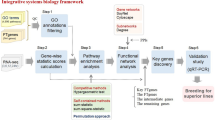
Abstract
Aluminum toxic stress in soybean (Glycine max L.) is a major abiotic stress that severely affects crop production on acidic soils. Understanding the molecular mechanisms of aluminum stress response as well as identifying the molecular targets are of paramount important in soybean molecular breeding. Thus, the utility of network biology and machine learning approaches were demonstrated on gene expression data to understand aluminum stress response mechanisms in soybean. Further, the major focus of the chapter is to demonstrate, first, the use of machine learning techniques to select informative genes for aluminum stress in soybean using high-dimensional gene expression data; second, the use of network biology approach to identify various gene modules responsible for aluminum stress tolerance; and third, the identification of hub and unique hub genes in constructed gene networks done through novel statistical approach. Moreover, the molecular characterization of various identified genes, hubs, and unique hubs revealed the underlying molecular mechanisms of aluminum toxic stress response in soybean. These identified genes can be used as molecular targets for aluminum stress response engineering in soybean.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
5.1 Introduction
Aluminum (Al) toxic stress is a major impediment to crop production on acidic soils that affects about 30–40% of the world’s arable lands (Kochian et al. 2004). Soybean (Glycine max L.), which provides major source of proteins, unsaturated fats, carbohydrates, and fibers, is one of the most important legume crops, capable of providing nutritional security to the global population. Soybean is preferably grown on acidic soil, and its productivity is significantly reduced by Al toxic stress. In acidic soil, Al stress causes rapid inhibition in root growth and subsequently inhibits water and nutrient uptake by plants. This increases the susceptibility of plants to other environmental stresses and results in reduction of crop productivity (Ma 2007). Under heavy pressure of population explosion and global warming, achieving nutritional security in general and protein security in particular through enhancing the productivity of soybean is of paramount importance. However, the underlying mechanisms for Al toxic stress response in plants in general and soybean in particular are not so clearly deciphered till now (Zeng et al. 2012).
Microarray data are used for gene selection and module detection in genetic network analysis, which suffers from the inherent limitation of its high-dimensionality, i.e., the number of genes is much larger than the number of subjects/samples (Guyon et al. 2002). Therefore, it is important to select most relevant genes related to stresses/conditions from thousand(s) of genes with the help of appropriate computational/machine learning approach(s). In this regard, volcano plot method (Cui and Churchill 2003) is quite popular among the researchers in which genes are selected by considering their relevance with their classes. However, such method may not be sufficient to discover some complex relationships among genes for a certain trait or condition (Liang et al. 2011). Besides, several statistical and machine learning methods, viz., t-score, F-score, information gain (IG) measure, random forest (RF), and support vector machine-recursive feature elimination (SVM-RFE) (Mao et al. 2006; Forman 2003; Díaz-Uriarte and de Andrés 2006; Lai et al. 2011; Guoyon and Elisseeff 2003), have also been used for gene selection. However, in these methods, genes are selected by considering only their relevance with classes. In such case, there is a possibility that genes which are spuriously associated with the classes may also get selected.
In order to understand the interrelationship among the selected genes, identification of gene modules and key genes responsible for a particular stress/condition and analysis of gene co-expression networks need to be carried out. Weighted gene co-expression network analysis (WGCNA) (Zhang and Horvath 2005) is a latest and popular technique used to decipher co-expression patterns among genes. The WGCNA approach typically deals with the identification of gene modules by using the gene expression levels that are highly correlated across samples (Zhang and Horvath 2005). This technique has been successfully utilized to detect gene modules in Arabidopsis, rice, maize, and poplar for various biotic and abiotic stresses (Childs et al. 2011; Downs et al. 2013; Zhang et al. 2012; Ficklin et al. 2010). Further, this approach also leads to the construction of gene co-expression network (GCN), a scale-free network, where genes are represented as nodes and edges depict associations among genes (Zhang and Horvath, 2005; Langfelder and Horvath 2008). In such network, highly connected genes are called hub genes, which are expected to play an important role in understanding the biological mechanism of response under stresses/conditions (Das et al. 2017; Barabasi and Oltvai 2004; Stumpf and Porter 2012). Identification of hub genes will also help in mitigating the stress in plants through genetic engineering. The existing approaches (Barabasi and Oltvai 2004; Stumpf and Porter 2012; Chen et al. 2008) have mainly focused on hub gene identification, based only on gene connection degrees in the GCN. Moreover, these techniques select such genes empirically without any statistical criterion.
In this chapter, a machine learning technique, i.e., Bootstrap SVM-RFE (Boot-SVM-RFE) (Das et al. 2017), is used for selection of informative genes. In this technique, genes are selected after reducing the effect of spurious associations between genes and classes, and it has better performance on real crop gene expression datasets. Further, a statistical approach for identification of hub genes in the GCN was also used, which found to be superior in terms of scale-free property of biological network (Das et al. 2017). Hub genes responsible for Al toxic stress have been identified, and their functional analysis has been reported.
5.2 Material and Methods
The soybean microarray experimental datasets under Al stress were collected from Gene Expression Omnibus with platform GPL4592 (http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GPL4592). This platform contains 3855 experimental samples on 37,593 probes generated using Affymetrix Soybean Genome Array. Out of these samples, 80 samples related to Al stress were used for further analysis. Initially, raw CEL files of these collected samples were processed using robust multichip average (RMA) algorithm available in affy bioconductor package of R (R Core Team 2015; Gentleman et al. 2004; Gautier et al. 2004). This includes background correction, quantile normalization, and summarization by the median polish approach.
5.2.1 Bootstrap Support Vector Machine-Recursive Feature Elimination Technique (Boot-SVM-RFE)
The Boot-SVM-RFE for selection of informative genes from high-dimensional gene expression dataset was developed by Das et al. (2017). In this approach, a nonparametric (NP) hypothesis testing procedure was used for the identification of informative genes based on their statistical significance. The details of Boot-SVM-RFE can be found in Das et al. (2017).
5.2.2 Gene Co-expression Network Analysis
GCNs were constructed by using gene co-expression measure that depicts association among genes (Zhang and Horvath 2005; Langfelder and Horvath 2008; Das et al. 2017). Let xi be the expression profile of ith gene, i.e., the expression values of ith gene across all the microarray samples. Then, gene co-expression similarity measure sij between ith and jth gene is computed as the absolute value of Pearson’s correlation coefficient (PCC) (Zhang and Horvath 2005; Das et al. 2017), which is given by:
The adjacency score (aij) between ith gene and jth gene is defined in terms of sij (Langfelder and Horvath 2008) as:
where β (≥1) is soft threshold power, determined by using the concept of scale-free property of biological networks (Zhang and Horvath 2005). The detail methodology for determination of the soft threshold power has been discussed elaborately by Zhang and Horvath (2005). This soft threshold approach leads to a weighted GCN that satisfies the scale-free property of biological networks. For both Al stress and control conditions, the value of β was taken as 8 for calculation of adjacency score, with best approximation to scale-free criteria (Barabasi and Oltvai 2004; Stumpf and Porter 2012) using R2 > 0.80 through fitting of power law model. In order to identify the gene modules (i.e., group of tightly co-expressed genes) within the selected informative genes, the topological overlap matrix was constructed based on the adjacency scores (Zhang and Horvath 2005). The blockwiseModules function available in WGCNA package of R was executed to identify these modules. For this purpose, various parameters like module size, deep split level, and tree merge cut height was set at 20–30, 4, and 0.15–0.25, respectively. In order to find the consensus modules showing co-expression patterns of genes across stress and control conditions, the function blockwiseConsensusModules was used with parameter settings 8, 30, and 0.15 as power, minimum module size, and merge cut height, respectively.
5.2.3 Statistical Approach for Identification of Hub Genes
In network theory, a node is defined as hub node (Das et al. 2017; Barabasi and Oltvai 2004; Stumpf and Porter 2012; Chen et al. 2008) if its connection degree is greater than the average connection degree of the network (Chen et al. 2008). In the existing approach, a gene is declared as hub gene based on an indicator function (Stumpf and Porter 2012; Chen et al. 2008), i.e., Hubi = [I(ki > τ)], and the number of hub genes (NHub) in the genetic network is calculated as \( NHub=\sum \limits_i\left[I\left({k}_i>t\right)\right], \) where Hubi is hub status of ith gene (i.e.,1 or 0), ki is connection degree of ith gene, and τ is a threshold value, i.e., average connection degree of the network. This technique selects hub genes empirically based on only observed gene connectivity without taking into account any statistical consideration. Therefore, an alternate statistical approach based on statistical significance of gene connectivity was developed by Das et al. (2017) for detection of hub genes in the GCN. This statistical approach is briefly produced here and its detail can be found in Das et al. (2017).
The weighted gene score (WGS) for ith gene in terms of weighted gene connectivity (aij) can be written as
where WGSi represents the relative importance of ith gene based on its connections to the remaining genes in GCN. For the purpose of hub gene identification, the following hypotheses are constructed:
-
H0:WGSi ≤ μ, i.e., ith gene in the GCN is not a hub gene
-
H1:WGSi > μ, i.e., ith gene in the GCN is a hub gene
where μ is average connection degree of the complete network model. Here, in order to get the distribution of the test statistic under H0, a resampling procedure was used. In this procedure, m microarray samples were selected randomly with equal probability from M microarray samples to construct one subsample (for one GCN) (m ≤ M). Then statistical measures (Eq. 5.1–5.3) were applied to get WGS for each gene in that GCN. This procedure was repeated large number of times say S to get S sets of WGS. In this study, S = 500 was taken to get 500 random GCNs under stress and control conditions separately. For testing H0 vs. H1, an NP test statistic was used to test the significance of the WGS for each gene, i.e., for testing whether WGS of a gene is greater than the average connection degree of the complete network or not. The algorithm of this approach is given below.
5.2.4 Algorithm
-
Step 1: Begin with all genes (nodes) in the GCN
-
Step 2: Construct a dataset say Tk with m samples randomly taken from M microarray samples
-
Step 3: Calculate WGS for all genes
-
Step 4: Repeat steps 2 and 3 S times to get S sets of WGS for each gene
-
Step 5: Take a particular gene (ith gene) along with its WGS
-
Step 6: Test the hypothesis for ith gene and obtain its p-value
-
Step 7: Repeat steps 5–6 for all genes (i = 1, 2, …, G)
-
Step 8: Rank the p-values and select the hub genes
The hub gene identification approach for the GCNs constructed under two contrasting conditions (stress vs. control) can be called as differential hub gene analysis (DHGA). By this approach, the identification of hub genes is possible in both these GCNs based on statistical test of significance. On the basis of p-values, genes in the GCNs under either condition can be grouped into various groups, viz., housekeeping hub genes (HHG), unique hub genes (UHG) for stress, UHG for control, and non-hub genes based on a decision matrix (Table 5.1).
5.3 Results
5.3.1 Selection of Informative Genes for Al Stress in Soybean
The Boot-SVM-RFE was superior compared to other gene selection techniques (Das et al. 2017), so it was employed to select informative genes for Al stress in soybean. In order to get a robust and minimal set of informative genes, the fold change in volcano plot was replaced with –log10 (p-values) obtained from Boot-SVM-RFE, and then a gene selection plot was constructed. The threshold values for Y- and X-axis of the gene selection plot were fixed as 4 and 2.5, respectively, which lead to the selection of 981 genes as shown in Fig. 5.1.
Gene selection plot for selection of informative genes for Al stress in soybean. The horizontal axis represents negative logarithm of statistical significance values obtained from Boot-SVM-RFE. The vertical axis shows the negative logarithm of statistical significance values from t-test. Green dots indicate selected probes with –log (p-value) from Boot-SVM-RFE ≥ threshold of 2.5 and t-test –log (p-value) ≥ threshold of 4. Red stars indicate the selected probes which have Arabidopsis orthologs. Blue dots indicate unselected probes
The consensus sequences of these 981 genes obtained from GeneChip Soybean Genome Array of Affymetrix were then used to identify the Arabidopsis orthologs (www.affymetrix.com/support), and it was found that 554 genes have unique orthologs in Arabidopsis (Fig. 5.1). Further, the annotations of these selected genes were obtained from SoyBase (http://soybase.org) (Grant et al. 2010).
5.3.2 Functional Analysis of Selected Genes for Al Stress in Soybean
The gene ontology (GO) enrichment analysis of the 981 selected genes was performed by using AgriGO (Du et al. 2010), a plant-specific GO term enrichment analysis tool, and the results are shown in Fig. 5.2a. It is observed that most of the selected genes are responsible for transition metal ion binding, metal ion binding, cation binding, ion binding, etc. (Fig. 5.2a). These molecular functions (MF) might be activated due to the high concentration of Al ions in water or soil. Two other MF, i.e., oxidoreductase (redox) and kinase, activities are also present in these selected genes (Fig. 5.2a). The significant behavior of the genes in redox activity might be related to electron transport in complex chemical reactions that balances the charges during ion transport. The redox activity might also be related to reactive oxygen species (ROS) that are produced in response to oxidative stress due to water deficit during abiotic stress like Al toxic stress (Miller et al. 2008).
Functional enrichment analysis of selected genes and hub genes under Al stress. The GO term enrichment analysis of 981 selected informative genes (a) and hub genes (b) for Al stress condition using Agrigo is shown for different gene ontology categories (CC, MF, and BP). For (a), the GO terms are chosen whose p-values <0. 008 and FDR values (false discovery rate) < 0.6. For (b), the GO terms are chosen whose p-values <0. 1 and FDR values <0.8
In biological process categories, such as cellular nitrogen compound metabolic process, amine metabolic process, cellular amino acid and derivative metabolic process, oxoacid metabolic process, organic acid metabolic process, carboxylic acid metabolic process, cellular ketone metabolic process, and ion transport activity, the number of selected genes is more as compared to other biological processes (Fig. 5.2a). It may be inferred that some of these chosen genes are involved in ion transport activities, i.e., involved in transporting the ions outside the cell to maintain the proper pH in the cell (Wang et al. 2013). In case of cellular components, chosen genes are related to transcription factor complex, cytoplasmic membrane-bounded vesicle, membrane-bounded vesicle, cytoplasmic vesicle, vesicle, and nucleoplasm part (Fig. 5.2a). It can be seen that the maximum number of the genes is related to vesicle and membrane, which is consistent with the detoxifying mechanism of metal ions available in Al stress condition, especially in sequestration by vacuole (Apse et al. 1999; Panda et al. 2009). Some of the selected genes present on membrane are found to be involved in transporting of metal ions outside the cell or to the vacuole to maintain pH and transmembrane proton gradient (Niu et al. 1996).
5.3.3 Gene Co-expression Network Analysis for Al Stress in Soybean
Using WGCNA, the selected 981 genes were divided into 19 and 18 modules (including gray color module, which is the module of the non-modular genes) for Al stress and control conditions, respectively, and the results are shown in Fig. 5.3.
In both cases, module represented by turquoise color contains maximum number of genes and hence designated as the largest module for either condition. Based on the expression profiles of these selected genes for both Al stress and control conditions, 23 consensus modules (set of genes with similar co-expression patterns) were obtained. The matching of various modules for either condition with the consensus modules can be visualized from Fig. 5.3 in terms of their colors. Further, the long length of branch in the dendrogram and high intensity of red color in heat maps showed that the genes belong to the same module have higher degree of co-expression as compared to genes present outside the module. The module memberships (number of genes present) of each module and their underlying molecular functions under Al stress condition are given in Table 5.2. It is observed that every module is significantly annotated with GO terms, except gene modules represented by green-yellow and gray color (Table 5.2). So, it can be inferred that functions of genes present within these two modules are still largely unknown.
5.3.4 Hub Gene Analysis for Al Stress Condition in Soybean
Using hub gene detection statistical approach, 228 and 187 genes were identified as hub genes whose p-values were ≤ 1E-10 for Al stress and control conditions of soybean, respectively (Table 5.3). From the DHGA result, it is seen that 98 hub genes are common, whereas 130 and 89 hub genes are unique for Al stress and control conditions, respectively (Table 5.3). The mapping of the HHG and UHG in soybean to Arabidopsis genome leads to the identification of corresponding Arabidopsis orthologous genes (Fig. 5.4c). The GCNs constructed for these two differential conditions (Al stress vs. control) in soybean along with the positions of hub genes and UHG are shown in Fig. 5.4.
Gene co-expression networks for two differential conditions in soybean. The GCNs are constructed for Al stress (a) and control (b) conditions, respectively. The nodes with red colors represent the housekeeping hub genes, green color nodes represent UHG, and blue color nodes represent the non-hub genes. (c) Venn diagram of hub genes in the GCNs constructed under Al stress (a) and control (b) conditions in soybean. The number of orthologous genes found in Arabidopsis corresponding to unique and common hub genes in soybean is also shown
The functional analysis of the selected hub genes under Al stress revealed their associated cellular mechanisms. From the GO analysis (in cellular components), it is observed that most of the hub genes are present in plastid, vacoule, membrane-bounded vacuole, and cytoplasmic vacuole (Fig. 5.2b), mainly responsible for pumping out ions from the cell. In MF category, majority of the hub genes were found to be involved in nucleic acid, cation ion, metal ion and zinc ion binding activities (Fig. 5.2b), which may be responsible for fixing metal ions. Further, a large portion of the hub genes were found to be responsible for nitrogen compound metabolic process, response to stress and chemical stimulus, defense response, and signal transduction under biological process category.
The module membership of the hub genes as well as UHG under Al stress showed that most of the hub genes under Al stress condition belong to turquoise (86), yellow (45), and brown (38) modules (Table 5.2). Similarly, out of 130 UHG, mainly 45, 32, and 26 are the members of turquoise, brown, and yellow color modules, respectively. Interestingly, it can be seen that the blue color module contains the second highest number of selected genes (137) but has no hub genes, while the brown color module is the third largest module (100 genes) which contains 38 hub genes, out of which 32 are UHG for Al stress. Further, brown color module is found to be associated with various important functions like ion binding, redox activity, kinase activity, and phosphotransferase activity (Table 5.2), which are important for the abiotic stress response in plants (Wang et al. 2013). From the molecular biology point of view, the brown color module along with its members seems to be very important for breeding Al stress-resistant varieties.
5.4 Discussion
The Boot-SVM-RFE technique was a superior technique for the selection of informative genes from high-dimensional gene expression data (Das et al. 2017). This approach is also advantageous over classical gene selection techniques like t-test and F-score, as it does not require any distributional assumptions about the data. In this technique, a p-value was assigned to each gene, and genes with lower p-values were considered as informative for the particular condition/trait under investigation. The selection of informative genes based on p-values is scientific as well as statistically meaningful to experimental biologists as compared to other techniques. Further, the bootstrap procedure used in this technique was expected to remove the spurious associations of the genes with their classes.
The statistical approach for hub gene identification allowed the ranking and selection of candidate hub genes in the GCN, based on an assessment of the statistical significance of the gene connections. This was done with a randomized resampling-based procedure where statistical significance values were calculated based on the NP test, which does not require Gaussian assumptions of data. Further, genes with lower p-values represent highly connected genes in the GCN and thus designated as hub genes. Moreover, the randomization procedure used in this approach allows one to test whether the observed gene connectivity is greater than expected gene connectivity value by chance (i.e., rejection of null hypothesis of random association). This was also able to remove the spurious association among genes, as these associations are measured on the basis of PCC. It seems to be more statistically convincing to select hub genes based on p-values rather than WGS alone, because in comparison to WGS, the p-values provide a reliable measure of gene connectivity based on a statistical criterion (lower p-value indicates high gene connectivity and vice-versa). Further, the detected hub genes tend to have higher connection degrees and are widely separated from the genes with low connection degrees in the GCN. Moreover, based on this approach, a few and important genes were identified as hubs in the GCN as compared to existing approach, which is in accordance with the scale-free property of biological networks.
Using the DHGA approach, genes in the GCN were grouped into various categories like HHG, non-hubs, and UHG for stress and control based on the computed p-values for these two contrasting conditions. These identified hub genes may be considered as biomarkers for further studies, including analysis of their involvement in diverse cellular mechanisms. Further, the HHG can be used for the maintenance of basal cellular functions that are essential for the existence of a cell (Eisenberg and Levanon 2013), whereas UHG can be used in stress response engineering in crops for developing stress-tolerant cultivars.
Understanding Al stress response mechanism in soybean is of paramount importance for plant breeders to develop Al stress-tolerant cultivars. In public domain databases, there are few samples available related to Al stress in soybean, which have been generated over varying experimental conditions by multiple studies. Then, machine learning and network biology techniques were applied to identify the responsible genes to understand stress response mechanism in this crop. It has been reported that there are two main processes involved in Al stress response in plants: (i) exclusion of Al ions from root cells and (ii) detoxification of Al ions in the plant cells (Kochian et al. 2005). Some selected genes were found to be involved in transporting of metal ions outside the cell, which might be associated with the first process. The function like redox activity related to electron transport under chemical reactions that balances the charges during metallic ions transport (Wang et al. 2013) might be associated with the second process. The redox activity might also be related to ROS generation that is produced in plants in response to Al stress. Further, ROS also seriously disrupts normal metabolism of cell through peroxidation of lipids (Wise and Naylor 1987), proteins, and nucleic acids (Imlay and Linn 1988). The increased redox activity is consistent with the activation of the antioxidative enzymes such as catalase, ascorbate peroxidase, and guaiacol peroxidase under abiotic stress condition (Tuteja 2007). The activities under BP taxonomy like cellular glucan and cellular amino acid metabolic processes are known to increase in plants in response to various abiotic stresses (Obata and Fernie 2012), and other reported biological processes need to be studied in the context of Al toxic stress. The role of phosphotranferase activity in conferring tolerance against abiotic stresses like drought and salt in rice and Arabidopsis is well established (Duan and Cai 2012). The role of stress-induced organic acid synthesis in conferring Al tolerance in higher plants is also well reported (Yang et al. 2013). These processes might be related to detoxification of Al ions, which occurs rapidly after exposure to Al stress in plants (Ryan et al. 2001).
5.5 Conclusions
In this chapter, machine learning and network biology techniques are used to understand the Al toxic stress response in soybean using gene expression data. Here, the main focus was as follows: first, the machine learning-based Boot-SVM-RFE technique was used for the selection of informative genes from high-dimensional soybean gene expression data. Second, a novel statistical approach was used for the identification of hub genes in a GCN. Third, the DHGA approach was used to group genes in the GCN into various categories based on their gene connectivity values. This chapter throws some light to understand the mechanism of Al stress response in soybean, and some key important genes were reported. Moreover, functional enrichment analysis of these key genes revealed their associated intracellular functions under Al stress. The information revealed in this article on various molecular mechanisms like biosynthesis of secondary metabolites and stress-specific roles of certain plant products may be useful for mitigation of Al stress in plants, particularly in soybean. These identified genes can act as potential targets for bioengineering of Al toxic stress response in soybean.
References
Apse M, Aharon G, Snedden W, Blumwald E (1999) Salt tolerance conferred by over expression of a vacuolar Na+/H+ antiport in Arabidopsis. Science 285(5431):1256–1258
Barabasi AL, Oltvai ZN (2004) Network biology: understanding the cell’s functional organization. Nat Rev Genet 5:101–113. https://doi.org/10.1038/nrg1272
Chen BS, Yang SK, Lan CY, Chuang YJ (2008) A systems biology approach to construct the gene regulatory network of systemic inflammation via microarray and databases mining. BMC Med Genet 1:46. https://doi.org/10.1186/1755-8794-1-46
Childs KL, Davidson RM, Buell CR (2011) Gene coexpression network analysis as a source of functional annotation for rice genes. PLoS One 6:e22196. https://doi.org/10.1371/journal.pone.0022196
Cui X, Churchill G (2003) Statistical tests for differential expression in cDNA microarray experiments. Genome Biol 4(4):210
Das S, Meher PK, Rai A, Bhar LM, Mandal BN (2017) Statistical approaches for gene selection, hub gene identification and module interaction in gene co-expression network analysis: an application to aluminum stress in soybean (Glycine max L.). PLoS One 12(1):e0169605. https://doi.org/10.1371/journal.pone.0169605
Díaz-Uriarte R, de Andrés SA (2006) Gene selection and classification of microarray data using random forest. BMC Bioinformatics 7:3. https://doi.org/10.1186/1471-2105-7-3
Downs GS, Bi YM, Colasanti J, Wu W, Chen X (2013) A developmental transcriptional network for Zea mays defines coexpression modules. Plant Physiol 161(4):1830–1843. https://doi.org/10.1104/pp.112.213231
Du Z, Zhou X, Ling Y, Zhang Z, Su Z (2010) AgriGO: a GO analysis toolkit for the agricultural community. Nucleic Acids Res 38:64–70. https://doi.org/10.1093/nar/gkq310
Duan J, Cai W (2012) Oslea3-2, an abiotic stress induced gene of rice plays a key role in salt and drought tolerance. PLoS One 7(9):e45117. https://doi.org/10.1371/journal.pone.0045117
Eisenberg E, Levanon EY (2013) Human housekeeping genes, revisited. Trends Genet 29(10):569–574
Ficklin SP, Luo F, Feltus FA (2010) The association of multiple interacting genes with specific phenotypes in rice using gene coexpression networks. Plant Physiol 154:13–24. https://doi.org/10.1104/pp.110.159459
Forman G (2003) An extensive empirical study of feature selection metrics for text classification. J Mach Learn Res 3:1289–1305
Gautier L, Cope L, Bolstad BM, Irizarry RA (2004) Affy – analysis of Affymetrix GeneChip data at the probe level. Bioinformatics 20:307–315. www.affymetrix.com/support
Gentleman RC, Carey VJ, Bates DM, Bolstad B, Dettling M, Dudoit S, Ellis B, Gautier L, Ge Y, Gentry J (2004) Bioconductor: open software development for computational biology and bioinformatics. Genome Biol 5:80
Grant D, Nelson RT, Cannon SB, Shoemaker RC (2010) SoyBase, the USDA-ARS soybean genetics and genomics database. Nucleic Acids Res 38:D843–D846. https://doi.org/10.1093/nar/gkp798
Guoyon I, Elisseeff A (2003) An introduction to variable and feature selection. J Mach Learn Res 3:1157–1182
Guyon I, Weston J, Barnhill S, Vapnik V (2002) Gene selection for cancer classification using support vector machines. Mach Learn 46:389–422
Imlay J, Linn S (1988) DNA damage and oxygen radical toxicity. Science 240:1302–1309
Kochian L, Pineros M, Hoekenga O (2005) The physiology, genetics and molecular biology of plant aluminum resistance and toxicity. Plant and Soil 274(1):175
Kochian LV, Hoekenga OA, Pineros MA (2004) How do crop plants tolerate acid soils? Mechanisms of aluminum tolerance and phosphorous efficiency. Annu Rev Plant Biol 55:459–493. PMID: 15377228
Lai H, Han B, Li L, Chen Y, Zhu L (2011) An integrated semi-random forests based approach to gene selection for Glioma classification. Acta Biophys Sin 26(9):833–845
Langfelder P, Horvath S (2008) WGCNA: an R package for weighted correlation network analysis. BMC Bioinformatics 9:559. https://doi.org/10.1186/1471-2105-9-559
Liang Y, Zhang F, Wang J, Joshi T, Wang Y et al (2011) Prediction of drought-resistant genes in Arabidopsis thaliana using SVM-RFE. PLoS One 6(7):e21750. https://doi.org/10.1371/journal.pone.0021750
Ma JF (2007) Syndrome of aluminum toxicity and diversity of aluminum resistance in higher plants. Int Rev Cytol 264:225–252
Mao K, Zhao P, Tan PH (2006) Supervised learning based cell image segmentation for p53 immunohistochemistry. IEEE Trans Biomed Eng 53(6):1153–1163
Miller G, Shulaev V, Mittler R (2008) Reactive oxygen signaling and abiotic stress. Physiol Plant 133(3):481–489
Niu X, Narasimhan M, Salzman R, Bressan R, Hasegawa P et al (1996) NaCl regulation of plasma membrane H+-ATPase gene expression in a Glycophyte and a halophyte. Plant Physiol 111:679–718
Obata T, Fernie AF (2012) The use of metabolomics to dissect plant responses to abiotic stresses. Cell Mol Life Sci 69:3225–3243. https://doi.org/10.1007/s00018-012-1091-5
Panda SK, Baluska F, Matsumoto H (2009) Aluminum stress signaling in plants. Plant Signal Behav 4(7):592–597
R Core Team (2015) R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna
Ryan PR, Delhaize E, Jones DL (2001) Function and mechanism of organic anion exudation from plant roots. Annu Rev Plant Physiol Plant Mol Biol 52:527–560
Stumpf MPH, Porter MA (2012) Critical truths about power laws. Science 335:665–666. https://doi.org/10.1126/science.1216142
Tuteja N (2007) Mechanisms of high salinity tolerance in plants. Methods Enzymol 428:419–438. https://doi.org/10.1016/s0076-6879(07)28024-3
Wang J, Chen L, Wang Y, Zhang J, Liang Y, Xu D (2013) A computational systems biology study for understanding salt tolerance mechanism in Rice. PLoS One 8(6):e64929. https://doi.org/10.1371/journal.pone.0064929
Wise R, Naylor A (1987) Chilling-enhanced photooxidation: evidence for the role of singlet oxygen and superoxide in the breakdown of pigments and endogenous antioxidants. Plant Physiol 83:278–282
Yang LT, Qi YP, Jiang HX, Chen LS (2013) Roles of organic acid anion secretion in aluminium tolerance of higher plants. Biomed Res Int:173682. https://doi.org/10.1155/2013/173682
Zeng QY, Yang CY, Ma QB, Li XP, Dong WW, Nian H (2012) Identification of wild soybean miRNAs and their target genes responsive to aluminum stress. BMC Plant Biol 12:182. https://doi.org/10.1186/1471-2229-12-182
Zhang B, Horvath S (2005) A general framework for weighted gene co-expression network analysis. Stat Appl Genet Mol Biol 4(17)
Zhang L, Yu S, Zuo K, Luo L, Tang K (2012) Identification of gene modules associated with drought response in rice by network-based analysis. PLoS One 7:e33748. https://doi.org/10.1371/journal.pone.0033748
Acknowledgments
Authors acknowledge the help and support obtained from the Indian Council of Agricultural Research (ICAR), New Delhi, India; ICAR-Indian Agricultural Statistics Research Institute, New Delhi, India; National Institutes of Health, USA; and University of Louisville, USA.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this chapter
Cite this chapter
Das, S., Maity, A. (2022). Utility of Network Biology Approaches to Understand the Aluminum Stress Responses in Soybean. In: Wani, S.H., Sofi, N.u.R., Bhat, M.A., Lin, F. (eds) Soybean Improvement. Springer, Cham. https://doi.org/10.1007/978-3-031-12232-3_5
Download citation
DOI: https://doi.org/10.1007/978-3-031-12232-3_5
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-12231-6
Online ISBN: 978-3-031-12232-3
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)