), di- and triacylglycerols (DAG and TAG, respectively; see Guelette et al. 2012; Benning et al. 2012; Tetyuk et al. 2013). Similarly, lipids have been detected in canola phloem as well (Madey et al. 2002). Several possibilities exist for the function of these lipids in the phloem:
-
I.
Lipids could be transported through the phloem as energy carriers, assimilates, or building blocks. However, most tissues including seeds are capable of making their own (storage) lipids, hence, transport as energy carriers or building blocks would be inefficient (Ohlrogge and Browse 1995).
-
II.
Lipids could be the product of regular membrane turnover within the sieve element (Lucas et al. 2013). When we compared leaf and phloem lipids, the lipid profiles were distinct, with several lipids being unique to the phloem (Guelette et al. 2012). It does not exclude the possibility though, that they are generated from other membrane lipids to serve as anchors for proteins or precursors for signaling molecules. For example, one of the PtdCho species found in phloem exudates contains a 18:3 acyl group. Polyunsaturated lipids are known precursors to signaling lipids, such as oxylipins, which are phloem mobile. However, the first step of oxylipins biosynthesis occurs in the chloroplast, a plastid that is not present in sieve elements. Moreover, Buseman et al. (2006) had proposed that galactolipids, which are plastidal lipids, are the substrate for the production of OPDA. This suggests that phloem lipids have phloem-specific functions and are likely not products of non-specific membrane turnover. Similarly, as part of the seasonal senescence-related mobilization, cellular components including lipids are metabolized and moved. However, this mobilization tends to occur in the form of sucrose rather than lipids (Thomas and Stoddard 1980).
-
III.
Phloem lipids could serve as long distance signals. Long-distance transport of hydrophobic compounds does occur in aqueous biological systems: For example, lipids in the blood are transported while bound to proteins. These lipid-protein complexes are transported to other tissues for storage, modification, or degradation (e.g. cholesterol; Glatz et al. 1995; Charbonneau et al. 2009), others transport vitamins (e.g., vitamin A: Blaner 1989) or serve as messengers and modulate transcription factor activity (Tontonez et al. 1994; Nagy and Szanto 2005; Musille et al. 2013). While these lipid-protein mechanisms are key to mammalian development their possible importance in plants is virtually unexplored.
All three of the above mechanisms would require the assistance/binding of proteins. As a result, we and others proposed that phloem (phospho-) lipids could act as long-distance developmental signals or in response to abiotic stress and that phloem lipid-binding proteins participate in different aspects of this signaling cascade (Benning et al. 2012; Zheng et al. 2012):
-
(i)
Proteins could release the lipid into the phloem, either by generating it in within the sieve element, possibly from the membrane and through a lipase-like activity; or they could transport the lipid into the phloem via plasmodesmata.
-
(ii)
Proteins could solubilize the lipid and transport it to its target organ. This would imply that the lipid is the mobile signal. Alternatively, the protein could be a signal which needs a bound lipid for functionality. An example for such a mechanism would be the Frizzled-Wnt interaction. Wnt is an important developmental regulator in animal systems, which binds to its receptor, Frizzled. It had long been known that Wnt can only solicit the downstream signaling cascade when bound to a lipid, palmitoleic acid. In 2012, crystallographic analysis showed that palmitoleic acid binding to Wnt is essential for Wnt-Frz interaction (Janda et al. 2012). A similar mechanism is conceivable in plants.
-
(iii)
Proteins either transport the lipid out of the sieve element or are part of a receptor that binds the signaling lipid and leads to a change in development (Fig. 14.1).
Fig. 14.1 Model of possible lipid signaling (derived from Hoffmann-Benning 2015)
To identify possible proteinaceous components of long-distance lipid signaling and transport in Arabidopsis we had used a proteomics (LC-ESI-MS/MS) approach. This allowed us to identify phloem proteins with predicted lipid-binding sites (Guelette et al. 2012). These findings together with the analysis of other phloem proteome databases led to the identification of 13 putative lipid-binding proteins (LBPs; Guelette et al. 2012; Benning et al. 2012) in Arabidopsis and additional plant species such as rice, canola, cucurbits, lupine, and Perilla (Hayashi et al. 2000; Hoffmann-Benning et al. 2002; Barnes et al. 2004; Giavalisco et al. 2006; Lin et al. 2009; Guelette et al. 2012). Genes encoding most of these proteins are expressed in companion cells (CC; Mustroph et al. 2009). References describing their first description in the phloem as well as their function according to the model above are summarized in Table 14.2.
The question arises whether any of the glycerolipids identified in phloem exudates are binding to any of the phloem lipid-binding proteins (LBPs)? And could they be signaling molecules? The answer is a definite yes. Six of these predicted phloem lipid-binding proteins have been studied further: DIR1 , a lipid transfer protein (LTP) plays a role in systemic acquired resistance in Arabidopsis and tomato (Maldonado et al. 2002; Lascombe et al. 2008; Mitton et al. 2009; see Shah et al. 2014). It has been discussed above. Annexin binds phospholipids and is involved in intracellular Ca2+-signaling as well as callose formation (Andrawis et al. 1993; Mortimer et al. 2008). FT signals the induction of flowering and seasonal leaf abscission (Corbesier et al. 2007; Tamaki et al. 2007; Lin et al. 2007). It binds PtdChos, however, whether this interaction occurs in the phloem/SE, leaf mesophyll cells or apical meristem, remains to be elucidated (Nakamura et al. 2014). 14-3-3 proteins are a component of the receptor system detecting the presence of FT in the phloem near the apical meristem (Taoka et al. 2011). While lipid-binding of these proteins has been shown, no connection has been made between these proteins and phloem lipids. Acyl-CoA-binding proteins (ACBPs) have been identified in phloem exudates from rice (Hayashi et al. 2000) and associated with vascular bundles in Arabidopsis (Chen et al. 2008; Zheng et al. 2012). ACBPs will be discussed in a separate chapter in this book. PLAFP , is a 20 kDa protein with a PLAT/LH2 domain (Polycystin-1, Lipoxygenase, Alpha-Toxin-domain). This domain is thought to mediate interaction with lipids or membrane-bound proteins (Bateman and Sandford 1999). Proteins containing this domain are typically induced by stress. PLAFP binds PtdOH, one of the lipid species found in phloem exudates (Benning et al. 2012). PLAFP and its possible function in long-distance lipid signaling will be discussed below.
The Role of Phospholipases in Phospholipid Signaling
Phospholipids are more than just membrane lipids. Phospholipid signaling cascades are normally grouped according to the phospholipase that catalyzes their formation (Zhu 2002; Wang 2004, ; Wang et al. 2007): phospholipase D (PLD), phospholipase C (PLC), phospholipase A1 (PLA1), and phospholipase A2 (PLA2). PLC and PLD cleave at the first and the second phosphodiester bond, respectively, while the PLAs cleave the acyl chains from the membrane lipid. Plant phospholipases participate in different cellular and physiological processes, which are classified into three categories: cellular regulation, membrane lipid remodeling, and lipid degradation; furthermore, some phospholipases can participate in the biosynthesis of storage lipids (Wang et al. 2012). Phospholipase D catalyzes the hydrolysis of glycerophospholipids into phosphatidic acid (PtdOH) and a free head group, such as choline or serine. Arabidopsis encodes 12 distinct PLDs: PLDα(1,2,3), PLDβ(1,2), PLDγ(1,2,3), PLDδ, PLDε, and PLDζ (Wang et al. 2012). Each of these subgroups is based on molecular and biochemical properties. Additionally, PLD can be classified into two groups based on their lipid binding domains; those with both pleckstrin homology (PH) and phox homology (PX) domains belong to the PLDζ class. The majority of PLDs fall into a class known as C2, which contain a C2 (calcium- and lipid- binding) domain. Phospholipase C cleaves phospholipids generating diacylglycerol (DAG) and a phosphorylated head group. Plants possess two distinct PLC families: phosphoinositide-specific PLC (PtdIns-PLC) and nonspecific PLC (NPC). PtdIns-PLC facilitates the hydrolysis of phosphatidylinositol 4, 5-bisphosphate (PtdInP2) to generate 1, 4, 5-triphosphate (InsP3) and DAG. The resulting DAG remains bound to the membrane, while InsP3 is released into the cytosol to act as a mediator (Wang et al. 2012). NPC functions in cleaving other membrane phospholipids, such as PtdCho and PtdEtn. PhospholipaseAs hydrolyze phospholipids leading to the production of lysophospholipids and free fatty acids. PLA1 and PLA2 cleave the fatty acid from sn-1 and sn-2 positions, respectively. In plants, there are four families of PLAs: the PC-hydrolyzing PLA1 (PLA1), the PA-preferring PLA1 (PA-PLA1), the secretory low molecular weight PLA2 (sPLA2), and the patatin-like PLA (pPLA; Wang et al. 2012).
In our phloem sap analysis, we detected PtdChos, PtdIns, PtdOHs, TAGs, and DAGs. While all of those play a role in intracellular signaling, PtdCho and PtdIns are likely precursors to the signaling molecules PtdOH and inositol phosphates. Their role in intracellular signaling and its possible implications for long-distance signaling are discussed below:
Phosphatidyl Inositol and Its Phosphates
Early findings in animal systems had suggested the existence of a phosphoinositide (PtdIns)/phospholipase C (PLC) intracellular signaling system: upon receptor stimulation, PLC hydrolyses phosphatidyl inositol 4,5-bisphosphate into the second messengers diacylglycerol and inositol- 1,4,5-trisphosphate (InsP3) and leads to a downstream change in developmental processes. The scenario is different in plants due to their low PtdInsP2 concentration (Munnik and Testerink 2009). Plant PLCs will hydrolyze phosphatidylinositol-4-phosphate but can also use phosphatidylinositol (PtdIns) as a substrate (Munnik et al. 1998). In addition, so far no InsP3 receptor has been found. However, InsP3 is an intermediate in the lipid-mediated biosynthesis as well as in the new lipid-independent biosynthesis of InsP6, a known signaling molecule in plants (Gillaspy 2013).
Phosphoinositides , the phosphorylated versions of PtdIns play an important role in lipid signaling, membrane signaling (through regulating ion channels and pumps), and vesicle trafficking. They appear to be localized in discrete signaling regions within the membrane (Boss et al. 2006). The lipid-soluble phosphatidylinositol phosphates and water-soluble inositolphosphates (InsPs) are interrelated in that InsP3 and diacylglycerol (DAG) are produced from PtdInsP2 through the action of PLCs (Berridge 1993). Some PLCs have the capability to hydrolyze PtdIns(4)P or PtdIns (Munnik et al. 1998). DAG remains in the membrane where, in animals, it can activate a kinase C localized near the membrane and lead to the phosphorylation of target proteins (PKC; Newton 2010). However, so far, DAG could not be shown to activate any of the plant PKC homologues (Gillaspy 2013). In plants, the resulting DAG is further converted to phosphatidic acid (PtdOH; Arisz et al. 2009) while InsP3 can be increasingly phosphorylated up to InsP6 (hexakisphosphate). InsP6 is a high-energy storage compound in plants, but inositol polyphosphates can also function as a signaling molecule, regulate plant hormone receptors like those for IAA (Tan et al. 2007) and JA (Mosblech et al. 2011), and affect trafficking (Nagy et al. 2009). In addition, it can be converted to inositol pyrophosphates (InsP7, InsP8; Bennett et al. 2006; Burton et al. 2009). These high-energy molecules have been linked to energy sensing in yeast and animal systems. In plants, their role is yet to be determined. The phosphoinositide pathway and inositol-1,4,5-triphosphate (InsP3) were shown to be involved in the response to many abiotic stresses (Im et al. 2007), however, more recent findings suggest a role in the signaling of biotic stress as well (Hung et al. 2014): One of the early responses in both pathogen- and effector-triggered immunity (PTI and ETI, respectively) is a transient increase in cytosolic Ca2+, likely mediated by an influx through a plasma membrane localized Ca2+ -channel/pump or release of intracellular stores of Ca2+. This is likely due to the PLC-dependent generation of InsP3 and InsP6: Arabidopsis plants expressing the human type I inositol polyphosphate 5-phosphatase (InsP5-ptase), which specifically hydrolyzes soluble inositol phosphates and thus, interferes with InsP3-mediated signaling, exhibit reduced InsP3 and InsP6 levels as well as a reduced response to gravity and impaired Ca signaling following salt and cold-stress (Perera et al. 2008). Hung et al. (2014) further showed that InsP5-ptase expressing plants lacked the pathogen-induced Ca2+ increase in response to the elicitor flagellin 22 (flg22). Expression of pathogenesis related genes, SA levels and SAR in response to P. syringae were reduced as well. These findings suggest that reduced levels of InsP3 (or InsP6) lead to a reduced intracellular Ca2+ release and impaired downstream responses. Interestingly, both PtdIns, as well as one of the PLC-cleavage products, DAG, have been found in the phloem exudates of Arabidopsis (Guelette et al. 2012). While similar responses within the sieve elements or the companion cells are possible, so far, no experimental evidence points at their long-distance function in biotic stress.
The same holds true for the role of PtdInsPs in abiotic stress: drought, ABA, salt stress, cold, anoxia, light, or gravitropic signals all lead to the production of, albeit different, InsPs from PtdInsPs. Plants contain a variety of PLCs, all of which appear to be regulated by different environmental stimuli (Dowd and Gilroy 2010) and can be, at least in response to flg22, post-translationally modified. These increases in InsPs, however, may not solely be due to an increase in PLC activity, but can also be founded in the increase of PtdInsP2 production. Contrary to animals, plants have a lower PtdIns(4,5)P2 to PtdIns(4)P ratio (1:20 instead of 1:2). It has been suggested that PtdIns(4)P may have a dual function as membrane anchor for proteins as well as PtdIns(4,5)P2 precursor. As the former, they may anchor proteins to the membrane and thus remove them from their signaling path. In addition, PtdInsPs may function as second messengers themselves (see Gillaspy 2013).
A recent report suggested that the regulatory functions of PtdIns(4,5)P 2 in plant cells is dependent on the phosphatidylinositol 4-phosphate 5-kinases isoform which generates the lipid (PtdIns4P 5-kinases/PIP5K; Stenzel et al. 2012). In plants, PtdIns4P 5-kinases belong to a family of 11 proteins, which act in pairs of closely related proteins (Müller-Röber and Pical 2002). PtdIns(4,5)P2 is membrane localized and appears to play a role in vesicle trafficking. For example it is necessary for the polarized distribution of the PIN1 auxin efflux carrier and thus, indirectly for the directional auxin transport and gravitropism in root and shoot (Salinas-Mondragon et al. 2010; Thole and Nielsen 2008; Heilmann 2009). It was proposed that PIN cycling is impaired in Arabidopsis pi5k2 mutants possibly due to interference with clathrin-mediated endocytosis (König et al. 2008; Mei et al. 2012). Indeed, Ischebeck et al. (2013) showed that pip5K1/pip5K2 double mutants showed significantly reduced PtdIns(4,5)P2 levels, which was accompanied by an attenuated gravitropic response and reduced shoot and root growth. The lack of PtdIns(4,5)P2 in pip5K1/pip5K2 double mutants affected PIN2 localization through the perturbation of clathrin-mediated endocytic trafficking.
While PtdIns has been identified in phloem exudates, the methodology employed could not unambiguously identify phosphoinositides. Hence, any possible function in long-distance signaling may be indirect, for example through the enhancement of trafficking of signaling molecules to the sieve element or through the conversion of PtdIns to DAG to PtdOH as signaling molecules.
Phosphatidic Acid
Phosphatidic acid (PtdOH ) is an important intermediate in lipid biosynthesis, a membrane component, and a signaling molecule. As membrane component it may affect the membrane curvature, and, as a consequence, can regulate trafficking and membrane biogenesis (Kooijman et al. 2003; Wang 2004). In addition, it participates in signaling pathways, often by tethering components of these pathways to the membrane, thus alternating their location and function.
PtdOH is rapidly and transiently generated in response to several biotic and abiotic stresses, such as pathogen infection, drought, salinity, wounding, cold, cell death, and oxylipin production by either the PLC-DGK pathway or directly by PLD (Munnik and Testerink 2009; Xue et al. 2009; Wang et al. 2007; Hong et al. 2010; Kim et al. 2013; Testerink and Munnik 2011; Wang et al. 2014; Julkowska et al. 2015). PLD cleaves structural phospholipids such as phosphatidylcholine (PtdCho) and phosphatidylethanolamine (PtdEtn) in order to generate PtdOH and a free head group. PLC, on the other hand, hydrolyzes PtdIns(4,5)P2 into Ins(1,4,5)P3 and DAG (Julkowska et al. 2015). InsP3 diffuses into the cytosol where it triggers Ca2+ release from intracellular stores, while DAG remains within the membrane and is phosphorylated to PtdOH by DAG kinase (DGK) (Testerink and Munnik 2005, 2011).
Different stresses elicit different PtdOH production paths. For example, PtdOH produced in response to cold stress is generated via the PLC-DGK pathway. However, mutations in individual DGK genes did not affect this increase in PtdOH, likely due to redundancy (Arisz et al. 2013). Similarly, pathogens appear to induce the PLC-DGK pathway: an infection-mimicking treatment with xylanase inducers leads to an increased transcription of StPLC5 and StPLC2 in tomato cell suspension cultures (Gonorazky et al. 2014). On the other hand, the responses to drought, nutrient deficiency, and salt stress are largely mediated by PLD pathways (McLoughlin and Testerink 2013): PLDζ2 is involved in the response to salt and phosphate stress by not only affecting PtdOH production but PIN redistribution and, as a result, root curvature and root growth (Li and Xue 2007; Oropeza-Aburto et al. 2011; Galvan-Ampudia et al. 2013). PLDα1 and PLDδ mediate drought/ABA-induced and ROS-induced PtdOH production, respectively (Uraji et al. 2012). They appear to play a role in freezing tolerance as well (Welti et al. 2002). The drought response will be discussed further below.
PtdOH binds a myriad of proteins, including transcription factors, protein kinases, lipid kinases, protein phosphatases, as well as proteins involved in vesicular trafficking and cytoskeletal rearrangements (Guo et al. 2011). Examples of PtdOH -interacting proteins include abscisic acid insensitive 1 (ABI1), phosphoinositide-dependent protein kinase (PDK1), constitutive triple response 1 (CTR1), trigalactosyldiacylglycerol 2 (TGD2), and NADPH oxidase (Guo et al. 2011). When PtdOH binds to ABI1, CTR1, and phosphoethanolamine N-methyltransferase (PEAMT), it prevents their phosphatase or kinase activity. On the other hand, through direct interaction, PtdOH stimulates the catalytic activity of PDK1, NADPH oxidases (RESPIRATORY BURST OXIDASE HOMOLOG D/F; RbohD/F), sphingosine kinases (SPHK1/2), mitogen activated protein kinase 6 (MPK6), and SNF1-related kinase 2 and glyceraldehyde-3-phosphate dehydrogenase (G3PDH; SnRK2; Kim et al. 2013; McLoughlin and Testerink 2013).
In addition to regulating protein activity through their active site, PtdOH has been shown to tether some of these proteins to the membrane. A well-known example is the function of PtdOH in the response to drought/ABA-signaling cascade: In response to ABA, PtdOH is produced by PLDα1. This phospholipase, in addition to PLDδ, participates in mediating the plant’s response to ABA as well as promoting stomatal closure, although the two lipases regulate different areas of the signaling pathway. PLDδ facilitates ROS response, while PLDα1 acts upstream of PLDδ and promotes ROS production (Uraji et al. 2012; Lu et al. 2013). Both, PLDα1 and PLDδ positively regulate ABA-induced stomatal closure and are involved in ABA-induced ROS and NO production. It appears that PLDα1 functions mainly under moderate environmental conditions while PLDα1 and PLDδ, together work under more severe environmental stress conditions (Uraji et al. 2012). Under drought stress, ABA binds to the pyrabactin resistance/pyr1-like protein/regulatory components of its receptor (PYR/PYL/RCAR), which results the inhibition of ABI1, a type 2C phosphatase (PP2C) and a negative regulator, allowing SNF1-related kinase 2 (SnRK2) to be activated, which mediates downstream signaling such as the phosphorylation of transcription factors and ABA-induced gene expression (Julkowska et al. 2015; Dorosh et al. 2013). In turn, PtdOH is produced by PLDα1 and interacts with SPHK (sphingosine kinase). Increased phyto-S1P (a phosphorylated long-chain base-1-phosphate) activates more PLDα1, leading to an increase in PtdOH levels. PtdOH regulates proteins such as ABI1, NADPH oxidase, and ion channels to facilitate stomatal closure (Guo et al. 2012). As part of this response, PtdOH binds and tethers ABI1 to the plasma membrane which prevents ABI1 from translocating to the nucleus thus preventing its binding to ATHB6, a transcription factor that negatively regulates ABA responses (Klingler et al. 2010; Kim et al. 2013; Zhang et al. 2013). A recent study noted that under drought conditions, PtdOH binds the MYB transcription factor WEREWOLF, and may tether it to the nuclear membrane, which would allow for the localization of the proteins from the cytosol to the nucleus (Yao et al. 2013). This could aid in further transcriptional regulation of other drought-responsive genes.
In addition to its effect on stomatal closure, PtdOH is important for normal root development and thus normal water and nutrient uptake in plants: PLDα knock-down mutants display shorter roots and reduced lateral roots (pldα3) as well as an altered PtdOH accumulation in response to water deficit and ABA (pldα1 and pldα3; Wang 2005; Singh et al. 2012). This suggests that under standard conditions, PtdOH regulates normal root development, possibly via the action of PIN proteins (Gao et al. 2013).
Outlook
So far, the described role of PtdOH in stress responses has solely been of intracellular nature. Recently, we have been able to show that PtdOH is present in phloem exudates (Guelette et al. 2012). Its function there could be to tether a signaling protein to the membrane and thus change its activity, it could tether a receptor to the membrane, or it could participate in the movement of essential components of ABA/drought-related signaling from the companion cell into the sieve element. In addition, it could be part of the signal itself, similar to the Wnt-palmitoleic acid interaction (Janda et al. 2012). We have identified at least one phloem-localized PtdOH-binding protein, PLAFP (phloem-localized lipid-associated protein), a 20 kDa uncharacterized protein with a PLAT/LH2 domain. In humans and animals, PLAT domains (Polycystin-1, Lipoxygenase, Alpha-Toxin-domains) are found in a variety of lipid- or membrane-associated proteins. The domain contains eight beta-strands and forms a beta-sandwich composed of two sheets of four strands each. While structurally similar, the PLAT domain of a group of PLAFPs is very different from that of humans: the plant-specific PLAT_plant_stress domain (cd01754) is loosely related but not identical to lipase and lipoxygenase domains. Homology to human lipoxygenases is less than 30 %. Plant proteins with this domain, on the contrary, are highly conserved in different plant species. Most importantly, these PLAFP proteins do not contain the lipoxygenase catalytic domain but only their PLAT domain with a role in protein-protein interaction. Contrary to larger, usually polymeric lipoxygenases, their size of only 20 kDa would allow for movement through plasmodesmata and through the phloem. Thus, PLAFP could function in the transport of lipid-signals into, out, or through the phloem sieve element, the detection of lipid-signals, or be part of a signal itself. PtdOH is an important signaling compound, yet the characteristics of the PtdOH-binding site in proteins remain elusive. SnRK2.4, a known PtdOH-binding protein, contains a 42-amino acid PtdOH-binding domain with five conserved basic amino acids (Julkowska et al. 2015). These conserved amino acids are essential for PtdOH binding but not sufficient to mediate salt-induced re-localization of the protein. Hence, additional factors may be necessary. If PtdOH anchors a protein to the membrane, a conserved hydrophilic pocket with basic amino acids as found in SnrRK.4 could interact with the head group while the remainder of the lipid remains integrated in the membrane. However, if the PtdOH-binding specificity extends beyond the binding of the head group and protein–lipid interaction is also based on the characteristics of the acyl groups, it would suggest that the protein binds the entire lipid, in which case the latter may not remain integrated in the membrane but instead is embedded in a hydrophobic groove on the surface of the protein . This is particularly interesting in the context of findings by other labs that different stresses are involving distinct PtdOH species (Guo et al. 2012). A dynamic modeling approach (Visual Molecular Dynamics – VMD; http://www.ks.uiuc.edu/Research/vmd/) using a ligand obtained from the RCSB-protein data bank (http://www.rcsb.org/; (R)-2-(formyloxy)-3-(phosphonooxy) propyl pentanoate) as approximation for an, albeit short-chain-, PtdOH suggested the presence of a hydrophobic grove on the surface of PLAFP bordered by amino acids with basic side chains (Fig. 14.2). These features would facilitate not only the interaction of the protein with the phosphatidic acid head group but with the acyl chains as well, suggesting interaction with the entire length of the molecule and thus possible PtdOH movement away from the membrane and intracellular or long-distance transport. While this model, at this point, is entirely speculative, it provides further avenues for studying possible mechanisms of phospholipid movement between membranes and throughout the plant.
Predicted ligand binding to PLAFP using vmd. C5-PA was used as a ligand. Glycerol backbone and phosphate headgroup are adjacent to hydrophilic (green) and basic (blue) amino acid side chains, respectively. The acyl groups extend into a hydrophobic area (grey) of the protein. A longer, monounsaturated acyl group could extend into the hydrophobic region at the top of the protein.
References
Andrawis A, Solomon M, Delmer DP (1993) Cotton fiber annexins – a potential role in the regulation of callose synthase. Plant J 3:763–772
Arisz SA, Testerink C, Munnik T (2009) Plant PA signaling via diacylglycerol kinase. Biochim Biophys Acta 1791(9):869–875
Arisz SA, van Wijk R et al (2013) Rapid phosphatidic acid accumulation in response to low temperature stress in Arabidopsis is generated through diacylglycerol kinase. Front Plant Sci 4:1. doi:10.3389/fpls.2013.00001. eCollection 2013
Ayre BG, Turgeon R (2004) Graft transmission of a floral stimulant derived from CONSTANS. Plant Physiol 135:2271–2278
Barnes A, Bale J et al (2004) Determining protein identity from sieve element sap in Ricinus communis L. by quadrupole time of flight (Q-TOF) mass spectrometry. J Exp Bot 55:1473–1481
Bateman A, Sandford R (1999) The PLAT domain: a new piece in the PKD1 puzzle. Curr Biol 9:R588–R590. doi:10.1016/S0960-9822(99)80380-7
Behmer ST, Grebenok RJ, Douglas AE (2010) Plant sterols and host plant suitability for a phloem feeding insect. Funct Ecol 25:484–491
Bennett M, Onnebo SM et al (2006) Inositol pyrophosphates: metabolism and signaling. Cell Mol Life Sci 63:552–564
Benning UF, Tamot B et al (2012) New aspects of phloem-mediated long-distance lipid signaling in plants. Front Plant Sci 3:53. doi:10.3389/fpls.2012.0053
Berridge MJ (1993) Inositol trisphosphate and calcium signaling. Nature 361:315–325
Blaner WS (1989) Retinol-binding protein: the serum transport protein for vitamin A. Endocr Rev 10:308–316
Boss WF, Davis AJ et al (2006) Phosphoinositide metabolism: towards an understanding of subcellular signaling. Subcell Biochem 39:181–205
Burton A, Hu X, Saiardi A (2009) Are inositol pyrophosphates signalling molecules? J Cell Physiol 220:8–15
Buseman CM, Tamura P et al (2006) Wounding stimulates the accumulation of glycerolipids containing oxophytodienoic acid and dinor-oxophytodienoic acid in Arabidopsis leaves. Plant Physiol 142:28–39
Champigny MJ, Isaacs M et al (2013) Long distance movement of DIR1 and investigation of the role of DIR1-like during systemic acquired resistance in Arabidopsis. Front Plant Sci 4:230
Chanda B, Xia Y et al (2011) Glycerol-3-phosphate is a critical mobile inducer of systemic immunity in plants. Nat Genet 43:421–427. doi:10.1038/ng.798. Epub 2011 Mar 27
Charbonneau D, Beauregard M, Tajmir-Riahi HA (2009) Structural analysis of human serum albumin complexes with cationic lipids. J Phys Chem B 113:1777–1781
Chaturvedi R, Shah J (2007) Salicylic acid in plant disease resistance. In: Hayat S, Ahmad A (eds) Salicylic acid – a plant hormone. Springer, Dordrecht, pp 335–370
Chaturvedi R, Krothapalli K et al (2008) Plastid x-3 desaturase-dependent accumulation of a systemic acquired resistance inducing activity in petiole exudates of Arabidopsis thaliana is independent of jasmonic acid. Plant J 54:106–117
Chaturvedi R, Venables B et al (2012) An abietane diterpenoid is a potent activator of systemic acquired resistance. Plant J 71:161–172
Chehab EW, Kaspi R et al (2008) Distinct roles of jasmonates and aldehydes in plant-defense responses. PLoS One 3, e1904
Chen S, Petersen BL et al (2001) Long-distance phloem transport of glucosinolates in Arabidopsis. Plant Physiol 127:194–201
Chen QF, Xiao S, Chye ML (2008) Overexpression of the Arabidopsis 10-kilodalton acyl-coenzyme A-binding protein ACBP6 enhances freezing tolerance. Plant Physiol 148:304–315
Chiou TJ, Lin SI (2011) Signaling network in sensing phosphate availability in plants. Annu Rev Plant Biol 62:185–206
Citovsky V, Zambryski P (2000) Systemic transport of RNA in plants. Trends Plant Sci 5:52
Corbesier L, Prinsen E et al (2003) Cytokinin levels in leaves, leaf exudate and shoot apical meristem of Arabidopsis thaliana during floral transition. J Exp Bot 54:2511–2517
Corbesier L, Vincent C et al (2007) FT protein movement contributes to long-distance signaling in floral induction of Arabidopsis. Science 316:1030–1033
Dave A, Graham IA (2012) Oxylipin signalling: a distinct role for the Jasmonic acid precursor 12-oxo-phytodienoic acid (OPDA). Front Plant Sci 3:42
Dave A, Hernández ML et al (2011) 12-Oxo-phytodienoicacid accumulation during seed development represses seed germination in Arabidopsis. Plant Cell 23:583–599
Ding B, Itaya A, Qi YJ (2003) Symplasmic protein and RNA traffic: regulatory points and regulatory factors. Curr Opin Plant Biol 6:596–602
Dorosh L, Kharenko OA et al (2013) Molecular mechanisms in the activation of abscisic acid receptor PYR1. PLoS Comput Biol 9:e1003114. doi:10.1371/journal.pcbi.1003114. Epub 2013 Jun 27
Dowd PE, Gilroy S (2010) The emerging roles of phospholipase C in plant growth and development. In: Munnik T (ed) Lipid signaling in plants. Springer, Berlin
Feussner I, Wasternack C (2002) The lipoxygenase pathway. Annu Rev Plant Biol 53:275–297
Fisher DB, Wu Y, Ku MSB (1992) Turnover of soluble-proteins in the wheat sieve tube. Plant Physiol 100:1433–1441
Froehlich DR, Mullendore DL et al (2011) Phloem ultrastructure and pressure flow: sieve-element-occlusion-related agglomerations do not affect translocation. Plant Cell 23:4428–4445
Galvan-Ampudia CS, Julkowska MM et al (2013) Halotropism is a response of plant roots to avoid a saline environment. Curr Biol 23:2044–2050. doi:10.1016/j.cub.2013.08.042. Epub 2013 Oct 3
Gao HB, Chu YJ, Xue HW (2013) Phosphatidic acid (PA) binds PP2AA1 to regulate PP2A activity and PIN1 polar localization. Mol Plant 6(5):1692–1702. doi:10.1093/mp/sst076. Epub 2013 May 17
Giavalisco P, Kapitza K et al (2006) Towards the proteome of Brassica napus phloem sap. Proteomics 6:896–909
Gillaspy GE (2013) The role of phosphoinositides and inositol phosphates in plant cell signaling. In: Capelluto GS (ed) Lipid-mediated protein signaling. Springer, Dordrecht, pp 141–157
Glatz JFC, Boerchers T et al (1995) Fatty acids in cell signalling: modulation by lipid binding proteins. Prostaglandins Leukot Essent Fatty Acids 52:121
Gonorazky G, Ramirez L et al (2014) The tomato phosphatidylinositol-phospholipase C2 (SlPLC2) is required for defense gene induction by the fungal elicitor xylanase. J Plant Physiol 171:959–965. doi:10.1016/j.jplph.2014.02.008. Epub 2014 Mar 12
Guelette BS, Benning UF, Hoffmann-Benning S (2012) Identification of lipids and lipid-binding proteins in phloem exudates from Arabidopsis thaliana. J Exp Bot 63:3603–3616. doi:10.1093/jxb/ers028
Guo L, Mishra G, Taylor K, Wang X (2011) Phosphatidic acid binds and stimulates Arabidopsis sphingosine kinases. J Biol Chem 286(15):13336–13345
Guo L, Mishra G et al (2012) Connections between sphingosine kinase and phospholipase D in the abscisic acid signaling pathway in Arabidopsis. J Biol Chem 287:8286–8296
Haebel S, Kehr J (2001) Matrix-assisted laser desorption/ionization time of flight mass spectrometry peptide mass fingerprints and post source decay: a tool for the identification and analysis of phloem proteins from Cucurbita maxima Duch. separated by two-dimensional polyacrylamide gel electrophoresis. Planta 214:332
Hayashi H, Fukuda A et al (2000) Proteins in the sieve element-companion cell complexes: their detection, localization and possible functions. Aust J Plant Physiol 27:489–496
Haywood V, Yu TS et al (2005) Phloem long-distance trafficking of gibberellic acid-insensitive RNA regulates leaf development. Plant J 42:49–68
Heilmann I (2009) Using genetic tools to understand plant phosphoinositide signalling. Trends Plant Sci 14:171–179
Hoad GV (1995) Transport of hormones in the phloem of higher plants. Plant Growth Reg 16:173–182
Hoffmann-Benning S, Gage DA et al (2002) Comparison of peptides in the phloem sap of flowering and non-flowering Perilla and lupine plants using microbore HPLC followed by matrix-assisted laser desorption/ionization time-of-flight mass spectrometry. Planta 216:140–147
Hoffmann-Benning S (2015) https://bmb.natsci.msu.edu/faculty/susanne-hoffmann-benning-assistant-professor/current-research
Hong Y, Zhang W, Wang X (2010) Phospholipase D and phosphatidic acid signaling in plant response to drought and salinity. Plant Cell Environ 33:627–635
Howe GA, Schilmiller AL (2002) Oxylipin metabolism in response to stress. Curr Opin Plant Biol 5:230–236
Hung CY, Aspesi P Jr et al (2014) Phosphoinositide-signaling is one component of a robust plant defense response. Front Plant Sci 5:267. doi:10.3389/fpls.2014.00267
Im YJ, Perera IY et al (2007) Increasing plasma membrane phosphatidylinositol (4,5)bisphosphate biosynthesis increases phosphoinositide metabolism in Nicotiana tabacum. Plant Cell 19:1603–1616
Ischebeck T, Werner S et al (2013) Phosphatidylinositol 4,5-bisphosphate influences PIN polarization by controlling clathrin-mediated membrane trafficking in Arabidopsis. Plant Cell 25:4894–4911. doi:10.1105/tpc.113.116582. Epub 2013 Dec 10
Janda CY, Waghray D et al (2012) Structural basis of Wnt recognition by frizzled. Science 337:59–64
Julkowska MM, McLoughlin F et al (2015) Identification and functional characterization of the Arabidopsis Snf1-related protein kinase SnRK2.4 phosphatidic acid-binding domain. Plant Cell Environ 38:614–624. doi:10.1111/pce.12421. Epub 2014 Sep 1
Jung HW, Tschaplinski TJ et al (2009) Priming in systemic plant immunity. Science 324:89–91
Kehr J, Haebel S et al (1999) Analysis of phloem protein patterns from different organs of Cucurbita maxima Duch. by matrix-assisted laser desorption/ionization time of flight mass spectroscopy combined with sodium dodecyl sulfate-polyacrylamide gel electrophoresis. Planta 207:612–619
Kim SC, Guo L, Wang X (2013) Phosphatidic acid binds to cytosolic glyceraldehyde-3-phosphate dehydrogenase and promotes its cleavage in Arabidopsis. J Biol Chem 288:11834–11844. doi:10.1074/jbc.M112.427229
Klingler JP, Batelli G, Zhu J-K (2010) ABA receptors: the START of a new paradigm in phytohormone signaling. J Exp Bot 61:3199–3210
König S, Ischebeck T et al (2008) Salt-stress-induced association of phosphatidylinositol 4,5-bisphosphate with clathrin-coated vesicles in plants. Biochem J 415:387–399
Kooijman EE, Chupin V et al (2003) Modulation of membrane curvature by phosphatidic acid and lysophosphatidic acid. Traffic 4:162–174
Kühn C, Franceschi VR et al (1997) Macromolecular trafficking indicated by localization and turnover of sucrose transporters in enucleate sieve elements. Science 275:1298–1300
Lascombe MB, Bakan B et al (2008) The structure of “defective in induced resistance” protein of Arabidopsis thaliana, DIR1, reveals a new type of lipid transfer protein. Protein Sci 17:1522–1530
Li G, Xue HW (2007) Arabidopsis PLD zeta2 regulates vesicle trafficking and is required for auxin response. Plant Cell 19:281–295. doi:10.1105/tpc.106.041426
Lin MK, Belanger H et al (2007) Flowering locus T protein may act as the long-distance florigenic signal in the cucurbits. Plant Cell 19:1488–1506
Lin MK, Lee YJ et al (2009) Analysis of the pumpkin phloem proteome provides functional insights into angiosperm sieve tube function. Mol Cell Proteomics 8:343–356
López-Bucio J, Hernández-Abreu E et al (2002) Phosphate availability alters architecture and causes changes in hormone sensitivity in the Arabidopsis root system. Plant Physiol 129:244–256
Lu S, Bahn SC et al (2013) Increased expression of phospholipase Dα1 in guard cells decreases water loss with improved seed production under drought in Brassica napus. Plant Biotechnol J 11:380–389
Lucas WJ, Groover A et al (2013) The plant vascular system: evolution, development and functions. J Integr Plant Biol 55:294–388. doi:10.1111/jipb.12041
Luna E, Ton J (2012) The epigenetic machinery controlling transgenerational systemic acquired resistance. Plant Signal Behav 7:615–618
Madey E, Nowack LM, Thompson JE (2002) Isolation and characterization of lipid in phloem sap of canola. Planta 214:625–634
Malamy J, Carr JP et al (1990) Salicylic acid: a likely endogenous signal in the resistance response of tobacco to viral infection. Science 250:1002–1004
Maldonado AM, Doerner P et al (2002) A putative lipid transfer protein involved in systemic acquired resistance signaling in Arabidopsis. Nature 419:399–403
Mandal M, Chanda B et al (2011) Glycerol-3-phosphate and systemic immunity. Plant Signal Behav 6:1871–1874
Marentes E, Grusak MA (1998) Mass determination of low-molecular-weight proteins in phloem sap using matrix-assisted laser desorption/ionization time-of-flight mass spectrometry. J Exp Bot 49:903–911
McLoughlin F, Testerink C (2013) Phosphatidic acid, a versatile water-stress signal in roots. Front Plant Sci 4:525. doi:10.3389/fpls.2013.00525
Mei Y, Jia WJ et al (2012) Arabidopsis phosphatidylinositol monophosphate 5-kinase 2 is involved in root gravitropism through regulation of polar auxin transport by affecting the cycling of PIN proteins. Cell Res 22:581–597
Métraux JP, Signer H et al (1990) Increase in salicylic acid at the onset of systemic acquired resistance in cucumber. Science 250:1004–1006
Mitton F, Pinedo ML, de la Canal L (2009) Phloem sap of tomato plants contains a DIR1 putative ortholog. J Plant Physiol 166:543–547
Mortimer JC, Laohavisit A et al (2008) Annexins: multifunctional components of growth and adaptation. J Exp Bot 59:533–544
Mosblech A, Thurow C et al (2011) Jasmonic acid perception by COI1 involves inositol polyphosphates in Arabidopsis thaliana. Plant J 65(6):949–957
Mueller-Roeber B, Pical C (2002) Inositol phospholipid metabolism in Arabidopsis. Characterized and putative isoforms of inositol phospholipid kinase and phosphoinositide-specific phospholipase C. Plant Physiol 130:22–46
Münch E (1930) Material flow in plants. Translated 2003 by Milburn JA and Kreeb KH, University of Bremen, Germany. Gustav Fischer Verlag, Jena
Munnik T, Testerink C (2009) Plant phospholipid signaling: “in a nutshell”. J Lipid Res 50:S.260–S265
Munnik T, Irvine RF, Musgrave A (1998) Phospholipid signalling in plants. Biochim Biophys Acta 1389:222–272
Musille PM, Kohn JA, Ortlund EA (2013) Phospholipid-driven gene regulation. FEBS Lett 587(8):1238–1246
Mustroph A, Zanetti ME et al (2009) Profiling translatomes of discrete cell populations resolves altered cellular priorities during hypoxia in Arabidopsis. Proc Natl Acad Sci U S A 106:18843–18848
Nagy L, Szanto A (2005) Roles for lipid activated transcription factors. Mol Nutr Food Res 49:1072–1074
Nagy R, Grob H et al (2009) The Arabidopsis ATP-binding cassette protein AtMRP5/AtABCC5 is a high affinity inositol hexakisphosphate transporter involved in guard cell signaling and phytate storage. J Biol Chem 284:33614–33622
Nakamura Y, Andrés F et al (2014) Arabidopsis florigen FT binds to diurnally oscillating phospholipids that accelerate flowering. Nat Commun 5:3553
Newton AC (2010) Protein kinase C: poised to signal. Am J Physiol Endocrinol Metab 298(3):E395–E402
Ohlrogge J, Browse J (1995) Lipid biosynthesis. Plant Cell 7:957–970
Oropeza-Aburto A, Cruz-Ramirez A et al (2011) Functional analysis of the Arabidopsis PLDZ2 promoter reveals an evolutionarily conserved low-Pi- responsive transcriptional enhancer element. J Exp Bot 63:2189–2202. doi:10.1093/jxb/err446
Park SW, Kaimoyo E et al (2007) Methyl salicylate is a critical mobile signal for plant systemic acquired resistance. Science 318:113–116
Perera IY, Hung CY et al (2008) Transgenic Arabidopsis plants expressing the type1 inositol 5-phosphatase exhibit increased drought tolerance and altered abscisic acid signaling. Plant Cell 20:2876–2893. doi:10.1105/tpc.108.061374
Raskin I (1992) Role of salicylic acid in plants. Annu Rev Plant Physiol Plant Mol Biol 43:439–463
Rescher U, Gerke V (2004) Annexins – unique membrane binding proteins with diverse functions. J Cell Sci 117:2631–2639
Ruiz-Medrano R, Xoconostle-Cázares B, Lucas WJ (1999) Phloem long-distance transport of CmNACP mRNA: implications for supracellular regulation in plants. Development 126:4405–4419
Salinas-Mondragon RE, Kajla JD et al (2010) Role of inositol 1,4,5-trisphosphate signaling in gravitropic and phototropic gene expression. Plant Cell Envrion 33:2041–2055
Savchenko T, Kolla VA et al (2014) Functional convergence of oxylipin and abscisic acid pathways controls stomatal closure in response to drought. Plant Physiol 164:1151–1160
Schilmiller AL, Howe GA (2005) Systemic signaling in the wound response. Curr Opin Plant Biol 8:369–377
Shah J, Chaturvedi R et al (2014) Signaling by small metabolites in systemic acquired resistance. Plant J 79:645–658
Singh A, Pandey et al (2012) Comprehensive expression analysis of rice phospholipase D gene family during abiotic stresses and development. Plant Signal Behav 7:847–855
Stenzel I, Ischebeck T et al (2012) Variable regions of PI4P 5-kinases direct PtdIns(4,5)P2 toward alternative regulatory functions in tobacco pollen tubes. Front Plant Sci 2:114
Tamaki S, Matsuo S et al (2007) Hd3a protein is a mobile flowering signal in rice. Science 316:1033–1036
Tan X, Calderon-Villalobos LI et al (2007) Mechanism of auxin perception by the TIR1 ubiquitin ligase. Nature 446:640–645
Taoka K et al (2011) 14-3-3 proteins act as intracellular receptors for rice Hd3a florigen. Nature 476:332–335
Testerink C, Munnik T (2005) Phosphatidic acid: a multifunctional stress signaling lipid in plants. Trends Plant Sci 10:368–375
Testerink C, Munnik T (2011) Molecular, cellular, and physiological responses to phosphatidic acid formation in plants. J Exp Bot 62(7):2349–2361
Tetyuk O, Benning UF, Hoffmann-Benning S (2013) Collection and analysis of Arabidopsis phloem exudates using the EDTA-facilitated method. J Vis Exp 80, e51111. doi:10.3791/51111. Movie and Text
Thole JM, Nielsen E (2008) Phosphoinositides in plants: novel functions in membrane trafficking. Curr Opin Plant Biol 11:620–631
Thomas B, Vince-Prue D (1997) Photoperiodism in plants. Academic, London
Thomas H, Stoddard JL (1980) Leaf senescence. Annu Rev Plant Physiol 31:83–111. doi:10.1146/annurev.pp.31.060180.000503
Thorpe MR, Ferrieri AP et al (2007) (11)C-imaging: methyl jasmonate moves in both phloem and xylem, promotes transport of jasmonate, and of photoassimilate even after proton transport is decoupled. Planta 226:541–551
Tontonez P, Hu E, Spiegelman BM (1994) Stimulation of adipogenesis in fibroblasts by PPARg2, a lipid-activated transcription factor. Cell 79:1147–1156
Truman W, Bennett MH et al (2007) Arabidopsis systemic immunity uses conserved defense signaling pathways and is mediated by jasmonates. Proc Natl Acad Sci U S A 104:1075–1080
Truman W, Bennett MH et al (2010) Arabidopsis auxin mutants are compromised in systemic acquired resistance and exhibit aberrant accumulation of various indolic compounds. Plant Physiol 152:1562–1573
Turgeon R, Wolfe S (2009) Phloem transport: cellular pathways, molecular trafficking. Annu Rev Plant Biol 60:207–221
Tuzun S, Kuc J (1985) Movement of a factor in tobacco infected with Peranospora tabacina Adam which systemically protects against blue mold. Physiol Plant Pathol 26:321–330
Tzen JTC, Huang AHC (1992) Surface structure and properties of plant seed oil bodies. J Cell Biol 117:327–335
Uraji M, Katagiri T et al (2012) Cooperative function of PLD delta and PLD alpha1 in ABA-induced stomatal closure in Arabidopsis. Plant Physiol 159:450–460. doi:10.1104/pp. 112.195578
van Bel AJE, Knoblauch M (2000) Sieve element and companion cell: the story of the comatose patient and the hyperactive nurse. Aust J Plant Physiol 27:477–487
Wang X (2004) Lipid signaling. Curr Opin Plant Biol 7:329–336
Wang X (2005) Regulatory functions of phospholipase D and phosphatidic acid in plant growth, development, and stress responses. Plant Physiol 139(2):566–573
Wang X, Zhang W, et al (2007) Phospholipid signaling in plant response to drought and salt stress. In Jenks MA et al (eds) Advances in molecular breeding toward drought and salt tolerant crops. Springer, Netherlands, pp 183–192
Wang G, Ryu S, Wang X (2012) Plant phospholipases: an overview. Methods Mol Biol 861:123–137
Wang X, Chapman KD (2013) Lipid signaling in plants. Front Plant Sci 4:1–2
Wang X, Guo L, Wang G, Li M (2014) PLD: phospholipase Ds in plant signaling phospholipases in plant signaling. Springer-Verlag Berlin Heidelberg, pp 3–26
Welti R, Li W et al (2002) Profiling membrane lipids in plant stress responses. Role of phospholipase D alpha in freezing-induced lipid changes in Arabidopsis. J Biol Chem 277:31994–32002. doi:10.1074/jbc.M205375200
Wu X, Dinneny JR et al (2003) Modes of intercellular transcription factor movement in the Arabidopsis apex. Development 130:3735–3745
Xoconostle-Cazares B, Yu X et al (1999) Plant paralog to viral movement protein that potentiates transport of mRNA into the phloem. Science 283:94–98
Xue H-W, Chen X, Mei Y (2009) Function and regulation of phospholipid signalling in plants. Biochem J 421:145–156
Yan Y, Christensen S et al (2012) Disruption of OPR7 and OPR8 reveals the versatile functions of JA in maize development and defense. Plant Cell 24:1420–1436
Yan Y, Borrego E, Kolomiets MV (2013) Jasmonate biosynthesis, perception and function in plant development and stress responses. In: Baez RV (ed) Lipid metabolism. Intech, pp 393–442. http://dx.doi.org/10.5772/52675
Yao H, Wang G et al (2013) Phosphatidic acid interacts with a MYB transcription factor and regulates its nuclear localization and function in Arabidopsis. The Plant Cell 25:5030–5042
Yoo BC, Kragler F et al (2004) A systemic small RNA signaling system in plants. Plant Cell 16:1979–2000
Yu K, Soares JM et al (2013) A feedback regulatory loop between G3P and lipid transfer proteins DIR1 and AZI1 mediates azelaic-acid-induced systemic immunity. Cell Rep 3:1266–1278
Zeevaart JAD (1976) Physiology of flower formation. Annu Rev Plant Physiol Plant Mol Biol 27:321–348
Zhang X, Jiang L et al (2013) Structural insights into the abscisic acid stereospecificity by the ABA receptors PYR/PYL/RCAR. PLoS One 8, e67477
Zheng SX, Xiao S, Chye ML (2012) The gene encoding Arabidopsis Acyl-CoA-binding protein 3 is pathogen-inducible and subject to circadian regulation. J Exp Bot 63:2985–3000
Zhu J-K (2002) Salt and drought stress signal transduction in plants. Annu Rev Plant Biol 53:247–273
Acknowledgements
The model in Fig. 14.2 was generated by Michael O’Keefe with the help of Michael Feig. This work was funded by NSF grant #1144391 and USDA-NIFA hatch project # MICL02233 to SHB. Lipid structures were obtained from Wikipedia.org, lipidlibrary.aocs.org, www.bio-protocol.org.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Barbaglia, A.M., Hoffmann-Benning, S. (2016). Long-Distance Lipid Signaling and its Role in Plant Development and Stress Response. In: Nakamura, Y., Li-Beisson, Y. (eds) Lipids in Plant and Algae Development. Subcellular Biochemistry, vol 86. Springer, Cham. https://doi.org/10.1007/978-3-319-25979-6_14
Download citation
DOI: https://doi.org/10.1007/978-3-319-25979-6_14
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-25977-2
Online ISBN: 978-3-319-25979-6
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)