Abstract
Targeted mutagenesis using CRISPR/Cas technology has become routine in elucidating biological processes or their application in breeding and agriculture. This means that the change to be achieved can be accurately predicted. However, knockout of a gene function is not always desirable, as reducing activity or affecting a protein domain can influence its properties and, thus, the phenotype. This chapter will therefore focus on precise genome modification in temperate cereals. The methods used, including some representative examples, are summarised here.
You have full access to this open access chapter, Download chapter PDF
Similar content being viewed by others
Time is an essential factor to consider in developing new agricultural varieties. Since domestication, plant breeders have steadily expanded their toolbox, but establishing a new cereal variety takes an average of 8–10 years [1]. Developments in genome sequencing (barley [2], wheat [3], rye [4]), oat [5], and molecular biology methods for genome-assisted breeding (marker-assisted breeding [6]) have provided tools and techniques for the breeding process that positively influence the process and workload.
In this chapter, temperate cereals of the botanical tribe Triticeae (barley, wheat, rye, and oats) grown in Europe will be considered. Due to limitations in the availability of genomic sequences (rye and oat only in the last two years) and the lack of efficient transformation protocols, there are currently only reports from barley and wheat. These are also among the more essential cereals in Europe in terms of cultivated area (FAO Stat). While diploid barley is mainly used for animal feed, beer, and whisky, tetra- and hexaploid wheat are essential for pasta and bakery products. Oat and rye are the main components of breakfast cereals.
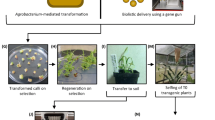
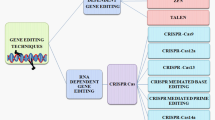
Targeted mutagenesis induced by endonucleases such as TALEN [7] and CRISPR/Cas [8] has enabled an incredible number of applications in a wide range of species since their first biotechnological application in 2012 [9]. Thus, the results have helped many new insights into basic research and show promise for applications. Targeting an organism’s genomic sequence has never been so easy. CRISPR/Cas technology is a two-component system in which a target-specific guide RNA guides a double-strand-inducing Cas enzyme to the desired location in the target genome [8]. The cell’s repair mechanisms then repair the induced double-strand breaks. A more detailed description of the technology and repair mechanisms is described in Chap. 1.
Applications of precise genome editing in plants have been summarised several times (for review, see [10,11,12]). These reach into all areas of modern plant research. Starting with pure basic research, i.e., the identification and characterization of genes and their phenotypic expressions, the methods are also used to master the current challenges of agriculture. Here, agronomic parameters play a similarly important role in improving product characteristics. Plants with enhanced resistance to fungal or viral pathogens are being developed and tested. Plants should be better adapted to changing climatic conditions and thus have better water and nutrient utilization. Plants should become heat or drought-tolerant but also be able to grow with an increased salt concentration in the soil. There are examples of all these experiments in barley and/or wheat (Fig. 9.1). The only publication on applying CRISPR/Cas technology in triticale, a cross between wheat and rye, shows its functionality exclusively in protoplasts [13].
Summary of barley and wheat genes with functions in agronomic features [14,15,16,17,18,19,20,21,22,23,24,25,26,27,28,29,30,31,32,33,34,35,36,37,38,39,40,41,42,43,44,45,46,47], gene discovery [48,49,50,51,52,53,54,55,56,57,58,59], product quality [21, 34, 60,61,62,63,64,65,66,67,68,69,70,71,72,73,74], and abiotic and biotic stress [46, 75,76,77,78,79,80,81,82,83,84,85,86,87,88,89,90,91,92]
Even if the targeted induction of a double-strand break can be carried out precisely, the result is random. According to previous reports, the most common outcome for CRISPR/Cas9 is a deletion of a few (≤10) nucleotides or the insertion of one nucleotide (InDel) [93]. However, it is impossible to predict precisely whether loss or insertion will occur, and it is also somewhat random which base is inserted. Although there are reports (for review, see [94]) that a certain percentage can be predicted using microhomology-dependent repair mechanisms, the outcome remains undetermined in most applications.
To achieve a prediction of the mutation result and thus precise genome editing, one can consider several possibilities. When using two gRNAs, one gets an exact deletion between the two induced double-strand breaks in a part of the mutated cells. In this case, the choice of gRNA binding sites can, for example, influence the function of a protein domain [49]. By not inducing a knockout, a reduced or altered functionality remains, similar to the RNAi effect but is genetically fixed in contrast.
Another possibility is to transfer a repair template with the desired sequence simultaneously with the double-strand-inducing reagent. However, the challenge here is to bring a sufficient number of repair templates to this site at the time of the double-strand break repair. One possibility is the biolistic transfer of the repaired DNA [95]. However, this method has all the previously described disadvantages that have led to the preferential use of Agrobacteria-mediated transformation [96].
Initial results at the cellular level in barley showed targeted allelic exchange of the fluorescence protein GFP. GFP and YFP differ in only one amino acid; thus, exchanging two nucleotides causes a change in the emission spectrum [97]. It was shown that 3% of the mutant epidermal cells had integrated the non-functional YFP fragment in the genome, thus exhibiting a shift in the lambda scan. These results were even surpassed when pre-assembled RNP complexes were biolistically transferred with Cas9 instead of plasmid DNA [95]. Here, it was shown that up to 8% of GFP-mutated epidermal cells exhibited such an allelic exchange. A typical application for allelic exchange is the creation of herbicide resistance [98]. Since this allows the selection of the correctly modified cells, such a method is easier to apply. However, the efficiency is expected to be lower if the modification has no selection advantage during creation.
An improvement here is the prime editing method [99]. In contrast to the Cas9 technology, the Moloney Murine Leukemia Virus reverse transcriptase (MLV-RT) domain was added to the Cas endonuclease. At the same time, the gRNA was extended by the part of the repair template. However, there have been few reports of plant applications so far, suggesting that the technology still needs improvement.
To precisely incorporate large DNA fragments in plants, a PrimeRoot-named method was recently described [100]. Third-generation PrimeRoot editors use optimized prime editing guide RNA designs, an improved plant prime editor, and superior recombinases to enable precise large DNA insertions of up to 11.1 kilobases into plant genomes. The authors describe using PrimeRoot to introduce gene regulatory elements into the rice. Applications in temperate cereals have not yet been described.
Base editing (BE) is another technology for the precise modification of genomes (DNA) or transcriptomes (RNA) of living cells at single-base resolution (for review, see [101]). BEs comprise a catalytically impaired cas nuclease fused with a nucleotide deaminase and sometimes DNA repair proteins. BEs can introduce single nucleotide variants at desired sites into the DNA (nuclear or organellar) or RNA of both dividing and non-dividing cells. There are two types of BEs – DNA BEs, which directly induce targeted point mutations in DNA, and RNA BEs, which convert one ribonucleotide to another in RNA. The currently available DNA BEs can be further divided into cytosine BEs (CBEs), adenine BEs (ABEs), C-to-G BEs (CGBEs), dual-base editors, and organellar BEs. These categories are discussed below [101]. After protoplast testing, C to T substitutions was successfully detected in two heterozygous wheat plants [102]. To increase the efficiency, further improvements such as the NLS, crRNA, LbCas12a nuclease, adenine deaminase, and linker were undertaken, achieving up to 55% efficiency in stable mutants (TaLOX and TaMLO) [103]. In other plants, further improvements, such as placing a N-terminal reverse transcriptase–Cas9 nickase fusion performed better in rice than the commonly applied C-terminal fusion [104]. In addition, introducing multiple-nucleotide substitutions in the reverse transcriptase template stimulated prime editing with enhanced efficiency. Additionally, it was shown that using two pegRNAs that encode the same edits but target complementary DNA strands highly promotes the desired outcome [105].
However, all the previously mentioned methods are still subject to particular challenges. These concern all parts of the process, such as selecting, using, and transferring appropriate gRNAs and Cas and proteins, general tissue culture, genotype dependence, detection of induced mutations, and identification of transgene-free, etc. homozygous progeny [106]. Further sequencing of genomes and enzyme evolution will undoubtedly lead to other plant improvements. It is crucial that the plants produced in this way also find use in European agriculture and that outdated regulations do not prevent their use.
References
Morrison, R.A., Evans, D.A.: Haploid plants from tissue culture: new plant varieties in a shortened time frame. Nat. Biotechnol. 6, 684–690 (1988). https://doi.org/10.1038/nbt0688-684
Mascher, M., Gundlach, H., Himmelbach, A., et al.: A chromosome conformation capture ordered sequence of the barley genome. Nature. 544, 427–433 (2017). https://doi.org/10.1038/nature22043
Shifting the limits in wheat research and breeding using a fully annotated reference genome. Science. 361 (2018) https://doi.org/10.1126/science.aar7191
Rabanus-Wallace, M.T., Hackauf, B., Mascher, M., et al.: Chromosome-scale genome assembly provides insights into rye biology, evolution and agronomic potential. Nat. Genet. 53, 564–573 (2021). https://doi.org/10.1038/s41588-021-00807-0
Kamal, N., Tsardakas Renhuldt, N., Bentzer, J., et al.: The mosaic oat genome gives insights into a uniquely healthy cereal crop. Nature. 606, 113–119 (2022). https://doi.org/10.1038/s41586-022-04732-y
Collard, B.C.Y., Mackill, D.J.: Marker-assisted selection: an approach for precision plant breeding in the twenty-first century. Philos. Trans. R. Soc. Lond. Ser. B Biol. Sci. 363, 557–572 (2008). https://doi.org/10.1098/rstb.2007.2170
Boch, J., Scholze, H., Schornack, S., et al.: Breaking the code of DNA binding specificity of TAL-type III effectors. Science. 326, 1509–1512 (2009). https://doi.org/10.1126/science.1178811
Jinek, M., Chylinski, K., Fonfara, I., et al.: A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science. 337, 816–821 (2012). https://doi.org/10.1126/science.1225829
Moon, S.B., Kim, D.Y., Ko, J.-H., et al.: Recent advances in the CRISPR genome editing tool set. Exp. Mol. Med. 51, 1–11 (2019). https://doi.org/10.1038/s12276-019-0339-7
Kumlehn, J., Pietralla, J., Hensel, G., et al.: The CRISPR/Cas revolution continues: from efficient gene editing for crop breeding to plant synthetic biology. J. Integr. Plant Biol. 60, 1127–1153 (2018). https://doi.org/10.1111/jipb.12734
Zhu, H., Li, C., Gao, C.: Applications of CRISPR-Cas in agriculture and plant biotechnology. Nat. Rev. Mol. Cell Biol. 21, 661–677 (2020). https://doi.org/10.1038/s41580-020-00288-9
Capdeville, N., Schindele, P., Puchta, H.: Getting better all the time – recent progress in the development of CRISPR/Cas-based tools for plant genome engineering. Curr. Opin. Biotechnol. 79, 102854 (2023). https://doi.org/10.1016/j.copbio.2022.102854
Michalski, K., Hertig, C., Mańkowski, D.R., et al.: Functional validation of cas9/guideRNA constructs for site-directed mutagenesis of Triticale ABA8′OH1 loci. Int. J. Mol. Sci. 22 (2021). https://doi.org/10.3390/ijms22137038
Gerasimova, S.V., Kolosovskaya, E.V., Vikhorev, A.V., et al.: WAX INDUCER 1 regulates β-Diketone biosynthesis by mediating expression of the Cer-cqu gene cluster in Barley. Int. J. Mol. Sci. 24 (2023). https://doi.org/10.3390/ijms24076762
Holubová, K., Hensel, G., Vojta, P., et al.: Modification of barley plant productivity through regulation of Cytokinin content by reverse-genetics approaches. Front. Plant Sci. 9, 1676 (2018). https://doi.org/10.3389/fpls.2018.01676
Gasparis, S., Przyborowski, M., Kała, M., et al.: Knockout of the HvCKX1 or HvCKX3 gene in Barley (Hordeum vulgare L.) by RNA-guided Cas9 nuclease affects the regulation of Cytokinin metabolism and root morphology. Cells. 8 (2019). https://doi.org/10.3390/cells8080782
Kirschner, G.K., Rosignoli, S., Guo, L., et al.: ENHANCED GRAVITROPISM 2 encodes a STERILE ALPHA MOTIF-containing protein that controls root growth angle in barley and wheat. Proc. Natl. Acad. Sci. U. S. A. 118 (2021). https://doi.org/10.1073/pnas.2101526118
Křenek, P., Chubar, E., Vadovič, P., et al.: CRISPR/Cas9-induced loss-of-function mutation in the barley mitogen-activated protein kinase 6 gene causes abnormal embryo development leading to severely reduced grain germination and seedling Shootless phenotype. Front. Plant Sci. 12, 670302 (2021). https://doi.org/10.3389/fpls.2021.670302
Lawrenson, T., Shorinola, O., Stacey, N., et al.: Induction of targeted, heritable mutations in barley and Brassica oleracea using RNA-guided Cas9 nuclease. Genome Biol. 16, 258 (2015). https://doi.org/10.1186/s13059-015-0826-7
Hisano, H., Hoffie, R.E., Abe, F., et al.: Regulation of germination by targeted mutagenesis of grain dormancy genes in barley. Plant Biotechnol. J. 20, 37–46 (2022). https://doi.org/10.1111/pbi.13692
Kamiya, Y., Abe, F., Mikami, M., et al.: A rapid method for detection of mutations induced by CRISPR/Cas9-based genome editing in common wheat. Plant Biotechnol. (Tokyo). 37, 247–251 (2020). https://doi.org/10.5511/plantbiotechnology.20.0404b
Abe, F., Haque, E., Hisano, H., et al.: Genome-edited triple-recessive mutation alters seed dormancy in wheat. Cell Rep. 28, 1362–1369.e4 (2019). https://doi.org/10.1016/j.celrep.2019.06.090
Zhu, Y., Lin, Y., Fan, Y., et al.: CRISPR/Cas9-mediated restoration of Tamyb10 to create pre-harvest sprouting-resistant red wheat. Plant Biotechnol. J. 21, 665–667 (2023). https://doi.org/10.1111/pbi.13981
Karunarathne, S.D., Han, Y., Zhang, X.-Q., et al.: CRISPR/Cas9 gene editing and natural variation analysis demonstrate the potential for HvARE1 in improvement of nitrogen use efficiency in barley. J. Integr. Plant Biol. 64, 756–770 (2022). https://doi.org/10.1111/jipb.13214
Zhang, J., Zhang, H., Li, S., et al.: Increasing yield potential through manipulating of an ARE1 ortholog related to nitrogen use efficiency in wheat by CRISPR/Cas9. J. Integr. Plant Biol. 63, 1649–1663 (2021). https://doi.org/10.1111/jipb.13151
Budhagatapalli, N., Halbach, T., Hiekel, S., et al.: Site-directed mutagenesis in bread and durum wheat via pollination by cas9/guide RNA-transgenic maize used as haploidy inducer. Plant Biotechnol. J. 18, 2376–2378 (2020). https://doi.org/10.1111/pbi.13415
Camerlengo, F., Frittelli, A., Sparks, C., et al.: CRISPR-Cas9 multiplex editing of the α-amylase/trypsin inhibitor genes to reduce allergen proteins in durum wheat. Front. Sustain. Food Syst. 4 (2020). https://doi.org/10.3389/fsufs.2020.00104
Guo, B., Jin, X., Chen, J., et al.: ATP-dependent DNA helicase (TaDHL), a novel reduced-height (Rht) gene in wheat. Genes (Basel). 13 (2022). https://doi.org/10.3390/genes13060979
Cheng, J., Hill, C., Han, Y., et al.: New semi-dwarfing alleles with increased coleoptile length by gene editing of gibberellin 3-oxidase 1 using CRISPR-Cas9 in barley (Hordeum vulgare L.). Plant Biotechnol. J. 21, 806–818 (2023). https://doi.org/10.1111/pbi.13998
Jiang, Y., Li, J., Liu, B., et al.: Novel Hina alleles created by genome editing increase grain hardness and reduce grain width in barley. Transgenic Res. 31, 637–645 (2022). https://doi.org/10.1007/s11248-022-00324-8
Zeng, Z., Han, N., Liu, C., et al.: Functional dissection of HGGT and HPT in barley vitamin E biosynthesis via CRISPR/Cas9-enabled genome editing. Ann. Bot. 126, 929–942 (2020). https://doi.org/10.1093/aob/mcaa115
Wang, W., Pan, Q., Tian, B., et al.: Gene editing of the wheat homologs of TONNEAU1-recruiting motif encoding gene affects grain shape and weight in wheat. Plant J. 100, 251–264 (2019). https://doi.org/10.1111/tpj.14440
Zhang, Y., Liang, Z., Zong, Y., et al.: Efficient and transgene-free genome editing in wheat through transient expression of CRISPR/Cas9 DNA or RNA. Nat. Commun. 7, 12617 (2016). https://doi.org/10.1038/ncomms12617
Zhang, S., Zhang, R., Song, G., et al.: Targeted mutagenesis using the agrobacterium tumefaciens-mediated CRISPR-Cas9 system in common wheat. BMC Plant Biol. 18, 302 (2018). https://doi.org/10.1186/s12870-018-1496-x
Singh, M., Kumar, M., Albertsen, M.C., et al.: Concurrent modifications in the three homeologs of Ms45 gene with CRISPR-Cas9 lead to rapid generation of male sterile bread wheat (Triticum aestivum L.). Plant Mol. Biol. 97, 371–383 (2018). https://doi.org/10.1007/s11103-018-0749-2
Li, J., Wang, Z., He, G., et al.: CRISPR/Cas9-mediated disruption of TaNP1 genes results in complete male sterility in bread wheat. J. Genet. Genomics. 47, 263–272 (2020). https://doi.org/10.1016/j.jgg.2020.05.004
Tang, H., Liu, H., Zhou, Y., et al.: Fertility recovery of wheat male sterility controlled by Ms2 using CRISPR/Cas9. Plant Biotechnol. J. 19, 224–226 (2021). https://doi.org/10.1111/pbi.13482
Zhang, R., Zhang, S., Li, J., et al.: CRISPR/Cas9-targeted mutagenesis of TaDCL4, TaDCL5 and TaRDR6 induces male sterility in common wheat. Plant Biotechnol. J. 21, 839–853 (2023). https://doi.org/10.1111/pbi.14000
de Souza Moraes, T., van Es, S.W., Hernández-Pinzón, I., et al.: The TCP transcription factor HvTB2 heterodimerizes with VRS5 and controls spike architecture in barley. Plant Reprod. 35, 205–220 (2022). https://doi.org/10.1007/s00497-022-00441-8
Wang, C., Yang, X., Zhang, Y., et al.: Barley FASCIATED EAR genes determine inflorescence meristem size and yield traits. Crop J. (2022). https://doi.org/10.1016/j.cj.2022.10.001
Chen, Z., Ke, W., He, F., et al.: A single nucleotide deletion in the third exon of FT-D1 increases the spikelet number and delays heading date in wheat (Triticum aestivum L.). Plant Biotechnol. J. 20, 920–933 (2022). https://doi.org/10.1111/pbi.13773
Errum, A., Rehman, N., Uzair, M., et al.: CRISPR/Cas9 editing of wheat Ppd-1 gene homoeologs alters spike architecture and grain morphometric traits. Funct. Integr. Genomics. 23, 66 (2023). https://doi.org/10.1007/s10142-023-00989-2
Wang, W., Yu, Z., He, F., et al.: Multiplexed promoter and gene editing in wheat using a virus-based guide RNA delivery system. Plant Biotechnol. J. 20, 2332–2341 (2022). https://doi.org/10.1111/pbi.13910
Miroshnichenko, D., Timerbaev, V., Klementyeva, A., et al.: CRISPR/Cas9-induced modification of the conservative promoter region of VRN-A1 alters the heading time of hexaploid bread wheat. Front. Plant Sci. 13, 1048695 (2022). https://doi.org/10.3389/fpls.2022.1048695
Gupta, A., Hua, L., Zhang, Z., et al.: CRISPR-induced miRNA156-recognition element mutations in TaSPL13 improve multiple agronomic traits in wheat. Plant Biotechnol. J. 21, 536–548 (2023). https://doi.org/10.1111/pbi.13969
Wang, W., Pan, Q., He, F., et al.: Transgenerational CRISPR-Cas9 activity facilitates multiplex gene editing in allopolyploid wheat. CRISPR J. 1, 65–74 (2018). https://doi.org/10.1089/crispr.2017.0010
Luo, J., Li, S., Xu, J., et al.: Pyramiding favorable alleles in an elite wheat variety in one generation by CRISPR-Cas9-mediated multiplex gene editing. Mol. Plant. 14, 847–850 (2021). https://doi.org/10.1016/j.molp.2021.03.024
Li, M., Hensel, G., Mascher, M., et al.: Leaf variegation and impaired chloroplast development caused by a truncated CCT domain gene in albostrians barley. Plant Cell. 31, 1430–1445 (2019). https://doi.org/10.1105/tpc.19.00132
Li, M., Hensel, G., Melzer, M., et al.: Mutation of the ALBOSTRIANS Ohnologous gene HvCMF3 impairs chloroplast development and thylakoid architecture in barley. Front. Plant Sci. 12, 732608 (2021). https://doi.org/10.3389/fpls.2021.732608
Han, Y., Broughton, S., Liu, L., et al.: Highly efficient and genotype-independent barley gene editing based on anther culture. Plant Commun. 2, 100082 (2021). https://doi.org/10.1016/j.xplc.2020.100082
Steckenborn, S., Cuacos, M., Ayoub, M.A., et al.: The meiotic topoisomerase VI B subunit (MTOPVIB) is essential for meiotic DNA double-strand break formation in barley (Hordeum vulgare L.). Plant Reprod. 36, 1–15 (2023). https://doi.org/10.1007/s00497-022-00444-5
Amanda, D., Frey, F.P., Neumann, U., et al.: Auxin boosts energy generation pathways to fuel pollen maturation in barley. Curr. Biol. 32, 1798–1811.e8 (2022). https://doi.org/10.1016/j.cub.2022.02.073
Huang, Y., Kamal, R., Shanmugaraj, N., et al.: A molecular framework for grain number determination in barley. Sci. Adv. 9, eadd0324 (2023). https://doi.org/10.1126/sciadv.add0324
Radchuk, V., Belew, Z.M., Gündel, A., et al.: SWEET11b transports both sugar and cytokinin in developing barley grains. Plant Cell. (2023). https://doi.org/10.1093/plcell/koad055
Yang, X., Wilkinson, L.G., Aubert, M.K., et al.: Ovule cell wall composition is a maternal determinant of grain size in barley. New Phytol. 237, 2136–2147 (2023). https://doi.org/10.1111/nph.18714
Li, J., Zhang, S., Zhang, R., et al.: Efficient multiplex genome editing by CRISPR/Cas9 in common wheat. Plant Biotechnol. J. 19, 427–429 (2021). https://doi.org/10.1111/pbi.13508
Hyde, L., Osman, K., Winfield, M., et al.: Identification, characterization, and rescue of CRISPR/Cas9 generated wheat SPO11-1 mutants. Plant Biotechnol. J. 21, 405–418 (2023). https://doi.org/10.1111/pbi.13961
Zhou, H., Xu, L., Li, F., et al.: Transcriptional regulation by CRISPR/dCas9 in common wheat. Gene. 807, 145919 (2022). https://doi.org/10.1016/j.gene.2021.145919
Howells, R.M., Craze, M., Bowden, S., et al.: Efficient generation of stable, heritable gene edits in wheat using CRISPR/Cas9. BMC Plant Biol. 18, 215 (2018). https://doi.org/10.1186/s12870-018-1433-z
Holme, I.B., Wendt, T., Gil-Humanes, J., et al.: Evaluation of the mature grain phytase candidate HvPAPhy_a gene in barley (Hordeum vulgare L.) using CRISPR/Cas9 and TALENs. Plant Mol. Biol. 95, 111–121 (2017). https://doi.org/10.1007/s11103-017-0640-6
Vlčko, T., Ohnoutková, L.: Allelic variants of CRISPR/Cas9 induced mutation in an inositol trisphosphate 5/6 kinase gene manifest different phenotypes in barley. Plants (Basel). 9 (2020). https://doi.org/10.3390/plants9020195
Kapusi, E., Corcuera-Gómez, M., Melnik, S., et al.: Heritable genomic fragment deletions and small Indels in the putative ENGase gene induced by CRISPR/Cas9 in Barley. Front. Plant Sci. 8, 540 (2017). https://doi.org/10.3389/fpls.2017.00540
Gasparis, S., Kała, M., Przyborowski, M., et al.: A simple and efficient CRISPR/Cas9 platform for induction of single and multiple, heritable mutations in barley (Hordeum vulgare L.). Plant Methods. 14, 111 (2018). https://doi.org/10.1186/s13007-018-0382-8
Gerasimova, S.V., Hertig, C., Korotkova, A.M., et al.: Conversion of hulled into naked barley by Cas endonuclease-mediated knockout of the NUD gene. BMC Plant Biol. 20, 255 (2020). https://doi.org/10.1186/s12870-020-02454-9
Zang, Y., Gong, Q., Xu, Y., et al.: Production of conjoined transgenic and edited barley and wheat plants for Nud genes using the CRISPR/SpCas9 system. Front. Genet. 13, 873850 (2022). https://doi.org/10.3389/fgene.2022.873850
Li, Y., Liu, D., Zong, Y., et al.: New D hordein alleles were created in barley using CRISPR/Cas9 genome editing. Cereal Res. Commun. 48, 131–138 (2020). https://doi.org/10.1007/s42976-020-00023-2
Yang, Q., Zhong, X., Li, Q., et al.: Mutation of the d-hordein gene by RNA-guided Cas9 targeted editing reducing the grain size and changing grain compositions in barley. Food Chem. 311, 125892 (2020). https://doi.org/10.1016/j.foodchem.2019.125892
Garcia-Gimenez, G., Barakate, A., Smith, P., et al.: Targeted mutation of barley (1,3;1,4)-β-glucan synthases reveals complex relationships between the storage and cell wall polysaccharide content. Plant J. 104, 1009–1022 (2020). https://doi.org/10.1111/tpj.14977
Zhong, Y., Blennow, A., Kofoed-Enevoldsen, O., et al.: Protein targeting to starch 1 is essential for starchy endosperm development in barley. J. Exp. Bot. 70, 485–496 (2019). https://doi.org/10.1093/jxb/ery398
Yang, Q., Ding, J., Feng, X., et al.: Editing of the starch synthase IIa gene led to transcriptomic and metabolomic changes and high amylose starch in barley. Carbohydr. Polym. 285, 119238 (2022). https://doi.org/10.1016/j.carbpol.2022.119238
Li, J., Jiao, G., Sun, Y., et al.: Modification of starch composition, structure and properties through editing of TaSBEIIa in both winter and spring wheat varieties by CRISPR/Cas9. Plant Biotechnol. J. 19, 937–951 (2021). https://doi.org/10.1111/pbi.13519
Raffan, S., Sparks, C., Huttly, A., et al.: Wheat with greatly reduced accumulation of free asparagine in the grain, produced by CRISPR/Cas9 editing of asparagine synthetase gene TaASN2. Plant Biotechnol. J. 19, 1602–1613 (2021). https://doi.org/10.1111/pbi.13573
Ibrahim, S., Saleem, B., Rehman, N., et al.: CRISPR/Cas9 mediated disruption of Inositol Pentakisphosphate 2-Kinase 1 (TaIPK1) reduces phytic acid and improves iron and zinc accumulation in wheat grains. J. Adv. Res. 37, 33–41 (2022). https://doi.org/10.1016/j.jare.2021.07.006
Hu, J., Yu, M., Chang, Y., et al.: Functional analysis of TaPDI genes on storage protein accumulation by CRISPR/Cas9 edited wheat mutants. Int. J. Biol. Macromol. 196, 131–143 (2022). https://doi.org/10.1016/j.ijbiomac.2021.12.048
Kim, S.-Y., Bengtsson, T., Olsson, N., et al.: Mutations in two aphid-regulated β-1,3-Glucanase genes by CRISPR/Cas9 do not increase barley resistance to Rhopalosiphum padi L. Front. Plant Sci. 11, 1043 (2020). https://doi.org/10.3389/fpls.2020.01043
Galli, M., Martiny, E., Imani, J., et al.: CRISPR/SpCas9-mediated double knockout of barley Microrchidia MORC1 and MORC6a reveals their strong involvement in plant immunity, transcriptional gene silencing and plant growth. Plant Biotechnol. J. 20, 89–102 (2022). https://doi.org/10.1111/pbi.13697
Hoffie, R.E., Otto, I., Perovic, D., et al.: Targeted knockout of eukaryotic translation initiation factor 4E confers Bymovirus resistance in winter barley. Front. Genome Editing. 3, 784233 (2021). https://doi.org/10.3389/fgeed.2021.784233
Hoffie, R.E., Perovic, D., Habekuß, A., et al.: Novel resistance to the Bymovirus BaMMV established by targeted mutagenesis of the PDIL5-1 susceptibility gene in barley. Plant Biotechnol. J. 21, 331–341 (2023). https://doi.org/10.1111/pbi.13948
Hahn, F., Sanjurjo Loures, L., Sparks, C.A., et al.: Efficient CRISPR/Cas-mediated targeted mutagenesis in spring and winter wheat varieties. Plants (Basel). 10 (2021). https://doi.org/10.3390/plants10071481
Kan, J., Cai, Y., Cheng, C., et al.: Simultaneous editing of host factor gene TaPDIL5-1 homoeoalleles confers wheat yellow mosaic virus resistance in hexaploid wheat. New Phytol. 234, 340–344 (2022). https://doi.org/10.1111/nph.18002
Kan, J., Cai, Y., Cheng, C., et al.: CRISPR/Cas9-guided knockout of eIF4E improves wheat yellow mosaic virus resistance without yield penalty. Plant Biotechnol. J. 21, 893–895 (2023). https://doi.org/10.1111/pbi.14002
Koide, H., Hisano, H., Yaeno, T.: CRISPR/Cas9-based generation of mlo mutants for allelic complementation experiments to elucidate MLO function in barley. J. Gen. Plant Pathol. 89, 153–158 (2023). https://doi.org/10.1007/s10327-023-01120-w
Zhang, Y., Bai, Y., Wu, G., et al.: Simultaneous modification of three homoeologs of TaEDR1 by genome editing enhances powdery mildew resistance in wheat. Plant J. 91, 714–724 (2017). https://doi.org/10.1111/tpj.13599
Sánchez-León, S., Gil-Humanes, J., Ozuna, C.V., et al.: Low-gluten, nontransgenic wheat engineered with CRISPR/Cas9. Plant Biotechnol. J. 16, 902–910 (2018). https://doi.org/10.1111/pbi.12837
Brambilla, A., Lenk, M., Ghirardo, A., et al.: Pipecolic acid synthesis is required for systemic acquired resistance and plant-to-plant-induced immunity in barley. J. Exp. Bot. (2023). https://doi.org/10.1093/jxb/erad095
Brauer, E.K., Balcerzak, M., Rocheleau, H., et al.: Genome editing of a Deoxynivalenol-induced transcription factor confers resistance to fusarium graminearum in wheat. Mol. Plant-Microbe Interact. 33, 553–560 (2020). https://doi.org/10.1094/mpmi-11-19-0332-r
He, F., Wang, C., Sun, H., et al.: Simultaneous editing of three homoeologues of TaCIPK14 confers broad-spectrum resistance to stripe rust in wheat. Plant Biotechnol. J. 21, 354–368 (2023). https://doi.org/10.1111/pbi.13956
Huang, X., Bai, X., Qian, C., et al.: TaUAM3, a UDP-Ara mutases protein, positively regulates wheat resistance to the stripe rust fungus. Food Energy Secur. (2023). https://doi.org/10.1002/fes3.456
Zheng, M., Lin, J., Liu, X., et al.: Histone acetyltransferase TaHAG1 acts as a crucial regulator to strengthen salt tolerance of hexaploid wheat. Plant Physiol. 186, 1951–1969 (2021). https://doi.org/10.1093/plphys/kiab187
Mohr, T., Horstman, J., Gu, Y.Q., et al.: CRISPR-Cas9 gene editing of the Sal1 gene family in wheat. Plants (Basel). 11 (2022). https://doi.org/10.3390/plants11172259
Tian, X., Qin, Z., Zhao, Y., et al.: Stress granule-associated TaMBF1c confers thermotolerance through regulating specific mRNA translation in wheat (Triticum aestivum). New Phytol. 233, 1719–1731 (2022). https://doi.org/10.1111/nph.17865
Kim, D., Alptekin, B., Budak, H.: CRISPR/Cas9 genome editing in wheat. Funct. Integr. Genomics. 18, 31–41 (2018). https://doi.org/10.1007/s10142-017-0572-x
Bortesi, L., Zhu, C., Zischewski, J., et al.: Patterns of CRISPR/Cas9 activity in plants, animals and microbes. Plant Biotechnol. J. 14, 2203–2216 (2016). https://doi.org/10.1111/pbi.12634
van Vu, T., Thi Hai Doan, D., Kim, J., et al.: CRISPR/Cas-based precision genome editing via microhomology-mediated end joining. Plant Biotechnol. J. 19, 230–239 (2021). https://doi.org/10.1111/pbi.13490
Becker, M., Hensel, G.: Ribonucleoprotein (RNP)-mediated allele replacement in barley (Hordeum vulgare L.) leaves. Methods Mol. Biol. 2653, 199–205 (2023). https://doi.org/10.1007/978-1-0716-3131-7_13
Harwood, W.A.: Advances and remaining challenges in the transformation of barley and wheat. J. Exp. Bot. 63, 1791–1798 (2012). https://doi.org/10.1093/jxb/err380
Budhagatapalli, N., Rutten, T., Gurushidze, M., et al.: Targeted modification of gene function exploiting homology-directed repair of TALEN-mediated double-strand breaks in barley. G3 (Bethesda). 5, 1857–1863 (2015). https://doi.org/10.1534/g3.115.018762
Svitashev, S., Schwartz, C., Lenderts, B., et al.: Genome editing in maize directed by CRISPR-Cas9 ribonucleoprotein complexes. Nat. Commun. 7, 13274 (2016). https://doi.org/10.1038/ncomms13274
Anzalone, A.V., Randolph, P.B., Davis, J.R., et al.: Search-and-replace genome editing without double-strand breaks or donor DNA. Nature. 576, 149–157 (2019). https://doi.org/10.1038/s41586-019-1711-4
Sun, C., Lei, Y., Li, B., et al.: Precise integration of large DNA sequences in plant genomes using PrimeRoot editors. Nat. Biotechnol. (2023). https://doi.org/10.1038/s41587-023-01769-w
Molla, K.A., Sretenovic, S., Bansal, K.C., et al.: Precise plant genome editing using base editors and prime editors. Nat. Plant. 7, 1166–1187 (2021). https://doi.org/10.1038/s41477-021-00991-1
Zong, Y., Wang, Y., Li, C., et al.: Precise base editing in rice, wheat and maize with a Cas9-cytidine deaminase fusion. Nat. Biotechnol. 35, 438–440 (2017). https://doi.org/10.1038/nbt.3811
Gaillochet, C., Peña Fernández, A., Goossens, V., et al.: Systematic optimization of Cas12a base editors in wheat and maize using the ITER platform. Genome Biol. 24, 6 (2023). https://doi.org/10.1186/s13059-022-02836-2
Xu, W., Yang, Y., Yang, B., et al.: A design optimized prime editor with expanded scope and capability in plants. Nat. Plant. 8, 45–52 (2022). https://doi.org/10.1038/s41477-021-01043-4
Lin, Q., Jin, S., Zong, Y., et al.: High-efficiency prime editing with optimized, paired pegRNAs in plants. Nat. Biotechnol. 39, 923–927 (2021). https://doi.org/10.1038/s41587-021-00868-w
Cardi, T., Murovec, J., Bakhsh, A., et al.: CRISPR/Cas-mediated plant genome editing: outstanding challenges a decade after implementation. Trends Plant Sci. 28, 1144–1165 (2023). https://doi.org/10.1016/j.tplants.2023.05.012
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Open Access This chapter is licensed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license and indicate if changes were made.
The images or other third party material in this chapter are included in the chapter's Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the chapter's Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
Copyright information
© 2024 The Author(s)
About this chapter
Cite this chapter
Pouramini, P., Hensel, G. (2024). Precise Gene Editing of Cereals Using CRISPR/Cas Technology. In: Ricroch, A., Eriksson, D., Miladinović, D., Sweet, J., Van Laere, K., Woźniak-Gientka, E. (eds) A Roadmap for Plant Genome Editing . Springer, Cham. https://doi.org/10.1007/978-3-031-46150-7_9
Download citation
DOI: https://doi.org/10.1007/978-3-031-46150-7_9
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-46149-1
Online ISBN: 978-3-031-46150-7
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)