Abstract
The cholesterol of the host cell plasma membrane and viral M2 protein plays a crucial role in multiple stages of infection and replication of the influenza A virus. Cholesterol is required for the formation of heterogeneous membrane microdomains (or rafts) in the budozone of the host cell that serves as assembly sites for the viral components. The raft microstructures act as scaffolds for several proteins. Cholesterol may further contribute to the mechanical forces necessary for membrane scission in the last stage of budding and help to maintain the stability of the virus envelope. The M2 protein has been shown to cause membrane scission in model systems by promoting the formation of curved lipid bilayer structures that, in turn, can lead to membrane vesicles budding off or scission intermediates. Membrane remodeling by M2 is intimately linked with cholesterol as it affects local lipid composition, fluidity, and stability of the membrane. Thus, both cholesterol and M2 protein contribute to the efficient and proper release of newly formed influenza viruses from the virus-infected cells.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
Introduction
Crossing the membrane is a significant physical barrier for viruses, and different strategies are employed to facilitate this process. The newly synthesized components of influenza A virus (IAV) in the infected host cell will come together and organize laterally in a region of the plasma membrane called the “budozone” (Fig. 16.1) (Leser and Lamb 2017). After this assembly occurs, the budozone becomes the site from which budding occurs. Importantly, the lipid composition in this region differs markedly from the average stoichiometric ratios found in the host cell as it is enriched in cholesterol and certain lipids (e.g., saturated phospholipids and sphingolipids), which help drive the assembly of viral components at the phase boundary of lipid microdomains and alter membrane biomechanics (Fig. 16.1). Some of the viral proteins (e.g., hemagglutinin (HA) and nucleoprotein) appear to concentrate spontaneously in the cholesterol-rich membrane environment whereas matrix protein 2 (M2) association with the budozone materializes in a fashion dependent on interactions with other proteins (matrix protein 1 (M1) and HA in the cluster (Leser and Lamb 2017). Notably, the lateral enrichment of certain lipid components persists through the entire process of budding and virus release, making the emerging viral envelope likewise enriched in cholesterol and sphingolipids (Lenard and Compans 1974; Nayak and Barman 2002). Lateral heterogeneity of lipids and (host as well as viral) proteins enable thermodynamically or dynamically stabilized microdomains of functional relevance to exist under the control of cellular regulatory processes (Mouritsen and Jorgensen 1992; Mouritsen 2016). The popular term membrane/lipid “raft” is frequently used to signify such structures of the membrane. Rafts appear to be an essential platform for viral and host components to interact with one another.
“Overview of the budding of influenza viruses, showing the coalescence of HA and NA containing lipid rafts (shown in yellow), the formation of a filamentous virion and membrane scission caused by M2 clustered at the neck of the budding virus.” Reprinted from Rossman and Lamb (2011), Copyright 2011, with permission from Elsevier
Viruses are compact and small by necessity. Their genetic material will typically contain a minimal set of genes encoding proteins strictly needed for functions they cannot hijack from the host cell. Therefore, the scission of enveloped viruses, the process in which the budding virus cuts the membrane in order to release from the host, is normally performed by using the endosomal sorting complex required for transport (ESCRT) machinery belonging to the host (Votteler and Sundquist 2013). Compelling evidence has led to the proposal that IAV is an exception to this principle. Microscopy data show that M2 is localized at the budding neck of the virus. Additionally, evidence from Nuclear magnetic resonance (NMR) and Electron paramagnetic resonance (EPR) spectroscopy, as well as computational studies, supports this hypothesis at the molecular level, although there are still many unresolved aspects.
This chapter will describe the roles and interactions between two of the key biomolecular components, cholesterol and M2, of the IAV-infected host cell lipid environment during budding and scission. Cholesterol is important in facilitating infection, replication, and budding of IAV (Sun and Whittaker 2003). M2 is important because of its evident role in facilitating the scission of the membrane to complete budding (Rossman et al. 2010b) in addition to its essential role as a proton channel during viral entry (Lamb et al. 1994; Helenius 1992; Pinto et al. 1992). We will set the frame by sketching out the key players and steps of IAV budding and scission (see below) and work toward a contemporary hypothesis for the roles of cholesterol and M2 in budding and scission based on recent discoveries. The essential biophysical techniques used to uncover the knowledge we have in this area are listed in the following section.
Methods
Experimental Techniques
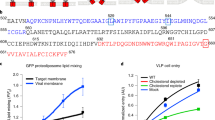
Microscopic techniques are used to visualize objects. Cells, microbes, and viruses are typically studied using light, fluorescence, or electron microscopy (Harry et al. 1995; Swanson et al. 1969; Goldsmith and Miller 2009; Scaffidi et al. 2021). Light microscopy illuminates the sample with visible light and uses arrangements of optical lenses to magnify the image. Fluorescence microscopy uses a high-intensity light source to excite a molecular tag (the fluorophore) attached to components of the sample (Fig. 16.2). The tag reemits light on a different wavelength once the excitation relaxes to a low-energy state, allowing for image magnification and effective resolution beyond what is possible with conventional light microscopy. This technique is also especially suited for highlighting certain components in order to see where they localize, including specific organelles of a cell or even individual proteins. In electron microscopy, on the other hand, the sample is exposed to a beam of accelerated electrons. The electron beam has a wavelength of around five orders of magnitude shorter than the photons of visible light and can therefore resolve much smaller structures. Often the technique of immunogold labeling is used in conjunction with electron microscopy to determine the localization of specific components. This labeling works by attaching gold particles to a secondary antibody that will bind to a primary antibody, which itself attaches to the target component (e.g., protein) (Slot and Geuze 1985; Leser et al. 1996). The high electron density of the gold particle gives rise to high levels of electron scattering, making them appear as distinct dark spots in the image. However, in contrast to light microscopy techniques, imaging by electron microscopy requires fixation and positive staining of the sample, preventing the examination of living samples (Leser et al. 1996).
Vesicle formation “(a) Deposition of lipid droplets onto the electrode surface. (b) Evaporation of organic solvent under vacuum. (c) Construction of the electroformation chamber. (d) Electroformation chamber filled with an internal solution and connected to an alternating current function generator. (e) An image of fluorescently labeled giant unilamellar vesicles (GUVs) obtained using fluorescence microscopy. The scale bar denotes 50 μm.” Reprinted from Membranes, Vol 11:11, Zvonimir Boban, Ivan Mardešić, Witold Karol Subczynski, Marija Raguz, Giant Unilamellar Vesicle Electroformation: What to Use, What to Avoid, and How to Quantify the Results, Article 860, Copyright 2021, under CC BY 4.0 license from MDPI (Boban et al. 2021)
In spectroscopic methods such as NMR and EPR, the goal is to measure quantities that will reveal properties of the sample by radiating it, for instance, with a strong magnetic field. The magnet excites the behavior of the atomic nuclei (in NMR) on the electron (in EPR). The results are quantifiable spectral lines or energies, which will often allow for making inferences about correlations or distances between components of the sample, for example, the distance between two amino acid residues of a protein (Klug and Feix 2008; Marion 2013; Fan and Lane 2016).
Theoretical Modeling and Simulation
The structural resolving of biomolecular components by experimental means such as X-ray crystallography, NMR, or cryo-electron microscopy enables interrogation of the molecules and their interplay using computational methods. Molecular dynamics (MD) simulation, affectionately called “the computational microscope,” is now routinely employed due to a relatively low entry barrier, both monetary and technical, and because the method enables investigation with unparalleled spatiotemporal resolution. MD simulations are used to study the behavior of complex molecular systems over time. A minimal, well-defined system called the simulation “box” containing all desired components (proteins, nucleic acids, lipids, ions, water, etc.) is constructed (Fig. 16.3). The interactions between all atoms in the system are described by physics-based potentials collectively known as the “force field” and the time evolution of the system can be simulated by integrating Newton’s equations of motion for the atoms in the system subject to the force field. From these data, a wide range of properties can be computed, including structural properties (e.g., distribution of distances and angles between atoms, and the organization of molecules), thermodynamic properties (e.g., temperature, entropy, and heat capacity), and kinetic properties (e.g., rates of reactions and transformations) (Allen and Tildesley 1989; Frenkel et al. 2002). For an accessible account of the more detailed aspects of the MD technique to nonexperts, we can refer to our recent review article (Ohkubo and Madsen 2021). In general, modern force fields demonstrate satisfactory accuracy, and there is a growing recognition of limitations in certain situations, such as those pertaining to cholesterol in lipid mixtures (Javanainen et al. 2023).
The general workflow of molecular dynamics simulation. A model structure of the M2 protein is selected together with the initial system setup configuration. We show a ribbon diagram of M2 resolved by solid-state NMR (PDB ID: 2L0J) (Sharma et al. 2010) embedded in a phospholipid bilayer consisting of 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine (POPC) and cholesterol. Water and ions fill the remainder of the box. The system is assembled, for example, using the CHARMM-GUI tool (Jo et al. 2008; Wu et al. 2014). From here, Newton’s equations of motion are solved, propagating the dynamics until the system’s properties stabilize over time. The resulting data are analyzed for the desired properties. Visualizations of the protein structure and simulation box components are created with VMD (Humphrey et al. 1996). Graph data graph is plotted with Matplotlib (Hunter 2007). The Fig. is created, in part, using BioRender (https://BioRender.com)
In certain studies, “coarse-grained” computational techniques are employed, which refers to methods where the individual biomolecular constituents of the simulation are represented at a lower level of detail than the atomistic level. These techniques can range from intermediate resolution methods (e.g., the Martini force field (Marrink et al. 2007)) to highly coarse-grained techniques (e.g., elastic network models (Madsen et al. 2017; Grime and Madsen 2019)) and are used to accelerate computational speed, enabling the exploration of larger systems over longer time scales. However, it is important to note that using such methods can exacerbate inaccuracies in the system (Ingolfsson et al. 2014; Jarin et al. 2021; Pak et al. 2019). Comprehensive validation to ensure that the results are reliable is required for any application of theoretical modeling and simulation.
Biomolecules and Interactions Driving Budding and Scission
IAV proteins and RNA interact intricately with each other and with components of the host cell in and around the plasma membrane in a complex, multistep process to assemble and bud-off progeny viruses (Lakadamyali et al. 2004; Noda and Kawaoka 2010). The key players in this process are the viral protein HA, neuraminidase (NA), M1, M2, the ribonucleoprotein (RNP) complex, and the arena of interaction at the heterogeneous plasma membrane environment. In this section, we review the fundamentals of budding and scission of IAV.
Initiation of Budding
Viral proteins HA and NA are transported to the plasma membrane where they cluster with cholesterol, sphingolipids, and saturated phospholipids, forming the budozone, a heterogeneous lipid domain on the plasma membrane where the virus assembles (Fig. 16.4A) (Leser and Lamb 2017). The budozone region serves as a scaffold for the recruitment of other viral proteins and the viral RNA. It is thought that this initial clustering and lipid domain formation initiates the process of viral budding by creating a lipid phase boundary, resulting in a local change in membrane curvature (Fig. 16.4A) (Wang et al. 2012). This change in curvature leads to bud formation, which subsequently grows and matures into a new virus particle in a process driven by the mutual interactions between the viral proteins (in particular, M1 polymerization), host cell proteins, and the plasma membrane.
“Model of influenza virus budding. (a) The initiation of virus budding caused by clustering of HA (shown in red) and NA (shown in orange) in lipid raft domains. M1 (shown in purple) is seen binding to the cytoplasmic tails of HA and NA and serves as a docking site for the vRNPs (shown in yellow). (b) Elongation of the budding virion caused by polymerization of the M1 protein, resulting in a polarized localization of the vRNPs. M2 (shown in blue) is recruited to the periphery of the budding virus though interactions with M1. (c) Membrane scission caused by the insertion of the M2 amphipathic helix at the lipid phase boundary, altering membrane curvature at the neck of the budding virus and leading to release of the budding virus.” Reprinted from Rossman and Lamb (2011), Copyright 2011, with permission from Elsevier
Virion Assembly
The cytoplasmic tails of clustered HA and NA, coupled with the lipid environment of the budozone, attract the internal structural components of the virus, in particular, M1 and the RNP complex (Fig. 16.4B). Polymerization of M1 at the budozone is thought to be responsible for the formation and elongation of the nascent virus, which can form spherical or filamentous virion morphologies depending on specific sequence motifs within certain viral proteins (Bourmakina and Garcia-Sastre 2005; Elleman and Barclay 2004; Roberts et al. 1998). The resulting virions contain HA and NA on their surface, a helical layer of polymerized M1 protein underlying the membrane and an internal collection of RNP complexes, each packaging a different segment of the viral genome (Fig. 16.4B) (Waterson et al. 1963; Nermut and Frank 1971; Schulze 1972). The presence of these different virus morphologies can have significant implications for viral budding, entry, host interactions, and may affect the severity of the disease caused by the virus (Taubenberger and Morens 2008; Kalil and Thomas 2019). Despite the formation of fully assembled virus particles, the virions remain attached to the plasma membrane by a small membrane neck that requires scission for their complete release (Fig. 16.4C) (McCown and Pekosz 2006; Rossman et al. 2010b).
Scission of the Budding Neck
The process of IAV membrane scission is thought to involve the action of the viral protein M2, which promotes the formation of curved lipid bilayer structures that can lead to membrane scission (Fig. 16.4C) (Rossman et al. 2010b; Schmidt et al. 2013; Martyna et al. 2017; Wang and Hong 2015). The mechanical forces necessary for scission may be contributed by cholesterol (Hubert et al. 2020; Anderson et al. 2021), which fittingly is also required for the stability of the virus envelope (Lenard and Compans 1974; Nayak and Barman 2002; Bajimaya et al. 2017). During the initial process of viral budding, the M2 protein is conspicuously excluded from the core of the budozone and may not be actively involved in virion formation (Leser and Lamb 2017). The first two steps of budding can even occur in viruses where the M2 protein is deleted or otherwise dysfunctional (Rossman et al. 2010b; Herneisen et al. 2017; McCown and Pekosz 2006). Intriguingly, the process comes to a complete stall in the absence of M2, leaving behind the uncut constricted budding neck from which a secondary budding can occur at the same site. This gives rise to a characteristic phenotype when several unsuccessful budding attempts occur in succession and a “beads-on-a-string” morphology emerges (Fig. 16.5, top panel) (McCown and Pekosz 2006; Rossman et al. 2010b). As virion assembly progresses, the planar budozone becomes folded and incorporated into the emerging virus. The M2 protein, located at the periphery of the budozone is then drawn to the constricted membrane neck of the budding virus (Rossman et al. 2010b). There is evidence that interactions between the M2 amphipathic helix M2(AH), which is a highly conserved region of the protein, and the plasma membrane can alter membrane curvature at the virion neck and are responsible, at least in part, for the role that M2 directly plays in membrane scission (Rossman et al. 2010b; Roberts et al. 2013; Schmidt et al. 2013; Rossman et al. 2010a; Martyna et al. 2017; Wang and Hong 2015; Andreas et al. 2015). During the process of scission, some M2 protein gets incorporated into the newly released virion, generating an infectious virus capable of starting the next round of infection (Fig. 16.5, top panel).
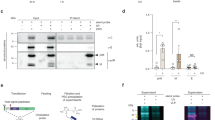
“The M2 Amphipathic Helix is Necessary for Membrane Scission and Virion Release. MDCK cells were infected with an MOI of 3 pfu/cell of (a) A/Udorn/72” or (b) “A/Udorn/72 M2(AH-Mut) for 18 hr and thin sections were analyzed by electron microscopy. The scale bars indicate 100 nm.” (c) “GUVs electroformed with 30 or 0.5 molar % of cholesterol, with 0.5 mg/ml of lucifer yellow (shown in blue) added to the resuspension buffer, were treated with 10 mM of the indicated peptide and imaged within 1 hr.” Reprinted from Cell, Vol 142:6, Jeremy S. Rossman, Xianghong Jing, George P. Leser, Robert A. Lamb, Influenza Virus M2 Protein Mediates ESCRT-Independent Membrane Scission, Pages 902–913, Copyright 2010, with permission from Cell Press (Rossman et al. 2010b)
A Contemporary Hypothesis on the Roles of Cholesterol and M2
M2 protein can form clusters in the plasma membrane and viral envelope (Sutherland et al. 2022; Madsen et al. 2018; Paulino et al. 2019; Elkins et al. 2017). Clustering is thought to be driven by hydrophobic interaction between M2 transmembrane (TM) domains, as well as the specific arrangements of the amino acids within the protein (Bao et al. 2022). The clustering process is believed to be highly dependent on lipid composition (Sutherland et al. 2022; Elkins et al. 2017; Elkins et al. 2018) and can be influenced by the microenvironment or the action of other proteins (Leser and Lamb 2017). Curvature sorting plays a role in membrane geometries where the curvature is commensurate with the wedge-shaped M2 (Madsen et al. 2018; Martyna et al. 2016; Ho et al. 2016). Proximity among multiple M2 proteins is thought to play a role in the functioning of the protein, with clustering thought to occur in a stepwise, hierarchical fashion (Fig. 16.6) (Sutherland et al. 2022; Chen et al. 2008).
“(a) M2 in a planar bilayer that is exposed to a constant, external compressive stress. (Top) (x, y)-plane projected area per lipid in the simulation box. Three time blocks are defined for further analysis, I, II, and III. (Bottom) Top view of protein tracer lines show how proteins gradually come together in the three time blocks (I, II, and III). (b) Final top view and side view snapshots after the membrane significantly deviates from planarity at t = 20 Mτ. Red arrows indicate the analogy with the in vitro system. (c) MDCK cells were infected with 3 pfu per cell of A/Udorn/72 for 18 h before fixation, immunogold labeling of M2, thin sectioning, and analysis by electron microscopy. (Scale bar: 100 nm.)” Reprinted from PNAS, Vol 115:37, Jesper J. Madsen, John M. A. Grime, Jeremy S. Rossman, Gregory A. Voth, Entropic forces drive clustering and spatial localization of influenza A M2 during viral budding, Pages E8595-E8603, with permission from PNAS (Madsen et al. 2018)
At the molecular level, M2 protein interacts with cholesterol in several ways to facilitate the budding and release of IAV particles. They are both integrated into the plasma membrane and can therefore interact by direct contact. Cholesterol molecules are embedded within the bilayer membrane, though typically asymmetrically distributed among the two leaflets in a nonstatic fashion that is subject to regulation by cellular processes (Wood et al. 2011). The functional consequences of cholesterol asymmetry likely include changes in membrane fluidity, alterations in lipid domains, lateral and transbilayer (“flip-flop”) diffusion, lipid packing, and perhaps even the function of some proteins (Wood et al. 2011). The M2 protein is seen to associate with cholesterol-enriched rafts in the plasma membrane primarily through association with other proteins such as M1 (Leser and Lamb 2017). Since the transmembrane domain of M2 traverses the membrane, it is tempting to speculate that cholesterol molecules can get in direct contact with this region of the M2 protein. Support for this idea comes from sequence analysis of M2, which reveals a cholesterol recognition/interaction amino acid consensus (CRAC) sequence motif thought to be involved with cholesterol recognition in other proteins, as well as the observation that M2 co-purifies with cholesterol (Schroeder et al. 2005). However, extensive mutagenesis studies have shown that the association between cholesterol and M2, and the proper functioning of M2 in facilitating virus replication, are not affected when the CRAC motif is deleted (Thaa et al. 2012; Stewart et al. 2010; Thaa et al. 2014; Thaa et al. 2011). In fact, some strains of IAV do not contain a M2 CRAC motif at all. We must therefore conclude that the CRAC motif is of no major functional importance in this context. Instead, the way that cholesterol and the M2 protein interact is likely through their ability to participate in lipid rafts or through the affinity of the M2 TM domain with cholesterol, positioning it to facilitate scission (Fig. 16.6) (Pan et al. 2019; Kolokouris et al. 2021; Rossman and Lamb 2010; Elkins et al. 2017). The presence of cholesterol in the plasma membrane might further have an impact on the structure and orientation of the M2 protein (Ekanayake et al. 2016; Wright et al. 2022; Claridge et al. 2013). Cholesterol has been shown to increase the insertion depth of the M2 protein in the lipid bilayer, bringing it closer to the hydrophobic core of the bilayer (Elkins et al. 2017; Martyna et al. 2020).
The key to elucidating the interplay between cholesterol and M2 in budding and scission will be to understand how their individual effects on membrane fluidity and mechanical properties add up when they colocalize in the budding neck. Is the sum greater than the parts, and how does the presence of M2 in low copy number (14 to 68 molecules per virion (Zebedee and Lamb 1988)) tip the balance to facilitate membrane scission? There is evidence that lipid phase separation phenomena and membrane scission are closely related (Roux et al. 2005; Ryu et al. 2014). Lipid phase separation refers to the spontaneous segregation of different species in the plasma membrane into distinct domains, which can form spontaneously due to differences in lipid composition or physical properties such as size, shape, and charge. We remark that this is distinct from the occurrence of lipid rafts, which are dynamic in nature and extend over much smaller spatial and temporal scales (and therefore cannot be resolved by microscopic methods) (Rajendran and Simons 2005; Lozano et al. 2016). The phase separation process results in the formation of nonuniform lipid structures, including liquid-ordered and liquid-disordered domains (Heberle and Feigenson 2011). The interface between the distinct phases is the focal point of mechanical forces such as tension, shear, and bending stresses potentially causing membrane scission (Liu et al. 2006). A variety of factors may contribute to membrane scission: Changes in lipid composition, to which both cholesterol and M2 protein contribute, changes in mechanical stress, likely caused by lipid phase boundary dynamics, and the action of additional specific proteins. Lipid raft structures have also been proposed to play a role in scission (Schroeder et al. 2005; Barman and Nayak 2007).
The AH domain within M2 acts as a membrane-sculpting tool (Huang et al. 2015). An AH is characterized by a Janus-faced physical/chemical surface consisting of both hydrophobic and hydrophilic amino acids residues concentrated on opposing sides of the α-helix structure. This arrangement allows the AH to insert into the plasma membrane right at the hydrophilic/hydrophobic interface region so that the hydrophobic residues interact with the lipid tails while the hydrophilic residues interact with the aqueous environment. The M2AH can facilitate budding and scission by inducing changes, primarily in composition, curvature, and stability, that help generate the appropriate lipid environment for budding and membrane scission to occur in model membrane systems, even in the absence of the other M2 domains (Fig. 16.5, lower panel) (Rossman et al. 2010b; Roberts et al. 2013; Martyna et al. 2017; Martyna et al. 2020). The effect appears most dependent on the chemical/physical properties of the AH domain and not dependent on the conserved amino acid motif per se, as one might have been tempted to assume (Hu et al. 2020). Furthermore, the conformation of the M2 protein is pH-dependent and can be altered when there are changes in pH or lipid composition, particularly when cholesterol is present, an effect that is likely facilitated by the protein’s AH domain (Torabifard et al. 2020; Liao et al. 2013; Liao et al. 2015; Kim et al. 2015; Thomaston et al. 2013; Saotome et al. 2015; Martyna et al. 2020; Kwon et al. 2015; Kyaw et al. 2023). While proteins, in general, can adapt conformations to suit different lipid environments, the opposite may be an important additional factor in some situations (Bacle et al. 2021).
Several other proteins and cellular processes have been shown to play a role in the processes of budding and membrane scission mediated by cholesterol and the M2 protein of IAV (Mao et al. 2022; Bruce et al. 2010; Han et al. 2021; Zhu et al. 2017; Wohlgemuth et al. 2018; Beale et al. 2014; Bhowmick et al. 2017; Vahey and Fletcher 2019). It is important to note that the exact roles of these factors are not fully understood, and additional research is needed to reveal the complex network of interactions that drive these processes.
Conclusion
The sophisticated interplay between cholesterol and the viral M2 protein plays a crucial role in multiple stages of the influenza virus life cycle. Cholesterol is essential for the formation of membrane raft microdomains that serve as virus assembly sites and contribute to the mechanical forces necessary for membrane scission during the last stage of virus budding. Furthermore, cholesterol helps to maintain the stability of the virus envelope. M2 protein also plays a critical role in influenza virus budding by promoting the formation of curved lipid bilayer structures that facilitate the process of membrane scission. The interaction between cholesterol and the M2 protein affects the local lipid composition, fluidity, and stability of the membrane, thus contributing to the efficient and proper release of newly formed influenza viruses from virus-infected cells.
In closing, investigation of the intricacies of the mechanistic steps of the influenza virus life cycle is critical because of the ongoing threat of influenza epidemics and pandemics. The knowledge gained from studying the interplay between cholesterol, the viral M2 protein and other associated components in influenza virus budding can contribute to the development of novel antiviral strategies and strengthen our efforts to combat viral infections and protect public health.
References
Allen MP, Tildesley DJ (1989) Computer simulation of liquids. Oxford science publications, Pbk. (with corr.). edn. Clarendon Press, Oxford
Anderson RH, Sochacki KA, Vuppula H, Scott BL, Bailey EM, Schultz MM, Kerkvliet JG, Taraska JW, Hoppe AD, Francis KR (2021) Sterols lower energetic barriers of membrane bending and fission necessary for efficient clathrin-mediated endocytosis. Cell Rep 37(7):110008. https://doi.org/10.1016/j.celrep.2021.110008
Andreas LB, Reese M, Eddy MT, Gelev V, Ni QZ, Miller EA, Emsley L, Pintacuda G, Chou JJ, Griffin RG (2015) Structure and mechanism of the Influenza A M218-60 dimer of dimers. J Am Chem Soc 137(47):14877–14886. https://doi.org/10.1021/jacs.5b04802
Bacle A, Buslaev P, Garcia-Fandino R, Favela-Rosales F, Mendes Ferreira T, Fuchs PFJ, Gushchin I, Javanainen M, Kiirikki AM, Madsen JJ, Melcr J, Milan Rodriguez P, Miettinen MS, Ollila OHS, Papadopoulos CG, Peon A, Piggot TJ, Pineiro A, Virtanen SI (2021) Inverse conformational selection in lipid-protein binding. J Am Chem Soc 143(34):13701–13709. https://doi.org/10.1021/jacs.1c05549
Bajimaya S, Frankl T, Hayashi T, Takimoto T (2017) Cholesterol is required for stability and infectivity of influenza A and respiratory syncytial viruses. Virology 510:234–241. https://doi.org/10.1016/j.virol.2017.07.024
Bao D, Lu C, Ma T, Xu G, Mao Y, Xin L, Niu S, Wu Z, Li X, Teng Q, Li Z, Liu Q (2022) Hydrophobic residues at the intracellular domain of the M2 protein play an important role in budding and membrane integrity of influenza virus. J Virol 96(9):e0037322. https://doi.org/10.1128/jvi.00373-22
Barman S, Nayak DP (2007) Lipid raft disruption by cholesterol depletion enhances influenza A virus budding from MDCK cells. J Virol 81(22):12169–12178. https://doi.org/10.1128/JVI.00835-07
Beale R, Wise H, Stuart A, Ravenhill BJ, Digard P, Randow F (2014) A LC3-interacting motif in the influenza A virus M2 protein is required to subvert autophagy and maintain virion stability. Cell Host Microbe 15(2):239–247. https://doi.org/10.1016/j.chom.2014.01.006
Bhowmick S, Chakravarty C, Sellathamby S, Lal SK (2017) The influenza A virus matrix protein 2 undergoes retrograde transport from the endoplasmic reticulum into the cytoplasm and bypasses cytoplasmic proteasomal degradation. Arch Virol 162(4):919–929. https://doi.org/10.1007/s00705-016-3153-8
Boban Z, Mardesic I, Subczynski WK, Raguz M (2021) Giant unilamellar vesicle electroformation: what to use, what to avoid, and how to quantify the results. Membranes (Basel) 11(11):860. https://doi.org/10.3390/membranes11110860
Bourmakina SV, Garcia-Sastre A (2005) The morphology and composition of influenza A virus particles are not affected by low levels of M1 and M2 proteins in infected cells. J Virol 79(12):7926–7932. https://doi.org/10.1128/JVI.79.12.7926-7932.2005
Bruce EA, Digard P, Stuart AD (2010) The Rab11 pathway is required for influenza A virus budding and filament formation. J Virol 84(12):5848–5859. https://doi.org/10.1128/JVI.00307-10
Chen BJ, Leser GP, Jackson D, Lamb RA (2008) The influenza virus M2 protein cytoplasmic tail interacts with the M1 protein and influences virus assembly at the site of virus budding. J Virol 82(20):10059–10070. https://doi.org/10.1128/JVI.01184-08
Claridge JK, Aittoniemi J, Cooper DM, Schnell JR (2013) Isotropic bicelles stabilize the juxtamembrane region of the influenza M2 protein for solution NMR studies. Biochemistry 52(47):8420–8429. https://doi.org/10.1021/bi401035m
Ekanayake EV, Fu R, Cross TA (2016) Structural influences: cholesterol, drug, and proton binding to full-length Influenza A M2 protein. Biophys J 110(6):1391–1399. https://doi.org/10.1016/j.bpj.2015.11.3529
Elkins MR, Williams JK, Gelenter MD, Dai P, Kwon B, Sergeyev IV, Pentelute BL, Hong M (2017) Cholesterol-binding site of the influenza M2 protein in lipid bilayers from solid-state NMR. Proc Natl Acad Sci U S A 114(49):12946–12951. https://doi.org/10.1073/pnas.1715127114
Elkins MR, Sergeyev IV, Hong M (2018) Determining cholesterol binding to membrane proteins by cholesterol (13) C labeling in yeast and dynamic nuclear polarization NMR. J Am Chem Soc 140(45):15437–15449. https://doi.org/10.1021/jacs.8b09658
Elleman CJ, Barclay WS (2004) The M1 matrix protein controls the filamentous phenotype of influenza A virus. Virology 321(1):144–153. https://doi.org/10.1016/j.virol.2003.12.009
Fan TW, Lane AN (2016) Applications of NMR spectroscopy to systems biochemistry. Prog Nucl Magn Reson Spectrosc 92-93:18–53. https://doi.org/10.1016/j.pnmrs.2016.01.005
Frenkel D, Smit B, ProQuest (2002) Understanding molecular simulation: from algorithms to applications, Computational science series, vol 1, 2nd edn. Academic Press, San Diego
Goldsmith CS, Miller SE (2009) Modern uses of electron microscopy for detection of viruses. Clin Microbiol Rev 22(4):552–563. https://doi.org/10.1128/CMR.00027-09
Grime JMA, Madsen JJ (2019) Efficient simulation of tunable lipid assemblies across scales and resolutions. arXiv:191005362. https://doi.org/10.48550/arXiv.1910.05362
Han J, Ganti K, Sali VK, Twigg C, Zhang Y, Manivasagam S, Liang CY, Vogel OA, Huang I, Emmanuel SN, Plung J, Radoshevich L, Perez JT, Lowen AC, Manicassamy B (2021) Host factor Rab11a is critical for efficient assembly of influenza A virus genomic segments. PLoS Pathog 17(5):e1009517. https://doi.org/10.1371/journal.ppat.1009517
Harry EJ, Pogliano K, Losick R (1995) Use of immunofluorescence to visualize cell-specific gene expression during sporulation in Bacillus subtilis. J Bacteriol 177(12):3386–3393. https://doi.org/10.1128/jb.177.12.3386-3393.1995
Heberle FA, Feigenson GW (2011) Phase separation in lipid membranes. Cold Spring Harb Perspect Biol 3(4):a004630. https://doi.org/10.1101/cshperspect.a004630
Helenius A (1992) Unpacking the incoming influenza virus. Cell 69(4):577–578. https://doi.org/10.1016/0092-8674(92)90219-3
Herneisen AL, Sahu ID, McCarrick RM, Feix JB, Lorigan GA, Howard KP (2017) A budding-defective M2 mutant exhibits reduced membrane interaction, insensitivity to cholesterol, and perturbed interdomain coupling. Biochemistry 56(44):5955–5963. https://doi.org/10.1021/acs.biochem.7b00924
Ho CS, Khadka NK, She F, Cai J, Pan J (2016) Influenza M2 transmembrane domain senses membrane heterogeneity and enhances membrane curvature. Langmuir 32(26):6730–6738. https://doi.org/10.1021/acs.langmuir.6b00150
Hu B, Siche S, Moller L, Veit M (2020) Amphipathic helices of cellular proteins can replace the helix in M2 of Influenza A virus with only small effects on virus replication. J Virol 94:3. https://doi.org/10.1128/JVI.01605-19
Huang S, Green B, Thompson M, Chen R, Thomaston J, DeGrado WF, Howard KP (2015) C-terminal juxtamembrane region of full-length M2 protein forms a membrane surface associated amphipathic helix. Protein Sci 24(3):426–429. https://doi.org/10.1002/pro.2631
Hubert M, Larsson E, Vegesna NVG, Ahnlund M, Johansson AI, Moodie LW, Lundmark R (2020) Lipid accumulation controls the balance between surface connection and scission of caveolae. elife 9:e55038. https://doi.org/10.7554/eLife.55038
Humphrey W, Dalke A, Schulten K (1996) VMD: visual molecular dynamics. J Mol Graph 14(1):33–38, 27-38. https://doi.org/10.1016/0263-7855(96)00018-5
Hunter JD (2007) Matplotlib: a 2D graphics environment. Comput Sci Eng 9(3):90–95
Ingolfsson HI, Lopez CA, Uusitalo JJ, de Jong DH, Gopal SM, Periole X, Marrink SJ (2014) The power of coarse graining in biomolecular simulations. Wiley Interdiscip Rev Comput Mol Sci 4(3):225–248. https://doi.org/10.1002/wcms.1169
Jarin Z, Newhouse J, Voth GA (2021) Coarse-grained force fields from the perspective of statistical mechanics: better understanding of the origins of a MARTINI hangover. J Chem Theory Comput 17(2):1170–1180. https://doi.org/10.1021/acs.jctc.0c00638
Javanainen M, Heftberger P, Madsen JJ, Miettinen MS, Pabst G, Ollila OHS (2023) Quantitative comparison against experiments reveals imperfections in force fields’ descriptions of popc–cholesterol interactions. J Chem Theory Comput. In press. https://doi.org/10.1021/acs.jctc.3c00648
Jo S, Kim T, Iyer VG, Im W (2008) CHARMM-GUI: a web-based graphical user interface for CHARMM. J Comput Chem 29(11):1859–1865. https://doi.org/10.1002/jcc.20945
Kalil AC, Thomas PG (2019) Influenza virus-related critical illness: pathophysiology and epidemiology. Crit Care 23(1):258. https://doi.org/10.1186/s13054-019-2539-x
Kim SS, Upshur MA, Saotome K, Sahu ID, McCarrick RM, Feix JB, Lorigan GA, Howard KP (2015) Cholesterol-dependent conformational exchange of the C-terminal domain of the Influenza A M2 protein. Biochemistry 54(49):7157–7167. https://doi.org/10.1021/acs.biochem.5b01065
Klug CS, Feix JB (2008) Methods and applications of site-directed spin labeling EPR spectroscopy. Methods Cell Biol 84:617–658. https://doi.org/10.1016/S0091-679X(07)84020-9
Kolokouris D, Kalenderoglou IE, Kolocouris A (2021) Inside and out of the pore: comparing interactions and molecular dynamics of Influenza A M2 viroporin complexes in standard lipid bilayers. J Chem Inf Model 61(11):5550–5568. https://doi.org/10.1021/acs.jcim.1c00264
Kwon B, Tietze D, White PB, Liao SY, Hong M (2015) Chemical ligation of the influenza M2 protein for solid-state NMR characterization of the cytoplasmic domain. Protein Sci 24(7):1087–1099. https://doi.org/10.1002/pro.2690
Kyaw A, Roepke K, Arthur T, Howard KP (2023) Conformation of influenza AM2 membrane protein in nanodiscs and liposomes. Biochim Biophys Acta Biomembr 1865(5):184152. https://doi.org/10.1016/j.bbamem.2023.184152
Lakadamyali M, Rust MJ, Zhuang X (2004) Endocytosis of influenza viruses. Microbes Infect 6(10):929–936. https://doi.org/10.1016/j.micinf.2004.05.002
Lamb RA, Holsinger LJ, Pinto LH (1994) Receptor-mediated virus entry into cells. Cold Spring Harbor Laboratory Press, Cold Spring Harbor
Lenard J, Compans RW (1974) The membrane structure of lipid-containing viruses. Biochim Biophys Acta 344(1):51–94. https://doi.org/10.1016/0304-4157(74)90008-2
Leser GP, Lamb RA (2017) Lateral organization of influenza virus proteins in the Budozone region of the plasma membrane. J Virol 91(9):e02104-16. https://doi.org/10.1128/JVI.02104-16
Leser GP, Ector KJ, Lamb RA (1996) The paramyxovirus simian virus 5 hemagglutinin-neuraminidase glycoprotein, but not the fusion glycoprotein, is internalized via coated pits and enters the endocytic pathway. Mol Biol Cell 7(1):155–172. https://doi.org/10.1091/mbc.7.1.155
Liao SY, Fritzsching KJ, Hong M (2013) Conformational analysis of the full-length M2 protein of the influenza A virus using solid-state NMR. Protein Sci 22(11):1623–1638. https://doi.org/10.1002/pro.2368
Liao SY, Yang Y, Tietze D, Hong M (2015) The influenza m2 cytoplasmic tail changes the proton-exchange equilibria and the backbone conformation of the transmembrane histidine residue to facilitate proton conduction. J Am Chem Soc 137(18):6067–6077. https://doi.org/10.1021/jacs.5b02510
Liu J, Kaksonen M, Drubin DG, Oster G (2006) Endocytic vesicle scission by lipid phase boundary forces. Proc Natl Acad Sci U S A 103(27):10277–10282. https://doi.org/10.1073/pnas.0601045103
Lozano MM, Hovis JS, Moss FR 3rd, Boxer SG (2016) Dynamic reorganization and correlation among lipid raft components. J Am Chem Soc 138(31):9996–10001. https://doi.org/10.1021/jacs.6b05540
Madsen JJ, Sinitskiy AV, Li J, Voth GA (2017) Highly coarse-grained representations of transmembrane proteins. J Chem Theory Comput 13(2):935–944. https://doi.org/10.1021/acs.jctc.6b01076
Madsen JJ, Grime JMA, Rossman JS, Voth GA (2018) Entropic forces drive clustering and spatial localization of influenza A M2 during viral budding. Proc Natl Acad Sci U S A 115(37):E8595–E8603. https://doi.org/10.1073/pnas.1805443115
Mao H, Cao L, Xu T, Xia X, Ren P, Han P, Li C, Hui X, Lin X, Huang K, Jin M (2022) YWHAG inhibits influenza a virus replication by suppressing the release of viral M2 protein. Front Microbiol 13:951009. https://doi.org/10.3389/fmicb.2022.951009
Marion D (2013) An introduction to biological NMR spectroscopy. Mol Cell Proteomics 12(11):3006–3025. https://doi.org/10.1074/mcp.O113.030239
Marrink SJ, Risselada HJ, Yefimov S, Tieleman DP, de Vries AH (2007) The MARTINI force field: coarse grained model for biomolecular simulations. J Phys Chem B 111(27):7812–7824. https://doi.org/10.1021/jp071097f
Martyna A, Gomez-Llobregat J, Linden M, Rossman JS (2016) Curvature sensing by a viral scission protein. Biochemistry 55(25):3493–3496. https://doi.org/10.1021/acs.biochem.6b00539
Martyna A, Bahsoun B, Badham MD, Srinivasan S, Howard MJ, Rossman JS (2017) Membrane remodeling by the M2 amphipathic helix drives influenza virus membrane scission. Sci Rep 7:44695. https://doi.org/10.1038/srep44695
Martyna A, Bahsoun B, Madsen JJ, Jackson F, Badham MD, Voth GA, Rossman JS (2020) Cholesterol alters the orientation and activity of the influenza virus M2 amphipathic helix in the membrane. J Phys Chem B 124(31):6738–6747. https://doi.org/10.1021/acs.jpcb.0c03331
McCown MF, Pekosz A (2006) Distinct domains of the influenza a virus M2 protein cytoplasmic tail mediate binding to the M1 protein and facilitate infectious virus production. J Virol 80(16):8178–8189. https://doi.org/10.1128/JVI.00627-06
Mouritsen OG (2016) Life - as a matter of fat. Softcover reprint of the original, 2nd 2016. edn. Springer International Publishing A+, Cham
Mouritsen OG, Jorgensen K (1992) Dynamic lipid-bilayer heterogeneity: a mesoscopic vehicle for membrane function? BioEssays 14(2):129–136. https://doi.org/10.1002/bies.950140211
Nayak DP, Barman S (2002) Role of lipid rafts in virus assembly and budding. Adv Virus Res 58:1–28. https://doi.org/10.1016/s0065-3527(02)58001-5
Nermut MV, Frank H (1971) Fine structure of influenza A2 (Singapore) as revealed by negative staining, freeze-drying and freeze-etching. J Gen Virol 10(1):37–51. https://doi.org/10.1099/0022-1317-10-1-37
Noda T, Kawaoka Y (2010) Structure of influenza virus ribonucleoprotein complexes and their packaging into virions. Rev Med Virol 20(6):380–391. https://doi.org/10.1002/rmv.666
Ohkubo YZ, Madsen JJ (2021) Uncovering membrane-bound models of coagulation factors by combined experimental and computational approaches. Thromb Haemost 121(9):1122–1137. https://doi.org/10.1055/s-0040-1722187
Pak AJ, Dannenhoffer-Lafage T, Madsen JJ, Voth GA (2019) Systematic coarse-grained lipid force fields with semiexplicit solvation via virtual sites. J Chem Theory Comput 15(3):2087–2100. https://doi.org/10.1021/acs.jctc.8b01033
Pan J, Dalzini A, Song L (2019) Cholesterol and phosphatidylethanolamine lipids exert opposite effects on membrane modulations caused by the M2 amphipathic helix. Biochim Biophys Acta Biomembr 1861(1):201–209. https://doi.org/10.1016/j.bbamem.2018.07.013
Paulino J, Pang X, Hung I, Zhou HX, Cross TA (2019) Influenza A M2 channel clustering at high protein/lipid ratios: viral budding implications. Biophys J 116(6):1075–1084. https://doi.org/10.1016/j.bpj.2019.01.042
Pinto LH, Holsinger LJ, Lamb RA (1992) Influenza virus M2 protein has ion channel activity. Cell 69(3):517–528. https://doi.org/10.1016/0092-8674(92)90452-i
Rajendran L, Simons K (2005) Lipid rafts and membrane dynamics. J Cell Sci 118(Pt 6):1099–1102. https://doi.org/10.1242/jcs.01681
Roberts PC, Lamb RA, Compans RW (1998) The M1 and M2 proteins of influenza A virus are important determinants in filamentous particle formation. Virology 240(1):127–137. https://doi.org/10.1006/viro.1997.8916
Roberts KL, Leser GP, Ma C, Lamb RA (2013) The amphipathic helix of influenza A virus M2 protein is required for filamentous bud formation and scission of filamentous and spherical particles. J Virol 87(18):9973–9982. https://doi.org/10.1128/JVI.01363-13
Rossman JS, Lamb RA (2010) Swine-origin influenza virus and the 2009 pandemic. Am J Respir Crit Care Med 181(4):295–296. https://doi.org/10.1164/rccm.200912-1876ED
Rossman JS, Jing X, Leser GP, Balannik V, Pinto LH, Lamb RA (2010a) Influenza virus m2 ion channel protein is necessary for filamentous virion formation. J Virol 84(10):5078–5088. https://doi.org/10.1128/JVI.00119-10
Rossman JS, Jing X, Leser GP, Lamb RA (2010b) Influenza virus M2 protein mediates ESCRT-independent membrane scission. Cell 142(6):902–913. https://doi.org/10.1016/j.cell.2010.08.029
Roux A, Cuvelier D, Nassoy P, Prost J, Bassereau P, Goud B (2005) Role of curvature and phase transition in lipid sorting and fission of membrane tubules. EMBO J 24(8):1537–1545. https://doi.org/10.1038/sj.emboj.7600631
Ryu YS, Lee IH, Suh JH, Park SC, Oh S, Jordan LR, Wittenberg NJ, Oh SH, Jeon NL, Lee B, Parikh AN, Lee SD (2014) Reconstituting ring-rafts in bud-mimicking topography of model membranes. Nat Commun 5:4507. https://doi.org/10.1038/ncomms5507
Saotome K, Duong-Ly KC, Howard KP (2015) Influenza A M2 protein conformation depends on choice of model membrane. Biopolymers 104(4):405–411. https://doi.org/10.1002/bip.22617
Scaffidi SJ, Shebes MA, Yu W (2021) Tracking the subcellular localization of surface proteins in Staphylococcus aureus by immunofluorescence microscopy. Bio Protoc 11(10):e4038. https://doi.org/10.21769/BioProtoc.4038
Schmidt NW, Mishra A, Wang J, DeGrado WF, Wong GC (2013) Influenza virus A M2 protein generates negative Gaussian membrane curvature necessary for budding and scission. J Am Chem Soc 135(37):13710–13719. https://doi.org/10.1021/ja400146z
Schroeder C, Heider H, Moncke-Buchner E, Lin TI (2005) The influenza virus ion channel and maturation cofactor M2 is a cholesterol-binding protein. Eur Biophys J 34(1):52–66. https://doi.org/10.1007/s00249-004-0424-1
Schulze IT (1972) The structure of influenza virus. II. A model based on the morphology and composition of subviral particles. Virology 47(1):181–196. https://doi.org/10.1016/0042-6822(72)90251-6
Sharma M, Yi M, Dong H, Qin H, Peterson E, Busath DD, Zhou HX, Cross TA (2010) Insight into the mechanism of the influenza A proton channel from a structure in a lipid bilayer. Science 330(6003):509–512. https://doi.org/10.1126/science.1191750
Slot JW, Geuze HJ (1985) A new method of preparing gold probes for multiple-labeling cytochemistry. Eur J Cell Biol 38(1):87–93
Stewart SM, Wu WH, Lalime EN, Pekosz A (2010) The cholesterol recognition/interaction amino acid consensus motif of the influenza A virus M2 protein is not required for virus replication but contributes to virulence. Virology 405(2):530–538. https://doi.org/10.1016/j.virol.2010.06.035
Sun X, Whittaker GR (2003) Role for influenza virus envelope cholesterol in virus entry and infection. J Virol 77(23):12543–12551. https://doi.org/10.1128/jvi.77.23.12543-12551.2003
Sutherland M, Tran N, Hong M (2022) Clustering of tetrameric influenza M2 peptides in lipid bilayers investigated by (19)F solid-state NMR. Biochim Biophys Acta Biomembr 1864(7):183909. https://doi.org/10.1016/j.bbamem.2022.183909
Swanson J, Hsu KC, Gotschlich EC (1969) Electron microscopic studies on streptococci. I. M antigen. J Exp Med 130(5):1063–1091. https://doi.org/10.1084/jem.130.5.1063
Taubenberger JK, Morens DM (2008) The pathology of influenza virus infections. Annu Rev Pathol 3:499–522. https://doi.org/10.1146/annurev.pathmechdis.3.121806.154316
Thaa B, Levental I, Herrmann A, Veit M (2011) Intrinsic membrane association of the cytoplasmic tail of influenza virus M2 protein and lateral membrane sorting regulated by cholesterol binding and palmitoylation. Biochem J 437(3):389–397. https://doi.org/10.1042/BJ20110706
Thaa B, Tielesch C, Moller L, Schmitt AO, Wolff T, Bannert N, Herrmann A, Veit M (2012) Growth of influenza A virus is not impeded by simultaneous removal of the cholesterol-binding and acylation sites in the M2 protein. J Gen Virol 93(Pt 2):282–292. https://doi.org/10.1099/vir.0.038554-0
Thaa B, Siche S, Herrmann A, Veit M (2014) Acylation and cholesterol binding are not required for targeting of influenza A virus M2 protein to the hemagglutinin-defined budozone. FEBS Lett 588(6):1031–1036. https://doi.org/10.1016/j.febslet.2014.02.014
Thomaston JL, Nguyen PA, Brown EC, Upshur MA, Wang J, DeGrado WF, Howard KP (2013) Detection of drug-induced conformational change of a transmembrane protein in lipid bilayers using site-directed spin labeling. Protein Sci 22(1):65–73. https://doi.org/10.1002/pro.2186
Torabifard H, Panahi A, Brooks CL 3rd (2020) M2 amphipathic helices facilitate pH-dependent conformational transition in influenza A virus. Proc Natl Acad Sci U S A 117(7):3583–3591. https://doi.org/10.1073/pnas.1913385117
Vahey MD, Fletcher DA (2019) Low-fidelity assembly of Influenza A virus promotes escape from host cells. Cell 176(1–2):281–294. e219. https://doi.org/10.1016/j.cell.2018.10.056
Votteler J, Sundquist WI (2013) Virus budding and the ESCRT pathway. Cell Host Microbe 14(3):232–241. https://doi.org/10.1016/j.chom.2013.08.012
Wang T, Hong M (2015) Investigation of the curvature induction and membrane localization of the influenza virus M2 protein using static and off-magic-angle spinning solid-state nuclear magnetic resonance of oriented bicelles. Biochemistry 54(13):2214–2226. https://doi.org/10.1021/acs.biochem.5b00127
Wang T, Cady SD, Hong M (2012) NMR determination of protein partitioning into membrane domains with different curvatures and application to the influenza M2 peptide. Biophys J 102(4):787–794. https://doi.org/10.1016/j.bpj.2012.01.010
Waterson AP, Hurrell JM, Jensen KE (1963) The fine structure of influenza A, B and C viruses. Arch Gesamte Virusforsch 12:487–495. https://doi.org/10.1007/BF01242156
Wohlgemuth N, Lane AP, Pekosz A (2018) Influenza A virus M2 protein apical targeting is required for efficient virus replication. J Virol 92(22):e01425-18. https://doi.org/10.1128/JVI.01425-18
Wood WG, Igbavboa U, Muller WE, Eckert GP (2011) Cholesterol asymmetry in synaptic plasma membranes. J Neurochem 116(5):684–689. https://doi.org/10.1111/j.1471-4159.2010.07017.x
Wright AK, Paulino J, Cross TA (2022) Emulating membrane protein environments horizontal line how much lipid is required for a native structure: influenza S31N M2. J Am Chem Soc 144(5):2137–2148. https://doi.org/10.1021/jacs.1c10174
Wu EL, Cheng X, Jo S, Rui H, Song KC, Davila-Contreras EM, Qi Y, Lee J, Monje-Galvan V, Venable RM, Klauda JB, Im W (2014) CHARMM-GUI membrane builder toward realistic biological membrane simulations. J Comput Chem 35(27):1997–2004. https://doi.org/10.1002/jcc.23702
Zebedee SL, Lamb RA (1988) Influenza A virus M2 protein: monoclonal antibody restriction of virus growth and detection of M2 in virions. J Virol 62(8):2762–2772. https://doi.org/10.1128/JVI.62.8.2762-2772.1988
Zhu P, Liang L, Shao X, Luo W, Jiang S, Zhao Q, Sun N, Zhao Y, Li J, Wang J, Zhou Y, Zhang J, Wang G, Jiang L, Chen H, Li C (2017) Host cellular protein TRAPPC6ADelta interacts with Influenza A virus M2 protein and regulates viral propagation by modulating M2 trafficking. J Virol 91(1):e01757-16. https://doi.org/10.1128/JVI.01757-16
Further Reading
Chlanda P, Zimmerberg J (2016) Protein-lipid interactions critical to replication of the influenza A virus. FEBS Lett 590:1940–1954
Heaton NS, Randall G (2011) Multifaceted roles for lipids in viral infection. Trends Microbiol 19:368–375
Lamb RA (2020) The structure, function, and pathobiology if the Influenza A and B virus ion channels. Cold Spring Harb Perspect Med 10:a038505. https://doi.org/10.1101/cshperspect.a038505
Martyna A, Rossman JS (2014) Alterations of membrane curvature during influenza virus budding. Biochem Soc Trans 42:1425–1428
Rossman JS, Lamb RA (2011) Influenza virus assembly and budding. Virology 411:229–236
Rossman JS, Lamb RA (2013) Viral membrane scission. Annu Rev. Cell Dev Biol 29:551–569
Schroeder C (2010) Cholesterol-binding viral proteins in virus entry and morphogenesis. Subcell Biochem 51:77–108
Veit M, Thaa B (2011) Association of influenza virus proteins with membrane rafts. Adv Virol 2011:370606
Zhou H-X, Cross TA (2013) Modeling the membrane environment has implications for membrane protein structure and function: Influenza A M2 protein. Protein Sci 22:381–394
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this chapter
Cite this chapter
Madsen, J.J., Rossman, J.S. (2023). Cholesterol and M2 Rendezvous in Budding and Scission of Influenza A Virus. In: Vijayakrishnan, S., Jiu, Y., Harris, J.R. (eds) Virus Infected Cells. Subcellular Biochemistry, vol 106. Springer, Cham. https://doi.org/10.1007/978-3-031-40086-5_16
Download citation
DOI: https://doi.org/10.1007/978-3-031-40086-5_16
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-40085-8
Online ISBN: 978-3-031-40086-5
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)