Abstract
Poly(ADP-ribose) polymerase-1 (PARP-1) is an abundant nuclear enzyme and the founding member of the PARP family of enzymes. Inhibition of PARP-1 has been the focus of drug discovery groups for over three decades in a wide range of therapeutic areas encompassing stroke, cardiac ischemia, inflammation, diabetes and most importantly cancer. Despite the great therapeutic potential for this target and over a decade of clinical studies, only recently have PARP inhibitors made headway in late stage clinical trials. After many tribulations, recent results from several PARP inhibitors in Phase II clinical trials for cancer therapy have reinvigorated the field. This chapter is structured to provide the readers with a brief summary of the rationale for PARP-1 as a therapeutic target for oncology and explain the genesis of the PARP inhibitor pharmacophore. In addition, this chapter will provide the optimization paradigms for each of the PARP inhibitors currently in clinical trials, analyzing some of the differentiating factors for the clinical compounds and a brief mention of the current clinical progress for each inhibitor.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 PARP-1 as a Therapeutic Target for Oncology
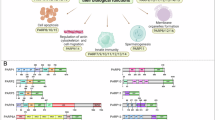
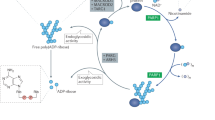
PARP-1 is the founding member of a family of 17 enzymes, many of which use nicotinamide adenine dinucleotide (1, NAD+, Fig. 7.1) as a substrate to form either mono- or polyADP(ribose) adducts on acceptor proteins. PARP-1 has three major domains, a catalytic domain, a DNA binding domain, and an automodification domain each of which play an active role in DNA repair, specifically base excision repair (BER) and maintenance of genomic function (see Chap. 3). To a lesser extent, these DNA repair functions are performed by PARP-2, the closest homolog to PARP-1 [1]. The zinc fingers of the DNA binding domain are critical for identifying DNA damage and binding the damaged site. This binding event results in a structural change that causes the automodification domain to be closely associated with the catalytic domain and thus become ADP-ribosylated using NAD+ as a substrate [2]. This event activates the catalytic machinery [3] of PARP-1/2, [4] prompting the construction of branched chain polymers of ADP-ribose onto nearby histone DNA binding proteins [5]. The large, negatively charged polymers act to relax the tertiary structure of the chromatin and provide a platform for the recruitment of DNA repair enzymes such as XRCC1, [6] and DNA ligase [7]. The ADP(ribose) polymers are broken down by poly (ADP-ribose) glycohydrolase (PARG) leading to further access to the damaged DNA by the repair enzymes. If this single strand repair does not occur, the single strand breaks can be converted to double strand breaks leading to further genomic destabilization ultimately resulting in apoptotic cell death [8]. This PARP mediated repair pathway is a major mechanism for DNA repair by many cancerous cell types leading to drug resistance by DNA damaging chemotherapeutics and continued tumor growth. Hence, a PARP-1/2 inhibitor in combination with the DNA damaging chemotherapeutics (Chap. 9 and 10) or radiation (Chap. 11) would compromise the cancer cell DNA repair mechanisms resulting in genomic dysfunction and cell death. Furthermore, PARP-1/2 inhibitors can be used as a monotherapy for tumor types that are already deficient in certain types of DNA repair mechanisms (e.g. homologous recombination, Chap. 13). This phenomenon is referred to as synthetic lethality, namely the loss of one DNA repair function will result in cell susceptibility, but the loss of both is lethal (e.g. BRCA1/2 deficient cells and a PARP-1 inhibitor). Over the past decade, the improvement in genotyping tumors [9] has allowed clinicians to more accurately identify specific tumor types or cell types that are susceptible to PARP-1 inhibitors (Chap. 21). This genotyping has played a major role the advancement PARP-1/2 inhibitors in the clinic by identifying cancer patients with the greatest likelihood to benefit from PARP inhibitor therapy (Chap. 21).
The breadth of PARP-1 oncological research along with several other factors led to an effective optimization paradigm that many of the medicinal chemistry programs followed to discover clinical candidate PARP-1 inhibitors. Many of the 1st generation PARP-1 inhibitors were discovered by optimization of several of the following parameters: (1) Enzymatic inhibition of PARP-1; (2) in vitro characterization in cancer cell lines and the ability to potentiate the cytotoxicity of chemotherapeutic agents; (3) ability to kill cells that are deficient in DNA repair mechanisms; (4) physicochemical properties (i.e. solubility, metabolic stability, oral bioavailability); and (5) ability to potentiate chemotherapeutic agents in vivo in xenograft models. Because PARP-1 inhibitors have been in the clinic since 2003, many medicinal chemistry groups have developed 2nd generation inhibitors which often include some of the following characteristics in the screening paradigms: (1) In vitro profiling of PARP inhibitors against other members of the PARP family, specifically PARP-2; (2) Head to head comparison with current benchmark PARP-1 inhibitors in vitro and in vivo; (3) ability to trap the PARP enzyme in a tight complex with DNA or ‘PARP trapping’ [10]; (4) ability to kill cancer cells that are resistant to other PARP-1 inhibitors. This chapter will outline the genesis of the PARP-1 pharmacophore and highlight the design and optimization of all of the PARP inhibitors currently in clinical trials.
2 Evolution of the Pharmacophore for NAD+ Competitive PARP Inhibitors
2.1 Substituted Benzamides
Some of the earliest PARP-1 inhibitors (a.k.a. Poly(ADP-ribose)synthetase inhibitors) were discovered in the 1970’s [11] and 1980’s [12]. At this time, over a decade of research had been conducted on ADP(ribosylation) , but the function of this type of protein modification was largely unknown. With the discovery that nicotinamide (2, NA, Fig. 7.1) was a modest PARP inhibitor, [11] studies were conducted to determine the physiological significance of PARP-1 inhibition using 2 as a tool compound [11]. However, nicotinamide has a variety of cellular functions not related to PARP inhibition, clouding the interpretation of the results of these studies. The need for more specific inhibitors of PARP-1 would become vital in delineating the role of this nuclear enzyme. Despite the relatively weak potency of 2 (IC50 = 210 µM), [13] it was a better lead as a substrate based inhibitor than adenine and other nucleoside and purine derivatives, [14] providing an more fruitful direction for identifying more specific PARP inhibitors (blue, Fig. 7.1). Structural analogs of 2 were next identified such as benzamide (3, 96 % inhibition of porcine PARP @50 µM) [15] and substituted benzamides (i.e. 3-aminobenzamide, 3-AB, 4, 90 % inhibition of porcine PARP @50 µM) [12].
The identification of substituted benzamides as some of the most potent PARP-1 inhibitors at the time prompted further synthetic efforts. Early benzamide analogs established basic PARP SAR [12] as 3-aminobenzoic acid (5), 3-amino acetophenone (6), and N-methyl-3-amino benzamide (7) were all inactive providing early indications of the importance of the aryl amide moiety to inhibition (Fig. 7.2). In addition, the 2- and 4-aminobenzamides (8 and 9, Fig. 7.2) had no inhibitory effect towards porcine PARP indicating that substitutions at the three position were preferred. Thus, 3-AB showed early distinction as a tool compound for the inhibition of PARP-1. As opposed to nicotinamide, 3-AB did not interact with other nicotinamide binding enzymes, thus it was a more desirable probe compound to study and understand PARP-1 inhibition. Indeed, shortly after identifying 3-AB, the role of PARP-1 in DNA base excision repair was discovered further advancing the field [3].
2.2 Refinement of PARP-1 Pharmacophore, Polycyclic Amides
The next advancement in the discovery of more potent PARP-1 inhibitors occurred in the early 1990’s, as bicyclic aryl amides started to emerge as sub micromolar inhibitors. Using 3-AB as a template, a group from Parke-Davis hypothesized that the orientation of the amide with respect to the substituent at the 3-position is important for optimal activity [16]. This group used 5- vs 7-substituted dihydroisoquinolinones to restrict the rotational energy of the benzamide to test this hypothesis (Fig. 7.3). The results of their work clearly demonstrated that 5-substituted derivatives 10b-d were 1–2 orders of magnitude more potent than their isomeric 7-substituted analogs 11a-c. This work refined the PARP-1 pharmacophore by demonstrating that constraining the aryl amide into another ring would restrict the degrees of freedom for the amide moiety, thus locking it into a conformation more beneficial for PARP-1 inhibitory potency. Another refinement of the PARP pharmacophore was unearthed with this publication, namely, the improvement in potency upon addition of a heteroatom in the 5-position (10a vs 10b-d, Fig. 7.3).
Perhaps the most comprehensive study to solidify the pharmacophore was conducted by Banasik and Ueda [13]. This group screened over 100 compounds from several structural classes against bovine PARP to discover multiple bicyclic and tricyclic lactams as low micromolar PARP inhibitors. Some of the compounds identified through this screen such as the isoquinolinones (10) and dihydroisoquionlinones (12) were previously identified, [16] but several other polycyclic aryl amide cores were also discovered. This ‘analoging by catalogue’ effort unearthed several basic PARP inhibitor scaffolds which were further optimized by future drug discovery groups such as phenanthridinones (14), [17] quinazolinones (15), [18] quinazoline diones (16), phthalazinones (17) (Fig. 7.4) [19]. Perhaps the most important of which is the phthalazinones (17, in box, Fig. 7.4), a bicyclic amide scaffold that is the core scaffold for three clinical candidates (vide infra).
2.3 Discovery of Benzimidazole Carboxamides
Another important ring system was identified shortly after the Banasik and Suto publications in the mid 1990’s. Inspired by the potent, cyclic aryl amide motifs, the group of Golding and Griffin designed ‘pseudocycles’ such as benzoxazole carboxamides 18a-b and more importantly, benzimidazole carboxamides 19a-b [20]. The benzimidazole carboxamides turned out to have remarkable enzymatic potency against PARP from L1210 cells when compared to the benzoxazole carboxamides (compare 19a vs 18a and 19b vs 18b). These series effectively lock the aryl amide in the desired conformation through an intramolecular hydrogen bond (in circle, Fig. 7.5) while incorporating a heteroatom three carbons from the aryl amide moiety (orange circle, Fig. 7.5). This potent, compact, easily derivatizable core scaffold was the inspiration for three clinical candidates as discussed below.
3 PARP Pharmacophore for NAD+ Competitive Inhibitors
With the discovery of many of the bicyclic aryl amides in the mid-1990s, the classical PARP pharmacophore was established with minor refinements still to come. Interestingly, this pharmacophore has remained largely unchanged throughout the decades of medicinal chemistry efforts and thousands of compounds tested [19]. With the advent of the first co-crystal structure of some 1st generation PARP inhibitors and chicken PARP [21] many of the aspects of this pharmacophore could be explained based on the active site binding interactions. The PARP-1 pharmacophore includes one or more of the following structural elements contributing to the inhibitory potency: (1) An aryl amide moiety fused within a bicyclic ring system or ‘pseudo bicyclic ring’ (e.g. 18 and 19) system as outlined in blue by rings A and B (Fig. 7.6), this amide is locked into a hydrogen bonding network with Ser904 and Gly863 of human PARP-1, [22] any disruption of this network such as nearby substituents resulted in loss of potency [16]. The multi-cyclic aryl moiety was preferred because two aryl residues, Tyr896 and Tyr907 form a π-electron pocket explaining the improvement in potency often seen with aryl amides versus aliphatic amides. (2) Hydrogen bond donors or acceptors on the opposite side of the A-ring from the amide (orange, Fig. 7.6). These heteroatoms are able to form either a direct or a water-mediated hydrogen bond with Glu988. (3) Small hydrophobic substituents on the A-ring, adjacent to the amide (pink, Fig. 7.6). The back wall of the nicotinamide subsite is bordered by Ala898 and Lys903 forming a tight pocket just large enough for a small substituent (e.g. CH3, F, Cl) on the aryl amide ring. (4) Large hydrophobic groups in the southeast portion of the pharmacophore (green, Fig. 7.6). These groups generally fill the large hydrophobic pocket adjacent to the nicotinamide binding site. This pocket is often referred to as the adenine-ribose binding site (AD site) and most series of PARP-1 inhibitors take advantage of this spacious pocket to improve potency, solubility and other pharmaceutical properties.
While there are examples of PARP inhibitors that are outside this pharmacophore, [23–26] profiling efforts of many of the 1st generation PARP inhibitors revealed that many of them also inhibit PARP-2, PARP-3 and PARP-4, indicating that this basic pharmacophore extends to other members of the PARP family of enzymes [27]. Several research efforts are ongoing to discover selective inhibitors for each member of the PARP family, including several successful examples of selective, NAD+ competitive inhibitors of TNKS1 and TNKS2 (PARP 5a/5b) that diverge significantly from this pharmacophore [28, 29].
4 PARP-1 Medicinal Chemistry Programs and Optimization Strategies Leading to Clinical Candidates
4.1 Newcastle/Agouron/Pfizer/Clovis (AG014699, PF-0137338, CO-338, Rucaparib)
Collaboration between the Newcastle group and Agouron Pharmaceuticals in the late 1990’s resulted in a series of tricyclic indole lactams derived from the optimization of tricyclic benzimidazole carboxamides as outlined in Fig. 7.7. The Newcastle group discovered that benzimidazole carboxamides proved to be a remarkably potent core structure (19c, Ki = 95 nM against human PARP-1), the most potent aryl amide scaffold identified at the time [30]. Optimization of these benzimidazole carboxamides led to several 2-aryl derivatives with low nanomolar potency. The first lead compound that emerged from this series was NU1085 (20, Ki = 6 nM, Fig. 7.7). In vitro, 10 μM NU1085 potentiated growth inhibition of Temozolomide and Topotecan over twofold in A2780 cells. However, this compound still suffered from poor aqueous solubility prompting an optimization pathway focused on better physicochemical properties.
Newcastle and Agouron (Pfizer) designed several series of compounds derived from benzimidazole carboxamides through structure based design in order to address the solubility issues and at the same time improve the structural novelty of their PARP-1 inhibitors [22, 31–33]. These groups deduced that the free carboxamide of 19c could be constrained within a 7-membered ring affording the same result as the intramolecular H-bond of benzimidazole carboxamides. Furthermore, keeping the aryl group in the same relative position and adding tertiary amine substituents in the 4-position (green, Fig. 7.7) they were able to improve the aqueous solubility, inhibitory potency and cell permeability [31]. The high potency of 21 (Ki = 5.6 nM) led to increased antiproliferative activity of TMZ against LoVo cells (PF50 = 7.8 when 21 was tested at 0.4 μM). This lead PARP inhibitor also displayed in vivo efficacy by causing complete regression of SW620 xenograft tumors (i.p daily at 5 or 15 mg/kg) in combination with TMZ (68 mg/kg p.o. daily for 5 days). Compound 21 served as the benchmark lead for the Agouron group as they designed other closely related series of PARP-1 inhibitors [33]. Much of the design for this series of compounds was made possible through cocrystal data (see Chap. 9). As expected, the constrained amide formed an H-bond network with Ser904 and Gly863 similar to many of the early PARP-1 inhibitors, in addition, the indole NH formed a water mediated H-bond with Glu988 (orange, Fig. 7.7), the aryl substituent formed a π–π interaction with Tyr889 and the tertiary amino group interacted with Asp766 in the AD pocket.
The eventual clinical candidate, (22, AG014699, PF01367338, CO-338, Rucaparib) was optimized from a series of 5,6,7 tricyclic indole lactams. Compound 22 displayed better in vitro potency (PARP-1 Ki = 1.4 nM) and in vivo efficacy than 21 (PF50 of 8.1 in LoVo cells) [34]. Because many of the [5–7] tricyclic lactam PARP-1 inhibitors had similar potency and potentiation factors, the selection strategy for the clinical candidate assessed the potency of lead inhibitors in rodent xenograft studies in the presence of TMZ. AG014699 , when dosed at 0.15 mg/kg/day i.p. exhibited a 50 % increase in tumor growth delay over many of the other closely related lead compounds in a 5 day xenograft study in conjunction with TMZ (68 mg/kg/day). AG014699 also displayed no toxicity alone or in combination with TMZ and no adverse effects on the PK of the co-administered anticancer agents. In 2003, this collaboration culminated in the first PARP-1 inhibitor in human clinical trials as a chemopotentiator.
Almost a decade has passed since Rucaparib entered into the clinic. During that time, research efforts have been conducted to further characterize Rucaparib in vitro and as well as the clinic. Most of the extra-clinical research has focused on using Rucaparib as a benchmark PARP inhibitor in comparison with some of the other 1st and 2nd generation PARP inhibitor clinical compounds [35]. Profiling efforts were conducted with over 150 PARP inhibitors to assess the relative selectivity against other members of the PARP family [27]. Interestingly, Rucaparib was singled out as being one of the least selective PARP inhibitors (i.e. a pan-PARP inhibitor). Rucaparib demonstrated significant binding potential (thermal shifts of 1.2–14.4 °C) towards several members of the PARP family. Furthermore, Rucaparib demonstrated some PARP-independent anti-tumor activity by stimulating phosphorylation of Akt/protein kinase B and decreasing phosphorylation at Stat3 [35].
In June 2011, despite some rather equivocal Phase 2 clinical results as a chemopotentiator, [36] the Clovis Oncology group licensed Rucaparib from Pfizer taking over global development and potential commercialization of the drug. Advancements in genotyping technology [37] provided the means for identifying patients more likely to respond to PARP inhibitors and a potential clinical path forward for the Clovis Oncology group. Clovis recently completed a Phase 2 biomarker study in which tumors were sequenced and Rucaparib response was correlated with genotype of patient. Using this information, Clovis identified several gene mutations correlative of efficacy which will be used in a recently initiated a Phase 3 placebo controlled clinical trial. The purpose of this trial is to determine which patients with ovarian, fallopian tube and primary peritoneal cancer will respond to oral Rucaparib (NCT01968213).
4.2 BASF/Abbott/AbbVie (ABT-888, Veliparib)
The BASF group initiated efforts to develop a PARP inhibitor in the late 1990’s covering a wide range of scaffolds, specifically 2-alkylamino benzimidazoles (19c), the eventual series from which Abbott’s clinical candidate emerged several years later. Abbott labs further advanced the progress that BASF made in the PARP field after the acquisition of BASF’s pharmaceutical division in 2001.
Attracted by the ligand efficiency of the benzimidazole carboxamide core (19c, Ki = 95 nM, MW = 161), the Abbott group aggressively synthesized and characterized several hundred 2-alkylamino derivatives of this scaffold [38, 39]. Their screening paradigm selected compounds with < 10 nM enzymatic potency and < 10 nM cellular potency (C41 peroxide damaged cellular assay) before advancing them into further in vivo studies. This optimization strategy led to the discovery of preclinical candidate A-620223 (23, IC50 = 8 nM, Fig. 7.8). As several other PARP medicinal chemistry groups discovered, having a secondary or tertiary amine provided adequate solubility (> 5 mg/mL for A-620223) while improving the cellular potency in a peroxide induced DNA damage cellular assay (EC50 = 3 nM). The alkyl substitution on the piperidine (green, Fig. 7.8) also contributed dramatically to an improvement in cellular potency over the unsubstituted analog. A-620223 displayed good oral bioavailability across species (32–82 %) and terminal elimination half-lives of 1.2–2.7 h in the same species. This compound demonstrated chemopotentiation of TMZ (74–83 % tumor growth inhibition vs 62 % for TMZ alone) in a B16F10 melanoma model at 1 mg/kg/day over 14 days. In addition, this compound potentiated the effect of cisplatin in an MX-1 breast cancer tumor model at slightly higher doses [38].
During the optimization process towards Veliparib (24, ABT-888, Veliparib, Ki = 5 nM, EC50 = 2 nM) , Abbott discovered that a quaternary carbon in the 2-position of the benzimidazole ring system was beneficial for both enzymatic potency and cellular efficacy (grey and light green, Fig. 7.8) [39]. Consistently, compounds with this feature were 2–13 times more potent in the C41 cellular assay than compounds with a tertiary carbon. While the enzymatic potency of 24 and its (S)-enantiomer were identical (Ki = 5 nM), stereochemistry played an important role in both the oral bioavailability and exposure leading to the selection of the (R)-enantiomer as the clinical candidate. This drug displayed excellent oral bioavailability across species (56–92 %) and a comparable terminal half-life to their other preclinical leads such as A-620223 (1.2 h). In addition, ABT-888 demonstrated excellent chemopotentiation in preclinical xenograft models. In a B16F10 melanoma model when administered orally in combination with TMZ demonstrated a 43–64 % tumor growth inhibition at 1, 5 and 12.5 mg/kg/day. Chemopotentiation was also observed in combination with carboplatin in an MX-1 breast cancer tumor model [39, 40].
Veliparib entered the clinic in 2006 and since that time over 60 clinical studies have been initiated with this compound. The most exciting results occurred in a recent ISPY2 Trial (Investigation of Serial Studies to Predict Your Therapeutic Response with Imaging And moLecular Analysis 2) a large scale breast cancer clinical study conducted by the Biomarkers Consortium, a creative public-private partnership which includes the FDA, NIH and several major pharmaceutical companies. The concept for this trial was to use a combination of MRI (Mammaprint score) and biomarkers (HER+/− and ER+/−) to categorize patients. Using this categorization data, the patients would then receive one of the 20+ participating drugs that are the most statistically likely to benefit the patient. Results from this trial are expected to assist in the design of Phase III studies. Veliparib was considered the first ‘graduate’ of the study after demonstrating a 52 % complete response rate in women with triple negative breast cancer when dosed with carboplatin [41]. The results of this trial have set the stage for Phase III breast cancer study by AbbVie.
4.3 Merck/Tesaro (MK-4827, Niraparib)
Niraparib (26, MK-4827) evolved from a series of heterocyclic benzamides related to benzimidazole carboxamides (Fig. 7.9). An initial optimization paradigm selected an indazole subseries over several other 5-membered heterocycles (grey, Fig. 7.9) because it had more desirable pharmacokinetics, enzymatic and cellular potency [42]. The 2-phenyl substituted indazole 25 demonstrated better potency (IC50 = 24 nM) than any of the other sub-series tested. It also displayed adequate cellular potency showing the ability to inhibit PAR formation after induction of DNA damage by peroxide in HeLa cells (EC50 = 3.7 μM). In addition, 25 demonstrated moderate stability in rat and human microsomes, acceptable oral BA (41 %) and terminal half-life (5.1 h) in rats. Further optimization of the this series was accomplished by incorporating a solubilizing amine group (green, Fig. 7.9) on the para position of the aryl ring leading to general improvements in enzymatic and cellular potency. Cellular potency of lead compounds was established in BRCA1 silenced HeLa cells for the ability to inhibit cell growth by 50 % (CC50) versus BRCA1 wild type HeLa cells. The (S)-piperidine was selected as the clinical candidate (blue, Fig. 7.9) because it afforded ~ 28 fold selectivity against BRCA1 silenced cells (CC50 = 33 nM) versus wild type (CC50 = 860 nM), while the (R)-enantiomer was only ~ 11 fold less selective. MK-4827 displayed excellent oral bioavailability in rats (65 %) as well as a high volume of distribution (Vdss = 6.9 L/kg) and a long terminal half-life (t½ = 3.4 h). MK-4827 demonstrated tumor regression in a BRCA-1 mutant MDA-MB-436 xenograft model orally at 100 mg/kg q.d. or 50 mg/kg b.i.d. with no overt weight loss or signs of toxicity.
Niraparib also achieved distinction for being an effective ‘PARP poison’ by being able to trap the PARP-DNA complex better than other PARP clinical compounds such as Olaparib and Veliparib [10]. Efficacy of PARP inhibitors can be achieved not only by inhibition of the catalytic activity, but also by trapping PARP-DNA complexes. This trapping prevents the dissociation of PARP from DNA resulting in inadequate DNA repair. Because all of the clinical PARP inhibitors have similar enzymatic potency, this trapping ability mechanism may account for some of the discrepancies in cellular toxicity and perhaps result in clinical differences as well, particularly if trapping does not occur at clinically relevant concentrations. From this standpoint, PARP inhibitors such as Niraparib may distinguish itself from some of the 1st generation inhibitors in the clinic.
Niraparib could be considered a relatively late entrant into the PARP-1 clinical scene in 2008 with a Phase I dose escalation study [43]. This study indicated that Niraparib was generally well tolerated at 300 mg per day, with a low rate of grade 3/4 toxicities and some grade 1/2 anemia, fatigue and nausea. At this dose, 75 % (three of four patients) with platinum sensitive high grade serous ovarian cancer achieved a RECIST response. In addition, a RECIST response rate of 50 % (5/10 patients) was achieved in patients with platinum sensitive ovarian cancer and germline BRCA mutations [44]. In 2012, Tesaro licensed Niraparib from Merck and rapidly propelled the compound into two Phase III trials, one in patients with HER2 negative, germline BRCA positive breast cancer (NCT 01905592) and another in platinum sensitive ovarian cancer (NCT01847274).
4.4 Maybridge/Kudos/AstraZeneca (KU 59436, AZD2281, Olaparib and 2nd Generation PARP Inhibitor AZD2461)
The discovery of Olaparib started with a collaboration between KuDOS pharmaceuticals and Maybridge in the early 2000’s. Through this collaboration, these groups unearthed several hits from the Maybridge chemical screening library, the most important of which was a 4-substituted benzyl phthalazinone 27 (PARP-1 IC50 = 770 nM, Fig. 7.10) [45]. The incorporation of the benzyl group improved the potency of the core phthalazinone (17, IC50 = 9 µM) identified almost a decade earlier by Banasik [13]. This ~ 10 fold improvement in potency along with an expedient synthetic route provided a definitive direction for medicinal chemistry optimization efforts. The optimization paradigm that identified Olaparib consisted of several key hurdles: (1) improving the PARP-1 in vitro potency (< 10 nM); (2) optimizing the cell killing activity in HeLa B cells treated with the alkylating agent methyl methanesulfonate (i.e. PF50); (3) obtaining good physicochemical properties (MW < 550, PSA < 140 Å, rotatable bonds < 7, hydrogen bond donors/acceptors < 10, solubility > 0.1 mg/mL); (4) demonstrating oral bioavailability; (5) demonstrating activity in BRCA-1/2 deficient cell lines; (6) demonstrating activity in vivo, in SW-620 tumor xenograft models [45, 46].
Initial improvements in PARP-1 potency and cellular activity were achieved through the addition of a succinimide group on the 3-position of the benzyl ring (blue, Fig. 7.10) [47]. An interesting trend was noted during the optimization of this series, namely, addition of a fluorine in the 4-position (orange, Fig. 7.10) as in lead compound 28 (KU 58684) had a rather dramatic effect on the cellular potency (PF50 = 5.6). On average, the addition of the fluorine in that position resulted in a > 2 fold improvement in the PF50 perhaps as the authors speculate it is due to restricted conformational rotation and molecular entropy leading to greater permeability. Replacement of the succinimide with a diacylpiperazine moiety (green, Fig. 7.10) improved the solubility and oral bioavailability culminating in the discovery of 29 (KU 59436, AZD2281, Olaparib), the eventual clinical candidate. This compound maintained the PARP-1 inhibitory potency (IC50 = 5 nM) and displayed a markedly improved cellular potency (PF50 = 25.8). In addition, 29 proved to have excellent pharmacokinetic data in rats (t1/2 = 53 min, Cl = 49 mL/min/kg, BA = 100 %) and dogs (t1/2 = 170 min, Cl = 5.4 mL/min/kg, BA = 100 %). The chemopotentiating activity of 29 was observed in SW620 cells when treated with MMS, showing a maximal effect at ~ 100 nM. The cellular potency in BRCA mutant cell lines (MDA-MD-463 and HCC1937) was also observed, plateauing at ~ 0.5 µM. When dosed orally at 10 mg/kg in combination with TMZ (50 mg/kg), compound 29 displayed a remarkable reduction in mean tumor volume throughout the terminal phase of the xenograft study without exacerbating the toxicological effects of TMZ [45].
In 2005, KU 59436 entered into the clinic and shortly thereafter the compound was acquired by AstraZeneca through the purchase of KuDOS pharmaceuticals. The clinical story of Olaparib, like many of the other PARP-1 clinical candidates, was a roller coaster with periods of both excitement [48] and disappointment [49] Currently, AstraZeneca has committed to its development by sponsoring two Phase 2 studies as a chemosensitizer for prostate cancer (NCT01972217), and gastric cancer (NCT01924533). In 2013, three Phase 3 trials began with Olaparib for the treatment of metastatic breast cancer patients with germline BRCA1/2 mutations (NCT02000622), patients with BRCA mutated ovarian cancer (NCT01844986) and platinum sensitive ovarian cancer patients (NCT01874353).
However, clinical studies with Olaparib and other PARP inhibitors clearly demonstrate that not all patients with BRCA1 or BRCA2 mutations respond to PARP inhibitor therapy [50]. Three mechanisms have been identified as contributing to this type of resistance: (1) restoration of BRCA function [51]; (2) efficient P-glycoprotein efflux of PARP inhibitors (i.e. Olaparib) by increased Mdr1 gene expression; and (3) restoration of HR mechanisms by (e.g.) inactivation of p53 binding protein 1 (53BP1) [52]. Acknowledging that Olaparib is a good substrate for PgP, AstraZeneca identified a backup compound, AZD2461 (30, Fig. 7.10), which displays much of the same characteristics as Olaparib, but is a substantially worse substrate for PgP. Indeed, AZD2461 demonstrated the ability to suppress the development of PARPi resistance in mice with KB1P tumors. Upon treatment of AZD2461 for 100 consecutive days, eight of nine mice engrafted did not develop refractory tumors within 300 days after treatment. Meanwhile, 100 consecutive days of Olaparib treatment resulted in six out of seven mice acquiring drug resistance. In addition, long term treatment of AZD2461 was well tolerated and doubled the median relapse-free survival from 64 to 132 days [52]. Despite being so structurally similar to Olaparib, AZD2461 possesses a profile significantly different enough to warrant a Phase 1 dose escalation study which was completed last year (NCT01247168).
4.5 Lead/Biomarin (BMN-673)
BioMarin Pharmaceuticals is the latest group to enter a PARP-1 inhibitor candidate into the clinic, yet it is perhaps one of the most interesting lead optimization stories. A medicinal chemistry group from Lead Pharmaceuticals serendipitously discovered their lead series of tricyclic phthalazinones (e.g. 35) as a byproduct during the attempted synthesis of pyrrolophthalazinones (31, Fig. 7.11). Because of their late start into the field of PARP-1 inhibitors, this group had the luxury of designing an inhibitor with a core scaffold that had already successfully led to a clinical compound (i.e. Olaparib). For this reason, the Biomarin group designed the pyrrolophthalazinones to mimic both the phthalazinones 17 and benzimidazole carboxamides 19c, two core scaffolds that are incorporated in many of the PARP-1 clinical compounds.
The synthetic route which led to the discovery of tricyclic phthalazinones is outlined in Fig. 7.12. The conversion of the lactone 32 was predicted to form compound 33 under basic conditions. Addition of hydrazine would have led to a precursor of the desired scaffold 31a through three more steps. However, the main product of the first reaction was actually the dihydroquinolinone methyl ester 34. Addition of hydrazine to this intermediate led to compound 35, a very potent PARP-1 inhibitor (IC50 = 6.1 nM) and a novel and unexplored scaffold. Additionally, this scaffold was primed for diversification through sequential addition of aldehydes using a three step synthetic route [53] .
With the ability to quickly generate a wide variety of potent PARP-1 inhibitors with this scaffold, compound 36 (racemate) was identified as a preliminary lead structure (Fig. 7.13) [54]. The in vitro potency of 36 was excellent (IC50 = 2.3 nM) and Cellular EC50 (ability to inhibit intracellular PARylation in peroxide damaged LoVo cells) was 16.9 nM. In addition, the incorporation of the imidazole group improved the aqueous solubility over 35. Compound 36 displayed good oral BA (F% = 59.7) and t1/2 (1.2 h) in rats as well. However, one important aspect in the optimization of this series was the ability to potentiate the cytotoxicity of TMZ and the ability to kill Capan-1 cells, a BRCA2-deficient cell line. For compound 36, the cellular potency in combination with TMZ was only 89 µM and against Capan-1 cells, the IC50 was only 2.0 µM, leaving room for improvement. All of these parameters were improved by the addition of two fluorines and a nitrogen atom (orange circles, Fig. 7.13) to obtain the clinical candidate, compound 37 (BMN-673). BMN-673 displayed improved enzymatic (IC50 = 0.57 nM) and cellular (EC50 = 2.51 nM) potency over its enantiomer (38, BMN-674). In addition, the in vitro metabolism was excellent across species (rat, dog and human) with > 90 % remaining after 2 h of incubation. More importantly, BMN-673 demonstrated the ability to selectively kill tumor cells with BRCA1, BRCA2 and PTEN mutations at low nanomolar concentrations [55]. The oral bioavailability in rats was determined to be > 40 % with a 2.72 h terminal t1/2. In vivo, BMN-673 was able to significantly inhibit the growth of MX-1 xenografts in mice when dosed orally (0.33 and 0.1 mg/kg) for 28 days. At the higher dose, four out of six mice achieved a complete response to this therapy. The chemosensitizing properties of BMN-673 were also apparent as the compound demonstrated significant potentiating effects with both TMZ and platinum drugs [54, 55].
As a late entry into the clinic, it was important for BMN-673 to differentiate itself from some of the earlier generation PARP-1 inhibitors. Interestingly, BMN-673 is ~ 3–8 fold more potent in vitro as a PARP1/2 inhibitor making this compound the most potent PARP inhibitor reported to date. However, this edge in potency does not account for the dramatic differences seen in cytotoxicity. In many BRCA1, BRCA2 and PTEN deficient cell lines, BMN-673 displayed from 20- to 200-fold greater potency over the 1st generation PARP-1 inhibitors Rucaparib , Olaparib and Veliparib [56]. It was recently reported that PARP-1 inhibitors can be cytotoxic not only by inhibition of the enzymatic activity of PARP, but also by ‘PARP trapping’ [10]. Effective inhibitors trap the PARP-DNA complexes , preventing the dissociation of PARP from DNA which is required for DNA repair. This alternative mechanism may account for some of the discrepancies in cellular toxicity amongst the PARP clinical compounds. Trapping studies demonstrated that BMN-673 was more effective at trapping damaged DNA/PARP complex by 40–100 fold in DT40 and DU145 prostate cancer cells damaged with MMS [56].
Clinical studies of BMN-673 indicate that the drug was well tolerated and the Phase 1 study supports a once daily oral dosing of 1 mg/day as the trough concentrations were over 10 nM (IC50 = 0.57 nM). BMN-673 also demonstrated substantial single agent anti-tumor activity in breast and ovarian patients with germline mutations. The RECIST response rate for germline BRCA breast cancer patients was 44 % and the clinical benefit response rate was 82 %. The RECIST response rate for germline BRCA ovarian patients was 39 % while the clinical benefit response rate was 67 % [57]. In October of 2013, BioMarin has initiated a Phase 3 trial with BMN-673 in patients with advanced or metastatic breast cancer with BRCA mutations (NCT01945775) .
4.6 Guilford/MGI/Eisai (E7016 and 2nd Generation E7449)
In 2005, a drug discovery group from Guilford/MGI designed a series of tetracyclic phthalazinones as a combination of two different scaffolds, phthalazinones 17 and tetracyclic isoquinolinones such as GPI 6150 (39, Fig. 7.14) [58, 59]. Combination of these two scaffolds led to an early lead compound GPI 15427 (40, Fig. 7.14, IC50 = 31 nM). Not surprisingly, the planar nature of this series posed solubility issues which were largely overcome by addition of a piperazine moiety (green, Fig. 7.14). Further refinements of the pendant amine led to the clinical candidate GPI 21016 (E7016, 41, Fig. 7.14) . This compound displayed significant chemopotentiating activity in a murine leukemia model in combination with cisplatin . When dosed 15 min pre-, 3 h post- and 6 h post-cisplatin, GPI 21016 (40 mg/kg i.p.) increased the life span by 160 % compared to cisplatin alone. GPI 21016 also demonstrated the ability to act as a chemo- and radiosensitizer in vivo in a glioblastoma xenograft model. GPI 21016 (40 mg/kg p.o.), when given in combination with radiation (4 Gy) and TMZ (3 mg/kg) provided a 32 % increase in life span over the TMZ/radiation control group. This model verifies that GPI 21016 is effective in a system that is relevant to the current clinical standard of care for glioblastoma (i.e. TMZ plus radiation).
Eisai finished a Phase 1 dose escalation trial with E7016 in combination with TMZ. The maximum tolerated dose of E7016 was established as 4 mg/kg p.o. per day for 7 days in combination with 150 mg/kg TMZ p.o. on days 1–5. Pharmacodynamic analysis indicated that E7016 was also able to inhibit PARP in PBMCs at this dose both alone and in combination [60]. In 2012, a Phase 2 trial of E7016 was initiated in combination with TMZ in patients with wild type BRAF Stage IV or unresectable Stage III melanoma (NCT01605162). While results from this trial have not been disclosed, Eisai has initiated another Phase 1 study with a 2nd generation PARP1/2 inhibitor, E7449 (structure not disclosed) [61].
5 Conclusions
After a long and arduous journey, the race to become the first PARP-1 inhibitor to obtain regulatory approval will be decided in the next 2–3 years. Interestingly, despite over 20 drug discovery groups actively designing PARP inhibitors, the current batch of clinical compounds possess a relative homogeneous set of characteristics (e.g. structural similarity, in vitro and in vivo potency, good physicochemical properties). Despite this similarity, there are still some factors that may differentiate the front runners such as PARP trapping ability and susceptibility to efflux transporters. Additionally, some of the first generation inhibitors like Rucaparib , Olaparib and Veliparib have undergone a lengthy clinical journey which has significantly shortened the patent life for each of these compounds. For this reason, groups such as AstraZeneca and Eisai have already progressed 2nd generation compounds into the clinic. While some of the more recent additions to the clinic such as Niraparib and BMN-673 not only have longer patent lives, but seem to have a greater ability to trap PARP-DNA complexes over some of the 1st generation PARP inhibitors. If PARP trapping has a profound effect on the cytotoxicity of PARP inhibitors in the clinic, then compounds like BMN-673 may have a slight advantage in efficacy/safety profile. To date, PARP inhibitors have not clearly demonstrated their usefulness in the clinic in combination therapy with chemotherapeutic agents. A more promising route to approval seems to be as single agents in patients with tumors deficient in certain DNA repair pathways. Indeed, every one of the Phase 3 trials are focused on patients with these deficiencies, a positive sign that PARP inhibitors may soon find their niche in the world of cancer treatment.
References
Yelamos J, Schreiber V, Dantzer F (2008) Toward specific functions of poly(ADP-ribose) polymerase-2. Trends Mol Med 14:169
Langelier MF, Planck JL, Roy S, Pascal JM (2012) Structural basis for DNA damage-dependent poly(ADP-ribosyl)ation by human PARP-1. Science 336:728
Durkacz BW, Omidiji O, Gray DA, Shall S (1980) (ADP-ribose)n participates in DNA excision repair. Nature 283:593
De Vos M, Schreiber V, Dantzer F (2012) The diverse roles and clinical relevance of PARPs in DNA damage repair: current state of the art. Biochem Pharmacol 84:137
Ogata N, Ueda K, Kagamiyama H, Hayaishi O (1980,) ADP-ribosylation of histone H1. Identification of glutamic acid residues 2, 14, and the COOH-terminal lysine residue as modification sites. J Biol Chem 255:7616
Caldecott KW (2003) XRCC1 and DNA strand break repair. DNA Repair 2:955
Audebert M, Salles B, Calsou P (2004) Involvement of poly(ADP-ribose) polymerase-1 and XRCC1/DNA ligase III in an alternative route for DNA double-strand breaks rejoining. J Biol Chem 279:55117
Schreiber V, Dantzer F, Ame JC, de Murcia G (2006) Poly(ADP-ribose): novel functions for an old molecule. Nat Rev Mol Cell Biol 7:517
La Thangue NB, Kerr DJ (2011) Predictive biomarkers: a paradigm shift towards personalized cancer medicine. Nat Rev Clin Oncol 8:587
Murai J et al (2012) Trapping of PARP1 and PARP2 by clinical PARP inhibitors. Cancer Res 72:5588
Clark JB, Ferris GM, Pinder S (1971) Inhibition of nuclear NAD nucleosidase and poly ADP-ribose polymerase activity from rat liver by nicotinamide and 5’-methyl nicotinamide. Biochim Biophys Acta 238:82
Purnell MR, Whish WJ (1980) Novel inhibitors of poly(ADP-ribose) synthetase. Biochem J 185:775
Banasik M, Komura H, Shimoyama M, Ueda K (1992) Specific inhibitors of poly(ADP-ribose) synthetase and mono(ADP-ribosyl)transferase. J Biol Chem 267:1569
Levi V, Jacobson EL, Jacobson MK (1978) Inhibition of poly(ADP-ribose) polymerase by methylated xanthines and cytokinins. FEBS Lett 88:144
Shall S (1975) Proceedings: experimental manipulation of the specific activity of poly(ADP-ribose) polymerase. J Biochem 77:2p
Suto MJ, Turner WR, Arundel-Suto CM, Werbel LM, Sebolt-Leopold JS (1991) Dihydroisoquinolinones: the design and synthesis of a new series of potent inhibitors of poly(ADP-ribose) polymerase. Anti-Cancer Drug Des 6:107
Ferraris D et al (2003) Design and synthesis of poly ADP-ribose polymerase-1 inhibitors. 2. Biological evaluation of aza-5[H]-phenanthridin-6-ones as potent, aqueous-soluble compounds for the treatment of ischemic injuries. J Med Chem 46:3138
Griffin RJ et al (1998) Resistance-modifying agents. 5. Synthesis and biological properties of quinazolinone inhibitors of the DNA repair enzyme poly(ADP-ribose) polymerase (PARP). J Med Chem 41:5247
Ferraris DV (2010) Evolution of poly(ADP-ribose) polymerase-1 (PARP-1) inhibitors. From concept to clinic. J Med Chem 53:4561
Griffin RJ et al (1995) Novel potent inhibitors of the DNA repair enzyme poly(ADP-ribose)polymerase (PARP). Anti-Cancer Drug Des 10:507
Ruf A, de Murcia G, Schulz GE (1998) Inhibitor and NAD+ binding to poly(ADP-ribose) polymerase as derived from crystal structures and homology modeling. Biochemistry 37:3893
Skalitzky DJ et al (2003) Tricyclic benzimidazoles as potent poly(ADP-ribose) polymerase-1 inhibitors. J Med Chem 46:210
Tao M et al (2006) Synthesis and structure-activity relationships of novel poly(ADP-ribose) polymerase-1 inhibitors. Bioorg Med Chem Lett 16:938
Zhang WT et al (2009) Design, synthesis, and cytoprotective effect of 2-aminothiazole analogues as potent poly(ADP-ribose) polymerase-1 inhibitors. J Med Chem 52:718
Gangloff AR et al (2013) Discovery of novel benzo[b][1,4]oxazin-3(4H)-ones as poly(ADP-ribose)polymerase inhibitors. Bioorg Med Chem Lett 23:4501
Ferrigno F et al (2010) Development of substituted 6-[4-fluoro-3-(piperazin-1-ylcarbonyl)benzyl]-4,5-dimethylpyridazin-3(2H)-ones as potent poly(ADP-ribose) polymerase-1 (PARP-1) inhibitors active in BRCA deficient cells. Bioorg Med Chem Lett 20:1100
Wahlberg E et al (2012) Family-wide chemical profiling and structural analysis of PARP and tankyrase inhibitors. Nat Biotechnol 30:283
Bregman H et al (2013) Discovery of novel, induced-pocket binding oxazolidinones as potent, selective, and orally bioavailable tankyrase inhibitors. J Med Chem 56:4320
Shultz MD et al (2013) Structure-efficiency relationship of [1,2,4]triazol-3-ylamines as novel nicotinamide isosteres that inhibit tankyrases. J Med Chem 56:7049
White AW et al (2000) Resistance-modifying agents. 9. Synthesis and biological properties of benzimidazole inhibitors of the DNA repair enzyme poly(ADP-ribose) polymerase. J Med Chem 43:4084
Canan Koch SS et al (2002) Novel tricyclic poly(ADP-ribose) polymerase-1 inhibitors with potent anticancer chemopotentiating activity: design, synthesis, and X-ray cocrystal structure. J Med Chem 45:4961
White AW et al (2004) Potentiation of cytotoxic drug activity in human tumour cell lines, by amine-substituted 2-arylbenzimidazole-4-carboxamide PARP-1 inhibitors. Bioorg Med Chem Lett 14:2433
Tikhe JG et al (2004) Design, synthesis, and evaluation of 3,4-dihydro-2H-[1,4]diazepino[6,7,1-hi]indol-1-ones as inhibitors of poly(ADP-ribose) polymerase. J Med Chem 47:5467
Thomas HD et al (2007) Preclinical selection of a novel poly(ADP-ribose) polymerase inhibitor for clinical trial. Mol Cancer Ther 6:945
Chuang HC, Kapuriya N, Kulp SK, Chen CS, Shapiro CL (2012) Differential anti-proliferative activities of poly(ADP-ribose) polymerase (PARP) inhibitors in triple-negative breast cancer cells. Breast Cancer Res Treat 134:649
Plummer R et al (2008) Phase I study of the poly(ADP-ribose) polymerase inhibitor, AG014699, in combination with temozolomide in patients with advanced solid tumors. Clin Cancer Res 14:7917
Sleijfer S, Bogaerts J, Siu LL (2013) Designing transformative clinical trials in the cancer genome era. J Clin Oncol 31:1834
Penning TD et al (2008) Discovery and SAR of 2-(1-propylpiperidin-4-yl)-1H-benzimidazole-4-carboxamide: a potent inhibitor of poly(ADP-ribose) polymerase (PARP) for the treatment of cancer. Bioorg Med Chem 16:6965
Penning TD et al (2009) Discovery of the Poly(ADP-ribose) polymerase (PARP) inhibitor 2-[(R)-2-methylpyrrolidin-2-yl]-1H-benzimidazole-4-carboxamide (ABT-888) for the treatment of cancer. J Med Chem 52:514
Donawho CK et al (2007) ABT-888, an orally active poly(ADP-ribose) polymerase inhibitor that potentiates DNA-damaging agents in preclinical tumor models. Clin Cancer Res 13:2728
Rugo HS et al (2013) In 36th Annual San Antonio Breast Cancer Symposium. (San Antonio, TX), vol. Abstract S5–02
Jones P et al (2009) Discovery of 2-{4-[(3S)-piperidin-3-yl]phenyl}-2H-indazole-7-carboxamide (MK-4827): a novel oral poly(ADP-ribose)polymerase (PARP) inhibitor efficacious in BRCA-1 and -2 mutant tumors. J Med Chem 52:7170
Sandhu SK et al (2013) The poly(ADP-ribose) polymerase inhibitor niraparib (MK4827) in BRCA mutation carriers and patients with sporadic cancer: a phase 1 dose-escalation trial. Lancet Oncol 14:882
Michie CO et al (2013) Final results of the phase I trial of niraparib (MK4827), a poly(ADP)ribose polymerase (PARP) inhibitor incorporating proof of concept biomarker studies and expansion cohorts involving BRCA1/2 mutation carriers, sporadic ovarian, and castration resistant prostate cancer (CRPC). J Clin Oncol 31:2513 (Meeting Abstracts)
Menear KA et al (2008) 4-[3-(4-cyclopropanecarbonylpiperazine-1-carbonyl)-4-fluorobenzyl]-2H-phthalazin- 1-one: a novel bioavailable inhibitor of poly(ADP-ribose) polymerase-1. J Med Chem 51:6581
Cockcroft XL et al (2006) Phthalazinones 2: optimisation and synthesis of novel potent inhibitors of poly(ADP-ribose)polymerase. Bioorg Med Chem Lett 16:1040
Loh VM, Jr. et al (2005) Phthalazinones. Part 1: the design and synthesis of a novel series of potent inhibitors of poly(ADP-ribose)polymerase. Bioorganic Med Chem Lett 15:2235
Fong PC et al (2009) Inhibition of poly(ADP-ribose) polymerase in tumors from BRCA mutation carriers. New Engl J Med 361:123
Khan OA et al (2011) A phase I study of the safety and tolerability of olaparib (AZD2281, KU0059436) and dacarbazine in patients with advanced solid tumours. Br J Cancer 104:750
Audeh MW et al (2010) Oral poly(ADP-ribose) polymerase inhibitor olaparib in patients with BRCA1 or BRCA2 mutations and recurrent ovarian cancer: a proof-of-concept trial. Lancet 376:245
Edwards SL et al (2008) Resistance to therapy caused by intragenic deletion in BRCA2. Nature 451:1111
Jaspers JE et al (2013) Loss of 53BP1 causes PARP inhibitor resistance in Brca1-mutated mouse mammary tumors. Cancer Discov 3:68
Wang BSJ, (CA, US), Chu, Daniel (Santa Clara, CA, US), Liu, Yongbo (Shanghai, CN), Jiang, Quan (Shanghai, CN), Lu, Lei (Zhejiang Province, CN). (United States, 2011). The international patent publication number is WO 2011/097602 A1
Wang B et al (2011) In Proceedings of the AACR-NCI-EORTC International Conference, San Francisco, CA. (Mol Cancer Ther), vol 10 (11 Suppl) Abstract B64
Shen Y et al (2013) BMN 673, a novel and highly potent PARP1/2 inhibitor for the treatment of human cancers with DNA repair deficiency. Clin Cancer Res 19:5003
Murai J et al (2014) Stereospecific PARP Trapping by BMN 673 and comparison with olaparib and rucaparib. Mol Cancer Ther 13(2):433–443
De Bono JS et al (2013) ASCO 31(suppl abstr 2580) (J Clin Oncol, 2013)
Zhang J (1999) PARP inhibition: a novel approach to treat ischemia/reperfusion and inflammation-related injuries. Exp Opin Emerg Drugs 4:209
Zhang J et al (2000) GPI 6150 prevents H(2)O(2) cytotoxicity by inhibiting poly(ADP-ribose) polymerase. Biochem Biophys Res Commun 278:590
Mita AC et al (2011) In AACR-NCI-EORTC International Conference: Molecular Targets and Cancer Therapeutics. (Mol Cancer Ther, San Francisco, CA), vol 10 (11 Suppl) Abstract B185
McGonigle S et al (2012) In Proceedings of the 103rd Annual Meeting of the American Association for Cancer Research. (Cancer Res, Chicago, IL), vol 72 (8 Suppl) Abstract 4688
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Ferraris, D. (2015). Overview of PARP Inhibitor Design and Optimization. In: Curtin, N., Sharma, R. (eds) PARP Inhibitors for Cancer Therapy. Cancer Drug Discovery and Development, vol 83. Humana Press, Cham. https://doi.org/10.1007/978-3-319-14151-0_7
Download citation
DOI: https://doi.org/10.1007/978-3-319-14151-0_7
Published:
Publisher Name: Humana Press, Cham
Print ISBN: 978-3-319-14150-3
Online ISBN: 978-3-319-14151-0
eBook Packages: MedicineMedicine (R0)