Abstract
Cancer cell heterogeneity is a feature of nearly all cancers and can be related to three major influences: (1) genetic and epigenetic heterogeneity due to clonal evolution/collapse; (2) microenvironmental influences; and (3) the underlying tissue hierarchy from which the tumor arises. Ongoing studies in whole genome sequencing and of the bone marrow, splenic and lymph node microenvironments demonstrate their contributions to tumor heterogeneity. In this chapter we will focus on the role of the hematopoietic hierarchy in blood cancer cellular heterogeneity; one of the most studied systems in mouse and human. Observations that myeloid and lymphoid malignancies harbor rare relatively quiescent therapy resistant cell populations date back over 30 years. Early studies in chronic myelogenous leukemia were consistent with a disease origin in the hematopoietic stem cell and subsequent studies have confirmed these findings. The publication by Bonnet et al. in 1994 offered the first prospective assessment of human cancer stem cell populations and established acute myeloid leukemia (AML) as a model system. In the intervening years, new technologies have allowed a continued reassessment of cancer stem cell populations in AML, myelodysplastic syndromes, multiple myeloma and acute lymphoblastic leukemia. While these studies have confirmed the existence of the rare cells capable of recapitulating the malignancy on transplantation, they have also identified considerable inter- and intra-patient heterogeneity with conflicting results on the ability to identify potential cancer stem cells using surface antigen profiles alone. Novel xenotransplantation models, whole genome sequencing and other technologies offer the tools to further refine this model in hematologic malignancies and develop rational therapies to target leukemia stem cells.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Hematopoietic stem cell s
- Leukemic stem cell s
- Pre-leukemic stem cells
- Chronic myelogenous leukemia
- Acute myelogenous leukemia
- Acute lymphoblastic leukemia
- Multiple myeloma
- Myelodysplastic syndromes
1 Introduction
Normal hematopoiesis relies on a highly regulated multi-tier hierarchy with a pool of rare, largely quiescent hematopoietic stem cells (HSCs) at the base (Rieger and Schroeder 2012; Morrison and Weissman 1994). Downstream are oligo-potent, lineage restricted progenitors with a capacity for robust proliferation in response to stress and terminally differentiated functional elements (Fig. 13.1). In hematopoiesis, the capacity for self-renewal is confined to mainly quiescent populations and is tightly regulated through cell autonomous and non-autonomous signals. Malignancies of the hematopoietic system are similarly heterogeneous and studies over several decades have consistently demonstrated that the capacity to repopulate the tumor on serial transplantation is confined to a rare population of cancer cells with the capacity to self-renew. Chronic phase CML is characterized by a limited mutation burden, a consistent leukemic stem cell (LSC) phenotype and an effective therapy. In contrast, acute myelogenous leukemia (AML), myelodysplastic syndromes (MDS ), B-cell acute lymphoblastic leukemia (B-ALL) and multiple myeloma (MM) result from multiple genetic and epigenetic events with increasing diversity in the cancer stem cell phenotypes. Studies of healthy aging donors as well as patients in remission have identified some of the potential early events and have identified possible limitations of targeting CSCs . Adding to the complexities, studies in human B-cell malignancies call into question the unidirectional nature of transitions between levels of the hierarchies.
2 Hematopoietic Stem Cells and Normal Hematopoiesis
2.1 Assays for the Study of Normal and Malignant Hematopoiesis
Study of normal and malignant hematopoiesis has been aided by a number of assays (Table 13.1). Commercially available standardized reagents and protocols permit comparison of results across labs. Historically suspension culture of HSC and early progenitor pools for more than a few days led to irreversible commitment and loss of stem cell function. Novel in vitro culture conditions extend the period of time prior to loss of HSC function and allow HSC expansion (Delaney et al. 2010). An adaption of the methylcellulose colony forming assays involves serial replating of methylcellulose colonies and serves as a surrogate marker for proliferative capacity with the number of replatings before CFU activity is extinguished related to self-renewal capacity. Cobblestone area forming (CAFC) and long term culture initiating cell (LTC-IC) assays allow the study of ST-HSC, MPPs and CMP/CLPs in vitro by co-culture with defined stroma. The readout for these assays is either scoring the number of CAFC beneath the stromal cell layer or transfer of hematopoietic cells into defined media methylcellulose to quantitate CFU activity. The CAFC assay has been adapted to allow large scale in vitro screening of potential LSC targeting molecules (Hartwell et al. 2013).
2.2 Identification of Normal Murine and Human Hematopoietic Stem Cells
A major strength of studying normal hematopoietic stem cell populations in mice is the availability of congenic strains that allow tracking donor and recipient contribution to the peripheral blood, spleen and marrow following transplantation. The C57bl/6 strain remains the standard for studying hematopoiesis. The cumulative results of mouse transplantation studies have resulted in a defined surface antigen phenotype that permits the transplantation of a single cell with hematopoietic reconstitution (Kiel et al. 2005) (Fig. 13.1).
Early models for human AML xenotransplatation suffered from an inability to work with small cell numbers, the need for cytokine supplementation and the impact of residual immune function. Increasingly immune deficient strains of mice have revolutionized the study of both normal and malignant hematopoiesis (Ito et al. 2002; Shultz et al. 2005). The NOG/NSG and also the NOD /SCID ß2mnull mice are among the most immune deficient strains of inbred laboratory mice described to date and demonstrate greater permissiveness in human tissue engraftment. These strains have been compared head to head with the NOD/SCID strain and permit engraftment of a greater percentage of primary AML patient samples as well as allow a higher level of engraftment. Direct transplantation of cells into the femur of the recipient further increases the efficiency of transfer (McKenzie et al. 2005). It is now possible to perform single cell xenotransplantation studies of normal human HSCs (Notta et al. 2011). Models attempting to maximize the capacity for human xenotransplantation studies in NOD, C57BL/6 and Balb/c strains are ongoing (Iwamoto et al. 2014). As xenotransplantation models improve results from prior LSC studies are being re-evaluated with regard to the impact of the assay on the observed phenotypes.
Application of the aforementioned assays has allowed careful delineation of the hematopoietic hierarchy in mouse and man. While an in depth review of normal hematopoiesis is outside the limits of this chapter several concepts are worth mentioning. First, the HSC pool size and makeup are tightly regulated through cell autonomous as well as extrinsic mechanisms many of which are still poorly understood (Fig. 13.1). There remains a mostly unexplained age dependent heterogeneity in the HSC pool with variation in the composition, kinetics, and progeny outputs when analyzed using clonal marking studies (Jordan and Lemischka 1990; Dick et al. 1985) or transplantation of highly purified populations (Uchida et al. 2003; Ema et al. 2005). Second, the assays employed to characterize stem cell and progenitor populations remain dependent upon the demonstration of a capacity for tissue regeneration following transplantation. The increasingly recognized heterogeneity of the populations in Fig. 13.1 and the instability of the available phenotypes have impaired efforts to circumvent the need for functional validation, i.e. transplantation.
3 The Role of Malignant Stem Cells in Myeloid Malignancies
3.1 Leukemic Stem Cells in CML
Chronic myeloid leukemia (CML) is a myeloproliferative disorder that can exist in three distinct phases, chronic phase, accelerated phase, and blast crisis, which is phenotypically identical to acute leukemia. The defining lesion of this disease, the Philadelphia chromosome (Ph), encodes the BCR/ABL proto-oncogene and results in a constitutively active protein tyrosine kinase product, p210BCR-ABL (Nowell and Hungerford 1960; Rowley 1973; de Klein et al. 1982). This event targets the normal HSC compartment resulting in inappropriate expansion of the granulocytic lineage. The LSC has been studied extensively in all phases of this disease as well as in the remission state i.e. minimal residual disease (MRD). Hence, CML is a near perfect paradigm for understanding the leukemia stem cell model.
In 1977, Philip Fialkow and colleagues provided some of the first data that CML is a clonal stem cell disease (Fialkow et al. 1977). They studied glucose-6-phosphate dehydrogenase (G-6-PD) isoenzymes in the granulocytes of eight woman with CML, and utilized X-linked polymorphisms to compare the enzyme types found in skin cells, normal granulocytes, and CML granulocytes. They found that the patients were heterozygous at the X-linked G-6-PD locus in the skin cells, but homozygous for G-6-PD enzyme type in the CML cells, as well as in the erythrocytes, platelets, and macrophages. This data suggested that these cells were derived from a common stem cell, i.e. the normal HSC compartment. Since this initial discovery and the works of others, the CML stem cell phenotype in chronic phase has now been defined as CD34 +CD38−CD90+Lin−Thy1+ (Ahuja et al. 1989; Jorgensen and Holyoake 2007; Eisterer et al. 2005; Holyoake et al. 1999).
Jamieson and colleagues evaluated samples from blast crisis CML to demonstrate that in CML blast crisis, secondary genetic events target the cell populations with surface antigen and gene expression profiles resembling a committed progenitor, CD34 +, CD38+ lineage−, and result in the acquisition of unlimited proliferative and self-renewal capacity, mostly through activation of the ß-catenin-signaling pathway (Jamieson et al. 2004). In a subsequent report, Abrahamsson et al. (2009) further identified glycogen synthase kinase 3beta missplicing events as contributors to expansion of self-renewal capacity to the GMP population. Follow-up studies have also identified a role for other mutations and pathways in the transition from chronic phase CML to blast crisis.
The CML blast crisis model suggests that while the “first hit” (i.e. acquisition of the oncogene BCR-ABL ), targets the normal HSC, secondary genetic events target the committed progenitor pool and result in unlimited proliferative and self-renewal potential. The secondary events that occur in blast crisis CML are the first clear evidence from human cancer that secondary events can instill the critical stem cell property of self-renewal in a population of cells that is normally non-self-renewing.
The CML stem cell has been studied in the minimal residual disease state as well. Targeted therapy with tyrosine kinase inhibitors (TKIs ), such as Imatinib , Dasatinib, Nilotinib, induces complete cytogenetic responses {i.e. no evidence of the (Ph) chromosome by FISH analysis} in more than 80 % of patients (Druker et al. 1996, 2006). The BCR-ABL transcript can still be detected by RT-PCR in most patients, however. In patients with undetectable transcript levels, approximately 60 % of patients will relapse when TKI therapy is discontinued (Rousselot et al. 2007). These clinical observations suggest that a quiescent LSC exists in CML and is resistant to TKI therapy and thus represents a reservoir for relapse of the disease. Corbin and colleagues examined how the LSC and progenitor cells survive during Imatinib therapy and asked whether the LSC is BCR-ABL dependent or independent. Utilizing a series of phosphorylation assays; the group demonstrated that the CML stem cells are not dependent on BCR-ABL activity for survival, and persist in patients with CML despite prolonged treatment with a TKI (Corbin et al. 2011). This field of research is active and focused on identifying the critical pathways of the residual CML leukemia cells.
3.2 Leukemic Stem Cells in AML
In 1994, Lapidot and colleagues published their work characterizing the leukemia initiating capacity of a sub-population of leukemic blasts (Lapidot et al. 1994). Using a SCID xenotransplantation model, they demonstrated that the ability to transplant human AML into primary and secondary recipients was confined to a population of cells defined by the expression of CD34 and the lack of expression of CD38 and markers of lineage commitment. This paper was one of many by the Dick lab examining the potential applications of xenotransplantation to the study of normal and malignant hematopoiesis. In the two decades since the report, the original phenotype has been extended as new antigens capable of enriching for LSC activities have been identified (Table 13.2). These studies have focused primarily on expanding the original CD34+CD38− phenotype.
Beginning in 2008, studies by the Bonnet lab called into question the reliance of the LSC field on the CD34 +CD38− phenotype (Taussig et al. 2008, 2010). First, Taussig et al. demonstrated that pre-treatment of unfractionated normal and AML bone marrow cells with antibodies against human CD38 impaired normal and leukemic engraftment in sub-lethally irradiated NOD -SCID recipients (Taussig et al. 2008). Pre-treatment of recipients with an antibody against the interleukin‐2 receptor β chain (CD122) partially restored human engraftment through elimination of residual NK cell activity. Transplantation of CD34+CD38− and CD34+CD38+ populations from seven AML samples into NSG or NOD/SCID/ß2mnullmice pretreated with IVIG or anti-CD122 demonstrated LSC activity in CD34 + CD38+ cells in all cases of AML demonstrating the impact of the model on the cancer stem cell phenotype.
Leukemic blasts from patients with AML associated with mutations in the NPM1 (NPM1mut) gene frequently lack surface expression of CD34 . Taussig et al. examined the capacity of CD34+CD38−, CD34+CD38+, CD34−CD38+ and the CD34−CD38− cells from patients with NPM1mut AML to engraft in their xenotransplantation assay (Taussig et al. 2010). LSC activity was demonstrated for either the CD34−CD38+, CD34−CD38− or both populations. NPMmut cases expressing CD34 demonstrated LSC activity most frequently in the CD34+CD38+ population. For most cases LSC activity was present in multiple populations demonstrating both inter-patient as well as intra-patient heterogeneity in LSC phenotypes. Using LDA they assessed LSC frequency for the different populations within a single patient. For two patients, the LIC frequency was similar among phenotypically distinct LSC populations. In a third patient, three populations demonstrated similar LSC frequencies while LSCs were less frequent in the fourth, CD34−CD38+, population.
Sarry et al. (2011) and Eppert et al. (2011) likewise identified inter- as well as intra-patient heterogeneity in LSC surface antigen phenotype, although Eppert et al. (2011) found that LSC frequency was greatest in the CD34 +CD38−population compared to the CD34+CD38+ population. In all of the above studies, LDA assays confirmed that LSCs make up a rare population of cells in the samples.
Multiple LSC populations within a patient suggest that the LSC pool evolves in response to the addition of other genetic/epigenetic events or occurs as a response to therapy. Cytogenetics, whole genome sequencing and DNA methylation studies have documented changes in the makeup of the malignancy following relapse (Welch et al. 2012; Kroeger et al. 2008). Our group carefully examined the impact of the patients’ clinical course on the LSC phenotype. Using either CD34 and CD38 or CD32 (CD34− cases) and CD38, we characterized the LSC activity in the four populations defined by these antigens prior to therapy and following relapse using an NSG xenotransplantation model. LDAs of samples obtained prior to therapy and following relapse (Ho et al. 2013) identified a 9–90-fold expansion of functional LSC activity at relapse. There was considerable inter-patient as well as intra-patient heterogeneity in LSC surface antigen phenotypes from diagnostic samples. We demonstrated expansion of LSC activity at relapse to populations lacking LSC activity in the pre-treatment sample. The molecular basis for this evolution and how it relates to the cell of origin is a work in progress.
3.3 Cell of Origin in AML
AML sequencing studies have demonstrated a median of 13 non-synonymous mutations per case (Cancer Genome Atlas Research 2013). How and where these mutations accumulate is an area of great interest. Miyamoto et al. isolated phenotypically distinct populations from patients with AML associated with t(8;21) following attainment of remission (Miyamoto et al. 2000). AML1/ETO transcripts were present in a fraction of stem cells, monocytes, and B-cells isolated from remission samples. Remission marrow cells were plated in methylcellulose and AML1/ETO transcripts were present in erythroid, granulocyte/macrophage, and megakaryocyte colonies. Thus, for AML associated with t(8;21), the cell of origin appears to be a stem cell population capable of giving rise to myeloid cells as well as B-cells. Recently, cell sorting and targeted mutation analysis were combined to investigate which stem or progenitor populations are targeted by specific mutations in AML (Shlush et al. 2014; Corces-Zimmerman et al. 2014; Jan et al. 2012). Following identification of each patient’s leukemia-specific profile, Corces-Zimmerman et al. analyzed leukemic cells, T-cells, and HSCs (CD34 +CD38−TIM3−CD99−) using targeted amplicon sequencing (Corces-Zimmerman et al. 2014). They confirmed the ability of isolated CD34+CD38−TIM3−CD99− HSCs to establish multi-lineage engraftment of NSG recipients. Mutations occurring in the HSC populations were termed “Pre-leukemic mutations” and represented a subset of the patient’s leukemia specific mutations. Leukemia -specific mutations not present in the HSC populations but present in the purified leukemic CD99+ TIM3+ cells were classified as “late events”. Combining this data with that of an earlier study (Jan et al. 2012), they identified 74 mutations in 16 patients with AML; 49 % were preleukemic and 52 % were late events. Mutations overrepresented in the preleukemic group included DNA methylation , chromatin modification, and chromatin topology (IDH2, DNMT3A, ASXL1, and IKZF1) while mutations in FLT3, NPM1 and genes involved in activated signaling were overrepresented in the late event group. CD34+ cells from remission samples identified patient specific pre-leukemic mutations in remission CD34+ progenitors and their progeny. Shlush et al. reported similar findings in a separate cohort of patients (Shlush et al. 2014).
To determine the frequency of commonly occurring mutations in non-leukemic hematopoiesis, (Xie et al. 2014) analyzed data from The Cancer Genome Atlas (TCGA) database. They selected patients with 11 different tumor types but no prior history of a hematologic malignancy or treatment. 2728 individuals were analyzed and a final list of 77 mutations in 58 cases was identified. Sixty-fiour of the events were in 19 genes including DNMT3A, TET2, JAK2, ASXL1, TP53, SF3B1, BCORL1, ASXL2 and SH2B3. Comparing this dataset with the TCGA datasets for myeloproliferative disorders, MDS , AML and chronic lymphocytic leukemia (CLL); DNMT3A, JAK2, TET2, ASXL1, TP53 and SF3B1 were consistently mutated in the TCGA blood dataset and at least two of the other datasets. The authors argued that these mutations were likely to be early events in the initiation of hematologic malignancies. Mutations of IDH1, RUNX1, NRAS, PHF6 were identified in the AML, MPN and MDS datasets but not in the TCGA blood dataset. CEBPA, WT1, PTPN11, KIT, SMC1A, and SMC3 were frequently mutated in the TCGA AML dataset but not in the TCGA blood dataset. These mutations were surmised to be later events.
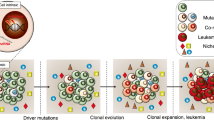
These studies support a model in which early mutations alter the “landscape” of the HSC pool (Corces-Zimmerman et al. 2014). This altered HSC pool is able to give rise to both myeloid and lymphoid progeny although skewed. Later events in the HSC or its progeny further modify the cell state and activate cell signaling resulting in the ability of cycling malignant progenitor populations to self-renew and completing leukemic transformation (Fig. 13.2). These studies rely on the deconstruction of the tumor and depend on an intact relationship between a cell’s surface antigen profile in cancer with that in normal hematopoiesis. With the advent of improved methods to target gene expression to normal cord blood stem cell populations and current xenotransplantation models the tools are in place to prospectively build human leukemias in vivo (Moriya et al. 2012; Chou et al. 2011). IPS technology provides an alternative approach to reverse engineering leukemia (Liu et al. 2014).
Model for chronic and acute myeloid leukemia. (a) Chronic myelogenous leukemia is the result of t(9;22) targeting the quiescent HSC. The LSC is a functioning HSC able to give rise to both myeloid and lymphoid progeny with expansion of the myeloid progenitor pools and a granulocytic predominance. Inhibition of the BCR-ABL fusion product restores normal hematopoiesis but does not eradicate the LSC. (b) AML. Early genetic events alter the “landscape” of the quiescent HSC pool without completing leukemic transformation. The altered HSC pool is able to give rise to both myeloid and lymphoid progeny although skewed. Later events in the progeny activate self-renewal and cell signaling resulting in the ability of cycling malignant progenitor populations to self-renew and completing the leukemic transformation. As the disease progresses, more committed progenitor populations acquire the potential for self-renewal. It has yet to be determined if these states are inter-exchangeable in human disease as demonstrated by Nolan et al. in a murine model
3.4 Lessons from Murine Models of Leukemia
Murine models of leukemia allow researchers to take full advantage of the extensive knowledge base of murine normal hematopoiesis and the power of murine genetics. As mutations in human AML are identified, they are modeled in the mouse. A complete review of murine models for leukemia is beyond the scope of this effort. We will highlight two stories where these systems were employed to address key concepts in LSC biology.
Early studies of LSCs in CML and AML suggested that the HSC pool was the primary target for early events with additional events in progenitors resulting in full transformations. Studies in mice have examined the potential of leukemia initiating events to transform HSCs as well as non-self-renewing progenitors. In 2006 Krivtsov et al. initiated acute leukemia in mice by targeting the expression of the MLL-AF9 fusion product to GMPs (Krivtsov et al. 2006). They demonstrated that the LSCs possess an immunophenotype and gene expression profile similar to that of normal GMPs. A subset of genes (363) in the MLL-AF9 LSC signature overlap with the expression profile of normal murine HSCs and the expression of profiles of human MLL associated AML. Subsequent studies have employed this model to refine the critical events in MLL mediated leukemia as well as identify novel therapeutics (Krivtsov et al. 2013; Wang et al. 2010; Hanoun et al. 2014; Miller et al. 2013). A similar capacity to activate a self-renewal program in progenitor populations has been has been demonstrated for the MLL-ENL and MOZ-TIF2 fusion products (Huntly et al. 2004; Cozzio et al. 2003). This capacity is not shared among all driver mutations BCR-ABL , Flt3 ITD mutations as well as the HOXA9-MEIS1 fusion product are not capable of fully transforming normal progenitors. Interestingly, MLL-AF9, when driven under the endogenous MLL promoter is unable to transform GMP populations.
Data from human studies supports the presence of multiple distinct LSC populations in patients with AML. Gibbs et al. (2012) applied Cytof technology to characterize LSCs populations in a murine model for AML driven by the HoxA9-Meis1 fusion. Three distinct LSC populations capable recapitulating the original immunophenotype on transplantation were identified, Lin−kit+, Gr1+kit+ and Lym+kit+. Detailed Cytof analyses of recipient marrow following transplantation of these independent LSC populations demonstrated shared signaling networks. The authors concluded that their data was not consistent with a unidirectional process of differentiation and that stemness may reflect a cellular state that exists independently of surface antigen definition.
3.5 Stem Cells in Myelodysplastic Syndromes
The myelodysplastic syndromes are a heterogeneous group of diseases characterized by bone marrow failure and a variable risk for transformation to acute leukemia. Intermediate and high risk MDS represent pre-leukemic states and are treated as such. MDS is characterized by recurrent cytogenetic and molecular abnormalities and sequencing efforts have identified overlap of the mutational spectrum in MDS with AML (Bejar et al. 2011). Recognized as a stem cell disorder early on (Raskind et al. 1984), clonal involvement of the HSC pool and its progeny is supported by studies in which HSC and early progenitors demonstrate the presence of patient specific genetic events (Nilsson et al. 2002; Tehranchi et al. 2010; Will et al. 2012). HSC and progenitor populations from primary patient samples demonstrate changes in the size of the HSC, CMP, CMP and MEP pools (Will et al. 2012). Studies evaluating the impact of therapy on the HSC pool in patients with MDS have established that despite clinical responses to lenalidomide or 5-azacytidine, including remission, the malignant HSC pool remains untargeted and serves as a reservoir for relapse (Will et al. 2012; Craddock et al. 2013; Tehranchi et al. 2010). Until recently, functional assessment of clonal HSCs and progenitors from patient samples has been hampered due to difficulties in achieving sustained engraftment of clonal hematopoiesis immune deficient mice (Nilsson et al. 2000, 2002). Two recent reports demonstrate sustained clonal engraftment of recipient mice following co-transplantation of sorted hematopoietic cells with bone marrow stromal cells via intrafemoral injection (Muguruma et al. 2011; Kerbauy et al. 2004).
With the identification of disease driving genetic events in MDS , the development of novel murine models for MDS that mirror human disease is also likely to help define the role of fully transformed stem cell populations in MDS. Early murine models for MDS were mainly limited to the less common CMML category while recent models demonstrate a phenotype more representative of the breadth of MDS (Barlow et al. 2010; Moran-Crusio et al. 2011). A recent model in which expression of the Dicer gene was targeted in bone marrow osteoblasts highlighted the role of the bone marrow microenvironment in the pathophysiology of MDS (Raaijmakers et al. 2010). These models will allow the investigation of the functional cancer stem cell populations in MDS.
3.6 Molecular Characterization of LSCs
While there have been a large number of reports applying gene expression analysis to characterize samples from large cohorts of patients with AML, only a few have studied the malignant stem cell populations (Ishikawa et al. 2007; Guzman et al. 2001b; Goardon et al. 2011; Gentles et al. 2010; Majeti et al. 2009; Eppert et al. 2011) . Guzman et al (2001b) examined the expression levels of 1400 genes related to cancer and apoptosis in leukemic CD34 +CD38− cells from primary patient samples and demonstrated aberrant expression of DAPK and IRF-1. Studies that followed applied microarray technology to compare leukemic CD34+CD38− cells to leukemic CD34+CD38+ cells (Ishikawa et al. 2007), leukemic CD123+CD34+ CD38low Lineage− cells to normal HSCs (Majeti et al. 2009; Gentles et al. 2010) and finally functionally defined LSC and normal HSC populations to leukemic populations lacking LSC activity (Eppert et al. 2011). Two of these signatures were shown to have prognostic significance (Eppert et al. 2011; Gentles et al. 2010). These differing approaches have yielded varying results with limited overlap but form a database for future queries. The LSC signature generated by Eppert et al. was enriched for an HSC signature. They also compared their LSC signature with data sets derived from embryonic stem cell as well as hematopoietic stem and progenitor populations. Their LSC signature was positively correlated with published HSC signatures and negatively correlated with more differentiated cell signatures. Majeti et al. compared normal HSCs to leukemic CD123+CD34+ CD38low Lineage− cells and identified the Adherens junction, Ribosome, Regulation of actin cytoskeleton, Tight junction and Focal adhesion pathways (KEGG) as the top five dysregulated pathways in leukemic progenitors. With the advent of new platforms to characterize miRNA levels as well as DNA methylation changes, integration of these datasets with functional data will be critical.
3.7 Therapeutic Targeting of LSCs in Myeloid Malignancies
Two decades have passed since publication of the study by Lapidot et al. (1994) that launched the stem cell model for AML. While there is agreement in the need to identify agents capable of eradicating the malignant stem cell population; increasing recognition of the heterogeneity of the LSC pool in and between patients complicates the design and implementation of LSC targeted therapies. The study of normal stem cell biology has identified a number of shared pathways that are responsible for self-renewal including sonic hedgehog, Wnt, Notch, and BMI1. These pathways are known to be necessary for malignant stem cell function in multiple tumor types. Efforts to target these pathways are in different phases of translation to the clinic.
A second approach to targeting cancer stem cells is to identify surface proteins present on cancer stem cell populations that are lacking or expressed at a lower level on normal stem cell and progenitor populations (Table 13.2). Anti-CD33 therapy has been in the clinical arena for over a decade; while there is a benefit to a small subset of patients; this approach overall has been disappointing (Burnett et al. 2011). Similar early phase clinical efforts for CD123, CLL1 are ongoing as are pre-clinical studies targeting CD44 , CD47, TIM-3 and IL1RAP (Majeti 2011; Askmyr et al. 2013; Kikushige and Miyamoto 2013). In addition to standard monoclonal antibody therapy, Bi-specific T-cell engagers (BiTEs) and CAR-T efforts using C33, CD123 and LeY antigens are in the pre-clinical and early clinical stages (Aigner et al. 2013; Ritchie et al. 2013; Gill et al. 2014; Dutour et al. 2012). A major limitation of the LSC antigen targeting approach is that none of the currently published LSC antigens appear to be both present on all of the LSC populations in a patient while lacking on normal HSCs and/or progenitors.
A third approach is to target unique cancer stem cell dependencies. The development of mutation specific therapies is one such approach although most mutations are limited to a small fraction of patients except for NPM1 and FLT3 mutations. An early study demonstrated constitutive NF Kappaβ activity in LSCs as compared to normal HSCs (Guzman et al. 2001a). This has been confirmed and a recent study demonstrated that NF Kappaβ activity was common to murine and human LSC populations and associated with expansion of the LSC pool (Kagoya et al. 2014). An ongoing Children’s Oncology Group trial is investigating the benefits of adding bortezomib to standard induction therapy. In 2005 Guzman et al. published a follow-up study in which they characterized the ability of a natural product, parthenolide, to selectively eradicate LSC populations (Guzman et al. 2005). Initially selected for its ability to inhibit NF Kappaβ signaling (Hehner et al. 1999), subsequent studies have identified alternative activities of this class of compounds including redox balance, heat shock protein response, proteasome signaling and glycolysis (Pei and Jordan 2012; Pei et al. 2013). Subsequent studies have confirmed the utility of this class of compounds in targeting the above pathways and identified other molecules with similar effects or that synergize with parthenolide (Lagadinou et al. 2013; Dai et al. 2010; Hassane et al. 2008, 2010).
4 The Role of Malignant Stem Cells in B-cell Malignancies
4.1 Normal B-cell Development
In the current model for B-cell development, multi-potent progenitors undergo a cell fate decision giving rise to a multipotent lymphoid progenitor that then give rise to either B-cells or T-cells with the early pro-B-cell the first stage of B-cell development (Fig. 13.1). Early B-cell development occurs in the bone marrow followed by exodus to the lymph organs. Following selection and maturation in the germinal centers of primary and secondary lymph organs, B-cells return to the bone marrow. Developmental stage can be identified using the expression of specific surface antigens and the rearrangement status of Ig H and L chains. CD34 +CD10+CD19− define the common lymphoid progenitors while pro-B-cells are defined as CD34+CD10+CD19+ and pre-B-cells are CD34−CD10+CD19+. Rearrangement of the VDJ H chain locus is characteristic of pro-B-cells while expression of a pre-BCR, composed of IgH chains and surrogate L chains marks the pre-B-cell population (Hystad et al. 2007). Memory B-cells, plasmablasts and mature plasma cells are similarly defined by their surface antigen expression profile (Memory B-cell: CD27+CD20+CD45++IL6R−CD138−CD38dim; Plasmablast: CD20−CD38++CD45++IL6R++CD138− and Mature plasma cells: CD20−CD38++CD45dimIL6R+CD138++).
Cells of the myeloid lineage including erythrocytes, megakaryocytes, and granulocytes have well defined and frequently quite short half-lives while a subset of mature B-cell populations are long lived. This is the basis for immunologic memory allowing for a more rapid antigen specific response following re-exposure by rapid expansion of memory B-cells. In 2006, Luckey et al. (2006) analyzed the expression profiles of naïve, effector and memory T-cells as well as naïve, germinal center, memory B-cells and plasma cells. Transcripts augmented in memory cell populations compared to naïve and effector cell populations were enriched in HSCs and lost following commitment. Likewise, transcripts down regulated in memory cell populations were down regulated in HSCs and increased with differentiation.
4.2 The Role of Leukemic Stem Cells in B-cell ALL
B-cell Acute Lymphoblastic leukemia (B-ALL) arises in an early B-cell progenitor (Teitell and Pandolfi 2009) from the accumulation of genetic events by hematopoietic stem cells and/or progenitor cells. Early immunoglobulin rearrangement studies demonstrated that a quarter of early B-ALL cases contain more than two IgH rearrangements. Sequential analyses of a few patients found that the patient specific pattern of IgH gene rearrangement may change over the course of the disease (Wright et al. 1987). TEL-AML1 B-ALL is a rare disease and cases of in-utero transfer of a pre-leukemic clone between twins have been reported. When one twin presents with B-ALL, the other twin may harbor the preleukemic clone. Hong et al. demonstrated that CD34 +CD38−CD19+ leukemic cells from four patients with TEL-AML1 B-ALL were capable of transplanting the leukemia in mice (Ma et al. 1998). The peripheral blood of the healthy twin of one patient contained found very rare circulating TEL-AML1 positive CD34+CD38−CD19+ pre-leukemic cells. Castor et al. (2005) examined the involvement of CD34+CD38−CD19− and CD34+CD38−CD19+ cells from children and adults with B-ALL for presence of the TEL-AML1, P190 BCR-ABL and P210 BCR-ABL translocations. TEL-AML1 and p190BCR-ABL involved the B-cell progenitor population but not the CD34+CD38−CD19− HSC pool while p210BCR-ABL cases involved both populations. Xenotransplantation studies revealed that only the CD34+CD38−CD19+ cells from TEL-AML1, P190 BCR-ABL and p210BCR-ABL cases gave rise to B-ALL upon transplantation. CD34+CD38−CD19− cells gave rise to normal reconstitution regardless of the type of B-ALL. This suggests that for these pediatric (TEL-AML1 and p190BCR-ABL) and adult (p210BCR-ABL) B-ALL, the cell of origin may differ but the functional LSC has a conserved phenotype, that of a committed B-cell progenitor. Cobaleda et al. (2000) isolated leukemic cells from B-ALL samples associated with t(9;22). In contrast to the study by Castor et al. (2005), only CD34 + CD38- cells were capable of engrafting NOD /SCID mice . Cox et al. (2004) employed a series of in vitro assays as well as xenotransplantation to characterize the surface antigen profile of B-ALL leukemia initiating cells. LSC activity was restricted to the CD34+CD10− and CD34−CD19− populations. This study did not include B-ALL associated with a t(9;22) or t(4;11).
The above studies relied on NOD /SCID assay and studies in AML have shown the potential impact of this assay on the phenotypes of LSC population(s) identified. Rau et al. (2014) undertook an extensive study of primary ALL samples using an NSG model with intrafemoral transplantation. The patients represented three distinct ALL risk groups. Despite using a number of surface antigens, they were unable to enrich for LSC activity. All populations demonstrated similar engraftment frequencies as well as kinetics of engraftment. LDA of the sorted populations demonstrated similar frequencies for LSC activity regardless of surface antigen phenotype. Their data was consistent with the lack of a hierarchy in acute B lymphoblastic leukemia and suggest a non-hierarchical model to account for tumor heterogeneity.
Efforts are underway to improve human B-ALL LSC models using cord blood stem cell targeting and xenotransplantation. Barabe et al. (2007) targeted the expression of the MLL-ENL and MLL AF9 fusion transcripts in primary cord blood progenitor cells. Recipient mice developed both B-ALL and AML that was transferrable into secondary recipients. Retroviral insertion analyses and IgH locus analyses demonstrated that the B-ALL recipients contained differing rearrangements of the IgH locus. 40 % of the cases were consistent with transformation of an early hematopoietic progenitor. Using serial transplantation to model disease progression, they showed that the contribution of clones with unarranged IgH loci diminished with passage while alternative LSC populations arising from more mature B-cell progenitors maintained the disease.
4.3 Myeloma Stem Cells
Multiple myeloma (MM) belongs to a spectrum of diseases that includes monoclonal gammopathy of undetermined significance (MGUS), smoldering myeloma and symptomatic myeloma. MGUS is present in approximately 3 % of the general population 50 years of age and older with a risk of transformation to multiple myeloma of 1 % per year. In 2009, it was demonstrated that in up to 75 % of myeloma patients a detectable M-protein was identifiable 8 or more years prior to diagnosis (Weiss et al. 2009). This is consistent with a clonal process maintained by a population(s) of cells that are long lived. Immunoglobulin sequencing studies in patients with MM demonstrates the presence of extensive hypermutation without intraclonal variation consistent with the development of the malignancy at the post-germinal center B-cell.
Myeloma was one of the cancer types in which tumor heterogeneity was initially assessed. In 1968, Bergsagel and Valeriote characterized the capacity of a murine plasma cell line to initiate a malignancy in recipient syngeneic mice (Bergsagel and Valeriote 1968). 3 × 104 malignant plasma cells were required to form at least one colony in the spleen of recipient mice. They used this model to show that most of the plasma cells were cycling and sensitive to vinblastine, a cell cycle dependent chemotherapeutic agent. In 1977, these findings were reproduced using human myeloma cells. More recent efforts have employed two different approaches to address heterogeneity in plasma cell tumors; NOD /SCID and NSG xenotransplantation models such as those noted above or models in which human (SCID-hu) or rabbit (SCID-rab) bone fragments are implanted directly into immunocompromised mice followed by injection of the myeloma cells into the implant.
In 1998, (Yaccoby et al. 1998) employed a SCID-Hu xenotransplantation model for studying primary MM samples. CD38++CD45−plasma cells engrafted the implanted human bone whereas the plasma cell depleted cells did not. In normal B-cell development, terminal plasma cell differentiation is characterized by loss of CD45 and increasing CD38 expression. Hosen et al. reported CFU activity as well sustained engraftment in the SCID-rab model for CD138−CD19−CD38++ cells (Hosen et al. 2012). CD19+ B-cells lacked CFU activity and engraftment potenital. CD138+ cells engrafted in a subset of the cases.
Matsui and colleagues used magnetic bead enrichment/depletion approaches to obtain CD138+ CD34 − and CD138−CD34− bone marrow cells from 24 MM patients. CD138+CD34− cells were unable to form colonies in vitro while the CD138−CD34− cells generated colonies of morphologically mature plasma cells expressing CD138. Engraftment of NOD /SCID mice by myeloma was observed only with CD138−CD34− cells which gave rise to CD138+ cells in the recipient mice. MM samples depleted of CD19, CD45 and CD20 cells lacked colony forming potential consistent with a memory B-cell-like progenitor. They refined the surface antigen profile of myeloma propagating populations in a follow-up report, CD138−CD27+CD19+, and demonstrated that this population was resistant to lenalidimide, bortezomib and cyclophosphamide; agents commonly used to treat patients with MM (Matsui et al. 2004; Matsui et al. 2008).
In 2013, Chaidos and colleagues addressed the disagreement as to the surface antigen phenotype of MM cells responsible for tumor maintenance (Chaidos et al. 2013). They extensively characterized the surface antigen profiles of MM blood and marrow samples coupled with IgH CDR3 characterization. They found that the MM clone did not include pre-germinal center B-cells but was comprised of a mixture of mature CD19+ B-cells (resting memory B-cells), plasmablasts, and CD138 low and CD138+ plasma cells. They also identified a CD138− population they termed the pre-plasma cell (pre-PC). They sorted and transplanted CD19+ B-cells, Pre-PCs and PCs from MM patients into sub lethally irradiated NSG mice. Recipients of CD138+ PCs displayed BM engraftment in 9 of 12 cases. Negatively enriched Pre-PCs also demonstrated engraftment while none of the mice receiving CD19+ cells showed evidence of engraftment. They proposed a model in which myeloma-stem cell activity was confined to two interconvertible populations of MM cells distinguishable only by the level of expression of CD138. It is unclear to what degree the heterogeneous results reported relate to clinical features of the samples studied, the approach to isolating populations (negative selection vs positive selection) or the different models employed for xenotransplantation assays. Alternatives to a reliance on surface antigen profiling have employed sorting based on ALDH expression or Hoechst 33,342 staining (Matsui et al. 2004). Additional studies will be necessary to further clarify the exact nature of the myeloma initiating cell using a uniform and standardized set of isolation approaches and xenotransplantation model.
4.4 Targeting Malignant Stem Cells in B-cell ALL and Myeloma
Similar to other cancers, pathways regulating normal stem cell self-renewal have been examined in B-ALL and MM. The sHH pathway is active in MM stem cells and early pre-clinical efforts have demonstrated some efficacy (Agarwal et al. 2014; Peacock et al. 2007).
CD19 is expressed on the cell of origin and the LSC for many cases of B-cell ALL and its expression is carried through late into B-cell development (see above). There is general excitement about several novel approaches to targeting CD19 in B-ALL and other B-cell malignancies. Monoclonal antibodies against CD19 have been conjugated to antineoplastic agents as well as to a single-chain variable region capable of binding to the CD3 T-cell receptor. These agents have shown activity in early phase trials and are now being tested in the upfront setting. Likewise, CAR-T-cells targeting CD19 have had success in relapsed B-ALL (Maude et al. 2014; Davila et al. 2014). These efforts have been expanded to most B-cell malignancies. The observation that MM propagating cells express CD20 led to a small clinical trial employing Rituximab, a monoclonal antibody against CD20 (Moreau et al. 2007). This trial demonstrated little efficacy for this approach in patients and this approach has not moved forward. Antibodies targeting CD138, present on some MM stem cell phenotypes but not others, has been studied and is currently undergoing phase 3 testing.
5 Conclusion and Future Directions
In 2011, Hanahan and Weinberg updated their treatise on the hallmarks of cancer which include sustaining proliferative signaling, evading growth suppressors, resisting cell death and enabling replicative immortality (self-renewal) (Hanahan and Weinberg 2011). These properties are shared by normal cells during normal development and homeostasis where they are compartmentalized and tightly regulated. Although initial studies in AML and CML demonstrated an overlap of the cancer stem cell phenotype with that of the normal HSC; subsequent studies have shown that in cancer the capacity for self-renewal can expand to less quiescent progenitor populations. As the disease progresses, the cancer stem cell phenotypes in AML, MM and ALL resemble late progenitors as “stemness” moves further out into the hierarchy. It will be critical to define the cancer specific mechanisms driving this expansion as targeting these pathways may restore control of self-renewal without restricting self-renewal in normal populations. Recently, constitutive activation of the nuclear factor-kappa B (NF-κB ) pathway was shown to expand functional LSC activity as assessed by LDA (Kagoya et al. 2014). As additional agents capable of targeting key pathways involved in self-renewal become available, studying their impact on all compartments in cancer and normal tissue will be critical.
Whole genome sequencing studies have outlined the genetic space for most hematologic malignancies. Interestingly, studies of normal appearing hematopoietic stem cells in the diagnosis and remission samples from patients with AML have identified cells retaining a capacity for multi-lineage differentiation with disease specific mutations. These may represent residual pre-leukemic stem cells with overlap of these mutations with those identified in peripheral blood and marrow samples from individuals without a hematologic malignancy. The frequency and nature of mutations in individuals without a hematologic malignancy is tightly associated with aging as are the diagnoses of AML, MDS and Myeloma. These findings will need to be validated in aged healthy individuals with follow-up analyses to ensure they affect a long-lived population. We will require a greater understanding of how these mutations alter the landscape of the stem cell pool prior to accumulating additional genetic events. A frequent statement in reviews and articles on cancer stem cells is that CSCs represent a reservoir for relapse hence the interest in identifying and phenotyping these populations. How to effectively target a pre-leukemic stem cell pool that may be indistinguishable from normal HSCs will likely serve as a task for the next decade. As demonstrated so well by chronic phase CML, it may not be necessary to eradicate pre-leukemic HSCs as long as there are effective therapies for their progeny.
References
Abrahamsson AE, Geron I, Gotlib J, Dao KH, Barroga CF, Newton IG, Giles FJ, Durocher J, Creusot RS, Karimi M, Jones C, Zehnder JL, Keating A, Negrin RS, Weissman IL, Jamieson CH (2009) Glycogen synthase kinase 3beta missplicing contributes to leukemia stem cell generation. Proc Natl Acad Sci USA 106(10):3925–3929
Agarwal JR, Wang Q, Tanno T, Rasheed Z, Merchant A, Ghosh N, Borrello I, Huff CA, Parhami F, Matsui W (2014) Activation of liver X receptors inhibits hedgehog signaling, clonogenic growth, and self-renewal in multiple myeloma. Mol Cancer Ther 13(7):1873–1881
Ahuja H, Bar-Eli M, Advani SH, Benchimol S, Cline MJ (1989) Alterations in the p53 gene and the clonal evolution of the blast crisis of chronic myelocytic leukemia. Proc Natl Acad Sci USA 86(17):6783–6787
Aigner M, Feulner J, Schaffer S, Kischel R, Kufer P, Schneider K, Henn A, Rattel B, Friedrich M, Baeuerle PA, Mackensen A, Krause SW (2013) T lymphocytes can be effectively recruited for ex vivo and in vivo lysis of AML blasts by a novel CD33/CD3-bispecific BiTE antibody construct. Leukemia 27(5):1107–1115
Askmyr M, Agerstam H, Hansen N, Gordon S, Arvanitakis A, Rissler M, Juliusson G, Richter J, Jaras M, Fioretos T (2013) Selective killing of candidate AML stem cells by antibody targeting of IL1RAP. Blood 121(18):3709–3713
Bakker AB, van den Oudenrijn S, Bakker AQ, Feller N, van Meijer M, Bia JA, Jongeneelen MA, Visser TJ, Bijl N, Geuijen CA, Marissen WE, Radosevic K, Throsby M, Schuurhuis GJ, Ossenkoppele GJ, de Kruif J, Goudsmit J, Kruisbeek AM (2004) C-type lectin-like molecule-1: a novel myeloid cell surface marker associated with acute myeloid leukemia. Cancer Res 64(22):8443–8450
Barabe F, Kennedy JA, Hope KJ, Dick JE (2007) Modeling the initiation and progression of human acute leukemia in mice. Science 316(5824):600–604
Barlow JL, Drynan LF, Hewett DR, Holmes LR, Lorenzo-Abalde S, Lane AL, Jolin HE, Pannell R, Middleton AJ, Wong SH, Warren AJ, Wainscoat JS, Boultwood J, McKenzie AN (2010) A p53-dependent mechanism underlies macrocytic anemia in a mouse model of human 5q- syndrome. Nat Med 16(1):59–66
Bejar R, Stevenson K, Abdel-Wahab O, Galili N, Nilsson B, Garcia-Manero G, Kantarjian H, Raza A, Levine RL, Neuberg D, Ebert BL (2011) Clinical effect of point mutations in myelodysplastic syndromes. N Engl J Med 364(26):2496–2506
Bergsagel DE, Valeriote FA (1968) Growth characteristics of a mouse plasma cell tumor. Cancer Res 28(11):2187–2196
Burnett AK, Hills RK, Milligan D, Kjeldsen L, Kell J, Russell NH, Yin JA, Hunter A, Goldstone AH, Wheatley K (2011) Identification of patients with acute myeloblastic leukemia who benefit from the addition of gemtuzumab ozogamicin: results of the MRC AML15 trial. J Clin Oncol 29(4):369–377
Cancer Genome Atlas Research N (2013) Genomic and epigenomic landscapes of adult de novo acute myeloid leukemia. N Engl J Med 368(22):2059–2074
Castor A, Nilsson L, Astrand-Grundstrom I, Buitenhuis M, Ramirez C, Anderson K, Strombeck B, Garwicz S, Bekassy AN, Schmiegelow K, Lausen B, Hokland P, Lehmann S, Juliusson G, Johansson B, Jacobsen SE (2005) Distinct patterns of hematopoietic stem cell involvement in acute lymphoblastic leukemia. Nat Med 11(6):630–637
Chaidos A, Barnes CP, Cowan G, May PC, Melo V, Hatjiharissi E, Papaioannou M, Harrington H, Doolittle H, Terpos E, Dimopoulos M, Abdalla S, Yarranton H, Naresh K, Foroni L, Reid A, Rahemtulla A, Stumpf M, Roberts I, Karadimitris A (2013) Clinical drug resistance linked to interconvertible phenotypic and functional states of tumor-propagating cells in multiple myeloma. Blood 121(2):318–328
Chou FS, Wunderlich M, Griesinger A, Mulloy JC (2011) N-Ras(G12D) induces features of stepwise transformation in preleukemic human umbilical cord blood cultures expressing the AML1-ETO fusion gene. Blood 117(7):2237–2240
Cobaleda C, Gutierrez-Cianca N, Perez-Losada J, Flores T, Garcia-Sanz R, Gonzalez M, Sanchez-Garcia I (2000) A primitive hematopoietic cell is the target for the leukemic transformation in human philadelphia-positive acute lymphoblastic leukemia. Blood 95(3):1007–1013
Corbin AS, Agarwal A, Loriaux M, Cortes J, Deininger MW, Druker BJ (2011) Human chronic myeloid leukemia stem cells are insensitive to imatinib despite inhibition of BCR-ABL activity. J Clin Invest 121(1):396–409
Corces-Zimmerman MR, Hong WJ, Weissman IL, Medeiros BC, Majeti R (2014) Preleukemic mutations in human acute myeloid leukemia affect epigenetic regulators and persist in remission. Proc Natl Acad Sci USA 111(7):2548–2553
Cox CV, Evely RS, Oakhill A, Pamphilon DH, Goulden NJ, Blair A (2004) Characterization of acute lymphoblastic leukemia progenitor cells. Blood 104(9):2919–2925
Cozzio A, Passegue E, Ayton PM, Karsunky H, Cleary ML, Weissman IL (2003) Similar MLL-associated leukemias arising from self-renewing stem cells and short-lived myeloid progenitors. Genes Dev 17(24):3029–3035
Craddock C, Quek L, Goardon N, Freeman S, Siddique S, Raghavan M, Aztberger A, Schuh A, Grimwade D, Ivey A, Virgo P, Hills R, McSkeane T, Arrazi J, Knapper S, Brookes C, Davies B, Price A, Wall K, Griffiths M, Cavenagh J, Majeti R, Weissman I, Burnett A, Vyas P (2013) Azacitidine fails to eradicate leukemic stem/progenitor cell populations in patients with acute myeloid leukemia and myelodysplasia. Leukemia 27(5):1028–1036
Dai Y, Guzman ML, Chen S, Wang L, Yeung SK, Pei XY, Dent P, Jordan CT, Grant S (2010) The NF (Nuclear factor)-kappaB inhibitor parthenolide interacts with histone deacetylase inhibitors to induce MKK7/JNK1-dependent apoptosis in human acute myeloid leukaemia cells. Br J Haematol 151(1):70–83
Davila ML, Riviere I, Wang X, Bartido S, Park J, Curran K, Chung SS, Stefanski J, Borquez-Ojeda O, Olszewska M, Qu J, Wasielewska T, He Q, Fink M, Shinglot H, Youssif M, Satter M, Wang Y, Hosey J, Quintanilla H, Halton E, Bernal Y, Bouhassira DC, Arcila ME, Gonen M, Roboz GJ, Maslak P, Douer D, Frattini MG, Giralt S, Sadelain M, Brentjens R (2014) Efficacy and toxicity management of 19-28z CAR T cell therapy in B cell acute lymphoblastic leukemia. Sci Transl Med 6(224):224ra225
de Klein A, van Kessel AG, Grosveld G, Bartram CR, Hagemeijer A, Bootsma D, Spurr NK, Heisterkamp N, Groffen J, Stephenson JR (1982) A cellular oncogene is translocated to the Philadelphia chromosome in chronic myelocytic leukaemia. Nature 300(5894):765–767
Delaney C, Heimfeld S, Brashem-Stein C, Voorhies H, Manger RL, Bernstein ID (2010) Notch-mediated expansion of human cord blood progenitor cells capable of rapid myeloid reconstitution. Nat Med 16(2):232–236
Dick JE, Magli MC, Huszar D, Phillips RA, Bernstein A (1985) Introduction of a selectable gene into primitive stem cells capable of long-term reconstitution of the hemopoietic system of W/Wv mice. Cell 42(1):71–79
Druker BJ, Tamura S, Buchdunger E, Ohno S, Segal GM, Fanning S, Zimmermann J, Lydon NB (1996) Effects of a selective inhibitor of the Abl tyrosine kinase on the growth of Bcr-Abl positive cells. Nat Med 2(5):561–566
Druker BJ, Guilhot F, O’Brien SG, Gathmann I, Kantarjian H, Gattermann N, Deininger MW, Silver RT, Goldman JM, Stone RM, Cervantes F, Hochhaus A, Powell BL, Gabrilove JL, Rousselot P, Reiffers J, Cornelissen JJ, Hughes T, Agis H, Fischer T, Verhoef G, Shepherd J, Saglio G, Gratwohl A, Nielsen JL, Radich JP, Simonsson B, Taylor K, Baccarani M, So C, Letvak L, Larson RA, Investigators I (2006) Five-year follow-up of patients receiving imatinib for chronic myeloid leukemia. N Engl J Med 355(23):2408–2417
Dutour A, Marin V, Pizzitola I, Valsesia-Wittmann S, Lee D, Yvon E, Finney H, Lawson A, Brenner M, Biondi A, Biagi E, Rousseau R (2012) In vitro and in vivo antitumor effect of anti-CD33 chimeric receptor-expressing EBV-CTL against CD33 acute myeloid leukemia. Adv Hematol 2012:683065
Eisterer W, Jiang X, Christ O, Glimm H, Lee KH, Pang E, Lambie K, Shaw G, Holyoake TL, Petzer AL, Auewarakul C, Barnett MJ, Eaves CJ, Eaves AC (2005) Different subsets of primary chronic myeloid leukemia stem cells engraft immunodeficient mice and produce a model of the human disease. Leukemia 19(3):435–441
Ema H, Sudo K, Seita J, Matsubara A, Morita Y, Osawa M, Takatsu K, Takaki S, Nakauchi H (2005) Quantification of self-renewal capacity in single hematopoietic stem cells from normal and Lnk-deficient mice. Dev Cell 8(6):907–914
Eppert K, Takenaka K, Lechman ER, Waldron L, Nilsson B, van Galen P, Metzeler KH, Poeppl A, Ling V, Beyene J, Canty AJ, Danska JS, Bohlander SK, Buske C, Minden MD, Golub TR, Jurisica I, Ebert BL, Dick JE (2011) Stem cell gene expression programs influence clinical outcome in human leukemia. Nat Med 17(9):1086–1093
Fialkow PJ, Jacobson RJ, Papayannopoulou T (1977) Chronic myelocytic leukemia: clonal origin in a stem cell common to the granulocyte, erythrocyte, platelet and monocyte/macrophage. Am J Med 63(1):125–130
Gentles AJ, Plevritis SK, Majeti R, Alizadeh AA (2010) Association of a leukemic stem cell gene expression signature with clinical outcomes in acute myeloid leukemia. Jama 304(24):2706–2715
Gibbs KD Jr, Jager A, Crespo O, Goltsev Y, Trejo A, Richard CE, Nolan GP (2012) Decoupling of tumor-initiating activity from stable immunophenotype in HoxA9-Meis1-driven AML. Cell Stem Cell 10(2):210–217
Gill S, Tasian SK, Ruella M, Shestova O, Li Y, Porter DL, Carroll M, Danet-Desnoyers G, Scholler J, Grupp SA, June CH, Kalos M (2014) Preclinical targeting of human acute myeloid leukemia and myeloablation using chimeric antigen receptor-modified T cells. Blood 123(15):2343–2354
Goardon N, Marchi E, Atzberger A, Quek L, Schuh A, Soneji S, Woll P, Mead A, Alford KA, Rout R, Chaudhury S, Gilkes A, Knapper S, Beldjord K, Begum S, Rose S, Geddes N, Griffiths M, Standen G, Sternberg A, Cavenagh J, Hunter H, Bowen D, Killick S, Robinson L, Price A, Macintyre E, Virgo P, Burnett A, Craddock C, Enver T, Jacobsen SE, Porcher C, Vyas P (2011) Coexistence of LMPP-like and GMP-like leukemia stem cells in acute myeloid leukemia. Cancer Cell 19(1):138–152
Guzman ML, Neering SJ, Upchurch D, Grimes B, Howard DS, Rizzieri DA, Luger SM, Jordan CT (2001a) Nuclear factor-kappaB is constitutively activated in primitive human acute myelogenous leukemia cells. Blood 98(8):2301–2307
Guzman ML, Upchurch D, Grimes B, Howard DS, Rizzieri DA, Luger SM, Phillips GL, Jordan CT (2001b) Expression of tumor-suppressor genes interferon regulatory factor 1 and death-associated protein kinase in primitive acute myelogenous leukemia cells. Blood 97(7):2177–2179
Guzman ML, Rossi RM, Karnischky L, Li X, Peterson DR, Howard DS, Jordan CT (2005) The sesquiterpene lactone parthenolide induces apoptosis of human acute myelogenous leukemia stem and progenitor cells. Blood 105(11):4163–4169
Hanahan D, Weinberg RA (2011) Hallmarks of cancer: the next generation. Cell 144(5):646–674
Hanoun M, Zhang D, Mizoguchi T, Pinho S, Pierce H, Kunisaki Y, Lacombe J, Armstrong SA, Duhrsen U, Frenette PS (2014) Acute myelogenous leukemia-induced sympathetic neuropathy promotes malignancy in an altered hematopoietic stem cell niche. Cell Stem Cell 15(3):365–375
Hartwell KA, Miller PG, Mukherjee S, Kahn AR, Stewart AL, Logan DJ, Negri JM, Duvet M, Jaras M, Puram R, Dancik V, Al-Shahrour F, Kindler T, Tothova Z, Chattopadhyay S, Hasaka T, Narayan R, Dai M, Huang C, Shterental S, Chu LP, Haydu JE, Shieh JH, Steensma DP, Munoz B, Bittker JA, Shamji AF, Clemons PA, Tolliday NJ, Carpenter AE, Gilliland DG, Stern AM, Moore MA, Scadden DT, Schreiber SL, Ebert BL, Golub TR (2013) Niche-based screening identifies small-molecule inhibitors of leukemia stem cells. Nat Chem Biol 9(12):840–848
Hassane DC, Guzman ML, Corbett C, Li X, Abboud R, Young F, Liesveld JL, Carroll M, Jordan CT (2008) Discovery of agents that eradicate leukemia stem cells using an in silico screen of public gene expression data. Blood 111(12):5654–5662
Hassane DC, Sen S, Minhajuddin M, Rossi RM, Corbett CA, Balys M, Wei L, Crooks PA, Guzman ML, Jordan CT (2010) Chemical genomic screening reveals synergism between parthenolide and inhibitors of the PI-3 kinase and mTOR pathways. Blood 116(26):5983–5990
Hehner SP, Hofmann TG, Droge W, Schmitz ML (1999) The antiinflammatory sesquiterpene lactone parthenolide inhibits NF-kappa B by targeting the I kappa B kinase complex. J Immunol 163(10):5617–5623
Ho TCL, O’Dwyer K, Mendler J, Liesveld J, Wetzler M, Wang ES, Guzman M, Jordan C, Becker MW (2013) Evolution of acute myelogenous leukemia stem cell properties following treatment and progression. Blood 122(21)
Holyoake T, Jiang X, Eaves C, Eaves A (1999) Isolation of a highly quiescent subpopulation of primitive leukemic cells in chronic myeloid leukemia. Blood 94(6):2056–2064
Hosen N, Park CY, Tatsumi N, Oji Y, Sugiyama H, Gramatzki M, Krensky AM, Weissman IL (2007) CD96 is a leukemic stem cell-specific marker in human acute myeloid leukemia. Proc Natl Acad Sci USA 104(26):11008–11013
Hosen N, Matsuoka Y, Kishida S, Nakata J, Mizutani Y, Hasegawa K, Mugitani A, Ichihara H, Aoyama Y, Nishida S, Tsuboi A, Fujiki F, Tatsumi N, Nakajima H, Hino M, Kimura T, Yata K, Abe M, Oka Y, Oji Y, Kumanogoh A, Sugiyama H (2012) CD138-negative clonogenic cells are plasma cells but not B cells in some multiple myeloma patients. Leukemia 26(9):2135–2141
Huntly BJ, Shigematsu H, Deguchi K, Lee BH, Mizuno S, Duclos N, Rowan R, Amaral S, Curley D, Williams IR, Akashi K, Gilliland DG (2004) MOZ-TIF2, but not BCR-ABL, confers properties of leukemic stem cells to committed murine hematopoietic progenitors. Cancer Cell 6(6):587–596
Hystad ME, Myklebust JH, Bo TH, Sivertsen EA, Rian E, Forfang L, Munthe E, Rosenwald A, Chiorazzi M, Jonassen I, Staudt LM, Smeland EB (2007) Characterization of early stages of human B cell development by gene expression profiling. J Immunol 179(6):3662–3671
Ishikawa F, Yoshida S, Saito Y, Hijikata A, Kitamura H, Tanaka S, Nakamura R, Tanaka T, Tomiyama H, Saito N, Fukata M, Miyamoto T, Lyons B, Ohshima K, Uchida N, Taniguchi S, Ohara O, Akashi K, Harada M, Shultz LD (2007) Chemotherapy-resistant human AML stem cells home to and engraft within the bone-marrow endosteal region. Nat Biotechnol 25(11):1315–1321
Ito M, Hiramatsu H, Kobayashi K, Suzue K, Kawahata M, Hioki K, Ueyama Y, Koyanagi Y, Sugamura K, Tsuji K, Heike T, Nakahata T (2002) NOD/SCID/gamma(c)(null) mouse: an excellent recipient mouse model for engraftment of human cells. Blood 100(9):3175–3182
Iwamoto C, Takenaka K, Urata S, Yamauchi T, Shima T, Kuriyama T, Daitoku S, Saito Y, Miyamoto T, Iwasaki H, Kitabayashi I, Itoh K, Kishimoto J, Kohda D, Matozaki T, Akashi K (2014) The BALB/c-specific polymorphic SIRPA enhances its affinity for human CD47, inhibiting phagocytosis against human cells to promote xenogeneic engraftment. Exp Hematol 42(3):163–171, e161
Jamieson CH, Ailles LE, Dylla SJ, Muijtjens M, Jones C, Zehnder JL, Gotlib J, Li K, Manz MG, Keating A, Sawyers CL, Weissman IL (2004) Granulocyte-macrophage progenitors as candidate leukemic stem cells in blast-crisis CML. N Engl J Med 351(7):657–667
Jan M, Chao MP, Cha AC, Alizadeh AA, Gentles AJ, Weissman IL, Majeti R (2011) Prospective separation of normal and leukemic stem cells based on differential expression of TIM3, a human acute myeloid leukemia stem cell marker. Proc Natl Acad Sci USA 108(12):5009–5014
Jan M, Snyder TM, Corces-Zimmerman MR, Vyas P, Weissman IL, Quake SR, Majeti R (2012) Clonal evolution of preleukemic hematopoietic stem cells precedes human acute myeloid leukemia. Sci Transl Med 4(149):149ra118
Jordan CT, Lemischka IR (1990) Clonal and systemic analysis of long-term hematopoiesis in the mouse. Genes Dev 4(2):220–232
Jordan CT, Upchurch D, Szilvassy SJ, Guzman ML, Howard DS, Pettigrew AL, Meyerrose T, Rossi R, Grimes B, Rizzieri DA, Luger SM, Phillips GL (2000) The interleukin-3 receptor alpha chain is a unique marker for human acute myelogenous leukemia stem cells. Leukemia 14(10):1777–1784
Jorgensen HG, Holyoake TL (2007) Characterization of cancer stem cells in chronic myeloid leukaemia. Biochem Soc Trans 35(Pt 5):1347–1351
Kagoya Y, Yoshimi A, Kataoka K, Nakagawa M, Kumano K, Arai S, Kobayashi H, Saito T, Iwakura Y, Kurokawa M (2014) Positive feedback between NF-kappaB and TNF-alpha promotes leukemia-initiating cell capacity. J Clin Invest 124(2):528–542
Kerbauy DM, Lesnikov V, Torok-Storb B, Bryant E, Deeg HJ (2004) Engraftment of distinct clonal MDS-derived hematopoietic precursors in NOD/SCID-beta2-microglobulin-deficient mice after intramedullary transplantation of hematopoietic and stromal cells. Blood 104(7):2202–2203
Kiel MJ, Yilmaz OH, Iwashita T, Yilmaz OH, Terhorst C, Morrison SJ (2005) SLAM family receptors distinguish hematopoietic stem and progenitor cells and reveal endothelial niches for stem cells. Cell 121(7):1109–1121
Kikushige Y, Miyamoto T (2013) TIM-3 as a novel therapeutic target for eradicating acute myelogenous leukemia stem cells. Int J Hematol 98(6):627–633
Krivtsov AV, Twomey D, Feng Z, Stubbs MC, Wang Y, Faber J, Levine JE, Wang J, Hahn WC, Gilliland DG, Golub TR, Armstrong SA (2006) Transformation from committed progenitor to leukaemia stem cell initiated by MLL-AF9. Nature 442(7104):818–822
Krivtsov AV, Figueroa ME, Sinha AU, Stubbs MC, Feng Z, Valk PJ, Delwel R, Dohner K, Bullinger L, Kung AL, Melnick AM, Armstrong SA (2013) Cell of origin determines clinically relevant subtypes of MLL-rearranged AML. Leukemia 27(4):852–860
Kroeger H, Jelinek J, Estecio MR, He R, Kondo K, Chung W, Zhang L, Shen L, Kantarjian HM, Bueso-Ramos CE, Issa JP (2008) Aberrant CpG island methylation in acute myeloid leukemia is accentuated at relapse. Blood 112(4):1366–1373
Lagadinou ED, Sach A, Callahan K, Rossi RM, Neering SJ, Minhajuddin M, Ashton JM, Pei S, Grose V, O’Dwyer KM, Liesveld JL, Brookes PS, Becker MW, Jordan CT (2013) BCL-2 inhibition targets oxidative phosphorylation and selectively eradicates quiescent human leukemia stem cells. Cell Stem Cell 12(3):329–341
Lapidot T, Sirard C, Vormoor J, Murdoch B, Hoang T, Caceres-Cortes J, Minden M, Paterson B, Caligiuri MA, Dick JE (1994) A cell initiating human acute myeloid leukaemia after transplantation into SCID mice. Nature 367(6464):645–648
Liu Y, Cheng H, Gao S, Lu X, He F, Hu L, Hou D, Zou Z, Li Y, Zhang H, Xu J, Kang L, Wang Q, Yuan W, Gao S, Cheng T (2014) Reprogramming of MLL-AF9 leukemia cells into pluripotent stem cells. Leukemia 28(5):1071–1080
Luckey CJ, Bhattacharya D, Goldrath AW, Weissman IL, Benoist C, Mathis D (2006) Memory T and memory B cells share a transcriptional program of self-renewal with long-term hematopoietic stem cells. Proc Natl Acad Sci USA 103(9):3304–3309
Ma SK, Chan GC, Wan TS, Lam CK, Ha SY, Lau YL, Chan LC (1998) Near-haploid common acute lymphoblastic leukaemia of childhood with a second hyperdiploid line: a DNA ploidy and fluorescence in-situ hybridization study. Br J Haematol 103(3):750–755
Majeti R (2011) Monoclonal antibody therapy directed against human acute myeloid leukemia stem cells. Oncogene 30(9):1009–1019
Majeti R, Becker MW, Tian Q, Lee TL, Yan X, Liu R, Chiang JH, Hood L, Clarke MF, Weissman IL (2009) Dysregulated gene expression networks in human acute myelogenous leukemia stem cells. Proc Natl Acad Sci USA 106(9):3396–3401
Matsui W, Huff CA, Wang Q, Malehorn MT, Barber J, Tanhehco Y, Smith BD, Civin CI, Jones RJ (2004) Characterization of clonogenic multiple myeloma cells. Blood 103(6):2332–2336
Matsui W, Wang Q, Barber JP, Brennan S, Smith BD, Borrello I, McNiece I, Lin L, Ambinder RF, Peacock C, Watkins DN, Huff CA, Jones RJ (2008) Clonogenic multiple myeloma progenitors, stem cell properties, and drug resistance. Cancer Res 68(1):190–197
Maude SL, Frey N, Shaw PA, Aplenc R, Barrett DM, Bunin NJ, Chew A, Gonzalez VE, Zheng Z, Lacey SF, Mahnke YD, Melenhorst JJ, Rheingold SR, Shen A, Teachey DT, Levine BL, June CH, Porter DL, Grupp SA (2014) Chimeric antigen receptor T cells for sustained remissions in leukemia. N Engl J Med 371(16):1507–1517
McKenzie JL, Gan OI, Doedens M, Dick JE (2005) Human short-term repopulating stem cells are efficiently detected following intrafemoral transplantation into NOD/SCID recipients depleted of CD122+ cells. Blood 106(4):1259–1261
Miller PG, Al-Shahrour F, Hartwell KA, Chu LP, Jaras M, Puram RV, Puissant A, Callahan KP, Ashton J, McConkey ME, Poveromo LP, Cowley GS, Kharas MG, Labelle M, Shterental S, Fujisaki J, Silberstein L, Alexe G, Al-Hajj MA, Shelton CA, Armstrong SA, Root DE, Scadden DT, Hynes RO, Mukherjee S, Stegmaier K, Jordan CT, Ebert BL (2013) In Vivo RNAi screening identifies a leukemia-specific dependence on integrin beta 3 signaling. Cancer Cell 24(1):45–58
Miyamoto T, Weissman IL, Akashi K (2000) AML1/ETO-expressing nonleukemic stem cells in acute myelogenous leukemia with 8;21 chromosomal translocation. Proc Natl Acad Sci USA 97(13):7521–7526
Moran-Crusio K, Reavie L, Shih A, Abdel-Wahab O, Ndiaye-Lobry D, Lobry C, Figueroa ME, Vasanthakumar A, Patel J, Zhao X, Perna F, Pandey S, Madzo J, Song C, Dai Q, He C, Ibrahim S, Beran M, Zavadil J, Nimer SD, Melnick A, Godley LA, Aifantis I, Levine RL (2011) Tet2 loss leads to increased hematopoietic stem cell self-renewal and myeloid transformation. Cancer Cell 20(1):11–24
Moreau P, Voillat L, Benboukher L, Mathiot C, Dumontet C, Robillard N, Herault O, Garnache F, Garand R, Varoqueaux N, Avet-Loiseau H, Harousseau JL, Bataille R, IFM group (2007) Rituximab in CD20 positive multiple myeloma. Leukemia 21(4):835–836
Moriya K, Suzuki M, Watanabe Y, Takahashi T, Aoki Y, Uchiyama T, Kumaki S, Sasahara Y, Minegishi M, Kure S, Tsuchiya S, Sugamura K, Ishii N (2012) Development of a multi-step leukemogenesis model of MLL-rearranged leukemia using humanized mice. PLoS One 7(6), e37892
Morrison SJ, Weissman IL (1994) The long-term repopulating subset of hematopoietic stem cells is deterministic and isolatable by phenotype. Immunity 1(8):661–673
Muguruma Y, Matsushita H, Yahata T, Yumino S, Tanaka Y, Miyachi H, Ogawa Y, Kawada H, Ito M, Ando K (2011) Establishment of a xenograft model of human myelodysplastic syndromes. Haematologica 96(4):543–551
Nilsson L, Astrand-Grundstrom I, Arvidsson I, Jacobsson B, Hellstrom-Lindberg E, Hast R, Jacobsen SE (2000) Isolation and characterization of hematopoietic progenitor/stem cells in 5q-deleted myelodysplastic syndromes: evidence for involvement at the hematopoietic stem cell level. Blood 96(6):2012–2021
Nilsson L, Astrand-Grundstrom I, Anderson K, Arvidsson I, Hokland P, Bryder D, Kjeldsen L, Johansson B, Hellstrom-Lindberg E, Hast R, Jacobsen SE (2002) Involvement and functional impairment of the CD34(+)CD38(−)Thy-1(+) hematopoietic stem cell pool in myelodysplastic syndromes with trisomy 8. Blood 100(1):259–267
Notta F, Doulatov S, Laurenti E, Poeppl A, Jurisica I, Dick JE (2011) Isolation of single human hematopoietic stem cells capable of long-term multilineage engraftment. Science 333(6039):218–221
Nowell PC, Hungerford DA (1960) Chromosome studies on normal and leukemic human leukocytes. J Natl Cancer Inst 25:85–109
Peacock CD, Wang Q, Gesell GS, Corcoran-Schwartz IM, Jones E, Kim J, Devereux WL, Rhodes JT, Huff CA, Beachy PA, Watkins DN, Matsui W (2007) Hedgehog signaling maintains a tumor stem cell compartment in multiple myeloma. Proc Natl Acad Sci USA 104(10):4048–4053
Pei S, Jordan CT (2012) How close are we to targeting the leukemia stem cell? Best Pract Res Clin Haematol 25(4):415–418
Pei S, Minhajuddin M, Callahan KP, Balys M, Ashton JM, Neering SJ, Lagadinou ED, Corbett C, Ye H, Liesveld JL, O’Dwyer KM, Li Z, Shi L, Greninger P, Settleman J, Benes C, Hagen FK, Munger J, Crooks PA, Becker MW, Jordan CT (2013) Targeting aberrant glutathione metabolism to eradicate human acute myelogenous leukemia cells. J Biol Chem 288(47):33542–33558
Raaijmakers MH, Mukherjee S, Guo S, Zhang S, Kobayashi T, Schoonmaker JA, Ebert BL, Al-Shahrour F, Hasserjian RP, Scadden EO, Aung Z, Matza M, Merkenschlager M, Lin C, Rommens JM, Scadden DT (2010) Bone progenitor dysfunction induces myelodysplasia and secondary leukaemia. Nature 464(7290):852–857
Raskind WH, Tirumali N, Jacobson R, Singer J, Fialkow PJ (1984) Evidence for a multistep pathogenesis of a myelodysplastic syndrome. Blood 63(6):1318–1323
Rau R, Magoon D, Greenblatt S, Li L, Annesley C, Duffield AS, Huso D, McIntyre E, Clohessy JG, Reschke M, Pandolfi PP, Small D, Brown P (2014) NPMc + cooperates with Flt3/ITD mutations to cause acute leukemia recapitulating human disease. Exp Hematol 42(2):101–113e105
Rieger MA, Schroeder T (2012) Hematopoiesis. Cold Spring Harb Perspect Biol 4(12):a008250
Ritchie DS, Neeson PJ, Khot A, Peinert S, Tai T, Tainton K, Chen K, Shin M, Wall DM, Honemann D, Gambell P, Westerman DA, Haurat J, Westwood JA, Scott AM, Kravets L, Dickinson M, Trapani JA, Smyth MJ, Darcy PK, Kershaw MH, Prince HM (2013) Persistence and efficacy of second generation CAR T cell against the LeY antigen in acute myeloid leukemia. Mol Ther 21(11):2122–2129
Rousselot P, Huguet F, Rea D, Legros L, Cayuela JM, Maarek O, Blanchet O, Marit G, Gluckman E, Reiffers J, Gardembas M, Mahon FX (2007) Imatinib mesylate discontinuation in patients with chronic myelogenous leukemia in complete molecular remission for more than 2 years. Blood 109(1):58–60
Rowley JD (1973) Letter: a new consistent chromosomal abnormality in chronic myelogenous leukaemia identified by quinacrine fluorescence and giemsa staining. Nature 243(5405):290–293
Sarry JE, Murphy K, Perry R, Sanchez PV, Secreto A, Keefer C, Swider CR, Strzelecki AC, Cavelier C, Recher C, Mansat-De Mas V, Delabesse E, Danet-Desnoyers G, Carroll M (2011) Human acute myelogenous leukemia stem cells are rare and heterogeneous when assayed in NOD/SCID/IL2Rgammac-deficient mice. J Clin Invest 121(1):384–395
Shlush LI, Zandi S, Mitchell A, Chen WC, Brandwein JM, Gupta V, Kennedy JA, Schimmer AD, Schuh AC, Yee KW, McLeod JL, Doedens M, Medeiros JJ, Marke R, Kim HJ, Lee K, McPherson JD, Hudson TJ, Brown AM, Yousif F, Trinh QM, Stein LD, Minden MD, Wang JC, Dick JE, Consortium HP-LGP (2014) Identification of pre-leukaemic haematopoietic stem cells in acute leukaemia. Nature 506(7488):328–333
Shultz LD, Lyons BL, Burzenski LM, Gott B, Chen X, Chaleff S, Kotb M, Gillies SD, King M, Mangada J, Greiner DL, Handgretinger R (2005) Human lymphoid and myeloid cell development in NOD/LtSz-scid IL2R gamma null mice engrafted with mobilized human hemopoietic stem cells. J Immunol 174(10):6477–6489
Taussig DC, Pearce DJ, Simpson C, Rohatiner AZ, Lister TA, Kelly G, Luongo JL, Danet-Desnoyers GA, Bonnet D (2005) Hematopoietic stem cells express multiple myeloid markers: implications for the origin and targeted therapy of acute myeloid leukemia. Blood 106(13):4086–4092
Taussig DC, Miraki-Moud F, Anjos-Afonso F, Pearce DJ, Allen K, Ridler C, Lillington D, Oakervee H, Cavenagh J, Agrawal SG, Lister TA, Gribben JG, Bonnet D (2008) Anti-CD38 antibody-mediated clearance of human repopulating cells masks the heterogeneity of leukemia-initiating cells. Blood 112(3):568–575
Taussig DC, Vargaftig J, Miraki-Moud F, Griessinger E, Sharrock K, Luke T, Lillington D, Oakervee H, Cavenagh J, Agrawal SG, Lister TA, Gribben JG, Bonnet D (2010) Leukemia-initiating cells from some acute myeloid leukemia patients with mutated nucleophosmin reside in the CD34(−) fraction. Blood 115(10):1976–1984
Tehranchi R, Woll PS, Anderson K, Buza-Vidas N, Mizukami T, Mead AJ, Astrand-Grundstrom I, Strombeck B, Horvat A, Ferry H, Dhanda RS, Hast R, Ryden T, Vyas P, Gohring G, Schlegelberger B, Johansson B, Hellstrom-Lindberg E, List A, Nilsson L, Jacobsen SE (2010) Persistent malignant stem cells in del(5q) myelodysplasia in remission. N Engl J Med 363(11):1025–1037
Teitell MA, Pandolfi PP (2009) Molecular genetics of acute lymphoblastic leukemia. Annu Rev Pathol 4:175–198
Uchida N, Dykstra B, Lyons KJ, Leung FY, Eaves CJ (2003) Different in vivo repopulating activities of purified hematopoietic stem cells before and after being stimulated to divide in vitro with the same kinetics. Exp Hematol 31(12):1338–1347
Wang Y, Krivtsov AV, Sinha AU, North TE, Goessling W, Feng Z, Zon LI, Armstrong SA (2010) The Wnt/beta-catenin pathway is required for the development of leukemia stem cells in AML. Science 327(5973):1650–1653
Weiss BM, Abadie J, Verma P, Howard RS, Kuehl WM (2009) A monoclonal gammopathy precedes multiple myeloma in most patients. Blood 113(22):5418–5422
Welch JS, Ley TJ, Link DC, Miller CA, Larson DE, Koboldt DC, Wartman LD, Lamprecht TL, Liu F, Xia J, Kandoth C, Fulton RS, McLellan MD, Dooling DJ, Wallis JW, Chen K, Harris CC, Schmidt HK, Kalicki-Veizer JM, Lu C, Zhang Q, Lin L, O’Laughlin MD, McMichael JF, Delehaunty KD, Fulton LA, Magrini VJ, McGrath SD, Demeter RT, Vickery TL, Hundal J, Cook LL, Swift GW, Reed JP, Alldredge PA, Wylie TN, Walker JR, Watson MA, Heath SE, Shannon WD, Varghese N, Nagarajan R, Payton JE, Baty JD, Kulkarni S, Klco JM, Tomasson MH, Westervelt P, Walter MJ, Graubert TA, DiPersio JF, Ding L, Mardis ER, Wilson RK (2012) The origin and evolution of mutations in acute myeloid leukemia. Cell 150(2):264–278
Will B, Zhou L, Vogler TO, Ben-Neriah S, Schinke C, Tamari R, Yu Y, Bhagat TD, Bhattacharyya S, Barreyro L, Heuck C, Mo Y, Parekh S, McMahon C, Pellagatti A, Boultwood J, Montagna C, Silverman L, Maciejewski J, Greally JM, Ye BH, List AF, Steidl C, Steidl U, Verma A (2012) Stem and progenitor cells in myelodysplastic syndromes show aberrant stage-specific expansion and harbor genetic and epigenetic alterations. Blood 120(10):2076–2086
Wright JJ, Poplack DG, Bakhshi A, Reaman G, Cole D, Jensen JP, Korsmeyer SJ (1987) Gene rearrangements as markers of clonal variation and minimal residual disease in acute lymphoblastic leukemia. J Clin Oncol 5(5):735–741
Xie M, Lu C, Wang J, McLellan MD, Johnson KJ, Wendl MC, McMichael JF, Schmidt HK, Yellapantula V, Miller CA, Ozenberger BA, Welch JS, Link DC, Walter MJ, Mardis ER, Dipersio JF, Chen F, Wilson RK, Ley TJ, Ding L (2014) Age-related mutations associated with clonal hematopoietic expansion and malignancies. Nat Med 20(12):1472–1478
Yaccoby S, Barlogie B, Epstein J (1998) Primary myeloma cells growing in SCID-hu mice: a model for studying the biology and treatment of myeloma and its manifestations. Blood 92(8):2908–2913
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Becker, M.W., O’Dwyer, K.M. (2015). Leukemic and Lymphoid Stem Cells. In: Babashah, S. (eds) Cancer Stem Cells: Emerging Concepts and Future Perspectives in Translational Oncology. Springer, Cham. https://doi.org/10.1007/978-3-319-21030-8_13
Download citation
DOI: https://doi.org/10.1007/978-3-319-21030-8_13
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-21029-2
Online ISBN: 978-3-319-21030-8
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)