Abstract
Among the insects, lepidopterans form the second most diverse group, with over 155,000 described species. Research on Lepidoptera has a long tradition in several fields, including taxonomy, phylogeny, ecology, population genetics, evolutionary biology, speciation, physiology, development and gene regulation, host–plant and insect–parasite interactions, and, in recent decades, genomics. These studies and genomic resources for them are widely distributed and often widespread in various databases. In this chapter, we analyze the state of the art for genomic resources for Lepidoptera in GenBank for the following genes: elongation factor-1α, wingless, cytochrome c oxidase I, ribosomal DNA and RNA, and in general a number of other protein and enzyme entries; complete mitochondrial genomes; complete nuclear genomes; and published work on barcode methodology. This information will help researchers find gaps in the available resources and direct research efforts in these areas.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
3.1 Introduction: Why Butterflies and Moths?
Lepidoptera is one of the largest groups of organisms in the world. This order comprises insects commonly known as butterflies and moths. Historically, the former have attracted the attention of professional and amateur entomologists, as well as the general public because of the beautiful colors and patterns present in their scaled wings. The moths are studied primarily not only because many species are economically important pests of agriculture and forestry but also for silk production, with the mulberry silkworm, Bombyx mori, considered one of the few “domesticated” insects [1], reared at least since 2600 BC [2].
The origin of the holometabolous order Lepidoptera is dated to the Late Carboniferous, but diversification occurred in the Early Cretaceous at the same time as the radiation of flowering plants [3]. Currently, the order Lepidoptera contains over 157,424 species including approximately 22 fossils; the living species (157,402) are classified into 45 superfamilies, 134 families, and 15,562 genera [4]. This is the second most diverse group of animals after Coleoptera.
Insects have long been used as model systems, and the fruit fly, Drosophila melanogaster, was the first choice historically, primarily because of its short life cycle and ease of rearing in the laboratory [5]. Nevertheless, the importance of model systems is that discoveries and implications can be extended far beyond the particular organism under study [6]. Certain phenomena such as evolution, coevolution, and biogeographic and ecological mechanisms are better documented and explained within Lepidoptera because there is significant background in the knowledge of this group, mostly due to its economic importance and attractiveness. This gives an advantage to Lepidoptera, as they are better known in many aspects than other diverse groups, and their genomic research will help to understand different kinds of processes.
3.2 GenBank Database: Lepidoptera Representation
In 1982, GenBank was officially released; by 1992, the National Center for Biotechnology Information (NCBI), which is part of the International Nucleotide Sequence Database Collaboration (INSDC), took responsibility for it. From August 2011 to 2012, GenBank had an annual increase in records of 33.1 %, but invertebrates had a decrease of 1.7 % in the same year. The GenBank Dataset is divided into two main groups, taxonomic and functional. The functional division in GenBank sequences makes the data easy to handle and reflects the methods used to obtain it [7]. Functional divisions in 2012 included transcriptome shotgun data, whole-genome shotgun (WGS) data, patented sequences, genome survey sequences, expressed sequence tags (ESTs), high-throughput genomics, sequence tagged sites, and high-throughput complementary DNA (cDNA). Transcriptome shotgun data was the fastest growing division, with more than 200 % growth that year [7]. The taxonomic division, GenBank Dataset, was useful only to know the species of Lepidoptera reported in GenBank. A search in GenBank with “Lepidoptera” on April 2, 2014, returned 1,093,006 sequences; 57,906 registers of these were not identified, yielding 1,035,100 sequences representing a comprehensive landscape of Lepidoptera genomics. According to a recent classification of Lepidoptera [4], 92 % of the 134 living families are represented in GenBank with at least one sequence (Fig. 3.1a), and only 10 families are not present (Anomosetidae, Schistonoeidae, Syringopaidae, Coelopoetidae, Epimarptidae, Whalleyanidae, Simaethistidae, Ratardidae, Peleopodidae, and Metarbelidae). As we go to lower taxonomic categories, the representation in GenBank is reduced to 41 % at the genus level (Fig. 3.1b) and only 13 % at the species level (Fig. 3.1c). Additionally, there is a dissimilarity in the proportion of representation of genera and species from different families or what Wilson [8] observed as uneven taxonomic distribution. Almost 20 % of the families have all their genera represented in GenBank (26 families with 100 %, Fig. 3.2a), including two butterfly families and the rest moths (Table 3.1). Nearly 20 % of the families have less than 20 % of their genera represented. At the species level, representation is very low, with just two families, Carthaeidae and Prodidactidae (Table 3.2), with 100 % representation for only one species each (Carthaea saturnioides and Prodidactis mystica, respectively), and 65 % of the families with less than 10 % of the species represented (Fig. 3.2b). The family Prodoxidae has proper representation with nearly 80 % of the species, and seven families are 56.8 % represented (Sphingidae, Aididae, Papilionidae, Agathiphagidae, Heterogynidae, Lophocoronidae, and Millieriidae) (Table 3.2).
Records of Lepidoptera in GenBank by taxonomic level. (a) Comparison between the number of families of Lepidoptera reported by Nieukerken et al. [4] and the families in GenBank as of April 2014. The data represent 92 % of the families of the order. (b) Representation at the genus level: only 41 % of the group is represented. (c) Representation at the species level: only 13 % of all species of Lepidoptera are represented in GenBank
In total, 124 families, 6336 genera, and 20,076 species of Lepidoptera are represented in GenBank; but a key question is, what functional sequences are documented for each one? We will present information on this using some well-represented sequences for Lepidoptera as a whole.
3.3 Global Lepidoptera Sequences
There are several uses for DNA sequences, such as phylogenetic studies, pest control applications, and analysis of evolutionary changes at the species level and even in particular gene families. Targets of analysis depend on the aims of the research. For instance, different regions of mitochondrial DNA such as cytochrome oxidase subunits I, II, and III (COI, COII, COIII), cytochrome b (cyt b), or nuclear DNA sequences, e.g., ribosomal RNA (rRNA), ribosomal DNA (rDNA), satellite DNA, introns, and nuclear protein-coding genes, can be used to delimit species, phylogeny, or functional genetics [9].
Knowing the nature of the DNA can provide new insights into the biology of this order. The most represented Lepidoptera genes in GenBank are elongation factor-1α (EF), wingless (Wg), rRNA, rDNA, COI, and selected proteins. In this chapter, proteins with a catalytic function are classified as enzymes and the rest remain as proteins.
3.4 Elongation Factor-1α
EF is a slowly evolving nuclear gene which is involved in the production of proteins, operating at the receptor site of the ribosome during the translation process [10]. In insects, when used in combination with mitochondrial genes [11, 12], it results in good resolution of high-level phylogenetic relationships, particularly in Lepidoptera [13–19]. Wahlberg et al. [20] resolved the polyphyletic nature of Limenitidinae in a cladistic analysis using one mitochondrial gene sequence (COI, 1450 bp) and two nuclear gene sequences (EF, 1064 bp and Wg, 412–415 bp).
The order Lepidoptera has 10,045 sequences of EF in GenBank; the Nymphalidae family is the most represented with 2982 entries, followed by Lycaenidae (850), Geometridae (704), Noctuidae (675), Gracillariidae (573), Prodoxidae (485), Erebidae (449), Papilionidae (397), Sphingidae (310), Nepticulidae (298), Pieridae (268), Cosmopterigidae (247), Hesperiidae (234), Tortricidae (173), Crambidae (156), Nolidae (141), and Saturniidae (120) (Fig. 3.3a, Table 3.3). Nymphalidae occupies the first place in the number of genera and species (450 and 1555, respectively). In the second place, Geometridae has only 25 % of the Nymphalidae species, with 390 species in 215 genera (Fig. 3.3a). Butterfly families Papilionidae, Pieridae, and Nymphalidae have a high percentage of genera with EF in GenBank, with 90 %, 85 %, and 80 %, respectively.
3.5 Wingless
Wg is a nuclear protein-coding gene involved in wing, gut, and nervous system development in insects. In Lepidoptera, it handles the color and spotted pattern of the wing and thus has a critical role in ecological and evolutionary processes [21–24]. It was thought that Wg contributed to mimicry, but Kunte et al. [25] recently showed that Doublesex (dsx) is a mimicry “supergene” involved in female-specific mimicry in Heliconius and Papilio spp.
Wg has been used to resolve species and subfamily relationships in Nymphalidae [26] and was useful at a tribe level in Riodinidae and Lycaenidae families [22]. For Hesperiidae, however, the resulting relationships are not congruent with those found using EF and COI [27]. In the Geometridae family, the use of Wg in combination with EF and three other nuclear genes helped to elucidate the evolution of female flightlessness in the tribe Operophterini [28].
GenBank has 6272 records of lepidopteran Wg sequences; Nymphalidae has approximately 40 % of the records, followed by Lycaenidae, Hesperiidae, Erebidae, and Pieridae, with just 5 %. The best-known families based on the number of genera and/or species with records of Wg in GenBank are Papilionidae, which have 78.1 % of their genera and 11.2 % of species, and Nymphalidae, with 77 % of their genera and 24 % of species (Fig. 3.3b).
3.6 Enzymes and Proteins
Work with nuclear coding genes such as acetylcholine esterase, alcohol dehydrogenase, actin, chorion, silk genes, and histones, among many others [9], has been significant in Lepidoptera for economic reasons, from silk production in B. mori [6, 29, 30] to biological control in pest species like the Asian rice borer, Chilo suppressalis [31], and the tobacco hornworm, Manduca sexta [32–34], another important lepidopteran model for basic research (see below). It has also been very important in the study of metabolism associated with life history traits such as diapause and eclosion, as well as the study of metabolic pathways and the structure of proteins [6]. However, even more importantly, protein-coding genes are essential for the resolution of deep phylogenetic branches in Lepidoptera [35–37] and study of evolution in families of genes or domestication events, as in the Bombyx genus [38].
GenBank contains 33,268 enzyme sequences for Lepidoptera; the family Nymphalidae is the most represented with 8053 sequences in 382 genera and 978 species, followed by Bombycidae (5581 sequences, 14 genera, and 18 species), Noctuidae (2581 sequences, 236 genera, and 329 species), Papilionidae (1855 sequences, 39 genera, and 225 species), Gracillariidae (1276 sequences, 48 genera, and 77 species), Crambidae (1100 sequences, 338 genera, and 799 species), and Pieridae (1078 sequences, 23 genera, and 85 species) (Fig. 3.4a).
Proteins other than enzymes are documented in GenBank with twice the number of enzyme sequences (67,334 sequences); again, the most represented is Nymphalidae, with 26,852 sequences corresponding to 73 genera and 215 species, followed by Bombycidae (26,515 sequences, 15 genera, and 19 species), Papilionidae (5387 sequences, 5 genera, and 21 species), Noctuidae (2064 sequences, 32 genera, and 53 species), and Crambidae (1072 sequences, 32 genera, and 40 species) (Fig. 3.4b).
3.7 Ribosomal DNA and RNA
Ribosomes are involved in protein synthesis. Eukaryotes contain two major cytoplasmic rRNA subunits, 28S and 18S; tandem arrays of rDNA genes encoding both subunits are located on the nuclear chromosomes, but there are also rDNA genes in the mitochondria (16S and 12S). Genes encoding rRNA have been widely used in phylogenetic analysis because their different regions have distinct rates of evolution, giving diverse resolution for phylogenetic inference [9, 39]. In Lepidoptera, diverse phylogenetic analyses have included mitochondrial rDNA to construct phylogeny [40–42]. The term ribosomal is used here to report either mitochondrial or nuclear rRNA and rDNA sequences,.
GenBank has 11,652 ribosomal accessions, but these include less than 5 % of the total sequences for Lepidoptera. Nymphalidae has the highest numbers of ribosomal records in GenBank (2891), followed by Lycaenidae (1143), Noctuidae (922), and Zygaenidae (895) (Fig. 3.5a). Additionally, Nymphalidae has the highest number of genera and species represented (383 and 1129, respectively), and Papilionidae has 90 % of their genera and 33 % of species represented in GenBank, followed by Nymphalidae (68.5 % genera and 18 % species). Being a small family, it is interesting that Zygenidae appears in the 4th place for the number of accessions for ribosomal sequences in GenBank, where it is represented by 18 genera and 108 species with 895 records. One genus, Zygaena, comprises 826 records of ribosomal sequences for 85 species, including 344 records for Zygaena transalpine and 125 records for Z. angelicae [43]; Niehuis et al. [44] contributed, with the complete sequences of mitochondrially encoded NADH dehydrogenase subunit 1 (MT-ND1), tRNA-leucine (tRNA-Leu), 16S rRNA, tRNA-valine (tRNA-Val), and, with large fragment of 12S rRNA, nuclear DNA of the small and large subunits ribosomal RNA (ncDNA-18S rRNA and ncDNA-28S rRNA) for a phylogenetic study of the zygaenoid group.
3.8 Cytochrome C Oxidase Subunit I (COI)
Cytochrome c oxidase is a protein complex (subunits 1–3) located in the mitochondria that plays an important role as a terminal enzyme in the respiratory chain, transferring electrons and reducing oxygen to water. This process is carried out by subunit 1 (COI) of the complex [45, 46]. Genes encoding COI form part of the mitogenome, and analysis of its complete sequence shows that different regions evolve at distinct rates, making COI very useful for insect phylogenetic studies [47]. In Lepidoptera, COI by itself has a better resolution at lower levels, such as species and species groups [48]. At higher levels, it is recommended to use COI together with other gene sequences (e.g., Wg, EF) for phylogenetic analysis and dating of divergence times [20, 42, 49, 50]. Given that COI has low intraspecific variability and high interspecific variability, it is suitable for species recognition, and in 2003, it was proposed to be used for a universal barcoding system in species identification [51, 52]. The critical sequence consists of an approximately 600 bp long fragment of COI which is amplified by PCR and sequenced. Then, this sequence is compared to a library of COI sequences of species identified previously by taxonomists. The advantages of using COI as a barcoding system include the large number of DNA copies per cell, its maternal inheritance, and lack of introns. In Lepidoptera, the barcoding system works very well, especially for the discovery of new species in groups with crypticism [53–57] and overlooked species [58]. Since the barcoding proposal in 2003, COI sequences have been increasing, and as of April 2014, GenBank had 215,074 accessions, which represent 22 % of all the sequences within families of Lepidoptera.
Wilson [8] used a fragment of COI (DNA barcode) and two other gene regions (EF and Wg) of 977 species from Lepidoptera to probe phylogenetic signal and concluded that the DNA barcode fragment has low signal for levels above genus. In the first quarter of 2014, there were 19,279 named species belonging to 6147 genera for COI alone; the huge increase in the number of species found in GenBank represents the widespread use of this marker in taxonomic and phylogenetic studies. In fact, GenBank contains 92 % of the lepidopteran families reported by Nieukerken et al. in 2011 [4] and 39.5 % of the genera, but just 12.25 % of the number of species. The Geometridae family has the largest number of genera represented by this gene, followed by Erebidae, Noctuidae, and Nymphalidae. Although Geometridae has the highest number of species, Nymphalidae has more species represented than Erebidae or Noctuidae (Fig. 3.6a). Considering the number of genera reported for each of the families with relatively high numbers of sequences registered in GenBank, coverage of Sphingidae is 99.5 %, followed by Papilionidae, Nymphalidae, Pieridae, and Noctuidae (94 %, 86 %, 77 %, and 67 %, respectively). This pattern is similar at the species level, but Erebidae, with the largest species number reported [4], has only 8.5 % representation in GenBank (Table 3.3 and Fig. 3.6a).
Records of lepidopteran mitochondrial sequences in GenBank. (a) COI records in GenBank by family of Lepidoptera as of April 2014. (b) Families that have a complete mitochondrial genome in GenBank. The first number in brackets refers to the number of genera, and the last is the number of species in each family
3.8.1 COI and Barcode Publications in ISI Web of Science and Scopus
In the period from 2003 to 2013, the total number of publications returned in the ISI Web of Science and Scopus based on a search using keywords “barcode/barcoding Lepidoptera” was 352. The year with the largest number of publications is 2012 (56 papers). The number of publications using barcodes appears to cycle, the first being bigger than the second, with a tendency to increase from 2003 to 2008 with 47 publications. The second cycle starts in 2009 with a reduction of 36 % and reaches the maximum in 2012 (Fig. 3.7a). These fluctuations are explained by the discovery of new species with crypticism using barcoding and the large inventories of newly detected species, all waiting for a taxonomist to name them in a publication. The type of journal confirms the latter hypothesis, with the largest number of articles on the subject published in Zootaxa (28), followed by Molecular Phylogenetics and Evolution (24) and Annals of the Entomological Society of America (20) (Fig. 3.7b).
The scientific publications of this information cover 73 families, with just 60 % of the families with sequences registered in GenBank. Families with the highest number of scientific publications are Nymphalidae (66) and Noctuidae (66). All butterfly families have publications (from Hedylidae with 4 to Nymphalidae with 66), but only 51 % of moth families are present in the barcode literature (66 families, Table 3.4). The Noctuidae family contains the majority of moth barcode publications (66), followed by Tortricidae, Geometridae, Erebidae, and Crambidae (39, 37, 25, and 18 studies, respectively).
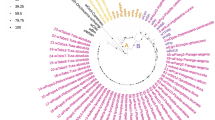
The publications with COI sequences for barcoding are mainly related to topics in taxonomy, evolution, biogeography, and biodiversity. Considering authors with the highest number of publications, 21 authors have five or more publications in this area (Fig. 3.8). N. Wahlberg currently has the most publications; his main area of research includes the systematics and evolution of the butterfly family Nymphalidae.
3.9 The Complete Mitochondrial Genome
The mitochondrial genome is the most extensively studied genomic system in insects because of its maternal inheritance, lack of recombination, small size, and an accelerated mutation rate compared to nuclear DNA. Mitochondrial DNA (mtDNA) is considerably smaller than nuclear DNA; animal mitochondria are 16–20 kb length, comprising 37 genes and lacking introns [9].
There are distinct regions within mtDNA that diverge at different rates (e.g., COI, COII, COIII, MT-ND4L [mitochondrially encoded NADH dehydrogenase 4L], Cyt b); as a result, it is very useful at diverse taxonomic levels, even to determine relationships among close species [59]. As noted previously, COI, a mitochondrial region of approximately 650 bp, was formally proposed as a barcode system for species identification in 2003 [51, 52]. This and other regions of mtDNA have been used extensively in studies of phylogenetics, comparative and evolutionary genomics, population genetics, molecular evolution, and phylogenomic analysis [60, 61].
Lepidoptera has 361 records of complete mtDNA in GenBank, representing 111 species (as accessed on April 2, 2014). Figure 3.6b shows the proportional representation for families that comprise 90 % of the accessions and the number of genera and species with a mitogenome: Nymphalidae (19/23), Bombycidae (2/3), Crambidae (10/12), Papilionidae (5/9), Noctuidae (7/9), Tortricidae (8/9), Saturniidae (6/8), Pieridae (8/9), Lycaenidae (7/7), and Erebidae (4/4). Nine families represent only 10 % of the accessions. The rapid increase of complete mitochondrial studies is important; in only 1 month, Wu et al. [62] contributed data for 29 recognized species of Nymphalidae, resulting in a total of 82 species for Papilionoidea and 58 for moths. Now, the largest number of species with complete mitochondrial genomes is Nymphalidae: Abrota ganga, Acraea issoria, Apatura ilia, A. metis, Argynnis childreni, A. hyperbius, Athyma asura, A. cama, A. kasa, A. opalina, A. perius, A. selenophora, A. sulpitia, Bhagadatta austenia, Bicyclus anynana, Calinaga davidis, Danaus plexippus, Dichorragia nesimachus, Dophia evelina, Euploea core, E. mulciber, Euthalia irrubesens, Fabriciana nerippe, Heliconius erato, H. melpomene, H. numata, Hipparchia autonoe, Issoria lathonia, Junonia almanac, J. orithya, Kallima inachus, Libythea celtis, Lexias dirtea, Melanitis leda, M. phedima, Melitaea cinxia, Neptis philyra, N. soma, Neope pulaha, Pandita sinope, Pantoporia hordonia, Parantica sita, Parasarpa dudu, Parthenos sylvia, Polyura arja, Sasakia charonda, S. funebris, Sumalia daraxa, Tanaecia julii, Timelaea maculate, Yoma sabina, and Ypthima akragas. The second largest family is Crambidae, a moth family with 12 species: C. suppressalis, Cnaphalocrocis medinalis, Diatraea saccharalis, Dichocrocis punctiferalis, Elophila interruptalis, Glyphodes quadrimaculalis, Maruca vitrata, Ostrinia furnacalis, O. nubilalis, Paracymoriza distinctalis, P. prodigalis, and Scirpophaga incertulas.
Nymphalidae represents the most diverse butterfly family, with 559 genera and 6152 species, which is one-third of all butterfly species [4]. This family has been extensively studied because it includes several species of economic importance as crop pests or potential agents for the biological control of weeds. It is widely distributed in diverse habitats worldwide, and several species have been used as models for ecological, conservation, evolutionary, and developmental studies [63–66]. Nevertheless, the relatively large number of genomic accessions for Nymphalidae is primarily due to many projects related to butterfly phylogeny [62].
Crambidae is a family with some pest species of sod grasses, maize, sugar cane, rice, and other Poaceae, including the sugarcane borer, D. saccharalis, which is an economically important pest of several major crops in North and South America. Whole mitogenome sequencing in 2011 was a major step providing molecular markers to monitor changes in population structure associated with acquisition of resistance to Bacillus thuringiensis, a class of bacterial endotoxins which is commonly used for pest control [67].
3.10 Genome Projects for Lepidoptera
Knowing the complete genome of Lepidoptera has made it a valuable model system in several ways, including the explanation of key processes such as the immune response, neurophysiology, olfaction, protein biochemistry, evolutionary mechanisms within species (e.g., evolving host–plant utilization) and between species and populations (e.g., wing pattern mimicry), the establishment of phylogenetic relationships, and as a reference for evolutionary comparisons with other insect orders. As of January 2015, eleven lepidopteran genome projects were reported: six butterflies, of which three are Nymphalidae (H. melpomene, D. plexippus, and M. cinxia) and three Papilionidae (P. glaucus, P. xuthus, and P. polytes), and five moths from diverse families (silk moth, B. mori [Bombycidae]; diamondback moth, Plutella xylostella [Plutellidae]; rice borer, C. suppressalis [Crambidae]; fall army worm, Spodoptera frugiperda [Noctuidae]; and tobacco hornworm, M. sexta [Sphingidae]) (Table 3.5). The M. sexta genome project will be published shortly, along with many other lepidopteran genome projects now in progress (Table 3.6).
Lepidopteran genomes comprise approximately 31 chromosomes [68, 69] with an average size of ~645 Mb, ranging from ~283 Mb (Danaus plexippus) to ~1897 Mb (Euchlaena irraria) [70]. Sequencing and assembling complete genomes from different lepidopteran species has taken considerable effort compared with the Drosophila genome, which has a genome size of ~180 Mb distributed on four chromosomes [71, 72]. Nevertheless, rapid improvements in the actual sequencing techniques and the significance of this group (economical, biological, and ecological) are likely to accelerate sequencing of lepidopteran genomes in order to use them in several ways, such as functional genomics, mutant analysis, bioinformatics, and other post-genomic applications that increase our biological and economical knowledge of Lepidoptera. However, it is important to solve the disaggregation of the community studying Lepidoptera as the great diversity of this group makes it difficult to consolidate operation of a Lepidoptera Consortium, limiting access to major funding [73].
3.10.1 Bombyx mori
The silkworm, B. mori (Bombycidae), has been domesticated for silk production for the past 5000 years. It is the most well-studied lepidopteran model system because of its relatively short life cycle [74, 75] and its rich repertoire of well-characterized mutations that affect virtually every aspect of the organism’s morphology, development, and behavior. Additionally, it has considerable economic importance. B. mori was the first lepidopteran insect genome to be fully sequenced.
In 2004, a Japanese and a Chinese group performed analyses of a WGS draft genome sequence of B. mori [76, 77], suggesting that the number of protein-coding genes was 18,000–20,000. The full genome of the silkworm was published in 2008 by the International Silkworm Genome Consortium [78], including a new genome assembly with 16,329 genes. This was made possible by the use of new fosmid- and BAC-end sequence data anchored to a fine genetic map, resulting in an increase in the scaffold size, which made possible a good assembly with low polymorphism (0.2 %) at the nucleotide level.
Based on an extensive database of expressed sequence tags (ESTs) [79] and full-length cDNAs [80], many Bombyx-specific genes have been found and annotated, showing the value of transcriptome sequencing for the molecular biology of the silkworm and the whole lepidopteran group.
3.10.2 Danaus plexippus
The monarch butterfly, D. plexippus (Nymphalidae), is the most well-recognized species of butterfly, which migrates up to 3000 km from central Mexico to eastern North America [81]. The initial assembly of the monarch genome was made by Zhan et al. in 2011 [82], reporting a genome draft of 273 Mb encoding 16,866 protein-coding genes and suggesting that Lepidoptera is the fastest evolving insect order. In 2013 Zhan et al. [83] established MonarchBase to make the genome data accessible. By 2014, Zhan et al. [84] reported the genetics of monarch butterfly migration and warning coloration, sequencing 80 genomes of D. plexippus and nine samples from four additional Danaus species. Among other findings, they noted that North American populations are the most basal lineages, with population structure indicating gene flow across North America, and likely origin in the southern USA or northern Mexico. They also found evidence for recurrent, divergent selection on flight muscle function and wing color variation mediated by a myosin gene with no prior known role in insect pigmentation, but an analogous effect in vertebrates. These studies illustrate the power of a genome project to enhance understanding of important biological processes.
3.10.3 Heliconius melpomene
For many years, researchers of the Heliconius group (Nymphalidae) have been searching for the mechanisms underlying adaptive radiation phenomena and Müllerian mimicry. Martin et al. [85] reported interspecific gene flow between sympatric and allopatric populations of H. melpomene, H. cydno, and H. timareta, addressing the idea of evolution without isolation. H. melpomene is a model for this type of study, and increased genome research provides the opportunity to explain some of the pathways of adaptive radiation related to the Müllerian mimicry process [86]. The Heliconius Genome Consortium published the H. melpomene genome sequence and predicted 12,657 gene models in 2012 [87] and, by comparison with D. plexippus and B. mori, found the chromosomal organization to be broadly conserved since the Cretaceous. Also, they reported [87] that the genomic region controlling the mimicry pattern has evidence of hybrid exchange of genes between H. melpomene, H. timareta, and H. elevatus. Establishment of this butterfly genome sequence has fuelled significant research, culminating in the recent publication of more robust models for the genetic and mechanistic basis of these phenomena [88].
3.10.4 Plutella xylostella
The diamondback moth, P. xylostella (Plutellidae), is one of the more serious pests of cultivated Brassicaceae worldwide [89, 90], which has rapidly evolved high resistance to conventional insecticides such as pyrethroids, organophosphates, fipronil, spinosad, B. thuringiensis toxin, and diamides. You et al. [91] published the first whole-genome sequence for this species in 2013, having 18,071 protein-coding and 1412 unique genes with an expansion of gene families related with perception and the detoxification of plant defense chemicals. They found higher levels of P. xylostella-specific genes compared with those from B. mori (463) and D. plexippus (1184). The P. xylostella-specific genes are associated with biological pathways essential to monitor and process environmental information, chromosomal replication and/or repair, transcriptional regulation, and carbohydrate and protein metabolism. These authors had to develop special techniques to deal with the extensive polymorphism in the DNA samples because they could not inbreed, as was possible in the other species, or use a cell line, as with S. frugiperda. Consequently, the genome was highly fragmented compared to other Lepidoptera genome assemblies. This will be a continuing problem as new sequences are developed for non-model Lepidoptera.
Jouraku et al. [92] developed KONAGAbase, a comprehensive transcriptome database for P. xylostella, which can assist researchers in the analysis of genes related to insecticide resistance, allowing the development of more efficient and less environmentally harmful insecticides through clarifying the mechanism of resistance.
3.10.5 Chilo suppressalis
The Asian rice stem borer, C. suppressalis (Crambidae), is one of the most economically important pests of rice crops in Northeast China [93]. C. suppressalis is a widespread species, extending from East Asia and Oceania into the Middle East and Europe [94]. Given its great economic importance, its metabolism and adaptation to xenobiotics have been extensively studied. In 2014 Yin et al. [95] obtained the first version of a draft genomic sequence for this species using an Illumina sequencing platform to generate WGS sequences that were subsequently assembled. They also established ChiloDB, a database which contains genome and transcriptome sequence data for C. suppressalis. In December 2013, they reported the following information was available in ChiloDB: 80,479 scaffolds (length ≥ 2 Kb), 10,221 annotated protein-coding sequences, 262 microRNAs, 82,639 predicted piwi-interacting RNAs, 37,040 midgut transcriptome sequences, 69,977 mixed sample transcriptome sequences, and 77 cytochrome p450 genes or gene fragments. ChiloDB group are working to improve the annotation quality to develop a comprehensive information system for the researchers [95].
3.10.6 Melitaea cinxia
The Glanville fritillary butterfly, M. cinxia, belongs to the Nymphalidae family and has been studied to understand the ecological, genetic, and evolutionary consequences of habitat fragmentation on metapopulation dynamics [96]. Vera et al. (2008) [97] reported one of the first studies using 454 pyrosequencing of cDNAs as an approach to genome sequencing for a non-model species and used relatively short sequence assemblies to create a microarray for large-scale functional genomics. However, it was not until 2014 that Ahola et al. [98] sequenced the complete genome of M. cinxia, from which they predicted 16,667 gene models. Somervuo et al. (2014) [99] found that a large number of genes were differentially expressed between the landscape types, based on RNA-sequence data. The genome sequence from this lepidopteran, which has the putative ancestral chromosome number (31), provides additional evidence for the evolutionary conservation of lepidopteran chromosomes.
3.10.7 Spodoptera frugiperda
The fall army worm, S. frugiperda (Noctuidae), is a polyphagous pest of economic importance in tropical and subtropical countries [100]. Casmuz et al. [101] conducted a literature review of records for this species in North and South America, reporting 186 host plants belonging to 42 different families. This species has devastating effects, damaging crops, and reducing food production [102].
In 2014, the International Centre for Genetic Engineering and Biotechnology (India) used a cell line (Sf9) from the ovary of S. frugiperda to obtain a draft sequence of this species. This novel approach gives good results but needs to be validated. Noctuidae is one of the largest families of Lepidoptera containing many of the agriculture pests, and this study represents the first complete genome publication in this family. The genomic DNA was sequenced and assembled into 37,243 scaffolds, 358 Mb in length, with 11,595 predicted genes, of which 36.4 % were assigned a functional characteristic. Repeat elements represent 20.28 % of the total genome. Having the complete genome sequence for this representative of a highly destructive taxonomic group will yield new insights into the evolution of such functions as host–plant specialization, detoxification of allelochemicals, insecticide resistance, and the existence of lepidopteran- and species-specific genes, ultimately helping to understand its biology for improving food production by controlling this species and its close relatives [102].
3.10.8 Papilio glaucus
Species of the genus Papilio have been the subject of many evolutionary studies that address issues ranging from population genetics, speciation, and conservation to phylogeny [50]. The North American butterfly, the Eastern tiger swallowtail, P. glaucus (Papilionidae), has remarkable morphological and behavioral features that have been described in evolutionary studies, such as Batesian mimicry [103, 104]. High levels of heterozygosity have been a problem in sequencing the genomes of species of Lepidoptera which cannot be easily inbred; the P. glaucus genome also has high levels of heterozygosity, similar to P. xylostella [105]. Nevertheless, in 2015 Cong et al. [105] succeeded in publishing the complete genome sequence for P. glaucus using a single wild-caught individual using a novel assembly strategy. Reporting a genome size of 376 Mb, they predicted 15,695 protein-coding genes and reported the function for 11,975 of them, with repeats constituting 22 % of the genome, values typical of other butterflies.
3.10.9 P. polytes and P. xuthus
The common Mormon swallowtail butterfly, Papilio polytes (Papilionidae), presents two adult forms, products of a female-limited Batesian mimicry: one mimetic form resembles Pachliopta aristolochiae and the other (cyrus) is non-mimetic [106]. In 2014, the dsx gene was reported by Kunte et al. [25] as a supergene that controls this mimetic expression. This was confirmed in 2015 by Nishikawa et al. [106], who determined whole-genome sequences of P. polytes (227 Mb, encoding 12,244 protein-coding genes) and the Asian swallowtail, P. xuthus (244 Mb, encoding 13,102 protein-coding genes). Comparison of the sequenced genomes of P. xuthus and P. polytes led to the discovery of an extended, highly heterozygous chromosomally inverted region encompassing the genetically mapped locus responsible for the mimetic polymorphism in P. polyetes females. The heterozygous, inverted region includes dsx, consistent with its proposed involvement in expression of the mimicry pattern. The Papilio genome projects are the most recent ones registered in GenBank and the first reports of an association of such a chromosome change with a historically significant phenotype in Lepidoptera. Such a phenomenon is unlikely to have been found without access to the genome sequences.
3.10.10 Manduca sexta
The tobacco hornworm, M. sexta (Sphingidae), has been used as a model system for many different fundamental studies of insect and lepidopteran biology, including behavior, immune response, transcription factors, olfaction, biochemistry, physiology, growth, and phylogenetic studies [33, 34, 107–112]. Recently, in 2012, a WGS genome project of M. sexta was registered in GenBank by M. Kanost, G. Blissard, J. Qu, S. Richards, et al. (accession number AIXA00000000.1) The genome sequence of this species will lead to an advanced understanding of many basic mechanisms in insect interactions with plants, other insects, and microbes, with potential applications in the areas of biomedicine (insect-vectored diseases) and agriculture (insect–plant interactions). As yet no publications concerning this sequencing project are available but are anticipated in the near future.
3.11 Lepidoptera Genomics Enlightens the Biological Sciences
Butterfly and moth sequences for individual mRNAs were first submitted to GenBank database in the early 1980s [113, 114]. Butterfly and moth genomes, particularly the B. mori genome, were among the first insect genomes to be sequenced; the B. mori genome was sequenced because of the importance of this insect in silk production, which researchers were focused on improving. Subsequent sequencing of Lepidoptera has targeted other economically significant species, such as S. frugiperda and P. xylostella. Despite the many GenBank entries (over one million) for the order Lepidoptera, the richness and biological diversity of this order remain underrepresented. The primary aim of current research is to explain complex processes, such as evolution, from a whole-genome perspective, for which lepidopterans are excellent models because many of their ecological and evolutionary traits are known. This potential has already been noticed, and now is the time to use deep genomics to understand these processes. New sequencing technologies are simplifying this task. Further work should focus on obtaining additional species with complete genomes to gain a better representation of the order Lepidoptera in the GenBank database. Additionally, taxonomists have an important task regarding sequenced specimens that remain unnamed because of the way in which barcoding with COI has accelerated the discovery of greater biodiversity. With greater collaborations among ecological, biological, biogeographical, evolutionary, and genomic researchers using Lepidoptera, new findings that will affect fundamental knowledge in all biological sciences can be discovered.
Abbreviations
- cDNA:
-
Complementary DNA
- BAC:
-
Bacterial artificial chromosome
- CDS:
-
Coding sequences
- COI, COII, COIII :
-
Cytochrome oxidase subunits I, II, III
- cyt b :
-
Cytochrome b
- dsx :
-
Doublesex
- EF :
-
Elongation factor-1α
- EST:
-
Expressed sequence tag
- mtDNA:
-
Mitochondrial DNA
- MT-ND4L :
-
Mitochondrially encoded NADH dehydrogenase 4L
- MT-ND1 :
-
Mitochondrially encoded NADH dehydrogenase subunit 1
- ncDNA-18S rRNA :
-
Nuclear DNA of the small subunit ribosomal RNA
- ncDNA-28S rRNA :
-
Nuclear DNA of the large subunit ribosomal RNA
- NCBI:
-
National Center for Biotechnology Information
- rDNA :
-
Ribosomal DNA
- rRNA :
-
Ribosomal RNA
- tRNA-Leu :
-
tRNA-leucine
- tRNA-Val :
-
tRNA-valine
- Wg :
-
Wingless
- WGS:
-
Whole-genome shotgun
References
Roe AD, Weller SJ, Baixeras J, Brown J, Cummings MP, Davis DR et al (2010) Evolutionary framework for Lepidoptera model systems. In: Goldsmith M, Marec F (eds) Genetics and molecular biology of Lepidoptera. CRC Press, Boca Raton, pp 1–24
Scoble MJ (1992) The Lepidoptera: form, function, and diversity. Oxford University Press, New York
Misof B, Liu S, Meusemann K, Peters RS, Donath A, Mayer C et al (2014) Phylogenomics resolves the timing and pattern of insect evolution. Science 346:763–767
Nieukerken EJV, Kaila L, Kitching IJ, Kristensen NP, Lees DC, Minet J et al (2011) Order Lepidoptera. In: Zhang Z-Q (ed) Animal biodiversity: an introduction to higher-level classification and taxonomic richness. Zootaxa 3148:212–221, Aukland, New Zealand
Rubin GM, Lewis EB (2000) A brief history of Drosophila’s contributions to genome research. Science 287(5461):2216–2218
Willis JH, Wilkins AS, Goldsmith MR (1995) A brief history of Lepidoptera as model systems. In: Goldsmith MR, Wilkins AS (eds) Molecular model systems in the Lepidoptera. Cambridge University Press, Cambridge, pp 1–20
Benson DA, Cavanaugh M, Clark K, Karsch-Mizrachi I, Lipman DJ, Ostell J, Sayers EW (2012) GenBank. Nucleic Acids Res:1–7. doi:10.1093/nar/gks1195
Wilson JJ (2010) Assessing the value of DNA barcodes and other priority gene regions for molecular phylogenetics of Lepidoptera. PLoS One 5(5):e10525. doi:10.1371/journal.pone.0010525
Hoy MA (2003) Insect molecular genetics. An introduction to principles and applications. Academic, Boston
Maroni G (1993) An atlas of Drosophila genes. Oxford University Press, Oxford
Lin CP, Danforth BN (2004) How do insect nuclear and mitochondrial gene substitution patterns differ? Insights from Bayesian analyses of combined datasets. Mol Phylogenet Evol 30:686–702
Kim M, Wan X, Kim MJ, Jeong HC, Ahn N, Kim K et al (2010) Phylogenetic relationships of true butterflies (Lepidoptera: Papilionoidea) inferred from COI, 16S rRNA and EF-1α sequences. Mol Cells 30:409–425
Cho S, Mitchell A, Regier JC, Mitter C, Poole RW, Friedlander TP et al (1995) A highly conserved nuclear gene for low-level phylogenetics: Elongation factor-1α recovers morphology-based tree for heliothine moths. Mol Biol Evol 12:650–656
Friedlander TP, Horst KR, Regier JC, Mitter C, Peigler RS, Fang QQ (1998) Two nuclear genes yield concordant relationship within Attacini (Lepidoptera: Saturniidae). Mol Phylogenet Evol 9:131–140
Mitchell A, Cho S, Regier JC, Mitter C, Poole RW, Matthews M (1997) Phylogenetic utility of elongation factor-1 alpha in noctuidae (Insecta: Lepidoptera): the limits of synonymous substitution. Mol Biol Evol 14(4):381–390
Mitchell A, Mitter C, Regier JC (2000) More taxa or more characters revisited: combining data from nuclear protein-encoding genes for phylogenetic analysis of Noctuoidea (Insecta: Lepidoptera). Syst Biol 49:202–224
Moulton JK (2000) Molecular sequence data resolves basal divergences within Simuliidae (Diptera). Syst Entomol 25:95–113
Regier JC, Mitter C, Peigler RS, Friedlander TP (2000) Phylogenetic relationship in Lasiocampidae (Lepidoptera): initial evidence from elongation factor-1 alpha sequences. Insect Syst Evol 31:179–186
Caterino MS, Cho S, Sperling FAH (2000) The current state of insect molecular systematics: a thriving Tower of Babel. Annu Rev Entomol 45:1–54
Wahlberg N, Weingartner E, Nylin S (2003) Towards a better understanding of the higher systematics of Nymphalidae (Lepidoptera: Papilionoidea). Mol Phylogenet Evol 28:473–484. doi:10.1016/S1055-7903(03)00052-6
Carroll SB, Gates J, Keys DN, Paddock SW, Panganiban GE, Selegue JE et al (1994) Pattern formation and eyespot determination in butterfly wings. Science 265(5168):109–114
Campbell DL, Brower AV, Pierce NE (2000) Molecular evolution of the wingless gene and its implications for the phylogenetic placement of the butterfly family Riodinidae (Lepidoptera: Papilionoidea). Mol Biol Evol 17(5):684–696
Beldade P, Brakefield PM (2002) The genetics and evo-devo of butterfly wing patterns. Nat Genet 3:442–452
Werner T, Koshikawa S, Williams TM, Carroll SB (2010) Generation of a novel wing colour pattern by the Wingless morphogen. Nature 464:1143–1148
Kunte K, Zhang W, Tenger-Trolander A, Palmer DH, Martin A, Reed RD et al (2014) Doublesex is a mimicry supergene. Nature 507(7491):229–232
Brower AVZ, DeSalle R (1998) Patterns of mitochondrial versus nuclear DNA sequence divergence among nymphalid butterflies: the utility of wingless as a source of characters for phylogenetic inference. Insect Mol Biol 7:1–10
Warren AD, Ogawa JR, Brower AVZ (2008) Phylogenetic relationships of subfamilies and circumscription of tribes in the family Hesperiidae (Lepidoptera: Hesperioidea). Cladistics 24:642–676
Snäll N, Tammaru T, Wahlberg N, Viidalepp J, Ruohomaki K, Savontaus ML et al (2007) Phylogenetic relationships of the tribe Operophterini (Lepidoptera, Geometridae): a case study of the evolution of female flightlessness. Biol J Linn Soc 92(2):241–252
Fedic R, Zurovec M, Sehnal F (2002) The silk of Lepidoptera. J Insect Biotechnol Sericol 71:1–15
Goldsmith MR, Shimada T, Abe H (2004) The genetics and genomics of the silkworm, Bombyx mori. Annu Rev Entomol 50:71–100
Gong ZJ, Zhou WW, Yu HZ, Mao CG, Zhang CX, Cheng JA et al (2009) Cloning, expression and functional analysis of a general odorant-binding protein 2 gene of the rice striped stem borer, Chilo suppressalis (Walker) (Lepidoptera: Pyralidae). Insect Mol Biol 18(3):405–417
Feng L, Prestwich GD (1997) Expression and characterization of a lepidopteran general odorant binding protein. Insect Biochem Mol Biol 27(5):405–412
Martin JP, Lei H, Riffell JA, Hildebrand JG (2013) Synchronous firing of antennal-lobe projection neurons encodes the behaviorally effective ratio of sex-pheromone components in male Manduca sexta. J Comp Physiol A 199:963–979
Vogt RG, Große-Wilde E, Zhou J-J (2015) The Lepidoptera odorant binding protein gene family: gene gain and loss within the GOBP/PBP complex of moths and butterflies. Insect Biochem Mol Biol. 62:142–153 http://dx.doi.org/10.1016/j.ibmb.2015.03.003
Mutanen M, Wahlberg N, Kalla L (2010) Comprehensive gene and taxon coverage elucidates radiation patterns in moths and butterflies. Proc R Soc B:277(1695):2839–2848. doi:10.1098/rspb.2010.0392
Regier JC et al (2009) Toward reconstructing the evolution of advanced moths and butterflies (Lepidoptera: Ditrysia): an initial molecular study. BMC Evol Biol 9:280. doi:10.1186/1471-2148-9-280
Regier JC et al (2013) A large-scale, higher-level, molecular phylogenetic study of the insect order Lepidoptera (moths and butterflies). PLoS One 8(3):e58568. doi:10.1371/journal.pone.0058568
Xia Q, Guo Y, Zhang Z, Li D, Xuan Z, Li Z et al (2009) Complete resequencing of 40 genomes reveals domestication events and genes in silkworm (Bombyx). Science 326(5951):433–436
Hills DM, Dixon MT (1991) Ribosomal DNA: molecular evolution and phylogenetic inference. Q Rev Biol 66(4):411–453
Pashley DP, Ke LD (1992) Sequence evolution in mitochondrial ribosomal and ND-1 genes in lepidoptera: implications for phylogenetic analyses. Mol Biol Evol 9(6):1061–1075
Wiegmann BM, Mitter C, Regier JC, Friedlander TP, Wagner DM, Nielsen ES (2000) Nuclear genes resolve Mesozoic-aged divergences in the insect order Lepidoptera. Mol Phylogenet Evol 15(2):242–259
Zimmermann M, Wahlberg N, Descimon H (2000) Phylogeny of Euphydryas checkerspot butterflies (Lepidoptera: Nymphalidae) based on mitochondrial DNA sequence data. Ann Entomol Soc Am 93(3):347–355
von Reumont BJ, Struwe J-F, Schwarzer J, Misof B (2011) Phylogeography of the burnet moth Zygaena transalpina complex: molecular and morphometric differentiation suggests glacial refugia in Southern France, Western France and micro-refugia within the Alps. J Zool Syst Evol Res 50(1):38–50. doi:10.1111/j.1439-0469.2011.00637.x
Niehuis O, Yen SH, Naumann CM, Misof B (2006) Higher phylogeny of zygaenid moths (Insecta: Lepidoptera) inferred from nuclear and mitochondrial sequence data and the evolution of larval cuticular cavities for chemical defence. Mol Phylogenet Evol 39:812–829
Capaldi RA, Malatesta F, Darley-Usmar VM (1983) Structure of cytochrome c oxidase. BBA Bioenergetics 726(2):135–148
Michel H (1998) The mechanism of proton pumping by cytochrome c oxidase. Proc Natl Acad Sci U S A 95:12819–12824
Lunt DH, Zhang DX, Szymura JM, Hewitt GM (1996) The insect cytochrome oxidase I gene: evolutionary patterns and conserved primers for phylogenetics studies. Insect Mol Biol 5(3):153–165
Caterino MS, Sperling FAH (1999) Papilio phylogeny based on mitochondrial cytochrome oxidase I and II genes. Mol Phylogenet Evol 11(1):122–137
Brower AVZ (1994) Phylogeny of Heliconius butterflies inferred from mitochondrial DNA sequences (Lepidoptera: Nymphalidae). Mol Phylogenet Evol 3(2):159–174
Zakharov E, Caterino MS, Sperling FAH (2004) Molecular phylogeny, historical biogeography, and divergence time estimates for swallowtail butterflies of the genus Papilio (Lepidoptera: Papilionidae). Syst Biol 53(2):193–215
Hebert PDN, Cywinska A, Ball SL, deWaard JR (2003) Biological identifications through DNA barcodes. P Roy Soc B Biol Sci 270:313–321
Hebert PDN, Ratnasingham S, deWaard JR (2003) Barcoding animal life: cytochrome c oxidase subunit 1 divergences among closely related species. P Roy Soc B Biol Sci 270:S596–S599
Hebert PDN, Penton EH, Burns JM, Janzen DH, Hallwachs W (2004) Ten species in one: DNA barcoding reveals cryptic species in the neotropical skipper butterfly Astraptes fulgerator. Proc Natl Acad Sci U S A 101:14812–14817
Janzen DH, Hajibabaei M, Burns J, Hallwachs W, Remigio E, Hebert PDN (2005) Wedding biodiversity inventory of a large and complex Lepidoptera fauna with DNA barcoding. Philos Trans R Soc Lond B Biol Sci 2005 Oct 29; 360(1462):1835–1845
Janzen DH, Hallwachs W, Blandin P, Burns JM, Cadiou JM, Chacon I et al (2009) Integration of DNA barcoding into an ongoing inventory of complex tropical biodiversity. Mol Ecol Resour 9:1–26
Burns JM, Janzen DH, Hajibabaei M, Hallwachs W, Hebert PDN (2007) DNA barcodes of closely related (but morphologically and ecologically distinct) species of skipper butterflies (Hesperiidae) can differ by only one to three nucleotides. J Lepid Soc 61:138–153
Burns JM, Janzen DH, Hajibabaei M, Hallwachs W, Hebert PDN (2008) DNA barcodes and cryptic species of skipper butterflies in the genus Perichares in Area de Conservacion Guanacaste, Costa Rica. Proc Natl Acad Sci U S A 105:6350–6355
Prado B, Pozo C, Valdez-Moreno M, Hebert PDN (2011) Beyond the colours: discovering hidden diversity in the nymphalidae of the Yucatan peninsula in Mexico through DNA barcoding. PLoS One 6(11):e27776. doi:10.1371/journal.pone.0027776
Beltran M, Jiggins CD, Bull V, Linares M, Mallet J, McMillan WO et al (2002) Phylogenetic discordance at the species boundary: comparative gene genealogies among rapidly radiating Heliconius butterflies. Mol Biol Evol 19(12):2176–2190
Hu J, Zhang D, Hao J, Huang D, Cameron S, Zhu C (2010) The complete mithocondrial genome of the yellow coaster, Craea issoria (Lepidoptera: Nymphalidae: Heliconiinae: Acraeini): sequence, gene organization and a unique tRNA translocation event. Mol Biol Rep 37:3431–3438
Cameron SL (2014) Insect mitocondrial genomics: implications for evolution and phylogeny. Annu Rev Entomol 59:95–117
Wu L, Lin L, Lees DC, Hsu Y (2014) Mitogenomic sequences effectively recover relationships within brush-footed butterflies (Lepidptera: Nymphalidae). BMC Genomics 15:468
Ackery PR, Vane-Wright RI (1984) Milkweed butterflies: their cladistics and biology, being an account of the natural history of the Danainae, a subfamily of the Lepidoptera, Nymphalidae. British Museum, London
Ehrlich PR, Hanski I (2004) On the wings of checkerspots: a model system for population biology. Oxford University Press, New York
Sheppard PM, Turner J, Brown K, Benson W, Singer M (1985) Genetics and the evolution of Muellerian mimicry in Heliconius butterflies. Philos Trans R Soc Lond B Biol Sci 308:433–610
Pollard E, Yates TJ (1993) Monitoring butterflies for ecology and conservation. Chapman and Hall, London
Li W, Zhang X, Fan Z, Yue B, Huang F, King E et al (2011) Structural characteristics and phylogenetic analysis of the mitochondrial genome of the sugarcane borer, Diatraea saccharalis (Lepidoptera: Crambidae). DNA Cell Biol 30(1):3–8
Saura A, von Schoultz B, Saura AO, Brown KS Jr (2013) Chromosome evolution in Neotropical butterflies. Hereditas 150:26–37
Robinson R (1971) Lepidoptera genetics. Pergamon, Oxford
Gregory TR, Hebert PDN (2003) Genome size variation in lepidopteran insects. Can J Zool 81:1399–1405
Adams MD, Celniker SE, Holt RA, Evans CA, Gocayne JD, Amanatides PG et al (2000) The genome sequence of Drosophila melanogaster. Science 287(5461):2185–2195
Celniker SE, Rubin GM (2003) The Drosophila melanogaster genome. Annu Rev Genom Hum G 4:89–117
Beldade P, McMillan WO, Papanicoloau A (2008) Butterfly genomics eclosing. Heredity 100:150–157
Goldsmith M, Marec F (2010) Genetics and molecular biology of Lepidoptera. CRC Press, Boca Raton
Bisch-Knaden S, Daimon T, Shimada T, Hansson BS, Sachse S (2014) Anatomical and functional analysis of domestication effects on the olfactory system of the silkmoth Bombyx mori. P Roy Soc B Biol Sci 281(1774):20132582
Mita K, Kasahara M, Sasaki S, Nagayasu Y, Yamada T, Kanamori H et al (2004) The genome sequence of silkworm, Bombyx mori. DNA Res 11(1):27–35
Xia Q, Zhou Z, Lu C, Cheng D, Dai F, Li B et al (2004) A draft sequence for the genome of the domesticated silkworm (Bombyx mori). Science 306(5703):1937–1940
The International Silkworm Genome Consortium (2008) The genome of a lepidopteran model insect, the silkworm Bombyx mori. Insect Biochem Mol Biol 38(12):1036–1045
Mita K, Morimyo M, Okano K, Koike Y, Nohata J, Kawasaki H et al (2003) The construction of an EST database for Bombyx mori and its application. Proc Natl Acad Sci U S A 100(24):14121–14126
Suetsugu Y, Futahashi R, Kanamori H, Kadono-Okuda K, Sasanuma S, Narukawa J et al (2013) Large scale full-length cDNA sequencing reveals a unique genomic landscape in a lepidopteran model insect, Bombyx mori. G3 (Bethesda) 3(9):1481–1492
Miller NG, Wassenaar LI, Hobson KA, Norris DR (2012) Migratory connectivity of the monarch butterfly (Danaus plexippus): patterns of spring re-colonization in eastern North America. PLoS One 7(3):e31891
Zhan S, Merlin C, Boore JL, Reppert SM (2011) The monarch butterfly genome yields insights into long-distance migration. Cell 147(5):1171–1185
Zhan S, Reppert SM (2013) MonarchBase: the monarch butterfly genome database. Nucleic Acids Res 41(D1):D758–D763
Zhan S, Zhang W, Niitepõld K, Hsu J, Haeger JF, Zalucki MP et al (2014) The genetics of monarch butterfly migration and warning colouration. Nature 514(7522):317–321
Martin SH, Dasmahapatra KK, Nadeau NJ, Salazar C, Walters JR, Simpson F et al (2013) Genome-wide evidence for speciation with gene flow in Heliconius butterflies. Genome Res 23(11):1817–1828
Cuthill JH, Charleston M (2012) Phylogenetic Codivergence supports coevolution of mimetic Heliconius butterflies. PLoS One 7(5):e36464. doi:10.1371/journal.pone.0036464
Heliconius Genome Consortium (2012) Butterfly genome reveals promiscuous exchange of mimicry adaptations among species. Nature 487(7405):94–98
Kronforst MR, Papa R (2015) The functional basis of wing patterning in Heliconius butterflies: the molecules behind mimicry. Genetics 200:1–19
Sarfraz M, Dosdall LM, Keddie BA (2006) Diamondback moth–host plant interactions: implications for pest management. Crop Prot 25(7):625–639
De Bortoli SA, Polanczyk RA, Vacari AM, De Bortoli CP, Duarte RT (2013) Plutella xylostella (Linnaeus, 1758) (Lepidoptera: Plutellidae): tactics for integrated pest management in Brassicaceae. In: Soloneski S (ed) Weed and pest control–conventional and new challenges. InTech. doi:5772/54110
You M, Yue Z, He W, Yang X, Yang G, Xie M et al (2013) A heterozygous moth genome provides insights into herbivory and detoxification. Nat Genet 45(2):220–225
Jouraku A, Yamamoto K, Kuwazaki S, Urio M, Suetsugu Y, Narukawa J et al (2013) KONAGAbase: a genomic and transcriptomic database for the diamondback moth, Plutella xylostella. BMC Genomics 14(1):464
Su JW, Xuan WJ, Sheng CF, Ge F (2003) Biology of overwintering larvae of the Asiatic rice borer, Chilo suppressalis, in paddy fields of Northeast China. Entomol Knowl 4:007
Khan ZR, Litsinger JA, Barrion AT, Villanueva FFD (1991) World bibliography of Rice Stem Borers 1794–1990. International Rice Research Institute, Makati
Yin C, Liu Y, Liu J, Xiao H, Huang S, Lin Y et al (2014) ChiloDB: a genomic and transcriptome database for an important rice insect pest Chilo suppressalis. Database:1–7. Published online 2005 Sep 14. doi:10.1098/rstb.2005.1715
Hanski I (2011) Eco-evolutionary spatial dynamics in the Glanville fritillary butterfly. Proc Natl Acad Sci U S A 108:14397–14404. doi:10.1073/pnas.1110020108
Vera JC, Wheat CW, Fescemyer HW, Frilander MJ, Crawford DL, Hanski I et al (2008) Rapid transcriptome characterization for a nonmodel organism using 454 pyrosequencing. Mol Ecol 17(7):1636–1647
Ahola V, Lehtonen R, Somervuo P, Salmela L, Koskinen P, Rastas P et al (2014) The Glanville fritillary genome retains an ancient karyotype and reveals selective chromosomal fusions in Lepidoptera. Nat Commun. 5:4737 doi:10.1038/ncomms5737
Somervuo P, Kvist J, Ikonen S, Auvinen P, Paulin L, Koskinen P et al (2014) Transcriptome analysis reveals signature of adaptation to landscape fragmentation. PLoS One 9(7):e101467
Valencia Cataño SJ, Rodríguez Chalarca J, Mesa Cobo NC (2014) Effect of varieties of cotton GM on Spodoptera frugiperda Smith (Lepidoptera: Noctuidae) larvae. Acta Agron 63:63–70
Casmuz A, Juárez ML, Socías MG, Murúa MG, Prieto S, Medina S et al (2010) Revisión de los hospederos del gusano cogollero del maíz, Spodoptera frugiperda (Lepidoptera: Noctuidae). Revista de la Sociedad Entomológica Argentina 69:209–231
Kakumani PK, Malhotra P, Mukherjee SK, Bhatnagar RK (2014) A draft genome assembly of the army worm, Spodoptera frugiperda. Genomics 104(2):134–143
Brower JVZ (1958) Experimental studies of mimicry in some North American butterflies: Part II. Battus philenor and Papilio troilus, P. polyxenes and P. glaucus. Evolution 12:123–136
Clarke CA, Sheppard PM (1962) The genetics of the mimetic butterfly Papilio glaucus. Ecology 43:159–161
Cong Q, Borek D, Otwinowski Z, Grishin NV (2015) Tiger swallowtail genome reveals mechanisms for speciation and caterpillar chemical defense. Cell Rep 10:910–919. doi:10.1016/j.celrep.2015.01.026
Nishikawa H, Iijima T, Kajitani R, Yamaguchi J, Ando T, Suzuki Y et al (2015) A genetic mechanism for female-limited Batesian mimicry in Papilio butterfly. Nat Genet 47(4):405–411
He Y, Cao X, Li K, Hu Y, Chen YR, Blissard G et al (2015) A genome-wide analysis of antimicrobial effector genes and their transcription patterns in Manduca sexta. Insect Biochem Mol Biol 62:23–37. doi:10.1016/j.ibmb.2015.01.015
Cao X, He Y, Hu Y, Wang Y, Chen YR, Bryant B et al (2015) The immune signaling pathways of Manduca sexta. Insect Biochem Mol Biol 62:64–74. doi:10.1016/j.ibmb.2015.03.006
Zhang X, He Y, Cao X, Gunaratna RT, Chen YR, Blissard G et al (2015) Phylogenetic analysis and expression profiling of the pattern recognition receptors: insights into molecular recognition of invading pathogens in Manduca sexta. Insect Biochem Mol Biol 62:38–50. doi:10.1016/j.ibmb.2015.02.001
Tobler A, Nijhout HF (2010) Developmental constraints on the evolution of wing-body allometry in Manduca sexta. Evol Dev 12(6):592–600
Thaler JS, Contreras H, Davidowitz G (2014) Effects of predation risk and plant resistance on Manduca sexta caterpillar feeding behaviour and physiology. Ecol Entomol 39(2):210–216
Zhang S, Cao X, He Y, Hartson S, Jiang H (2014) Semi-quantitative analysis of changes in the plasma peptidome of Manduca sexta larvae and their correlation with the transcriptome variations upon immune challenge. Insect Biochem Mol Biol 47:46–54
Ohshima Y, Suzuki Y (1977) Cloning of the silk fibroin gene and its flanking sequences. Proc Natl Acad Sci U S A 74(12):5363–5367
Lecanidou R, Eickbush TH, Rodakis GC, Kafatos FC (1983) Novel B family sequence from an early chorion cDNA library of Bombyx mori. Proc Natl Acad Sci U S A 80(7):1955–1959
Acknowledgments
We thank the reviewers of this chapter, M. R. Goldsmith, A. A. Tolulope, and R. Chandrasekar, who helped us to improve it. In particular, M. R. Goldsmith helped us on the incorporation of information in a relevant way.
We would like to thank Jovana Jasso Martínez, Karen Fernanda Real Salazar, Ana Karina Cruz Galindo, and Azalea Guadalupe Acosta Carreón, who have contributed to this work.
This work was supported by El Colegio de la Frontera Sur and Facultad de Ciencias, Universidad Nacional Autónoma de México.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Pozo, C., Prado, B., Castañeda-Sortibrán, A.N. (2015). Updating Genomic Data of Lepidoptera. In: Raman, C., Goldsmith, M., Agunbiade, T. (eds) Short Views on Insect Genomics and Proteomics. Entomology in Focus, vol 3. Springer, Cham. https://doi.org/10.1007/978-3-319-24235-4_3
Download citation
DOI: https://doi.org/10.1007/978-3-319-24235-4_3
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-24233-0
Online ISBN: 978-3-319-24235-4
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)