Abstract
Crohn’s disease (CD) is a chronic inflammatory bowel disease with a complex aetiology that includes genetic susceptibility and the gastrointestinal microbiome and results in an aberrant Th17 inflammatory response. NOD2 is an intracellular sensor that responds to bacterial cell wall peptidoglycan and contributes to immune defense. Polymorphisms in the NOD2 gene predispose to Crohn’s disease, with the largest effect of any of the known genetic risk factors. We have found that wild-type NOD2 controls the expression of miR-29 in human dendritic cells (DCs). miR-29 regulates the expression of a number of immune mediators including the IL-23 cytokine subunits IL-12p40 and IL-23p19. CD patient DCs expressing NOD2 polymorphisms fail to induce miR-29 and show enhanced IL-12p40 release on exposure to adherent invasive E. coli. Moreover in a murine model deficient in miR-29, a more severe Th17-driven colitis is established after DSS administration. Therefore, we suggest that the loss of miR-29-mediated immunoregulation in CD-variant NOD2 DCs contributes to elevated IL-23 and aberrant Th17 response in this disease.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Paneth Cell
- Muramyl Dipeptide
- Innate Immune Receptor
- Bacterial Cell Wall Peptidoglycan
- Inflammatory Bowel Disease Genetic
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
1 Introduction
Crohn’s disease (CD) is a form of chronic inflammatory bowel disease (IBD). It typically presents in young people (peak onset between years 10 and 20), with the highest incidence in the western world with a rising incidence as nations move towards a western lifestyle and environment [1]. CD usually manifests clinically with a combination of abdominal pain, weight loss, and diarrhoea. The clinical course is variable, from a fairly mild self-limiting illness through to severe disease resulting in multiple surgeries, short gut syndrome and severe malabsorption. Although there have been substantial improvements in medical therapy, there exists a significant therapeutic gap as evidenced by the modest sustained remission rates with anti-TNF therapy [2] (currently the most potent available therapy), plus the requirement for at least one surgery in the majority of patients (up to 80 % in ileal disease [3, 4]). The transmural granulomatous inflammation typical of Crohn’s can involve any part of the gastrointestinal (GI) tract from mouth to anus, but usually affects the terminal ileum, colon and/or perianal regions.
The aetiology of Crohn’s disease is complex and multifactorial, with the best evidence suggesting that it involves a combination of genetic susceptibility, a disordered GI tract microflora (dysbiosis), and possibly a third as yet poorly characterised environmental factor, or factors.
There is good observational evidence for a genetic component to disease susceptibility, from the documentation of familial clustering by Burrell Crohn himself [5] to the monozygotic twin disease concordance rate of nearly 50 % [6] and the high percentage of patients with a positive family history [7]. Over the past 15 years genetic studies have identified an increasing number of polymorphisms associated with CD, and currently there are 163 loci associated with IBD [8]. The modern era of IBD genetics was initiated by the discovery of mutations in the caspase recruitment domain family member containing 15 (CARD15), or nucleotide-binding oligomerisation domain containing 2 (NOD2), gene in the IBD1 locus on chromosome 16 [9].
CARD15/NOD2 (henceforth NOD2) encodes a cytosolic innate immune receptor and is a member of the NOD-like receptor (NLR) family. NOD2 is triggered by bacterial cell wall muramyl dipeptide (MDP) [10, 11], which is expressed by both gram +ve and gram −ve organisms. This aligns with another central tenet of CD susceptibility—that of the influence of the GI tract bacteria or microbiome. The GI tract is home to a staggering quantity of bacteria—at their most dense the bacteria number 1012/ml. Coupled with this is the startling fact that these bacteria are only physically separated from GI tissue residence/invasion by a single layer of epithelial cells and a covering of mucus (varying in depth from approximately 200 to 700 μm). In many respects, it is interesting that we do not develop overt inflammation of the GI tract more frequently.
Nevertheless, we are dependent on the development and the presence of a normal gut flora for a healthy gut epithelium, for digestion and for the development and function of the immune system [12]. Defining what constitutes a ‘normal’ gut flora is still a work in progress and depends in part on what is sampled (faeces or mucosa-associated bacteria) [13] and on other factors such as ones underlying genotype [14], diet [15], age [16] and any recent antibiotic use. The colonisation of the GI tract begins at birth (with commensals from the mother’s vagina or skin, depending on delivery via birth canal or C-section respectively), and then slowly develops over 2–3 years into that consistent with an adult faecal signature. Some elements of the commensal bacterial flora, such as Faecalibacterium prausnitzii [17, 18] (or F. prau) in humans, or certain Clostridial species in mice [19], appear to have crucial anti-inflammatory properties. Conversely, the presence of ‘pathobionts’ such as adherent-invasive Escherichia coli (AIEC) [20–22] correlates with an increased risk of ileal CD.
There is observational and experimental evidence that the presence of the microbiota is necessary in driving, if not definitively initiating, GI inflammation. In humans, antibiotics can ameliorate CD [23], faecal stream diversion is an effective therapy for Crohn’s [24, 25], and lamina propria T cells from CD patients are reactive to gut flora [26]. Murine models of colitis are for the most part dependent on the presence of the microbial flora to develop colitis (an exception being the anti-CD40 model). Moreover, under certain conditions, it is possible to transfer a colitogenic flora from one mouse model to another [27].
Patients with Crohn’s disease have an altered microbiome or ‘dysbiosis’, and disease severity corresponds with the degree of difference from healthy controls [28]. The main difference in CD appears to be that the microbial community is more unstable over time and less diverse, for example, with the loss of Bacteroidetes and certain Firmicutes, and an increase in Proteobacteria [29]. It remains unclear how much of this dysbiosis is a primary aetiological event, and how much is secondary to an already established inflammation.
The nature of the aberrant inflammatory response in Crohn’s has been demonstrated genetically, in murine models and in human studies. The significance of IL-23 and the Th17 pathway is highlighted by polymorphisms in IL23R, IL12B (encoding for IL-12p40), STAT3, JAK2 and TYK2 [30]. Both innate and T cell-dependent models of murine colitis demonstrate a key role for IL-23. In innate colitis, IL-23 directs the expression of IL-17 from innate lymphoid cells [31]. In T cells, IL-23R signalling leads to enhanced Th17 polarisation and reduced FoxP3+ T cell differentiation and IL-10 expression [32]. Finally, in human studies, IL-23 expression is increased in the mucosa of IBD patients [33].
2 NOD2 and Crohn’s Disease
The role of NOD2 within this complex GI environment is an area of active research, and some progress has been made in the 14 years since the gene was identified. NOD2 is constitutively expressed in GI epithelial cells [34], as well as in myelomonocytic cells such as dendritic cells (DCs) [35, 36]. NOD2 has two N-terminal caspase-recruitment domains (CARDs), a central nucleotide-binding oligomerisation domain (NOD) and a C-terminal leucine-rich repeat (LRR) domain. The polymorphisms that predispose to CD occur in (or adjacent to in the case of R702W) the LRR domain that functions as the MDP ligand-recognition domain. Three single nucleotide polymorphisms (one frame-shift: FS1007insC and two missense: G908R and R702W), account for over 80 % of those identified [37]. Homozygosity, or compound heterozygosity, confer a 17-fold increased risk of developing CD, whilst around 40 % of European CD patients carry at least one mutation [38] versus 14 % of healthy controls. Mutations in NOD2 predispose to terminal ileal CD [38], early-onset disease [37, 39] and possibly a stricturing phenotype [40].
NOD2 activation, on recognition of MDP, is thought to result in oligomerisation [41, 42] and recruitment of an adaptor protein RIPK-2 [43] via a CARD–CARD domain interaction. This leads, via an incompletely understood signalling cascade, to NFκB activation [36] and pro-inflammatory cytokine production. NOD2 is just one of a number of innate immune pattern recognition receptors (PRRs) expressed by antigen-presenting cells such as DCs. Others include the membrane-bound toll-like receptors (TLRs), the C-type lectin receptors (CLRs) and the cytosolic Nod-like receptors (NLRs) of which NOD2 is a member. These PRRs typically respond to pathogen-associated molecular patterns (PAMPs) or damage-associated molecular patterns (DAMPs). Immune activation by whole microbes is therefore complex, and it is the combination of PRR triggering by a given organism, plus the nature of the cellular microenvironment, that will influence the nature of the subsequent immune response.
Although there is now a large literature on the function of NOD2, the key findings can be summarised reasonably succinctly. Firstly, NOD2 induces autophagy on bacterial recognition in both dendritic [44] and epithelial cells [45], helping to clear the invasive bacteria and (in DCs) facilitating MHC class II antigen presentation. Autophagy is an intrinsic cytosolic process in which the formation of double-membrane vesicles facilitates the degradation of intracellular proteins (macro-autophagy) [46], bacteria (xenophagy) [47] or mitochondria (mitophagy) [48]. The role of autophagy in Crohn’s pathogenesis only became apparent when genetic studies revealed polymorphisms in both ATG16L1 and IRGM [30]. The subsequent studies linking NOD2 and xenophagy are important in that they coalesce apparently unrelated CD polymorphisms around the same defective pathway.
Secondly, NOD2 is highly expressed in specialised small intestinal epithelial cells, called Paneth cells [34, 49], which are themselves most numerous in the terminal ileum [50]. Paneth cells are located in the crypts of Lieberkühn, and amongst other functions they secrete anti-microbial peptides, such as defensins. NOD2 polymorphisms are associated with decreased alpha defensin production (HD-5 and HD-6) from Paneth cells [51, 52], raising the question as to whether this is a primary defect in CD pathogenesis. However, there is some continued debate as to whether this apparent reduction owes more to inflammation rather than NOD2 expression [53] or whether the reverse is true [54]. In certain cases, the lack of defensins is due to other pathways altogether, such as defective Wnt signalling [55, 56].
Lastly, as already described, NOD2 is only one of a broad range of innate immune receptors. Large-scale gene-expression studies reveal that NOD2 can synergise with other PRRs, and that this synergy is lost in the presence of CD-variant NOD2 [57–59]. Wild-type NOD2, which in isolation has a relatively weak effect [57], has a key role in amplifying the release of certain pro-inflammatory cytokines, particularly interleukin-1β (IL-1β), IL-6 and IL-23 from DCs and macrophages [60, 61]. It is mooted that this NOD2-driven amplification of TLR responses is in keeping with the cytosolic expression of the NOD2 protein, because MDP stimulation may be indicative of invasive bacterial infection.
3 NOD2 and miR-29
One of the questions that arise from the cytokine data is why the CD-variant NOD2 should predispose to a pro-inflammatory condition. PRR signalling pathways that induce effector responses, including cytokines, need to be tightly regulated in order to limit bystander damage and terminate the immune response. microRNAs (miRNAs), the main function of which is to target messenger RNA (mRNA), are key regulators of gene expression in mammalian cells. Prior to our investigation, a number of studies of TLR signalling had established a role for miRNAs in the negative regulation of innate immune responses [62–66]. We hypothesised that wild-type NOD2 triggering would contribute to miRNA expression and that the absence of this in CD-variant NOD2 would lead to aberrant cytokine expression and the immunopathology seen in Crohn’s.
We used human monocyte-derived dendritic cells (MDDCs) to explore miRNA expression initially after NOD2 and/or TLR2 stimulation [67]. TLR2 responds to Pam3CSK4 that, along with MDP, is a component of bacterial cell wall peptidoglycan, meaning that TLR2 is very likely to be co-triggered with NOD2. An miRNA array identified not only the already-described miR-155 and miR-146 as downstream of TLR2 triggering but also miR-29 expression as a novel miRNA in relation to innate immune signalling. Further investigation established that wild-type NOD2 stimulation, and the presence of the adaptor molecule RIPK-2, is critical for the up-regulation of miR-29 and that this is amplified by the co-stimulation of TLR2 or TLR5. Interestingly, this effect is not dependent on the TLR-signalling adaptor molecule MyD88. miR-29 is part of a miRNA family expressed from two clusters on chromosomes 1 and 7 (mir-29a, b and c), and these miRNAs possess identical seed sequences, therefore targeting the same mRNAs. miR-29 expression in MDDCs is detectable from 12 h after NOD2 + TLR2 stimulation and peaks at around day 3. This delayed expression pattern would be in keeping with a role as a negative regulator of immune response, allowing effector mechanisms to work before appropriate termination.
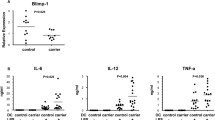
In order to identify potential miR-29 targets in MDDCs, we transfected NOD2- and TLR2-stimulated DCs with miR-29 premiR to artificially increase miR-29 expression. We used large-scale gene-expression microarrays to subsequently identify a number of differentially regulated genes and went on to validate these by quantitative PCR (qPCR). These genes included a number of pro-inflammatory and immune pathway mediators, and one of the most strongly down-regulated genes identified by this methodology was IL-12p40. IL-12p40 is a predicted target of miR-29 (Targetscan) and is a cytokine subunit of both IL-12 (with IL-12p35) and IL-23 (with IL-23p19). We established that miR-29 directly targets IL-12p40 mRNA via the 3′UTR and indirectly down-regulates IL-23p19, but not IL-12p35.
We went on to demonstrate that NOD2-directed control of miR-29 is relevant at physiological miRNA expression levels. Firstly, we utilised an IL-12p40 3′UTR seed target protector that was designed for miR-29 binding sites. MDDCs transfected with this protector and then stimulated with NOD2 and TLR2 ligands for 24 h express elevated levels of IL-12p40. Secondly, we identified patients with ileal CD and who are homozygous for CD-variant NOD2 polymorphisms. MDDCs from these patients fail to up-regulate miR-29 when NOD2 is triggered and, perhaps more importantly, they express higher levels of IL-12p40 after AIEC infection than their wild-type counterparts. This discrepancy is rescued by the transfection, or replacement, of miR-29 using a premiR.
We next explored the role of miR-29 in vivo using mice with a targeted deletion of the miR-29a/b-1 locus (henceforth mir-29 KO mice). We investigated whether a lack of miR-29 in a murine model alters the development or susceptibility to colitis in a DSS model. miR-29 KO mice show an increased susceptibility to colitis (1.7-fold higher compared to WT littermates) and exhibit more severe pathological scores and weight loss. In keeping with the human in vitro data, we demonstrated a marked Th17 transcriptional signature from inflamed colonic tissue. This included elevated expression of the miR-29-targeted Il12b and Il23a, as well as the mRNA encoding cytokine IL-17A, and the Th17 subset-determining transcription factor RORγt. This contrasts with the transcription factors GATA-3, T-bet and Foxp3 which are essentially unchanged. Moreover, there is no general change in pro- or anti-inflammatory mediators between wild-type and mir-29 KO mice, with similar colonic expression of Il1b, Tnfa and Il6 and Il10. In other murine models, miR-29 targets IFN-γ in NK cells, CD4+ and CD8+ T cells [68], and through targeting of T-bet and Eomes in T cells influences Th1 bias [69], but in our model we found no difference in IFN-γ expression or Th1 cell numbers in colonic tissue.
4 Conclusion
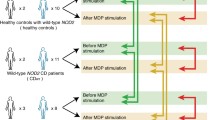
In summary, our data suggest that wild-type NOD2 has an important role, via the expression of miR-29, in ensuring that the critical balance of pro- and anti-inflammatory responses is maintained within the GI tract (Fig. 1). There is evidence that IL-23 expression is particularly prominent within the terminal ileum [70], which would align with the known phenotype of NOD2-associated Crohn’s. We would propose that the defective expression of miR-29 observed in the presence of CD-variant NOD2 should be placed in context of other NOD2 data [71]. In this model excess inflammation in the terminal ileum would result from a dysbiosis (possibly related to defensin deficiency), aberrant bacterial handling and persistence due to defective autophagy and an inability to appropriately arrest the resultant Th17 response due to deficient miR-29 expression.
A representation of the role of NOD2-stimulated miR-29 expression in antigen presenting cells (APCs) such as gastrointestinal DCs. In the left-hand panel the cell is challenged by bacterial infection and mounts an appropriate Th17 immune response. The central panel details appropriate resolution of this response through delayed miR-29 expression in the presence of wild-type NOD2. The right-hand panel depicts the problems that occur in the presence of CD-variant NOD2, with initial bacterial persistence due to defective autophagy, and a subsequently unchecked Th17 response due to lack of miR-29 up-regulation
Abbreviations
- AIEC:
-
Adherent-invasive Escherichia coli
- APC:
-
Antigen presenting cell
- ATG16L1:
-
Autophagy related 16-like 1
- CARD15:
-
Caspase recruitment domain family member 15
- CD:
-
Crohn’s disease
- Chr:
-
Chromosome
- CLR:
-
C-type lectin receptors
- DAMP:
-
Damage-associated molecular pattern
- DC:
-
Dendritic cell
- DSS:
-
Dextran sodium sulphate
- F. prau :
-
Faecalibacterium prausnitzii
- Foxp3:
-
Forkhead box P3
- GI:
-
Gastrointestinal
- GWAS:
-
Genome-wide association study
- HD:
-
Human defensin
- IBD:
-
Inflammatory bowel disease
- IFN:
-
Interferon
- IL:
-
Interleukin
- IL-12B:
-
Interleukin 12B/IL-12p40
- IL-23R:
-
Interleukin-23 receptor
- IRGM:
-
Immunity-related GTPase family, M
- JAK2:
-
Janus kinase 2
- KO:
-
Knock-out
- LPS:
-
Lipopolysaccharide
- LRR:
-
Leucine-rich repeat
- MDDC:
-
Monocyte-derived dendritic cell
- MDP:
-
Muramyl dipeptide
- miR:
-
microRNA
- MyD88:
-
Myeloid differentiation primary response gene 88
- NFκB:
-
Nuclear factor kappa B
- NK:
-
Natural killer
- NLR:
-
NOD-like receptor
- NOD2:
-
Nucleotide-binding oligomerisation domain containing 2
- Pam3CSK4 :
-
Synthetic triacylated lipoprotein—TLR1/2 ligand
- PAMP:
-
Pathogen-associated molecular pattern
- PCR:
-
Polymerase chain reaction
- PGN:
-
Peptidoglycan
- PRR:
-
Pattern recognition receptor
- qPCR:
-
Quantitative polymerase chain reaction
- RIPK-2:
-
Receptor-interacting protein kinase 2
- RORγt:
-
RAR-related orphan receptor gamma
- STAT3:
-
Signal transducer and activator of transcription 3
- T-bet:
-
T-box transcription factor
- Th1/17:
-
T helper 1/17
- TLR:
-
Toll-like receptor
- TNF:
-
Tumour necrosis factor
- Wnt:
-
Wingless
- WT:
-
Wild-type
References
Ng SC (2014) Epidemiology of inflammatory bowel disease: focus on Asia [Review]. Best Pract Res Clin Gastroenterol 28(3):363–372
Hanauer SB, Feagan BG, Lichtenstein GR, Mayer LF, Schreiber S, Colombel JF et al (2002) Maintenance infliximab for Crohn’s disease: the ACCENT I randomised trial. Lancet 359(9317):1541–1549
Goldberg PA, Wright JP, Gerber M, Claassen R (1993) Incidence of surgical resection for Crohn’s disease. Dis Colon Rectum 36(8):736–739
Higgens CS, Allan RN (1980) Crohn’s disease of the distal ileum. Gut 21(11):933–940
Crohn G (1932) Regional ileitis: a pathologic and clinical entity. JAMA 99:1323–1329
Halme L, Paavola-Sakki P, Turunen U, Lappalainen M, Farkkila M, Kontula K (2006) Family and twin studies in inflammatory bowel disease. World J Gastroenterol 12(23):3668–3672
Monsen U, Bernell O, Johansson C, Hellers G (1991) Prevalence of inflammatory bowel disease among relatives of patients with Crohn’s disease. Scand J Gastroenterol 26(3):302–306
Jostins L, Ripke S, Weersma RK, Duerr RH, McGovern DP, Hui KY et al (2012) Host-microbe interactions have shaped the genetic architecture of inflammatory bowel disease. Nature 491(7422):119–124
Hugot JP, Chamaillard M, Zouali H, Lesage S, Cezard JP, Belaiche J et al (2001) Association of NOD2 leucine-rich repeat variants with susceptibility to Crohn’s disease. Nature 411(6837):599–603
Girardin SE, Boneca IG, Viala J, Chamaillard M, Labigne A, Thomas G et al (2003) Nod2 is a general sensor of peptidoglycan through muramyl dipeptide (MDP) detection. J Biol Chem 278(11):8869–8872
Inohara N, Ogura Y, Fontalba A, Gutierrez O, Pons F, Crespo J et al (2003) Host recognition of bacterial muramyl dipeptide mediated through NOD2. Implications for Crohn’s disease. J Biol Chem 278(8):5509–5512
Hooper LV, Gordon JI (2001) Commensal host-bacterial relationships in the gut. Science 292(5519):1115–1118
Zoetendal EG, von Wright A, Vilpponen-Salmela T, Ben-Amor K, Akkermans AD, de Vos WM (2002) Mucosa-associated bacteria in the human gastrointestinal tract are uniformly distributed along the colon and differ from the community recovered from feces. Appl Environ Biol 68:3401–3407
Stewart JA, Chadwick VS, Murray A (2005) Investigations into the influence of host genetics on the predominant eubacteria in the faecal microflora of children. J Med Microbiol 54(Pt 12):1239–1242
Muegge BD, Kuczynski J, Knights D, Clemente JC, Gonzalez A, Fontana L et al (2011) Diet drives convergence in gut microbiome functions across mammalian phylogeny and within humans. Science 332(6032):970–974
Tiihonen K, Ouwehand AC, Rautonen N (2010) Human intestinal microbiota and healthy ageing [Review]. Ageing Res Rev 9(2):107–116
Sokol H, Pigneur B, Watterlot L, Lakhdari O, Bermudez-Humaran LG, Gratadoux JJ et al (2008) Faecalibacterium prausnitzii is an anti-inflammatory commensal bacterium identified by gut microbiota analysis of Crohn disease patients. Proc Natl Acad Sci USA 105(43):16731–16736
Sokol H, Seksik P, Furet JP, Firmesse O, Nion-Larmurier I, Beaugerie L et al (2009) Low counts of Faecalibacterium prausnitzii in colitis microbiota. Inflamm Bowel Dis 15(8):1183–1189
Atarashi K, Tanoue T, Shima T, Imaoka A, Kuwahara T, Momose Y et al (2011) Induction of colonic regulatory T cells by indigenous Clostridium species. Science 331(6015):337–341
Baumgart M, Dogan B, Rishniw M, Weitzman G, Bosworth B, Yantiss R et al (2007) Culture independent analysis of ileal mucosa reveals a selective increase in invasive Escherichia coli of novel phylogeny relative to depletion of Clostridiales in Crohn’s disease involving the ileum. ISME J 1(5):403–418
Darfeuille-Michaud A, Boudeau J, Bulois P, Neut C, Glasser AL, Barnich N et al (2004) High prevalence of adherent-invasive Escherichia coli associated with ileal mucosa in Crohn’s disease. Gastroenterology 127(2):412–421
Martin HM, Campbell BJ, Hart CA, Mpofu C, Nayar M, Singh R et al (2004) Enhanced Escherichia coli adherence and invasion in Crohn’s disease and colon cancer. Gastroenterology 127(1):80–93
Feller M, Huwiler K, Schoepfer A, Shang A, Furrer H, Egger M (2010) Long-term antibiotic treatment for Crohn’s disease: systematic review and meta-analysis of placebo-controlled trials. Clin Infect Dis 50(4):473–480
Harper PH, Truelove SC, Lee EC, Kettlewell MG, Jewell DP (1983) Split ileostomy and ileocolostomy for Crohn’s disease of the colon and ulcerative colitis: a 20 year survey. Gut 24(2):106–113
Rutgeerts P, Goboes K, Peeters M, Hiele M, Penninckx F, Aerts R et al (1991) Effect of faecal stream diversion on recurrence of Crohn’s disease in the neoterminal ileum. Lancet 338(8770):771–774
Duchmann R, May E, Heike M, Knolle P, Neurath M, Meyer zum Buschenfelde KH (1999) T cell specificity and cross reactivity towards enterobacteria, bacteroides, bifidobacterium, and antigens from resident intestinal flora in humans. Gut 44(6):812–818
Garrett WS, Lord GM, Punit S, Lugo-Villarino G, Mazmanian SK, Ito S et al (2007) Communicable ulcerative colitis induced by T-bet deficiency in the innate immune system. Cell 131(1):33–45
Ott SJ, Musfeldt M, Wenderoth DF, Hampe J, Brant O, Folsch UR et al (2004) Reduction in diversity of the colonic mucosa associated bacterial microflora in patients with active inflammatory bowel disease. Gut 53(5):685–693
Frank DN, St Amand AL, Feldman RA, Boedeker EC, Harpaz N, Pace NR (2007) Molecular-phylogenetic characterization of microbial community imbalances in human inflammatory bowel diseases. Proc Natl Acad Sci USA 104(34):13780–13785
Franke A, McGovern DP, Barrett JC, Wang K, Radford-Smith GL, Ahmad T et al (2010) Genome-wide meta-analysis increases to 71 the number of confirmed Crohn’s disease susceptibility loci. Nat Genet 42(12):1118–1125
Buonocore S, Ahern PP, Uhlig HH, Ivanov II, Littman DR, Maloy KJ et al (2010) Innate lymphoid cells drive interleukin-23-dependent innate intestinal pathology. Nature 464(7293):1371–1375
Ahern PP, Schiering C, Buonocore S, McGeachy MJ, Cua DJ, Maloy KJ et al (2010) Interleukin-23 drives intestinal inflammation through direct activity on T cells. Immunity 33(2):279–288
Liu Z, Yadav PK, Xu X, Su J, Chen C, Tang M et al (2011) The increased expression of IL-23 in inflammatory bowel disease promotes intraepithelial and lamina propria lymphocyte inflammatory responses and cytotoxicity. J Leukoc Biol 89(4):597–606
Ogura Y, Lala S, Xin W, Smith E, Dowds TA, Chen FF et al (2003) Expression of NOD2 in Paneth cells: a possible link to Crohn’s ileitis. Gut 52(11):1591–1597
Gutierrez O, Pipaon C, Inohara N, Fontalba A, Ogura Y, Prosper F et al (2002) Induction of Nod2 in myelomonocytic and intestinal epithelial cells via nuclear factor-kappa B activation. J Biol Chem 277(44):41701–41705
Ogura Y, Inohara N, Benito A, Chen FF, Yamaoka S, Nunez G (2001) Nod2, a Nod1/Apaf-1 family member that is restricted to monocytes and activates NF-kappaB. J Biol Chem 276(7):4812–4818
Lesage S, Zouali H, Cezard JP, Colombel JF, Belaiche J, Almer S et al (2002) CARD15/NOD2 mutational analysis and genotype-phenotype correlation in 612 patients with inflammatory bowel disease. Am J Hum Genet 70(4):845–857
Cuthbert AP, Fisher SA, Mirza MM, King K, Hampe J, Croucher PJ et al (2002) The contribution of NOD2 gene mutations to the risk and site of disease in inflammatory bowel disease. Gastroenterology 122(4):867–874
Ahmad T, Armuzzi A, Bunce M, Mulcahy-Hawes K, Marshall SE, Orchard TR et al (2002) The molecular classification of the clinical manifestations of Crohn’s disease. Gastroenterology 122(4):854–866
Seiderer J, Brand S, Herrmann KA, Schnitzler F, Hatz R, Crispin A et al (2006) Predictive value of the CARD15 variant 1007fs for the diagnosis of intestinal stenoses and the need for surgery in Crohn’s disease in clinical practice: results of a prospective study. Inflamm Bowel Dis 12(12):1114–1121
Hu Y, Ding L, Spencer DM, Nunez G (1998) WD-40 repeat region regulates Apaf-1 self-association and procaspase-9 activation. J Biol Chem 273(50):33489–33494
Srinivasula SM, Ahmad M, Fernandes-Alnemri T, Alnemri ES (1998) Autoactivation of procaspase-9 by Apaf-1-mediated oligomerization. Mol Cell 1(7):949–957
Kobayashi K, Inohara N, Hernandez LD, Galan JE, Nunez G, Janeway CA et al (2002) RICK/Rip2/CARDIAK mediates signalling for receptors of the innate and adaptive immune systems. Nature 416(6877):194–199
Cooney R, Baker J, Brain O, Danis B, Pichulik T, Allan P et al (2010) NOD2 stimulation induces autophagy in dendritic cells influencing bacterial handling and antigen presentation. Nat Med 16(1):90–97
Travassos LH, Carneiro LA, Ramjeet M, Hussey S, Kim YG, Magalhaes JG et al (2010) Nod1 and Nod2 direct autophagy by recruiting ATG16L1 to the plasma membrane at the site of bacterial entry. Nat Immunol 11(1):55–62
Feng Y, He D, Yao Z, Klionsky DJ (2014) The machinery of macroautophagy. Cell Res 24(1):24–41
Gomes LC, Dikic I (2014) Autophagy in antimicrobial immunity. Mol Cell 54(2):224–233
MacVicar T (2013) Mitophagy [Review]. Essays Biochem 55:93–104
Lala S, Ogura Y, Osborne C, Hor SY, Bromfield A, Davies S et al (2003) Crohn’s disease and the NOD2 gene: a role for paneth cells. Gastroenterology 125(1):47–57
Porter EM, Bevins CL, Ghosh D, Ganz T (2002) The multifaceted Paneth cell. Cell Mol Life Sci 59(1):156–170
Elphick D, Liddell S, Mahida YR (2008) Impaired luminal processing of human defensin-5 in Crohn’s disease: persistence in a complex with chymotrypsinogen and trypsin. Am J Pathol 172(3):702–713
Wehkamp J, Salzman NH, Porter E, Nuding S, Weichenthal M, Petras RE et al (2005) Reduced Paneth cell alpha-defensins in ileal Crohn’s disease. Proc Natl Acad Sci USA 102(50):18129–18134
Simms LA, Doecke JD, Walsh MD, Huang N, Fowler EV, Radford-Smith GL (2008) Reduced alpha-defensin expression is associated with inflammation and not NOD2 mutation status in ileal Crohn’s disease. Gut 57(7):903–910
Bevins CL, Stange EF, Wehkamp J (2009) Decreased Paneth cell defensin expression in ileal Crohn’s disease is independent of inflammation, but linked to the NOD2 1007fs genotype. Gut 58(6):882–883, discussion 883–884
Perminow G, Beisner J, Koslowski M, Lyckander LG, Stange E, Vatn MH et al (2010) Defective paneth cell-mediated host defense in pediatric ileal Crohn’s disease. Am J Gastroenterol 105(2):452–459
Wehkamp J, Wang G, Kubler I, Nuding S, Gregorieff A, Schnabel A et al (2007) The Paneth cell alpha-defensin deficiency of ileal Crohn’s disease is linked to Wnt/Tcf-4. J Immunol 179(5):3109–3118
Uehara A, Yang S, Fujimoto Y, Fukase K, Kusumoto S, Shibata K et al (2005) Muramyldipeptide and diaminopimelic acid-containing desmuramylpeptides in combination with chemically synthesized Toll-like receptor agonists synergistically induced production of interleukin-8 in a NOD2- and NOD1-dependent manner, respectively, in human monocytic cells in culture. Cell Microbiol 7(1):53–61
van Heel DA, Ghosh S, Hunt KA, Mathew CG, Forbes A, Jewell DP et al (2005) Synergy between TLR9 and NOD2 innate immune responses is lost in genetic Crohn’s disease. Gut 54(11):1553–1557
Yang S, Tamai R, Akashi S, Takeuchi O, Akira S, Sugawara S et al (2001) Synergistic effect of muramyldipeptide with lipopolysaccharide or lipoteichoic acid to induce inflammatory cytokines in human monocytic cells in culture. Infect Immun 69(4):2045–2053
Kobayashi KS, Chamaillard M, Ogura Y, Henegariu O, Inohara N, Nunez G et al (2005) Nod2-dependent regulation of innate and adaptive immunity in the intestinal tract. Science 307(5710):731–734
van Beelen AJ, Zelinkova Z, Taanman-Kueter EW, Muller FJ, Hommes DW, Zaat SA et al (2007) Stimulation of the intracellular bacterial sensor NOD2 programs dendritic cells to promote interleukin-17 production in human memory T cells. Immunity 27(4):660–669
Bazzoni F, Rossato M, Fabbri M, Gaudiosi D, Mirolo M, Mori L et al (2009) Induction and regulatory function of miR-9 in human monocytes and neutrophils exposed to proinflammatory signals. Proc Natl Acad Sci USA 106(13):5282–5287
Ceppi M, Pereira PM, Dunand-Sauthier I, Barras E, Reith W, Santos MA et al (2009) MicroRNA-155 modulates the interleukin-1 signaling pathway in activated human monocyte-derived dendritic cells. Proc Natl Acad Sci USA 106(8):2735–2740
Sheedy FJ, Palsson-McDermott E, Hennessy EJ, Martin C, O’Leary JJ, Ruan Q et al (2010) Negative regulation of TLR4 via targeting of the proinflammatory tumor suppressor PDCD4 by the microRNA miR-21. Nat Immunol 11(2):141–147
Taganov KD, Boldin MP, Chang KJ, Baltimore D (2006) NF-kappaB-dependent induction of microRNA miR-146, an inhibitor targeted to signaling proteins of innate immune responses. Proc Natl Acad Sci USA 103(33):12481–12486
Tili E, Michaille JJ, Cimino A, Costinean S, Dumitru CD, Adair B et al (2007) Modulation of miR-155 and miR-125b levels following lipopolysaccharide/TNF-alpha stimulation and their possible roles in regulating the response to endotoxin shock. J Immunol 179(8):5082–5089
Brain O, Owens BM, Pichulik T, Allan P, Khatamzas E, Leslie A et al (2013) The intracellular sensor NOD2 induces microRNA-29 expression in human dendritic cells to limit IL-23 release. Immunity 39(3):521–536
Ma F, Xu S, Liu X, Zhang Q, Xu X, Liu M et al (2011) The microRNA miR-29 controls innate and adaptive immune responses to intracellular bacterial infection by targeting interferon-gamma. Nat Immunol 12(9):861–869
Steiner DF, Thomas MF, Hu JK, Yang Z, Babiarz JE, Allen CD et al (2011) MicroRNA-29 regulates T-box transcription factors and interferon-gamma production in helper T cells. Immunity 35(2):169–181
Becker C, Wirtz S, Blessing M, Pirhonen J, Strand D, Bechthold O et al (2003) Constitutive p40 promoter activation and IL-23 production in the terminal ileum mediated by dendritic cells. J Clin Invest 112(5):693–706
Wehkamp J, Harder J, Weichenthal M, Schwab M, Schaffeler E, Schlee M et al (2004) NOD2 (CARD15) mutations in Crohn’s disease are associated with diminished mucosal alpha-defensin expression. Gut 53(11):1658–1664
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Brain, O., Simmons, A. (2015). The Relationship Between miR-29, NOD2 and Crohn’s Disease. In: Greene, C. (eds) MicroRNAs and Other Non-Coding RNAs in Inflammation. Progress in Inflammation Research. Springer, Cham. https://doi.org/10.1007/978-3-319-13689-9_10
Download citation
DOI: https://doi.org/10.1007/978-3-319-13689-9_10
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-13688-2
Online ISBN: 978-3-319-13689-9
eBook Packages: MedicineMedicine (R0)