Abstract
Inclusion bodies (IBs) are protein aggregates formed under recombinant protein production processes in microbial cell factories. Their characterization has shown that they are self-assembling and biologically active protein nanoparticles with promising properties for a wide range of applications, including biocatalysis, tissue engineering, and therapy. Besides, different protocols have also been developed to obtain soluble protein from IBs using non-denaturing conditions.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Inclusion bodies
- Functional nanoparticles
- Protein nanoparticles
- Protein aggregates
- Recombinant protein production
1 Introduction
Inclusion bodies (IBs) are protein aggregates formed during recombinant protein production in microbial cell factories (Fig. 1). For decades they have been considered a useless byproduct, and various strategies have been developed aimed to increase protein solubility thereby reducing the aggregation phenomenon. However, for over 15 years now, this vision has started to change. Different research groups have demonstrated that bacterial IBs are far from being merely inactive recombinant protein deposits (de Marco et al. 2019; Rinas et al. 2017). They have been characterized as self-assembling protein nanoparticles with a dual composition: (1) an amyloid-like scaffold and (2) folded or partially folded protein species with β-amyloid structure (Cano-Garrido et al. 2013; Cano-Garrido et al. 2016). The folded or partially folded protein conformers are biologically active and can be easily released from the scaffold under physiological conditions (Seras-Franzoso et al. 2016; Carratalá et al. 2021a). In addition, the formation of these protein aggregates is a general phenomenon (Villaverde et al. 2015), rather than being a specific trait of Escherichia coli, observed in different microbial cell factories, including Lactococcus lactis (Cano-Garrido et al. 2016) and Pichia pastoris (Rueda et al. 2016; Carratalá et al. 2020a). Thus, IBs can be easily produced through a scalable process, and straightforward purification protocols have been optimized consisting in multiple centrifugation and washing steps (Seras-Franzoso et al. 2015). Altogether, IB features have radically changed the concept of protein aggregation and become an attractive protein-based biomaterial with a variety of applications, including biocatalysis, biomedical therapy, and tissue engineering (de Marco et al. 2019; Rinas et al. 2017; Gifre-Renom et al. 2020a; Hrabárová et al. 2015; Liovic et al. 2012; García-Fruitós et al. 2012). Moreover, the demonstration that IBs are nanoparticles containing proteins with their native structure has led to a modification of the protocols used to obtain soluble protein from IBs which proven that denaturing and resolubilization steps are not necessary (Gifre-Renom et al. 2018; Singhvi et al. 2020; Singh et al. 2015a; Ferrer-Miralles et al. 2018). Alternatively, mild or non-denaturing solubilization protocols have been developed by different research groups to solubilize proteins from IBs maintaining protein structure and activity (Fig. 1).
Recombinant protein production and purification process. Recombinant bacteria produce the protein of interest in both soluble and aggregated (nanoparticle or inclusion bodies) forms. After that, cells are disrupted and soluble protein and protein nanoparticles are purified. If necessary, the soluble form can be obtained from inclusion bodies through a solubilization process
2 Protein Production and IB Formation
The bioactivity of releasable proteins forming IBs coupled with the high stability of these protein nanoparticles has made IBs an appealing alternative form to their soluble counterpart (Gifre-Renom et al. 2020a; Gifre-Renom et al. 2020b; Pesarrodona et al. 2019). Thus, contrary to what has been done for years, several research groups have focused on optimizing the formation and purification of IBs (Seras-Franzoso et al. 2015). Production time and temperature are the two cultivation parameters that can significantly influence the formation of aggregates (Vera et al. 2007; Garcia-Fruitós 2009). Increased production temperature has an impact not only on the size of IBs, i.e., larger size at higher growth temperatures (García-Fruitós et al. 2007), but also on the conformational quality of the aggregated proteins (Vera et al. 2007). Lower growth temperatures improve the conformational quality of both soluble and insoluble protein fractions (Vera et al. 2007; Jevsevar et al. 2005), meaning that IBs containing more active proteins can be easily obtained. The bacterial strain used to produce the recombinant protein of interest can also determine the properties of these proteins. Whereas in most cases IBs are spherical-like nanoparticles (Cano-Garrido et al. 2016; Garcia-Fruitós 2009; García-Fruitós et al. 2010), some E. coli strains produced tear-shaped aggregates with particular characteristics for tissue engineering applications (García-Fruitós et al. 2010). IBs of the same proteins produced in different bacterial systems (E. coli and L. lactis) also show differences in size and surface functional group density when used for micropatterned surface decoration (Martínez-Miguel et al. 2020).
Another strategy that has been used to optimize the formation of IBs is based on the use of protein tags based on specific aggregation-prone proteins, including the foot-and-mouth disease virus capsid protein (VP1) (García-Fruitós et al. 2005a), a variant of the human β-amyloid peptide (Morell et al. 2008), the maltose-binding protein mutant (Arié et al. 2006), poxB from Paenibacillus polymyxa (Park et al. 2012), and the cellulose-binding domain of Clostridium cellulovorans (Nahalka and Nidetzky 2007). Also, shorter aggregation peptides (Carratalá et al. 2020a; Carratalá et al. 2021b; Wang et al. 2015a; Wu et al. 2011; Zhou et al. 2012; Jiang et al. 2019; Küsters et al. 2021), such as coiled-coil domains (Küsters et al. 2021; Jäger et al. 2019; Jäger et al. 2018; Gil-Garcia et al. 2020; Lamm et al. 2020) and leucine zippers (Choi et al. 2014; Roca-Pinilla et al. 2020a), have been successfully used to promote protein aggregation. Although all these tags have been described to promote aggregation, a screening for each specific protein would be needed to gain optimal results.
3 Structure, Composition, and Activity of IBs
The study and characterization of IBs have been mainly focused on their structure and composition. Regarding the structure, as already mentioned, there are two main parts conforming the aggregates: the β-sheet skeleton, which is a common structure in all IBs (García-Fruitós et al. 2011; de Groot et al. 2009; Castillo et al. 2011), and the fractions formed by the native or native-like recombinant proteins, which is protein-dependent (Rinas et al. 2017). The former provides the mechanical and chemical stability, while the latter is responsible for the specific activity of the IBs (Carratalá et al. 2020a; García-Fruitós et al. 2005a).
Despite all IBs present a common structural pattern, their composition varies. It depends on the recombinant protein that is being produced but also on the specific recombinant cell factory used. Thus, other molecules or impurities such as host cell proteins like chaperones, lipids, lipopolysaccharide (LPS), and/or nucleic acids, can be accumulated inside the IBs increasing the variability of their composition (Roca-Pinilla et al. 2020a; Rinas and Bailey 1992; Valax and Georgiou 1993). Therefore, the molecular complexity of IBs requires a huge variety of techniques for their characterization (Rinas et al. 2017). In this section, techniques extensively used to describe IBs physicochemical and biological features are provided in detail (Fig. 2).
Transmission electron microscopy (TEM) and some of its variants, such as Cryo-TEM, are techniques used for the visualization of intracellular structures, such as organelles or membranes, at high resolution. Furthermore, it is possible to detect the cell components, including IBs, at nanoscale. Various publications have shown that these microscopical techniques allow to identify the presence of the IBs and to determine their exact location inside the recombinant cell factories (Rinas et al. 2017; Cano-Garrido et al. 2016; Rueda et al. 2016; Zhou et al. 2012). Normally, IBs present a polar distribution inside the producer cells, showing an electrodense pattern under the microscope. Moreover, using purified IBs as a sample, it is possible to determine their size and shape (Cano-Garrido et al. 2013; Garcia-Fruitós 2009; García-Fruitós et al. 2010; Zhou et al. 2012; Wang et al. 2008). The size of these protein aggregates usually ranges between 50 and 500 nm with a pseudospherical shape in most of the cases.
Scanning electron microscopy (SEM) is another technique widely used to determine physical parameters of IBs like shape, roughness, volume, and size (Cano-Garrido et al. 2016; Pesarrodona et al. 2019; Díez-Gil et al. 2010), which complements TEM analysis. SEM allows to work directly with IBs samples without any preparation prior to their visualization. For that, both SEM and TEM are usually used together in many IBs studies, obtaining a complete physical characterization of these protein aggregates (Carratalá et al. 2020a; Garcia-Fruitós 2009; Torrealba et al. 2016a). With 3D visualization, it has been possible to observe a porous structure in IBs, which is an interesting property not only for its application as biocatalysts but also as biomaterials to decorate surfaces and promote cell adhesion and proliferation (see the following sections).
Atomic force microscopy (AFM) is a microscopy technique with a wide applicability in IBs research (Garcia-Fruitós 2009; Díez-Gil et al. 2010; Sanagavarapu et al. 2019). This microscopy technique perfectly completes the information obtained by TEM and SEM, capable of characterizing morphology and stiffness of the samples directly under environmental conditions.
Dynamic light scattering (DLS) is used to measure the size distribution of nanoparticles or protein samples in solution. It allows researchers to determine the volume size distribution of soluble proteins in either unassembled or monomeric form (Unzueta et al. 2020), as well as the size and zeta potential of IBs (Carratalá et al. 2020a; Garcia-Fruitós 2009; Díez-Gil et al. 2010). The zeta potential provides information on the superficial charge of IBs, in which negatively charged IBs indicate the aggregation-prone nature of these protein-based aggregates (Garcia-Fruitós 2009; Díez-Gil et al. 2010). Furthermore, in some cases, DLS is used in the stability studies of IBs, by performing the measurements at different time points to see if the physical properties are maintained. DLS is very useful to obtain the particles’ mean diameter when the sample is homogenous, and there is no important variance in the diameter of the particles. Moreover, DLS can differentiate populations of different particle sizes within the sample if the size of each population is homogenous (Martínez-Miguel et al. 2020). Thus, DLS offers many possibilities for the study of IBs, as it is relatively easy to measure under environmental conditions (i.e., buffers, solution, etc.).
Other techniques have allowed studying in detail the secondary structure of proteins forming these aggregates. Fourier transform infrared spectroscopy (FTIR) is the most widely used technique for this purpose, and different articles have proven that IBs contain correctly folded proteins but also an amyloid structure inside the IBs (Jevsevar et al. 2005; Natalello and Doglia 2015; Ami et al. 2006). FTIR is capable to detect and differentiate between β-sheet and α-helix structures, opening the possibility to define the structure and the organization of the proteins inside the aggregates (Garcia-Fruitós 2009). FTIR also allows researchers to establish comparisons between the soluble and aggregated forms of the same protein, elucidating differences in the structure or the appearance of new patterns during the aggregation process (Cano-Garrido et al. 2016; Wang 2009). This fact is an important advantage compared to other techniques used for secondary structure analyses, such as circular dichroism (CD), that only permits the analysis of soluble protein samples. Specifically, the pattern described in IBs by FTIR comprises a part of β-sheet structure which corresponds to the IB scaffold and serves as the link to α-helix parts conforming to the native-folded protein inside the aggregates (Roca-Pinilla et al. 2020a; Ami et al. 2006). In addition, FTIR has been used for the in vivo detection of IBs in bacteria, following their rate of formation at different temperatures (Ami et al. 2005). The secondary structure evolves from α-helixes predomination at earlier stages of the aggregation process, corresponding to the native-folded protein, to β-sheet forms showing the growth of the IBs inside the cells.
To determine the specific composition of IBs, techniques such as 2D sodium dodecyl sulfate-polyacrylamide gel electrophoresis (2D SDS-PAGE) have been applied. As previously mentioned, recombinant protein coexists with other cell proteins. This has been studied in detail using 2D SDS-PAGE, alloweing researchers to take a step forward to elucidate the heterogeneity inside these aggregates (Jevsevar et al. 2005; Rinas and Bailey 1992; Rinas et al. 1993; Jürgen et al. 2010). Alternatively, matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-ToF MS) can be used to identify cell proteins present in the aggregate (Gardner et al. 2019). Besides, western blot (WB) is widely used to specifically quantify the amount of recombinant protein present in IBs. Protocols for protein quantification in IBs have been well established, and this has allowed not only to determine IB recombinant protein yields but also to determine the soluble/aggregated protein ratio during recombinant protein production processes (Gifre-Renom et al. 2018; García-Fruitós et al. 2007).
However, IBs conformational heterogeneity is not only determined by different types of proteins; many molecules can be embedded inside these aggregates. In this context, recombinant protein production using E. coli as the cell factory has a big concern: the presence of LPS in the recombinant products. LPS, even at low amounts, can induce an endotoxic immune response in mammals, which greatly limits their biomedical applicability. Thus, its detection is crucial for knowing possible drawbacks or alterations to the final activity of the proteins that conform the aggregates (Rueda et al. 2014). Due to the possible obstacles that this could cause, the use of LPS-free systems or generally recognized as safe (GRAS) microorganisms, such as lactic acid bacteria, has increased significantly in the last few years (García-Fruitós 2012; Song et al. 2017). In addition to LPS impurities, carbohydrates or lipids could be present inside the IBs, being possible to determine the specific amounts in each case (Roca-Pinilla et al. 2020a).
Beyond the physicochemical, structural, and morphological characterization of IBs, different research groups have been working on the analysis of the activity of recombinant proteins in form of IBs. It has been widely demonstrated that IBs are formed (at least partially) by active proteins in a very important percentage. It is well-known that the composition of the IBs is directly related to the recombinant protein produced and that is why the assays to study the activity are protein-dependent. Thus, different protocols have been optimized for the determination of IBs activity including enzymatic activity assays (Hrabárová et al. 2015; García-Fruitós et al. 2005a; Tokatlidis et al. 1991; García-Fruitós et al. 2005b; Worrall and Goss 1989; Gifre-Renom et al. 2020c), cytokine activity (Carratalá et al. 2020a), antimicrobial activity (Roca-Pinilla et al. 2020b), and also fluorescence emission (Garcia-Fruitós 2009). The presence of activity in these protein aggregates has opened a large number of possibilities in terms of applicability, as detailed in the following sections.
4 Stability of IBs
IBs are protein aggregates with a notable mechanical stability (Rinas et al. 2017). It has been widely described that their amyloid structure allows them to preserve their integrity and morphology upon mechanical, chemical, and enzymatic cell disruption and upon long-term storage under different conditions. Recent research has correlated these in vitro observations with the in vivo efficiency of this new biomaterial while comparing the effect of other types of nanoparticles. Matrix metalloproteinase-9 (MMP-9) protein, which has an important role in facilitating the migration of immune cells, has been used as a model protein in these studies (Gifre-Renom et al. 2020b). The MMP-9 IBs were compared with their soluble counterpart and MMP-9 encapsulated in polymeric-based micelles (PM) through ionic and covalent binding. The soluble MMP-9 and the MMP-9-ionic PM showed the highest activity values in vitro, whereas IBs showed the lowest activity values. However, the in vitro stability test in 50% bovine serum at room temperature proved that the IBs were the most stable format. Interestingly, the data were well correlated in vivo using an intra-dermal air-pouch model in mice. MMP-9 IBs appeared to be the biomaterial with the highest in vivo activity compared to the soluble MMP-9 form, that was associated with a low and a transitory peak of activity. These results demonstrated that, although the IBs are not always the most active format in vitro, their stability can switch this biomaterial in the most active form once administrated in vivo thanks to their slow-release properties and resilience to protein degradation.
5 Inclusion Bodies as Active Nanoparticles: Applications
Acknowledging IBs as functional protein nanoparticles (Jevsevar et al. 2005; García-Fruitós et al. 2005a; Peternel and Komel 2011) boosted a new perspective in research that aimed to exploit the different properties of these nanoparticles in a wide range of applications. As such, during the last decades, IBs have exhibited an important economic potential in industrial catalysis and an intrinsic biological interest in tissue engineering and in therapeutic research, including regenerative medicine, drug delivery in cancer and immunotherapy, and their antimicrobial use as alternatives to classic antibiotics (Fig. 3).
5.1 IBs in Biocatalysis
Pharmaceuticals, food-related products, biofuels, detergents, and other everyday goods, such as paper and textile, are being produced at high rates by engineered biocatalysts. The use of enzymes at industrial scale was enhanced after the DNA technology discovery, which provided the tools to obtain recombinant enzymes on demand. The main limitations were then the lifetime and stability of these enzymes in which, together with the high costs involved in protein purification processes, they made industrial scale-up expensive. The immobilization of the enzymes with either organic or inorganic carriers, or through carrier-free cross-linked enzyme aggregation (further reviewed methods in Wang et al. (2015b)), allowed the recyclability of these catalysts for several enzymatic reactions, importantly reducing the overall costs. However, laborious and expensive chromatographic purification of the enzymes and covalent or non-covalent attachments to carriers are inherent in these approaches. In fact, these are still in the focus of biocatalysts optimization.
In this regard, IBs have demonstrated to be a highly convenient type of carrier-free immobilizing method for biocatalysts (Fig. 4a). The fact that these aggregates do not require any chromatographic purification steps makes them already appealing from their production process, which is important for the reduction in industrial scaling-up costs (Kloss et al. 2018). Hence, abundant research has been dedicated to the study of IBs as biocatalysts, as summarized in the book chapter by Hrabárová et al. (2015) and in the mini-review by Jäger et al. (2020). However, the benefits of using IBs go beyond purification-related processes. On the one hand, since IBs are spontaneously formed as aggregates (otherwise promoted by fusion of aggregation-inducing tags (Jäger et al. 2019)), no further immobilization procedures are required afterward, reducing not only any adsorption efficiency issue but also time and enzyme manipulation. Moreover, these new class of biocatalysts are reusable, maintaining their activity after multiple cycles of catalysis (Fig. 4a) (Rinas et al. 2017; Nahálka et al. 2008). On the other hand, IBs can be easily tailored to fit a desired size, resistance to solubilizing agents, porosity, purity, or specific activity by simply modulating the production conditions (e.g., temperature, pH, etc.) (de Marco et al. 2019) or the expression system. The porous nature of IBs, for instance, can benefit its adhesion in case of solid phase catalysis. Moreover, improved IB biocatalysts can be obtained by combining different proteins or by fusion of specific protein domains (Jäger et al. 2019). For example, the fusion of an hydroxynitrile lyase to a coiled-coil domain improved the enzyme resistance to acidic pH (Diener et al. 2016), or two-step cascade reactions could be facilitated by colocalizing different enzymes within the IBs, further improving the stability of the involved enzymes (Jäger et al. 2018).
Examples of IBs activity. (a) Yields of substrate to product conversion using cross-linked IBs. Reprinted from Nahalka et al., 2008, copyright (2008) with permission from Elsevier. (b) Cell growth of 1671, HeLa, HepG2, and PC12 cell cultures on FGF-2 IBs and nonfunctional IBs (IBs IR). Reprinted from Seras-Franzoso et al., 2014, copyright (2014) with permission from Elsevier. (c) Quantification of apoptotic nuclei in treated tumors with (FN-GFP-H6) and FN-p31-H6 IBs. Reprinted from Pesarrodona et al., 2019 (open access article distributed under the terms of the Creative Commons CC BY license). (d) Antibiofilm activity of JAMF1 IBs against carbapenem-resistant Klebsiella pneumoniae (KPC). Reprinted from Roca-Pinilla et al., 2020 (open access article licensed under a Creative Commons Attribution 4.0 International License)
Thus, so far, many enzymes have proven to form catalytically active IBs exploitable as industrial biocatalysts. Some examples are hydrolases (Park et al. 2012; Tokatlidis et al. 1991; Worrall and Goss 1989; Li et al. 2013; Dong et al. 2014), oxidoreductases (García-Fruitós et al. 2005a), oxidases (Hrabárová et al. 2015), lyases (Kloss et al. 2018), phosphatases (Huang et al. 2013), kinases (Nahálka et al. 2006), aldolases (Nahálka et al. 2008; Sans et al. 2012), phosphorylases (Nahálka 2008), and transferases (Korovashkina et al. 2012; Mestrom et al. 2020). Nevertheless, IBs do not guarantee per se the best (although still high) catalytic performance, as described by a recent work in which IBs and carrier-immobilization methods are compared for trehalose transferase (Mestrom et al. 2020); instead, this might be an enzyme-to-enzyme matter. The benefits of using IBs as biocatalysts are, however, not only drastically reducing the overall procedure costs and providing stability and tailorable properties to the enzyme but also conferring the possibility to design cascade-like bioreactions relatively easily.
5.2 IBs in Therapy/Nanopills
Another remarkable potential of IBs is their use for drug delivery and regenerative medicine approaches in biomedicine and animal science. The fact that the nanoparticle form of proteins provides stability to the embedded proteins, along with their slow-release properties in physiological conditions and the capability of engineering the proteins for targeting purposes, motivated an extensive study of IBs to identify their therapeutic value as nanoscale aggregates with releasable protein (nanopills) (Vázquez et al. 2012). For example, IBs formed by biologically irrelevant proteins (i.e., green fluorescent protein (GFP)) were used to decorate surfaces for in vitro cell culture and demonstrated to increase human mesenchymal stem cells adherence and to promote their differentiation to osteoblasts (Seras-Franzoso et al. 2014). This bottom-up approach was further investigated with IBs composed of extracellularly and/or intracellularly acting biologically relevant proteins (e.g., fibroblast growth factor 2 and human chaperone Hsp70), with results suggesting the cellular internalization of the slowly released proteins from the immobilized IBs (Fig. 4b) (Seras-Franzoso et al. 2014; Seras-Franzoso et al. 2013a; Seras-Franzoso et al. 2013b). These results are highly valuable for tissue engineering and regenerative medicine applications (Martínez-Miguel et al. 2020). This example has already been materialized in in vitro wound healing studies accomplished by protein hormone-releasing IBs (Stamm et al. 2018). In addition, IBs in suspension can also be internalized by mammalian cells with a high efficiency (Seras-Franzoso et al. 2016; Vázquez et al. 2012), suggesting their potential for drug delivery in vivo and inspiring tumor-targeting research, as will be discussed later on. In the context of the aforementioned applications of IBs, it is relevant to point out the absence of toxicity signs neither when injected (intraperitoneal or intratumorally) nor when administrated orally in mice and zebrafish (Vázquez et al. 2012; Torrealba et al. 2016b). Further, orally administered tumor necrosis factor (TNF)-α IBs protected zebrafish from a lethal infection through their immunostimulant action (Torrealba et al. 2016b). Other proteins forming IBs have been studied also for animal science applications. The low costs of IB production and their versatility make these nanomaterials especially appealing for veterinary applications.
5.3 IBs in Cancer
IBs have several properties that make them also attractive as peptide/protein drug delivery systems (DDS) for cancer (Vázquez et al. 2012). The IBs stability, penetrability, and slow-release properties, therefore, might be useful for the delivery of the proteins to tumoral tissues, if the proteins can target a specific biomarker that is only present or overexpressed in malignant cells. Taking advantage of this, researchers sought to target CD184+ colorectal cancer cells, which overexpress the chemokine receptor CD184 (Unzueta et al. 2018). They designed two modular constructs based on the T22 and R9 proteins using GFP as a fusion partner. These two proteins, T22 and R9, are highly specific ligands of the CD184 receptor and, thus, are able to home in cells that overexpress it. In addition, T22 and R9, when fused to GFP, and in combination with a poly-His tag, can self-assemble in protein nanoparticles (pNPs) that can efficiently direct and increase pNPs cell penetration in tumoral cells. These pNPs can also aggregate forming IBs and provide an immobilized DDS that maintained the sustained release of these tumor-targeting pNPs both in vitro and in vivo. Remarkably, T22-GFP IBs provided high amounts of pNPs for up to 10 days after subcutaneous administration. This work only proved the appealing slow-release properties of IBs for cancer applications, with an efficient accumulation in the desired malignant tissue, yet it did not use any therapeutic proteins.
Moving a step further, Pesarrodona et al. designed two different modular proteins formed by the polypeptides p31 and Omomyc (Pesarrodona et al. 2019). The first protein (p31) is based on a fragment from p130cas which can promote apoptosis (Casanova et al. 2006). The second protein (Omomyc) is a dominant negative mutant of the Myc protein which is involved in the cell cycle and possesses antitumoral properties (Whitfield et al. 2017). These two proteins were each fused to a tumor-homing peptide (FN) that binds to the CD44 receptor, which is a widely recognized tumoral marker and is related to tumor progression and metastasis. In this study, these fusion antitumoral constructs were produced recombinantly as IBs, which were able to selectively enter and kill CD44+ cells in vitro in cell culture, and in vivo in a mice model of human breast cancer (Fig. 4c). Moreover, an important conclusion of both studies was that, in any case, there was no off-target accumulation and toxicity.
5.4 Antimicrobial IBs
Antibiotic-resistant bacteria (ARB) are one of the biggest threats to global health today. Many infectious diseases that could be successfully treated with antibiotic drugs have acquired resistance to most or nearly all of the available compounds, including a growing list of infections, such as pneumonia, tuberculosis, gonorrhea, and foodborne diseases (Laxminarayan et al. 2013). These infections are becoming harder, and sometimes impossible, to treat with the current generation of medicines. To make matter worse, no new class of antibiotics has been discovered since the 1980s, and few antibiotics are being developed to face the challenge of ARBs. So, there is an urgent need for the development of new antimicrobials to enable the treatment for those infections effectively.
Among the potential alternatives to conventional antibiotics, antimicrobial IBs might be a promising one, as they are highly stable and have slow-release properties likely able to maintain antimicrobial activity. Constant administration of antimicrobial compounds that have a short half-life is required, or otherwise, the use of concentrations that are under the minimum inhibitory concentration (MIC) will probably increase antimicrobial resistances (AMRs) (Gao et al. 2011). A slow-release profile seems to be vital to maintain constant antimicrobial levels for long periods of time, to obtain the optimal therapeutic benefits while minimizing AMRs. Another benefit might be that one long-lasting administration is more advantageous than multiple short-lived ones. Further, IBs offer all these properties without the need to manipulate them further (i.e., encapsulation processes or protein embedding in a matrix), because they are produced in a one-step process.
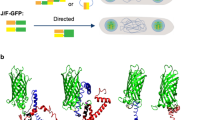
A study published in 2020 found out that the antimicrobial peptide (AMP) GWH1 had a therapeutic effect in a mouse mastitis model (Carratalá et al. 2020b). The GWH1 peptide was fused to the N-terminus of either of GFP or IFN-γ in a form of IBs and resulted in diminished bacterial loads by five- and sixfold, respectively. The study also found that the IB form per se can be antimicrobial, as GFP or IFN-γ IBs showed no antimicrobial activity and, in some cases, achieve a similar reduction in bacterial loads when compared to GWH1 fusion constructs (Carratalá et al. 2020b). Another study found that a multidomain AMP named JAMF1 displayed a clear antimicrobial effect against several ARB, such as Klebsiella pneumoniae, E. coli, Enterococcus spp., and E. faecalis, in both planktonic and biofilm state (Fig. 4d) (Roca-Pinilla et al. 2020b). The JAMF1 construct was based on the combination of human α-defensin-5 (HD5), human XII-A secreted phospholipase A2 (sPLA2), and a gelsolin-based bacterial-binding domain. Two aggregation-seeding domains based on leucine zippers were also added to promote IB formation during the recombinant production of JAMF1.
5.5 IBs a Source of Soluble Protein
Inclusion bodies have also been used for years as a source to obtain soluble proteins when these cannot be obtained directly from the cell cytoplasm or from the growth media if secreted. Although some heterologous proteins are produced mainly in the soluble form, most of them are in the aggregated form. In some cases, the aggregation levels are so high that the only option to obtain the soluble form is through the solubilization of IBs. One example of this is matrix metalloproteinase 9 (MMP-9); when using L. lactis as the cell factory, MMP-9 is only produced in an aggregated form (Gifre-Renom et al. 2018). Other proteins of interest are produced in a soluble form, but they are co-purified with host cell proteins. This occurs, for example, with mammary serum amyloid protein A3 (M-SAA3), and their solubilization from IBs is the only protocol that allows to have good purity levels (Gifre-Renom et al. 2018). Thus, for both aggregation-prone and difficult-to-produce/purify proteins, the solubilization of IBs is the only strategy for their purification.
The strategy that has been traditionally used to recover soluble proteins from IBs has been based on the following sequential steps: IB isolation and purification from the recombinant culture, IB solubilization with harsh detergents and chaotropic agents (e.g., urea or guanidine hydrochloride) at high concentration (i.e., 6–8 M), refolding of the solubilized protein to reach their native conformation, and purification by chromatography methods (Singh et al. 2015a). However, the use of these solubilization agents lead to a protein denaturation, and refolding steps are necessary to recover the original structure and function of the target proteins. Solubilization and refolding are critical steps limiting the yield of recovered protein, and have been subjected to an intense and expanding research (Burgess 2009; Singh et al. 2015b). Usually, proteins can be refolded upon removal of denaturing agents jointly providing favorable conditions to reach their native state (Singhvi et al. 2020). The underlying principle in the refolding process is that the proteins are able to switch back into their native conformation from a denatured condition. However, during this process, non-desirable aggregated intermediates can be formed. In the refolding process, quality and quantity of the folded protein rely on the buffer, protein concentration, and the method used (Singhvi et al. 2020). Alternative and simpler strategies considering the biological nature of bacterial IBs have been developed over the last few years. The new description of IBs as protein aggregates containing properly folded and biologically active recombinant proteins has permitted to advance significantly in the development of new protocols without the need of using denaturing agents to solubilize proteins from IBs (Ferrer-Miralles et al. 2018). For that, mild solubilization agents, such as n-lauroylsarcosine, dimethyl sulfoxide (DMSO), and organic solvents such as n-propanol and isopropanol are used, which are strong enough to solubilize IBs without disturbing the native structure of the protein (Singhvi et al. 2020; Ferrer-Miralles et al. 2018; Peternel et al. 2008; Sarker et al. 2019). The use of n-lauroylsarcosine to recover soluble protein from IBs has allowed the development of a non-denaturing protocol for IBs that are produced from E. coli (Roca-Pinilla et al. 2020b; Peternel et al. 2008; Park et al. 2018; Francis et al. 2012) and L. lactis (Gifre-Renom et al. 2018). Target proteins are gradually solubilized in a single step, keeping their native structure. In addition, mild detergents enhance membrane protein recovery due to stabilization of their hydrophobic groups, playing both roles of solubilization and retaining its folded and active state (Francis et al. 2012). Once solubilized, the protein can be further purified by chromatography. An alternative to mild detergents are organic solvents, which have been proven to be plausible alternative to obtain high-quality protein from IBs. The use of n-propanol (Singh et al. 2012), for example, has been reported to be an efficient bioactive protein solubilization method. In addition, trifluoroethanol (TFE) has also demonstrated a remarkable potential to achieve functional protein from IBs (Upadhyay et al. 2016). Nevertheless, organic solvents can generate non-desirable chemical modifications, and in many cases they need to be used in combination with low concentrations of chaotropic agents (Singhvi et al. 2020).
Another strategy is based in the use of high hydrostatic pressure to disaggregate the insoluble aggregates and refold them back in their native structure (Ferrer-Miralles et al. 2018; St John et al. 2001). In addition, the use of heat (Cai et al. 2020), pH oscillations (Panda 2003), and freeze-thaw cycles (Qi et al. 2015) with the combination of low amounts of denaturant agents has been postulated as efficient methods to solubilize and recover bioactive protein from IBs (Ferrer-Miralles et al. 2018).
6 Conclusions
Inclusion bodies arise as a new protein-based biomaterial that is spontaneously formed in bacterial hosts during recombinant protein production. Several characterization procedures based on electron microscopy, molecular techniques, and physicochemical analyses can be used to finely determine their structure and composition. IBs have been demonstrated to be a promising biomaterial for biocatalysis, tissue regeneration, drug delivery, and antimicrobial therapy applications. It is likely that the IBs’ stability along with their slow-release properties is the basis for its potential and successful use. Although the soluble version of a recombinant protein is needed in some cases, the target protein can be easily obtained from IBs under mild solubilization protocols and free of host cell proteins and other impurities. In short, IBs are valuable in the world of recombinant proteins and new protein-based biomaterials, which motivates to deeply study new microorganisms as IBs producers to tune those features that can be further optimized.
References
Ami D, Natalello A, Gatti-Lafranconi P, Lotti M, Doglia SM (2005) Kinetics of inclusion body formation studied in intact cells by FT-IR spectroscopy. FEBS Lett 579(16):3433–3436
Ami D, Natalello A, Taylor G, Tonon G, Maria DS (2006) Structural analysis of protein inclusion bodies by Fourier transform infrared microspectroscopy. Biochim Biophys Acta 1764(4):793–799
Arié JP, Miot M, Sassoon N, Betton JM (2006) Formation of active inclusion bodies in the periplasm of Escherichia coli. Mol Microbiol 62(2):427–437
Burgess RR (2009) Refolding solubilized inclusion body proteins. Methods Enzymol 463:259–282
Cai H, Chen G, Yu H, Tang Y, Xiong S, Qi X (2020) One-step heating strategy for efficient solubilization of recombinant spider silk protein from inclusion bodies. BMC Biotechnol 20(1):37
Cano-Garrido O, Rodríguez-Carmona E, Díez-Gil C, Vázquez E, Elizondo E, Cubarsi R et al (2013) Supramolecular organization of protein-releasing functional amyloids solved in bacterial inclusion bodies. Acta Biomater 9(4):6134–6142
Cano-Garrido O, Sanchez-Chardi A, Pares S, Giro I, Tatkiewicz WI, Ferrer-Miralles N et al (2016) Functional protein-based nanomaterial produced in microorganisms recognized as safe: a new platform for biotechnology. Acta Biomater 43:230–239
Carratalá JV, Cano-Garrido O, Sánchez J, Membrado C, Pérez E, Conchillo-Solé O et al (2020a) Aggregation-prone peptides modulate activity of bovine interferon gamma released from naturally occurring protein nanoparticles. New Biotechnol 57:11–19
Carratalá JV, Brouillette E, Serna N, Sánchez-Chardi A, Sánchez JM, Villaverde A et al (2020b) In vivo bactericidal efficacy of GWH1 antimicrobial peptide displayed on protein nanoparticles, a potential alternative to antibiotics. Pharmaceutics. 12(12):1217
Carratalá JV, Gifre-Renom L, Roca-Pinilla R, Villaverde A, Arís A, Garcia-Fruitós E et al (2021a) Selecting subpopulations of high-quality protein conformers among conformational mixtures of recombinant bovine MMP-9 solubilized from inclusion bodies. Int J Mol Sci 22(6):3020
Carratalá JV, Cisneros A, Hellman E, Villaverde A, Ferrer-Miralles N (2021b) Title: insoluble proteins catch heterologous soluble proteins into inclusion bodies by intermolecular interaction of aggregating peptides. Microb Cell Factories 20(1):30
Casanova I, Parreño M, Farré L, Guerrero S, Céspedes MV, Pavon MA et al (2006) Celecoxib induces anoikis in human colon carcinoma cells associated with the deregulation of focal adhesions and nuclear translocation of p130Cas. Int J Cancer 118(10):2381–2389
Castillo V, Graña-Montes R, Sabate R, Ventura S (2011) Prediction of the aggregation propensity of proteins from the primary sequence: aggregation properties of proteomes. Biotechnol J 6(6):674–685
Choi SL, Lee SJ, Yeom SJ, Kim HJ, Rhee YH, Jung HC et al (2014) Controlled localization of functionally active proteins to inclusion bodies using leucine zippers. PLoS One 9(6):e97093
de Groot NS, Sabate R, Ventura S (2009) Amyloids in bacterial inclusion bodies. Trends Biochem Sci 34(8):408–416
de Marco A, Ferrer-Miralles N, Garcia-Fruitós E, Mitraki A, Peternel S, Rinas U et al (2019) Bacterial inclusion bodies are industrially exploitable amyloids. FEMS Microbiol Rev 43(1):53–72
Diener M, Kopka B, Pohl M, Jaeger K-E, Krauss U (2016) Fusion of a coiled-coil domain facilitates the high-level production of catalytically active enzyme inclusion bodies. ChemCatChem 8(1):142–152
Díez-Gil C, Krabbenborg S, García-Fruitós E, Vazquez E, Rodríguez-Carmona E, Ratera I et al (2010) The nanoscale properties of bacterial inclusion bodies and their effect on mammalian cell proliferation. Biomaterials 31(22):5805–5812
Dong Q, Yan X, Zheng M, Yang Z (2014) Characterization of an extremely thermostable but cold-adaptive β-galactosidase from the hyperthermophilic archaeon Pyrococcus furiosus for use as a recombinant aggregation for batch lactose degradation at high temperature. J Biosci Bioeng 117(6):706–710
Ferrer-Miralles N, Saccardo P, Garcia-Fruitós E. Chapter 6. Protein purification from protein aggregates. In: Publishers NS, editor. Handbook on Protein Purification: Industry Challenges and Technological Developments. 2018
Francis VG, Majeed MA, Gummadi SN (2012) Recovery of functionally active recombinant human phospholipid scramblase 1 from inclusion bodies using N-lauroyl sarcosine. J Ind Microbiol Biotechnol 39(7):1041–1048
Gao P, Nie X, Zou M, Shi Y, Cheng G (2011) Recent advances in materials for extended-release antibiotic delivery system. J Antibiot 64(9):625–634
García-Fruitós E (2012) Lactic acid bacteria: a promising alternative for recombinant protein production. Microb Cell Factories 11:157
García-Fruitós E, González-Montalbán N, Morell M, Vera A, Ferraz RM, Arís A et al (2005a) Aggregation as bacterial inclusion bodies does not imply inactivation of enzymes and fluorescent proteins. Microb Cell Factories 4:27
García-Fruitós E, Carrió MM, Arís A, Villaverde A (2005b) Folding of a misfolding-prone beta-galactosidase in absence of DnaK. Biotechnol Bioeng 90(7):869–875
García-Fruitós E, Martínez-Alonso M, Gonzàlez-Montalbán N, Valli M, Mattanovich D, Villaverde A (2007) Divergent genetic control of protein solubility and conformational quality in Escherichia coli. J Mol Biol 374(1):195–205
García-Fruitós E, Sabate R, de Groot NS, Villaverde A, Ventura S (2011) Biological role of bacterial inclusion bodies: a model for amyloid aggregation. FEBS J 278(14):2419–2427
García-Fruitós E, Seras-Franzoso J, Vazquez E, Villaverde A (2010) Tunable geometry of bacterial inclusion bodies as substrate materials for tissue engineering. Nanotechnology 21(20):205101
García-Fruitós E, Vázquez E, Díez-Gil C, Corchero JL, Seras-Franzoso J, Ratera I et al (2012) Bacterial inclusion bodies: making gold from waste. Trends Biotechnol 30(2):65–70
Garcia-Fruitós E-C (2009) EscarlataDíez-Gil, CésarFerraz, Rosa MVázquez, EstherCorchero, José Luis Cano-Sarabia, Mary Ratera, Imma, Ventosa N, Veciana J. Villaverde A Surface cell growth engineering assisted by a novel bacterial nanomaterial Adv Mater 21:1–5
Gardner QA, Hassan N, Hafeez S, Arif M, Akhtar M (2019) Exploring the nature of inclusion bodies by MALDI mass spectrometry using recombinant proinsulin as a model protein. Int J Biol Macromol 139:647–653
Gifre-Renom L, Cano-Garrido O, Fàbregas F, Roca-Pinilla R, Seras-Franzoso J, Ferrer-Miralles N et al (2018) A new approach to obtain pure and active proteins from Lactococcus lactis protein aggregates. Sci Rep 8(1):13917
Gifre-Renom L, Ugarte-Berzal E, Martens E, Boon L, Cano-Garrido O, Martínez-Núñez E et al (2020a) Recombinant protein-based nanoparticles: elucidating their inflammatory effects in vivo and their potential as a new therapeutic format. Pharmaceutics. 12(5):450
Gifre-Renom L, Seras-Franzoso J, Rafael D, Andrade F, Cano-Garrido O, Martinez-Trucharte F et al (2020b) The biological potential hidden in inclusion bodies. Pharmaceutics 12(2):157
Gifre-Renom L, Carratalá JV, Parés S, Sánchez-García L, Ferrer-Miralles N, Villaverde A et al (2020c) Potential of MMP-9 based nanoparticles at optimizing the cow dry period: pulling apart the effects of MMP-9 and nanoparticles. Sci Rep 10(1):11299
Gil-Garcia M, Navarro S, Ventura S (2020) Coiled-coil inspired functional inclusion bodies. Microb Cell Factories 19(1):117
Hrabárová E, Achbergerová L, Nahálka J (2015) Insoluble protein applications: the use of bacterial inclusion bodies as biocatalysts. Methods Mol Biol 1258:411–422
Huang Z, Zhang C, Chen S, Ye F, Xing XH (2013) Active inclusion bodies of acid phosphatase PhoC: aggregation induced by GFP fusion and activities modulated by linker flexibility. Microb Cell Factories 12:25
Jäger VD, Kloss R, Grünberger A, Seide S, Hahn D, Karmainski T et al (2019) Tailoring the properties of (catalytically)-active inclusion bodies. Microb Cell Factories 18(1):33
Jäger VD, Lamm R, Kloß R, Kaganovitch E, Grünberger A, Pohl M et al (2018) A synthetic reaction cascade implemented by colocalization of two proteins within catalytically active inclusion bodies. ACS Synth Biol 7(9):2282–2295
Jäger VD, Lamm R, Küsters K, Ölçücü G, Oldiges M, Jaeger KE et al (2020) Catalytically-active inclusion bodies for biotechnology-general concepts, optimization, and application. Appl Microbiol Biotechnol 104(17):7313–7329
Jevsevar S, Gaberc-Porekar V, Fonda I, Podobnik B, Grdadolnik J, Menart V (2005) Production of nonclassical inclusion bodies from which correctly folded protein can be extracted. Biotechnol Prog 21(2):632–639
Jiang L, Xiao W, Zhou X, Wang W, Fan J (2019) Comparative study of the insoluble and soluble Ulp1 protease constructs as carrier free and dependent protein immobilizates. J Biosci Bioeng 127(1):23–29
Jürgen B, Breitenstein A, Urlacher V, Büttner K, Lin H, Hecker M et al (2010) Quality control of inclusion bodies in Escherichia coli. Microb Cell Factories 9:41
Kloss R, Limberg MH, Mackfeld U, Hahn D, Grünberger A, Jäger VD et al (2018) Catalytically active inclusion bodies of L-lysine decarboxylase from E. coli for 1,5-diaminopentane production. Sci Rep 8(1):5856
Korovashkina AS, Rymko AN, Kvach SV, Zinchenko A (2012) Enzymatic synthesis of c-di-GMP using inclusion bodies of Thermotoga maritima full-length diguanylate cyclase. J Biotechnol 164(2):276–280
Küsters K, Pohl M, Krauss U, Ölçücü G, Albert S, Jaeger KE et al (2021) Construction and comprehensive characterization of an EcLDCc-CatIB set-varying linkers and aggregation inducing tags. Microb Cell Factories 20(1):49
Lamm R, Jäger VD, Heyman B, Berg C, Cürten C, Krauss U et al (2020) Detailed small-scale characterization and scale-up of active YFP inclusion body production with Escherichia coli induced by a tetrameric coiled coil domain. J Biosci Bioeng 129(6):730–740
Laxminarayan R, Duse A, Wattal C, Zaidi AK, Wertheim HF, Sumpradit N et al (2013) Antibiotic resistance-the need for global solutions. Lancet Infect Dis 13(12):1057–1098
Li S, Lin K, Pang H, Wu Y, Xu J (2013) Production, characterization, and application of an organic solvent-tolerant lipase present in active inclusion bodies. Appl Biochem Biotechnol 169(2):612–623
Liovic M, Ozir M, Zavec AB, Peternel S, Komel R, Zupancic T (2012) Inclusion bodies as potential vehicles for recombinant protein delivery into epithelial cells. Microb Cell Factories 11:67
Martínez-Miguel M, Kyvik AR, Ernst LM, Martínez-Moreno A, Cano-Garrido O, Garcia-Fruitós E et al (2020) Stable anchoring of bacteria-based protein nanoparticles for surface enhanced cell guidance. J Mater Chem B 8(23):5080–5088
Mestrom L, Marsden SR, McMillan DGG, Schoevaart R, Hagedoorn P-L, Hanefeld U (2020) Comparison of enzymes immobilised on immobeads and inclusion bodies: a case study of a trehalose transferase. ChemCatChem 12(12):3249–3256
Morell M, Bravo R, Espargaró A, Sisquella X, Avilés FX, Fernàndez-Busquets X et al (2008) Inclusion bodies: specificity in their aggregation process and amyloid-like structure. Biochim Biophys Acta 1783(10):1815–1825
Nahálka J (2008) Physiological aggregation of maltodextrin phosphorylase from Pyrococcus furiosus and its application in a process of batch starch degradation to alpha-D-glucose-1-phosphate. J Ind Microbiol Biotechnol 35(4):219–223
Nahálka J, Gemeiner P, Bucko M, Wang PG (2006) Bioenergy beads: a tool for regeneration of ATP/NTP in biocatalytic synthesis. Artif Cells Blood Substit Immobil Biotechnol 34(5):515–521
Nahalka J, Nidetzky B (2007) Fusion to a pull-down domain: a novel approach of producing Trigonopsis variabilis D-amino acid oxidase as insoluble enzyme aggregates. Biotechnol Bioeng 97(3):454–461
Nahálka J, Vikartovská A, Hrabárová E (2008) A crosslinked inclusion body process for sialic acid synthesis. J Biotechnol 134(1–2):146–153
Natalello A, Doglia SM (2015) Insoluble protein assemblies characterized by Fourier transform infrared spectroscopy. Methods Mol Biol 1258:347–369
Panda AK (2003) Bioprocessing of therapeutic proteins from the inclusion bodies of Escherichia coli. Adv Biochem Eng Biotechnol 85:43–93
Park AR, Jang SW, Kim JS, Park YG, Koo BS, Lee HC (2018) Efficient recovery of recombinant CRM197 expressed as inclusion bodies in E. coli. PLoS One 13(7):e0201060
Park SY, Park SH, Choi SK (2012) Active inclusion body formation using Paenibacillus polymyxa PoxB as a fusion partner in Escherichia coli. Anal Biochem 426(1):63–65
Pesarrodona M, Jauset T, Díaz-Riascos ZV, Sánchez-Chardi A, Beaulieu ME, Seras-Franzoso J et al (2019) Targeting antitumoral proteins to breast cancer by local administration of functional inclusion bodies. Adv Sci 6(18):1900849
Peternel S, Grdadolnik J, Gaberc-Porekar V, Komel R (2008) Engineering inclusion bodies for non denaturing extraction of functional proteins. Microb Cell Factories 7:34
Peternel S, Komel R (2011) Active protein aggregates produced in Escherichia coli. Int J Mol Sci 12(11):8275–8287
Qi X, Sun Y, Xiong S (2015) A single freeze-thawing cycle for highly efficient solubilization of inclusion body proteins and its refolding into bioactive form. Microb Cell Factories 14:24
Rinas U, Bailey JE (1992) Protein compositional analysis of inclusion bodies produced in recombinant Escherichia coli. Appl Microbiol Biotechnol 37(5):609–614
Rinas U, Boone TC, Bailey JE (1993) Characterization of inclusion bodies in recombinant Escherichia coli producing high levels of porcine somatotropin. J Biotechnol 28(2–3):313–320
Rinas U, Garcia-Fruitós E, Corchero JL, Vázquez E, Seras-Franzoso J, Villaverde A (2017) Bacterial inclusion bodies: discovering their better half. Trends Biochem Sci 42(9):726–737
Roca-Pinilla R, Fortuna S, Natalello A, Sánchez-Chardi A, Ami D, Arís A et al (2020a) Exploring the use of leucine zippers for the generation of a new class of inclusion bodies for pharma and biotechnological applications. Microb Cell Factories 19(1):175
Roca-Pinilla R, López-Cano A, Saubi C, Garcia-Fruitós E, Arís A (2020b) A new generation of recombinant polypeptides combines multiple protein domains for effective antimicrobial activity. Microb Cell Factories 19(1):122
Rueda F, Cano-Garrido O, Mamat U, Wilke K, Seras-Franzoso J, García-Fruitós E et al (2014) Production of functional inclusion bodies in endotoxin-free Escherichia coli. Appl Microbiol Biotechnol 98(22):9229–9238
Rueda F, Gasser B, Sanchez-Chardi A, Roldan M, Villegas S, Puxbaum V et al (2016) Functional inclusion bodies produced in the yeast Pichia pastoris. Microb Cell Factories 15:1–12
Sanagavarapu K, Nüske E, Nasir I, Meisl G, Immink JN, Sormanni P et al (2019) A method of predicting the in vitro fibril formation propensity of Aβ40 mutants based on their inclusion body levels in E. coli. Sci Rep 9(1):3680
Sans C, García-Fruitós E, Ferraz RM, González-Montalbán N, Rinas U, López-Santín J et al (2012) Inclusion bodies of fuculose-1-phosphate aldolase as stable and reusable biocatalysts. Biotechnol Prog 28(2):421–427
Sarker A, Rathore AS, Gupta RD (2019) Evaluation of scFv protein recovery from E. coli by in vitro refolding and mild solubilization process. Microb Cell Factories 18(1):5
Seras-Franzoso J, Peebo K, Luis Corchero J, Tsimbouri PM, Unzueta U, Rinas U et al (2013a) A nanostructured bacterial bioscaffold for the sustained bottom-up delivery of protein drugs. Nanomedicine 8(10):1587–1599
Seras-Franzoso J, Steurer C, Roldán M, Vendrell M, Vidaurre-Agut C, Tarruella A et al (2013b) Functionalization of 3D scaffolds with protein-releasing biomaterials for intracellular delivery. J Control Release 171(1):63–72
Seras-Franzoso J, Peternel S, Cano-Garrido O, Villaverde A, García-Fruitós E (2015) Bacterial inclusion body purification. Methods Mol Biol 1258:293–305
Seras-Franzoso J, Sanchez-Chardi A, Garcia-Fruitos E, Vazquez E, Villaverde A (2016) Cellular uptake and intracellular fate of protein releasing bacterial amyloids in mammalian cells. Soft Matter 12(14):3451–3460
Seras-Franzoso J, Tsimbouri PM, Burgess KV, Unzueta U, Garcia-Fruitos E, Vazquez E et al (2014) Topographically targeted osteogenesis of mesenchymal stem cells stimulated by inclusion bodies attached to polycaprolactone surfaces. Nanomedicine 9(2):207–220
Singh A, Upadhyay V, Upadhyay AK, Singh SM, Panda AK (2015a) Protein recovery from inclusion bodies of Escherichia coli using mild solubilization process. Microb Cell Factories 14:41
Singh A, Upadhyay V, Panda AK (2015b) Solubilization and refolding of inclusion body proteins. Methods Mol Biol 1258:283–291
Singh SM, Sharma A, Upadhyay AK, Singh A, Garg LC, Panda AK (2012) Solubilization of inclusion body proteins using n-propanol and its refolding into bioactive form. Protein Expr Purif 81(1):75–82
Singhvi P, Saneja A, Srichandan S, Panda AK (2020) Bacterial inclusion bodies: a treasure trove of bioactive proteins. Trends Biotechnol 38(5):474–486
Song AA, In LLA, Lim SHE, Rahim RA (2017) A review on Lactococcus lactis: from food to factory. Microb Cell Factories 16(1):55
St John RJ, Carpenter JF, Balny C, Randolph TW (2001) High pressure refolding of recombinant human growth hormone from insoluble aggregates. Structural transformations, kinetic barriers, and energetics. J Biol Chem 276(50):46856–46863
Stamm A, Strauß S, Vogt P, Scheper T, Pepelanova I (2018) Positive in vitro wound healing effects of functional inclusion bodies of a lipoxygenase from the Mexican axolotl. Microb Cell Factories 17(1):57
Tokatlidis K, Dhurjati P, Millet J, Béguin P, Aubert JP (1991) High activity of inclusion bodies formed in Escherichia coli overproducing clostridium thermocellum endoglucanase D. FEBS Lett 282(1):205–208
Torrealba D, Seras-Franzoso J, Mamat U, Wilke K, Villaverde A, Roher N et al (2016a) Complex particulate biomaterials as immunostimulant-delivery platforms. PLoS One 11(10):e0164073
Torrealba D, Parra D, Seras-Franzoso J, Vallejos-Vidal E, Yero D, Gibert I et al (2016b) Nanostructured recombinant cytokines: a highly stable alternative to short-lived prophylactics. Biomaterials 107:102–114
Unzueta U, Cespedes MV, Sala R, Alamo P, Sánchez-Chardi A, Pesarrodona M et al (2018) Release of targeted protein nanoparticles from functional bacterial amyloids: a death star-like approach. J Control Release 279:29–39
Unzueta U, Roldán M, Pesarrodona M, Benitez R, Sánchez-Chardi A, Conchillo-Solé O et al (2020) Self-assembling as regular nanoparticles dramatically minimizes photobleaching of tumour-targeted GFP. Acta Biomater 103:272–280
Upadhyay V, Singh A, Jha D, Panda AK (2016) Recovery of bioactive protein from bacterial inclusion bodies using trifluoroethanol as solubilization agent. Microb Cell Factories 15:100
Valax P, Georgiou G (1993) Molecular characterization of beta-lactamase inclusion bodies produced in Escherichia coli. 1. Composition Biotechnol Prog 9(5):539–547
Vázquez E, Corchero JL, Burgueño JF, Seras-Franzoso J, Kosoy A, Bosser R et al (2012) Functional inclusion bodies produced in bacteria as naturally occurring nanopills for advanced cell therapies. Adv Mater 24(13):1742–1747
Vera A, González-Montalbán N, Arís A, Villaverde A (2007) The conformational quality of insoluble recombinant proteins is enhanced at low growth temperatures. Biotechnol Bioeng 96(6):1101–1106
Villaverde A, Corchero JL, Seras-Franzoso J, Garcia-Fruitos E (2015) Functional protein aggregates: just the tip of the iceberg. Nanomedicine 10(18):2881–2891
Wang L (2009) Towards revealing the structure of bacterial inclusion bodies. Prion 3(3):139–145
Wang L, Maji SK, Sawaya MR, Eisenberg D, Riek R (2008) Bacterial inclusion bodies contain amyloid-like structure. PLoS Biol 6(8):e195
Wang X, Zhou B, Hu W, Zhao Q, Lin Z (2015a) Formation of active inclusion bodies induced by hydrophobic self-assembling peptide GFIL8. Microb Cell Factories 14:88
Wang M, Qi W, Su R, He Z (2015b) Advances in carrier-bound and carrier-free immobilized nanobiocatalysts. Chem Eng Sci 135:21–32
Whitfield JR, Beaulieu ME, Soucek L (2017) Strategies to inhibit myc and their clinical applicability. Front Cell Dev Biol 5:10
Worrall DM, Goss NH (1989) The formation of biologically active beta-galactosidase inclusion bodies in Escherichia coli. Aust J Biotechnol 3(1):28–32
Wu W, Xing L, Zhou B, Lin Z (2011) Active protein aggregates induced by terminally attached self-assembling peptide ELK16 in Escherichia coli. Microb Cell Factories 10:9
Zhou B, Xing L, Wu W, Zhang XE, Lin Z (2012) Small surfactant-like peptides can drive soluble proteins into active aggregates. Microb Cell Factories 11:10
Acknowledgments
The authors acknowledge the financial support granted to A.A and E.G.F from Instituto Nacional de Investigación y Tecnología Agraria y Alimentaria, Spain (RTA2015-00064-C02-01) and Ministerio de Ciencia e Innovación (MICINN), Spain (grant PID2019-107298RB-C21). This work was also supported by Marató de TV3 foundation (201812-30-31-32-33). We are also indebted to CERCA Programme (Generalitat de Catalunya) and European Social Fund for supporting our research. R.B.F. and A.L.C received a predoctoral fellowship from INIA (FPI-INIA, MINECO) and Generalitat de Catalunya (FI-AGAUR), respectively.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this chapter
Cite this chapter
Baltà-Foix, R., Roca-Pinilla, R., López-Cano, A., Gifre-Renom, L., Arís, A., Garcia-Fruitós, E. (2022). Functional Inclusion Bodies. In: Rehm, B.H.A., Wibowo, D. (eds) Microbial Production of High-Value Products. Microbiology Monographs, vol 37. Springer, Cham. https://doi.org/10.1007/978-3-031-06600-9_11
Download citation
DOI: https://doi.org/10.1007/978-3-031-06600-9_11
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-06599-6
Online ISBN: 978-3-031-06600-9
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)