Abstract
Fungi and Oomycetes are notorious plant pathogens and use similar strategies to infect plants. The majority of plants, however, is not infected by pathogens as they recognize pathogen-associated molecular patterns (PAMPs) by pattern recognition receptors that mediate PAMP-triggered immunity (PTI) , a basal defense response effective against potential pathogens. Successful pathogens secrete effectors to suppress PTI and alter host plant physiology. In turn, plants have evolved immune receptors that recognize effectors, resulting in effector-triggered immunity (ETI) . ETI includes the hypersensitive response which is effective against biotrophic plant pathogens that require living cells to feed on. Other pathogens are hemi-biotrophic, which start infection as a biotroph, but after having colonized the host tissue can also feed on death tissue. Necrotrophic pathogens kill host tissue before they start to feed on it. Co-evolution between pathogens and their hosts had led to the development of numerous effectors produced by pathogens and corresponding resistance proteins in host plants, which has generated an arms race genetically described by the gene-for-gene concept. Resistance genes can now successfully be transferred to crop plants by classical breeding or as transgenes stapled into one cultivar.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
1 Fungal and Oomycetous Pathogens and Their Life Styles
Fungi and Oomycetes are uni- or multicellular eukaryotic heterotrophic organisms producing filamentous structures, designated hyphae, that show tip growth, except for yeasts that multiply by budding. Hyphae are usually 1–2 μm in diameter, but may reach more than 100 μm for some fungi. Cells of hyphae contain one, two or multiple nuclei often divided by septa. Until the 1990s, Oomycetes were considered true fungi, but based on genome analyses they are now classified among the Chromista (Chap. 39). Pathogenic Oomycetes are still treated as fungi because they have many properties in common, including their filamentous growth and their mode of plant infection. Fungal cell walls contain chitin , α- and β- glucans and (glyco)proteins, but no cellulose, while those of Oomycetes contain cellulose and glucans but lack chitin . Several fungi and Oomycetes are important for industrial production of enzymes, bread, cheese, alcohol and organic acids. As producers of antibiotics they can have important medical applications. On the other hand, many fungi and Oomycetes can cause disease to humans, animals and plants. Around 150,000 fungal species have now been described, which represent around 10 % all fungi estimated to be present on earth (Hawksworth 1991) . The most important fungal plant pathogens belong to (i) Ascomycetes , producing sexual spores (ascospores) in a sac-like structure, the ascus, and asexual spores (conidia), (ii) Basidiomycetes producing sexual spores (basidiospores) on a basidium, dikaryotic vegetative mycelium and asexual spores, (iii) Oomycetes producing sexual spores (oospores) and asexual spores (sporangia that can germinate directly or produce zoospores) (Agrios 2005) . In the remainder of this chapter I will jointly discuss fungi and Oomycetes as fungi, unless phenomena are specific to Oomycetes only. Fungi are one of the largest living organisms; the soil-borne fungus Armillaria ostoyae is estimated to be 2400 years old, covering 8.4 km2 of soil in Oregon, USA (Burdsall and Volk 2008) . Pathogenic fungi enter plants via natural openings (e.g. stomata) or penetrate directly by a penetration peg produced by an appressorium on a host cell (see Chap. 25). Many fungi produce haustoria in plant cells, specialized feeding organs for retrieval of nutrients. Extracellular pathogens grow as epiphytes on the outside of plants or in the apoplastic space between cells without producing haustoria. Ascomytous fungi are haploid for the major part of their life and produce haploid spores (1N). During the sexual stage haploid hyphae of Ascomycetes may fuse to produce a dikaryon inside the ascogenous hyphae where the nuclei soon fuse to produce a zygote (2N) that quickly divides meiotically to produce eight haploid ascospores per ascus. Oomycetes are diploid for the major part of their life and have a life cycle that is very similar to that of algae. During the sexual stage Oomycetes produce gametangia in which meiosis occurs, followed by fertilization and production of the diploid zygote, the oospore, that produces a sporangium that geminates directly or indirectly by producing zoospores. Basidiomycetes are dikaryotic (N + N) for the major part of their life. During the sexual stage in the basidium the paired nuclei fuse and form a zygote (2N) that quickly divides meiotically to produce four haploid basidiospores (Agrios 2005). In most fungi the asexual cycle repeats multiple times during the growth season and is most damaging to plants, whereas the sexual cycle usually occurs only once a year at the end of the growth season when host plants senesce and nutrients become limiting.
Diseases Caused by Oomycetes
The most important oomycetous plant pathogens comprising species belonging to the genera Pythium, Phytophthora, Peronospora and Plasmopara. The late blight disease of potato is an important oomycetous pathogen discussed in Chap. 39. Downy mildew of grape is another important disease caused by Plasmopara viticola (Fig. 10.1a), that almost completely destroyed the grape and wine industry in France soon after it was imported into Europe from the United States around 1875; the first fungicide, Bordeaux mixture, effective against P. viticola was discovered by accident in 1885. Downy mildews are obligate biotrophic pathogens that overwinter as oospores. Many resistance genes against downy mildews have been cloned (Takken and Goverse 2012) . They belong to the cytoplasmic nucleotide binding, leucine-rich repeat protein receptors (NB-LRRs) (discussed later).
Symptoms of economically important plant diseases caused by fungi and Oomycetes. a Plasmopara viticola, downy mildew of grape (http://www.biolib.cz/cz/image/id100651/). b Blumeria graminis f.sp. tritici, powdery mildew of wheat (http://www.apsnet.org/publications/imageresources/Pages/fi00189.aspx). c Nectria galligena, apple canker (http://www.downgardenservices.org.uk/cankerapp.htm). d Venturia inaequalis, apple scab (http://www.nature.com/news/us-regulation-misses-some-gm-crops-1.13580). e Ophiostoma ulmi, Dutch elm disease (http://commons.wikimedia.org/wiki/File:Ceratocystis_ulmi_1_beentree.jpg). f Melampsora lini, flax rust (http://www.sciencearchive.org.au/events/frontiers/frontiers2008/dodds.html). g Puccinia graminis f.sp. tritici, wheat leaf rust (http://www.mississippi-crops.com/2012/03/02/wheat-leaf-rust-and-stripe-rust-update-march-2-2012/). h Ustilago tritici, loose smut of wheat (http://www.bayercropscience.cl/soluciones/fichaproblema.asp?id=27). i Ustilago maydis, corn smut (http://commons.wikimedia.org/wiki/File:Maisbrand,_Maisbeulenbrand_(Ustilago_maydis)_-_hms(1).jpg)
Diseases Caused by Ascomycetes
Powdery mildews can infect virtually all plant species and are easy recognizable by the white powdery overlay on infected plant leaves (Fig. 10.1b). Powdery mildews are obligate biotrophic pathogens that can only infect epidermis cells from which they retrieve nutrients by multiple haustoria . Short conidiophores are produced on the plant surface producing chains of conidia. At the end of the growth season the fungus may produce cleistothecia with asci and ascospores. Many resistance genes against powdery mildews have been cloned. They belong to the cytoplasmic nucleotide binding NB-LRRs (Takken and Goverse 2012) (discussed later).
Tree Cankers
These are caused by ascomycetous fungi . Cankers generally begin at a wound from which they expand in all directions, but the host may survive the disease by producing callus tissue around the dead areas thereby limiting the canker. In subsequent years the fungus invades additional healthy tissue, and new concentric ridges of callus tissue are produced every year, resulting in a typical canker. Canker of apple is caused by Nectria galligena (Fig. 10.1c), one of the most important diseases of apple worldwide. Apple scab is caused by Venturia inaequalis (Fig. 10.1d), and occurs in areas with cool, moist springs and summers. Infected fruits develop scab lesions and the cuticle is ruptured at the margin of these lesions. The mycelium in living tissues is located between the cuticle and epidermal cells, and produces short conidiophores bearing conidia. The fungus is a hemi-biotroph starting infection as a biotroph, but at the end of the growth season, the mycelium grows through dead leaf tissues and produces pseudothecia with asci and ascospores. In spring, when pseudothecia become thoroughly wet, the asci forcibly discharge the ascospores that infect young apple leaves. Apple varieties resistant against apple scab exist, and the resistance genes encode typical receptor-like proteins (RLPs) (discussed later) .
Vascular Wilts
These are widespread, very destructive plant diseases and are caused by fungal pathogen residing in the xylem vessels of plants that may be clogged with mycelium, spores, or polysaccharides produced by the fungus and gels and gums produced by plant cells upon attack by the fungus. In some hosts, tyloses are produced by parenchyma cells adjoining xylem vessels. The disease can sometimes be controlled by using disease-resistant cultivars.
Dutch Elm Disease
This disease is caused by Ophiostoma ulmi (Fig. 10.1e). It is a very destructive wilt disease that affects all elm species. Usually trees that become infected in spring or early summer die quickly, whereas those infected in late summer are less seriously affected and may recover. Perithecia with asci and ascospores may be produced. The spread of the Dutch elm disease fungus depends on bark beetles belonging to the genus Scolytus that carry fungal spores from infected wood to healthy elm trees. The fungus overwinters in the bark of dead elm trees as mycelium or spores. Adult female beetles tunnel through the bark and lay eggs that, after hatching, develop into adult beetles that carry thousands of spores. The beetles feed on healthy elms and carry fungal spores that infect healthy xylem vessels. Control of Dutch elm disease depends primarily on removal and destruction of diseased elm trees. Inoculation of the xylem vessels with a spore suspension of the wilt fungus Verticillium dahliae offers protection through induced resistance (see Chap. 14), whereas protection can also be achieved by inoculating trees with particular strains of Pseudomonas bacteria. Resistance genes against wilt diseases have been cloned; they encoded either NB-LRRs or RLPs (discussed later).
Diseases Caused by Basidiomycetes
Basidiomycetes produce their sexual spores, basidiospores, on a club shaped basidium. Most Basidiomycetes are either saprobes or fungi causing wood decay including root and stem rots of trees. However, they also include plant pathogens that cause rust and smut diseases.
Rusts
Rusts are obligate biotrophic pathogens that have caused many famines in the history of many countries by destroying cereal crops. Presently the rust strain Ug99 causes a dramatic epidemic on wheat in eastern Africa from which it has spread into Arabia (Ward 2007). There are about 5000 species of rusts, mostly belonging to the genus Puccinia. They are very specialized pathogens attacking only particular genera or only certain species or even only some varieties of particular cereals. In the latter case they are called races that can be identified only by a set of differential varieties carrying different resistance genes. The gene-for-gene system proposed by Flor in the USA was based on research performed on the rust fungus of flax caused by Melampsora lini (Fig. 10.1f), (Flor 1971) . Rust fungi can produce up to five different types of spores: uredospores, teliospores and basidiospores (on wheat), spermatia and aeciospores (on barbary) (Agrios 2005) .
Stem Rust of Wheat
This disease is caused by Puccinia graminis f.sp. trititici (Fig. 10.1g), and affects wheat wherever it is grown. It attacks all aboveground parts of wheat and the alternate host barberry (Berberis vulgaris). Symptoms on wheat appear as pustules and uredia. The latter produce red-colored uredospores. Later in the season two-celled black teliospores are produced in telia. On barberry, spermagonia with spermatia and receptive hyphae are produced on the upper side of leaves, and on the lower side orange cup-like aecia are produced containing aeciospores. The most effective means of control of the rust pathogen is by growing resistant wheat varieties. The resistance proteins are typical NB-LRR receptor proteins (Takken and Goverse 2012) (discussed later).
Smut Fungi
More than 1200 species of smut fungi exist that occur throughout the world. Most smut fungi attack the ovaries of grains and grasses and develop in the kernels, which they destroy completely. Some smuts infect seeds or seedlings before they emerge from the soil, in which they grow systemically until they reach the inflorescence. Cells in affected tissues are either destroyed and replaced by black smut spores, or they are first stimulated to divide and enlarge to produce a swelling or gall that is subsequently destroyed and replaced by the black smut spores. Most smut fungi produce only teliospores and basidiospores. When haploid basidiospores germinate, the germ tubes fuse to produce dikaryotic infectious mycelium. Important smut fungi are Ustilago tritici (Fig. 10.1h), causing smut on wheat and Ustilago maydis (Fig. 10.1i), causing smut galls on corn. Together with Flor in the USA, Oort in the Netherlands proposed the gene-for-gene hypothesis based on research performed on Ustilago tritici and wheat (Oort 1944) .
Root Rots
Root rots of trees are often caused by species of Armillaria that occur worldwide and affect hundreds of species of trees. The pathogen Armillaria mellea is one of the most common fungi in forest soils. Diagnostic characteristics of Armillaria root rot appear at decayed areas in the bark, at the root-stem junction, and on the roots. White mycelial mats are formed between the bark and wood. Another characteristic sign of the disease is the formation of rhizomorphs or “shoe strings”, cordlike threads of mycelium 1–3 mm in diameter that can grow long distances and infect healthy trees (Agrios 2005) .
2 Characteristcis of Fungal and Oomycetous Plant Pathogens
These organisms can be obligate biotrophic, biotrophic, hemi-biotrophic or necrotrophic pathogens. Biotrophic pathogens thrive on living host cells. Some biotrophs cannot be cultured on synthetic media and are called obligate biotrophic pathogens like rusts, downy mildews and powdery mildews. However, many biotrophic pathogens grow on host plants under natural conditions but can still be cultured on synthetic media like the tomato pathogen Cladosporium fulvum (Joosten and De Wit 1999) . Hemi-biotrophic pathogens start infection as a biotroph, but later in the growth season they can live as a saprophyte on death host tissue. A defense response that is very efficient against obligate biotrophic and biotrophic pathogens is the hypersensitive response (HR) , death of a few plant cells at the site of infection. In contrast, necrotrophic fungal pathogens kill host tissue before they retrieve nutrients, and the HR is not effective against them.
Extracellular and Intracellular Pathogens
Many fungi live on plants as epiphytes of which some have developed into pathogens. They usually enter plants through stomata and thrive in the apoplastic space surrounding cells without producing haustoria. Most extracellular pathogens are slow growers and show long latent periods as the apoplast often contains antimicrobial compounds and antimicrobial enzymes, and is poor in nutrients. Intracellular pathogen often produce haustoria .
3 Fungal and Oomycetous Infection Strategies and Host Defense Mechanisms
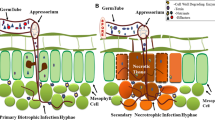
Infection Strategies
Infection strategies employed by fungal pathogens depend strongly on where they thrive in their host plants. The cell wall is a major obstacle for plant pathogens which contains physical and chemical barriers like the plant waxy cuticle, (lignified) cell walls and antimicrobial metabolites and proteins (Balmer et al. 2013) . Many fungal genomes have been sequenced showing that pathogenic fungi contain many different classes of cell-wall degrading enzymes. The expression of these genes is highly up-regulated during infection by necrotrophic plant pathogens (Zhao et al. 2013) . Biotrophic pathogens often contain similar numbers of genes encoding cell-wall degrading enzymes as necrotrophs, but they are often lowly expressed by these pathogens or only at specific sites and phases of infection. Necrotrophic fungi often also produce secondary metabolites that are toxic to plant cells thereby facilitating their necrotrophic lifestyle, while those metabolites are often absent in (obligate) biotrophic pathogens. For example, the powdery mildew Blumeria graminis contains hardly any genes encoding secondary metabolites (Spanu 2012) and in the tomato leaf pathogen C. fulvum these genes are down-regulated during infection (De Wit et al. 2012) . Apart from producing enzymes that enable pathogens to degrade plant cell walls and to retrieve the released mono/oligosaccharides, fungi also need to protect themselves against antifungal proteins that are present in plant-cell walls and the apoplast by detoxifying enzymes like tomatinase produced by Cladosporium fulvum that detoxifies the toxic saponin, α-tomatine occurring at high concentrations in tomato (Okmen et al. 2013) .
Basal Defense Strategies
Plants have developed sophisticated defense strategies to recognize pathogens and to defend themselves against fungal pathogens. All plants can recognize pathogen-associated molecular patterns (PAMPs) , like chitin from fungi, by pattern recognition receptors (PRRs) that mediate PAMP-triggered immunity (PTI), a response that protects plants against potential microbial pathogens (Jones and Dangl 2006; Liebrand et al. 2014) . PRRs are extracellular LRR-containing receptor-like transmembrane proteins with a cytoplasmic kinase signaling domain known as RLKs. They mediate PAMP-triggered basal structural and chemical defense responses including callose deposition, accumulation of reactive oxygen species (ROS), cell wall enforcements and accumulation of pathogenesis-related (PR) proteins including chitinases, proteases and glucanases.
Effector-Triggered Susceptibility
Although plants have developed basal defense strategies against microbes, successful pathogens have found ways to overcome basal defense responses. They can suppress PTI by secreting different types of effectors that target various components of PTI (Stergiopoulos and de Wit 2009) . By suppressing PTI they cause effector-triggered susceptibility (ETS). Various types of effectors have been described in various pathogenic fungi (Stergiopoulos and De Wit 2009) . Overall they manipulate host defenses and host physiology to facilitate virulence in various ways. Here I discuss the intrinsic functions of two Cladosporium fulvum effectors that can serve as an example of many other fungal effectors. C. fulvum secretes the Avr2 effector protein which inhibits plant cysteine proteases including Rcr3pim (required for C. fulvum resistance 3) (Rooney et al. 2005) . Heterologous expression of Avr2 in tomato and Arabidopsis resulted in increased susceptibility to different fungal pathogens including Botrytis cinerea and Verticillium dahliae (Van Esse et al. 2008) . Mutation studies on Rcr3pim showed it also to be active against Phytophthora infestans, which secretes EPIC1 and EPIC2B proteins that also bind and inhibit Rcr3pim. Also the root parasitic nematode Globodera rostochiensis secretes a venom allergen-like effector protein, Gr-VAP1, that targets and inhibits Rcr3 perturbing its active site (Lozano-Torres et al. 2012) .
C. fulvum also secretes the cysteine-rich Avr4 protein which is a chitin-binding lectin that protects fungal cell walls against plant chitinases, providing a defensive role during infection (Van den Burg et al. 2006) . In addition, silencing of the Avr4 gene in C. fulvum reduces virulence on tomato plants (Van Esse et al. 2007) . Protection against plant chitinases is important for virulence of most plant pathogenic fungi because chitin is a basic component of fungal cell walls. Indeed homologs of Avr4 have been identified in several dothideomycetous fungi, including Mycosphaerella fijiensis and Dothistroma septosporum (Stergiopoulos et al. 2010) . Interestingly many fungi, including C. fulvum, also secrete proteins that contain LysM carbohydrate-binding domain proteins (Kombrink and Thomma 2013) . The LysM protein Ecp6 of C. fulvum also binds to chitin, but does not protect the fungus against basic plant chitinases. Ecp6 mainly binds small chitin fragments (PAMPs) that are released from the fungal cell wall in planta and prevents them from being recognized by chitin receptors present in plants to induce PTI (De Jonge et al. 2010) . Ecp6 is a virulence factor because silencing of Ecp6 results in reduced virulence (Kombrink and Thomma 2013) . Many homologs of Ecp6 are identified in various fungal species (De Jonge and Thomma 2009; Kombrink and Thomma 2013) , suggesting an important function in preventing chitin-triggered immunity in many plant-fungus interactions. Recently, it was shown that Mycosphaerella graminicola secretes three LysM effectors called Mg1LysM, Mg3LysM and MgxLysM (Marshall et al. 2011) of which both Mg1LysM and Mg3LysM bind chitin, whereas only Mg3LysM prevents chitin-triggered immunity. However, only the Mg3LysM deletion mutant showed significantly reduced virulence . Magnaporthe oryzae LysM effector Slp1 also binds and scavenges small chitin fragments. In rice, the chitin elicitor binding protein, CEBiP, recognizes chitin and activates PTI. Indeed it was found that Slp1 prevents chitin-triggered immunity by competing with the CEBiP receptor for chitin binding (Mentlak et al. 2012) .
Effector-Triggered Immunity
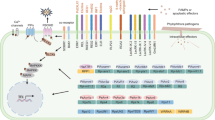
In turn, plants have evolved sophisticated ways to recognize and respond to effectors. In addition to PRRs that recognize PAMPs, plants have developed immune receptors that recognize effectors or host plant targets manipulated by effectors resulting in effector-triggered immunity (ETI) (Jones and Dangl 2006) . One of the most typical characteristics of ETI is the HR. At the host species and cultivar level, co-evolution between hosts and their pathogens has caused an arms race that has led to the development of numerous novel effectors and corresponding resistance proteins, which are described by the gene-for-gene concept (De Wit et al. 2009) (Fig. 10.2a, 10.2b, 10.2c, 10.2d) .
Schematic overview of an evolutionary scenario for adaptation of the leaf mould pathogen Cladosporium fulvum to tomato. a Chitin fragments (pattern-associated molecular pattern; PAMP) are recognized by the tomato chitin receptor (SlCERK) that triggers PAMP-triggered immunity (PTI) providing basal defense against the pathogen. b To become pathogenic, C. fulvum secretes the Ecp6 effector that scavenges chitin fragments to prevent PTI. c C. fulvum secretes additional effectors to increase virulence including Avr2 inhibiting apoplastic cysteine protease Rcr3, and Avr4 protecting chitin present in the cell wall of the fungus against tomato chitinases. d Tomato responds by developing RLP immune receptors Cf-Ecp6, Cf-2 and Cf-4 to recognize Ecp6, Avr2 and Avr4, respectively, and to mediate effector-triggered immunity (ETI) leading to the hypersensitive response (HR) and resistance against fungal strains secreting these effectors. C. fulvum is supposed to produce numerous effectors and tomato a comparable number of immune receptors
4 Resistance Against Fungi and Oomycetes
The cloning of effector genes speeded up the cloning of the matching resistance genes that encode either cell surface-localized receptor-like proteins known as RLPs (Liebrand et al. 2014) or cytoplasmic nucleotide binding, leucine-rich repeat (NB-LRR) or NLR proteins (Takken and Goverse 2012) . RLPs are integral plant membrane proteins containing an extracellular leucine-rich repeat or LRR domain, a membrane spanning domain, and a short cytoplasmic tail without signaling domain. RLPs recognize effectors of extracellular fungal pathogens and mediate ETI. However, cytoplasmic pathogens exploit the cytoplasm of plant cells by injecting effectors into host cells that interact with cytoplasmic targets to suppress PTI. The cytoplasmic effectors are usually recognized by cytoplasmic NLR immune receptors. Pathogens can develop mutations in effectors or simply loose them to escape recognition by RLP and NRL immune receptors or develop new effectors for compensation. Some immune receptors can work in concert in receptor complexes active against more than one pathogen (Macho and Zipfel 2014) . Defense systems of plants are able to respond to different types of PAMPs and effectors, but they cannot always be distinguished easily and downstream defense pathways in plants activated during PTI and ETI often overlap and operate against a broad spectrum of pathogens (Thomma et al. 2011) . These responses include the generation of reactive oxygen species (ROS), antimicrobial phytoalexins, chitinases, glucanases, proteases, and often the HR. Many immune receptors encoded by resistance genes have been cloned in the last decade. They can now be introduced in many copies by breeders in crops by classical breeding or by cis/transgenesis (Zhu et al. 2012) . It is important to introduce multiple immune receptor genes in plants against multiple pathogen effectors in order to obtain durable plant resistance (Brunner et al. 2010; Vleeshouwers and Oliver 2014) . Overcoming multiple immune receptors by a pathogen by modification or loss of effectors is expected to cause a fitness cost and decrease in virulence.
References
Agrios GN (2005) Plant pathology handbook, 5th edn. Elsevier, Amsterdam, 922 p
Balmer D, Planchamp C, Mauch-Mani B (2013) On the move: induced resistance in monocots. J Exp Bot 64:1249–1261
Brunner S, Hurni S, Streckeisen P et al (2010) Intragenic allele pyramiding combines different specificities of wheat Pm3 resistance alleles. Plant J 64:433–445
Burdsall HH, Volk TJ (2008) Armillaria solidipes, an older name for the fungus called Armillaria ostoyae. N Am Fungi 3:261–267
De Jonge R, Thomma BPHJ (2009) Fungal LysM effectors: extinguishers of host immunity? Trends Microbiol 17:151–157
De Jonge R, Van Esse HP, Kombrink A et al (2010) Conserved fungal LysM effector Ecp6 prevents chitin-triggered immunity in plants. Science 329:953–955
De Wit PJGM, Mehrabi R, Van den Burg HA et al (2009) Fungal effector proteins: past, present and future. Mol Plant Pathol 10:735–747
De Wit PJGM, Van der Burgt A, Ökmen B et al (2012) The genomes of the fungal plant pathogens Cladosporium fulvum and Dothistroma septosporum reveal adaptation to different hosts and lifestyles but also signatures of common ancestry. PLoS Genet. doi:10.1371/journal.pgen.1003088
Flor HH (1971) Current status of the gene-for-gene concept. Annu Rev Phytopathol 9:275–296
Hawksworth DL (1991) The fungal dimension of biodiversity -magnitude, significance, and conservation. Mycol Res 95:641–655
Jones JDG, Dangl JL (2006) The plant immune system. Nature 444:323–329
Joosten MHAJ, De Wit PJGM (1999) The tomato-Cladosporium fulvum interaction: a versatile experimental system to study plant-pathogen interactions. Annu Rev Phytopathol 37:335–367
Kombrink A, Thomma BPHJ (2013) LysM effectors: secreted proteins supporting fungal life. Plos Pathog 9:e1003769. doi:10.1371/journal.ppat.1003769
Liebrand TWH, van den Burg HA, Joosten MHAJ (2014) Two for all: receptor-associated kinases SOBIR1 and BAK1. Trends Plant Sci 19:123–132
Lozano-Torres JL, Wilbers RHP, Gawronski P et al (2012) Dual disease resistance mediated by the immune receptor Cf-2 in tomato requires a common virulence target of a fungus and a nematode. Proc Natl Acad Sci U S A 109:10119–10124
Macho AP, Zipfel C (2014) Plant PRRs and the activation of innate immune signaling. Mol Cell 54:263–272. doi:10.1016/j.molcel.2014.03.028
Marshall R, Kombrink A, Motteram J et al (2011) Analysis of two in planta expressed LysM effector homologs from the fungus Mycosphaerella graminicola reveals novel functional properties and varying contributions to virulence on wheat. Plant Physiol 156:756–769
Mentlak TA, Kombrink A, Shinya T et al (2012) Effector-mediated suppression of chitin-triggered immunity by Magnaporthe oryzae is necessary for rice blast disease. Plant Cell 24:322–335
Okmen B, Etalo DW, Joosten MHAJ et al (2013) Detoxification of α-tomatine by Cladosporium fulvum is required for full virulence on tomato. New Phytol 198:1203–1214
Oort AJP (1944) Onderzoekingen over stuifbrand. II. Overgevoeligheid van tarwe voor stuifbrand (Ustilago tritici) with a summary: hypersensitiviness of wheat to loose smut. Tijdschr Planteziekten 50:73–106
Rooney HCE, Van’t Klooster JW, Van der Hoorn RAL et al (2005) Cladosporium Avr2 inhibits tomato Rcr3 protease required for Cf-2-dependent disease resistance. Science 308:1783–1786
Spanu PD (2012) The genomics of obligate (and nonobligate) biotrophs. Annu Rev Phytopathol 50:91–109
Stergiopoulos I, De Wit PJGM (2009) Fungal effector proteins. Annu Rev Phytopathol 47:233–263
Stergiopoulos I, van den Burg HA, Okmen B et al (2010) Tomato Cf resistance proteins mediate recognition of cognate homologous effectors from fungi pathogenic on dicots and monocots. Proc Natl Acad Sci U S A 107:7610–7615
Takken FLW, Goverse A (2012) How to build a pathogen detector: structural basis of NB-LRR function. Curr Opin Plant Biol 15:375–384
Thomma B, Nurnberger T, Joosten M (2011) Of PAMPs and effectors: the blurred PTI-ETI dichotomy. Plant Cell 23:4–15
Van den Burg HA, Harrison SJ, Joosten MHAJ et al (2006) Cladosporium fulvum Avr4 protects fungal cell walls against hydrolysis by plant chitinases accumulating during infection. Mol Plant Microbe Interact 19:1420–1430
Van Esse HP, Bolton MD, Stergiopoulos I et al (2007) The chitin-binding Cladosporium fulvum effector protein Avr4 is a virulence factor. Mol Plant Microbe Interact 20:1092–1101
Van Esse HP, Van’t Klooster JW, Bolton MD et al (2008) The Cladosporium fulvum virulence protein Avr2 inhibits host proteases required for basal defense. Plant Cell 20:1948–1963
Vleeshouwers VGAA, Oliver RP (2014) Effectors as tools in disease resistance breeding against biotrophic, hemibiotrophic, and necrotrophic plant pathogens. Mol Plant Microbe Interact 27:196–206
Ward R (2007) The global threat posed by Ug99. Phytopathol 97: S136
Zhao ZT, Liu HQ, Wang CF et al (2013) Comparative analysis of fungal genomes reveals different plant cell wall degrading capacity in fungi. BMC Genomics 14:274. doi:10.1186/1471-2164-14-274
Zhu SX, Li Y, Vossen JH et al (2012) Functional stacking of three resistance genes against Phytophthora infestans in potato. Transgenic Res 21:89–99
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer International Publishing Switzerland
About this chapter
Cite this chapter
de Wit, P. (2015). Plant Pathogenic Fungi and Oomycetes. In: Lugtenberg, B. (eds) Principles of Plant-Microbe Interactions. Springer, Cham. https://doi.org/10.1007/978-3-319-08575-3_10
Download citation
DOI: https://doi.org/10.1007/978-3-319-08575-3_10
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-08574-6
Online ISBN: 978-3-319-08575-3
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)