Abstract
Cancer treatments such as radiotherapy and chemotherapy induce DNA damage, which can be a factor determining therapeutic efficacy. A DNA double strand break (DSB) is considered to be the most critical type of DNA lesion, since DSBs cause cell death when they are unrepaired and generate mutations if they are misrepaired. Ionising radiation (IR) produces a broad spectrum of DNA damage, including DSBs, single strand breaks (SSBs) and base damages. Specific poly(ADP-ribose)polymerase (PARP) inhibitors, currently being tested in clinical trials, compromise SSB repair after IR, resulting in the accumulation of replication-associated DSBs. Since replication-associated DSBs are effectively repaired by homologous recombination, PARP inhibition sensitizes cells that are defective in homologous recombination. In addition, PARP inhibition effectively blocks backup DSB repair in cells defective in non-homologous end joining (NHEJ) following IR. Importantly, the sensitization in NHEJ-defective cells occurs independently of DNA replication. In this chapter, we discuss the multiple effects of PARP inhibition in DSB repair-defective cells in the context of the potential availability of PARP inhibitor in clinical use. We further discuss how a PARP inhibitor influences the type of cell death, which may affect prognosis following cancer treatment. In cancer therapy using PARP inhibitors, a comprehensive understanding of PARP signaling from DNA damage to cell death may be required to augment DNA damage-induced cell death and to direct restrained cell death in order to reduce inflammation responses in surrounding tissues.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Involvement of PARP in DNA Double Strand Break Repair
1.1 The Role of PARP in DNA Damage Responses
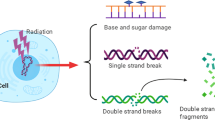
Ionizing radiation (IR) causes several types of damage to DNA strands including DNA double strand breaks (DSBs), single strand breaks (SSBs), base damage, and DNA cross-links. DSBs are a type of critical lesion that determine cellular fate, because they may cause severe genomic instability through such means as a deletion or chromosomal translocation leading to carcinogenesis . In addition, cells with unrepaired DSBs undergo cell death if excessive amounts of DNA damage are generated or cell cycle checkpoint arrest is not fully functional. DNA damage-dependent PARP (PARP-1, PARP-2 and PARP-3) are activated following binding to SSBs, gaps or DSBs [1–5]. Activated PARP-1 and PARP-2 at SSBs or gaps recruits SSB repair proteins, e.g. XRCC1, DNA polymerase β, ATPX, and LIG3, via polyADP-ribosylation on the target proteins, including PARP itself and the histones [6–8]. Since SSBs are also formed during base excision repair (BER), PARP-1 and PARP-2 also participate in this repair process (see also other chapters) (Fig. 15.1) [9, 10]. These repair pathways are compromised by selective PARP inhibitors against PARP-1 and -2 activities, which are currently being examined in a clinical trial. Although endogenous levels of SSBs and base damage (or even IR-induced SSBs) do not largely influence cell viability, these types of damage become toxic lesions if unrepaired before DNA replication , i.e. unrepaired SSBs and base damage are converted to DSBs when a replication fork encounters this damage in the S phase (Fig. 15.2) [11, 12].
Role of PARP in base excision repair (BER) and SSB repair after IR. IR induces base damage and SSBs as well as DSBs. Generated base damage is removed by DNA glycosylases. Following the removal of damaged base, apyrimidinic/apurinic (AP) endonuclease creates a nick with 3’- hydroxyl and 5’-deoxyribose termini. The nick is recognized and repaired by SSB repair proteins such as PARP-1, PARP-2 and XRCC1
Induction of replication-associated DSBs by PARP inhibition. IR causes SSBs directly or via the BER pathway (Fig. 15.1). In the presence of PARP inhibitor, SSB repair is delayed. If the SSBs are not repaired prior to the entry of S phase, replication-associated DSBs are generated when replication forks encounter SSBs. Since PARP keeps binding SSBs until SSB repair is completed, DNA ends at replication-associated DSBs may be masked by PARP proteins in the presence of PARP inhibitor, which may block the access of other DSB repair proteins. To allow access, the PARP-DNA ends may be removed by Artemis nuclease activity [15.1]. Since the DSB end at the replication fork does not have a partner, these breaks are preferentially repaired by HRR
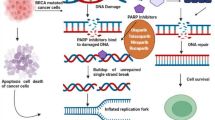
The living cell generates oxidative species during cellular metabolism. It has therefore been suggested that more than 10,000 SSBs per cell arise from oxidative stress per day. In normal cells and even tumor cells, this damage is repaired without affecting cell growth. However, if SSB repair or BER is compromised, unrepaired SSB will be converted into toxic DSBs at replication forks. Replication-associated DSB have a unique damage structure compared to DSBs that are directly induced by IR: replication-associated DSBs have one end, while IR-induced DSBs have two ends (Fig. 15.2) [13–15]. Since a replication-associated DSB does not represent a ‘break’, the DNA structure is also known as a DNA double strand end (DSE) [15]. At two-ended DSBs directly induced by IR, Ku protein, which is a central non-homologous end joining (NHEJ) factor, binds to both DSB ends, tethers and promotes the rejoining of the DSB ends. In contrast, in replication-associated DSB, the DSB is not effectively rejoined by NHEJ because Ku cannot bind the other side of the break end (Fig. 15.2). Therefore, replication-associated DSBs are preferentially repaired by homologous recombination repair (HRR), which is another DSB repair pathway. Consistent with the model in DSB repair pathway choice at replication associated DSBs, an increasing number of reports demonstrate that HRR defective cells show hypersensitivity to PARP inhibitor [16]. Among tumor cells defective in HRR, BRCA1 or BRCA2 defective tumor cells exhibit an exceptional increase in sensitivity to PARP inhibitor [11, 12]. The exceptional effect can be also explained by the finding that PARP-1 is involved in DNA repair, which requires BRCA2 , at stalled replication forks [17, 18]. Thus, it is likely that the strong synthetic lethal interaction between PARP and BRCA1/2 is regulated by multiple unrevealed mechanisms (for detailed descriptions, see other chapters in this volume).
1.2 Replication-Dependent Radiosensitization by PARP Inhibition in Cells Defective in HRR, Including Artemis and ATM
Inhibition of DNA damage-dependent PARP activities by PARP inhibitor enhances the sensitivity of radiotherapy in tumor cells [19, 20]. IR induces ~ 1000 SSBs per Gy. The SSBs, however, are rapidly repaired without cellular toxicity if SSB repair is intact [21]. Numerous reports demonstrate that the radiosensitizing effect of PARP inhibition requires DNA replication, because the enhanced sensitivity is attenuated by the blocking of DNA replication polymerases [11, 12, 22]. The mechanism of radiosensitization is therefore explained by the enhanced conversion of unrepaired SSBs to DSBs during the S phase. Thus, the radiosensitization by PARP inhibition is explained by the formation of toxic DSBs at the replication fork.
Generally PARP inhibitors cause a modest increase in radiosensitivity in tumors [19, 23]. Interestingly, however, some radioresistant tumor cells exhibit hypersensitivity to IR in combination with PARP inhibitor [23]. The difference is explained by the rate of cell growth, i.e. PARP inhibition preferentially sensitizes tumor cells by virtue of their elevated replication rates. This aspect is well investigated in actively dividing glioma cells [23]. Consistent with the above notion, such actively dividing cell lines show an apparent increase in radiosensitivity, even after low doses of IR [24]. Hence, a PARP inhibitor can be a potent radiation sensitizer in highly replicating tumor cells that do not contain mutations in the HRR genes. As a similar mechanism, PARP inhibition exacerbates the cytotoxicity of the alkylating agents in highly replicating cells, since SSBs are also generated during BER following treatment with alkylating agents as described above [25].
Interestingly, there is a report that PARP inhibitors augment radiosensitivity in Artemis or ATM defective cells [25]. The data shows that, in the presence of PARP inhibitor IR- or alkylating agent- induced replication-associated DSBs are not effectively repaired in Artemis or ATM-defective cells. Artemis and ATM are involved in a subset of DSBs, which potentially require DNA end processing in NHEJ and HRR [26]. Whilst core NHEJ components are essential for the overall NHEJ process, Artemis and ATM are required for the repair of IR-induced DSBs, which arise in the heterochromatin (HC) region [27, 28]. ATM facilitates chromatin remodeling at the HC region via phosphorylation of KAP-1 (KRAB-associated protein-1), which relieves the compact chromatin structure to allow the access of DSB repair proteins at the HC region [28, 29]. The involvement of Artemis in HC-DSB repair is apparent under epistatic analysis, however, the precise role of Artemis is yet to be revealed. Nevertheless, the current model proposes that, because of its nuclease activity, Artemis is involved in the processing of a “dirty” DSB end, e.g. a DSB with base damages [26]. Notably, the enhanced radiosensitivity in Artemis and ATM cells with PARP inhibitors is attenuated by the blocking of DNA replication, suggesting that a subset of toxic DSBs, which arise in DNA replication, require Artemis and ATM [25]. In the presence of a PARP inhibitor, PARP is kept at the DNA ends, preventing the access of other repair proteins required for further repair steps [30]. Thus, Artemis may function in removing the PARP-DSB complexes recognized as “dirty” ends, which potentially become toxic lesions at DNA replication. As an alternative possibility, the repair of DSB arising at the HC region may require Artemis and ATM. Artemis and ATM may be also involved in removing “dirty” ends prior to the initiation of HRR at the replication forks. Hence, IR-induced SSBs can become toxic DNA lesions following the formation of PARP-DSB complex in the presence of PARP inhibitor if Artemis/ATM is downregulated. This also implies that the nature of IR-induced SSBs could be different from endogenous SSBs, since PARP inhibition alone does not influence cell viability in Artemis/ATM defective cells [25, 27].
1.3 Replication Independent Effects by PARP Inhibition in NHEJ Defective Cells: Ionizing Radiation
PARP inhibitors also enhance sensitization to IR in cells that are defective in the core components of NHEJ, irrespective of DNA replication [25]. Classical NHEJ (C-NHEJ) is comprised of the core components Ku, DNA-PKcs, DNA Ligase IV, XRCC4, and XLF, which are responsible for the repair of most radiation-induced DSBs [31, 32]. Besides the C-NHEJ pathway, an alternative NHEJ (A-NHEJ) pathway (also known as the backup NHEJ), has been identified in the absence of C-NHEJ factors [33]. In the A-NHEJ pathway, DSB ends undergo short range of end resection, leading to deletion mutations (Fig. 15.3) [34]. In normal cells, Ku can tether and stabilize two DSB ends, however, without the protection by Ku, the DNA ends are resected by the nuclease activity of MRE11 and CtIP [35]. The precise mechanism for this resection remains to be elucidated. To rejoin DSB ends without Ku, two DSBs frequently utilize microhomology sequences to tether and anneal both ends. An SSB or gap is subsequently sealed by SSB repair proteins, including PARP, XRCC1 and DNA Ligase III. Since PARP inhibition prevents the SSB repair pathway, the A-NHEJ pathway is effectively compromised by PARP inhibitor [25, 33].
PARP inhibitor compromises A-NHEJ in the lack of Ku protein. In wild type cells, Ku binds both DSB ends. Subsequently, Ku tethers both ends and recruit core NHEJ factors. The NHEJ pathway in wild type cells are called classical NHEJ (C-NHEJ). Conversely, in the absence of Ku, DSB ends are resected due to the lack of DSB end protection by Ku. Following the resection, when DSB ends find microhomology sequences, they are annealed and rejoined by SSB repair factors. This pathway is called alternative NHEJ (A-NHEJ). Thus, PARP inhibition effectively compromises A-NHEJ pathway
The A-NHEJ pathway is active throughout the cell cycle phases. The usage is enhanced due to increased resection activity in the S/G2 phase [37]. Nevertheless, it is noteworthy that the activity of A-NHEJ is largely regulated by Ku status, rather than by NHEJ status. This idea is supported by the data showing that A-NHEJ pathway is not able to compensate C-NHEJ in cells defective in DNA-PKcs or XRCC4, because Ku can bind to DSB ends in these cells [38]. In G1 cells, most DSBs are rejoined by NHEJ since HRR cannot function due to a lack of sister chromatid and downregulated HRR proteins. Thus, it has been proposed that the A-NHEJ pathway is the exclusive repair pathway in the absence of C-NHEJ. Consistent with this notion, PARP inhibition perfectly blocks all DSB repair in G0/G1-arrested Ku80 null cells [25]. Although null mutations in NHEJ genes are rarely reported in humans because NHEJ is the essential repair mechanism for cell survival, NHEJ is downregulated in some tumors with high-grade malignancies such as carcinoma of the bladder and glioblastoma multiforme [39–41]. Indeed, it has been reported that bladder tumor extracts fail to conduct accurate NHEJ and instead use an error-prone mechanism, which is likely regulated by A-NHEJ [42]. Furthermore, it has been shown that the expression of NHEJ proteins is downregulated following treatment with a histone deacetylase inhibitor, which is in clinical use [43]. Hence, a PARP inhibitor can be a potent radiosensitizer, irrespective of replication status. Moreover, PARP inhibition may be utilized to exterminate tumors because it has been suggested that cancer stem cells exist in a quiescent state as G0/G1 if the repair pathway can be manipulated from C-NHEJ to A-NHEJ.
PARP-3 is effectively activated by DSB ends, but not SSBs [44]. Similar to PARP-1 and -2, PARP-3 ADP-ribosylates PARP-3 itself and the histones [44]. The ADP-ribosylated histones recruit APLF, which interacts with Ku. Thus, the inhibition of PARP-3 attenuates NHEJ activity, resulting in the delay of DSB repair. However, the inhibition of PARP-3 or loss of APLF does not reduce cell viability following irradiation, because DSBs are finally repaired in PARP-3/APLF-defective cells, despite the delay of repair. It is therefore highly likely that the enhanced sensitivity in radiation or chemotherapeutic drugs with a PARP inhibitor is dependent on the inhibition of PARP-1 and -2 in the SSB repair process. In summary, the radiosensitizing effects of PARP inhibitors are manifested in replicating cells, and are augmented under defects in HRR. In addition, radiosensitivity in non-replicating cells is enhanced if C-NHEJ is downregulated.
1.4 Replication Independent Effects of PARP Inhibition in NHEJ Defective Cells: Alkylating Agents
Alkylating agents are favorably utilized in radioresistant tumors, for instance, temozolomide is usually given to treat glioblastoma, a type of brain tumor. Alkylating agents form adducts at N- and O-atoms of DNA. N 7-methylation, which is known to be a major DNA adduct, is removed by BER. PARP inhibition therefore causes accumulation of SSBs following the administration of alkylating agents, which also results in the formation of replication-associated DSBs. A combination of an alkylating agent and PARP inhibitor enhances the sensitivity particularly in cells defective in HRR as described in the above model. Interestingly, PARP inhibition exhibits gross sensitization of Ligase IV deficient cells to methylmethane sulfonate (MMS: an alkylating agent) [25]. It is important to note, however, that the enhanced sensitivity is replication-independent. PARP inhibition promotes the replication-independent DSB formation from MMS-induced SSBs in close proximity, i.e. DSBs arising from overlapping SSBs or from interactions between transcription and obstructed SSBs (Fig. 15.4) [45]. For instance, ~ 20 DSBs arise by PARP inhibition if NHEJ is deficient following 1 mM MMS treatment, although these DSBs are rapidly repaired in NHEJ-proficient normal cells [25]. Thus, the combined usage of an alkylating agent and a PARP inhibitor may be a useful chemotherapy in non-cycling radioresistant tumors.
Replication-independent DSB formation by PARP inhibition following treatment with alkylating agent. A high dose of alkylating agent causes multiple alkylations on DNA strands. As described in Fig. 15.1, an SSB is formed during BER. If two SSBs are ambilaterally generated in close proximity, e.g. within < 30 base pairs, duplex DNA is denatured and a DSB is formed. The DSB is immediately repaired by NHEJ, since any DSBs are not observed in wild type cells even in the presence of PARP inhibitor. However, the DSB formation becomes evident in NHEJ defective cells
Hence, studies reveal the molecular mechanisms of sensitization by PARP inhibition following DNA damage induction. (1) Inhibition of PARP exhibits dramatic synthetic lethality in BRCA1 and BRCA2 defective tumor cells. Increased lethality is also observed in cells defective in other HRR components, including accessory components, (2) Artemis and ATM, albeit to a lesser extent. The enhanced sensitization is highly dependent on DNA replication. (3) Furthermore, PARP inhibitor radiosensitizes tumor cells defective in NHEJ by blocking the backup NHEJ pathway, independently of DNA replication. Radiotherapy is able to target tumor cells, however, DNA damage in surrounding normal tissues is a cause for concern. It would therefore be important to reduce radiation doses with PARP inhibition by referring to a patient database with consideration of DSB repair capability.
The accumulation of DSBs in repair-defective cells switches the cellular fate from survival to cell death. It would therefore be important to consider the effect of PARP inhibition on downstream signaling that results in cell death. In the next section, we discuss the involvement of PARP in DNA damage-induced cell death and suggest a potential plan for manipulating the course of cell death to direct a restrained cell death, which may reduce inflammation responses in surrounding normal tissues.
2 PARP-Dependent Cell Death
2.1 PARP Overactivation-Induced Cell Death Triggered by DNA Damage
DNA damage-dependent PARPs are activated in order to maintain genomic stability following ionizing radiation. Cells irradiated with lethal levels of ionizing radiation undergo various types of cell death including apoptosis, necrosis , and mitotic catastrophe. Among such types of cell death induced by DNA damage, there are two types of PARP-dependent cell death involving PARP overactivation following lethal levels of DNA damage (Fig. 15.5). In these processes, the involvement of PARP-1 in PARP-dependent cell death is evident. The first is PARP-dependent cell death from depletion of intracellular NAD+/ATP level by PARP overactivation. Since poly(ADP-ribosyl)ation requires NAD+ for the reaction, DNA damage-induced overactivation of PARP decreases cellular NAD+ levels [46, 47]. Cellular NAD+ is used as a coenzyme for ATP synthesis by biochemical reactions such as glycolysis, the TCA cycle, and the electron transport chain. Massive PARP overactivation therefore induces ATP depletion via NAD+ overconsumption [48]. The intracellular ATP level is closely involved with the types of cell death. The execution of the apoptotic cell death mechanism, which is the most common type of programmed cell death, requires ATP. On the other hand, excessive DNA damage, followed by a breakdown of intracellular ATP level leads the cells to necrotic cell death. In the past, PARP overactivation-induced cell death following the DNA damage response had been therefore regarded as necrosis.
PARP-Dependent Cell Death induced by DNA Damage. a Typical DNA strand breakers induce apoptosis, which is dependent on caspase activation. b Massive DNA damage caused by DNA alkylators and high dose of DNA strand breakers triggers necrosis-like programmed cell death that is PARP-dependent cell death
However some groups have recently reported evidence that PARP overactivation- cell death is another type of programmed cell death, necroptosis. In necroptosis, certain molecular mechanisms actively regulate the execution [49, 50]. Necroptosis is dictated by the RIP1/RIP3 kinase complex and is involved in the regulation of physiological responses such as the response of the immune system. Although it has been reported that PARP-1 is involved in the RIP1/RIP3 signaling pathway, the mechanism seems to be different according to the types of the cell death inducers [50–52]. In an alkylating agent-induced necroptosis cascade, PARP-1 activation precedes RIP1/RIP3 activation, because RIP1 downregulation attenuates PARP-1-mediated cell death without inhibition of PARP-1 activation [50]. However, the contribution of PARP-1 to RIP1/RIP3 activation in death receptor-induced necroptosis remains controversial [51, 52]. The relationship between necrosis and necropstosis in PARP overactivation-induced cell death remains unclear, because of the indecisive discrimination between necrosis and necroptosis, and the definition of necroptosis itself, appears to confuse many researchers. Nevertheless, it is noteworthy that that certain types of PARP-dependent cell death are regulated based on molecular mechanisms.
2.2 Poly(ADP-Ribose) as a Death Signal Molecule
The second type of PARP-dependent cell death is caused by a direct death signal from the poly(ADP-ribose) polymer itself or from the poly(ADP-ribosyl)ation of other targets. Since the PARG family members possess both endoglycosidic and exoglycosidic activity, PARG reactions likely control the length of poly(ADP-ribose) chain on the poly(ADP-ribosyl)ated proteins, and concurrently produce various sizes of acceptor-free poly(ADP-ribose), including monomer, oligomer, and polymer [53–55]. It has been shown that acceptor-free poly(ADP-ribose) can function directly as a death signal [56, 57]. The execution of this type of cell death requires the binding of poly(ADP-ribose) to AIF, followed by the translocation of AIF from the mitochondrial membrane into the nucleus. The nuclear form of AIF triggers the activation of DNA degrading complex, including CypA [58]. These findings suggest that this type of PARP-dependent cell death could be a type of necroptosis. It has also been shown that poly(ADP-ribose) polymer itself functions as a death signal, although it remains unclear whether poly(ADP-ribosyl)ation of AIF can mediate similar signal transduction.
3 Involvement of PARP in the Cell Death “Execution” Process
3.1 PARP Cleavage by Cell Death-Involved Proteases
PARP-1 and PARP-2 are death substrates that are cleaved during apoptosis [59, 60]. PARP-1 cleavage has been well-characterized (Fig. 15.6). During apoptotic cell death, full-length PARP-1 (theoretical molecular weight, MW: 113 kDa) is cleaved mainly by the apoptotic proteases, caspase-3 and -7, into p89 (MW: 89 kDa) and p24 (MW: 24 kDa) fragments [61–63]. The main recognition sequence in PARP-1, 210DEVD213, is cleaved after the 213D residue by caspase-3 and caspase-7 [64]. p89 fragment inhibits homodimerization of intact PARP-1 molecules in a dominant negative manner, resulting in a decrease of cellular PARP-1 activity [65]. The tight binding of p24 fragment to DNA breaks leads to the inhibition of further activation of uncleaved PARP-1 [66]. In any case, cleaved PARP-1 can function as a strong inhibitor of PARP on DNA strands, because the p24 fragment can bind DNA breaks and act as a dominant negative inhibitor of PARP-dependent repair. The blockade of PARP-1 activity by cleaved fragments inhibits DNA repair pathways, and also diminishes secondary PARP-1 activation in response to DNA fragmentation caused during apoptosis by cell death-related nucleases such as CAD. Furthermore, the study of a cleavage-resistant mutant of PARP-1 has revealed that inhibition of PARP-1 cleavage causes resistance to DNA damage-induced cell death [55, 67]. Importantly, the presence of cleavage-resistant PARP-1 causes the converse type of cell death; apoptosis-type programmed cell death becomes necrosis or necrosis-like programmed cell death [36, 68, 69]. Taken together with these findings, the physiological significance of PARP-1 cleavage during apoptosis appears to be secure completion of classical apoptosis, which is considered “restrained cell death” in terms of the elimination of dead cells. While the physiological functions of PARP-1 fragments from apoptotic cleavage have been well studied, it has also been reported that PARP-1 is cleaved by cathepsins and calpains during non-apoptotic cell death [70–73]. However the exact means by which PARP-1 cleavage occurs during non-apoptotic cell death are currently under investigation.
3.2 Use of PARP Inhibitors for “Restrained Cell Death”
Increasing evidence reveals the role of PARP in DNA repair and cell death execution. To understand the exact effects of PARP inhibitors on cells or individuals, the “change of death forms” that occur following PARP inhibition should be considered. The first is a change in the type of programmed cell death following a switch of the dominant repair pathway caused by PARP inhibition. Programmed cell death does not always occur at a specific cell cycle phase [74, 75]. Furthermore, considering the existence of mitotic catastrophe, which is another pathway of programmed cell death that occurs at mitosis , it is possible that distinct types of cell death exist at each interphase [75, 76]. The second is a change in the type of programmed cell death following an alteration of the dominant execution process of cell death. When DNA damage triggers cell death, PARP inhibition leads to the enhancement of both the lethality caused by the inhibition of PARP-mediated DNA repair and apoptosis via the cell death-specific inhibition of PARP activity. On the other hand, when cell death induction is DNA damage-independent, PARP inhibition increases apoptosis while decreasing necrosis or necrosis-like programmed cell death. Considering radiation therapy for cancer, the type of cell death following the DNA damage response can be variable if DNA damage responsive genes are upregulated or downregulated in tumors and it can show tissue specificity. Since PARPs have multiple functions in the DNA damage response and cell death processes, the eventual outcome in irradiated cells cannot always be predicted. Nevertheless if both steps are comprehensively controlled under radiotherapy with PARP inhibitor treatment, the cancer prognosis will be significantly improved by switching cell death pathway to the restrained cell death of cancer cells, which does not induce massive inflammation in surrounding normal cells as occurs during necrosis.
4 Summary
In this chapter, we discussed about the role of PARPs in DNA repair and cell death following cancer treatment. Since PARPs have multiple functions in SSB, BER and DSB repair, the comprehensive understanding of the role of PARPs in DNA repair should be required for proposing the idea to augment therapeutic efficacy in chemo- and radiotherapy with PARP inhibition. Further, we propose that it is also important to consider the type of cell death in cancer cells following cancer treatment. The type of cell death may be controllable by manipulating the activity of PARPs. If an optimal mode of cell death is manipulated in cancer therapy, side effects such as over inflammation response in surrounding normal tissue may be avoidable. Collectively, a comprehensive understanding of PARP signaling from DNA damage to cell death will be important to optimize a therapeutic protocol in cancer treatment.
References
Alvarez-Gonzalez R, Althaus FR (1989) Poly(ADP-ribose) catabolism in mammalian cells exposed to DNA-damaging agents. Mutat Res 218:67–74
Benjamin RC, Gill DM (1980) ADP-ribosylation in mammalian cell ghosts. Dependence of poly(ADP-ribose) synthesis on strand breakage in DNA. J Biol Chem 255:10493–10501
Benjamin RC, Gill DM (1980) Poly(ADP-ribose) synthesis in vitro programmed by damaged DNA. A comparison of DNA molecules containing different types of strand breaks. J Biol Chem 255:10502–10508
Boehler C, Gauthier LR, Mortusewicz O, Biard DS, Saliou JM, Bresson A, Sanglier-Cianferani S, Smith S, Schreiber V, Boussin F, Dantzer F (2011) Poly(ADP-ribose) polymerase 3 (PARP3), a newcomer in cellular response to DNA damage and mitotic progression. Proc Natl Acad Sci U S A 108:2783–2788
Ikejima M, Noguchi S, Yamashita R, Suzuki H, Sugimura T, Miwa M (1989) Expression of human poly(ADP-ribose) polymerase with DNA-dependent enzymatic activity in Escherichia coli. Biochem Biophys Res Commun 163:739–745
El-Khamisy SF, Masutani M, Suzuki H, Caldecott KW (2003) A requirement for PARP-1 for the assembly or stability of XRCC1 nuclear foci at sites of oxidative DNA damage. Nucleic Acids Res 31:5526–5533
Le Page F, Schreiber V, Dherin C, De Murcia G, Boiteux S (2003) Poly(ADP-ribose) polymerase-1 (PARP-1) is required in murine cell lines for base excision repair of oxidative DNA damage in the absence of DNA polymerase beta. J Biol Chem 278:18471–18477
Masson M, Niedergang C, Schreiber V, Muller S, Menissier-de Murcia J, de Murcia G (1998) XRCC1 is specifically associated with poly(ADP-ribose) polymerase and negatively regulates its activity following DNA damage. Mol Cell Biol 18:3563–3571
de Murcia JM, Niedergang C, Trucco C, Ricoul M, Dutrillaux B, Mark M, Oliver FJ, Masson M, Dierich A, LeMeur M, Walztinger C, Chambon P, de Murcia G (1997) Requirement of poly(ADP-ribose) polymerase in recovery from DNA damage in mice and in cells. Proc Natl Acad Sci U S A 94:7303–7307
Schreiber V, Ame JC, Dolle P, Schultz I, Rinaldi B, Fraulob V, Menissier-de Murcia J, de Murcia G (2002) Poly(ADP-ribose) polymerase-2 (PARP-2) is required for efficient base excision DNA repair in association with PARP-1 and XRCC1. J Biol Chem 277:23028–23036
Bryant HE, Schultz N, Thomas HD, Parker KM, Flower D, Lopez E, Kyle S, Meuth M, Curtin NJ, Helleday T (2005) Specific killing of BRCA2-deficient tumours with inhibitors of poly(ADP-ribose) polymerase. Nature 434:913–917
Farmer H, McCabe N, Lord CJ, Tutt AN, Johnson DA, Richardson TB, Santarosa M, Dillon KJ, Hickson I, Knights C, Martin NM, Jackson SP, Smith GC, Ashworth A (2005) Targeting the DNA repair defect in BRCA mutant cells as a therapeutic strategy. Nature 434:917–921
Furuta T, Takemura H, Liao ZY, Aune GJ, Redon C, Sedelnikova OA, Pilch DR, Rogakou EP, Celeste A, Chen HT, Nussenzweig A, Aladjem MI, Bonner WM, Pommier Y (2003) Phosphorylation of histone H2AX and activation of Mre11, Rad50, and Nbs1 in response to replication-dependent DNA double-strand breaks induced by mammalian DNA topoisomerase I cleavage complexes. J Biol Chem 278:20303–20312
Saleh-Gohari N, Bryant HE, Schultz N, Parker KM, Cassel TN, Helleday T (2005) Spontaneous homologous recombination is induced by collapsed replication forks that are caused by endogenous DNA single-strand breaks. Mol Cell Biol 25:7158–7169
Shrivastav M, De Haro LP, Nickoloff JA (2008) Regulation of DNA double-strand break repair pathway choice. Cell Res 18:134–147
Helleday T (2010) Homologous recombination in cancer development, treatment and development of drug resistance. Carcinogenesis 31:955–960
Schlacher K, Christ N, Siaud N, Egashira A, Wu H, Jasin M (2011) Double-strand break repair-independent role for BRCA2 in blocking stalled replication fork degradation by MRE11. Cell 145:529–542
Ying S, Hamdy FC, Helleday T (2012) Mre11-dependent degradation of stalled DNA replication forks is prevented by BRCA2 and PARP1. Cancer Res 72:2814–2821
Chalmers AJ (2009) The potential role and application of PARP inhibitors in cancer treatment. Br Med Bull 89:23–40
Curtin N (2014) PARP inhibitors for anticancer therapy. Biochem Soc Trans 42:82–8
Caldecott KW (2001) Mammalian DNA single-strand break repair: an X-ra(y)ted affair. Bioessays 23:447–455
Dungey FA, Caldecott KW, Chalmers AJ (2009) Enhanced radiosensitization of human glioma cells by combining inhibition of poly(ADP-ribose) polymerase with inhibition of heat shock protein 90. Mol Cancer Ther 8:2243–2254
Chalmers AJ (2007) Radioresistant glioma stem cells-therapeutic obstacle or promising target? DNA Repair (Amst) 6:1391–1394
Chalmers A, Johnston P, Woodcock M, Joiner M, Marples B (2004) PARP-1, PARP-2, and the cellular response to low doses of ionizing radiation. Int J Radiat Oncol Biol Phys 58:410–419
Loser DA, Shibata A, Shibata AK, Woodbine LJ, Jeggo PA, Chalmers AJ (2010) Sensitization to radiation and alkylating agents by inhibitors of poly(ADP-ribose) polymerase is enhanced in cells deficient in DNA double-strand break repair. Mol Cancer Ther 9:1775–1787
Riballo E, Kuhne M, Rief N, Doherty A, Smith GC, Recio MJ, Reis C, Dahm K, Fricke A, Krempler A, Parker AR, Jackson SP, Gennery A, Jeggo PA, Lobrich M (2004) A pathway of double-strand break rejoining dependent upon ATM, Artemis, and proteins locating to gamma-H2AX foci. Mol Cell 16:715–724
Beucher A, Birraux J, Tchouandong L, Barton O, Shibata A, Conrad S, Goodarzi AA, Krempler A, Jeggo PA, Lobrich M (2009) ATM and Artemis promote homologous recombination of radiation-induced DNA double-strand breaks in G2. EMBO J 28:3413–3427
Goodarzi AA, Noon AT, Deckbar D, Ziv Y, Shiloh Y, Lobrich M, Jeggo PA (2008) ATM signaling facilitates repair of DNA double-strand breaks associated with heterochromatin. Mol Cell 31:167–177
Goodarzi AA, Kurka T, Jeggo PA (2011) KAP-1 phosphorylation regulates CHD3 nucleosome remodeling during the DNA double-strand break response. Nat Struct Mol Biol 18:831–839
Hochegger H, Dejsuphong D, Fukushima T, Morrison C, Sonoda E, Schreiber V, Zhao GY, Saberi A, Masutani M, Adachi N, Koyama H, de Murcia G, Takeda S (2006) PARP-1 protects homologous recombination from interference by Ku and Ligase IV in vertebrate cells. EMBO J 25:1305–1314
Jeggo P, Lavin MF (2009) Cellular radiosensitivity: how much better do we understand it? Int J Radiat Biol 85:1061–1081
Lieber MR (2010) The mechanism of double-strand DNA break repair by the nonhomologous DNA end-joining pathway. Annu Rev Biochem 79:181–211
Wang M, Wu W, Wu W, Rosidi B, Zhang L, Wang H, Iliakis G (2006) PARP-1 and Ku compete for repair of DNA double strand breaks by distinct NHEJ pathways. Nucleic Acids Res 34:6170–618250
Betermier M, Bertrand P, Lopez BS (2014) Is non-homologous end-joining really an inherently error-prone process? PLoS Genet 10:e1004086
Zhang Y, Jasin M (2011) An essential role for CtIP in chromosomal translocation formation through an alternative end-joining pathway. Nat Struct Mol Biol 18:80–84
Los M, Mozoluk M, Ferrari D, Stepczynska A, Stroh C, Renz A, Herceg Z, Wang ZQ, Schulze-Osthoff K (2002) Activation and caspase-mediated inhibition of PARP: a molecular switch between fibroblast necrosis and apoptosis in death receptor signaling. Mol Biol Cell 13:978–988
Wu W, Wang M, Mussfeldt T, Iliakis G (2008) Enhanced use of backup pathways of NHEJ in G2 in Chinese hamster mutant cells with defects in the classical pathway of NHEJ. Radiat Res 170:512–20
Mansour WY, Borgmann K, Petersen C, Dikomey E, Dahm-Daphi J (2013) The absence of Ku but not defects in classical non-homologous end-joining is required to trigger PARP1-dependent end-joining. DNA Repair (Amst) 12:1134–1142
Diggle CP, Bentley J, Knowles MA, Kiltie AE (2005) Inhibition of double-strand break non-homologous end-joining by cisplatin adducts in human cell extracts. Nucleic Acids Res 33:2531–2539
Greiner TC, Dasgupta C, Ho VV, Weisenburger DD, Smith LM, Lynch JC, Vose JM, Fu K, Armitage JO, Braziel RM, Campo E, Delabie J, Gascoyne RD, Jaffe ES, Muller-Hermelink HK, Ott G, Rosenwald A, Staudt LM, Im MY, Karaman MW, Pike BL, Chan WC, Hacia JG (2006) Mutation and genomic deletion status of ataxia telangiectasia mutated (ATM) and p53 confer specific gene expression profiles in mantle cell lymphoma. Proc Natl Acad Sci U S A 103:2352–2357
Liu Y, Zhou K, Zhang H, Shugart YY, Chen L, Xu Z, Zhong Y, Liu H, Jin L, Wei Q, Huang F, Lu D, Zhou L (2008) Polymorphisms of LIG4 and XRCC4 involved in the NHEJ pathway interact to modify risk of glioma. Hum Mutat 29:381–389
Bentley J, Diggle CP, Harnden P, Knowles MA, Kiltie AE (2004) DNA double strand break repair in human bladder cancer is error prone and involves microhomology-associated end-joining. Nucleic Acids Res 32:5249–5259
Groselj B, Kerr M, Kiltie AE (2013) Radiosensitisation of bladder cancer cells by panobinostat is modulated by Ku80 expression. Radiother Oncol 108:429–433
Rulten SL, Fisher AE, Robert I, Zuma MC, Rouleau M, Ju L, Poirier G, Reina-San-Martin B, Caldecott KW (2011) PARP-3 and APLF function together to accelerate nonhomologous end-joining. Mol Cell 41:33–45
Ho EL, Satoh MS (2003) Repair of single-strand DNA interruptions by redundant pathways and its implication in cellular sensitivity to DNA-damaging agents. Nucleic Acids Res 31:7032–7040
Skidmore CJ, Davies MI, Goodwin PM, Halldorsson H, Lewis PJ, Shall S, Zia’ee AA (1979) The involvement of poly(ADP-ribose) polymerase in the degradation of NAD caused by gamma-radiation and N-methyl-N-nitrosourea. Eur J Biochem 101:135–142
Smulson ME, Schein P, Mullins DW Jr, Sudhakar S (1977) A putative role for nicotinamide adenine dinucleotide-promoted nuclear protein modification in the antitumor activity of N-methyl-N-nitrosourea. Cancer Res 37:3006–3012
Sims JL, Berger SJ, Berger NA (1983) Poly(ADP-ribose) Polymerase inhibitors preserve nicotinamide adenine dinucleotide and adenosine 5’-triphosphate pools in DNA-damaged cells: mechanism of stimulation of unscheduled DNA synthesis. Biochemistry 22:5188–5194
Degterev A, Huang Z, Boyce M, Li Y, Jagtap P, Mizushima N, Cuny GD, Mitchison TJ, Moskowitz MA, Yuan J (2005) Chemical inhibitor of nonapoptotic cell death with therapeutic potential for ischemic brain injury. Nat Chem Biol 1:112–119
Xu Y, Huang S, Liu ZG, Han J (2006) Poly(ADP-ribose) polymerase-1 signaling to mitochondria in necrotic cell death requires RIP1/TRAF2-mediated JNK1 activation. J Biol Chem 281:8788–8795
Jouan-Lanhouet S, Arshad MI, Piquet-Pellorce C, Martin-Chouly C, Le Moigne-Muller G, Van Herreweghe F, Takahashi N, Sergent O, Lagadic-Gossmann D, Vandenabeele P, Samson M, Dimanche-Boitrel MT (2012) TRAIL induces necroptosis involving RIPK1/RIPK3-dependent PARP-1 activation. Cell Death Differ 19:2003–2014
Sosna J, Voigt S, Mathieu S, Lange A, Thon L, Davarnia P, Herdegen T, Linkermann A, Rittger A, Chan FK, Kabelitz D, Schutze S, Adam D (2014) TNF-induced necroptosis and PARP-1-mediated necrosis represent distinct routes to programmed necrotic cell death. Cell Mol Life Sci 71:331–348
Braun SA, Panzeter PL, Collinge MA, Althaus FR (1994) Endoglycosidic cleavage of branched polymers by poly(ADP-ribose) glycohydrolase. Eur J Biochem 220:369–375
Brochu G, Duchaine C, Thibeault L, Lagueux J, Shah GM, Poirier GG (1994) Mode of action of poly(ADP-ribose) glycohydrolase. Biochim Biophys Acta 1219:342–350
Hatakeyama K, Nemoto Y, Ueda K, Hayaishi O (1986) Purification and characterization of poly(ADP-ribose) glycohydrolase. Different modes of action on large and small poly(ADP-ribose). J Biol Chem 261:14902–14911
Andrabi SA, Kim NS, Yu SW, Wang H, Koh DW, Sasaki M, Klaus JA, Otsuka T, Zhang Z, Koehler RC, Hurn PD, Poirier GG, Dawson VL, Dawson TM (2006) Poly(ADP-ribose) (PAR) polymer is a death signal. Proc Natl Acad Sci U S A 103:18308–18313
Yu SW, Andrabi SA, Wang H, Kim NS, Poirier GG, Dawson TM, Dawson VL (2006) Apoptosis-inducing factor mediates poly(ADP-ribose) (PAR) polymer-induced cell death. Proc Natl Acad Sci U S A 103:18314–18319
Artus C, Boujrad H, Bouharrour A, Brunelle MN, Hoos S, Yuste VJ, Lenormand P, Rousselle JC, Namane A, England P, Lorenzo HK, Susin SA (2010) AIF promotes chromatinolysis and caspase-independent programmed necrosis by interacting with histone H2AX. EMBO J 29:1585–1599
Benchoua A, Couriaud C, Guegan C, Tartier L, Couvert P, Friocourt G, Chelly J, Menissier-de Murcia J, Onteniente B (2002) Active caspase-8 translocates into the nucleus of apoptotic cells to inactivate poly(ADP-ribose) polymerase-2. J Biol Chem 277:34217–34222
Kaufmann SH, Desnoyers S, Ottaviano Y, Davidson NE, Poirier GG (1993) Specific proteolytic cleavage of poly(ADP-ribose) polymerase: an early marker of chemotherapy-induced apoptosis. Cancer Res 53:3976–3985
Fernandes-Alnemri T, Takahashi A, Armstrong R, Krebs J, Fritz L, Tomaselli KJ, Wang L, Yu Z, Croce CM, Salveson G et al (1995) Mch3, a novel human apoptotic cysteine protease highly related to CPP32. Cancer Res 55:6045–6052
Lazebnik YA, Kaufmann SH, Desnoyers S, Poirier GG, Earnshaw WC (1994) Cleavage of poly(ADP-ribose) polymerase by a proteinase with properties like ICE. Nature 371:346–347
Tewari M, Quan LT, O’rourke K, Desnoyers S, Zeng Z, Beidler DR, Poirier GG, Salvesen GS, Dixit VM (1995) Yama/CPP32 beta, a mammalian homolog of CED-3, is a CrmA-inhibitable protease that cleaves the death substrate poly(ADP-ribose) polymerase. Cell 81:801–809
Nicholson DW, Ali A, Thornberry NA, Vaillancourt JP, Ding CK, Gallant M, Gareau Y, Griffin PR, Labelle M, Lazebnik YA et al (1995) Identification and inhibition of the ICE/CED-3 protease necessary for mammalian apoptosis. Nature 376:37–43
Kim JW, Kim K, Kang K, Joe CO (2000) Inhibition of homodimerization of poly(ADP-ribose) polymerase by its C-terminal cleavage products produced during apoptosis. J Biol Chem 275:8121–8125
D’amours D, Sallmann FR, Dixit VM, Poirier GG (2001) Gain-of-function of poly(ADP-ribose) polymerase-1 upon cleavage by apoptotic proteases: implications for apoptosis. J Cell Sci 114:3771–3778
Oliver FJ, de la Rubia G, Rolli V, Ruiz-Ruiz MC, de Murcia G, Murcia JM (1998) Importance of poly(ADP-ribose) polymerase and its cleavage in apoptosis. Lesson from an uncleavable mutant. J Biol Chem 273:33533–33539
Boulares AH, Yakovlev AG, Ivanova V, Stoica BA, Wang G, Iyer S, Smulson M (1999) Role of poly(ADP-ribose) polymerase (PARP) cleavage in apoptosis. Caspase 3-resistant PARP mutant increases rates of apoptosis in transfected cells. J Biol Chem 274:22932–22940
Herceg Z, Wang ZQ (1999) Failure of poly(ADP-ribose) polymerase cleavage by caspases leads to induction of necrosis and enhanced apoptosis. Mol Cell Biol 19:5124–5133
Casiano CA, Ochs RL, Tan EM (1998) Distinct cleavage products of nuclear proteins in apoptosis and necrosis revealed by autoantibody probes. Cell Death Differ 5:183–1903
Gobeil S, Boucher CC, Nadeau D, Poirier GG (2001) Characterization of the necrotic cleavage of poly(ADP-ribose) polymerase (PARP-1): implication of lysosomal proteases. Cell Death Differ 8:588–594
McGinnis KM, Gnegy ME, Park YH, Mukerjee N, Wang KK (1999) Procaspase-3 and poly(ADP)ribose polymerase (PARP) are calpain substrates. Biochem Biophys Res Commun 263:94–99
Shah GM, Shah RG, Poirier GG (1996) Different cleavage pattern for poly(ADP-ribose) polymerase during necrosis and apoptosis in HL-60 cells. Biochem Biophys Res Commun 229:838–844
Ishida M, Gomyo Y, Tatebe S, Ohfuji S, Ito H (1996) Apoptosis in human gastric mucosa, chronic gastritis, dysplasia and carcinoma: analysis by terminal deoxynucleotidyl transferase-mediated dUTP-biotin nick end labelling. Virchows Arch 428:229–235
Tao D, Wu J, Feng Y, Qin J, Hu J, Gong J (2004) New method for the analysis of cell cycle-specific apoptosis. Cytometry A 57:70–74
Okita N, Yoshimura M, Watanabe K, Minato S, Kudo Y, Higami Y, Tanuma S (2013) CHK1 cleavage in programmed cell death is intricately regulated by both caspase and non-caspase family proteases. Biochim Biophys Acta 1830:2204–2213
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Okita, N., Shibata, A. (2015). Other Determinants of Sensitivity. In: Curtin, N., Sharma, R. (eds) PARP Inhibitors for Cancer Therapy. Cancer Drug Discovery and Development, vol 83. Humana Press, Cham. https://doi.org/10.1007/978-3-319-14151-0_15
Download citation
DOI: https://doi.org/10.1007/978-3-319-14151-0_15
Published:
Publisher Name: Humana Press, Cham
Print ISBN: 978-3-319-14150-3
Online ISBN: 978-3-319-14151-0
eBook Packages: MedicineMedicine (R0)