Abstract
The Cytomatrix Assembled at the active Zone (CAZ) of a presynaptic terminal displays electron-dense appearance and defines the center of the synaptic vesicle release. The protein constituents of CAZ are multiple-domain scaffolds that interact extensively with each other and also with an ensemble of synaptic vesicle proteins to ensure docking, fusion, and recycling. Reflecting the central roles of the active zone in synaptic transmission, CAZ proteins are highly conserved throughout evolution. As the nervous system increases complexity and diversity in types of neurons and synapses, CAZ proteins expand in the number of gene and protein isoforms and interacting partners. This chapter summarizes the discovery of the core CAZ proteins and current knowledge of their functions.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Presynaptic active zone
- Munc13
- UNC-13
- Rim
- UNC-10
- RIM-BP
- ELKS
- Bruchpilot
- Fife
- CLA-1
- Bassoon
- Piccolo
- Liprin-α
- SYD-2
- SYD-1
- CASK
1 Introduction
The appearance of an electron-dense matrix associated with patches of the axonal plasma membrane and surrounded by small clusters of vesicles under electron microscopy has been taken as the morphological landmark of a presynaptic terminal, often named active zone (Chapter “The Architecture of the Presynaptic Release Site”). Molecular identification of presynaptic components began with the ingenious invention of biochemical preparation of synaptosomes, developed by Victor Whittaker and coworkers [1]. Combined with technology advances in mass spectrometry proteomics, thousands of distinct presynaptic proteins have been identified, culminating to a landmark study by Reinhart Jahn and coworkers, which reports 410 proteins associated with a single synaptic vesicle [2]. However, the constituents of the Cytomatrix Assembled at the active Zone (CAZ) tend to be insoluble in biochemical purifications, and some proteins may be present in selective types of synapses or transiently associate with synapses. It took additional approaches, such as antibody-based protein expression screening, protein-interaction screening, and molecular genetics in model organisms, to unveil the identities of CAZ proteins.
It is generally agreed that CAZ proteins fall into three main functional categories. First are the classical cytoskeletal proteins corresponding to actin, tubulin, myosin, spectrin α chain and β chain, and β-catenin. They are the fundamental elements of the cytoskeletal framework of active zone cytomatrix. Second are the adaptor and scaffold proteins, such as SAP90/PSD95/DLG4, SAP97/DLG1, and CASK/LIN-2. These proteins are not restricted to presynaptic active zones, also participate in clustering postsynaptic receptors, and are involved in the organization of a variety of cell junctions. If the cytoskeleton proteins form a grid-like structure at the active zone, these proteins probably link the ion channels and the synaptic vesicle fusion machinery onto the grid to ensure proper active zone function. Third are the active-zone-specific CAZ proteins, represented by six evolutionarily conserved families known as Munc13/UNC-13, RIM, RIM-BP (RIM-binding protein), ELKS, Bassoon and Piccolo, and Liprin-α [3]. This chapter will focus on the discovery and function of these active-zone-specific CAZ components.
2 Experimental Approaches Used in the Identification of CAZ Proteins
We begin by offering a brief overview of the key approaches used to identify CAZ proteins.
2.1 Antibody-Based Protein Expression Screen
When researchers realized that the protein constituents of the electron-dense matrix in presynaptic terminals were low in abundance and problematic with solubility in biochemical purification, they sought to obtain antibodies against brain synaptic junctional proteins. The antibodies were used in immunocytochemistry on either brain tissues or cells to determine if the corresponding antigens were localized to presynaptic terminals [4, 5]. To search for the molecules that encode the antigens, researchers relied on a powerful technique, developed in the late 1980s, that enabled the production of any proteins in bacteriophage lambda [6]. In essence, mRNAs isolated from brain tissues were made into cDNAs, which were cloned into special expression vectors for protein production in bacteriophage lambda. The synapse-specific antibodies were used to recognize proteins produced from lambda. The amino acid sequences for candidate proteins were then deduced from the DNA sequences of the corresponding cDNA. This approach led to the identification of the CAZ proteins Bassoon and Piccolo in mammals [7, 8], and Bruchpilot in Drosophila [9].
2.2 Protein-Interaction-Based Screen
Around late 1980s, another powerful technique was developed to detect protein–protein interactions in yeast, named yeast-two-hybrid (Y2H) interaction assay [10]. The Y2H design was based on the finding that the transcriptional activity of the yeast protein Gal4 requires two modular protein domains, a DNA-binding (DB) domain and a transcription-activation domain (AD). When the DB and AD domains are in close proximity, transcription of genes encoding enzymes of galactose utilization can be activated, thereby allowing yeast to grow in galactose selection media. In a Y2H assay, a bait protein X, which can be either the full length or a fragment of the protein of interest, is fused to the Gal4(DB) domain, and potential prey proteins (Y) are fused to the Gal4(AD) domain. Upon co-expression in yeast, if protein X binds to protein Y, it will lead to reconstitution of Gal4 transcriptional activity. Thus, the Y2H assay does not rely on either solubility or abundance of target proteins and can be carried out on a large scale when a library of prey is used. However, the resulting candidate binding partners need to be verified using other biochemical assays and validated for expression in brain tissues. This approach led to the identification of the CAZ proteins RIM [11], RIM-Binding Protein [12], and ELKS [13].
2.3 Forward Genetic Screen for Mutants Affecting Synaptic Transmission
Around 1960s, the nematode Caenorhabditis elegans was chosen by the Nobel Laureate Sydney Brenner to study the development and function of the nervous system. He carried out the first forward genetic screen and isolated a large number of mutants that displayed a variety of abnormal patterns of movement, categorized as uncoordinated [14]. Subsequent molecular cloning of the genes related to the unc phenotypes and physiological studies began to uncover the synaptic basis of the uncoordinated movement [15]. By early 1990s, it became clear that genes acting in synaptic transmission are evolutionarily conserved. This notion fueled the efforts to search for homologs of C. elegans unc genes in mammals and other species based on DNA sequence similarity. For example, C. elegans unc-13 (uncoordinated-13) mutants are paralyzed and resistant to drugs that perturb synaptic transmission. Molecular cloning of unc-13 revealed that the predicted UNC-13 protein contains domains, known as C1 and C2, that can bind to Ca2+, phospholipids, and diacylglycerol [16]. Using unc-13 cDNA to screen a rat brain cDNA library then led to the discovery of its mammalian member named as Munc13 [17]. Protein expression studies further showed that Munc13 and UNC-13 localize to presynaptic active zone.
The nervous system of C. elegans is fully reconstructed at the ultrastructural level, providing the precise knowledge on the synapse number, position, and pattern for each neuron [18]. C. elegans is also optically transparent. With the advent of using GFP and other fluorescent proteins as non-invasive reporters in living C. elegans [19], researchers can observe any cellular morphology and compartment. In particular, transgenic reporters expressing chimeric proteins, in which GFP is fused in-frame to synaptic vesicle proteins, such as Synaptobrevin-1 (SNB-1::GFP), enabled the visualization of synapses [20, 21]. Combined with genome-wide chemical mutagenesis, mutants that displayed abnormal synapse morphology, position, and number were subsequently isolated [22]. Molecular cloning and expression studies of the corresponding genes showed that many proteins are localized to sub-compartments of presynaptic terminals. This approach led to the identification of the CAZ protein SYD-2/Liprin-α [23].
3 Summary of CAZ Proteins and Function
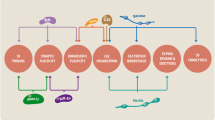
The active-zone-specific CAZ proteins are composed of multiple domains known for protein–protein, protein–lipid, and protein–ion bindings (Fig. 1a). They exhibit homomeric interactions and also bind extensively with other CAZ proteins, the synaptic plasma membrane, components of the synaptic cytoskeleton, and the synaptic vesicle recycling machinery (Fig. 1b). Like the conserved nature of synapse ultrastructure (Chapter “The Architecture of the Presynaptic Release Site”), CAZ proteins are highly conserved from invertebrates to human. Each family of CAZ proteins is typically encoded by a single gene in invertebrates, but multiple genes in vertebrates, reflecting the expansion of genomes in gene number and regulatory capacity. Regardless of species, each CAZ gene can produce several protein isoforms through the use of alternative promoters and alternative splicing, and these protein isoforms often show distinct dynamics and binding interactions depending on synapse type and neuronal activity state. Functional investigation of CAZ proteins using genetic malleable invertebrates has offered key insights into evolutionarily conserved mechanisms, while studies of CAZ proteins in mammalian nervous systems have both validated the commonality and also uncovered additional divergent themes. Here, we summarize general knowledge of each CAZ protein family (Fig. 1).
(a) Schematics of presynaptic CAZ proteins, using a representative full-length protein isoform for each family. Functional domains are marked following conventional designation (see main text and Ref. [3]). (b) Graphic illustration of CAZ protein-interacting network at the presynaptic active zone. (Modified from the graphic abstract in Ref. [47], provided by Mingjie Zhang)
3.1 Munc13/UNC-13
The Munc13/UNC-13 proteins are characterized by an ordered arrangement of three C2 domains (designated as C2A, C2B, C2C), a calmodulin-binding domain, a C1 domain that binds lipid, and an MUN domain that binds to the SNARE protein syntaxin and the SM protein Munc18/UNC-18 [24, 25] (Fig. 1a). Mammals have five Munc13 genes, with Munc13-1, -2, and -3 being abundantly expressed in CNS synapses. Alternative protein isoforms produced from each gene can vary in the amino acid linker sequences between the identified domains and in the number of C2 domains. C. elegans and Drosophila each has only one such gene, which also produces several protein isoforms. Munc13/UNC-13 proteins decorate the center of the presynaptic active zone.
The key function of this protein family is to prime synaptic vesicles for fast exocytosis. The first evidence came from electrophysiological studies of the unc-13 mutants, which revealed a complete abolishment of neurotransmitter release [26]. Subsequent studies of Drosophila unc-13 mutants and mouse Munc13-1 knockout supported their essential role in synaptic vesicle exocytosis [27, 28]. Knockout of other Munc13 genes also resulted in similar effects in a variety of synapses [3]. However, in the absence of any Munc13/UNC-13 member, the morphological organization of synapses and the assembly of dense projection are grossly normal, except that the precise docking pattern of synaptic vesicles is altered in a way that is consistent with changes in exocytosis dynamics [29].
Biochemical studies of Munc13/UNC-13 proteins have uncovered a complex protein-interaction network involving each domain of Munc13/UNC-13 [3]. For example, the most N-terminal C2A domain binds the Zinc Finger of RIM [30]. In different synapses and organisms, C2A domain is shown to be important for synapse vesicle docking and priming [31, 32], release probability [33], and kinetics [34], partly through regulating the spatial proximity of Munc13/UNC-13 to the calcium channels [35]. The C2B domain binds Ca2+ and anionic phospholipid, and works together with the C1 domain, which binds diacylglycerol and phorbol esters, to inhibit Ca2+-dependent neurotransmitter release [36]. The C2C domain at the C-terminus is shown to function as a vesicle or endosome adaptor [37]. With the many protein isoforms that often display subtle differences in their binding affinities and binding partners, a major remaining puzzle is how Munc13/UNC-13 protein diversity endorses the physiological specificity of the synaptic vesicle release.
3.2 RIM
The first member of RIM (for Rab3-interacting molecule) proteins, RIM1, was identified in a yeast-two-hybrid protein-interaction screen using an activated form of the small GTPase RAB3 [11]. Vertebrates have four Rim genes, with Rim1 and Rim2 broadly expressed in synapses, and C. elegans and Drosophila has one gene each, known as unc-10 and Rim, respectively. RIM proteins contain a Zinc Finger at N-terminus, a PDZ domain in the middle, two C2 domains at the C-terminus, and a proline-rich region in between the C2 domains (Fig. 1a). Each domain binds to specific proteins. The N-terminus of RIM binds to GTP-bound RAB3 associated with the synaptic vesicles, the Zinc Finger binds to the C2A domain of Munc13 [30], the PDZ domain binds to the CAZ protein ELKS [13], the proline-rich region binds to the CAZ protein RIM-BP [12], and the C2 domains mediate interactions with SNAREs, calcium channels [38], and the CAZ protein Liprin-α [39]. Thus, Rim acts as scaffolds at the presynaptic cytomatrix to organize synaptic vesicles and other proteins in the presynaptic release machinery (Fig. 1b).
Studies of C. elegans unc-10 mutants provided first functional evidence for a role of RIM in synaptic vesicle release. In unc-10 mutants the morphology of the presynaptic density and the docking pattern of synaptic vesicles are grossly normal, but there is greatly diminished neurotransmitter release [40]. Analyses of Drosophila Rim mutants revealed similar synaptic transmission deficits, and further showed reduced readily release pool of synaptic vesicles and reduced clustering of calcium channels at the active zone [41, 42]. In mice Rim 1 and Rim 2 each produces at least two protein isoforms through alternative splicing, with RIM1a being more abundant than RIM1b at presynaptic sites. Knockout of Rim1a caused a selective reduction in Munc13-1 expression at synapses, altered release probability, and short-term synaptic plasticity, but normal synaptic morphology [39]. Conditional knockout of both Rim 1 and Rim 2 led to further reduced readily releasable pool of vesicles and calcium channels at the Calyx of Held synapse [43]. These functional effects of RIM are consistent with the extensive molecular interactions between RIM and other CAZ proteins (Fig. 1b).
3.3 RIM-BP
As implied by its name, RIM-BP (RIM-binding protein) binds to RIM, originally isolated by yeast-two-hybrid screening [12]. RIM-BP proteins contain three SRC homology 3 (SH3) domains and three fibronectin III domains (Fig. 1a). The two C-terminal SH3 domains bind the proline-rich motifs of RIM and a number of voltage-gated calcium channels [44, 45], and the N-terminal SH3 domain binds the proline-rich motif of the CAZ protein Bassoon [46]. Mammals express three Rim-BP genes, while C. elegans and Drosophila each has one gene known as rimb-1 and Rim-BP. In vitro biochemical studies show that the binding between RIM and RIM-BP displays liquid–liquid phase separation, forming dynamic and condensed assemblies. In the presence of voltage-gated calcium channels, the RIM and RIM-BP condensates can enrich the channels [47]. Such mode of protein interactions may underlie the appearance of small clusters of vesicle release sites observed in neuronal synapses [48]. Genetic knockout studies with mice, Drosophila, and C. elegans have supported the functional significance of protein binding between Rim-BP and calcium channels, such that the mutant synapses have reduced number of calcium channels. Double mutants of Rim-BP and Rim show more severe synaptic deficits, revealing some overlapping roles of Rim and Rim-BP in synaptic vesicle docking, the morphology of presynaptic dense projections, and the number of calcium channels at the active zone [49,50,51]. In mice, a complete loss of Rim-BP1 leads to motor abnormalities reminiscent of dystonia, decreased Purkinje cell dendritic arborization, and a reduced number of cerebellar synapses. Interestingly, several loss-of-function mutations in human TSPOAP1/Rim-BP are recently reported to cause autosomal recessive dystonia [52]. These findings form the basis for further broadening our understanding of RIM-BPs.
3.4 ELKS
The name of ELKS proteins reflects the fact that these proteins are rich in E (Glutamate), L (Leucine), K (Lysine), and S (Serine) amino acid residues. ELKS1 was initially identified as a fusion protein with the receptor tyrosine kinase RET in thyroid carcinomas [53]. Mammals have two Elks genes that produce several isoforms; and C. elegans expresses a single ortholog ELKS-1, whereas Drosophila expresses an ELKS-like molecule called Bruchpilot (Fig. 1a). ELKS proteins contain mostly coiled-coil domains, and interact with a multitude of proteins, hence given different names in the literatures, including Rab6IP2 (Rab6-interacting protein 2) [54], CAST (CAZ-associated structural protein) [55], and ERC (ELKS/Rab6IP2/CAST) [13]. At synapses, ELKS/ELKS-1/Bruchpilot can bind to multiple CAZ proteins, including Rim via the coiled-coil region in C-termini [13], Bassoon and Piccolo via the central coiled-coil region [55], and Liprin-α via the N-terminal coiled-coil region [56].
Functional studies of individual ELKS genes using genetic knockout animals in different species show no major synapse defects. C. elegans elks-1 null animals have normal synapse architecture and CAZ protein expression [57]. However, as described below, in a gain-of-function SYD-2/Liprin-α mutant, ELKS-1 is required for the function and morphological integrity of certain synapses [58]. Elks single knockout mice also show no major synapse defects [59]. However, when both Rim1 and Rim2 and both Elks1 and Elks2 were deleted, neurons showed overall normal synaptic organization, but an absence of docked synaptic vesicles and a strong reduction in Munc13, Bassoon, Piccolo, and RIM-BP at the active zone, indicating disassembly of the presynaptic active zone [60]. These data show that ELKS proteins alone are not essential for formation of presynaptic terminals, but can modulate presynaptic active zone under specific conditions.
Drosophila Bruchpilot is a large protein that contains an ELKS homology region at the N-terminus and lacks the RIM-interacting domain, instead acquires a unique large C-terminal extension that bears features of cytoskeletal proteins such as plectin and myosin (Fig. 1a) [9]. Bruchpilot has received extensive attention as it was the first CAZ protein with its precise localization revealed by the STimulated Emission Depletion (STED) super-resolution microscopy. At the neuromuscular junction (NMJ), the N-terminus of Bruchpilot is close to the presynaptic plasma membrane, and its C-terminus extends into synaptic vesicle clusters. The ring-like T-bar structure is formed by two protein isoforms of Bruchpilot arranged in an alternating pattern in a circular array. Such an array creates “slots” for calcium channels and synaptic vesicle docking sites allowing efficient neurotransmission [61]. Loss of Bruchpilot alone causes dramatic effects on the formation of the platform of the T-bar structure at the presynaptic active zone (see diagram in Chapter “The Architecture of the Presynaptic Release Site”, Fig. 1), and is required for the clustering of calcium channels at the “pedestal” of T-bar at the center of active zone, ensuring the close proximity of calcium influx to the synaptic vesicle fusion machinery [62, 63]. It is possible that the different effects of eliminating Bruchpilot and ELKS on synaptic morphology are due to their differences in molecular structures.
3.5 Bassoon and Piccolo
These two very large proteins of greater than 450 kDa were identified using an antibody-based expression cloning method, and were among the first group of proteins to define the CAZ of presynaptic terminals in the vertebrate nervous system [7, 8, 64]. Bassoon and Piccolo share related protein structures, namely, repeated homologous regions called Piccolo Bassoon homology domain (PBH domains), of unknown function, coiled-coil regions, and two Zinc Finger domains (Fig. 1a). Additionally, Piccolo has a single PDZ domain and two C2 domains at its C-terminal (Fig. 1a). Given their large sizes, it is not surprising that they have many binding partners, ranging from components of the synaptic actin cytoskeleton to synaptic vesicle-associated proteins, such as the prenylated Rab acceptor PRA1 [64], other CAZ proteins (e.g., Rim, Rim-BP, ELKS, and Munc13) [65], and voltage-gated calcium channels [46]. The CC2 domain Bassoon also directly binds to an ubiquitin E3 ligase molecule Atg5 [66].
Although Bassoon and Piccolo were initially thought to be unique to CNS synapses, subsequent analyses show that they are also present at neuromuscular junctions, ribbon synapses, and other peripheral synapses [67, 68]. Bassoon and Piccolo show differential distributions within presynaptic terminals and play distinct roles in different types of synapses. In glutamatergic synapses of mammalian CNS, Bassoon clusters detected by immune-EM are often present above the filaments emanating from the plasma membrane at the active zone [69]. Super-resolution imaging using STED microscopy reveals that the C-terminus of Bassoon is close to the presynaptic plasma membrane and N-terminus extended into the presynaptic cytoplasm [70] (Fig. 1b). The mammalian photoreceptor ribbon synapse has two sub-compartments: a dense projection from the plasma membrane, and an electron-dense ribbon extending from the SV release site into the presynaptic cytoplasm [71]. Bassoon localizes at the junction between these two sub-compartments, while Piccolo associates with the ribbon, and Rim, Munc13, and ELKS exist at the presynaptic density [72].
At least two mutant mouse strains of Bassoon (Bsn) have been reported. A Bsn in-frame deletion mutant, which expresses a protein of 180 kd that includes the N-terminal and C-terminal regions but lacks the central part of Bassoon, shows unanchored ribbons in the presynaptic terminal of retina photoreceptors [73]. In these Bsn mutant mice, the inner hair cell ribbon synapses in the cochlea also exhibit loss of fast neurotransmitter release [74], while CNS excitatory synapses exhibit impaired synaptic transmission but apparently normal synaptic morphology [75]. These data establish the importance of Bassoon in ribbon synaptic architecture and suggest Bassoon’s role may vary depending on synapse type. In a Bsn null mutant, cerebellar mossy fiber synapses show enhanced short-term synaptic depression but largely normal basal synaptic transmission and the number of synaptic vesicles [76]. In the endbulb synapses of auditory nerve fibers, the replenishment of synaptic vesicles at the release sites is significantly reduced [77]. Bassoon promotes vesicle replenishment in part through inhibiting presynaptic autophagy [66].
Piccolo also exhibits synapse-type specific effects. For example, a short isoform of Piccolo, Piccolino, is found to be predominantly expressed at sensory ribbon synapses in the eye and ear [78]. Piccolino is required for the formation of the plate-shaped ribbons, as loss of Piccolino in rodents results in spherical ribbons and a disruption of the maturation of ribbons [79, 80]. At the rat calyx of the Held synapse, Piccolo deficiency results in a defect in replenishment of readily releasable synaptic vesicles during prolonged and intense firing activities, and smaller synapses [81]. The Piccolo knockout rats (Pclotgt/gt) also exhibit abnormal brain morphology and altered cerebellar neural circuitry [82]. Interestingly, loss-of-function mutations of human Piccolo (PCLO) have been associated with type 3 pontocerebellar hypoplasia (PCH3), also known as cerebellar atrophy with progressive microcephaly [83]. Knockdown of both Bassoon and Piccolo in hippocampal and cortical neurons led to a reduction in synaptic vesicles, but did not alter synapse physiology, supporting their partially overlapping functions [84].
For a period of time, it was thought that proteins similar to Bassoon and Piccolo were not present in synapses of invertebrates. Through careful molecular phylogeny analysis, in combination with expression and genetic studies, the Drosophila Fife and C. elegans CLA-1/Clarinet are reported to share features similar to Piccolo and Rim [85]. Both Fife and CLA-1 are large proteins that contain Zinc Finger, C2, and PDZ domains, along with numerous unique repeats. Both genes produce multiple protein isoforms, which exhibit distinctive localization in the presynaptic cytomatrix, forming nanodomains and interacting with other CAZ proteins [86, 87]. Genetic studies of fife and cla-1 mutants show that different isoforms display synapse-type specificity to regulate presynaptic terminal structural and functional integrity [85,86,87]. CLA-1 has recently been linked to autophagy (bioRxiv 2021.08.19.457026), supporting mechanistic conservation with Bassoon.
3.6 Liprin-Alpha
Liprin (for Lar-interacting-protein-related protein) proteins were named because of their initial identification by Y2H assay as proteins interacting with the intracellular phosphatase domain of LAR and closely related receptor tyrosine phosphatases [88]. Liprin proteins include three subfamilies: alpha, beta, and gamma; however, only alpha proteins are extensively studied for their roles in synapses. The N-termini of Liprin-α proteins are characterized by multiple coiled-coil structures, with a stretch of ~100 amino acids, known as the LH1 (Liprin Homology) domain, which shares near 90% sequence identity from C. elegans to mammals; and the C-terminal half of Liprin-α contains three SAM domains (Fig. 1). The middle region of different isoforms of Liprin-α generally has low-complexity domains. C. elegans and Drosophila each expresses one Liprin-α gene, known as syd-2 and DLiprin-α, respectively [23, 89]. Both are exclusively localized to presynaptic terminals. Vertebrates express four Liprin-α genes. Expression of Liprin-α1 and Liprin-α4 is seen in limited area in the brain and also outside of the nervous system. Liprin-α2 and Liprin-α3 are specifically and broadly expressed in the brain. Liprin-α3 shows strong localization to the presynaptic active zone [70], and Liprin-α2 is present at both pre- and postsynaptic sites [90].
Genetic studies of mutants in C. elegans and Drosophila provided the first evidence for roles of Liprin-α in structural integrity of the presynaptic active zone. C. elegans syd-2 was identified from a forward genetic screen using an SNB-1::GFP reporter, because loss of function in syd-2 caused both reduced and diffused accumulation of SNB-1::GFP at neuromuscular synapses [23]. Subsequent findings from Drosophila showed that loss of DLiprin-α also altered morphology of neuromuscular synapses [89]. Ultrastructural analysis of both syd-2 and DLiprin-α mutants revealed a primary effect on the length and shape of the presynaptic dense projection. Moreover, the CAZ proteins ELKS and RIM show diffuse and reduced accumulation at the presynaptic active zone [58, 91, 92]. In mouse hippocampal neuron synapses, knockout of Liprin-α3 alone causes subtle but significant alternation in presynaptic ultrastructure and CAZ protein compositions [70]. Knockout of both Liprin-α2 and α3 leads to stronger reductions in the protein machinery for docking and priming and in the pool of releasable vesicles at excitatory and inhibitory synapses, and increases in Ca2+ channels and release probability at excitatory but not inhibitory synapses [93].
Studies of a gain-of-function (gf) mutation in the C. elegans SYD-2 have provided an important clue to the understanding of how Liprin-α organizes presynaptic cytomatrix. The syd-2(gf) mutation changes Arg184 to Cys in the highly conserved LH1 domain and causes enlarged presynaptic dense projections [58, 94]. In biochemical studies, purified wild type LH1 domain forms dimers, whereas LH1 domain with the Arg184Cys mutation forms multimers [95]. The LH1 domain is predicted to form a coiled-coil structure. Crystal structure studies of the LH1 domain of the vertebrate Liprin-α2 reveals that the helix containing Arg194 (corresponding to Arg184 in SYD-2) forms a homo-tetramer [96]. Arg194 faces away from the tetramerization surface and stabilizes intramolecular interaction between orderly arranged helical dimers. The Arg194Cys mutation disrupts the interaction among positively charged amino acid residues and enhances the dimer to oligomer transition of the LH1 domain. Moreover, in both biochemical assays and living cells, oligomerized Liprin-α2 promotes formation of ELKS condensates via liquid–liquid phase separation, and such ELKS condensates recruit RIM1a, RIM-BP [58]. As described in Chapter “The Architecture of the Presynaptic Release Site”, functional evidence from C. elegans and mice has supported the idea that the liquid–liquid phase separation property of CAZ proteins is important for presynaptic active zone assembly.
4 Intracellular Regulators of CAZ Proteins in Synapse Formation
The extensive interactive nature among CAZ proteins underlies the complex regulation of presynaptic assembly. Live imaging of CAZ proteins has probed into the dynamic processes of assembly of a presynaptic terminal. CAZ proteins translated in the soma must be sorted into the axon, and transported and delivered to the presynaptic terminal [97,98,99]. The transport and delivery of CAZ components involve diverse vesicular carriers that interact with Golgi apparatus, endosomes, and lysosomal pathways [100, 101]. For example, ELKS can interact with Rab6 to mediate transport between endosome and Golgi [54]. Bassoon and Piccolo are transported in large dense core vesicles that contain other CAZ proteins and SNAREs [98]. Liprin-α interacts with the axonal motor proteins kinesin-1 and KIF1A/Unc104 [92, 102] as well as many intracellular and cell surface molecules to contribute to the delivery of other CAZ proteins and the nucleation of nascent active zone [62, 91, 103]. Here, we highlight two intracellular pathways that coordinate with CAZ proteins in synapse formation.
4.1 SYD-1/dSYD-1/SYDE
Molecular genetic studies from C. elegans and Drosophila have supported a functional hierarchy involving the conserved SYD-1/dSYD-1 proteins as an upstream regulator in the assembly of presynaptic active zone. C. elegans syd-1 was identified from the forward genetic screen that yielded syd-2, based on similarly altered SNB-1::GFP patterns [104]. The invertebrate SYD-1 and dSYD-1 full-length proteins have an N-terminus PDZ domain, followed by a C2 domain and a Rho-GAP (GTPase Activating Protein) domain (Fig. 1a) [104, 105]. Mouse expresses two homologs, known as SYDE1 and SYDE2, containing only C2 and Rho-GAP domains [106], resembling a short isoform of C. elegans SYD-1. The C2 domain facilitates protein association with the membrane. However, the function of the GAP domain varies between species. Drosophila dSYD-1 has GAP activity on Rac1 [107] and mouse SYDE1/mSYD1A acts on CDC42 [106], whereas the GAP domain in C. elegans SYD-1 may not be active [104].
Evidence from both C. elegans and Drosophila supports a conclusion that syd-1/dsyd-1 and syd-2/Dliprin-α act in a common molecular pathway to promote synapse formation [58, 105, 108, 109]. In C. elegans, double mutants of syd-1 and syd-2 resemble each single mutant. In the absence of syd-1, the syd-2(gf) mutation can induce synapse formation [58]. Synapse imaging studies in both C. elegans and Drosophila show that SYD-1/dSYD-1 arrives early at nascent sites of the presynaptic terminals and facilitates the accumulation of SYD-2/DLiprin-α and other CAZ proteins, such as ELKS-1/Bruchpilot [35, 110,111,112]. SYD-1/dSYD-1 can interact with neurexin via the PDZ domain [110], and also bind to other synaptic proteins such as spinophilin [113], neurabin [111], and RSY-1/PNISR [114]. Such complex interactions orchestrate early assembly processes at the Drosophila NMJs [110, 115]. Mice lacking Syde1/mSYD1A exhibit reduced docked vesicles and synaptic activities [106]. An intrinsically disordered region of SYDE1/mSYD1A interacts with multiple synapse proteins, including CAZ protein Liprin-α2, Munc18-1, and presynaptic receptor tyrosine phosphatases. Conceivably, such multi-protein interactions play a key role in tethering synaptic vesicles.
4.2 Tripartite Complex of CASK, Mint, and Veli
This tightly associated PDZ domain-containing protein complex has been studied in many cellular contexts and is present at both pre- and postsynaptic sites [116]. A presynaptic role for CASK/Mint/Veli was initially hinted at by the interactions with neurexins [117]. CASK/Mint/Veli also bind directly or indirectly to other synaptic proteins such as voltage-gated calcium channels, Liprin-α, and Cdk5 [118, 119], as well as several synaptic adhesion molecules [116]. A single member of each protein family is present in invertebrates; in mammals, Mint and Veli each is encoded by three genes. Knockout mice of either Mint or Veli die at an early stage, precluding a full examination of their roles in synapse formation [120, 121]. Nonetheless, it is generally agreed that these three proteins play regulatory roles in synaptic vesicle release, and may indirectly contribute to the transport of synaptic proteins and interaction with synaptic surface proteins to facilitate the development and maintenance of synaptic architecture.
5 Summary
Through consorted efforts using combinatorial approaches, research in the past two decades has revealed the shared core CAZ components. Despite their precise localization to the presynaptic active zone, functional studies have placed different families of CAZ proteins in a wide spectrum for their essentiality in synapse formation and function. Munc13/Unc-13 is essential for synaptic vesicle release, whereas ELKS has the least functional significance yet provides maladaptive regulation to many actions of presynaptic release. The interwoven protein-interaction network at CAZ remains a wonderland where conceptual creativity and technology innovation continue to push the boundary of our knowledge about the mystery of synapses.
References
Nagy A, Baker RR, Morris SJ, Whittaker VP. The preparation and characterization of synaptic vesicles of high purity. Brain Res. 1976;109:285–309. https://doi.org/10.1016/0006-8993(76)90531-x.
Takamori S, Holt M, Stenius K, Lemke EA, Gronborg M, Riedel D, et al. Molecular anatomy of a trafficking organelle. Cell. 2006;127:831–46. https://doi.org/10.1016/j.cell.2006.10.030.
Sudhof TC. The presynaptic active zone. Neuron. 2012;75:11–25. https://doi.org/10.1016/j.neuron.2012.06.012.
Kistner U, Wenzel BM, Veh RW, Cases-Langhoff C, Garner AM, Appeltauer U, et al. SAP90, a rat presynaptic protein related to the product of the Drosophila tumor suppressor gene dlg-A. J Biol Chem. 1993;268:4580–3.
Langnaese K, Seidenbecher C, Wex H, Seidel B, Hartung K, Appeltauer U, et al. Protein components of a rat brain synaptic junctional protein preparation. Brain Res Mol Brain Res. 1996;42:118–22. https://doi.org/10.1016/s0169-328x(96)00147-7.
Huse WD, Sastry L, Iverson SA, Kang AS, Alting-Mees M, Burton DR, et al. Generation of a large combinatorial library of the immunoglobulin repertoire in phage lambda. Science. 1989;246:1275–81. https://doi.org/10.1126/science.2531466.
tom Dieck S, Sanmarti-Vila L, Langnaese K, Richter K, Kindler S, Soyke A, et al. Bassoon, a novel zinc-finger CAG/glutamine-repeat protein selectively localized at the active zone of presynaptic nerve terminals. J Cell Biol. 1998;142:499–509. https://doi.org/10.1083/jcb.142.2.499.
Cases-Langhoff C, Voss B, Garner AM, Appeltauer U, Takei K, Kindler S, et al. Piccolo, a novel 420 kDa protein associated with the presynaptic cytomatrix. Eur J Cell Biol. 1996;69:214–23.
Wagh DA, Rasse TM, Asan E, Hofbauer A, Schwenkert I, Durrbeck H, et al. Bruchpilot, a protein with homology to ELKS/CAST, is required for structural integrity and function of synaptic active zones in Drosophila. Neuron. 2006;49:833–44. https://doi.org/10.1016/j.neuron.2006.02.008.
Fields S, Song O. A novel genetic system to detect protein-protein interactions. Nature. 1989;340:245–6. https://doi.org/10.1038/340245a0.
Wang Y, Okamoto M, Schmitz F, Hofmann K, Sudhof TC. Rim is a putative Rab3 effector in regulating synaptic-vesicle fusion. Nature. 1997;388:593–8. https://doi.org/10.1038/41580.
Wang Y, Sugita S, Sudhof TC. The RIM/NIM family of neuronal C2 domain proteins. Interactions with Rab3 and a new class of Src homology 3 domain proteins. J Biol Chem. 2000;275:20033–44. https://doi.org/10.1074/jbc.M909008199.
Wang Y, Liu X, Biederer T, Sudhof TC. A family of RIM-binding proteins regulated by alternative splicing: implications for the genesis of synaptic active zones. Proc Natl Acad Sci U S A. 2002;99:14464–9. https://doi.org/10.1073/pnas.182532999.
Brenner S. The genetics of Caenorhabditis elegans. Genetics. 1974;77:71–94. https://doi.org/10.1093/genetics/77.1.71.
Rand JB, Nonet ML. Synaptic transmission. In: Riddle DL, Blumenthal T, Meyer BJ, Priess JR, editors. C. elegans II. Cold Spring Harbor (NY); 1997.
Maruyama IN, Brenner S. A phorbol ester/diacylglycerol-binding protein encoded by the unc-13 gene of Caenorhabditis elegans. Proc Natl Acad Sci U S A. 1991;88:5729–33. https://doi.org/10.1073/pnas.88.13.5729.
Brose N, Hofmann K, Hata Y, Sudhof TC. Mammalian homologues of Caenorhabditis elegans unc-13 gene define novel family of C2-domain proteins. J Biol Chem. 1995;270:25273–80. https://doi.org/10.1074/jbc.270.42.25273.
White JG, Southgate E, Thomson JN, Brenner S. The structure of the nervous system of the nematode Caenorhabditis elegans. Philos Trans R Soc Lond Ser B Biol Sci. 1986;314:1–340. https://doi.org/10.1098/rstb.1986.0056.
Chalfie M, Tu Y, Euskirchen G, Ward WW, Prasher DC. Green fluorescent protein as a marker for gene expression. Science. 1994;263:802–5. https://doi.org/10.1126/science.8303295.
Nonet ML. Visualization of synaptic specializations in live C. elegans with synaptic vesicle protein-GFP fusions. J Neurosci Methods. 1999;89:33–40. https://doi.org/10.1016/s0165-0270(99)00031-x.
Jorgensen EM, Hartwieg E, Schuske K, Nonet ML, Jin Y, Horvitz HR. Defective recycling of synaptic vesicles in synaptotagmin mutants of Caenorhabditis elegans. Nature. 1995;378:196–9. https://doi.org/10.1038/378196a0.
Jin Y. Synaptogenesis. WormBook. 2005; https://doi.org/10.1895/wormbook.1.44.1.
Zhen M, Jin Y. The liprin protein SYD-2 regulates the differentiation of presynaptic termini in C. elegans. Nature. 1999;401:371–5. https://doi.org/10.1038/43886.
Betz A, Okamoto M, Benseler F, Brose N. Direct interaction of the rat unc-13 homologue Munc13-1 with the N terminus of syntaxin. J Biol Chem. 1997;272:2520–6. https://doi.org/10.1074/jbc.272.4.2520.
Lai Y, Choi UB, Leitz J, Rhee HJ, Lee C, Altas B, et al. Molecular mechanisms of synaptic vesicle priming by Munc13 and Munc18. Neuron. 2017;95:591–607 e510. https://doi.org/10.1016/j.neuron.2017.07.004.
Richmond JE, Davis WS, Jorgensen EM. UNC-13 is required for synaptic vesicle fusion in C. elegans. Nat Neurosci. 1999;2:959–64. https://doi.org/10.1038/14755.
Aravamudan B, Fergestad T, Davis WS, Rodesch CK, Broadie K. Drosophila UNC-13 is essential for synaptic transmission. Nat Neurosci. 1999;2:965–71. https://doi.org/10.1038/14764.
Augustin I, Rosenmund C, Sudhof TC, Brose N. Munc13-1 is essential for fusion competence of glutamatergic synaptic vesicles. Nature. 1999;400:457–61. https://doi.org/10.1038/22768.
Imig C, Min SW, Krinner S, Arancillo M, Rosenmund C, Sudhof TC, et al. The morphological and molecular nature of synaptic vesicle priming at presynaptic active zones. Neuron. 2014;84:416–31. https://doi.org/10.1016/j.neuron.2014.10.009.
Lu J, Machius M, Dulubova I, Dai H, Sudhof TC, Tomchick DR, et al. Structural basis for a Munc13-1 homodimer to Munc13-1/RIM heterodimer switch. PLoS Biol. 2006;4:e192. https://doi.org/10.1371/journal.pbio.0040192.
Deng L, Kaeser PS, Xu W, Sudhof TC. RIM proteins activate vesicle priming by reversing autoinhibitory homodimerization of Munc13. Neuron. 2011;69:317–31. https://doi.org/10.1016/j.neuron.2011.01.005.
Camacho M, Basu J, Trimbuch T, Chang S, Pulido-Lozano C, Chang SS, et al. Heterodimerization of Munc13 C2A domain with RIM regulates synaptic vesicle docking and priming. Nat Commun. 2017;8:15293. https://doi.org/10.1038/ncomms15293.
Zhou K, Stawicki TM, Goncharov A, Jin Y. Position of UNC-13 in the active zone regulates synaptic vesicle release probability and release kinetics. elife. 2013;2:e01180. https://doi.org/10.7554/eLife.01180.
Hu Z, Tong XJ, Kaplan JM. UNC-13L, UNC-13S, and Tomosyn form a protein code for fast and slow neurotransmitter release in Caenorhabditis elegans. elife. 2013;2:e00967. https://doi.org/10.7554/eLife.00967.
Bohme MA, Beis C, Reddy-Alla S, Reynolds E, Mampell MM, Grasskamp AT, et al. Active zone scaffolds differentially accumulate Unc13 isoforms to tune Ca(2+) channel-vesicle coupling. Nat Neurosci. 2016;19:1311–20. https://doi.org/10.1038/nn.4364.
Michelassi F, Liu H, Hu Z, Dittman JS. A C1-C2 module in Munc13 inhibits calcium-dependent neurotransmitter release. Neuron. 2017;95:577–590 e575. https://doi.org/10.1016/j.neuron.2017.07.015.
Padmanarayana M, Liu H, Michelassi F, Li L, Betensky D, Dominguez MJ, et al. A unique C2 domain at the C terminus of Munc13 promotes synaptic vesicle priming. Proc Natl Acad Sci U S A. 2021;118 https://doi.org/10.1073/pnas.2016276118.
Coppola T, Magnin-Luthi S, Perret-Menoud V, Gattesco S, Schiavo G, Regazzi R. Direct interaction of the Rab3 effector RIM with Ca2+ channels, SNAP-25, and synaptotagmin. J Biol Chem. 2001;276:32756–62. https://doi.org/10.1074/jbc.M100929200.
Schoch S, Castillo PE, Jo T, Mukherjee K, Geppert M, Wang Y, et al. RIM1alpha forms a protein scaffold for regulating neurotransmitter release at the active zone. Nature. 2002;415:321–6. https://doi.org/10.1038/415321a.
Koushika SP, Richmond JE, Hadwiger G, Weimer RM, Jorgensen EM, Nonet ML. A post-docking role for active zone protein Rim. Nat Neurosci. 2001;4:997–1005. https://doi.org/10.1038/nn732.
Graf ER, Valakh V, Wright CM, Wu C, Liu Z, Zhang YQ, et al. RIM promotes calcium channel accumulation at active zones of the Drosophila neuromuscular junction. J Neurosci. 2012;32:16586–96. https://doi.org/10.1523/JNEUROSCI.0965-12.2012.
Muller M, Liu KS, Sigrist SJ, Davis GW. RIM controls homeostatic plasticity through modulation of the readily-releasable vesicle pool. J Neurosci. 2012;32:16574–85. https://doi.org/10.1523/JNEUROSCI.0981-12.2012.
Han Y, Kaeser PS, Sudhof TC, Schneggenburger R. RIM determines Ca(2)+ channel density and vesicle docking at the presynaptic active zone. Neuron. 2011;69:304–16. https://doi.org/10.1016/j.neuron.2010.12.014.
Hibino H, Pironkova R, Onwumere O, Vologodskaia M, Hudspeth AJ, Lesage F. RIM binding proteins (RBPs) couple Rab3-interacting molecules (RIMs) to voltage-gated Ca(2+) channels. Neuron. 2002;34:411–23. https://doi.org/10.1016/s0896-6273(02)00667-0.
Kaeser PS, Deng L, Wang Y, Dulubova I, Liu X, Rizo J, et al. RIM proteins tether Ca2+ channels to presynaptic active zones via a direct PDZ-domain interaction. Cell. 2011;144:282–95. https://doi.org/10.1016/j.cell.2010.12.029.
Davydova D, Marini C, King C, Klueva J, Bischof F, Romorini S, et al. Bassoon specifically controls presynaptic P/Q-type Ca(2+) channels via RIM-binding protein. Neuron. 2014;82:181–94. https://doi.org/10.1016/j.neuron.2014.02.012.
Wu X, Cai Q, Shen Z, Chen X, Zeng M, Du S, et al. RIM and RIM-BP form presynaptic active-zone-like condensates via phase separation. Mol Cell. 2019;73:971–984 e975. https://doi.org/10.1016/j.molcel.2018.12.007.
Tang AH, Chen H, Li TP, Metzbower SR, MacGillavry HD, Blanpied TA. A trans-synaptic nanocolumn aligns neurotransmitter release to receptors. Nature. 2016;536:210–4. https://doi.org/10.1038/nature19058.
Petzoldt AG, Gotz TWB, Driller JH, Lutzkendorf J, Reddy-Alla S, Matkovic-Rachid T, et al. RIM-binding protein couples synaptic vesicle recruitment to release sites. J Cell Biol. 2020;219 https://doi.org/10.1083/jcb.201902059.
Acuna C, Liu X, Gonzalez A, Sudhof TC. RIM-BPs mediate tight coupling of action potentials to Ca(2+)-triggered neurotransmitter release. Neuron. 2015;87:1234–47. https://doi.org/10.1016/j.neuron.2015.08.027.
Kushibiki Y, Suzuki T, Jin Y, Taru H. RIMB-1/RIM-binding protein and UNC-10/RIM redundantly regulate presynaptic localization of the voltage-gated Calcium Channel in Caenorhabditis elegans. J Neurosci. 2019;39:8617–31. https://doi.org/10.1523/JNEUROSCI.0506-19.2019.
Mencacci NE, Brockmann MM, Dai J, Pajusalu S, Atasu B, Campos J, et al. Biallelic variants in TSPOAP1, encoding the active-zone protein RIMBP1, cause autosomal recessive dystonia. J Clin Invest. 2021;131 https://doi.org/10.1172/JCI140625.
Nakata T, Kitamura Y, Shimizu K, Tanaka S, Fujimori M, Yokoyama S, et al. Fusion of a novel gene, ELKS, to RET due to translocation t(10;12)(q11;p13) in a papillary thyroid carcinoma. Genes Chromosomes Cancer. 1999;25:97–103. https://doi.org/10.1002/(sici)1098-2264(199906)25:2<97::aid-gcc4>3.0.co;2-l.
Monier S, Jollivet F, Janoueix-Lerosey I, Johannes L, Goud B. Characterization of novel Rab6-interacting proteins involved in endosome-to-TGN transport. Traffic. 2002;3:289–97. https://doi.org/10.1034/j.1600-0854.2002.030406.x.
Ohtsuka T, Takao-Rikitsu E, Inoue E, Inoue M, Takeuchi M, Matsubara K, et al. Cast: a novel protein of the cytomatrix at the active zone of synapses that forms a ternary complex with RIM1 and munc13-1. J Cell Biol. 2002;158:577–90. https://doi.org/10.1083/jcb.200202083.
Ko J, Na M, Kim S, Lee JR, Kim E. Interaction of the ERC family of RIM-binding proteins with the liprin-alpha family of multidomain proteins. J Biol Chem. 2003;278:42377–85. https://doi.org/10.1074/jbc.M307561200.
Deken SL, Vincent R, Hadwiger G, Liu Q, Wang ZW, Nonet ML. Redundant localization mechanisms of RIM and ELKS in Caenorhabditis elegans. J Neurosci. 2005;25:5975–83. https://doi.org/10.1523/JNEUROSCI.0804-05.2005.
Dai Y, Taru H, Deken SL, Grill B, Ackley B, Nonet ML, et al. SYD-2 Liprin-alpha organizes presynaptic active zone formation through ELKS. Nat Neurosci. 2006;9:1479–87. https://doi.org/10.1038/nn1808.
Kaeser PS, Deng L, Chavez AE, Liu X, Castillo PE, Sudhof TC. ELKS2alpha/CAST deletion selectively increases neurotransmitter release at inhibitory synapses. Neuron. 2009;64:227–39. https://doi.org/10.1016/j.neuron.2009.09.019.
Wang SSH, Held RG, Wong MY, Liu C, Karakhanyan A, Kaeser PS. Fusion competent synaptic vesicles persist upon active zone disruption and loss of vesicle docking. Neuron. 2016;91:777–91. https://doi.org/10.1016/j.neuron.2016.07.005.
Matkovic T, Siebert M, Knoche E, Depner H, Mertel S, Owald D, et al. The Bruchpilot cytomatrix determines the size of the readily releasable pool of synaptic vesicles. J Cell Biol. 2013;202:667–83. https://doi.org/10.1083/jcb.201301072.
Fouquet W, Owald D, Wichmann C, Mertel S, Depner H, Dyba M, et al. Maturation of active zone assembly by Drosophila Bruchpilot. J Cell Biol. 2009;186:129–45. https://doi.org/10.1083/jcb.200812150.
Kittel RJ, Wichmann C, Rasse TM, Fouquet W, Schmidt M, Schmid A, et al. Bruchpilot promotes active zone assembly, Ca2+ channel clustering, and vesicle release. Science. 2006;312:1051–4. https://doi.org/10.1126/science.1126308.
Fenster SD, Chung WJ, Zhai R, Cases-Langhoff C, Voss B, Garner AM, et al. Piccolo, a presynaptic zinc finger protein structurally related to bassoon. Neuron. 2000;25:203–14. https://doi.org/10.1016/s0896-6273(00)80883-1.
Wang X, Hu B, Zieba A, Neumann NG, Kasper-Sonnenberg M, Honsbein A, et al. A protein interaction node at the neurotransmitter release site: domains of Aczonin/Piccolo, Bassoon, CAST, and rim converge on the N-terminal domain of Munc13-1. J Neurosci. 2009;29:12584–96. https://doi.org/10.1523/JNEUROSCI.1255-09.2009.
Okerlund ND, Schneider K, Leal-Ortiz S, Montenegro-Venegas C, Kim SA, Garner LC, et al. Bassoon controls presynaptic autophagy through Atg5. Neuron. 2018;97:727. https://doi.org/10.1016/j.neuron.2018.01.010.
Dick O, Hack I, Altrock WD, Garner CC, Gundelfinger ED, Brandstatter JH. Localization of the presynaptic cytomatrix protein Piccolo at ribbon and conventional synapses in the rat retina: comparison with Bassoon. J Comp Neurol. 2001;439:224–34. https://doi.org/10.1002/cne.1344.
Juranek J, Mukherjee K, Rickmann M, Martens H, Calka J, Sudhof TC, et al. Differential expression of active zone proteins in neuromuscular junctions suggests functional diversification. Eur J Neurosci. 2006;24:3043–52. https://doi.org/10.1111/j.1460-9568.2006.05183.x.
Siksou L, Rostaing P, Lechaire JP, Boudier T, Ohtsuka T, Fejtova A, et al. Three-dimensional architecture of presynaptic terminal cytomatrix. J Neurosci. 2007;27:6868–77. https://doi.org/10.1523/JNEUROSCI.1773-07.2007.
Wong MY, Liu C, Wang SSH, Roquas ACF, Fowler SC, Kaeser PS. Liprin-alpha3 controls vesicle docking and exocytosis at the active zone of hippocampal synapses. Proc Natl Acad Sci U S A. 2018;115:2234–9. https://doi.org/10.1073/pnas.1719012115.
Gray EG, Pease HL. On understanding the organisation of the retinal receptor synapses. Brain Res. 1971;35:1–15. https://doi.org/10.1016/0006-8993(71)90591-9.
tom Dieck S, Altrock WD, Kessels MM, Qualmann B, Regus H, Brauner D, et al. Molecular dissection of the photoreceptor ribbon synapse: physical interaction of Bassoon and RIBEYE is essential for the assembly of the ribbon complex. J Cell Biol. 2005;168:825–36. https://doi.org/10.1083/jcb.200408157.
Dick O, tom Dieck S, Altrock WD, Ammermuller J, Weiler R, Garner CC, et al. The presynaptic active zone protein bassoon is essential for photoreceptor ribbon synapse formation in the retina. Neuron. 2003;37:775–86. https://doi.org/10.1016/s0896-6273(03)00086-2.
Khimich D, Nouvian R, Pujol R, Tom Dieck S, Egner A, Gundelfinger ED, et al. Hair cell synaptic ribbons are essential for synchronous auditory signalling. Nature. 2005;434:889–94. https://doi.org/10.1038/nature03418.
Altrock WD, tom Dieck S, Sokolov M, Meyer AC, Sigler A, Brakebusch C, et al. Functional inactivation of a fraction of excitatory synapses in mice deficient for the active zone protein bassoon. Neuron. 2003;37:787–800. https://doi.org/10.1016/s0896-6273(03)00088-6.
Hallermann S, Fejtova A, Schmidt H, Weyhersmuller A, Silver RA, Gundelfinger ED, et al. Bassoon speeds vesicle reloading at a central excitatory synapse. Neuron. 2010;68:710–23. https://doi.org/10.1016/j.neuron.2010.10.026.
Mendoza Schulz A, Jing Z, Sanchez Caro JM, Wetzel F, Dresbach T, Strenzke N, et al. Bassoon-disruption slows vesicle replenishment and induces homeostatic plasticity at a CNS synapse. EMBO J. 2014;33:512–27. https://doi.org/10.1002/embj.201385887.
Regus-Leidig H, Ott C, Lohner M, Atorf J, Fuchs M, Sedmak T, et al. Identification and immunocytochemical characterization of Piccolino, a novel Piccolo splice variant selectively expressed at sensory ribbon synapses of the eye and ear. PLoS One. 2013;8:e70373. https://doi.org/10.1371/journal.pone.0070373.
Regus-Leidig H, Fuchs M, Lohner M, Leist SR, Leal-Ortiz S, Chiodo VA, et al. In vivo knockdown of Piccolino disrupts presynaptic ribbon morphology in mouse photoreceptor synapses. Front Cell Neurosci. 2014;8:259. https://doi.org/10.3389/fncel.2014.00259.
Muller TM, Gierke K, Joachimsthaler A, Sticht H, Izsvak Z, Hamra FK, et al. A multiple Piccolino-RIBEYE interaction supports plate-shaped synaptic ribbons in retinal neurons. J Neurosci. 2019;39:2606–19. https://doi.org/10.1523/JNEUROSCI.2038-18.2019.
Parthier D, Kuner T, Korber C. The presynaptic scaffolding protein Piccolo organizes the readily releasable pool at the calyx of Held. J Physiol. 2018;596:1485–99. https://doi.org/10.1113/JP274885.
Falck J, Bruns C, Hoffmann-Conaway S, Straub I, Plautz EJ, Orlando M, et al. Loss of Piccolo function in rats induces cerebellar network dysfunction and pontocerebellar hypoplasia type 3-like phenotypes. J Neurosci. 2020;40:2943–59. https://doi.org/10.1523/JNEUROSCI.2316-19.2020.
Ahmed MY, Chioza BA, Rajab A, Schmitz-Abe K, Al-Khayat A, Al-Turki S, et al. Loss of PCLO function underlies pontocerebellar hypoplasia type III. Neurology. 2015;84:1745–50. https://doi.org/10.1212/WNL.0000000000001523.
Mukherjee K, Yang X, Gerber SH, Kwon HB, Ho A, Castillo PE, et al. Piccolo and bassoon maintain synaptic vesicle clustering without directly participating in vesicle exocytosis. Proc Natl Acad Sci U S A. 2010;107:6504–9. https://doi.org/10.1073/pnas.1002307107.
Bruckner JJ, Gratz SJ, Slind JK, Geske RR, Cummings AM, Galindo SE, et al. Fife, a Drosophila Piccolo-RIM homolog, promotes active zone organization and neurotransmitter release. J Neurosci. 2012;32:17048–58. https://doi.org/10.1523/JNEUROSCI.3267-12.2012.
Bruckner JJ, Zhan H, Gratz SJ, Rao M, Ukken F, Zilberg G, et al. Fife organizes synaptic vesicles and calcium channels for high-probability neurotransmitter release. J Cell Biol. 2017;216:231–46. https://doi.org/10.1083/jcb.201601098.
Xuan Z, Manning L, Nelson J, Richmond JE, Colon-Ramos DA, Shen K, et al. Clarinet (CLA-1), a novel active zone protein required for synaptic vesicle clustering and release. elife. 2017;6 https://doi.org/10.7554/eLife.29276.
Serra-Pages C, Medley QG, Tang M, Hart A, Streuli M. Liprins, a family of LAR transmembrane protein-tyrosine phosphatase-interacting proteins. J Biol Chem. 1998;273:15611–20. https://doi.org/10.1074/jbc.273.25.15611.
Kaufmann N, DeProto J, Ranjan R, Wan H, Van Vactor D. Drosophila liprin-alpha and the receptor phosphatase Dlar control synapse morphogenesis. Neuron. 2002;34:27–38. https://doi.org/10.1016/s0896-6273(02)00643-8.
Wyszynski M, Kim E, Dunah AW, Passafaro M, Valtschanoff JG, Serra-Pages C, et al. Interaction between GRIP and liprin-alpha/SYD2 is required for AMPA receptor targeting. Neuron. 2002;34:39–52. https://doi.org/10.1016/s0896-6273(02)00640-2.
Ackley BD, Harrington RJ, Hudson ML, Williams L, Kenyon CJ, Chisholm AD, et al. The two isoforms of the Caenorhabditis elegans leukocyte-common antigen related receptor tyrosine phosphatase PTP-3 function independently in axon guidance and synapse formation. J Neurosci. 2005;25:7517–28. https://doi.org/10.1523/JNEUROSCI.2010-05.2005.
Miller KE, DeProto J, Kaufmann N, Patel BN, Duckworth A, Van Vactor D. Direct observation demonstrates that Liprin-alpha is required for trafficking of synaptic vesicles. Curr Biol. 2005;15:684–9. https://doi.org/10.1016/j.cub.2005.02.061.
Emperador-Melero J, Wong MY, Wang SSH, de Nola G, Nyitrai H, Kirchhausen T, et al. PKC-phosphorylation of Liprin-alpha3 triggers phase separation and controls presynaptic active zone structure. Nat Commun. 2021;12:3057. https://doi.org/10.1038/s41467-021-23116-w.
Kittelmann M, Hegermann J, Goncharov A, Taru H, Ellisman MH, Richmond JE, et al. Liprin-alpha/SYD-2 determines the size of dense projections in presynaptic active zones in C. elegans. J Cell Biol. 2013;203:849–63. https://doi.org/10.1083/jcb.201302022.
Taru H, Jin Y. The Liprin homology domain is essential for the homomeric interaction of SYD-2/Liprin-alpha protein in presynaptic assembly. J Neurosci. 2011;31:16261–8. https://doi.org/10.1523/JNEUROSCI.0002-11.2011.
Liang M, Jin G, Xie X, Zhang W, Li K, Niu F, et al. Oligomerized liprin-alpha promotes phase separation of ELKS for compartmentalization of presynaptic active zone proteins. Cell Rep. 2021;34:108901. https://doi.org/10.1016/j.celrep.2021.108901.
Ahmari SE, Buchanan J, Smith SJ. Assembly of presynaptic active zones from cytoplasmic transport packets. Nat Neurosci. 2000;3:445–51. https://doi.org/10.1038/74814.
Zhai RG, Vardinon-Friedman H, Cases-Langhoff C, Becker B, Gundelfinger ED, Ziv NE, et al. Assembling the presynaptic active zone: a characterization of an active one precursor vesicle. Neuron. 2001;29:131–43. https://doi.org/10.1016/s0896-6273(01)00185-4.
Garner CC, Zhai RG, Gundelfinger ED, Ziv NE. Molecular mechanisms of CNS synaptogenesis. Trends Neurosci. 2002;25:243–51. https://doi.org/10.1016/s0166-2236(02)02152-5.
Maas C, Torres VI, Altrock WD, Leal-Ortiz S, Wagh D, Terry-Lorenzo RT, et al. Formation of Golgi-derived active zone precursor vesicles. J Neurosci. 2012;32:11095–108. https://doi.org/10.1523/JNEUROSCI.0195-12.2012.
Vukoja A, Rey U, Petzoldt AG, Ott C, Vollweiter D, Quentin C, et al. Presynaptic biogenesis requires axonal transport of lysosome-related vesicles. Neuron. 2018;99:1216–1232 e1217. https://doi.org/10.1016/j.neuron.2018.08.004.
Wagner OI, Esposito A, Kohler B, Chen CW, Shen CP, Wu GH, et al. Synaptic scaffolding protein SYD-2 clusters and activates kinesin-3 UNC-104 in C. elegans. Proc Natl Acad Sci U S A. 2009;106:19605–10. https://doi.org/10.1073/pnas.0902949106.
Shin H, Wyszynski M, Huh KH, Valtschanoff JG, Lee JR, Ko J, et al. Association of the kinesin motor KIF1A with the multimodular protein liprin-alpha. J Biol Chem. 2003;278:11393–401. https://doi.org/10.1074/jbc.M211874200.
Hallam SJ, Goncharov A, McEwen J, Baran R, Jin Y. SYD-1, a presynaptic protein with PDZ, C2 and rhoGAP-like domains, specifies axon identity in C. elegans. Nat Neurosci. 2002;5:1137–46. https://doi.org/10.1038/nn959.
Owald D, Fouquet W, Schmidt M, Wichmann C, Mertel S, Depner H, et al. A Syd-1 homologue regulates pre- and postsynaptic maturation in Drosophila. J Cell Biol. 2010;188:565–79. https://doi.org/10.1083/jcb.200908055.
Wentzel C, Sommer JE, Nair R, Stiefvater A, Sibarita JB, Scheiffele P. mSYD1A, a mammalian synapse-defective-1 protein, regulates synaptogenic signaling and vesicle docking. Neuron. 2013;78:1012–23. https://doi.org/10.1016/j.neuron.2013.05.010.
Spinner MA, Walla DA, Herman TG. Drosophila Syd-1 Has RhoGAP activity that is required for presynaptic clustering of Bruchpilot/ELKS but not Neurexin-1. Genetics. 2018;208:705–16. https://doi.org/10.1534/genetics.117.300538.
Patel MR, Lehrman EK, Poon VY, Crump JG, Zhen M, Bargmann CI, et al. Hierarchical assembly of presynaptic components in defined C. elegans synapses. Nat Neurosci. 2006;9:1488–98. https://doi.org/10.1038/nn1806.
Li L, Tian X, Zhu M, Bulgari D, Bohme MA, Goettfert F, et al. Drosophila Syd-1, liprin-alpha, and protein phosphatase 2A B' subunit Wrd function in a linear pathway to prevent ectopic accumulation of synaptic materials in distal axons. J Neurosci. 2014;34:8474–87. https://doi.org/10.1523/JNEUROSCI.0409-14.2014.
Owald D, Khorramshahi O, Gupta VK, Banovic D, Depner H, Fouquet W, et al. Cooperation of Syd-1 with Neurexin synchronizes pre- with postsynaptic assembly. Nat Neurosci. 2012;15:1219–26. https://doi.org/10.1038/nn.3183.
Chia PH, Patel MR, Shen K. NAB-1 instructs synapse assembly by linking adhesion molecules and F-actin to active zone proteins. Nat Neurosci. 2012;15:234–42. https://doi.org/10.1038/nn.2991.
Fulterer A, Andlauer TFM, Ender A, Maglione M, Eyring K, Woitkuhn J, et al. Active Zone Scaffold protein ratios tune functional diversity across brain synapses. Cell Rep. 2018;23:1259–74. https://doi.org/10.1016/j.celrep.2018.03.126.
Muhammad K, Reddy-Alla S, Driller JH, Schreiner D, Rey U, Bohme MA, et al. Presynaptic spinophilin tunes neurexin signalling to control active zone architecture and function. Nat Commun. 2015;6:8362. https://doi.org/10.1038/ncomms9362.
Patel MR, Shen K. RSY-1 is a local inhibitor of presynaptic assembly in C. elegans. Science. 2009;323:1500–3. https://doi.org/10.1126/science.1169025.
Ramesh N, Escher MJF, Mampell MM, Bohme MA, Gotz TWB, Goel P, et al. Antagonistic interactions between two Neuroligins coordinate pre- and postsynaptic assembly. Curr Biol. 2021;31:1711–1725 e1715. https://doi.org/10.1016/j.cub.2021.01.093.
Hsueh YP. The role of the MAGUK protein CASK in neural development and synaptic function. Curr Med Chem. 2006;13:1915–27. https://doi.org/10.2174/092986706777585040.
Butz S, Okamoto M, Sudhof TC. A tripartite protein complex with the potential to couple synaptic vesicle exocytosis to cell adhesion in brain. Cell. 1998;94:773–82. https://doi.org/10.1016/s0092-8674(00)81736-5.
Olsen O, Moore KA, Fukata M, Kazuta T, Trinidad JC, Kauer FW, et al. Neurotransmitter release regulated by a MALS-liprin-alpha presynaptic complex. J Cell Biol. 2005;170:1127–34. https://doi.org/10.1083/jcb.200503011.
Samuels BA, Hsueh YP, Shu T, Liang H, Tseng HC, Hong CJ, et al. Cdk5 promotes synaptogenesis by regulating the subcellular distribution of the MAGUK family member CASK. Neuron. 2007;56:823–37. https://doi.org/10.1016/j.neuron.2007.09.035.
Atasoy D, Schoch S, Ho A, Nadasy KA, Liu X, Zhang W, et al. Deletion of CASK in mice is lethal and impairs synaptic function. Proc Natl Acad Sci U S A. 2007;104:2525–30. https://doi.org/10.1073/pnas.0611003104.
Ho A, Liu X, Sudhof TC. Deletion of Mint proteins decreases amyloid production in transgenic mouse models of Alzheimer's disease. J Neurosci. 2008;28:14392–400. https://doi.org/10.1523/JNEUROSCI.2481-08.2008.
Acknowledgments
We thank many researchers for their beautiful work in the studies of CAZ proteins and apologize for selective citations due to space limit. We acknowledge funds from NIH that have supported the research in our laboratories.
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this chapter
Cite this chapter
Jin, Y., Zhai, R.G. (2023). Presynaptic Cytomatrix Proteins. In: Wang, ZW. (eds) Molecular Mechanisms of Neurotransmitter Release. Advances in Neurobiology, vol 33. Springer, Cham. https://doi.org/10.1007/978-3-031-34229-5_2
Download citation
DOI: https://doi.org/10.1007/978-3-031-34229-5_2
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-34228-8
Online ISBN: 978-3-031-34229-5
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)