Abstract
Bioluminescence is one of the most fascinating features of basidiomycetes. In this chapter, we have reviewed a total of 105 bioluminescent fungi that have been documented to date. We show that these species are restricted to three major lineages, including mycenoid, Armillaria, and Omphalotus, and display different bioluminescent patterns and geographical distributions. A global phylogeny shows that these species also exhibit patchy phylogenetic placement with their non-bioluminescent relatives. We review previous work on their genomes, which contain the fungal luciferin biosynthetic cluster. The dynamics of this cluster provide an evolutionary scenario since the last common ancestor of the family Mycenaceae and the marasmioid clade of Agaricales. Finally, we discuss potential functions and possible research directions of bioluminescence in these fungi.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
Bioluminescence is the emission of light by a living organism, generated by a chemical reaction that involves proteins called luciferases that catalyze the oxidation of substrates called luciferins. It is a trait that has evolved at least 94 times independently across the tree of life (Lau and Oakley 2021), with over 700 genera of bioluminescent organisms recorded from freshwater, terrestrial, and marine environments (Haddock et al. 2010; Widder 2010; Lau and Oakley 2021; Delroisse et al. 2021). Bioluminescence can be further categorized into two types, autogenic and bacteriogenic, in which light is produced either by the organism itself or bioluminescent bacterial symbionts, respectively (Lau and Oakley 2021). Bioluminescence in fungi is autogenic, which means fungi emit light using their own luciferin and luciferase (Oliveira et al. 2012; Oliveira and Stevani 2009; Airth and Mc 1959). Fungal luciferin is different from the other 13 known luciferins (Lau and Oakley 2021; Purtov et al. 2015) and all fungal luciferases are homologous and distinct from other known luciferases used by other organisms based on sequence similarity (Delroisse et al. 2021; Kotlobay et al. 2018).
All bioluminescent fungi belong to the Agaricales of Basidiomycetes, which is a major order comprising around 13,000 gilled mushrooms (Kirk et al. 2008). These fungi belong to ten different unrelated genera although some of which have taxonomic synonyms. The apparent patchy phylogenetic distribution of bioluminescence in fungi and the conserved genetic mechanisms underlying phenotype can be explained by either convergent evolution or an intricate loss/gain history. Recent phylogenomic progress has resolved the phylogenetic relationships among species and helps us answer those questions.
This review focuses on the evolution of bioluminescence in mushrooms in the Agaricales order. The recent identification of the luciferin biosynthetic pathway genes and new phylogenomic analyses of bioluminescent fungi and their sister taxa have shown that fungal bioluminescence occurred in the last common ancestor of the mycenoid and the marasmioid clade. We will review the features of bioluminescent fungi, the known genes involved in the bioluminescence, and their evolutionary history.
2 Diversity of Bioluminescent Fungi
Bioluminescence has long captured the imagination of scientists. The taxonomy and evolution of bioluminescent fungi was first compiled and discussed 70 years ago (Wassink 1978; Harvey 1952). The number of taxonomically reported bioluminescent fungi continues to increase with improvement over higher sensitivity in light detection initially from the naked eye to digital camera, luminometer, or photomultiplier. Two reviews have updated the accounts to 64 luminescent species in 2008 (Desjardin et al. 2008) and 81 species in 2015 (Chew et al. 2015). We have removed the phylogenetically uncertain species and merged synonyms where possible, as well as compiled and assigned a list of 28 additional bioluminescent species (Mihail 2015; Desjardin et al. 2016; Cortes-Perez et al. 2019; Chang et al. 2020; Karunarathna et al. 2020; Dauner et al. 2021; Terashima et al. 2016). The current number of recorded bioluminescent fungi is 105. In recent years, molecular data were generated using barcode DNA sequences including rRNA ITS (Internal transcribed spacer), large subunit rRNA (LSU), and RNA polymerase II (rpb2) to revise the phylogenetic positions of these species, which can be classified into three lineages (Fig. 12.1): 80 in the mycenoid (78 Mycenaceae and two from the Lucentipes lineage) (Table 12.1), 11 in the Armillaria lineage and 13 in the Omphalotus lineage, and one species (Pleurotus nitidus Har. Takah. & Taneyama) with uncertain position (Table 12.2). With the availability of molecular markers, the term mycenoid was originally intended to describe various morphological features that are now coherently grouped in species belonging to the Mycenaceae family. A fourth lineage (Lucentipes) containing the bioluminescent Gerronema viridilucens Desjardin, Capelari & Stevani (Desjardin et al. 2005) and Mycena lucentipes Desjardin, Capelari & Stevani (Desjardin et al. 2007) which were classified as mycenoid species is excluded in this review because of the lack of available marker sequences. The three lineages shared a common origin in the last common ancestor of Mycenaceae and the marasmioid clade. For more details on the relationships, see Ke et al. 2020. Here, we summarize the properties of these three major bioluminescent fungal lineages.
Overview of bioluminescent fungi lineages and their sister groups. Basidiomes of Mycena kentingensis in light (a-1) and dark (a-2) exposures and its mycelium in light (a-3) and dark (a-4) cultured in compost. Basidiomes of Mycena luguensis (b-1) and its mycelium in light (b-2) and dark (b-3) grown on wood. Basidiomes of Armillaria mellea (c-1) and its cultured mycelium in light (c-2) and dark (c-3). Basidiomes of Omphalotus japonicus on natural host tree (d-1) and the fruiting body in light (d-2) and dark (d-3)
2.1 Mycenaceae Family
The Mycenaceae family contains most species originally part of the mycenoid lineage (Desjardin et al. 2008), which are a type of mushrooms whose gills adhere to cartilaginous and central stipes and lack a ring or volva (Ulloa and Hanlin 2017). The mycenoid lineage includes the majority of recorded bioluminescent fungi (80) belonging to the Mycenaceae family with the exception of two species (Gerronema viridilucens and Mycena lucentipes; Desjardin et al. 2008). In this review, we focus on the diversity of Mycenaceae. Mycenaceae contains over 600 species including Atheniella, Dictyopanus, Favolaschia, Mycena, Mycenoporella, Panellus, Resinomycena, Roridomyces, and Xeromphalina (Redhead et al. 2012; Kirk et al. 2008). Members of this family are known primarily to colonize leaves or wood as litter- or wood-decaying saprophytes (Moncalvo et al. 2002). Still, other members of this family were reported to use different nutritional strategies such as orchid mycorrhizae (Ogura-Tsujita et al. 2009; Martos et al. 2009; Zhang et al. 2012), the coffee pathogen Mycena citricolor (Rao and Tewari 1987; Quesada-Chanto and Jimenez-Ulate 1996), a plant root invader (Harder et al. 2021), and tissues in moss and root system in healthy plant (Davey et al. 2013; Liao et al. 2014; Botnen et al. 2014; Lorberau et al. 2017; Kernaghan and Patriquin 2011). ITS is the most common marker used in this lineage. We downloaded available ITS sequences (>500 bp) from the National Center for Biotechnology Information (NCBI) according to taxId including Mycenaceae, Favolaschia, Filoboletus, Dictyopanus, and Roridomyces. We constructed a phylogentic tree of 287 non-redundant sequences with species information using IQ-TREE (Nguyen et al. 2015). An alignment length of 1220 bp (last updated 2022.09.11; https://github.com/HueiMien/bioluminescent-fungi.git) was produced with MAFFT (Katoh and Standley 2013) and trimming with trimal (Capella-Gutierrez et al. 2009). (Fig. 12.2). We found that the bioluminescent trait was scattered along the Mycenaceae phylogeny and did not group to a specific genus. This observation is consistent with previous patchy distributions observed in smaller datasets using different molecular markers and focusing on different closely related species (Chew et al. 2015; Karunarathna et al. 2020; Dauner et al. 2021; Shih et al. 2014; Chang et al. 2020).
ITS tree of 287 Mycenaceae sequences with two Armillaria species as outgroup which was constructed by IQ-TREE (Nguyen et al. 2015) with an alignment length of 1220 bp without redundant species downloaded from NCBI. Numbers on each branch denote support values (SH-aLRT support (%) / ultrafast bootstrap support (%)). Dark green nodes annotated with a star denote the bioluminescent species. Note that the sequences are not necessary the same as the holotype, and not all bioluminescent Mycenaceae species were listed here since not all of them have been recorded in NCBI. The inset indicates the lineage with species belong to sect. Calodontes
Among different species, luminescence also showed diverse patterns in different tissues, including bioluminescent mycelia, caps and stipes of basidiomes (Table 12.1).
Our revision was consistent with the observations of Desjardin et al. 2008. Overall, the species with bioluminescent basidiomes mostly have bioluminescent mycelia (26/32) but some have non-bioluminescent mycelia (6/32). The variations can usually be found between closely related species, indicating that the expression of bioluminescent is not restricted to specific tissues. An example was the Mycena sect. Calodontes shown in Fig. 12.2. These 13 species with >85.8 ITS nucleotide identity form a monophyletic clade with seven non-bioluminescent species and six bioluminescent species, including two species with bioluminescent mycelia but non-bioluminescent fruiting bodies, as well as four species with bioluminescent fruiting bodies with different intensity in the stipes and caps. Different species also displayed different degrees of endemism (Fig. 12.3). M. pura is the most widely distributed across different continents. By contrast, M. chlorophos is restricted in Asia, and M. kentingensis has only been found in Taiwan so far.
2.2 Armillaria lineage
This lineage includes all members of genus Armillaria comprising more than 40 species (Baumgartner et al. 2011; Anderson and Stasovski 1992; Piercey-Normore et al. 1998; Coetzee et al. 2001; Koch et al. 2017). The Armillaria lineage and its sister group, the physalacrioid lineage (all of which are non-bioluminescent), form the family Physalacriaceae (Moncalvo et al. 2002), which includes 11 genera and 169 species (Kirk et al. 2008). Most of them can decompose organic material as saprotrophs and infect plants as necrotrophic pathogens when they are in a suitable environment (Sipos et al. 2018; Shaw et al. 1991; Smith et al. 1992; Kedves et al. 2021). Some of them are associated with orchid mycorrhizas (Martos et al. 2009; Cha and Igarashi 1995; Sekizaki et al. 2008; Guo et al. 2016).
Eleven species in the Armillaria lineage have been confirmed to have a luminescent property in their mycelium (Table 12.2), and it has been believed that the mycelium of most Armillaria species is luminescent (Desjardin et al. 2008). Among them, only Desarmillaria ectypa (Synonym A. ectypa) emit light in both mycelium and fruitbodies (mainly from the gills) so far in report. Based on the classification of a 2018 review using a translation elongation factor subunit 1-alpha (tef-1α) (Coetzee et al. 2018), nine sub-lineages were proposed (Fig. 12.4). Most of the known luminescent species according to phenotypic observations belong to the Holarctic lineages including the A. gallica, A. solidipes/ostoyae, A. mellea, A. mexicana, and Exannulated (subgenus Desarmillaria) lineages. In addition, based on the recent genomic content analysis, all of the analyzed 14 species have conserved luciferin biosynthetic pathway genes (mentioned later), which extend bioluminescence to two more lineages of Armillaria (A. luteobubalina and A. novae-zelandiae Lineages).
The abbreviation of a phylogeny from (Coetzee et al. 2018), created using tef-1α sequences. Species with dark green denote the bioluminescent fungi according to observation of light emission, those with light green denotes the bioluminescent fungi with prediction of luciferase cluster in genome. The figure was adapted with permission and redrawn
Even when based just on those recorded bioluminescent fungi, it does not seem that the ability to emit light is related to geographic distribution. For example, the known luminescent species include A. gemina which is confined to North America, A. fuscipes which is confined to Africa, and A. ostoyae which has transcontinental distribution across Europe, North America, and Asia. Light emission was recorded in Desarmillaria ectypa and D. tabescens, which belongs to the monophyletic group of the Exannulated lineage basal to all Armillaria species, but no light emission has been recorded in the closely related genus Guyanagaster. Interestingly, the genome of G. necrorhizus maintains the luciferase cluster but with a truncated luciferase gene (Sect. 12.3).
2.3 Omphalotus Lineage
The Omphalotus lineage includes the genera Omphalotus and Neonothopanus (Kirchmair et al. 2004). According to a 2019 review (Oliveira et al. 2019) and Mycobank and Index Fungorum, in addition to Omphalotus and Neonothopanus, Omphalotaceae includes the other nine genera (Anthracophyllum, Connopus, Gymnopanella, Gymnopus, Lentinula, Marasmiellus, Marasmius, Mycetinis, Pusillomyces, and Rhodocollybia). It has been suspected that all species from the Omphalotus lineage form luminescent fruiting bodies but that their mycelia can be either luminescent or non-luminescent. They are known for large mushrooms and Neonothopanus gardneri and N. nambi were well-characterized by bioluminescent mechanisms (Oliveira et al. 2015) and the relevant enzyme activity (Kotlobay et al. 2018).
Three new species, Marasmiellus lucidus, M. venosus, and Pleurotus nitidus, with bioluminescent fruiting bodies were recently discovered outside the current three major bioluminescent lineages (Terashima et al. 2016). We successfully amplified and sequenced the ITS sequences of Marasmiellus lucidus and M. venosus (submitted to NCBI; accession pending) and found both of them were part of Anthracophyllum lineage which is sister to the Omphalotus lineage (Fig. 12.5). As it stands, Pleurotus nitidus remains the only record of bioluminescent fungus in the entire Pleurotaceae family. We speculate it may be misclassified that warrants further investigation as this family often was confused with species belonging in the Omphalotus lineage.
(a), ITS tree of 34 Omphalotaceae sequences with five species of Marasmiaceae as an outgroup. The tree was constructed by IQ-TREE (Nguyen et al. 2015) with an alignment length of 1373 bp and without redundant species downloaded from NCBI. Numbers on each branch denote support values (SH-aLRT support (%) / ultrafast bootstrap support (%)). Nodes with dark green color denote bioluminescent species. Note that the sequences not necessary the same as the holotype, and not all bioluminescent species were listed here since not all of them have been recorded in NCBI. The clade name in the front of species name according to previous review (Oliveira et al. 2019). (b), The geographic distribution of bioluminescent fungi recorded by extracting ITS (Internal transcribed spacer) from NCBI and converting the collected country to six continents
3 Genes Responsible for Bioluminescence; Luciferase Cluster: Luz and Luciferin Biosynthesis Genes
Green light (~530 nm) emission in fungi was confirmed to be a result of an enzyme-mediated reaction in which a hot extract containing heat-stable substrates (e.g., luciferin) mixed with a cold extract containing enzymes (e.g., luciferase) (Oliveira and Stevani 2009).
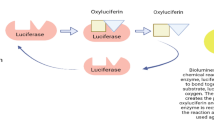
Later, one single bioluminescent system shared by all known bioluminescent fungal lineages was validated by detecting bioluminescence from combinations of substrate/enzyme extract from bioluminescent species from four lineages (Gerronema viridilucens from Lucentipes lineage, Armillaria mellea from Armillaria lineage, Mycena luxaeterna from Mycenaceae lineage, and Neonothopanus gardneri from Omphalotus lineage) and comparing them to nonluminous species (Oliveira et al. 2012). The structure of fungal luciferin and its precursor were identified in 2015 as 3-hydroxyhispidin and hispidin in the bioluminescent Neonothopanus nambi, Mycena citricolor, Panellus stipticus, and Armillaria borealis (Purtov et al. 2015). In 2017, the light-emitting process was demonstrated by reporting the structure of the emitter, oxyluciferin, and oxyluciferin is enzymatically hydrolyzed yielding caffeic acid (Kaskova et al. 2017). The biosynthetic cycle was finalized and the four enzymes involved were identified in N. nambi (Fig. 12.6a): Hispidin synthase (hisps) converts caffeic acid to hispidin; hispidin-3-hydroxylase (h3h) hydroxylates hispidin to give 3-hydroxyhispidin, the fungal luciferin; and the oxidation of luciferin by luciferase (luz) yields an unstable high-energy intermediate, which emits light while turning into oxyluciferin, a substance that can be recycled to caffeic acid by caffeylpyruvate hydrolase (cph) (Kotlobay et al. 2018). The luz, h3h, and hispS were introduced in yeast Pichia pastoris for autoluminescence.
(a), Fungal luciferin biosynthetic pathway proposed by (Kotlobay et al. 2018). (b), Tree topology from species with genomes and their gene organization of the luciferase cluster inferred from (Ke et al. 2020) and (Kotlobay et al. 2018). Block arrows indicate genes and their orientation. The dashed block arrows denote the loss of a gene. All the species with dark green are bioluminescent species with records in the literature. The species with light green are the species believed to illuminate
4 The Origin of Fungal Bioluminescence
Like other bioluminescent species, the discovery of bioluminescent fungi, with their minor presence and patchy phylogenetic placement in gilled mushrooms, initially suggested they evolved independently multiple times in a convergent manner. However, recent advances in the genome sequencing of Agaricales have pointed to an alternative scenario. One study found that the genes responsible for luciferin biosynthetic cycles were physically adjacent in genomes (luciferase cluster; Fig. 12.6b) by comparing genomes from ten bioluminescent fungi and their sister species (Kotlobay et al. 2018). By generating reference genomes of bioluminescent fungal genomes from five Mycenaceae (Ke et al. 2020), eight Armillaria lineages, and two Omphalotus lineages, we were able to investigate the evolutionary dynamics around the luciferase cluster. Members of the luciferase cluster were clearly homologous across distant bioluminescent fungi, suggesting that bioluminescence was not independently evolved. Both studies concluded that bioluminescence originated from the last common ancestor of the Mycenaceae family and marasmioid clade of Agaricales, although the species trees in these two studies were not congruent. The latter showed that the Schizophyllaceae was the closest lineage with the bioluminescent clade and had a genome encoding a homologous luciferase. Through phylogenetic reconciliation, it was found that the species tree was congruent to the gene trees in the luciferase cluster, suggesting the patchy distribution was a result of a loss of the luciferase cluster in the majority of Agaricales rather than horizontal gene transfer (Ke et al. 2020). However, the origin of these luciferin biosynthetic pathway genes within Schizophyllaceae is still unknown. Our experiment of adding hispidin (luciferin precursor) to Schizophyllum commune (unpublished data) showed that it could not be converted to light.
A fungal gene cluster can be differentially maintained due to differences in genome plasticity (Marcet-Houben and Gabaldon 2019). The luciferase cluster was located in different low synteny regions of Mycena genomes, which may suggest that they were dispensable and could be lost in most species in the Mycenaceae lineage. In contrast, the luciferase cluster was located in the conserved syntenic region in all eight of the bioluminescent fungal genomes from the Armillaria lineage. Ke et al. proposed an evolutionary scenario that the luciferase cluster was originally located at the dispensable region of the last common ancestor and the cluster was translocated to different genomic locations through rearrangement (Ke et al. 2020). In the Armillaria lineage, the luciferase cluster was translocated to the core region and became more conserved.
5 Biological Function of Bioluminescent Fungi
The roles of bioluminescence in various organisms have been extensively studied (for review please see (Lau and Oakley 2021)), and often involve mating, luring prey or defense. At the molecular level, negative selection was detected on the luciferase cluster, and it could explain why the luciferase cluster had been maintained across hundreds of millions of years (Ke et al. 2020) suggesting that bioluminescence may play a role in environmental adaptation but in a species-specific way. Several pieces of evidence support different possible hypotheses for the function of bioluminescence. First, several studies proposed that fungal bioluminescence may facilitate spore dispersal by attracting potential vectors. More recently, in the Brazilian forest, a biological circadian control of luminescence was found by measuring the profiles of the luciferase-, reductase-, and luciferin-rich extract from mycelia of N. gardneri, as well as insects captured using an artificial mushroom with light similar to the bioluminescence of dark mushrooms (Oliveira et al. 2015). It was proposed that the bioluminescent N. gardneri of the Omphalotus lineage increased spore dispersal by attracting arthropods in the evening. However, the insects’ attraction has not been proven in other bioluminescent species in the same lineage, such as Omphalotus nidiformis (Weinstein et al. 2016).
Second, fungal bioluminescence is a by-product of an unknown biological process. This hypothesis has been proposed two decades ago according to the existence of luminous and nonluminous populations of the same species or the continuous light production in bioluminescent fungi (Herring 1994; Deheyn and Latz 2007). Recent studies revealing the expression of bioluminescence also indicate a biological function. It has been found that the synthesis of luciferin precursors and H3H was inhibited during the Armillaria development from mycelium to mushroom (Purtov et al. 2017; Mihail et al. 2018; Puzyr et al. 2017). This suppression resulted in decreased luminescence indicating a possible form of regulation and biological role. There is a divergent intensity in cap, stipe, or mycelium of species of Mycenaceae lineage. Several genes that could be responsible for this regulation were enriched when investigating the differential expression of genes among different stages of Mycena kentingensis (Ke et al. 2020). In addition, a diversity of intensities was also found in cultured mycelium among species in the Mycenaceae lineage (Fig. 12.7). Since regulation implies an adaptive function, these observations and analyses provide indirect evidence for a biological role. However, further validation will be necessary to clear up the biological role for each species.
6 Conclusions
Bioluminescence in fungi is an intriguing feature restricted to gilled mushrooms that utilizes a single type of biochemical reaction. We have been able to understand the evolution of bioluminescent mushrooms better recently thanks to the accumulated record of which fungi are bioluminescent, molecular studies on luciferin biosynthetic pathways, and genome research. When the first fungal luciferase gene cluster of Neonothopanus nambi was identified in 2018, this successfully heterogeneously expressed biosynthetic pathway advanced the further study and application of fungal bioluminescence ((Kotlobay et al. 2018); for a detailed review on potential applications see (Ke and Tsai 2021)). The homologous luciferase found from the genomic sequencing of several bioluminescent fungi and their sister groups allowed us to trace the evolutionary history to a last common ancestor of the Mycenaceae family and the marasmioid clade within Agaricales. Despite a great deal of progress, many important questions still remain. Important goals for future work include developing a more complete understanding of bioluminescent features, ecological function and its gene regulation.
References
Ainsworth M (2004) Searching for luminous mushrooms of the marsh fungus Armillaria ectypa. Field Mycol 5(4):142–144
Airth RL, Mc EW (1959) Light emission from extracts of luminous fungi. J Bacteriol 77(2):249–250. https://doi.org/10.1128/jb.77.2.249-250.1959
Anderson JB, Stasovski E (1992) Molecular phylogeny of northern-hemisphere species of Armillaria. Mycologia 84(4):505–516. https://doi.org/10.2307/3760315
Aravindakshan DM, Kumar TKA, Manimohan P (2012) A new bioluminescent species of Mycena sect. Exornatae from Kerala State, India. Mycosphere 3(5):556–561. https://doi.org/10.5943/mycosphere/3/5/4
Baumgartner K, Coetzee MP, Hoffmeister D (2011) Secrets of the subterranean pathosystem of Armillaria. Mol Plant Pathol 12(6):515–534. https://doi.org/10.1111/j.1364-3703.2010.00693.x
Berkeley MJ (1844) Decades of fungi London J Bot 3:185–194
Berliner MD (1961a) Diurnal periodicity of luminescence in three basidiomycetes. Science 134(3481):740. https://doi.org/10.1126/science.134.3481.740
Berliner MD (1961b) Studies in fungal luminescence. Mycologia 53(1):84–90. https://doi.org/10.2307/3756133
Bermudes D, Petersen RH, Nealson KH (1992) Low-level bioluminescence detected in Mycena-Haematopus Basidiocarps. Mycologia 84(5):799–802. https://doi.org/10.2307/3760392
Bigelow HE, Miller JOK, Thiers HD (1976) A new species of Omphalotus. Mycotaxon 3(3):363–372
Bothe F (1930) Ein neuer einheimischer Leuchtpilz, Mycena tintinnabulum. Ber Deut Bot Ges 48:394–399
Bothe F (1931) Über das Leuchten verwesender Blätter und seine Erreger. Planta 14:752–765. https://doi.org/10.1007/bf01917160
Botnen S, Vik U, Carlsen T, Eidesen PB, Davey ML, Kauserud H (2014) Low host specificity of root-associated fungi at an Arctic site. Mol Ecol 23(4):975–985. https://doi.org/10.1111/mec.12646
Capelari M, Desjardin DE, Perry BA, Asai T, Stevani CV (2011) Neonothopanus gardneri: a new combination for a bioluminescent agaric from Brazil. Mycologia 103(6):1433–1440. https://doi.org/10.3852/11-097
Capella-Gutierrez S, Silla-Martinez JM, Gabaldon T (2009) trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25(15):1972–1973. https://doi.org/10.1093/bioinformatics/btp348
Cha JY, Igarashi T (1995) Armillaria species associated with Gastrodia elata in Japan. Eur J For Pathol 25(6–7):319–326
Chang CC, Chen CY, Lin WW, Kao HW (2020) Mycena jingyinga, Mycena luguensis, and Mycena venus: three new species of bioluminescent fungi from Taiwan. Taiwania 65(3):396–406. https://doi.org/10.6165/tai.2020.65.396
Chew ALC, Desjardin DE, Tan YS, Musa MY, Sabaratnam V (2015) Bioluminescent fungi from Peninsular Malaysia-a taxonomic and phylogenetic overview. Fungal Divers 70(1):149–187. https://doi.org/10.1007/s13225-014-0302-9
Chew ALC, Tan YS, Desjardin DE, Musa MY, Sabaratnam V (2013) Taxonomic and phylogenetic re-evaluation of Mycena illuminans. Mycologia 105(5):1325–1335. https://doi.org/10.3852/13-009
Chew AL, Tan YS, Desjardin DE, Musa MY, Sabaratnam V (2014) Four new bioluminescent taxa of Mycena sect. Calodontes from Peninsular Malaysia. Mycologia 106(5):976–988. https://doi.org/10.3852/13-274
Coetzee MPA, Wingfield BD, Bloomer P, Ridley GS, Kile GA, Wingfield MJ (2001) Phylogenetic relationships of Australian and New Zealand Armillaria species. Mycologia 93(5):887–896. https://doi.org/10.2307/3761754
Coetzee MPA, Wingfield BD, Wingfield MJ (2018) Armillaria root-rot pathogens: species boundaries and global distribution. Pathogens 7(4):83. https://doi.org/10.3390/pathogens7040083
Corner EJH (1950) Descriptions of two luminous tropical agarics (Dictyopanus and Mycena). Mycologia 42:423–431
Corner EJH (1954) Further descriptions of luminous agarics. Trans Br Mycol Soc 37:256–271
Corner EJH (1981) The agaric genera Lentinus, Panus, and Pleurotus: with particular reference to Malaysian species. Beihefte zur Nova Hedwigia; Heft 69. J. Cramer, Vaduz
Cortes-Perez A, Desjardin DE, Perry BA, Ramirez-Cruz V, Ramirez-Guillen F, Villalobos-Arambula AR, Rockefeller A (2019) New species and records of bioluminescent Mycena from Mexico. Mycologia 111(2):319–338. https://doi.org/10.1080/00275514.2018.1554172
Dauner LAP, Karunarathna SC, Tibpromma S, Xu JC, Mortimer PE (2021) Bioluminescent fungus Roridomyces viridiluminus sp. nov. and the first Chinese record of the genus Roridomyces, from Southwestern China. Phytotaxa 487(3):233–250. https://doi.org/10.11646/phytotaxa.487.3.4
Davey ML, Heimdal R, Ohlson M, Kauserud H (2013) Host- and tissue-specificity of moss-associated Galerina and Mycena determined from amplicon pyrosequencing data. Fungal Ecol 6(3):179–186. https://doi.org/10.1016/j.funeco.2013.02.003
Deheyn DD, Latz MI (2007) Bioluminescence characteristics of a tropical terrestrial fungus (Basidiomycetes). Luminescence 22(5):462–467. https://doi.org/10.1002/bio.985
Delroisse J, Duchatelet L, Flammang P, Mallefet J (2021) Leaving the dark side? Insights into the evolution of luciferases. Front Mar Sci 8:ARTN 673620. https://doi.org/10.3389/fmars.2021.673620
Desjardin DE, Braga-Neto R (2007) Mycena lacrimans, a rare species from Amazonia, is bioluminescent. Edinb J Bot 64(3):275–281
Desjardin DE, Capelari M, Stevani CV (2005) A new bioluminescent agaric from Sao Paulo, Brazil. Fungal Divers 18:9–14
Desjardin DE, Capelari M, Stevani C (2007) Bioluminescent Mycena species from Sao Paulo, Brazil. Mycologia 99(2):317–331. https://doi.org/10.3852/mycologia.99.2.317
Desjardin DE, Oliveira AG, Stevani CV (2008) Fungi bioluminescence revisited. Photochem Photobiol Sci 7(2):170–182. https://doi.org/10.1039/b713328f
Desjardin DE, Perry BA, Lodge DJ, Stevani CV, Nagasawa E (2010) Luminescent Mycena: new and noteworthy species. Mycologia 102(2):459–477
Desjardin DE, Perry BA, Stevani CV (2016) New luminescent mycenoid fungi (Basidiomycota, Agaricales) from Sao Paulo State, Brazil. Mycologia 108(6):1165–1174. https://doi.org/10.3852/16-077
Guo T, Wang HC, Xue WQ, Zhao J, Yang ZL (2016) Phylogenetic analyses of Armillaria reveal at least 15 phylogenetic lineages in China, seven of which are associated with cultivated Gastrodia elata. PLoS One 11(5):e0154794. https://doi.org/10.1371/journal.pone.0154794
Haddock SH, Moline MA, Case JF (2010) Bioluminescence in the sea. Annu Rev Mar Sci 2:443–493. https://doi.org/10.1146/annurev-marine-120308-081028
Haneda Y (1955) Luminous organisms of Japan and the Far East. The Luminescence of Biological Systems. American Association for the Advancement of Science, Washington DC
Harder CB, Hesling E, Botnen SS, Dima B, von Bonsdorff-Salminen T, Niskanen T, Jarvis SG, Lorberau KE, Ouimette A, Hester A, Hobbie EA, Taylor AFS, Kauserud H (2021) Mycena species can be opportunist-generalist plant root invaders. bioRxiv. https://doi.org/10.1101/2021.03.23.436563
Harvey EN (1952) Bioluminescence. Academic Press, New York
Herring PJ (1994) Luminous fungi. s.n., S.l
Horak E (1978) Mycena rorida (Fr.) Quél. And related species from the southern hemisphere. Ber Schweiz Bot Ges 88(1/2):20–29
Josserand M (1953) Sur la luminescence de “Mycena rorida” en Europe occidentale. Bull Mens Soc Linn Lyon 22(4):99–102
Karunarathna SC, Mortimer PE, Tibpromma S, Dutta AK, Paloi S, Hu YW, Baurah G, Axford S, Marciniak C, Luangharn T, Madawala S, Lin C, Chen JZ, Acharya K, Kobmoo N, Samarakoon MC, Karunarathna A, Gao SY, Xu JC, Lumyong S (2020) Roridomyces phyllostachydis (Agaricales, Mycenaceae), a new bioluminescent fungus from Northeast India. Phytotaxa 459(2):155–167. https://doi.org/10.11646/phytotaxa.459.2.6
Kaskova ZM, Dorr FA, Petushkov VN, Purtov KV, Tsarkova AS, Rodionova NS, Mineev KS, Guglya EB, Kotlobay A, Baleeva NS, Baranov MS, Arseniev AS, Gitelson JI, Lukyanov S, Suzuki Y, Kanie S, Pinto E, Di Mascio P, Waldenmaier HE, Pereira TA, Carvalho RP, Oliveira AG, Oba Y, Bastos EL, Stevani CV, Yampolsky IV (2017) Mechanism and color modulation of fungal bioluminescence. Sci Adv 3(4):e1602847. https://doi.org/10.1126/sciadv.1602847
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30(4):772–780. https://doi.org/10.1093/molbev/mst010
Kawamura S (1910) Studies on the luminous fungus, Pleurotus japonicus,sp. nov. 24:275–281
Ke HM, Lee HH, Lin CI, Liu YC, Lu MR, Hsieh JA, Chang CC, Wu PH, Lu MJ, Li JY, Shang G, Lu RJ, Nagy LG, Chen PY, Kao HW, Tsai IJ (2020) Mycena genomes resolve the evolution of fungal bioluminescence. Proc Natl Acad Sci U S A 117(49):31267–31277. https://doi.org/10.1073/pnas.2010761117
Ke HM, Tsai IJ (2021) Understanding and using fungal bioluminescence – Recent progress and future perspectives. Curr Opin Green Sustain Chem 33(100570):1–6. https://doi.org/10.1016/j.cogsc.2021.100570
Kedves O, Shahab D, Champramary S, Chen L, Indic B, Boka B, Nagy VD, Vagvolgyi C, Kredics L, Sipos G (2021) Epidemiology, biotic interactions and biological control of Armillarioids in the northern hemisphere. Pathogens 10(1):76. https://doi.org/10.3390/pathogens10010076
Kernaghan G, Patriquin G (2011) Host associations between fungal root endophytes and boreal trees. Microb Ecol 62(2):460–473. https://doi.org/10.1007/s00248-011-9851-6
Kirchmair M, Morandell S, Stolz D, Poder R, Sturmbauer C (2004) Phylogeny of the genus Omphalotus based on nuclear ribosomal DNA-sequences. Mycologia 96(6):1253–1260. https://doi.org/10.2307/3762142
Kirk PM, Ainsworth GC, Bisby GR, Bioscience C (2008) Dictionary of the fungi. 10th / prepared by CABI Bioscience, 10th edn. CAB International, Wallingford
Kobayasi Y (1951) Contributions to the luminous fungi from Japan. J Hattori Bot Lab 5:1–6
Koch RA, Wilson AW, Sene O, Henkel TW, Aime MC (2017) Resolved phylogeny and biogeography of the root pathogen Armillaria and its gasteroid relative, Guyanagaster. BMC Evol Biol 17(1):33. https://doi.org/10.1186/s12862-017-0877-3
Kotlobay AA, Sarkisyan KS, Mokrushina YA, Marcet-Houben M, Serebrovskaya EO, Markina NM, Gonzalez Somermeyer L, Gorokhovatsky AY, Vvedensky A, Purtov KV, Petushkov VN, Rodionova NS, Chepurnyh TV, Fakhranurova LI, Guglya EB, Ziganshin R, Tsarkova AS, Kaskova ZM, Shender V, Abakumov M, Abakumova TO, Povolotskaya IS, Eroshkin FM, Zaraisky AG, Mishin AS, Dolgov SV, Mitiouchkina TY, Kopantzev EP, Waldenmaier HE, Oliveira AG, Oba Y, Barsova E, Bogdanova EA, Gabaldon T, Stevani CV, Lukyanov S, Smirnov IV, Gitelson JI, Kondrashov FA, Yampolsky IV (2018) Genetically encodable bioluminescent system from fungi. Proc Natl Acad Sci U S A 115(50):12728–12732. https://doi.org/10.1073/pnas.1803615115
Lau ES, Oakley TH (2021) Multi-level convergence of complex traits and the evolution of bioluminescence. Biol Rev Camb Philos Soc 96(2):673–691. https://doi.org/10.1111/brv.12672
Léveillé JH (1844) Champignons exotiques. Annales des Sciences Naturelles Botanique 3:167–221
Li J, Hu X (1993) A new species of Lampteromyces from Hunan. Act Sci Nat Univ Norm Hunan 16(2):188–189
Liao HL, Chen Y, Bruns TD, Peay KG, Taylor JW, Branco S, Talbot JM, Vilgalys R (2014) Metatranscriptomic analysis of ectomycorrhizal roots reveals genes associated with Piloderma-Pinus symbiosis: improved methodologies for assessing gene expression in situ. Environ Microbiol 16(12):3730–3742. https://doi.org/10.1111/1462-2920.12619
Liu PG (1995) Luminous fungi. Biodivers Sci 3(2):109–112
Lorberau KE, Botnen SS, Mundra S, Aas AB, Rozema J, Eidesen PB, Kauserud H (2017) Does warming by open-top chambers induce change in the root-associated fungal community of the arctic dwarf shrub Cassiope tetragona (Ericaceae)? Mycorrhiza 27(5):513–524. https://doi.org/10.1007/s00572-017-0767-y
Maas Geesteranus RA (1992) Filoboletus manipularis and some related species. Proceedings of the Koninklijke Nederlandse Akademie van Wetenschappen 95(2):267–274
Marcet-Houben M, Gabaldon T (2019) Evolutionary and functional patterns of shared gene neighbourhood in fungi. Nat Microbiol 4(12):2383–2392. https://doi.org/10.1038/s41564-019-0552-0
Martos F, Dulormne M, Pailler T, Bonfante P, Faccio A, Fournel J, Dubois MP, Selosse MA (2009) Independent recruitment of saprotrophic fungi as mycorrhizal partners by tropical achlorophyllous orchids. New Phytol 184(3):668–681. https://doi.org/10.1111/j.1469-8137.2009.02987.x
Mihail JD (2015) Bioluminescence patterns among North American Armillaria species. Fungal Biol 119(6):528–537. https://doi.org/10.1016/j.funbio.2015.02.004
Mihail JD, Bilyeu L, Lalk SR (2018) Bioluminescence expression during the transition from mycelium to mushroom in three north American Armillaria and Desarmillaria species. Fungal Biol 122(11):1064–1068. https://doi.org/10.1016/j.funbio.2018.08.007
Mihail JD, Bruhn JN (2007) Dynamics of bioluminescence by Armillaria gallica, A mellea and A tabescens. Mycologia 99(3):341–350
Miller OK Jr (1994) Observations on the genus Omphalotus in Australia. Mycol Helv 6:91–100
Moncalvo JM, Vilgalys R, Redhead SA, Johnson JE, James TY, Catherine Aime M, Hofstetter V, Verduin SJ, Larsson E, Baroni TJ, Greg Thorn R, Jacobsson S, Clemencon H, Miller OK Jr (2002) One hundred and seventeen clades of euagarics. Mol Phylogenet Evol 23(3):357–400. https://doi.org/10.1016/S1055-7903(02)00027-1
Nguyen LT, Schmidt HA, von Haeseler A, Minh BQ (2015) IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol Biol Evol 32(1):268–274. https://doi.org/10.1093/molbev/msu300
Ogura-Tsujita Y, Gebauer G, Hashimoto T, Umata H, Yukawa T (2009) Evidence for novel and specialized mycorrhizal parasitism: the orchid Gastrodia confusa gains carbon from saprotrophic Mycena. Proc Biol Sci 276(1657):761–767. https://doi.org/10.1098/rspb.2008.1225
Oliveira AG, Desjardin DE, Perry BA, Stevani CV (2012) Evidence that a single bioluminescent system is shared by all known bioluminescent fungal lineages. Photochem Photobiol Sci 11(5):848–852. https://doi.org/10.1039/c2pp25032b
Oliveira AG, Stevani CV (2009) The enzymatic nature of fungal bioluminescence. Photochem Photobiol Sci 8(10):1416–1421. https://doi.org/10.1039/b908982a
Oliveira AG, Stevani CV, Waldenmaier HE, Viviani V, Emerson JM, Loros JJ, Dunlap JC (2015) Circadian control sheds light on fungal bioluminescence. Curr Biol 25(7):964–968. https://doi.org/10.1016/j.cub.2015.02.021
Oliveira JJS, Vargas-Isla R, Cabral TS, Rodrigues DP, Ishikawa NK (2019) Progress on the phylogeny of the Omphalotaceae: Gymnopus s. str., Marasmiellus s. str., Paragymnopus gen. nov. and Pusillomyces gen. nov. Mycol Prog 18(5):713–739. https://doi.org/10.1007/s11557-019-01483-5
Piercey-Normore MD, Egger KN, Berube JA (1998) Molecular phylogeny and evolutionary divergence of North American biological species of Armillaria. Mol Phylogenet Evol 10(1):49–66. https://doi.org/10.1006/mpev.1997.0485
Purtov KV, Petushkov VN, Baranov MS, Mineev KS, Rodionova NS, Kaskova ZM, Tsarkova AS, Petunin AI, Bondar VS, Rodicheva EK, Medvedeva SE, Oba Y, Oba Y, Arseniev AS, Lukyanov S, Gitelson JI, Yampolsky IV (2015) The chemical basis of fungal bioluminescence. Angew Chem Int Ed Engl 54(28):8124–8128. https://doi.org/10.1002/anie.201501779
Purtov KV, Petushkov VN, Rodionova NS, Gitelson JI (2017) Why does the bioluminescent fungus Armillaria mellea have luminous mycelium but nonluminous fruiting body? Dokl Biochem Biophys 474(1):217–219. https://doi.org/10.1134/S1607672917030176
Puzyr AP, Medvedeva SE, Bondar VS (2017) Biochemical changes causes lack of bioluminescence in fruiting bodies of Armillaria. Mycosphere 8(1):9–17. https://doi.org/10.5943/mycosphere/8/1/2
Quesada-Chanto A, Jimenez-Ulate F (1996) Short communication: in vitro evaluation of a Bacillus sp. for the biological control of the coffee phytopathogen Mycena citricolor. World J Microbiol Biotechnol 12(1):97–98. https://doi.org/10.1007/BF00327810
Rao DV, Tewari JP (1987) Production of oxalic acid by Mycena citricolor, causal agent of the American leaf spot of coffee. Phytopathology 77(6):780–785. https://doi.org/10.1094/Phyto-77-780. Index Fungorum 14: 1–1 (2012)
Redhead SA, Moncalvo JM, Vilgalys R, Desjardin DE, Perry BA (2012) Index Fungorum 14:1–1
Rishbeth J (1986) Some characteristics of English Armillaria species in culture. T Brit Mycol Soc 86:213–218. https://doi.org/10.1016/S0007-1536(86)80147-4
Saccardo PA (1887) Sylloge Hymenomycetum, Vol. I. Agaricineae. Sylloge Fungorum 5:1–1146
Sekizaki H, Kuninaga S, Yamamoto M, Asazu SN, Sawa S, Kojoma M, Yokosawa R, Yoshida N (2008) Identification of Armillaria nabsnona in gastrodia tubers. Biol Pharm Bull 31(7):1410–1414. https://doi.org/10.1248/bpb.31.1410
Shaw CG, Kile GA, United States. Forest Service (1991) Armillaria root disease. Agriculture handbook, vol 691. Forest Service, U.S. Dept. of Agriculture, Washington, DC
Shih YS, Chen CY, Lin WW, Kao HW (2014) Mycena kentingensis, a new species of luminous mushroom in Taiwan, with reference to its culture method. Mycol Prog 13(2):429–435. https://doi.org/10.1007/s11557-013-0939-x
Singer R (1947) New genera of Fungi.3. Mycologia 39(1):77–89. https://doi.org/10.2307/3755289
Sipos G, Anderson JB, Nagy LG (2018) Armillaria. Curr Biol 28(7):R297–R298. https://doi.org/10.1016/j.cub.2018.01.026
Smith ML, Bruhn JN, Anderson JB (1992) The fungus Armillaria-Bulbosa is among the largest and oldest living organisms. Nature 356(6368):428–431. https://doi.org/10.1038/356428a0
Terashima Y, Takahashi H, Taneyama Y (2016) The fungal flora in southwestern Japan: agarics and boletes. Tokai University Press, Japan
Treu R, Agerer R (1990) Culture characteristics of some Mycena species. Mycotaxon 38:279–309
Ulloa M, Hanlin RT (2017) Illustrated dictionary of mycology, 2nd edn. APS Press, St. Paul, MN
Wassink EC (1948) Observations on the luminescence in fungi, 1, including a critical review of the species mentioned as luminescent in literature. Recueil des travaux botaniques néerlandais 41(1):150–212
Wassink EC (1978) Luminescence in fungi. In: Herring PJ (ed) Bioluminescence in action. Academic Press, London, pp 171–197
Wassink EC (1979) On fungus luminescence. Meded Landbouwhogeschool Wageningen 79 (5)
Weinstein P, Delean S, Wood T, Austin AD (2016) Bioluminescence in the ghost fungus Omphalotus nidiformis does not attract potential spore dispersing insects. IMA Fungus 7(2):229–234. https://doi.org/10.5598/imafungus.2016.07.02.01
Widder EA (2010) Bioluminescence in the ocean: origins of biological, chemical, and ecological diversity. Science 328(5979):704–708. https://doi.org/10.1126/science.1174269
Zang M (1979) Some new species of higher fungi from Xizang (Tibet) of China. Acta Bot Yunnanica 1(2):101–105
Zhang L, Chen J, Lv Y, Gao C, Guo S (2012) Mycena sp., a mycorrhizal fungus of the orchid Dendrobium officinale. Mycol Prog 11:395–401. https://doi.org/10.1007/s11557-011-0754-1
Acknowledgment
We thank Yu-Shen Shih and Hua-Te Fang for providing photos of bioluminescent fungi, and Eiji Hadano and Atsuko Hadano for specimen collection to validate the phylogenetic position of Marasmiellus lucidus and M. venosus. I.J.T was supported by Career Development Award AS-CDA-107-L01, Academia Sinica, Taiwan, and the Ministry of Science and Technology, Taiwan under Grant 110-2628-B-001-027.
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this chapter
Cite this chapter
Ke, HM. et al. (2023). Evolution and Diversity of Bioluminescent Fungi. In: Pöggeler, S., James, T. (eds) Evolution of Fungi and Fungal-Like Organisms. The Mycota, vol 14. Springer, Cham. https://doi.org/10.1007/978-3-031-29199-9_12
Download citation
DOI: https://doi.org/10.1007/978-3-031-29199-9_12
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-29198-2
Online ISBN: 978-3-031-29199-9
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)