Abstract
A non-invasive technique for skin cancer diagnostics that reliably classifies lesions as malignant or benign is analyzed and preferred using machine-learning and deep-learning algorithms. The different stages of diagnostics involve using machine learning: a collection of data images, filtering the images to remove unwanted details and noise, segmenting the images using various clustering algorithms. Feature extraction methods have been used to accomplish classification. Five distinctive classifiers have been trained and their efficiency has been compared. K-nearest neighbor, support vector machine, decision trees, multi-layer perceptron, and random forest are used to classify the skin lesion as malignant or benign. An effective comparison of two different deep-learning architectures, such as AlexNet and GoogLeNet, has been carried out. The dataset, which contains 900 images, is subjected to various identification techniques. The accuracy, F-measure precision, and recall were used to evaluate the effectiveness of the classification scheme. As a result of the findings, when compared with the random forest classifier, AlexNet has a high accuracy of 95%. The number of training samples seems to have a serious influence on the ability of deep-learning strategies. Although having a small number of training samples, the presented scheme was able to precisely discriminate among healthy and diseased lesions. Hence, the proposed method will enhance the effectiveness of early detection for skin cancer and could be used in computer-assisted systems to help dermatologists discover cancerous lesions.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keyword
1 Introduction
Machine learning (ML) and artificial intelligence (AI) are quickly developing fields, particularly adversely influencing numerous conventional organizations and enterprises, and offer to rebuild numerous parts of day-to-day existence. Such rebuilding will be especially helpful in medicine, where life or death choices could be altogether further developed utilizing information and calculations. High-level clinical image examination is progressively fundamental in the visualization, treatment, and analytical assessment of illness. A perspective of machine-learning and deep-learning algorithms is extended to investigate and prefer a non-invasive technique for skin cancer diagnosis that accurately classifies the lesions as malignant or benign melanoma.
Earlier recognition of skin malignancy is critical. Skin cancer is now considered to be a major hazardous form of cancer observed in humans. One of the biggest causes of skin cancer is the sun’s ultraviolet (UV) emission. Continuous exposure to sun can affect ageing and pave the way for cancer development. The sun’s UV light may damage the elastin fibers present in the skin, and when these fibers break down they continue to sag and stretch and finally lose the ability to get back to the original place [1]. Skin malignancy develops when the melanocytes mutate and become cancerous. Skin malignancy is commonly categorized as malignant or benign melanoma. When a group of melanocytes gather together and form a lesion, owing to elevated concentration of melanin, a brown pigmented patch appears on the skin. These melanocytic lesions may consist of cells that are benign or malignant. It is possible to divide non-melanocytic lesions into benign and malignant neoplasms. Seborrheic keratoses, vascular lesions, and dermatofibroma are examples of the former. The malignant neoplasm is termed basal cell carcinoma (BCC). It is the prevalent type of fatal skin disease, but owing to its slow growth, it is regarded as less hazardous than melanoma [2].
Melanoma is an assortment of melanocytic injuries that is dangerous. This injury progresses more quickly than BCC, profoundly fit for attacking tissues and metastasizing to different organs. The deadliest type of skin malignant growth is one of these melanomas [3]. Recuperating can be effective when melanoma malignant growth is recognized at the beginning phase. One of the strategies utilized by dermatologists to analyze melanomas is an imaging strategy called dermoscopy, where an amplification apparatus and a light source are utilized to review the skin injury. This enables the dermatologist to detect subcutaneous patterns that would require extensive preparation to be undetectable [4]. Furthermore, the determination is abstract and often difficult to imitate. Hence, programmed strategies should be created to help dermatologists give a more exact conclusion. Clinical image determination can be successfully performed utilizing Personal Computer vision. A computer-based demonstrative framework for the skin image has significant screening and disease-finding potential. Improvement in the determination of the progress of melanoma is accomplished utilizing computer-based object recognition system. As a visual framework frequently causes fault, the requirement for better accuracy and second opinions is featured. On the other hand, it decreases a doctor’s assignments and obligations. Many investigations in the programmed recognition of melanoma have been created. The imminent advantages of such examinations are significant and immense. In addition, the interdependence of difficulties is high, and the new contributions in the area are highly valued. Then again, it is generally perceived that better precision is expected by the more certain and proficient identification frameworks [5].
2 Analogous Performance
To improve the computational capability of standard ABCD assessment, a computer-assisted diagnostic system is adopted. Melanin production and surface (photodynamic therapy [PDT]) qualities are characterized by features gained from local investigation of lesion intensity. The findings demonstrate that PDT structures are hopeful qualities that, when combined with standard ABCD features, can increase the detection efficiency of pigmented skin lesions [6].
A Boltzman Entropy novel technique is employed for categorizing carcinogenic and noncarcinogenic skin lesions. DullRazor performs hair removal, whereas lesion texture and color information are used to enhance lesion contrast. A hybrid method is introduced in lesion segmentation and outcomes are combined using the addition law of probability. Subsequently, the serial-based technique is implemented to extract and fuse attributes such as color, texture, and histogram of oriented gradients (shape). The merged attributes are then chosen using a novel Boltzmann entropy technique. Last, support vector machine (SVM) classifies the chosen features. Compared with current techniques, the suggested method detects and classifies melanoma relatively well [7].
A multi parameter artificial neural networks on basis of manageable personal health info with elevated sensitivity and specificity for early identification of nonmelanoma skin cancer, even in the lack of known exposure to UV rays was generated [7].
A further approach had two phases: an initial step used a kernel- and region-based convolutional strategy to consistently crop the particular object on dermatological imaging, and the next segment used the ResNet152 framework to discriminate potentially cancerous abnormalities. The efficacy of the categorization methodology has been enhanced [8].
A deep convolutional neural network (CNN) based on a deep-learning strategy is also employed for appropriate identification of normal and infected dermatitis. The deep CNN paradigm is tested with transfer learning approaches such as AlexNet, DenseNet, MobileNet, ResNet, and VGG-16 to determine overall effectiveness. The eventual findings of the current deep CNN model are described as being much more effective than authorization learning techniques [9].
Furthermore, a CNN with a dynamic GoogLeNet topology is constructed. The eight performance indicators assessed were polygon region, kappa, categorization efficiency, sensitivity, F-score measurement, specificity, area under the curve, and time complexity. According to the observations, the generated CNN had the best calculation efficiency with the least amount of time to accomplish the assignment [10].
3 Evaluation of Skin Malignancy Using Machine-Learning Methodologies
In the development of computer-based detection methods for melanoma diagnosis, different classification algorithms were used. Whether one technique outperforms the other, however, is not evident. As there are robust and fragile points in each category process, selecting only one method to carry out all comparisons of features and descriptors is not simple. Therefore, five distinct algorithms were implemented in this work. An appropriate classification scheme for melanoma images is developed using methods of machine learning to characterize skin lesions as harmless or cancerous. Figure 1 uses machine-learning methods to demonstrate the flow chart of the classification of skin lesions.
3.1 Anisotropic Diffusion Filtering
Dermoscopic images usually contain some artifacts. Powerful approaches to eliminate artifacts and enhance the appearance of the initial images are therefore required. The basic motive behind this pre-processing is to improve melanoma image quality by evacuating irrelevant portions and noise for further processing in the background of an image. Using 2D anisotropic diffusion filter, noise and artifacts were removed at the original point [11]. ADF method was applied to minimize image noises, assuring essential elements of image detail, generally borders as well as outlines / equivalent points are not disturbed from image view. On three channels (red green blue [RGB]) the anisotropic filters were implemented individually. Unsharp masking was implemented on an entire image only after denoising. The image was sharpened using gray world normalization. After that, color constancy was implemented on the three channels together. Hairs behave as an ambiguity on dermoscopic images. Gray world normalization is used to identify the hair. An inpainting technique was used to separate the identified hairs. Figure 2 shows the results of the steps used for pre-processing.
3.2 Melanoma Segmentation Analysis
Numerous segmentation forms of algorithms such as Otsu’s threshold, k, fuzzy c, and adaptive k-means were used for the segmentation of melanoma. Maximizing interclass variability and minimizing intraclass variability is performed using Otsu’s thresholding method. A threshold limit is fixed and the value above the limit is regarded in the forefront and the value under the limit is taken in the background [12].
The variance of the inside class is described in Eq. 1 as:
Only the centroid defines all cluster. The closest centroid classifies each pixel. In k-means clustering [13], there were two clusters (Eq. 2):
The centroid needs an update under each iteration’s end, wherein the succeeding equation is used to update the centroid. If the value does not change further, the iteration stops (Eq. 3):
Fuzzy c means algorithm functions through assigning each pixel to the segment. The comparison depends on the distance of particular pixel from multiple clusters. The Euclidean division between two points states that the correlated condition that can characterize i and j in Eq. 4 is
There are two clusters: one cluster denotes the foreground whereas the background is denoted by the other one, m indicates the fuzziness factor, μ(i, j) represent the membership variable, d(i, j) is Euclidean distance within ith data and the center of jth form of the data set. The outcome produced showcases the ground truth provided. The Dice similarity index (DSI) facilitates determination of image segmentation accuracy (Eq. 5):
To quantitatively assess performance of the segmentation method, the work also utilizes the Dice similarity coefficient. All targeted areas are effectively segmented using the above-mentioned segmentation techniques. The focus of this procedure is to evaluate the execution of segmentation with radiotherapy conveyance control of the distinct techniques for treating the targeted region. Abdel and Allan [14] provided analysis parameters on the basis of a unique class pertaining to region from the calculated DSI shown (Table 1).
In segmenting lesions, the k-means and Otsu’s Dice coefficients appeared lower than FCM and adaptive k-means coefficients. Findings (Table 1) indicated that the Dice coefficient of adaptive k-means appeared significantly high and much more appropriate for region separation of images. Figure 3 represents the outcomes of the different segmentation processes.
3.3 Feature Extraction
To categorize the images, feature extraction techniques are used to obtain features. Three elements of structure are obtained from binary differentiated images: irregularity, shape, and circularity signal.
Equation 6 shows how to calculate the irregularity:
where BI is the binary image. The fast Fourier transformed the shape signal and split it into ten rays. Each ray was considered an element. There were 13 shape elements in all. Binary object circularity is calculated in Eq. 7:
Texture-derived attributes were obtained through three distinct channels (R, G, and B) from segmented images. Using mean and standard deviation the first-order statistics of an image may be acquired. These are associated with separate pixel characteristics. Second-order image statistics obtained via the gray-level co-occurrence matrix (GLCM) accounting for spatial interdependence of two pixels at particular relative places. Contrast, correlation, power, homogeneity and entropy were five Haralick attributes acquired from the GLCM. The following formula is used to measure average (Eq. 8) and standard deviation (SD) (Eq. 9):
Ten local binary pattern features were also calculated [15].
3.4 Benign and Malignant Classification
Classifiers have been trained via obtained attributes. Five distinctive classifiers have been learned and their precision has been compared: k-nearest neighbor (k-NN), support vector machine (SVM), decision tree (DT), multi-layer perceptron (MLP), and random forest (RF) [16]. The condition of all classifiers has been enhanced by ten-fold cross-validation. Of the total images, 60% were used as training samples and the testing set utilized the remaining 40%.
3.5 K-Nearest Neighbor
This computation depends on a pseudo-parametric identification methodology. The output is determined as the category with the maximum malignancy from the k-most comparative events at the stage where k-NN is used for interpretation. The value of k has been maintained as five. The melanoma that is categorized as harmless or cancerous will be identified as the primary vote it gets from its nearest neighbor.
3.6 Support Vector Machine
It is selective. With labeled learning information being supplied, a hyperplane is drawn that chooses the boundaries of selection. To categorize images using SVM, the hyperplane separates item sets with completely unpredicted forms of memebership. The analysis of the hyperplane classifies the images as cancerous and non-cancerous.
3.7 Decision Tree
This classifier supports the algorithmic principle of supervised learning. The goal of using DT is to produce a training model that is used by learning data to predict category or estimate target variables by learning choice rules. By using tree delineation, the DT resolves the problem. The internal node of each tree is comparable with a quality. Each leaf node is associated with a category tag. In decision tree, it is typical to start at the base of the tree, predict a class label, and examine the root features with actual data. During examination, the algorithm compares the branch to successive nodes and moves forward. Once it reaches the leaf node of the expected class, the algorithm classifies as harmless (benign)/cancerous (malignant).
3.8 Multilayer Perceptron
This classifier relies on a neural mechanism (feed forward) made up of three layers. Each layer is entirely connected to the layers above in the system. The primary is the layer of input, the hidden level represents the second, and the tertiary is the yield layer. The input data are represented by nodes within the primary layer. All distinct node points of input layer are processed by using linear input mixture with node w weights linked to bias b and using activation function. It could be formed with K + 1 layers (Eq. 10) in a network frame for the MLP classifier as needed. The sigmoid operator is used by nodes in hidden layers (Eq. 11).
Nodes in the yield layer use the softmax function (Eq. 12):
To train MLP, the back propagation method is utilized. The number of neural network nodes equivalent to number of categories in the yield layer.
3.9 Random Forest
This creates a DT group from an arbitrarily selected sub-set of the training set. It then summarizes the votes from various trees of selection to settle on the test object’s ultimate category. It is made up of the number of DTs. There were 100 trees in this analysis. The principal distinction between DT and RF is that the single tree is represented by DT, whereas RF consists of multiple trees [17].
Receiver-operating characteristics (ROC) curve indicates sensitivity/specificity for testing to evaluate the consistency of five classifiers. The ROC curve is nothing but the true-positive (TP) rate and the false-positive (FP) rate relation. TP, FP, false negative (FN), and true negative (TN) are the four parameters that are utilized to figure out the accuracy, sensitivity, and specificity of the classifiers. The positive qualities effectively estimated by the model define the true-positive rate, and the false-positive rate is positively misidentified by negative attributes. The corresponding condition measured the accuracy of the different computational models (Eq. 13), their sensitivity (Eq. 14), and their specificity (Eq. 15):
Figure 4 revealed the effects of classification using the ROC curve of five distinct classifiers.
The accuracy of different classifiers is mentioned in Table 2. The general accuracy of RF can be obviously noted to be the highest. The confusion matrix of five classifiers is depicted in Table 3, for DT, out of 900 (458 benign and 269 malignant) 727 are properly classified and 173 misclassified (benign), 99 are categorized as malignant and 74 as benign. For k-NN, out of 900, 727 are properly categorized (420 benign and 307 malignant) and 173 are found to be misclassified, 71 as malignant and 102 as benign. For MLP, 727 out of 900 are correctly classified (393 benign and 334 malignant) and 173 are misidentified, 54 as malignant and 118 as benign. For SVM, 727 out of 900 are properly classified (411 benign and 316 malignant) and 173 misclassified, 47 as malignant and 126 as benign. For RF, 727 are properly categorized (613 benign and 114 malignant) and 173 misclassified, 121 as malignant and 52 as benign.
The average calculation time for pre-processing to classification was found to be 2.043 ± 0.122 min. Overall computation interval in 20 images is graphically denoted in Fig. 5.
Table 4 demonstrates that the highest level of learning and test efficiency is generated by RF. A cross-validity score of 93.47% was estimated for RF.
3.10 Summary of Melanoma Classification Using Machine Learning
An effective melanoma image classification scheme has been developed to classify a noncancerous (benign) form and a similarly cancerous (malignant) type of lesion. Different segmentation algorithms employed over 900 dataset images. The DSI was used to validate the segmentation technique, and adaptive k-means clustering outperformed the other clustering algorithms in terms of precision. The estimation of the efficiency of the five classifiers is determined. The best of five classifiers is assessed on the basis of precision, specificity, and sensitivity. The ROC plot is used for further analysis. From the observational outcome, the precision of the classifier is 93%, 86.9%, 75%, 71.5%, and 69% respectively for RF, DT, MLP, k-NN, and SVM. From this it could be surmised that the classifier with the greatest accuracy is RF. Thus, it served as an effective classifier for the detection of benign/malignant forms of skin lesions.
4 Deep-Learning Approaches to Skin Cancer Diagnosis
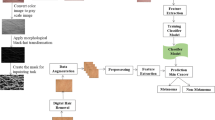
Deep-learning strategies are now employed to categorize harmless and cancerous lesions [18]. Using a similar sample, transfer learning techniques such as AlexNet are being used to assess effectiveness. The layout of the intended work is presented in Fig. 6 as a schematic drawing.
4.1 Image Enhancement
Many strategies exist for downsizing, hiding, filtering, hair elimination, and converting RGB shading to gray resolution images. They are implemented to greatly reduce noise and reflective aberrations. The median window is used to de-clutter the image, disguise the undesirable traits, and eradicate them. It is frequently employed to remove the error without diminishing the image quality, thereby improving the image clarity [19].
4.2 Augmentation of Images
Augmentation is a technique for increasing the volume of data without generating new data by introducing slightly altered imagery into old training samples. The training sample number could be considerably increased, or the system could be protected against overfitting, through oversampling. To minimize overfitting, augmentation parameters such as rotation, shear, zoom, channel shift, height shift, and width shift are applied [20].
4.3 AlexNet Topology
Krizhevsky designed AlexNet, which uses the ReLu function. AlexNet provides multi-general processing unit (GPU) learning, in which half of a net neuron is handled on one GPU whereas the remaining neurons are processed on the other. AlexNet is composed of eight layers: five convolutional layers with a combination of max-pooling layers, and three fully linked layers [21]. This primarily enables larger-scale training, thereby also reducing the training process [22].
4.4 Experimental Findings
The effectiveness of skin cancer screening is improved by employing deep neural networks. Melanoma malignancy is diagnosed through images from the International Skin Imaging Collaboration (ISIC) repository dataset. Initially, the image is loaded and normalized. It is processed via image augmentation, and the architecture and layers of the network are constructed. The CNN uses AlexNet [23, 24]. The system is then trained using supervised learning after the loss function of the dataset is created. During testing and training, the data are equally divided. Finally, the validation is performed by computing accuracy (Eq. 16), F-measure (Eq. 17) and recall (Eq. 18):
The deep neural module in MATLAB R2020b is employed to construct and validate the network. The dataset aggregation is categorized into two major groups: 80% data trained and 20% data utilized for testing. The learning rate is set at 0.0001 and the number of epochs is limited to six. In elements of accuracy, F-measure, precision, and recall, the relevant formulas are utilized to analyze and evaluate the results of the network procedure.
The ISIC dataset (http://www.isic-archive.com) has been used to collect 900 pictures (600 benign and 300 malignant) for this proposed assessment [25]. Eighty percent of the lesions in each category were selected at random and utilized as training examples, whereas the leftover data have been used as a testing set. Both malignant and benign presentations are displayed in Fig. 7. The use of AlexNet to characterize benign and diseased lesions is a high priority of our conceptual framework.
The confusion matrix of AlexNet is given in Table 5. Table 6 depicts its performance when examining quantitative metrics such as accuracy, F-measure, precision, and recall evaluation outcomes. The efficiency of the AlexNet framework training and testing processes is depicted in Fig. 8.
To validate the efficacy of the AlexNet architecture, F-measure, precision, accuracy, and recall parameters are estimated. Specific factors such as TN, FP, TP, and FN were employed to compute the performance of the AlexNet system [26]. The TP factor refers to the percentage of positive traits correctly identified by the system, whereas the FP score refers to the percentage of negative traits misappropriated as positive.
Table 5 indicates that AlexNet correctly categorized 855 images out of 900 datasets, whereas 45 were inaccurately categorized (295 malignant and 560 benign). Table 5 shows the quantifiable parameters used by AlexNet. AlexNet is shown to have a 95% accuracy level. As an outcome, AlexNet may be used by specialists to categorize dermoscopy images and generate appropriate predictions.
As a result, larger sample sources are set to increase the significance of the findings. The approach can be implemented in a clinician’s computer-assisted sensing devices to aid in the identification of skin malignancy. It can also be applied to images of lesions taken from patients and delivered on handheld devices. It therefore allows a quick cancer diagnosis, which dramatically streamlines therapy and improves chances of recovery.
5 Conclusion
The significant impacts of work in this field are summed up concerning portions of the framework, potential strategies, intercessions, and insightful results. A viewpoint on machine learning and deep learning is described in the above review to propel a skin injury acknowledgment technique for characterization on dermatoscopic images of threatening and harmless lesions. A thorough examination data set is produced by gathering dermoscopic images from different chroniclers such as the International Society for Digital Imaging of the Skin and ISIC. To empower similar examinations on dermoscopic image division and characterization calculation for research and benchmarking purposes, the PH2 dataset has been made. The main attribute of the examination work is that around 900 dermoscopic image tests are chosen for the exploratory work. Thus, the handling speed is essentially expanded. A systematic evaluation was successfully carried out between different machine-learning techniques, such as DT, MLP, SVM, k-NN, RF, and deep-learning techniques such as AlexNet. The experimental results illustrate the importance and main achievements of this work, which has an estimated classification accuracy of 93% for the RF model and 95% for the AlexNet model. Therefore, the deep-learning system shows an automated diagnostic technique for constant and accurate determination of skin malignancy with an extraordinary ability to carry out treatment strategies using non-invasive methods.
References
Sriram RD, Subrahmanian E. Transforming health care through digital revolutions. J Indian Inst Sci. 2020;100(4):753–72.
Siegel RL, Miller KD, Jemal A. Cancer statistics, 2019. CA Cancer J Clin. 2019;69(1):7–34.
Xu YG, Aylward JL, Swanson AM, Spiegelman VS, Vanness ER, Teng JM, Snow SN, Wood GS. Nonmelanoma skin cancers: basal cell and squamous cell carcinomas. In: Abeloff’s clinical oncology. Philadelphia: Elsevier; 2020. p. 1052–73.
Gershenwald JE, Scolyer RA, Hess KR, Sondak VK, Long GV, Ross MI, Lazar AJ, Faries MB, Kirkwood JM, McArthur GA, Haydu LE. Melanoma staging: evidence-based changes in the American Joint Committee on Cancer eighth edition cancer staging manual. CA Cancer J Clin. 2017;67(6):472–92.
Han SS, Kim MS, Lim W, Park GH, Park I, Chang SE. Classification of the clinical images for benign and malignant cutaneous tumors using a deep learning algorithm. J Investig Dermatol. 2018;138(7):1529–38.
Aljanabi M, Özok YE, Rahebi J, Abdullah AS. Skin lesion segmentation method for dermoscopy images using artificial bee colony algorithm. Symmetry. 2018;10(8):347.
Zakeri A, Hokmabadi A. Improvement in the diagnosis of melanoma and dysplastic lesions by introducing ABCD-PDT features and a hybrid classifier. Biocybern Biomed Eng. 2018;38(3):456–66.
Nasir M, Attique Khan M, Sharif M, Lali IU, Saba T, Iqbal T. An improved strategy for skin lesion detection and classification using uniform segmentation and feature selection based approach. Microsc Res Tech. 2018;81(6):528–43.
Roffman D, Hart G, Girardi M, Ko CJ, Deng J. Predicting non-melanoma skin cancer via a multi-parameterized artificial neural network. Sci Rep. 2018;8(1):1–7.
Jojoa Acosta MF, Caballero Tovar LY, Garcia-Zapirain MB, Percybrooks WS. Melanoma diagnosis using deep learning techniques on dermatoscopic images. BMC Med Imaging. 2021;21(1):1–1.
Ali MS, Miah MS, Haque J, Rahman MM, Islam MK. An enhanced technique of skin cancer classification using deep convolutional neural network with transfer learning models. Mach Learn Appl. 2021;5:100036.
Yilmaz ER, Trocan MA. A modified version of GoogLeNet for melanoma diagnosis. J Inf Telecommun. 2021;5(3):395–405.
Meskini E, Helfroush MS, Kazemi K, Sepaskhah M. A new algorithm for skin lesion border detection in dermoscopy images. J Biomed Phys Eng. 2018;8(1):117.
Al Mamun M, Uddin MS. A comparative study among segmentation techniques for skin disease detection systems. In: Proceedings of international conference on trends in computational and cognitive engineering. Singapore: Springer; 2021. p. 155–67.
Anas M, Gupta K, Ahmad S. Skin cancer classification using K-means clustering. Int J Tech Res Appl. 2017;5(1):62–5.
Taha AA, Hanbury A. Metrics for evaluating 3D medical image segmentation: analysis, selection, and tool. BMC Med Imaging. 2015;15(1):1–28.
Guo Z, Zhang L, Zhang D. A completed modeling of local binary pattern operator for texture classification. IEEE Trans Image Process. 2010;19(6):1657–63.
Ozkan IA, Koklu M. Skin lesion classification using machine learning algorithms. Int J Intell Syst Appl Eng. 2017;5(4):285–9.
Amutha S, Devi RM. Early detection of malignant melanoma using random forest algorithm. Int J Trends Eng Technol. 2016;13(1):1–14
Esteva A, Kuprel B, Novoa RA, Ko J, Swetter SM, Blau HM, Thrun S. Dermatologist-level classification of skin cancer with deep neural networks. Nature. 2017;542(7639):115–8.
Roslin SE. Classification of melanoma from dermoscopic data using machine learning techniques. Multimed Tools Appl. 2020;79(5):3713–28.
Shorten C, Khoshgoftaar TM. A survey on image data augmentation for deep learning. J Big Data. 2019;6(1):1–48.
Li Y, Shen L. Skin lesion analysis towards melanoma detection using deep learning network. Sensors. 2018;18(2):556.
Dildar M, Akram S, Irfan M, Khan HU, Ramzan M, Mahmood AR, Alsaiari SA, Saeed AH, Alraddadi MO, Mahnashi MH. Skin cancer detection: a review using deep learning techniques. Int J Environ Res Public Health. 2021;18(10):5479.
Nasiri S, Helsper J, Jung M, Fathi M. DePicT Melanoma Deep-CLASS: a deep convolutional neural networks approach to classify skin lesion images. BMC Bioinformatics. 2020;21(2):1–3.
Favole F, Trocan M, Yilmaz E. Melanoma detection using deep learning. In: International conference on computational collective intelligence. Cham: Springer; 2020. p. 816–24.
Gautam D, Ahmed M, Meena YK, Ul HA. Machine learning-based diagnosis of melanoma using macro images. Int J Numer Method Biomed Eng. 2017;34(5):1–8.
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this chapter
Cite this chapter
Bethanney Janney, J., Divakaran, S., Sudhakar, T., Grace Kanmani, P., Hemalatha, R.J., Nag, M. (2023). Emerging Soft Computation Tools for Skin Cancer Diagnostics. In: Ram Kumar, C., Karthik, S. (eds) Translating Healthcare Through Intelligent Computational Methods. EAI/Springer Innovations in Communication and Computing. Springer, Cham. https://doi.org/10.1007/978-3-031-27700-9_16
Download citation
DOI: https://doi.org/10.1007/978-3-031-27700-9_16
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-27699-6
Online ISBN: 978-3-031-27700-9
eBook Packages: EngineeringEngineering (R0)