Abstract
Data-independent acquisition (DIA) implemented in a method called MSE can be performed in a massively parallel, time-based schedule rather than by sampling masses sequentially in shotgun proteomics. In MSE alternating low and high energy spectra are collected across the full mass range. This approach has been very successful and stimulated the development of variants modeled after the MSE protocol, but over narrower mass ranges. The massively parallel MSE and other DIA methodologies have enabled effective label-free quantitation methods that have been applied to a wide variety of samples including affinity pulldowns and studies of cells, tissues, and clinical samples. Another complementary technology matches accurate mass and retention times of precursor ions across multiple chromatographic runs. This further enhances the impact of MSE in counteracting the stochastic nature of mass spectrometry as applied in proteomics. Otherwise significant amounts of data in typical large-scale protein profiling experiments are missing. A variety of software packages perform this function similar in concept to matching of accurate mass tags. Another enhancement of this method involves a variation of MSE coupled with traveling wave ion mobility spectrometry to provide separations of peptides based on cross-sectional area and shape in addition to mass/charge (m/z) ratio. Such a two-dimensional separation in the gas phase considerably increases protein coverage as well as typically a doubling of the number of proteins detected. These developments along with advances in ultrahigh pressure liquid chromatography have resulted in the evolution of a robust and versatile platform for label-free protein profiling.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Graphic Processing Unit
- Shotgun Proteomics
- Full Mass Range
- Retention Time Information
- Isotopic Label Approach
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
4.1 MSE and Other Data-Independent Acquisition Strategies
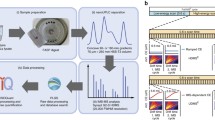
Shotgun proteomics is a strategy with broad applicability, and is based on digestion of proteins with proteolytic enzymes and analysis of the resulting peptides by liquid chromatography–mass spectrometry (LC/MS). The method of choice for data collection has long been a data-dependent acquisition (DDA) method in which acquisition parameters are modified in real time by selecting a narrow mass window in a quadrupole analyzer to allow precursor ions to pass through for fragmentation after an initial survey scan [1]. This was an important enhancement to the initial approach of using stored mass and retention time information from previous liquid chromatography runs [2]. More recently, a strategy has been proposed [3, 4] and finally implemented [5, 6] using data-independent acquisition (DIA) in a method called MSE to perform data collection in a massively parallel, time-based schedule rather than attempting to sample masses sequentially. MSE has been extensively developed as a method in which alternating low and high energy scans are recorded across the full mass range and was implemented on quadrupole time-of-flight mass spectrometers from Waters Corporation [5, 6]. The original approach has been very successful and has stimulated the development of variants modeled after the MSE protocol. These include sequential proteomic method precursor acquisition independent from ion count (PAcIFIC) [7], windowed data-independent acquisition of total high-resolution (SWATH) [8], all-ion fragmentation (AIF) [9], and multiplexed data-independent acquisition (MSX) [10]. These recent adaptations can be utilized on other instrument platforms, although only MSE is unique in that it covers the entire mass range in each scan. Our group has used MSE effectively on a routine basis for cells, tissues, and affinity preparations in a label-free approach to large-scale protein profiling [11–15]. The validity, power, and effectiveness of MSE are now firmly established and extensively validated independently by many groups [16–18]. One example is our work on human somatic stem cells, where we were able to use this technique to demonstrate clear differential expression of proteins such as aldose reductase and many other proteins (Fig. 4.1) which responded to the combination of growth factor priming and increased osmotic pressure [13]. The roles of some of these proteins are characteristic of the physiological states studied [13]. In that experiment, 5′-nucleotidase and transgelin were detected as differentially expressed, and have been previously linked to cell differentiation state. Data-independent label-free profiling was demonstrated in that work to be a useful tool in characterizing cellular responses to treatment regimes, and as an aid to optimization of cell priming protocols for cartilage tissue engineering.
Example of expression data for selected proteins in agglomerative hierarchical cluster of Z-score transformed intensity data as processed by the Rosetta Elucidator program. These proteins have at least 1.5-fold response to the treatment, and are represented by at least two peptides for both identification and quantification at ratio P-values as calculated by Elucidator. Mean relative abundance of each protein in control and treated as well as abundance ratio is listed. Cluster coloration indicates protein abundance in the sample (red indicates higher abundance; green indicates lower abundance and black equal abundance). Reprinted with permission from Oswald et al. [13]. Copyright (2011) American Chemical Society
4.2 DIA Strategies Enhanced by Accurate Mass and Retention Time Matching Across Multiple Chromatographic Runs
Our work on adipose-derived stem cells [13] uses the Elucidator Protein Expression Data Analysis System from Rosetta (and Ceiba Solutions), a commercial program to match accurate mass and retention time intensities of precursor ions across multiple chromatographic runs. This enabled measurement of the intensities of precursor peptides even if fragmentation is not particularly successful for a particular peptide in a particular chromatographic run. This approach to a large extent counters the limitation imposed by the stochastic nature of mass spectrometry that otherwise results in a large amounts of missing data in large-scale experiments. A similar strategy is used by the TransOmics Informatics for Progenesis QI (TOIP) from Waters Corporation [19], a program also specifically enhanced to take advantage of ion mobility separations (see below). Coupling mass and retention time matching programs such as these with DIA technologies including MSE provides a particularly powerful label-free platform that enables large and complex experiments otherwise difficult to conduct by other methods.
These software approaches are enabled by new faster instruments capable of data-independent scanning, but owe their conceptual inspiration to the concept of an accurate mass tags database originally conceived by Smith’s group [20].
4.3 Enhancement of MSE: Coupling to Traveling Wave ion Mobility Spectrometry (TWIMS)
A variation of MSE coupled with traveling wave ion mobility spectrometry (TWIMS) provides the capability to perform separations of peptides based on cross-sectional area and shape in addition to their mass/charge (m/z) ratio. The use of TWIMS for shotgun proteomics is now well established and has been validated independently by a number of groups [21–23]. A two-dimensional separation is achieved in the gas phase, providing considerably increased protein coverage, typically a doubling of the number of proteins detected, a significant advantage [12, 24]. The complex TWIMS data requires a powerful computing platform, typically with a graphics processing unit (GPU)-equipped computer with 448 cores or more [12, 24]. The TWIMS principle is illustrated in Fig. 4.2 where integrating ion mobility drift time enhances identification of peptides [25]. In a study of a bacteriophage virion proteome [26], MSE with TWIMS was consistently more effective than more conventionally employed DDA method. Current versions of the ProteinLynx Global Server commercial program [5, 6] (Waters Corp.) have been enabled to process TWIMS data.
Diagram illustrating the role of ion mobility spectrometry in addressing the challenge of peptides with both the same elution time in liquid chromatography (co-eluting) and having the same mass (isobaric). Without ion mobility, fragmentation of isobaric peptides in the collision cell results in a chimeric spectrum which is difficult to interpret. When ion mobility is activated, peptides are separated in the mobility cell on the basis of their drift time. The end result is cleaner spectra with reduced chimeracy [26]. Reprinted from Journal of Virological Methods, Vol 195, Moran, Deborah; Cross, Trevor; Brown, Lewis M.; Colligan, Ryan M; Dunbar, David, Data-independent acquisition (MSE) with ion mobility provides a systematic method for analysis of a bacteriophage structural proteome, pp. 9–17, 2014, with permission from Elsevier
We have applied these techniques in many other systems [15, 26] including patient samples [27]. Such analyses include fold-change and p-value determinations, providing an unbiased view of phenotype or biological responses to experimental treatments. An example is shown in Fig. 4.3 (mouse hippocampus) [12]. This figure represents fragmentation spectra collected with and without TWIMS activated. It can be clearly seen from this example that the interfering peaks from other peptides (derived from other proteins, magenta) and unassigned peaks (grey) are largely separated out by TWIMS. These sorts of comparisons can be made in data where spectra with or without TWIMS can be readily compared for the same peptide.
Fragment ion spectra of peptide ADQLADESLESTR with TWIMS off and TWIMS on. Purple and grey peaks represent interferences and noise, respectively [12]
With this technique it is possible to quantify large numbers of proteins and generate abundances (examples in Table 4.1) for each LC/MS/MS run and for each biological replicate, and calculate means and standard deviations with coefficient of variation as low as 10–20 % for replicate analyses [12]. Large-scale experiments can be done on a routine basis with recording of many thousands of protein abundance values for multiple biological and technical replicates.
MSE combined with TWIMS is being applied to a remarkable variety of practical biological problems including the interactome of the RNA-binding protein RALY [28], analysis of chaperonins and biosynthetic enzymes in the unculturable bacterial endosymbiont Blochmannia, [29] and quantitative analysis of human embryonic kidney cells proteome following sialic acid overproduction [30].
In another practical example of this technique, Drosophila melanogaster was evaluated for the role of dMyc in the larval fat body (Table 4.2). The fat body is a tissue that functions as a sensor of circulating nutrients to control the release of Drosophila insulin-like peptides (Dilps) influencing growth and development. Using MSE and TWIMS, it was demonstrated that dMyc affected expression of hexokinase C and pyruvate kinase, key regulators of glycolysis, as well as of stearoyl-CoA desaturase (Desat1). Desat1 is an enzyme that is necessary for monosaturation and production of fatty acids, and its reduction affects dMyc and the ability to induce fat storage and resistance to animal survival.
These techniques can also be applied very effectively to chemoproteomic affinity experiments. For example, we have identified a protein target of a small molecule (RSL3) that is an inducer of a novel form of cell death (ferroptosis) and a potential novel anticancer agent [24]. In that work we also used DIA (MSE) with TWIMS. A total of 1,353 proteins were detected in 27 HDMSE chromatograms. Analysis focused on 979 proteins with identification and quantitation supported by two or more unique peptides. Most detected proteins were not differentially bound to fluorescein-RSL3 beads compared to controls. Only one protein, glutathione peroxidase 4 (GPX4_HUMAN), was significantly enriched (P < 0.01) and had the highest fold-change (26-fold in the affinity preparation compared to the inactive analogue and 13-fold compared to the control of preincubation with free RSL3). These studies identified GPX4 as an essential regulator of ferroptotic cancer cell death using data-independent profiling with TWIMS and these results were confirmed by an extensive series of corroborative experiments [24]. Isotopic clusters of an example peptide from GPX4 are given in Fig. 4.4
3D visualization of an isotopic cluster of an example peptide ILAFPCNQFGK from GPX4 from an affinity preparation and two controls. Cell lysates were prepared from cells treated with active probe, inactive probe, or active probe in the presence of competitor. Reprinted from Cell Vol. 156, Yang, Wan S.; SriRamaratnam, R.; Welsch, Matthew E.; Shimada, K.; Skouta, R.; Viswanathan, Vasanthi S.; Cheah, Jaime H.; Clemons, Paul A.; Shamji, Alykhan F.; Clish, Clary B.; Brown, Lewis M.; Girotti, Albert W.; Cornish, Virginia W.; Schreiber, Stuart L.; Stockwell, Brent R., Regulation of Ferroptotic Cancer Cell Death by GPX4, pp., 317–331, 2014, with permission from Elsevier
4.4 Conclusions
In mass spectrometry-based proteomics, there has been an evolution from initial pioneering data-dependent approaches where slower instruments could not collect enough precursor intensity data to allow robust label-free quantitation. The massively parallel MSE and other DIA methodologies have enabled label-free quantitation and have been applied to a wide variety of sample types from cells to tissues, and have been applied to viral, microbial, plant, animal, and patient samples. This evolution has continued with the introduction of software to match accurate mass and retention time data in large experiments, increasing sophistication of ultrapressure liquid chromatography separations and finally through orthogonal separation with technologies such as TWIMS that significantly increase the number of proteins detected in an experiment. This pipeline provides exceptional flexibility for large and complex experiments and complements isotopic labeling approaches described in other chapters of this volume.
References
Stahl DC, Swiderek KM, Davis MT, Lee TD (1996) Data-controlled automation of liquid chromatography/tandem mass spectrometry analysis of peptide mixtures. J Am Soc Mass Spectrom 7(6):532–540
Eng JK, McCormack AL, Yates JR III (1994) An approach to correlate tandem mass spectral data of peptides with amino acid sequences in a protein database. J Am Soc Mass Spectrom 5(11):976–989
Masselon C, Anderson GA, Harkewicz R, Bruce JE, Pasa-Tolic L, Smith RD (2000) Accurate mass multiplexed tandem mass spectrometry for high-throughput polypeptide identification from mixtures. Anal Chem 72(8):1918–1924
Purvine S, Eppel J-T, Yi EC, Goodlett DR (2003) Shotgun collision-induced dissociation of peptides using a time of flight mass analyzer. Proteomics 3(6):847–850
Silva JC, Denny R, Dorschel C, Gorenstein MV, Li GZ, Richardson K, Wall D, Geromanos SJ (2006) Simultaneous qualitative and quantitative analysis of the Escherichia coli proteome—a sweet tale. Mol Cell Proteomics 5(4):589–607
Silva JC, Gorenstein MV, Li GZ, Vissers JPC, Geromanos SJ (2006) Absolute quantification of proteins by LCMSE—a virtue of parallel MS acquisition. Mol Cell Proteomics 5(1):144–156
Panchaud A, Scherl A, Shaffer SA, von Haller PD, Kulasekara HD, Miller SI, Goodlett DR (2009) Precursor acquisition independent from ion count: how to dive deeper into the proteomics ocean. Anal Chem 81(15):6481–6488
Gillet LC, Navarro P, Tate S, Rost H, Selevsek N, Reiter L, Bonner R, Aebersold R (2012) Targeted data extraction of the MS/MS spectra generated by data-independent acquisition: a new concept for consistent and accurate proteome analysis. Mol Cell Proteomics 11(6): O111.016717.doi: 10.1074/mcp.O111.016717
Geiger T, Cox J, Mann M (2010) Proteomics on an Orbitrap benchtop mass spectrometer using all-ion fragmentation. Mol Cell Proteomics 9(10):2252–2261
Egertson JD, Kuehn A, Merrihew GE, Bateman NW, MacLean BX, Ting YS, Canterbury JD, Marsh DM, Kellmann M, Zabrouskov V, Wu CC, MacCoss MJ (2013) Multiplexed MS/MS for improved data-independent acquisition. Nat Methods 10(8):744–746
Brown LM, Boël G, Gangadhar N, Stockwell BR, Firestein S, Hunt JF (2009) Label-free proteomics with MSE: applications to protein functional biology and the biology of adult stem cells. In: Proceedings of the 57th annual conference on mass spectrometry and allied topics, May 31—June 4, 2009, Philadelphia, PA
Brown LM, Rayman JB, Gornstein A, Kandel ER (2011) Quantitative label-free protein profiling of mouse hippocampus. In: Proceedings of the 59th annual conference on mass spectrometry and allied topics, June 5–9, 2011, American Society for Mass Spectrometry, Denver, CO
Oswald ES, Brown LM, Bulinski JC, Hung CT (2011) Label-free protein profiling of adipose-derived human stem cells under hyperosmotic treatment. J Proteome Res 10(7):3050–3059
Oswald ES, Brown LM, Burke M, Schwartz S, Bulinski JC, Hung CT (2010) Proteomics of adipose-derived human stem cells under hyperosmotic treatment. In: Proceedings of the 58th annual conference on mass spectrometry and allied topics, May 23–27, 2010, Salt Lake City, UT
Parisi F, Riccardo S, Zola S, Lora C, Grifoni D, Brown LM, Bellosta P (2013) dMyc expression in the fat body affects DILP2 release and increases the expression of the fat desaturase Desat1 resulting in organismal growth. Dev Biol 379(1):64–75
Cyr DD, Lucas JE, Thompson JW, Patel K, Clark PJ, Thompson A, Tillmann HL, McHutchison JG, Moseley MA, McCarthy JJ (2011) Characterization of serum proteins associated with IL28B genotype among patients with chronic hepatitis C. PLoS One 6(7):e21854s
Levin Y, Hradetzky E, Bahn S (2011) Quantification of proteins using data-independent analysis (MS(E)) in simple and complex samples: a systematic evaluation. Proteomics 11(16):3273–3287
Reidel B, Thompson JW, Farsiu S, Moseley MA, Skiba NP, Arshavsky VY (2011) Proteomic profiling of a layered tissue reveals unique glycolytic specializations of photoreceptor cells. Mol Cell Proteomics 10(3):M110.002469
Brown LM, Tang G, Colligan RM, Sulzer D (2013) Data-independent acquisition with ion mobility (HDMSE) for analysis of the mouse synaptosome proteome. In: Proceedings of the 61st annual conference on mass spectrometry and allied topics, American Society for Mass Spectrometry, Minneapolis, MN
Lipton MS, Pasa-Tolic’ L, Anderson GA, Anderson DJ, Auberry DL, Battista JR, Daly MJ, Fredrickson J, Hixson KK, Kostandarithes H, Masselon C, Markillie LM, Moore RJ, Romine MF, Shen Y, Stritmatter E, Tolic’ N, Udseth HR, Venkateswaran A, Wong K-K, Zhao R, Smith RD (2002) Global analysis of the Deinococcus radiodurans proteome by using accurate mass tags. Proc Natl Acad Sci 99(17):11049–11054
Ge H, Wu GS, Moore R, Rao N, Lee T, Gugiu G (2011) Label free quantitative proteomics of formalin fixed embedded (FFPE) tissue sections from temporal giant cell arteritis patients. In: Proceedings of the 59th annual conference on mass spectrometry and allied topics, June 5–9, 2011, American Society for Mass Spectrometry, Denver, CO
Ibrahim Y, Danielson WF, Prior D, Baker E, Kurulugama R, Anderson G, Seim T, Meka D, Smith RD, Belov M (2011) Performance of a new sensitive LC-IMS-QTOF platform for proteomics measurements. In: Proceedings of the 59th annual conference on mass spectrometry and allied topics, June 5–9, 2011, American Society for Mass Spectrometry, Denver, CO
Thompson JW, Geromanos S, Stapels MD, Dubois LD, Fadgen K, Chepanoske C, Soderblom EJ, Moseley MA (2011) Ion mobility-assisted data independent analysis with inter-analysis alignment provides improved depth of proteome coverage. In: Proceedings of the 59th annual conference on mass spectrometry and allied topics, June 5–9, 2011, American Society for Mass Spectrometry, Denver, CO
Yang WS, SriRamaratnam R, Welsch ME, Shimada K, Skouta R, Viswanathan VS, Cheah JH, Clemons PA, Shamji AF, Clish CB, Brown LM, Girotti AW, Cornish VW, Schreiber SL, Stockwell BR (2014) Regulation of ferroptotic cancer cell death by GPX4. Cell 156(1–2):317–331
Valentine SJ, Ewing MA, Dilger JM, Glover MS, Geromanos S, Hughes C, Clemmer DE (2011) Using ion mobility data to improve peptide identification: intrinsic amino acid size parameters. J Proteome Res 10(5):2318–2329
Moran D, Cross T, Brown LM, Colligan RM, Dunbar D (2014) Data-independent acquisition (MSE) with ion mobility provides a systematic method for analysis of a bacteriophage structural proteome. J Virol Methods 195:9–17
Yang J, Li Y, Chan L, Tsai Y-T, Wu W-H, Nguyen HV, Hsu C-W, Li X, Brown LM, Egli D, Sparrow JR, Tsang SH (2014) Validation of genome-wide association study (GWAS)-identified disease risk alleles with patient-specific stem cell lines. Human Molecular Genetics. first published online February 4, 2014 doi:10.1093/hmg/ddu053
Tenzer S, Moro A, Kuharev J, Francis AC, Vidalino L, Provenzani A, Macchi P (2013) Proteome-wide characterization of the RNA-binding protein RALY-interactome using the in vivo-biotinylation-pulldown-quant (iBioPQ) approach. J Proteome Res 12(6):2869–2884
Fan Y, Thompson JW, Dubois LG, Moseley MA, Wernegreen JJ (2013) Proteomic analysis of an unculturable bacterial endosymbiont (Blochmannia) reveals high abundance of chaperonins and biosynthetic enzymes. J Proteome Res 12(2):704–718
Parviainen VI, Sakari J, Tohmola N, Renkonen R (2013) Label-free mass spectrometry proteome quantification of human embryonic kidney cells following 24 hours of sialic acid overproduction. Proteome Sci
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Brown, L.M. (2014). Quantitative Shotgun Proteomics with Data-Independent Acquisition and Traveling Wave Ion Mobility Spectrometry: A Versatile Tool in the Life Sciences. In: Woods, A., Darie, C. (eds) Advancements of Mass Spectrometry in Biomedical Research. Advances in Experimental Medicine and Biology, vol 806. Springer, Cham. https://doi.org/10.1007/978-3-319-06068-2_4
Download citation
DOI: https://doi.org/10.1007/978-3-319-06068-2_4
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-06067-5
Online ISBN: 978-3-319-06068-2
eBook Packages: Chemistry and Materials ScienceChemistry and Material Science (R0)