Abstract
In neurological intensive care units (NICUs) we are collecting an ever increasing quantity of data. These range from patient demographics and physiological monitoring to treatment strategies and outcomes. The BrainIT database is an example of this type of rich data source. It contains validated data on 264 patients who suffered traumatic brain injury (TBI) admitted to 22 NICUs in 11 European countries between March 2003 and July 2005 [1, 6].
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Introduction
In neurological intensive care units (NICUs) we are collecting an ever increasing quantity of data. These range from patient demographics and physiological monitoring to treatment strategies and outcomes. The BrainIT database is an example of this type of rich data source. It contains validated data on 264 patients who suffered traumatic brain injury (TBI) admitted to 22 NICUs in 11 European countries between March 2003 and July 2005 [1, 6].
The application of data mining techniques may allow us to identify patterns or relationships in clinical databases and lead to new insights that improve patient care [9]. Cluster analysis is a form of data mining that allows researchers to analyse a data set in its entirety and provides the opportunity to identify patterns within the data [4]. The aim of cluster analysis, unlike simply categorising data, is to identify linked groups based on multivariate factors. Reasons for the formation of these groups can then be hypothesised and tested in a clinically appropriate scientific manner.
Cluster analysis has previously been applied to high-frequency physiological data collected from adult patients following TBI or polytrauma and from paediatric patients following TBI [2, 8, 10]. It has been possible to identify complex patient states that may be used to predict patient outcome [10]. However, these previous studies have been restricted to small numbers of patients and focussed only on continuous variables. The BrainIT database contains data on a much larger number of patients and the data are continuous, ordinal and categorical. We therefore describe a pilot study using cluster analysis to identify distinct groups of patients within the BrainIT database.
Materials and Methods
Data Processing
The BrainIT database is organised into nine large data tables: demographic and one-off clinical data; daily management data; laboratory data; event data; surgical procedures; monitoring data summary; neurological event summary; targeted therapies; vital monitoring data. For the purposes of this pilot study we decided to focus on the demographic and one-off clinical data. As mentioned above, this would mean dealing with a mixture of continuous, ordinal and categorical data. Also, the quantity of data involved compared with the vital monitoring was far less computationally demanding. The table contains 109 variables including age, gender, hospital admission and transfer times, diagnosis, admission physiological variables and Glasgow Coma Scale (GCS) score, admission laboratory results and Glasgow Outcome Scale (GOS) score.
The data table was processed to optimise its clinical relevance and suitability for cluster analysis. For example, dates and times were converted to “time to” values, with time of trauma being time zero. Any negative or unrealistically large values identified were examined and manually corrected if an obvious date/time input error was present, or else made blank. Additionally, multiple units used for hydrogen ion, haemoglobin and glucose concentrations were converted to a standard. An appendix documenting all of the alterations made to the data table and their rationale will be made available with the BrainIT database.
We further processed the data table to create an increased data density for cluster analysis. To achieve this, we limited analysis to variables with at least 70 % data entered and patients with at least 65 % of these variables measured. Ultimately, we created two final tables, with one containing a combination of categorical, ordinal and continuous data and another containing continuous data only.
Cluster Analysis
Gower’s dissimilarity metric was calculated to determine the difference between patients based on each of the measured variables [3]. It is regarded as the most appropriate measure when performing analysis on mixed data types. The resulting metric was then used to perform agglomerative hierarchical cluster analysis. This is illustrated in Fig. 1.
All processing of data tables and cluster analysis was performed in r studio version 0.95.258 with r statistics version 2.13.1. The Gower dissimilarity metric was calculated using the gower.dist function and agglomerative hierarchical cluster analysis using the function hclust.
Results
Data Processing
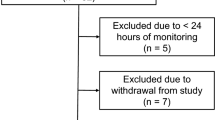
Processing of the demographic and one-off clinical data table resulted in the creation of one data table containing 42 variables for 251 patients and another containing 10 continuous variables for 203 patients (Fig. 2).
Cluster Analysis
Agglomerative hierarchical clustering of the combined data revealed five clusters selected based on visual inspection of the resulting dendrogram (Fig. 3). Cluster B contained most patients (160), clusters A, C and D were of similar size (33, 22 and 33) and cluster E was an outlier with only three patients (Table 1). The features of clusters A and D were the most clinically interesting. Patients in cluster A tended to be older and were most likely to have: consumed alcohol; fallen or been assaulted; previous dysfunction; mass lesion on CT (computed tomography). Despite having reasonable GCS motor scores on admission they had the highest mortality. Patients in cluster D tended to be younger and were most likely to have: been in a road traffic accident; associated multi-trauma; hypoxia and hypotension on admission; lowest GCS motor score; dilated and non-reactive pupils; lowest GOS.
Hierarchical cluster analysis of the continuous data table revealed three clusters based on visual inspection of the dendrogram. Cluster A contained most of the patients (175), whereas clusters B and C were of equal size (14; Table 2). Cluster C seems to have the most unique features with a tendency to lower SaO2 and PaO2 values, associated with increased transfer time from the pre-neurosurgical hospital (PNSH) to the neurosurgical hospital (NSH).
Discussion
This is a pilot project of cluster analysis of the BrainIT database and is the first analysis of its type in this rich data source. However, results produced through cluster analysis must be interpreted with caution. The clusters formed are dependent upon the dissimilarity metric and the method of cluster analysis chosen [4]. In this study, we used the Gower dissimilarity metric owing to the inclusion of categorical, ordinal and continuous data. Similar to previous studies in this domain, we elected to perform agglomerative hierarchical cluster analysis [2, 8, 10].
With the above caution in mind, we believe that our analyses have identified clusters of patients that are physiologically and clinically sensible. In analysing the table containing combined data types, we have identified two clinically relevant clusters of patients. The first is an older group, who have a higher rate of falls and are more likely to have mass lesions on CT. The second is a younger group, who have suffered multi-trauma and have markers of severe injuries on admission with more hypoxia, hypotension, pupil abnormalities and lower GCS scores. As would be expected from existing predictive models [5, 7], these patients tended to have poorer outcomes as assessed by the GOS.
In addition, analysis of the table containing only continuous data revealed a relevant cluster of patients who tended to have lower SaO2 and PaO2 values. It can be reasoned that this physiological derangement led to them also having longer transfer times from the PNSH to the NSH.
This project has successfully demonstrated the feasibility of using cluster analysis in the exploration of a large, mixed database of TBI patients. The unsupervised identification of clinically recognisable groups of patients supports the validity of the results presented. We now plan to apply similar analyses to other tables in the BrainIT database that contain high-frequency physiological data including blood pressure and intracranial pressure. The ultimate aim is to identify demographically and physiologically distinct groups of patients who will be amenable to specific treatment strategies.
References
BrainIT Research Group. Brain monitoring with information technology. http://www.brain-it.eu. Accessed 15 December 2012
Cohen MJ, Grossman AD, Morabito D, Knudson MM, Butte AJ, Manley GT (2010) Identification of complex metabolic states in critically injured patients using bioinformatic cluster analysis. Crit Care 14:R10
Gower JC (1971) A general coefficient of similarity and some of its properties. Biometrics 27:857–871
Jain AK, Murty MN, Flynn PJ (1999) Data clustering: a review. ACM Comput Surv 31:264–323
McHugh GS, Engel DC, Butcher I et al (2007) Prognostic value of secondary insults in traumatic brain injury: results from the IMPACT study. J Neurotrauma 24:287–293
Piper I, Citerio G, Chambers I et al (2003) The BrainIT group: concept and core dataset definition. Acta Neurochir (Wien) 145:615–628; discussion 628–629
Steyerberg EW, Mushkudiani N, Perel P et al (2008) Predicting outcome after traumatic brain injury: development and international validation of prognostic scores based on admission characteristics. PLoS Med 5:165
Sorani MD, Hemphill JC, Morabito D, Rosenthal G, Manley GT (2007) New approaches to physiological informatics in neurocritical care. Neurocrit Care 7(1):45–52
Sullivan R (2012) Introduction to data mining for the life sciences. Springer Science + Business Media, New York
Wainwright MS, Lewandowski R (2012) Bioinformatics analysis of mortality associated with elevated intracranial pressure in children. Acta Neurochir Suppl 114:67–73
Acknowledgements
The authors would like to acknowledge the work of the BrainIT group of investigators and participating centres in the BrainIT data set without whom this work could not have been conducted: Barcelona, Spain: Prof Sahuquillo; Cambridge, UK: Prof Pickard; Edinburgh, UK: Prof Whittle; Glasgow, UK: Mr Dunn; Gothenburg, Sweden: Dr Rydenhag; Heidelberg, Germany: Dr Kiening; Iasi, Romania: Dr Iencean; Kaunas, Lithuania: Prof Pavalkis; Leipzig, Germany: Prof Meixensberger; Leuven, Belgium: Prof Goffin; Mannheim, Germany: Prof Vajkoczy; Milan, Italy: Prof Stocchetti; Monza, Italy: Dr Citerio; Newcastle upon Tyne, UK: Dr Chambers; Novara, Italy: Prof Della Corte; Southampton, UK: Dr Hell; Uppsala, Sweden: Prof Enblad; Torino, Italy: Dr Mascia; Vilnius, Lithuania: Prof Jarzemaskas; Zurich, Switzerland: Prof Stocker.
Conflict of Interest Statement
We declare that we have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2016 Springer International Publishing Switzerland
About this chapter
Cite this chapter
McLennan, F., Hawthorne, C., Shaw, M., Piper, I. (2016). Identification of Clinically Relevant Groups of Patients Through the Application of Cluster Analysis to a Complex Traumatic Brain Injury Data Set. In: Ang, BT. (eds) Intracranial Pressure and Brain Monitoring XV. Acta Neurochirurgica Supplement, vol 122. Springer, Cham. https://doi.org/10.1007/978-3-319-22533-3_10
Download citation
DOI: https://doi.org/10.1007/978-3-319-22533-3_10
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-22532-6
Online ISBN: 978-3-319-22533-3
eBook Packages: MedicineMedicine (R0)