Abstract
Automated Machine Learning (AutoML) tools are being increasingly used by researchers to solve a large number of pattern recognition tasks. A typical AutoML tool efficiently evaluates the performance of classifiers (e.g., SVM, Neural Network, Decision Tree, etc.) on an input dataset and determines the best performing classifier, along with the corresponding optimized parameters. In this work, we explore the use of a fusion scheme to combine the outputs of multiple classifier models generated and optimized by an AutoML tool known as Auto Tuned Models (ATM). We show that this approach provides a flexible framework that can be successfully applied to a wide variety of medical image classification tasks. Because each individual classification model is optimized for the given dataset, regardless of the size of the dataset, there is no requirement to identify and transfer knowledge from other domains. We generate up to 3,600 models for each dataset and explore the efficiency of fusion on subsets of models, as well as present two methods to automatically sort the classifier models for fusion. Experiments conducted on three medical imaging datasets, viz., stem cell, brain tumor and prostate cancer, convey the benefits of the proposed fusion scheme in improving classification accuracy.
Access provided by Autonomous University of Puebla. Download conference paper PDF
Similar content being viewed by others
Keywords
1 Introduction
A classical pattern recognition system maps input data (e.g., an image) to an output label referred to as a class (e.g., “face” or “bicycle” or “stem-cell”). This mapping is accomplished using a classifier such as a neural network or a support vector machine, with each classifier having a number of tunable parameters. Determining the best classifier and the associated optimized parameters for a given pattern recognition task is often relegated to a trial-and-error approach that can be both laborious and sub-optimal. Further, a classification model (i.e., a classifier along with its parameters) that is optimal on one dataset may not work well on other datasets.
To address this issue, there is a need for developing techniques that can automatically determine the best classifier for a given dataset. Over the years, a suite of such automated machine learning (AutoML) tools have been created and shown to have the ability to surpass the state of the art performances of previously hand-selected models. One such tool, Auto Tuned Models (ATM), not only surpassed performance of human-tuned models on 30% of 420 datasets in OpenML, but also completed this work in 1/100th of the time [21].
These tools have the ability to produce many different optimized classifier models based on different classifier types and different parameter combinations. In this work, we explore the possibility of combining multiple classifier models generated by an AutoML tool in order to further improve classification accuracy. The fusion scheme is tested on three medical imaging datasets corresponding to stem cell detection, brain tumor classification and prostate cancer classification.
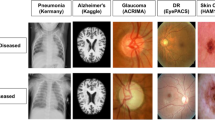
Medical image classification is often challenging due to the costs of the imaging machines, the required expertise to annotate images, and patient privacy laws. These challenges often result in limited data availability which can restrict the ability to train deep neural networks, which often require large annotated datasets. Adaptations to deep neural networks such as transfer learning [4] or self-supervised learning techniques [11] have been proposed to address these challenges. However, these approaches rely on the ability to appropriately select an existing architecture or the creation of custom image transformations. Thus, the use of classification techniques that do not require copious amounts of annotated training data can be pertinent in such cases (Fig. 1).
2 Background
We define a classifier as the type of classification algorithm, such as K-Nearest Neighbors (KNN), support vector machine (SVM), or multiple-layered perceptron (MLP). Once a classifier’s parameters have been set, either by human intuition or via an AutoML tool, the resulting instance is referred to as a classifier model (see Table 1).
2.1 Pattern Recognition with AutoML
Given labeled data, a supervised classifier learns a model that can accurately map the input data to the correct output class label. The performance of a model depends on many factors such as the amount of training data available, the balance of data across classes, and the distribution of the data itself. A classifier model that performs well on a given dataset may not achieve the same performance on a different dataset for the same task.
Software packages used to generate these models have started to offer automated methods for classifier selection and parameter tuning. These methods are referred to as automated machine learning (AutoML) tools, and have provided a way to intelligently find classifier models optimized for specific datasets [12]. Popular AutoML tools include Auto-WEKA [14], Auto-Sklearn [10], and TPOT (Tree-based Pipeline Optimization Tool for Automating Data Science) [19]. In this work, we utilized a recently developed AutoML tool known as Auto Tuned Models (ATM) [21]. We choose ATM for its ability to tune models in parallel, as well as the ability to archive all the generated models.
ATM is constructed with Python’s SciKit-Learn, and allows the same classifiers that are supported in SciKit-Learn. An example of the SVM classifier, its parameters, and the resulting tuned model are given in Table 1.
The above approaches focus on classical pattern recognition approaches. Newer AutoML tools such as Auto-DeepLab [17] and Auto-Keras [13] focus on tuning deep learning models. However, we do not focus on the latter set of tools as our goal is to demonstrate how simple classification models, trained on small-sized datasets, can be judiciously selected and combined to improve classification performance in the context of medical image classification.
2.2 Fusion
An AutoML tool typically outputs multiple classifier models, including the one with the highest accuracy on the training set. While one model may have the highest accuracy, is it possible to leverage multiple models to achieve higher performance? Combining multiple sources of information to improve performance is generally referred to as fusion, and there are multiple levels at which fusion can be performed. This includes fusion at the raw data level [8], feature level, score level [20], rank level [20], and decision level [16].
This work considers fusion of multiple classifier models generated by ATM at the score level. Since this technique relies only on the score reported, it is possible to use this approach without knowledge of the features used to produce the score. In this work, the score generally denotes the confidence of each classifier model when it renders an output class label based on the input feature vector.
There are many approaches to fusion, such as the simple sum rule where scores are averaged [20]. The fusion in this paper is performed with an SVM, where the scores from multiple models are input to an SVM which then maps this score vector to an output class or label. The question we raise is the following: how do we determine the models whose scores have to be fused?
3 Datasets
Three different medical imaging datasets are utilized in this work. These datasets include stem cells, brain tumors, and prostate cancer grade classifications for magnetic resonance imaging (MRI) scans. In our analysis, the MRI scans are comprised of a stack of 16-bit tiff images. These images are generated from DICOM (Digital Imaging and Communications in Medicine standard), a common file format generated by medical imaging machines such as MRIs.
3.1 MSU Stem Cell Dataset
Afridi et al. created a dataset of 6 in vivo MRI scans of rat brains that were hand-labeled for stem cells by radiologists [1, 2]. This dataset was generated from 2 different MRI machines of varying field strengths, and is intended to evaluate the generalizability and robustness of potential stem cell detection algorithms. Each scan is composed of 16-bit tiff images (Fig. 2, left), which radiologists manually reviewed to identify and label stem cells. Once a stem cell is identified, a 9 \(\times \) 9 pixel patch is extracted around the stem cell (Fig. 2, right).
Using handcrafted features and Bayesian Classifiers, Afridi et al. [1] obtained an AUC accuracy of 89.1%. They were able to further improve performance using a CNN-based approach that incorporated the information about the time radiologists spent identifying stem cells (referred to as Labeling Latency). This approach achieved an accuracy of 94.6% [2]. For our analysis, we divide the labeled data into 80% training and 20% testing. This is summarized in Table 2 (Figs. 3).
3.2 Brain Tumor Classification
Cheng et al. [6] generated a dataset comprised of brain scans of 233 patients with identified tumors. Figure 4 provides examples of four slices (top) and the annotated tumors (bottom). In addition to the manually segmented tumor masks (highlighted in yellow on the bottom row), we display the bounding box surrounding the tumors in red. Three types of tumors are identified: Glioma, Meningioma, and Pituitary. Table 3 gives the breakdown of the 3,064 MRI slices. For our analysis, we divide the labeled data into 80% training and 20% testing. This is summarized in Table 3.
The authors of this dataset explored classification of tumors through region augmentation by including the segmented tumor and the regions surrounding it [7]. They obtained a classification accuracy of 82.3%. Afshar et al. further improved classification performance to 86.56% by designing a Capsule Network [3]. In our work, however, we only use the segmented tumor without surrounding pixels for classification.
3.3 Prostate Cancer Classification
The American Association of Physicists in Medicine, the International Society for Optics and Photonics, and the National Cancer Institute collaborated to create the PROSTATEx Challenges for classifying the aggressiveness of prostate cancer [5]. The PROSTATEx challenge dataset provides information for 182 subjects. This information includes multiple MRI scans with multiple weighting agents, and a cancer diagnosis based on Gleason Score grouping. A low Gleason score indicates small, well formed and uniform glands, i.e., mostly normal cells. A high score indicates irregular sizes, glands, or masses (abnormal cells). These scores are categorized into 5 groups defined in Table 4. Additionally, a set of KTrans images (a measure of capillary permeability is provided for each patient. This data is already divided into training and test sets. The best results of this competition was an AUC accuracy score of 95% (https://prostatex.grand-challenge.org/evaluation/results/) (Fig. 5).
4 Proposed Approach
The contribution of this work is the principled use of fusion to combine the multiple classifier models produced by an AutoML tool such as ATM. Questions we explore include which data representations to use, how many models are necessary to fuse, and how to select models for fusion. In this section, the proposed approach is described in detail. The proposed approach implements the following steps:
-
1.
Extract Data: For each of the datasets described in Sect. 2, image patches are extracted from the MRI scan and given a class label.
-
2.
Form Feature-Sets: While the pixels in a patch can be directly used as raw input to ATM, we also consider alternate inputs where a patch is subjected to different feature extraction techniques via texture descriptors. These alternate representations along with the original patch representation are referred to as Feature-Sets.
-
3.
Generate Models with ATM: ATM [21] generates multiple classifier models from the training set of extracted patches and the corresponding feature-sets.
-
4.
Collect Model Scores: Test patches are given as input to the models to obtain a score (described later in Sect. 4.4).
-
5.
Perform Fusion: The scores of multiple classifier models are fused using an SVM classifier.
For the rest of this section, let \(m_{ijk}\) represent the \(i^{th}\) model pertaining to the \(j^{th}\) classifier tuned on the \(k^{th}\) feature-set (\(i\in Z\), \(j \in J\), \(k \in K\)), and let FM represent the subset of models to be used in fusion. Here, K represents the feature-sets (raw, scaled, HOG, LBP, LBP-Uniform, LBP-Rotation Invariant), and J represents the collection of classifiers (Logistic Regression, Support Vector Machine, Linear Classifier with Stochastic Gradient Descent, Decision Tree, Extra Trees, Random Forest, Gaussian Naive Bayes, Multinomial Naive Bayes, Bernoulli Naive Bayes, Gaussian Process, Passive Aggressive, K Nearest Neighbors, Multi-Layer Perceptron).
4.1 Data Extraction
For the stem cell dataset, we use the same method proposed by Afridi et al. [2] to extract image patches. We center patches around regions of interest (stem cell and non-stem cell), as shown in Fig. 2. The resulting patches are 9 \(\times \) 9 pixels.
For the brain tumor dataset, we first find the smallest width and also the smallest height of all the bounding boxes within the dataset. This worked out to be 14 pixels on both dimensions. We then locate the center pixel of the tumor’s bounding box, and extract a 14 \(\times \) 14 pixel patch surrounding the center pixel. This approach ensures that the patches contain only tumor information and nothing of the surrounding brain tissue.
For the prostate dataset, locations of legions were provided in the same manner as the stem cell data. That is, an expert identified point-of-interest is labeled within an MRI scan. We extract a 15 \(\times \) 15 pixel patch of the slice around the point of interest.
4.2 Data Descriptors Through Feature-Sets
In addition to the raw pixel values within the patches, scaled pixel intensities and texture descriptors are also considered. These methods produce the following feature-sets: Raw pixel intensities (in the range [0, 65,535]), Scaled Intensities (in the range of [0, 255]), Histogram of Oriented Gradients (HOG) [9], DAISY feature descriptors [22], and Local Binary Pattern (Classical, Rotation-Invariant, Uniform) [18].
4.3 Model Generation
Once all the feature-sets are generated, the labeled training data for each feature-set in K is run through ATM. We set ATM’s model budget, B, to 600 models for each feature-set in K. With \(|K| = 6\), this means ATM produces up to 3,600 tuned models. Since the number of tuned models is very large, we would like a principled manner to form a subset of candidate models, FM, that will reliably improve performance while efficiently selecting only the models necessary to achieve good results. This selection process is described in Sect. 4.5.
4.4 Model Scores
Given an input feature-set, each tuned classifier model predicts an output class label. Additionally, each model produces a score, which in many cases, corresponds to a probability value. For example, a multi layered perceptron (MLP) model classifies the input data by using a softmax score, which is the probability that the input data belongs to class y. If scores are not readily available from a tuned model, other representations of prediction strengths are used to estimate a score. To obtain scores for the KNN classifier, for example, we average the Euclidean distance of the K neighbors. The smaller the distance, the stronger the evidence is that the point is correctly classified. For the Extra Trees and Random Forest classifiers, the score denotes the percentage of trees that correctly classify the sample, and for the singular decision tree, the score corresponds to the percentage of nodes in agreement.
4.5 Model Selection for Fusion
With so many optimized models generated, a question arises. Should all models be included in fusion? It is possible that several optimized models are only slight variations of a dominant classifier’s parameters, resulting in redundant score data and biased classification predictions. Before we propose methods to automatically select models for fusion, we first explore various methods to partitioning the models to form subset FM and address potential bias.
Forming FM: The first obvious method to subsetting M into FM is to select the top n models reporting the highest classification accuracy (Top). This approach is straightforward and is effective at drastically cutting the size of M. However, as before, we find that if a certain classifier performs particularly well on a dataset, a large portion of the n models in FM will belong to the same classifier and make similar classification errors. Therefore, we next explore forming FM by partitioning M based on the classifier type (CS), the feature-set (FS), and the classifier-feature-set combination (CFS). To form FM, we select the highest performing model from each partition. For example, in the CS stratification approach we select the model with the highest reported AUC accuracy for each classifier type. Lastly, we form FM by considering every optimized model produced by ATM (FM == M). In summary, the following are the approaches we apply to form FM: Top, CS, FS, CFS, and the entire set of models.
Once the models in FM have been identified using one of the aforementioned methods, the next step is to design an algorithm to effectively select the minimal number of classifier models to fuse. Intelligently selecting models for fusion begins with deciding which model best complements the models already selected for fusion. To facilitate this, we compute the pair-wise score correlation between all pairs of candidate models. Typically, combining models which are least correlated can be beneficial [15]. In this work, we develop two techniques for selecting models to fuse. In both approaches, we start with the best performing model generated by ATM and then use pairwise correlation value to guide model selection. Then, we compare the performance of these two approaches, examining how the accuracy and efficiency are impacted by the number of models fused.
Static Selection: The first selected model is the best model produced by ATM. The remainder of the models in FM that are not selected form the unselected model set. The next model chosen is the unselected model with the least correlation to the last selected model. This process is repeated until there are no more unselected models and the selection is complete. By using the Pearson correlation between models’ scores across training samples in order to select models, we are selecting models whose scores contain new information not captured by the previously selected models.


Dynamic Selection: We next explore sorting models by dynamically updating the correlation values after selecting a new model for fusion. Once the first 2 candidate models are selected as described in the static sort above, we immediately fuse the selected models to obtain a new model with the new set of fused scores. We update the correlation coefficients with the unselected models to account for the scores from the newly fused model. This selection method is dynamic and allows for the flexibility to select the next model based on the current models already selected for fusion.
5 Results and Analysis
The proposed method is able to improve the AUC accuracies, both over any individual AutoML generated model and over the previous state-of-the-art accuracies described in Sect. 3. We focus on the Prostate and Brain datasets, as ATM produced significantly optimized models for the Stem Cell dataset with individual AUC accuracies over 99%. While no single stratification approach universally performed the best, we find that using a stratification approach reports higher AUC accuracies than randomly selecting the same number of models (repeated 5 times), as shown in Table 5, with the class-feature stratification (CFS) technique providing the best results with the fewest models for the prostate dataset, and selecting models from the top-performing stratification (TOP) provides the best results for the brain dataset. Figure 6 shows how the AUC accuracies change as models are added to the fusion.
6 Summary
We applied score fusion to combine multiple classification models generated by an AutoML tool to produce improved accuracy on three medical imaging datasets. Further, we developed two methods to select models for fusion based on Pearson’s correlation coefficient. This case study focused on medical image datasets, and obtained several classification models via an AutoML tool (ATM) to fuse together. We observe that fusing models improves the accuracy beyond the best individual classification model produced by ATM, and that the highest accuracy achieved through fusion of these models surpasses the accuracy of even deep learning approaches. Furthermore, these results are achieved with small training data sizes without auxiliary information, such as labeling behavior or pixels surrounding the area of interest. This underscores the importance of judiciously selecting models for fusion in order to improve classification accuracy.
References
Afridi, M.J., Liu, X., Shapiro, E., Ross, A.: Automatic in vivo cell detection in MRI. In: Navab, N., Hornegger, J., Wells, W.M., Frangi, A.F. (eds.) MICCAI 2015. LNCS, vol. 9351, pp. 391–399. Springer, Cham (2015). https://doi.org/10.1007/978-3-319-24574-4_47
Afridi, M.J., Ross, A., Shapiro, E.M.: L-CNN: exploiting labeling latency in a CNN learning framework. In: Pattern Recognition (ICPR), pp. 2156–2161 (2016)
Afshar, P., Mohammadi, A., Plataniotis, K.N.: Brain tumor type classification via capsule networks. In: 25th IEEE International Conference on Image Processing (ICIP), pp. 3129–3133 (2018)
Alzubaidi, L., et al.: Novel transfer learning approach for medical imaging with limited labeled data. Cancers 13(7), 1590 (2021)
Armato, S.G., et al.: PROSTATEx challenges for computerized classification of prostate lesions from multiparametric magnetic resonance images. J. Med. Imaging 5(4), 044501 (2018)
Cheng, J.: Brain Tumor Dataset, April 2017. https://doi.org/10.6084/m9.figshare.1512427.v5. https://figshare.com/articles/brain_tumor_dataset/1512427/5
Cheng, J., et al.: Enhanced performance of brain tumor classification via tumor region augmentation and partition. PLoS ONE 10(10), e0140381 (2015)
Chitroub, S.: Classifier combination and score level fusion: concepts and practical aspects. Int. J. Image Data Fusion 1(2), 113–135 (2010). https://doi.org/10.1080/19479830903561944
Dalal, N., Triggs, B.: Histograms of oriented gradients for human detection. In: IEEE Computer Society Conference on Computer Vision and Pattern Recognition, vol. 1, pp. 886–893 (2005)
Feurer, M., et al.: Efficient and robust automated machine learning. In: Advances in Neural Information Processing Systems, pp. 2962–2970. Curran Associates, Inc. (2015)
Ghesu, F.C., et al.: Self-supervised learning from 100 million medical images (2022)
Hutter, F., Kotthoff, L., Vanschoren, J.: Automated Machine Learning: Methods, Systems, Challenges. Springer, Cham (2019). https://doi.org/10.1007/978-3-030-05318-5
Jin, H., Song, Q., Hu, X.: Auto-keras: efficient neural architecture search with network morphism. arXiv preprint arXiv:1806.10282 (2018)
Kotthoff, L., Thornton, C., Hoos, H.H., Hutter, F., Leyton-Brown, K.: Auto-WEKA: automatic model selection and hyperparameter optimization in WEKA. In: Hutter, F., Kotthoff, L., Vanschoren, J. (eds.) Automated Machine Learning. TSSCML, pp. 81–95. Springer, Cham (2019). https://doi.org/10.1007/978-3-030-05318-5_4
Kuncheva, L.I., Whitaker, C.J.: Measures of diversity in classifier ensembles and their relationship with the ensemble accuracy. Mach. Learn. 51(2), 181–207 (2003)
Lam, L., Suen, S.: Application of majority voting to pattern recognition: an analysis of its behavior and performance. IEEE Trans. Syst. Man Cybern. Part A Syst. Hum. 27(5), 553–568 (1997)
Liu, C., et al.: Auto-deeplab: hierarchical neural architecture search for semantic image segmentation. In: Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, pp. 82–92 (2019)
Ojala, T., Pietikainen, M., Maenpaa, T.: Multiresolution gray-scale and rotation invariant texture classification with local binary patterns. IEEE Trans. Pattern Anal. Mach. Intell. 24(7), 971–987 (2002)
Olson, R.S., et al.: Automating biomedical data science through tree-based pipeline optimization. In: Squillero, G., Burelli, P. (eds.) EvoApplications 2016. LNCS, vol. 9597, pp. 123–137. Springer, Cham (2016). https://doi.org/10.1007/978-3-319-31204-0_9
Ross, A.A., Nandakumar, K., Jain, A.K.: Handbook of Multibiometrics. Springer, Boston (2006). https://doi.org/10.1007/0-387-33123-9
Swearingen, T., et al.: ATM: A distributed, collaborative, scalable system for automated machine learning. In: IEEE International Conference on Big Data, Boston, MA, USA, 11–14 December, pp. 151–162 (2017). https://doi.org/10.1109/BigData.2017.8257923
Tola, E., Lepetit, V., Fua, P.: Daisy: an efficient dense descriptor applied to wide-baseline stereo. IEEE Trans. Pattern Anal. Mach. Intell. 32(5), 815–830 (2010)
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 Springer Nature Switzerland AG
About this paper
Cite this paper
Dale, M., Ross, A., Shapiro, E.M. (2022). Fusing AutoML Models: A Case Study in Medical Image Classification. In: El Yacoubi, M., Granger, E., Yuen, P.C., Pal, U., Vincent, N. (eds) Pattern Recognition and Artificial Intelligence. ICPRAI 2022. Lecture Notes in Computer Science, vol 13364. Springer, Cham. https://doi.org/10.1007/978-3-031-09282-4_15
Download citation
DOI: https://doi.org/10.1007/978-3-031-09282-4_15
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-09281-7
Online ISBN: 978-3-031-09282-4
eBook Packages: Computer ScienceComputer Science (R0)