Abstract
The cystic fibrosis transmembrane conductance regulator (CFTR), also known as ABCC7, is an unusual member of the ATP-binding cassette family in that it is an ion channel rather than a transporter. Here, ATP binding is thought to initiate channel opening, with hydrolysis of the nucleotide being associated with channel closure. Loss of CFTR function through mutation leads to the life-threatening and -shortening condition known as cystic fibrosis. Recently, the importance of external factors affecting CFTR function has also been reported. In this chapter I have focussed on the basic structural biology and biochemistry of the protein but have also attempted to link this knowledge with the understanding of the disease. I have employed a bottom-up approach, starting with what can be learned from the primary structure of the protein, leading on to an examination of the secondary and tertiary structure of CFTR. The mapping of common CF-causing missense mutations within the CFTR 3D structure will also be addressed here. Lastly, the chapter finishes with some discussion of the prospects for future research on CFTR and how data for the channel function of CFTR can inform the ATP-binding cassette field of study in general.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
Introduction
As discussed at the end of this chapter, ABCC7 (hereafter termed CFTR) is one of the best-studied ABC transporters. It should be stated from the outset that, as far as we know, CFTR is not a transporter, but rather a channel allowing the flux of negatively charged anions such as chloride and bicarbonate. However, the fact that we know so much about it means that it sits quite comfortably in this book alongside more robust, active transporters that do a serious haulage job—transporting molecules out of, or into the cell. There is a wealth of data on CFTR and an abundance of excellent reviews available to the interested PhD student or researcher. This makes the job of doing justice to the breadth and depth of CFTR research well beyond the capacity of the author. Instead, the challenge of reviewing a relatively narrow window of CFTR research was chosen. In so doing there are hopefully some original insights in this chapter, or at least something to provoke the reader into a new direction of thought. As a structural biologist particularly interested in membrane proteins, the ABC transporter field is an ideal playground. With the exception of the GPCR family, the ABC transporter field is perhaps the most richly endowed with structural data for full-length transmembrane proteins. If one is allowed to count structures for the (mostly) water-soluble nucleotide-binding and substrate-binding domain structures of ABC transporters, then the ABC family dominates at the pinnacle of structural data for membrane proteins. This chapter will spend some time examining the primary structure (i.e. the amino acid sequence) of CFTR and some recent bioinformatics analyses. It is with good reason that we should start a review of CFTR with its primary structure: Not only do we have outstanding databases for CFTR missense mutations that set this ABC family member apart from its sibs, but also because in CFTR significant portions of its polypeptide chain are disordered. Hence primary structure is directly linked to function for these regions. The chapter then looks at the secondary and tertiary structure of CFTR and finally tries to place the mutational information within the context of the tertiary structure.
I wish to thank colleagues in the CFF CFTR 3D structure consortium who have contributed hugely to ideas and insights into this chapter via discussions and with the sharing of reagents, in particular: Jack Riordan (University of North Carolina), John Hunt (Columbia University), Julie Forman-Kay (University of Toronto), Christie Brouillette, Larry DeLucas, John Kappes (University of Alabama), Ina Urbatsch (Lubbock University), Patrick Thibodeau (University of Pittsburgh) and Hanoch Senderowitz (Bar-Ilan University). I also thank my colleagues here in Manchester who have provided advice and help, in particular Natasha Cant, Mark Rosenberg and Naomi Pollock. Much of the structural biology research on CFTR done in the laboratory of RCF is and was funded by the CFF.
CFTR and Disease
Loss of CFTR channel function as a result of mutations in the cftr gene can cause the disease cystic fibrosis (Cant et al. 2014; Sosnay et al. 2013). This life-threatening and life-shortening condition affects mainly people of European origins, though why this population in particular is affected remains a mystery. Presumably at some point in human evolution, being a heterozygote with one mutated copy of the cftr gene carried a selective advantage. There have been many attempts to explain this, including theories around the past high mortality rates in children due to poor hygiene and intestinal infections. However no single explanation has yet been accepted by the majority of the cystic fibrosis (CF) research community (Cant et al. 2014).
CFTR dysfunction affects many organs in the body, but is arguably most serious in the lungs, where the lack of chloride efflux from the apical surface of epithelial airway cells causes shrinkage of the airway surface liquid layer and a rise in the viscosity of the mucus. The subsequently sticky mucus becomes the site of persistent bacterial and fungal infections, leading to inflammation and eventual remodelling and fibrosis of the airways (Riordan 2008). After many rounds of infection and inflammation, the lungs become more and more inefficient at oxygen exchange and have less capacity for inhaling air, leading to serious life-threatening disability. CFTR dysfunction has also been associated with other diseases where it is thought that CFTR activity is inhibited by external factors. For example, cigarette smoking has been associated with CFTR dysfunction (Clunes et al. 2012).
CFTR Primary Structure
CFTR (ABCC7) is a 1480-residue membrane protein, with the typical ABC transporter domain architecture of two transmembrane domains (TMDs), and two nucleotide-binding domains (NBDs) that are all fused in a single polypeptide chain (Higgins 1992). This is described elsewhere in detail in this book and will not be re-worked here. Unique features to CFTR are an additional long (circa 200-residue) regulatory (R) region, as well as N- and C-terminal regions about 80 and 40 residues in length, respectively (Hunt et al. 2013). These regions are arranged in order from N- to C-terminus: N-term-TMD1–NBD1–R–TMD2–NBD2–C-term. It is important to recognise that having a long linker region between NBD1 and TMD2 is not unique to CFTR in the ABC transporter family. In plants, for example, the first member of the ABCA family also has a linker region between NBD1 and TMD2 that is over 150 residues long. However, it displays no significant sequence homology with the CFTR R region. Similarly, the ABCC family in mammals has several members with an additional transmembrane domain, often called the TMD0. This TMD of 3–5 transmembrane helices is joined to the standard TMD1 domain by a cytoplasmic linker that is also quite long (120 residues). From phylogenetic analysis, CFTR must have evolved from the ABCC subfamily, although there is little evidence to suggest any residual ABCC-type function in the present-day protein (Dean and Allikmets 2001). This ‘C’ family consists of several drug transporters with wide-ranging substrates. Glutathione transport is often associated with this family, and glutathione-conjugated drugs are common substrates.
Co-evolution of CFTR Residues
Given that CFTR has diverged from the ABC transporter family as a channel, then some insights into its channel function may be derived by bioinformatics analysis of its primary structure and the co-evolution of residues. A few attempts have been made to carry out such a bioinformatics analysis. A study of 2000 ABC transporter sequences that were extracted (by BLAST) on the basis of similarity to the CFTR sequence was carried out after filtering and processing by multiple sequence alignment. The data can be downloaded at http://cftrfolding.org/Mendoza_Thomas_Cell_2012/Mendoza_Thomas_Cell_2012.html (Mendoza et al. 2012). The resulting matrices show correlations between positions separated in the sequence but likely linked in terms of functional interactions or closely linked in terms of 3D space. The authors also employed the data to look for residues that could be mutated to rescue the effects of the most common disease-causing mutation in CF patients (F508 deletion).
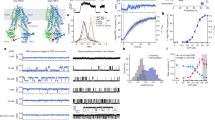
Figure 1 gives an illustration of a subset of data from the 2D matrix employing one of the four statistical tests (statistical coupling analysis, SCA) (Lockless and Ranganathan 1999) used by the authors. Here, the numerical score provides a quantitation of the degree to which the probability of finding an individual residue at position i in the sequence is dependent on a given perturbation at a separate position j in the sequence. Correlation between three residues likely involved in CFTR channel properties in the sixth transmembrane helix of TMD1 (R334, I336, T338) and all other residues in the CFTR sequence is plotted. Regions of very low correlation (co-evolution) correspond to unique parts of the CFTR primary structure, as discussed above, as well as the C-terminal region which also shows little similarity to any other ABC transporter. Interestingly, peaks in the SCA scores are distributed throughout the primary structure of CFTR and there are no immediately obvious clues as to the evolution of CFTR as a channel except in the regions of very low SCA score. Regions with low scores correspond to regions that are likely to be specialised in CFTR channel and maturation functions, and show little or no conservation with other ABC proteins. As mentioned above, a significant proportion of CFTR contains regions that are also likely to be structurally disordered and these all fall into low SCA score regions shown in Fig. 1. Interestingly the N-terminal region is an exception to this: It is thought to be disordered, but it does not display scores close to zero, implying more homology with other ABC proteins and that it may not be specialised for CFTR function. The opposite applies to the TMD–NBD linker regions, which are not disordered in the available high-resolution ABC protein exporter structures (where the TMD and NBD are fused). Here, the low SCA scores probably relate to the observation that CFTR linkers are quite different from those of other ABC proteins in terms of length as well as sequence (Hunt et al. 2013). This specialisation of the linkers in CFTR may be related to their location, which is along the ‘shoulders’ of the NBDs close to the inner leaflet of the membrane. Structural data for CFTR imply that this region may also be important as a location for the unique regulatory (R)—region of CFTR. Hence the linkers may have evolved to interact with the CFTR regulatory machinery and perhaps to act as a loose binding site for this otherwise disordered part of the CFTR structure. In the first half of the protein, the TMD–NBD linker region is closely followed by a disordered region in NBD1, the so-called regulatory insertion (RI). This region seems to be specialised to CFTR, but deletion of the RI results in a NBD1 that behaves better in terms of solubility (Lewis et al. 2005; Protasevich et al. 2010).
Linear representation of the CFTR sequence with the major domains indicated above. The plot shows the SCA score for three selected residues in transmembrane helix 6 versus every other residue (see main text), (Mendoza et al. 2012). Key: R334 (blue), I336 (red), T338 (green). Regions with low scores correspond to regions that are likely to be specialised in CFTR channel and maturation functions, hence showing little or no conservation with other ABC proteins. Some of these regions are also likely to be structurally disordered. Interestingly the N-terminal region is thought to be disordered, but does not display scores close to zero hence may show more homology with other ABC proteins. The TMD–NBD linker regions are not disordered in available ABC protein exporter structures, but the CFTR linkers are quite different from other ABC proteins in terms of length as well as sequence (Hunt et al. 2013). In the first half of the protein, the TMD–NBD linker is closely followed by a disordered region in NBD1, the so-called regulatory insertion (RI)
Secondary and Tertiary CFTR Structure
To date, there is no high-resolution X-ray crystal structure of full-length CFTR nor for any other ABCC family member. However, electron crystallography methodology has been used to generate a Coulomb density map for CFTR at 9 Å resolution (or more correctly 1/9 Å−1) (Rosenberg et al. 2011), and single-particle analysis has been used to solve a number of low-resolution structures (Mio et al. 2008; Zhang et al. 2009, 2010). The electron crystallography methodology has also revealed the structure of another ABCC family member (ABCC1/MRP1) (Rosenberg et al. 2010), and single-particle analysis combined with electron microscopy has provided structural data for the hetero-octameric ABCC8/SUR1 complex (Mikhailov et al. 2005). A number of computational atomic models of CFTR have been generated, based on similarity at the primary structure level with a bacterial homologue Sav1866 or the eukaryotic P-gp/ABCB1 (Mornon et al. 2008; Rahman et al. 2013; Serohijos et al. 2008). Structures of the isolated soluble domains of CFTR have also been reported, with the X-ray crystal structures of NBD1 (Lewis et al. 2004) and NBD2 (PDB: 3GD7, unpublished). Nuclear magnetic resonance (NMR) structures of NBD1 (Baker et al. 2007; Bozoky et al. 2013a, b; Hudson et al. 2012; Kanelis et al. 2010) and the (mostly disordered) isolated R region have also been reported (Baker et al. 2007).
The greatest detail in current CFTR structural knowledge comes from the X-ray structures of its NBDs, as reviewed recently (Hunt et al. 2013). Overall, the NBDs of CFTR have similar structural folds to those of other ABC proteins, including the common Walker A, Walker B and signature motifs and this is covered elsewhere in this book in detail (Lewis et al. 2004). The NBDs of CFTR also have a few unique features compared to other ABC proteins: Firstly, CFTR NBD1 has an additional ~35-residue regulatory insertion (RI) and the NBD1 structure probably includes a short extension that represents the first segment of the regulatory (R) region. Note that the atomic coordinates are available for NBD2 of CFTR (PDBID = 3GD7), however there is currently no publication describing the work behind the structural analysis. The NBDs of CFTR are not very symmetrical in terms of their primary structure, with less than 30 % identity between NBD1 and NBD2 sequences (Klein et al. 1999), compared to above 70 % identity in other ABC proteins (Higgins 1992). CFTR has changes in the signature sequence in NBD2 (LSGGQ to LSHGH) resulting in only one active ATP hydrolytic site, although both sites still have the ability to tightly bind ATP (Aleksandrov et al. 2002; Lewis et al. 2004; Thibodeau et al. 2005). Such strong NBD asymmetry is found with several other ABC proteins, e.g. MRP1/ABCC1 (Hou et al. 2000; Jones and George 2004). Structural and biophysical studies have suggested that the NBDs of CFTR form the same ATP-dependent head-to-tail sandwich dimer conformation as other ABC proteins (Aleksandrov et al. 2009; Jones and George 2004; Mense et al. 2006; Rosenberg et al. 2011; Vergani et al. 2005).
The two TMDs of CFTR each consist of the usually encountered six transmembrane α-helices with three extracellular loops (ECLs) and two long intracellular loops (ICLs). The longest ECL is ECL4 in TMD2 which is also N-glycosylated (Chang et al. 2008). In contrast to other ABC transporters, the TMDs of CFTR are expected to form a continuous pore through which ions can pass passively (Gadsby et al. 2006; Riordan 2008). For other ABC transporters the TMDs form a binding site (or binding sites) for the transported substrates that can be alternately exposed to the outside or inside milieu via conformational changes (Jardetzky 1966). Comparison of the CFTR electron crystallography map to the X-ray crystal structure of a homologous bacterial ABC transporter Sav1866 has revealed a lozenge-shaped density between TM helices 3, 6, 9 and 12 that may account for the location of a channel pore-regulating gate (Rosenberg et al. 2011). This position correlates with the expected locations of biochemically identified pore-lining residues (Linsdell 2005; Smith et al. 2001) in CFTR structural models (Mornon et al. 2008; Serohijos et al. 2008). The pore appears to have a deep, wide vestibule on the extracellular side, which may be at least partially accessible via the outer leaflet of the lipid bilayer.
The ICLs are predicted to form part of the typical ABC protein coupling helices that form the interface between TMDs and NBDs (Serohijos et al. 2008). The isolated R region is a highly dynamic and disordered structure that is mostly predicted to be random coil and with around 5 % α-helical secondary structure, measured by NMR studies (Baker et al. 2007; Ostedgaard et al. 2000). NMR methods have also detected interactions between the R region and NBD1, NBD2 and the C-terminus when the R region is mixed with the other domain in vitro (Baker et al. 2007; Bozoky et al. 2013b; Kanelis et al. 2010). Electron microscopy studies have tentatively assigned at least part of the R region to a location on the ‘shoulder’ of one NBD and close to the lipid bilayer (Rosenberg et al. 2011; Zhang et al. 2010). The R region is highly charged and contains several target sites for protein kinase A (PKA) phosphorylation (Cheng et al. 1991; Csanady et al. 2005; Riordan et al. 1989), as well as being a substrate for other kinases (Chappe et al. 2003; French et al. 1995). Phosphorylation of the R region regulates CFTR channel function (Csanady et al. 2005; Dahan et al. 2001; Gregory et al. 1990; Seibert et al. 1999), but no single phosphorylation site seems to be indispensible. The disordered structure of the R region is likely to be important to maximise accessibility of kinases for phosphorylation (Chong et al. 2013), however how phosphorylation of a region that is disordered then goes on to promote the activity of the channel is not well understood.
Electron microscopy studies of full-length CFTR showed that the protein is homologous to X-ray crystal structures of other ABC exporters, in particular Sav1866 (Dawson and Locher 2006). Such data have also shown that CFTR can display an outward-facing conformation in the absence of ATP (Rosenberg et al. 2004, 2011; Zhang et al. 2009, 2010). This is somewhat at odds with the idea that the inward- and outward-facing conformations of CFTR represent the closed and open channel configurations, respectively (Vergani et al. 2005) and (Wang and Linsdell 2012). At first sight, therefore, the structural data seem to indicate a structural miscoupling in CFTR between ATP binding and hydrolysis and switching between outward-facing and inward-facing conformations, respectively. However, other structural studies of ABC transporters have also revealed a slightly less than obligatory coupling between nucleotide status and conformation. For example, inward-facing conformations of the mitochondrial ABC transporter ABCB10 have been observed in the presence of nucleotide (Shintre et al. 2013). Similarly, outward-facing conformations in a pretranslocation state have been reported for a nucleotide-free state of the maltose ABC transporter MalFGK (Oldham et al. 2013).
The quaternary structure of CFTR has also been inferred from biophysical data. CFTR monomers were crystallised in two-dimensional arrays that were two molecular layers thick (Rosenberg et al. 2011). Conversely, single-particle analysis revealed dimeric CFTR particles (Awayn et al. 2005; Mio et al. 2008; Zhang et al. 2009) in the same detergent, whilst more recent studies of CFTR in novel facial detergent showed a monomeric state (Hildebrandt et al. 2014). It is important to note that all these structures were obtained for CFTR in different detergent micelles and may not reflect the oligomeric status of the protein in a biological membrane. The quaternary structure of CFTR in vivo still remains a matter of debate, especially as both monomers (Chen et al. 2002; Haggie and Verkman 2008; Ramjeesingh et al. 2001) and dimers (Eskandari et al. 1998; Ramjeesingh et al. 2001; Schillers et al. 2004) have been proposed for CFTR that was integrated into a lipid membrane environment.
CFTR Tertiary Structure Explored by Electron Crystallography
The highest resolution experimentally determined structure for the full-length CFTR protein has been derived from electron crystallography of two-dimensional crystals. These are crystals that are a single or few molecules thick, hence are very fragile and must be supported on a flat surface provided by the carbon film of an electron microscopy grid. The spatial resolution of the CFTR map derived from these crystals was reported to be at about 1/9 Å−1 (i.e. densities about 9 Å apart can just be resolved from each other). The thickness of the crystals (circa 300 Å, composed of two CFTR molecular layers) as well as the low symmetry (C1 symmetry) required a lengthy and painstaking data collection and processing. Nevertheless, at this level of resolution (1/9 Å−1), domains should be readily identifiable and some long transmembrane helices should also be distinguishable (especially if they are well separated from the main helical bundle). For reference, a theoretical study using the ABC transporter BtuCD (Ford and Holzenburg 2008) showed that even with the usual signal: noise and missing cone defects in electron crystallography data, it should be possible to resolve transmembrane helices at about 1/8 Å−1 and that at 1/10 Å−1 well-separated and long helices in the BtuCD structure could still be discerned. On the other hand, beta strands and short helices in the NBD region could not be discerned—even at 1/8 Å−1 resolution.
A good example of the restriction of resolution on the interpretation of the CFTR map is shown in Fig. 2. Elongated segments of density from the experimental CFTR map can be fitted with transmembrane alpha helices. In this case the helices are shown well separated from surrounding densities. Interpretation of the map using a published homology model for CFTR (Mornon et al. 2009, 2014) assigns the longest helical density to the seventh transmembrane helix, residues 860–891. This helix and the nearby transmembrane helix 8 could be more confidently assigned because they extend much further on the extracellular side of the protein than the other transmembrane helices. The large extracellular loop that links these two transmembrane helices (ECL4) is the site of glycosylation in CFTR (arrows, centre panel, Fig. 2, see also Fig. 1). In contrast, short helices, closely packed helices and beta sheets are not easily distinguishable at this resolution (see Fig. 2, all panels, for examples of this).
a Experimental outward-facing CFTR structure derived from electron crystallography (yellow mesh, left) with gaping TMDs at the top and closely contacting NBDs at the bottom. The map is at a resolution where long and well-separated transmembrane helices can be distinguished, as shown in the 1-nm thick slice through the map (right). CFTR closely resembles other structures for ABC transporters in the outward-facing state, and the map can be interpreted using a homology model based on the Sav1866 transporter (Dawson and Locher 2006; Mornon et al. 2009) shown in the left. The blue ribbon trace is from the N-terminal half of the model, green trace is from the C-terminal half. Potential flexible regions (N-terminal 80 residues, R region, RI region) were removed from the model before fitting to the map. The final correlation coefficient between the CFTR map and a suitably resolution-adjusted map calculated from the model was about 0.65 (where 1.0 would be perfect correspondence, and Sav1866 fitting gave 0.56), reflecting the systematic deviation from the symmetric Sav1866 structure during evolutionary timescales. Models can be further refined with some local flexible fitting (Senderowitz, Bar-Ilan University, unpublished data) with a correlation coefficient approaching 0.9. b Illustration of the pivotal position occupied by F508 (red space-filling atoms) in the model. It sits at the interface between ICL4 in TMD2 (green ribbon) and NBD1 (blue ribbon, bottom). The position of F508 in the model corresponds to a small cavity in the experimental map, again pointing to the local discrepancies between the Sav1866-based homology model and CFTR, as discussed below
CFTR Mutations
To date, there are over 2000 different CFTR mutations that are logged as CF-causing. However, a high proportion of CF patients have protein with deletion of phenylalanine at position 508 (F508del) on at least one chromosome (Cystic Fibrosis Mutations Database, available at http://www.genet.sickkids.on.ca/cftr/). All heterozygotes with one WT copy of the gene display no disease (Cutting 2005; Riordan et al. 1989). CFTR mutations are functionally classified into several groups (Prickett and Jain 2013): Class 1 mutations cause a defect in CFTR protein synthesis, such as the premature stop codon W1282X, that results in little or no CFTR at the plasma membrane (PM). Class 2 mutations, including the common F508del, are translated into full-length nascent polypeptide chains but are defective in folding and are thus targeted for degradation rather than trafficked to the PM. Class 3 mutants of CFTR are able to reach the PM but have channel gating defects that decrease channel open time and decrease chloride flux, e.g. the second most commonly encountered mutation, G551D. Class 4 mutants reach the PM, but with decreased channel conductance. Class 5 mutants represent a fully functional CFTR at the PM but with reduced levels due to defective mRNA splicing. Classes 1–3 are associated with more severe disease phenotypes, whereas classes 4 and 5 are thought to be milder disease-causing mutations. It should be noted that some CFTR mutations may have more than one effect, for example the F508del mutation has reduced channel activity and shorter PM half-life in addition to the maturation and processing defects (Hwang et al. 1997).
As the vast majority of patients harbour F508del CFTR, this mutation has received the most attention. The ΔF508 mutation destabilises CFTR structure and folding (Lewis et al. 2005) and during protein synthesis, most of the expressed F508del CFTR is targeted by the endoplasmic-reticulum-associated protein degradation (ERAD) pathway for degradation by the proteasome, with only 1 % of translated protein reaching the PM (Kopito 1999; Ward et al. 1995). The small amount of ΔF508-CFTR that reaches the PM is a functional channel, although operating with slightly altered gating, namely with more time spent in the closed state (Cui et al. 2006). Significant levels of misfolding might cause gross changes in the overall F508del-CFTR structure, which could be detected by e.g. increased susceptibility to limited proteolysis (Hoelen et al. 2010; Peters et al. 2011), see also (Zhang et al. 1998). Surprisingly, however, the X-ray crystal structures of isolated F508del-NBD1 showed little structural differences compared to wild-type CFTR (Lewis et al. 2005, 2010). This has led to the hypothesis that F508 is important in inter-domain stabilisation and assembly, rather than the folding of NBD1 itself (Lewis et al. 2005, 2010; Loo et al. 2010; Rabeh et al. 2012; Serohijos et al. 2008; Thibodeau et al. 2005). The F508 residue lies on the surface of the NBD1 in an area that, in the X-ray crystal structures of other ABC proteins, forms crucial interactions with the second intracytoplasmic loop of the opposing TMD region (Aller et al. 2009; Dawson and Locher 2006). Hence the F508-mediated interaction in CFTR is between NBD1 and ICL4 of TMD2 (Cui et al. 2007; Lewis et al. 2010; Loo et al. 2010; Rabeh et al. 2012; Serohijos et al. 2008; Thibodeau et al. 2005). It has also been suggested that the peptide backbone at position 508 is important for CFTR folding but that the phenyalanine side chain is necessary for inter-domain contacts and stability of the folded protein (Thibodeau et al. 2005).
As well as the destabilisation of NBD1–TMD2 interactions, F508del-NBD1 itself appears to be globally destabilised. Recombinant isolated F508del-NBD1 has a thermal unfolding transition about 6 °C lower than its wild-type counterpart (Protasevich et al. 2010). The F508del mutation is proposed to thermodynamically favour the formation of a molten-globule state that is prone to aggregation (Wang et al. 2010). The thermal destabilisation effect of the F508del mutation is also inferred from in vivo experiments showing that low temperature growth conditions are able to rescue F508del CFTR allowing it to progress to the PM (Denning et al. 1992). Molecules that can bind and thermostabilise F508del NBD1 in vitro have also been effective in correcting full-length F508del CFTR in cells (Sampson et al. 2011). However, there is some controversy over whether F508del CFTR can be rescued by stabilising NBD1 alone (Aleksandrov et al. 2012), or whether the correction of NBD1–TMD2 interactions is also required (Rabeh et al. 2012). A combination of both may be required.
Mapping of Common CF-Causing Missense Mutations Within the CFTR Structure
The large majority of mutations catalogued in the CF mutation database have not been well characterised in terms of their functional effects on the protein, and the rarer mutations inevitably occur in combination with a more commonly occurring mutation on the other copy of the cftr gene. In response to this deficit in our knowledge, the most commonly encountered mutations in CF patients have been included in a study of their effects at the cellular level—the so-called CFTR2 study (http://www.cftr2.org/). The common missense mutations in this study can be readily mapped on to the existing CFTR homology models and within the electron crystallography-derived Coulomb density map as shown in Fig. 2, hence it is possible for the first time to begin to correlate the functional effects of mutations with their likely structural context.
F508del is the most common disease-causing mutation by far. Does the CFTR structure explain why the loss of this single residue causes such a major temperature-dependent defect on the protein? Panel b in Fig. 2 shows the location of F508 in the density map (arrow). This view emphasises the importance of the F508 residue in mediating interactions between the first NBD (blue) and the second TMD, in particular ICL4 (green). Loss of the F508 residue would remove a contact point between the domains, which lies in a narrow bridge of density with little surrounding density. Removal of this bridging residue would be predicted to destabilise the tertiary structure of CFTR to a greater extent than its local effects on the stability of the isolated NBD1 (Protasevich et al. 2010). It is also interesting to note that the expected position of F508 in the CFTR density map corresponds approximately with a small cavity in the map that would not be expected to be there from the homology model. Such density voids in experimental maps may arise for the obvious reason—that this part of the map is filled with solvent rather than protein. However, voids can also occur if there are variable conformations for the protein in this particular region. This arises because the map is derived from the summation of data from many millions of molecules. Hence a region where the protein contributes density but where each molecule has a slightly different configuration could result in a lower (but not zero) density. Since maps such as the one shown in Fig. 2 are generally ‘binary’ in the sense that a given density threshold is chosen as a cut-off for displaying the map, such subtleties in the map can often be lost and a solvent-filled void may not be distinguishable from a region of disorder. On the other hand, use of a low density threshold runs the risk of including noisy features in the interpretation of the map. Tentative inspection of the map in this region with a lower density threshold implies that there is still a small cavity formed between F508 and residues that would contribute density in this region from ICL4 of CFTR, hence arguing for better homology models as well as better experimental data.
Of the well-characterised missense mutations listed in the CFTR2 database, there are more disease-causing mutations in the first half of the protein and especially in NBD1, but none in the linking regulatory region. It would seem reasonable that missense mutations in the less ordered R region could be tolerated with little impact on the functioning of the protein. Perhaps unsurprisingly, there is a clustering of disease-causing missense mutations in the three-dimensional structure, which may provide insights into the likely consequences of these mutations as well as into the most crucial regions of CFTR for its function. For example, if just the 30 most common disease-causing missense mutations in humans are considered, then two discrete clusters become obvious (Fig. 3a, b): One cluster, mostly in NBD1, probably relates to the previously discussed sensitive position of the ICL4/NBD1 interface where F508 is positioned. However, the homology model based on a bacterial transporter may not completely reflect the arrangement of residues in the CFTR structure, hence it is useful to superimpose the experimental map for CFTR on to the model. For example, Fig. 3c displays the map around the cluster of missense mutations close to F508, which is composed of F508, I507, V520, A559 and R560 in NBD1 and R1066, L1065 and L1077 in ICL4. As mentioned above, F508 sits close to a cavity in the map, suggesting contacts in this region between NBD1 and ICL4 in CFTR are much less robust than in the Sav1866 structure (Dawson and Locher 2006; Mornon et al. 2009, 2014).
a Location of residues in the N-terminal half of CFTR that, when mutated to other residues, causes CF. Of the 20 most commonly encountered mutations, 18 are displayed. P67 and R74 are N-terminal residues missing from the fitting, but present in the originally published model, and probably reside in the ‘elbow’ helix that lies parallel to the surface of the cytoplasmic side of the membrane. Residues in the first half of CFTR are coloured light blue and disease-causing mutated residues in space filling/dark blue. b Location of residues in the C-terminal half of CFTR that when mutated causes disease. The 10 most commonly encountered are shown as space-filling representations in yellow. c Zoom into the cluster of residues related to disease at the interface between NBD1 and ECL4 in TMD2 close to F508 (space filling) within the context of the experimental map (red surface with slice plane transparent grey). Note the small cavity in the map in the region of the F508 residue that is discussed in the text. d Focus on residues associated with disease-causing missense mutations in ICL2 in TMD1 that are close to N1303 in NBD2. e Stereo image looking down the central pore formed by the TMDs and from the extracellular side, with residues associated with disease-causing mutations shown as space-filling representations. A cluster of residues associated with channel gating dysfunction (when mutated) is present top left. The experimental map shows a cleft on the right-hand side that is not present on the left
A second cluster of more common missense mutations causing the CF disease is found mainly in TMD1 and appears to relate to pore-lining or gating residues that follow a path from one side of the membrane to the other. This cluster is composed of R334, I336 and T338 on the extracellular side and L206, R347, R352 towards the centre of the membrane whilst D1152 in TMD2 is further on the cytoplasmic side of the membrane. A stereo image is included to be able to visualise this pathway through the CFTR channel (Fig. 3e). Note that the majority of the disease-causing mutations are on the left-hand side of the figure and cluster in TMD1 (blue residues). Few commonly occurring missense mutations appear on the right (TMD2, yellow residues). The experimental map shows a more significant deviation from twofold symmetry than the homology model (which is biased by the twofold symmetry of the Sav1866 transporter used as the model). The most obvious manifestation of this asymmetry is the lack of a pronounced gap on the left-hand side of the protein. In the Sav1866 structure, gaps communicating to the extracellular leaflet of the lipid bilayer exist on both sides of the transporter and these gaps have been reported for other ABC transporters (Ward et al. 2007). This asymmetry in the CFTR structure may be a reflection of the evolution from a transporter to a channel, and may explain why common disease-causing mutations cluster on one side of the transmembrane region rather on the side that resembles more closely the Sav1866 transporter.
Is there a structural and functional equivalent of F508 in NBD2 or is the asymmetry in CFTR sufficient to distinguish between the interaction sites at NBD1/ICL4 versus NBD2/ICL2? The most obvious analogue for F508 in NBD2 is an asparagine residue, N1303. Mutation of this residue to lysine (N1303 K) is the third most common missense mutation, present in 1200 patients worldwide, or about 1.5–2 % of alleles. Like F508del, N1303K leads to failure of maturation of the mutated protein (Sosnay et al. 2011). N1303 deletion does not appear in the CF mutation database, however. There are several residues in ICL2 of TMD1 that are predicted to pack closely with N1303 and these also result in disease when mutated or deleted. Methionine 281 is most closely associated with N1303 in the modelling (Fig. 3d). The M281T mutation has been associated with pancreatitis (de Cid et al. 2010), but because of its rarity, is not one of the mutations studied so far in the first sweep of the CFTR2 study (Sosnay et al. 2013). Nearby residues in ICL2 have also been reported as CF-causing (W277R, I285F) and their locations with respect to N1303 are shown in Fig. 3d. A deletion of a residue in this region of CFTR can also be disease-causing (E278del), although in this case the deleted residue is found in ICL2 rather than in NBD2 and a glutamate residue is much less bulky than phenylalanine. These ICL2/NBD2 interface residues lie in regions that show a significant co-evolution in ABC proteins (Mendoza et al. 2012) as shown in Figs. 1 and 4 panels a and c. Interestingly, two of the main peaks in the co-evolution scoring for N1303 correspond to residues that become disease-causing when mutated to other residues (e.g. W277R, M281T—but not S269). Worthy of mention is the striking period of 4 oscillation that is displayed when several residues flanking N1303 are included in the correlation plots versus ICL2. This probably arises because of the helical secondary structure flanking ICL2 and reflects the likelihood that every fourth residue in a helix will occupy the same position, approximately, along the length of the helix.
Co-evolution of residues in ICL2 versus N1303 in NBD2 (panel a) and in ICL4 versus F508 in NBD1 (panel c). Mutations of W277 and M281 in ICL2 are CF-causing (see main text). Panel c Correlation scores for several residues in the vicinity of N1303 versus residues in ICL2. A period of 4 oscillation is apparent in the SCA scores (cylinders) that is probably reflecting the helical secondary structure flanking the linking non-helical connecting loop (double headed arrow). A similar profile is observed for residues in ICL4 versus residues around F508 in NBD1 (panels b, d), although with a less clear-cut pattern, perhaps reflecting the relative looseness of this interface as discussed in the main text. Data extracted from the database is described in (Mendoza et al. 2012)
Similar co-evolution scoring patterns exist for F508 and ICL4 residues (see panel b, Fig. 4). In this case, the main peak in the scoring corresponds to L1065 (can be mutated to F/R/P) and R1066 (can be mutated to C/S/H/L). Both of these residues when mutated can cause CF. The second noticeable peak corresponds to G1061 which is also disease-causing when mutated to arginine. Surprisingly, R1070 does not score highly, despite it being a common disease-causing locus. For example, R1070 W is disease-causing on its own, but when in combination with F508del, W at position 1070 can alleviate the effects of the nearby phenylalanine loss. The period of 4 oscillation in the correlation with ICL4 residues for corresponding residues around F508 del is also apparent in panel d of Fig. 4, but the oscillation is noticeably weaker. This may be a reflection of deviation of a strict helical secondary structure in this region of the protein. It is also possible that this is a reflection of the weaker density in this region of the experimental CFTR density map (Figs. 2 and 3) and the apparent void close to F508 and ICL4. It has been speculated that this region of CFTR is the ‘Achilles Heel’ of the CFTR protein (Rosenberg et al. 2011).
Conclusions
CFTR is one of the best-studied ABC transporters (but with the caveat that it is not a transporter). The intense research efforts on CFTR are being driven in response to the devastating effects of mutations in the CFTR gene and their relative frequency in one part of the human population. Fortunately, these large-scale efforts into understanding CFTR have been generously funded by charities such as the US—based Cystic Fibrosis Foundation. Encouragingly, there are now some new drugs that have proven to be very effective at treating the basic defect in the disease—and at least for some mutations and patients, a cure seems very likely (Sosnay et al. 2013; Van Goor et al. 2009, 2011).
It could be argued that CFTR can give us a unique insight into structure–function relationships for ABC transporters and so merits a place in this book despite its lack of transport activity. The huge database that exists for CF-causing mutations is invaluable in the ABC transporter field,—see http://www.genet.sickkids.on.ca—and the work of the CFTR2 consortium will build on that knowledge with more direct functional data for disease-causing mutations (Sosnay et al. 2013). We should also not forget that, uniquely in the ABC transporter field, the activity of CFTR could be monitored at the single molecule level and with very high temporal resolution (Aleksandrov et al. 2007; Hwang and Sheppard 2009). This has allowed a detailed dissection of some aspects of its function that are the envy of researchers working on other ABC transporters.
In some other areas, however, progress in CFTR research has been very difficult compared to other ABC proteins. Expression levels of CFTR in normal epithelial cells are quite low. The most enriched source of CFTR has been reported to be from the rectal gland of sharks (Riordan et al. 1994) and, perhaps not too surprisingly, isolation of the protein from naturally expressing cells has been a highly specialized activity. Overexpression of the protein in stably- or transiently transformed cells has been the main approach for the subsequent physical isolation of the protein. In these cell types a high proportion of even the wild-type protein is degraded before it can reach the plasma membrane, and the large flexible regions of CFTR such as the regulatory region (see Fig. 1) are prime targets for intrinsic proteases. The inherent instability of the protein is compounded by a lack of solubility in many detergents. Some detergents based on lysolipids have been reported to efficiently solubilise the protein (Huang et al. 1998; Matar-Merheb et al. 2011; Pollock et al. 2014), but these may be too harsh for the full preservation of CFTR activity (Matar-Merheb et al. 2011; Pollock et al. 2014).
As a result of the difficulties in biochemical isolation, only a few studies have been carried out on the (relatively modest) ATPase activity of purified CFTR and our understanding of how it correlates with channel opening is still sketchy. Unlike other ABC transporters, there is no substrate-induced stimulation of ATPase activity in CFTR that would allow one to distinguish its activity from contaminating ATPases. There is some evidence that phosphorylation of the protein by protein kinase A (PKA) stimulates the ATPase activity of CFTR, and this would make some sense in terms of the prerequisite of phosphorylation for channel opening (and subsequent closing) (Eckford et al. 2012). However PKA is, itself, a good ATPase and hence must be removed from CFTR before the latter protein’s activity can be assayed. This adds to the difficulty of the assay. Like many other ABC transporters, the ATPase activity of CFTR preparations is strongly influenced by the detergent employed in the purification scheme and depends on whether the protein is reconstituted into proteoliposomes or not (Pollock et al. 2014). Despite these difficulties, there has been significant progress over the last few years. New yeast expression systems for the protein have been reported to yield milligram quantities of the purified protein (Pollock et al. 2014) and these reagents have been made available to the general CF research community. Technology for mammalian cell expression continues to improve, making it possible to use these systems for production of significant quantities of the protein. Thus it seems promising that CFTR biochemistry will soon become a standard approach in many laboratories around the world, leading to a much better understanding of its function and of the evolution of the ABC transporter family in general.
Abbreviations
- ABC:
-
ATP-binding cassette
- C:
-
Carboxy-terminal
- CF:
-
Cystic fibrosis
- CFF:
-
Cystic fibrosis foundation
- CFTR:
-
Cystic fibrosis transmembrane conductance regulator
- DDM:
-
Dodecyl maltoside
- EM:
-
Electron microscopy
- ECL:
-
Extracellular loop
- GPCR:
-
G-protein coupled receptor
- ICL:
-
Intracytoplasmic loop
- N:
-
Amino-terminal
- NBD:
-
Nucleotide-binding domain
- PDB:
-
Protein data bank
- PM:
-
Plasma membrane
- RI:
-
Regulatory insertion
- R:
-
Regulatory (region or domain)
- SCA:
-
Statistical coupling analysis
- TMD:
-
Transmembrane domain
References
Aleksandrov L, Aleksandrov AA, Chang XB, Riordan JR (2002) The first nucleotide binding domain of cystic fibrosis transmembrane conductance regulator is a site of stable nucleotide interaction, whereas the second is a site of rapid turnover. J Biol Chem 277:15419–15425
Aleksandrov AA, Aleksandrov LA, Riordan JR (2007) CFTR (ABCC7) is a hydrolyzable-ligand-gated channel. Pflugers Arch 453:693–702
Aleksandrov AA, Cui L, Riordan JR (2009) Relationship between nucleotide binding and ion channel gating in cystic fibrosis transmembrane conductance regulator. J Physiol 587:2875–2886
Aleksandrov AA, Kota P, Cui L, Jensen T, Alekseev AE, Reyes S, He L, Gentzsch M, Aleksandrov LA, Dokholyan NV, Riordan JR (2012) Allosteric modulation balances thermodynamic stability and restores function of ΔF508 CFTR. J Mol Biol 419:41–60
Aller S, Yu J, Ward A, Weng Y, Chittaboina S, Zhuo R, Harrell P, Trinh Y, Zhang Q, Urbatsch I, Chang G (2009) Structure of P-glycoprotein reveals a molecular basis for poly-specific drug binding. Science 1718–1722
Awayn NH, Rosenberg MF, Kamis AB, Aleksandrov LA, Riordan JR, Ford RC (2005) Crystallographic and single-particle analyses of native- and nucleotide-bound forms of the cystic fibrosis transmembrane conductance regulator (CFTR) protein. Biochem Soc Trans 33:996–999
Baker JM, Hudson RP, Kanelis V, Choy WY, Thibodeau PH, Thomas PJ, Forman-Kay JD (2007) CFTR regulatory region interacts with NBD1 predominantly via multiple transient helices. Nat Struct Mol Biol 14:738–745
Bozoky Z, Krzeminski M, Chong PA, Forman-Kay JD (2013a) Structural changes of CFTR R region upon phosphorylation: a plastic platform for intramolecular and intermolecular interactions. FEBS J 280:4407–4416
Bozoky Z, Krzeminski M, Muhandiram R, Birtley JR, Al-Zahrani A, Thomas PJ, Frizzell RA, Ford RC, Forman-Kay JD (2013b) Regulatory R region of the CFTR chloride channel is a dynamic integrator of phospho-dependent intra- and intermolecular interactions. In: Proceedings of the National Academy of Sciences of the United States of America
Cant N, Pollock N, Ford RC (2014) CFTR structure and cystic fibrosis. Int J Biochem Cell Biol 52:15–25
Chang X, Mengos A, Hou Y, Cui L, Jensen T, Aleksandrov A, Riordan J, Gentzsch M (2008) Role of N-linked oligosaccharides in the biosynthetic processing of the cystic fibrosis membrane conductance regulator. J Cell Sci 121:2814–2823
Chappe V, Hinkson DA, Zhu T, Chang XB, Riordan JR, Hanrahan JW (2003) Phosphorylation of protein kinase C sites in NBD1 and the R domain control CFTR channel activation by PKA. J Physiol 548:39–52
Chen JH, Chang XB, Aleksandrov AA, Riordan JR (2002) CFTR is a monomer: biochemical and functional evidence. J Membr Biol 188:55–71
Cheng SH, Rich DP, Marshall J, Gregory RJ, Welsh MJ, Smith AE (1991) Phosphorylation of the R domain by cAMP-dependent protein kinase regulates the CFTR chloride channel. Cell 66:1027–1036
Chong PA, Kota P, Dokholyan NV, Forman-Kay JD (2013) Dynamics intrinsic to cystic fibrosis transmembrane conductance regulator function and stability. Cold Spring Harb Perspect Med 3:a009522
Clunes LA, Davies CM, Coakley RD, Aleksandrov AA, Henderson AG, Zeman KL, Worthington EN, Gentzsch M, Kreda SM, Cholon D, Bennett WD, Riordan JR, Boucher RC, Tarran R (2012) Cigarette smoke exposure induces CFTR internalization and insolubility, leading to airway surface liquid dehydration. FASEB J (official publication of the Federation of American Societies for Experimental Biology) 26:533–545
Csanady L, Seto-Young D, Chan KW, Cenciarelli C, Angel BB, Qin J, McLachlin DT, Krutchinsky AN, Chait BT, Nairn AC, Gadsby DC (2005) Preferential phosphorylation of R-domain Serine 768 dampens activation of CFTR channels by PKA. J Gen Physiol 125:171–186
Cui L, Aleksandrov L, Hou YX, Gentzsch M, Chen JH, Riordan JR, Aleksandrov AA (2006) The role of cystic fibrosis transmembrane conductance regulator phenylalanine 508 side chain in ion channel gating. J Physiol 572:347–358
Cui L, Aleksandrov L, Chang XB, Hou YX, He L, Hegedus T, Gentzsch M, Aleksandrov A, Balch WE, Riordan JR (2007) Domain interdependence in the biosynthetic assembly of CFTR. J Mol Biol 365:981–994
Cutting GR (2005) Modifier genetics: cystic fibrosis. Annu Rev Genomics Hum Genet 6:237–260
Dahan D, Evagelidis A, Hanrahan JW, Hinkson DA, Jia Y, Luo J, Zhu T (2001) Regulation of the CFTR channel by phosphorylation. Pflugers Arch 443(Suppl 1):S92–S96
Dawson RJ, Locher KP (2006) Structure of a bacterial multidrug ABC transporter. Nature 443:180–185
de Cid R, Ramos MD, Aparisi L, Garcia C, Mora J, Estivill X, Farre A, Casals T (2010) Independent contribution of common CFTR variants to chronic pancreatitis. Pancreas 39:209–215
Dean M, Allikmets R (2001) Complete characterization of the human ABC gene family. J Bioenerg Biomembr 33:475–479
Denning GM, Anderson MP, Amara JF, Marshall J, Smith AE, Welsh MJ (1992) Processing of mutant cystic fibrosis transmembrane conductance regulator is temperature-sensitive. Nature 358:761–764
Eckford PD, Li C, Ramjeesingh M, Bear CE (2012) Cystic fibrosis transmembrane conductance regulator (CFTR) potentiator VX-770 (ivacaftor) opens the defective channel gate of mutant CFTR in a phosphorylation-dependent but ATP-independent manner. J Biol Chem 287:36639–36649
Eskandari S, Wright EM, Kreman M, Starace DM, Zampighi GA (1998) Structural analysis of cloned plasma membrane proteins by freeze-fracture electron microscopy. Proc Natl Acad Sci USA 95:11235–11240
Ford RC, Holzenburg A (2008) Electron crystallography of biomolecules: mysterious membranes and missing cones. Trends Biochem Sci 33:38–43
French PJ, Bijman J, Edixhoven M, Vaandrager AB, Scholte BJ, Lohmann SM, Nairn AC, de Jonge HR (1995) Isotype-specific activation of cystic fibrosis transmembrane conductance regulator-chloride channels by cGMP-dependent protein kinase II. J Biol Chem 270:26626–26631
Gadsby DC, Vergani P, Csanady L (2006) The ABC protein turned chloride channel whose failure causes cystic fibrosis. Nature 440:477–483
Gregory RJ, Cheng SH, Rich DP, Marshall J, Paul S, Hehir K, Ostedgaard L, Klinger KW, Welsh MJ, Smith AE (1990) Expression and characterization of the cystic fibrosis transmembrane conductance regulator. Nature 347:382–386
Haggie PM, Verkman AS (2008) Monomeric CFTR in plasma membranes in live cells revealed by single molecule fluorescence imaging. J Biol Chem 283:23510–23513
Higgins C (1992) ABC transporters: from microorganisms to man. Annu Rev Cell Biol 8:67–113
Hildebrandt E, Zhang Q, Cant N, Ding H, Dai Q, Peng L, Fu Y, DeLucas LJ, Ford R, Kappes JC, Urbatsch IL (2014) A survey of detergents for the purification of stable, active human cystic fibrosis transmembrane conductance regulator (CFTR). Biochim Biophys Acta 1838:2825–2837
Hoelen H, Kleizen B, Schmidt A, Richardson J, Charitou P, Thomas PJ, Braakman I (2010) The primary folding defect and rescue of ΔF508 CFTR emerge during translation of the mutant domain. PLoS ONE 5:e15458
Hou Y, Cui L, Riordan JR, Chang X (2000) Allosteric interactions between the two non-equivalent nucleotide binding domains of multidrug resistance protein MRP1. J Biol Chem 275:20280–20287
Huang P, Liu Q, Scarborough GA (1998) Lysophosphatidylglycerol: a novel effective detergent for solubilizing and purifying the cystic fibrosis transmembrane conductance regulator. Anal Biochem 259:89–97
Hudson RP, Chong PA, Protasevich II, Vernon R, Noy E, Bihler H, An JL, Kalid O, Sela-Culang I, Mense M, Senderowitz H, Brouillette CG, Forman-Kay JD (2012) Conformational changes relevant to channel activity and folding within the first nucleotide binding domain of the cystic fibrosis transmembrane conductance regulator. J Biol Chem 287:28480–28494
Hunt JF, Wang C, Ford RC (2013) Cystic fibrosis transmembrane conductance regulator (ABCC7) structure. Cold Spring Harb Perspect Med 3:a009514
Hwang TC, Sheppard DN (2009) Gating of the CFTR Cl-channel by ATP-driven nucleotide-binding domain dimerisation. J Physiol 587:2151–2161
Hwang TC, Wang F, Yang IC, Reenstra WW (1997) Genistein potentiates wild-type and delta F508-CFTR channel activity. Am J Physiol 273:C988–C998
Jardetzky O (1966) Simple allosteric model for membrane pumps. Nature 211:969–970
Jones PM, George AM (2004) The ABC transporter structure and mechanism: perspectives on recent research. Cell Mol Life Sci 61:682–699
Kanelis V, Hudson RP, Thibodeau PH, Thomas PJ, Forman-Kay JD (2010) NMR evidence for differential phosphorylation-dependent interactions in WT and DeltaF508 CFTR. EMBO J 29:263–277
Klein I, Sarkadi B, Varadi A (1999) An inventory of the human ABC proteins. Biochim Biophys Acta 1461:237–262
Kopito RR (1999) Biosynthesis and degradation of CFTR. Physiol Rev 79:S167–S173
Lewis HA, Buchanan SG, Burley SK, Conners K, Dickey M, Dorwart M, Fowler R, Gao X, Guggino WB, Hendrickson WA, Hunt JF, Kearins MC, Lorimer D, Maloney PC, Post KW, Rajashankar KR, Rutter ME, Sauder JM, Shriver S, Thibodeau PH, Thomas PJ, Zhang M, Zhao X, Emtage S (2004) Structure of nucleotide-binding domain 1 of the cystic fibrosis transmembrane conductance regulator. EMBO J 23:282–293
Lewis HA, Zhao X, Wang C, Sauder JM, Rooney I, Noland BW, Lorimer D, Kearins MC, Conners K, Condon B, Maloney PC, Guggino WB, Hunt JF, Emtage S (2005) Impact of the deltaF508 mutation in first nucleotide-binding domain of human cystic fibrosis transmembrane conductance regulator on domain folding and structure. J Biol Chem 280:1346–1353
Lewis HA, Wang C, Zhao X, Hamuro Y, Conners K, Kearins MC, Lu F, Sauder JM, Molnar KS, Coales SJ, Maloney PC, Guggino WB, Wetmore DR, Weber PC, Hunt JF (2010) Structure and dynamics of NBD1 from CFTR characterized using crystallography and hydrogen/deuterium exchange mass spectrometry. J Mol Biol 396:406–430
Linsdell P (2005) Location of a common inhibitor binding site in the cytoplasmic vestibule of the cystic fibrosis transmembrane conductance regulator chloride channel pore. J Biol Chem 280:8945–8950
Lockless SW, Ranganathan R (1999) Evolutionarily conserved pathways of energetic connectivity in protein families. Science 286:295–299
Loo TW, Bartlett MC, Clarke DM (2010) The V510D suppressor mutation stabilizes DeltaF508-CFTR at the cell surface. Biochemistry 49:6352–6357
Matar-Merheb R, Rhimi M, Leydier A, Huche F, Galian C, Desuzinges-Mandon E, Ficheux D, Flot D, Aghajari N, Kahn R, Di Pietro A, Jault JM, Coleman AW, Falson P (2011) Structuring detergents for extracting and stabilizing functional membrane proteins. PLoS ONE 6:e18036
Mendoza JL, Schmidt A, Li Q, Nuvaga E, Barrett T, Bridges RJ, Feranchak AP, Brautigam CA, Thomas PJ (2012) Requirements for efficient correction of DeltaF508 CFTR revealed by analyses of evolved sequences. Cell 148:164–174
Mense M, Vergani P, White DM, Altberg G, Nairn AC, Gadsby DC (2006) In vivo phosphorylation of CFTR promotes formation of a nucleotide-binding domain heterodimer. EMBO J 25:4728–4739
Mikhailov MV, Campbell JD, de Wet H, Shimomura K, Zadek B, Collins RF, Sansom MS, Ford RC, Ashcroft FM (2005) 3-D structural and functional characterization of the purified KATP channel complex Kir6.2-SUR1. EMBO J 24:4166–4175
Mio K, Ogura T, Mio M, Shimizu H, Hwang TC, Sato C, Sohma Y (2008) Three-dimensional reconstruction of human cystic fibrosis transmembrane conductance regulator chloride channel revealed an ellipsoidal structure with orifices beneath the putative transmembrane domain. J Biol Chem 283:30300–30310
Mornon JP, Lehn P, Callebaut I (2008) Atomic model of human cystic fibrosis transmembrane conductance regulator: membrane-spanning domains and coupling interfaces. Cell Mol Life Sci 65:2594–2612
Mornon JP, Lehn P, Callebaut I (2009) Molecular models of the open and closed states of the whole human CFTR protein. Cell Mol Life Sci CMLS 66:3469–3486
Mornon JP, Hoffmann B, Jonic S, Lehn P, Callebaut I (2014) Full-open and closed CFTR channels, with lateral tunnels from the cytoplasm and an alternative position of the F508 region, as revealed by molecular dynamics. Cell Mol Life Sci CMLS
Oldham ML, Chen S, Chen J (2013) Structural basis for substrate specificity in the Escherichia coli maltose transport system. Proc Natl Acad Sci USA 110:18132–18137
Ostedgaard LS, Baldursson O, Vermeer DW, Welsh MJ, Robertson AD (2000) A functional R domain from cystic fibrosis transmembrane conductance regulator is predominantly unstructured in solution. Proc Natl Acad Sci USA 97:5657–5662
Peters KW, Okiyoneda T, Balch WE, Braakman I, Brodsky JL, Guggino WB, Penland CM, Pollard HB, Sorscher EJ, Skach WR, Thomas PJ, Lukacs GL, Frizzell RA (2011) CFTR folding consortium: methods available for studies of CFTR folding and correction. Methods Mol Biol 742:335–353
Pollock N, Cant N, Rimington T, Ford RC (2014) Purification of the cystic fibrosis transmembrane conductance regulator protein expressed in Saccharomyces cerevisiae. J Vis Exp JoVE
Prickett M, Jain M (2013) Gene therapy in cystic fibrosis. Transl Res 161:255–264
Protasevich I, Yang Z, Wang C, Atwell S, Zhao X, Emtage S, Wetmore D, Hunt JF, Brouillette CG (2010) Thermal unfolding studies show the disease causing F508del mutation in CFTR thermodynamically destabilizes nucleotide-binding domain 1. Protein Sci 19:1917–1931
Rabeh WM, Bossard F, Xu H, Okiyoneda T, Bagdany M, Mulvihill CM, Du K, di Bernardo S, Liu Y, Konermann L, Roldan A, Lukacs GL (2012) Correction of both NBD1 energetics and domain interface is required to restore ΔF508 CFTR folding and function. Cell 148:150–163
Rahman KS, Cui G, Harvey SC, McCarty NA (2013) Modeling the conformational changes underlying channel opening in CFTR. PLoS ONE 8:e74574
Ramjeesingh M, Li CH, Kogan I, Wang YC, Huan LJ, Bear CE (2001) A monomer is the minimum functional unit required for channel and ATPase activity of the cystic fibrosis transmembrane conductance regulator. Biochemistry 40:10700–10706
Riordan JR (2008) CFTR function and prospects for therapy. Annu Rev Biochem 77:701–726
Riordan J, Rommens J, Kerem B, Alon N, Rozmahel R, Grzelczak Z, Zielenski J, Lok S, Plavsic N, Chou J (1989) Identification of the cystic fibrosis gene: cloning and characterization of complementary DNA. Science 245:1066–1073
Riordan JR, Forbush B, Hanrahan JW (1994) The molecular basis of chloride transport in shark rectal gland. J Exp Biol 196:405–418
Rosenberg MF, Kamis AB, Aleksandrov LA, Ford RC, Riordan JR (2004) Purification and crystallization of the cystic fibrosis transmembrane conductance regulator (CFTR). J Biol Chem 279:39051–39057
Rosenberg MF, Oleschuk CJ, Wu P, Mao Q, Deeley RG, Cole SP, Ford RC (2010) Structure of a human multidrug transporter in an inward-facing conformation. J Struct Biol 170:540–547
Rosenberg MF, O’Ryan LP, Hughes G, Zhao Z, Aleksandrov LA, Riordan JR, Ford RC (2011) The cystic fibrosis transmembrane conductance regulator (CFTR): three-dimensional structure and localization of a channel gate. J Biol Chem 286:42647–42654
Sampson HM, Robert R, Liao J, Matthes E, Carlile GW, Hanrahan JW, Thomas DY (2011) Identification of a NBD1-binding pharmacological chaperone that corrects the trafficking defect of F508del-CFTR. Chem Biol 18:231–242
Schillers H, Shahin V, Albermann L, Schafer C, Oberleithner H (2004) Imaging CFTR: a tail to tail dimer with a central pore. Cell Physiol Biochem 14:1–10
Seibert FS, Chang XB, Aleksandrov AA, Clarke DM, Hanrahan JW, Riordan JR (1999) Influence of phosphorylation by protein kinase A on CFTR at the cell surface and endoplasmic reticulum. Biochim Biophys Acta 1461:275–283
Serohijos AW, Hegedus T, Aleksandrov AA, He L, Cui L, Dokholyan NV, Riordan JR (2008) Phenylalanine-508 mediates a cytoplasmic-membrane domain contact in the CFTR 3D structure crucial to assembly and channel function. Proc Natl Acad Sci USA 105:3256–3261
Shintre CA, Pike AC, Li Q, Kim JI, Barr AJ, Goubin S, Shrestha L, Yang J, Berridge G, Ross J, Stansfeld PJ, Sansom MS, Edwards AM, Bountra C, Marsden BD, von Delft F, Bullock AN, Gileadi O, Burgess-Brown NA, Carpenter EP (2013) Structures of ABCB10, a human ATP-binding cassette transporter in apo- and nucleotide-bound states. Proc Natl Acad Sci USA 110:9710–9715
Smith SS, Liu X, Zhang ZR, Sun F, Kriewall TE, McCarty NA, Dawson DC (2001) CFTR: covalent and noncovalent modification suggests a role for fixed charges in anion conduction. J Gen Physiol 118:407–431
Sosnay PR, Castellani C, Corey M, Dorfman R, Zielenski J, Karchin R, Penland CM, Cutting GR (2011) Evaluation of the disease liability of CFTR variants. Methods Mol Biol 742:355–372
Sosnay PR, Siklosi KR, Van Goor F, Kaniecki K, Yu H, Sharma N, Ramalho AS, Amaral MD, Dorfman R, Zielenski J, Masica DL, Karchin R, Millen L, Thomas PJ, Patrinos GP, Corey M, Lewis MH, Rommens JM, Castellani C, Penland CM, Cutting GR (2013) Defining the disease liability of variants in the cystic fibrosis transmembrane conductance regulator gene. Nat Genet 45:1160–1167
Thibodeau PH, Brautigam CA, Machius M, Thomas PJ (2005) Side chain and backbone contributions of Phe508 to CFTR folding. Nat Struct Mol Biol 12:10–16
Van Goor F, Hadida S, Grootenhuis PD, Burton B, Cao D, Neuberger T, Turnbull A, Singh A, Joubran J, Hazlewood A, Zhou J, McCartney J, Arumugam V, Decker C, Yang J, Young C, Olson ER, Wine JJ, Frizzell RA, Ashlock M, Negulescu P (2009) Rescue of CF airway epithelial cell function in vitro by a CFTR potentiator, VX-770. Proc Natl Acad Sci USA 106:18825–18830
Van Goor F, Hadida S, Grootenhuis PD, Burton B, Stack JH, Straley KS, Decker CJ, Miller M, McCartney J, Olson ER, Wine JJ, Frizzell RA, Ashlock M, Negulescu PA (2011) Correction of the F508del-CFTR protein processing defect in vitro by the investigational drug VX-809. Proc Natl Acad Sci USA 108:18843–18848
Vergani P, Lockless SW, Nairn AC, Gadsby DC (2005) CFTR channel opening by ATP-driven tight dimerization of its nucleotide-binding domains. Nature 433:876–880
Wang W, Linsdell P (2012) Alternating access to the transmembrane domain of the ATP-binding cassette protein cystic fibrosis transmembrane conductance regulator (ABCC7). J Biol Chem
Wang C, Protasevich I, Yang Z, Seehausen D, Skalak T, Zhao X, Atwell S, Spencer Emtage J, Wetmore DR, Brouillette CG, Hunt JF (2010) Integrated biophysical studies implicate partial unfolding of NBD1 of CFTR in the molecular pathogenesis of F508del cystic fibrosis. Protein Sci 19:1932–1947
Ward CL, Omura S, Kopito RR (1995) Degradation of CFTR by the ubiquitin-proteasome pathway. Cell 83:121–127
Ward A, Reyes CL, Yu J, Roth CB, Chang G (2007) Flexibility in the ABC transporter MsbA: Alternating access with a twist. Proc Natl Acad Sci USA 104:19005–19010
Zhang F, Kartner N, Lukacs GL (1998) Limited proteolysis as a probe for arrested conformational maturation of delta F508 CFTR. Nat Struct Biol 5:180–183
Zhang L, Aleksandrov LA, Zhao ZF, Birtley JR, Riordan JR, Ford RC (2009) Architecture of the cystic fibrosis transmembrane conductance regulator protein and structural changes associated with phosphorylation and nucleotide binding. J Struct Biol 167:242–251
Zhang L, Aleksandrov LA, Riordan JR, Ford RC (2010) Domain location within the cystic fibrosis transmembrane conductance regulator protein investigated by electron microscopy and gold labelling. Biochim Biophys Acta
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Ford, R.C. (2016). ABCC7/CFTR. In: George, A. (eds) ABC Transporters - 40 Years on. Springer, Cham. https://doi.org/10.1007/978-3-319-23476-2_13
Download citation
DOI: https://doi.org/10.1007/978-3-319-23476-2_13
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-23475-5
Online ISBN: 978-3-319-23476-2
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)