Abstract
Climate change causes diverse impacts on agricultural ecosystems and crop productivities. Being exposed to the external environment and sessile in nature, crop plants are thus most vulnerable to the ever-changing climate conditions. Variations in the external environment often result in different abiotic stresses, like heat and cold/chilling stresses due to temperature fluctuations, severe drought periods, waterlogging, and floods owing to variations in rainfall patterns. In this regard, scientists over the globe are striving to develop climate-resilient crops through integration of classical and modern crop breeding techniques in order to ensure food security for ever-increasing population around the world. In this chapter, we summarized the general impacts of climate change on food crops, importance of conventional breeding and novel approaches, and roles of multi-omics techniques, i.e., transcriptomics, genomics, metabolomics, phenomics, and ionomics for crop improvement under climate change scenario. Furthermore, we also elaborated on the recent use of multi-omics approaches to induce abiotic stress tolerance in crop plants. Overall, interactive use of omics approaches with systems biology may be favorable for improving crop resilience and productivity under climate change on a sustainable basis.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
3.1 Introduction
Variations in climatic conditions generally disturb the agro-ecosystems and food supply chain (Hasegawa et al. 2018). As agriculture significantly depends on the prevailing climatic conditions, thus crops often undergo external environmental and/or abiotic stresses due to climate variations (Hussain et al. 2018). Crop plants may suffer individual and concurrent effects of many abiotic stresses, such as salinity, chilling, drought, waterlogging, heavy metals, and temperature fluxes, which extensively impede the growth and development and reduce agricultural productivity over the globe (Ashraf et al. 2015; Raza et al. 2020; Ashraf et al. 2017a,b; Anjum et al. 2017). Moreover, global warming due to emissions of greenhouse gases also resulted in an increase in temperature and drought conditions that also threaten crop productivity, e.g., 20–30% yield loss was noticed in two major cereals, i.e., wheat and maize (Wang et al. 2019). Previously, traditional plant breeding techniques have produced several widely used high-yielding crop cultivars throughout the world for decades; however, requirements of longer time span on a variety of development and breeding cycles are the major drawbacks regarding implementation of such techniques. In addition, it is a roadblock in plant breeders’ ability to respond quickly to rising food supply demands under changing climate scenarios (Lenaerts et al. 2019).
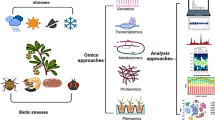
In the current era, the focus of plant breeding is to develop high-yielding crop varieties for the traits under consideration with minimal interventions during crop growth period. This has been accomplished by adopting modern biotechnological techniques which helped introduce genes with novel properties by breaking the inter-species and inter-genus barriers (Moose and Mumm 2008; Shiferaw et al. 2013). This all aims to furnish crops at a faster rate with the properties which their ancestors possessed to tolerate adverse environmental conditions. This does not mean that conventional breeding is ruled out from the agricultural system due to its inappropriateness. Actually, the current century is based on techniques to realize maximum output from the existing agricultural system due to huge food demand for rapidly increasing population from the existing set of agricultural land (Moose and Mumm 2008; Voss-Fels et al. 2019). The said task that increased production from limited land area with better adaptability for adverse environmental conditions could not be accomplished by depending merely upon conventional breeding. Although conventional breeding techniques are classical, most promising with least controversies associated with them, but, the time required to get the objective, blindness to ascertain the intrusion of target trait and transfer of only a fraction of genetic variations among crossing species, hindered its efficacy (Swamy and Sarla 2008). These factors majorly paved the way in the adoption of modern plant breeding and multi-omics approaches to address the debate of “sustainable agricultural intensification” by advocating ever-increasing global food demand under climate change conditions (Tester and Langridge 2010; Cheema 2018). The general impacts of abiotic stresses due to changing climate and importance of multi-omics approaches for crop improvement have been depicted in Fig. 3.1.
3.2 Impacts of Changing Climate on Food Crops
Crop production is vulnerable to climate change, which linked to an increase in carbon dioxide levels, shifting rainfall patterns, and rising temperatures (Mo et al. 2017). Enhancing agricultural output in one way while also addressing climate change risks on the other war is a difficult task. As a result, we need to pay greater attention to adaptation and mitigation. Climate change has substantial impacts on food crops; therefore, changes in policies, as well as the assistance of national adaptation funds and other resources, are required to alleviate negative influences of climate change on crop plants especially food crops. The growth responses, yield variability, and crop performance of vegetables, cereals, and pulses have been discussed below:
3.2.1 Vegetables
Vegetables are the best source of micronutrients and provide more revenue to smallholder farmers than staple crops (Rojas et al. 2013). Among vegetables tomato, cabbage, onion, hot pepper, and eggplant are widely cultivated (Julkowska and Testerink 2015). In Asia, the highest yields are generally obtained in the east region, where the temperature is mostly temperate or sub-temperate. Because most vegetables prefer lower temperatures, thus the production is the lowest in hot and humid lowlands of Southeast Asia (Ali 2000). Average tomato yields in sub-Saharan Africa (excluding South Africa) and tropical Asia are only approximately 10–12 MT/hectare, well below those in temperate areas. The intensity of environmental stress placed on vegetable crops will be influenced by climate change. For instance, rising temperatures, decreasing irrigation water supply, floods, and salt will be key stumbling blocks to maintaining and growing vegetable output. Temperature fluctuations hider vegetable growth and productivity, e.g., high temperatures are common in the tropics during the growing season, and as the climate changes, crops in this region would be susceptible to high-temperature stress. Temperatures are rising, according to climatic trends in tomato-growing areas, and the intensity and regularity of above-optimal temperature events will rise in the future decades (Tao et al. 2008). Temperature, alone or in combination with other environmental variables, has a significant impact on tomato vegetative and reproductive activities (Makeen et al. 2021). Plants’ metabolic processes that are essential for regular cell and photosynthetic activity are disrupted by high temperatures (Ergin et al. 2021). High temperatures can impair tomato output by reducing fruit set and producing smaller, lower-quality fruits (Solankey et al. 2021). Moreover, high-temperature stress shortens the time of photo-assimilation, triggers the developmental stages in determinate vegetables, and hampers fruit-set of fruiting vegetables (Bisbis et al. 2018).
3.2.2 Cereals
3.2.2.1 Maize
Temperatures are predicted to increase 1.5–4 °C globally during the next century (Vescio et al. 2021). Temperatures that are higher than the optimum for plant development are thought to be harmful for successful crop production (Battisti and Naylor 2009). Warmer temperatures and greater vapor pressure deficit (VPD) reduced yield of maize by 8.5 ± 1.4% and 12.9 ± 1.8%, respectively, as compared to the normal conditions, while applying both variables at the same time reduced yield by 24.2 ± 2.8%. Higher CO2 concentrations, on the other hand, increased maize production by 17.2 ± 3.5%. When increasing CO2 levels were combined with warmer temperatures or greater VPD, this rise in yield reduced, and the combination of the three climatic impacts (Temp, VPD, and CO2) largely cancelled each other out, resulting in a lesser drop in yield of −4 ± 3.4% (Resop et al. 2016). The onset of drought conditions impaired the normal metabolism, photosystems, and chlorophyll pigments in maize (Efeoğlu et al. 2009). Drought conditions hampered the early establishment of maize, whereas seed priming with Zn and Se enhanced germination and antioxidant activities (Nawaz et al. 2021). Moreover, Dong et al. (2021) reported that heat tolerance in long-season than short-season maize lines is subjected to more kernel formation by sustaining comparatively stable anthesis-silking interval. Based on meteorological and phenological data for 30 years, i.e., 1981–2010, it was observed that the vegetative growth interval has extended in spring-planted and inter-cropping maize and shortened in the summer season, whereas reproductive period has extended in spring- and summer-planted maize (Liu et al. 2021a, b). Overall, quantification of the changes in crop phenological development stages owing to climate change and anthropogenic management activities may assist to design promising climate change adaptation strategies in crop plants.
3.2.2.2 Rice
Rice, a staple crop in Asia, nourishes roughly 557 million people and is critical for maintaining food security, promoting rural employment, and generating export money (Xu et al. 2012; Le 2016). Rice yield growth in Asia declined at the start of the twenty-first century, and it is unable to keep up with the regional rapid population increase, resulting in high demands, supply shortages, and high prices. Rice is typically very susceptible to climatic fluctuations; thus recent trends of climate change would have serious impacts on its growth and productivity (Maruyama et al. 2014a, b; Kong et al. 2017). Future climate change is unknown, which would exacerbate the food security issues in Asia and other vulnerable regions (Palazzo et al. 2017; Carpena 2019).
Climatic change is responsible for a third of worldwide rice, maize, wheat, and soybean production variability, i.e., 32–39% (Ray et al. 2015), whereas the effects on rice yield in various locations have been estimated using external factors that include temperature, CO2 fertilization, rainfall, and other climatic aspects. Without accounting for CO2 fertilization effects, predictions using an “InfoCrop rice model” under the A1b, A2, B1, and B2 scenarios suggested that rice yields in India would decline by 4% in 2020, 7% in 2050, and 10% in 2080 (Soora et al. 2013). Rice yields in China will reduce by 6.1 to 18.6%, 13.5 to 31.9%, and 23.6 to 40.2%, respectively, if world temperatures rise by 1, 2, or 3°C (Tao et al. 2008). Under the representative concentration route (RCP) 8.5, Cambodia would see the greatest loss in rice yields in Southeast Asia (about 45% in the 2080s) (Chun et al. 2016). Climate change would have a detrimental influence on rice production. Other researches, on the other hand, show that climate change has a favorable influence on rice production. For example, according to the A2a, A2b, A2c, and B2a scenarios, rice yields in Bangladesh would improve by 7.58%, 8.31%, 4.57%, and 1.67%, respectively.
Variations in rice output in major rice-producing nations such as Bangladesh, China, India, and Myanmar (BCIM) were observed as a result of climate change. The BCIM countries have been identified as the world’s most climate-vulnerable areas because they account for 60.63% of world rice production and 59.74% of global rice consumption; these four nations play critical roles in the global rice trade pattern. A study conducted in Nepal reported that climate change has a significant impact on rice production, with an increase in severe precipitation having a negative impact. According to the extreme climate condition model, a 1% increase in days with extreme rainfall variation (i.e., three standard deviations above or below the long-term average) reduces rice output by 0.28% or 5.34 kg per family (Baldos et al. 2019). However, there is little indication that the rise in severe temperature days has had a similar impact. The average climate condition model, on the other hand, shows that a rise in long-term average monsoon temperature has a considerable negative influence on rice production. They further reported that a 1 °C rise in average summer temperature reduces rice output by 0.48% (4183 kg) per season; nevertheless, with increasing average temperatures, this impact diminishes. However, no evidence was found about the linkage of rise in average monsoon precipitation with increased rice output (Janes et al. 2019).
3.2.2.3 Wheat
Climate change may cause prolonged dry spells or more intense heat waves in some areas, thus severely reducing agricultural productivity by inducing water and temperature stress, especially during critical crop growth stages (Rounsevell et al. 1999). Dry land areas, such as dry and semi-arid parts of South Asia, are particularly susceptible to climate change, especially given their current water scarcities and higher temperatures approaching tolerance limits (Liu et al. 2019). Land degradation and restricted water supplies have hampered current agricultural output and put the food security of most of the countries in this region in danger.
Monsoon patterns, driven by monsoon lows and depressions in the Arabian Sea and the Bay of Bengal, dominate the weather system in Swat (Pakistan). The average annual temperature in Chitral district is 16 °C, and this area receives 451 mm of total annual rainfall, with more than 60% of that falling during the wheat growing season (October–April). The average annual temperature in Swat district is 19 °C and it receives 966 mm of rain annually, with 41% of that falling during the wheat growing season (October–March). These two districts were chosen to enable a comparison of the effects of climate change on wheat production at various elevations and in different weather systems. Wheat cultivated in the highlands and foothills is generally utilized for both food and fodder. Wheat is cultivated for food up to a height of around 1500 meters. On the other hand, wheat is used as fodder above this temperature since lower temperatures prevent the crop from ripening into edible grain (Hussain 2003). For temperature rises of 1.5 °C and 3.0 °C, yields in Chitral district (Pakistan) are anticipated to increase by 14% and 23%, respectively (Hussain and Mudasser 2007). Furthermore, direct and indirect impacts of climate change on wheat productivity were assessed using crop models by Daloz et al. (2021) and reported that the direct impacts (via fluctuations in temperature and precipitation) caused –1 to –8% wheat yield loss, whereas indirect impacts (via changes in water availability) led to –4% to –36% yield loss in wheat. Moreover, climate change conditions were found to substantially affect the phenology and yield of wheat, while these effects can be minimized by some agronomic strategies by modifying sowing times and irrigation scheduling (Azmat et al. 2021). Rise in temperature could enhance water deficit and the chances of drought in winter wheat (Zhang et al. 2021a, b). Hence, the uncertainty and complexity of climate change, it is important to modify agronomic, crop management, and breeding strategies to induce the crop tolerance and resilience in food crops to cope against deleterious effects of climate change.
3.2.3 Pulses
Climate change has been one of the most widely debated topics in recent decades, owing to its direct influence on food supply and security. Changes in the statistical distribution of weather patterns have a significant influence on agricultural crops, especially rain-fed crops such as pulses (Gorim and Vandenberg 2017). Pulse crops suffer from forced maturity and reduced yields as a result of very high temperatures throughout the growing season, particularly during flowering and pod development. Drought, flood, excessive salinity, soil acidification, waterlogging, and other abiotic stressors are made easier by variations in rainfall patterns connected with humidity and their associated consequences (Basu et al. 2009). The impacts of abiotic stresses induced by changing climatic conditions on vegetables, cereals, and pulses have been presented in Table 3.1.
3.3 Conventional Breeding vs Novel Approaches for Crop Improvement
3.3.1 Modern Breeding Approaches for Crop Improvement
Crop breeding has been boosted by the addition of mutagenesis techniques which further expanded on a larger scale with the emergence of recombinant DNA technologies, both contributed in bio-diversification of crop plants to a great extent (Sammina et al. 2021; Wang et al. 2012). Genetic alteration of crop plant for one or multiple genes by recombinant DNA technology relies on Agrobacterium-mediated transformation. This technique which remained in use since the 1990s has marked several success stories in the production of major food and oil crops with improved resistance to biotic and abiotic environmental conditions. These crops include wheat, rice, maize, soybean, cotton, rapeseed, sunflower, etc., grown worldwide as major transgenic crops with improved yield and least use of agrochemicals (Young et al. 2019). However, besides the immense contribution of transgene technology in crop improvement, in the recent decade, attention is paid to precise editing of plant genome. Introduction of novel genome sequencing, genotyping, and genome editing tools has revolutionized plant breeding worldwide. These methods allow for targeted and precise DNA alteration/modification in a short time and are considered to have a greater impact on plant genetics and breeding program (Chen et al. 2019; Li et al. 2019; Young et al. 2019). These novel technologies include next-generation sequencing (NGS) (Varshney et al. 2009), high-throughput marker genotyping technology (Kumar et al. 2012; Mammadov et al. 2012, Chen et al. 2015; Singh et al. 2016), and most recently cutting-edge technique called clustered regularly interspaced short palindromic repeats/CRISPR-associated protein 9 (CRISPR/Cas9) (Mao et al. 2013; Zhang et al. 2016; Lee et al. 2019). All these technologies are based on detection of genetic variation within a gene pool.
3.3.2 Genome Applications by Next-Generation Sequencing (NGS) Technology
Next-generation sequencing (NGS) in the current era is meant for a platform which provides nucleotide information of millions of sequence reads in a cost-effective manner within a short time (Varshney et al. 2009). Three major approaches are currently being used under NGS which includes applied biosystems such as SOLiD, genome sequencer FLX (Roche/454), and Illumina genome analyzer each having their own mechanism, benefits, and drawbacks (Gupta 2008; Mardis 2008; Shendure and Ji 2008). All three approaches are in use with differential impacts and effects based on objective, target species, and resource availability. The common among all is to generate nucleotide sequence of millions of reads within a short time with a variable potential. The genome sequence obtained has revolutionized the field of molecular marker development and marker-assisted selection (MAS) in plant breeding programs. Several success stories have been marked in multiple plant species, e.g., soybean (Hyten et al. 2008), maize (Barbazuk et al. 2007), sorghum (Paterson et al. 2009), rice (Goff et al. 2002), potato (Diambra 2011), apple (Velasco et al. 2010), etc. Apart from its role in rapid molecular marker development, other applications of NGS include association and comparative mapping, de novo sequencing, polymorphism detection, whole-genome analysis, transcriptome expression analysis, evolutionary biology, and population genetics (Varshney et al. 2009).
Decreasing the cost of sequencing with the passage of time is promoting its use for the crops having large genome size, polyploids, and underutilized crops which otherwise are not being paid attention before. The NGS data obtained could be manipulated in various ways by the use of data analysis tools for various applications. These applications could be structural/functional genomics, DNA-protein interaction, polygenetics, meta-genomics, and single-cell genomics, and the knowledge obtained from these studies could be utilized to decipher the complex network of genes with various biological functions, like never before. In the future, NGS technology has the potential to boost the crop improvement programs to its next level with more number of sequenced plant genomes (Kumar et al. 2012; Mammadov et al. 2012; Chen et al. 2015; Singh et al. 2016).
3.3.3 High-Throughput Marker Genotyping Technology Role in Crop Improvement
Genome sequencing paved the way to the development of molecular markers in high-throughput manner as per the sequencing data obtained and analyzed by the use of genome analysis tools. The success of marker development also depends upon its cost-effectiveness and high-throughput development with enhanced accuracy within a short period of time. Unlike before where few reads were obtained and analyzed for marker presence, NGS made it possible to develop thousands of molecular markers within a short time. Molecular markers exhibit a variety of basic and applied applications in crop improvement programs. These applications associated with successful and presence of abundant identifiable molecular markers include linkage map development, quantitative trait loci (QTLs) mapping in population, genome finger printing, marker-assisted selection (MAS), genetic diversity, and evolutionary analysis (Zalapa et al. 2012). Several types of molecular markers have been developed which include AFLP, SSR, ESTs, SNP, etc., each having variable merits, de-merits, uses, and benefits. A large number of SSRs (simple sequence repeats) and EST (expressed sequence tags) SSRs were developed in multiple crops by using NGS successfully (Kumari et al. 2013; Chen et al. 2015; Hao et al. 2015; Singh et al. 2016).
In addition, the SNP (single-nucleotide polymorphism) are found the most promising and innovative marker as it has the potential to detect the variation at the single-nucleotide level among the species (Kumar et al. 2012; Mammadov et al. 2012). For genotyping of this most promising marker, two assays are mostly used, i.e., (1) illumine golden gate assay and (2) whole-genome genotyping infinium assay (Rostoks et al. 2006; Hyten et al. 2008; Akhunov et al. 2009; Hyten et al. 2009). Both assays have different base mechanisms to get activated and the processes of development which is called high-throughput marker genotyping. This high-throughput marker genotyping is particularly important for diversity, characterization, and genome/association/comparative mapping (Rostoks et al. 2006; Hyten et al. 2009).
3.3.4 Genome Editing by CRISPR/Cas9 Approach
CRISPR/Cas9 is a nature-occurred gene editing tool and a high-throughput cutting-edge genomics technique having extensive applications in gene transformation, knockout mutation, monitoring of gene expression, and its regulation at the genomic and epigenomic level and in drug discovery (Bortesi et al. 2016; Fiaz et al. 2021). With the addition of CRISPR/Cas9 in the field of molecular biology, it is possible to manipulate or edit an organism’s genome in a precise and specific way, leading toward precision breeding. CRISPR/Cas9 is categorized into two main classes and six types: class I (types I, III, IV) and class II (types II, V, VI), with type II being the most widely used system in genome editing (Zhang et al. 2016; Lee et al. 2019). This system is discovered in bacterial and archaeal immune system in which CRISPR loci contained a cluster of consecutive repeats separated by spacer sequence. Other parts of the functional system include a Cas9 encoding operon and transcription setup. In response to immune system activation, these specific repeats get incorporated into bacterial genome by the activity of foreign invader or any viral genome (Jinek et al. 2012; Wang et al. 2012; Zhang et al. 2014). After infection, bacteria produce the same DNA which gets back incorporated into an invader virus along with one nuclease, called “Cas.” “Cas” then eventually acts as internal molecular scissors and cuts the invader/virus DNA into pieces. Based on its mechanism of action, it is also known as acquired immune defense system in prokaryotes against viruses (Zhang et al. 2014).
In the recent decade, the CRISPR/Cas9 system has two major targets with respect to crop breeding. Firstly, to find the efficient delivery method to enhance the potential of this system by eliminating off-target effects. Secondly, tracing and modification of key genes controlling growth and development in plants. This happens with the aid of genome sequencing, which is helpful in recognizing and locating specific sequences within the genome, which could be altered by the use of genome editors (Li et al. 2017). In a short period of time, CRISPR/Cas9 has a promising achievement in crop improvement.
Under natural conditions, plants are exposed to various biotic and abiotic stresses simultaneously, which results in reduction of yield and crop quality (Pandey et al. 2017; Ashraf et al. 2020; Hussain et al. 2020). Many of these stresses are under quantitative gene action, and it’s very difficult to figure out the exact mechanism of action and control. A remarkable achievement had been attained by creating resistant crop cultivars against diseases through knockout mutants by the use of the CRISPR/Cas9 system. Few success stories include Potyvirus in Arabidopsis, causing Turnip mosaic virus (TuMV) which is a major issue controlled by eIF(iso)4E; thus the knockout mutants using CRISPR/Cas9 have been generated which showed resistance to TuMV (Porter and Grills 2016). Likewise in watermelon, Clpsk1 gene editing showed enhanced resistance against Fusarium oxysporum f. sp. niveum (Zhang et al. 2020a, b), as well as Os8n3 and OsERF922 mutations exhibited resistance to Xanthomonas oryzae pv. oryzae (Xoo) and Magnaporthe oryzae, respectively (Wang et al. 2016; Kim et al. 2019). Other success examples include genes in tomato for SiMlo (Nekrasov et al. 2017) and SlJAZ2 (Ortigosa et al. 2019) and mlo in wheat (Diambra 2011; Hückelhoven and Panstruga 2011; Wang et al. 2014). All given genes are effective in imparting resistance against bacterial and fungal diseases which are located and manipulated by the use of CRISPR/Cas9 and resistance created against the pathogen successfully.
Common abiotic stresses which a plant copes with under natural conditions include drought, heat, salinity, and environmental pollution. Lots of structural and regulatory genes are found associated with a single stress condition and not a single gene which plays a dominant role in stress tolerance against any of these abiotic stress conditions (Zhang 2015; Zafar et al. 2020). Recently, ARGOS8 gene regulating ethylene production in maize is manipulated by the use of CRISPR/Cas9 and as a result enhanced drought tolerance observed in mutant plants under field conditions (Shi et al. 2017). Likewise in tomato knockout mutant for the gene SIAGAMOUS-like6 allowed the plant to tolerate heat stress successfully (Klap et al. 2017). Water use efficiency in tomato enhanced under drought and salinity stress in knockout mutant of ARF transcription factor (Bouzroud et al. 2020). The “G protein” mutants gs3 and dep1 in rice enhanced tolerance against salt stress (Cui et al. 2020), while ppa6 to alkaline stress positively (Wang, Wang et al. 2019). Thus, CRISPR/Cas9 is not only useful in creating knockout mutants but also useful in activation and regulation of gene expression (Meng et al. 2015; Wang et al. 2017; Paixão et al. 2019; Peng et al. 2020). Overall, to cope with fluctuating adverse environmental conditions is a complicated task and controlled by multiple genes or gene network with different magnitude of effects. In the current era, plant breeders had made a remarkable progress in a very short time by the use of the CRISPR/Cas9 approach which otherwise was nearly impossible. The quickly developed and adopted CRISPR/Cas9 approach provided a perfect tool to realize yield increments with improved tolerance to adverse environmental conditions. Current multiple omics, including whole-genome sequencing (DNA, RNA), further aid into the success stories in CRISPR/Cas9-based crop improvement.
3.4 Omics Approaches for Crop Improvement Under Climate Change Scenario
Plant molecular biology is concerned with the study of biological processes, their genetic regulation, and interactions with the environment (Yadav et al. 2020). In recent years, advances in omics technologies like genomics have made molecular biology research more efficient. As a result, the linkages between molecular machineries may be examined in great detail, allowing researchers to incorporate genes, proteins, and a wide range of other key regulatory components into their research (Muthamilarasan et al. 2019). The term “omics” refers to this large-scale research. The genomic, transcriptomic, proteomic, and metabolomic analyses are all important aspects of omics (Subramanian et al. 2020). These omics methods are often utilized in many agricultural and plant research areas. As technology develops, omics techniques have advanced dramatically in the previous decade (Anders et al. 2021). All these aspects of omics are discussed below:
3.4.1 Genomic Approach
In genomics, genes and whole genomes are analyzed in depth. It is now possible to read an organism’s whole genome using high-throughput sequencing practices such as next-generation sequencing (NGS) (Tanaka et al. 2020). The stress-related genes may now be predicted with the use of genome-wide association studies, or GWAS, and next-generation sequencing (NGS) (Saidi and Hajibarat 2020). The use of genomics techniques has grown in relevance for the discovery of genes involved in the tolerance of multiple abiotic stresses. Researchers have made great progress in the genomics era in scanning the genomes of several crop species during the last few years (Brozynska et al. 2016).
Furthermore, genomics is critical in identifying genetic diversity that underpins improved performance and breeding efficiency, therefore contributing to crop species’ genetic improvement. Quantitative trait locus (QTL) mapping is an outstanding mode to explore the genetically complex systems for abiotic stress tolerance in plants (Shen et al. 2018). Through QTL mapping, specific resistance loci were studied which indicates the presence of genes involved a development process of the plant during stress conditions (Adhikari et al. 2019). For instance, “B73” and “Mo17,” the two inbred maize lines commonly used in American maize germplasm, have varying levels of resistance to low temperatures during the seedling stage. In addition, seedling reaction to low-temperature exposure varied greatly among different inbred maize lines. Quantitative trait loci (QTLs) were identified to study this variation from the recombinant line population of maize. Two QTLs with several candidate genes were identified which also involved in hormone response and abiotic stress response (Qiu et al. 2020).
Different types of markers are involved in genome-wide analysis of the desired traits and use them in crop improvement during the breeding process as RFLP (restriction fragment length polymorphism), RAPD (rapid amplified polymorphic DNA), AFLP (amplified fragment length polymorphism), SAMPL (selectively amplified microsatellite polymorphic microsatellite loci), SRAP (sequence-related amplified polymorphism), and TRAP (target region amplification polymorphism) (Singh et al. 2021; Soltabayeva et al. 2021). However, with the introduction of next-generation sequencing (NGS) for large-scale genotyping and genomics-assisted breeding, SNPs have been the most preferred markers (Thabet et al. 2021).
Recently, Liu et al. (2020) identified 26 QTLs from rice cultivars of Vietnam genotyped with SNP markers and found that several genes present in QTLs were involved in salinity stress tolerance and also in signal transduction and hormonal response. These QTLs are further used in the breeding process for crop improvement. Alkali-tolerant gene loci were studied in two varieties of japonica rice under alkali treatment. Four candidate genes were identified on chromosomes 2 and 3 using next-generation sequencing technology (Sun et al. 2021). Furthermore, 302 wild landraces and modified soybean accessions were sequenced, and 230 selective sweeps and 162 copy number variations were found. By studying genome-wide correlations between novel sequences and domestication or improvement characteristics, 13 previously unidentified loci for agronomic variables were identified which include oil content, plant height, and pubescence shape. Using quantitative trait locus (QTL) data, 230 areas were studied, from which 96 correspond to previously discovered oil QTLs and 21 contain genes involved in fatty acid production (Zhou et al. 2015).
3.4.2 Transcriptomic Approach
It is possible to discover genes that respond to stress through the use of genome-wide transcriptome analysis methods. Through plant breeding and/or gene editing methods, these genes might serve as prospective targets for improving drought stress tolerance (Dudziak et al. 2019). Transcriptional responses to abiotic stress have been studied using a variety of techniques, including expressed sequence tags (ESTs), superSAGE, and microarrays. These findings show that abiotic stress alters the expression patterns of numerous genes involved in diverse biological processes (Alonge et al. 2020). It is possible to quantify transcriptome alterations underlying developmental transitions and stress responses using RNA sequencing at the genome scale (Ding et al. 2020).
It is via ESTs that new gene structures, such as expression systems, gene maps, and cDNA sequencing initiatives, may be discovered and built upon. The ESTs have long been regarded as the ideal method for revealing sequence-related information under adverse environmental circumstances (Khazaei et al. 2020). It has been shown that SAGE is a powerful tool for studying gene expression. The SAGE technique generates a tag that identifies a gene transcriptome product. SAGE is regarded as the most cost-effective and high-throughput method for mRNA isolation, cloning, and sequencing (Kumar et al. 2015). The “tag” used in the SAGE method is a short nucleotide sequence with a pointed head-to-head specific restriction enzyme. In other words, SAGE represents digitally expressed gene expression. Several SAGE applications are found in plants, such as pathogen-host interactions, plant responses to a variety of abiotic stressors, transcriptome profiling, and the metabolism of numerous hazardous chemicals (Jain et al. 2021).
To identify salt-responsive genes, a variety of methods are being used including cDNA microarrays, serial gene expression analysis, suppression subtractive hybridization, differential display RT-PCR, and complementary DNA-amplified fragment length polymorphism (cDNA-AFLP) (Bajwa and Khan 2021). The cDNA-AFLP’s specificity and sensitivity allow it to differentiate between homologous sequences and find even the low-expression genes. However, because of the great sensitivity of this technique, even uncommon transcripts can be identified (Kamyab et al. 2021). In addition, this is a less labor-intensive and very effective mRNA fingerprinting technique for identifying genes that exhibit differential expression in stressed plants/crops (Zhang et al. 2020a, b).
Gene expression may be observed using cDNA microarrays, which are an effective tool for evaluating the possible roles of many genes. Microarray cDNA studies of gene expression in different plants under a variety of abiotic factors have been well published (ul Qamar et al. 2020). RNA-sequencing is a new technology that makes use of and provides a new perspective on the transcriptome sequence by allowing total access to transcripts. As a result, RNA-seq may be used to replace a variety of approaches for quantifying transcripts, with the added benefit of increased sensitivity and the capacity to distinguish between related gene paralogs that differ by only a few nucleotides (Xu et al. 2021).
The thermotolerant germplasm of rice, SDWG005, was identified, and their anther structures that are involved in thermotolerance were studied. By transcriptome analysis, 3559 differentially expressed genes (DEGs) were identified and divided into categories according to their expressions. The major gene categories involved in thermoregulation include transcription factors and protein and nucleic acid metabolism-related genes (Liu et al. 2020). Differentially expressed genes (DEGs) were also identified from the wheat cultivar, TW004, under salt stress. Expression analysis of DEGs revealed that they are mostly involved in metabolic pathways that are related to salt stress. Different types of transcription factors involved in salt stress tolerance were also identified, such as MYB (Deng et al. 2020).
3.4.3 Proteomic and Metabolomic Approaches
In proteomics, the proteome in a complicated biological system under a particular circumstance is studied in detail. Plant proteomics offers a unique chance to study the process and production of various proteins in agricultural plants under distinct environmental conditions. Because of this, it’s possible to better understand gene regulation and metabolic pathways within plant species because it offers more information about the proteins’ interactions with one another (Jorrin-Novo 2020).
Because of the activation of stress-responsive pathways in response to biotic and non-biotic stressors, the plant proteome changes significantly. Heat-shock proteins, abundant proteins in late embryogenesis, kinases and phosphatases, redox enzymes, secondary metabolism enzymes, osmolyte biosynthetic enzymes, photosynthesis, and enzymatic reactive oxygen species (ROS) scavenger proteins are proteins that are known to play a function in the abiotic stress response (Kosová et al. 2018).
Salt-induced proteome changes were studied in barely landraces using the two-dimensional polyacrylamide gel electrophoresis (2D-PAGE) proteomic study. Results revealed that under salt stress, the accumulation of 11 salt-responsive proteins have differed between the two accessions, while 43 proteins were genotype-specific (Jardak et al. 2021). Proteomes of anthers of two soybean cultivars, heat-resistant and heat-sensitive, were studied using the iTRAQ approach. Results showed that most of the proteins related to heat tolerance are involved in carbohydrate and energy metabolism, protein synthesis and degradation, nitrogen assimilation, and ROS detoxification (Li et al. 2020).
In Arabidopsis, drought tolerance is controlled by jasmonate ZIM-domain protein 7 (JAZ7) utilizing TMT-based quantitative proteomics and targeted metabolomics (Meng et al. 2019). Arabidopsis AT-Hook-Like10 phosphorylation needed for stress growth control is elucidated using more sophisticated proteomics techniques, such as quantitative phosphoproteomics (Wong et al. 2019). To understand the function of genes in eukaryotic organisms, including plants, RNA silencing has proven to be a novel and potentially useful reverse genetics tool (Rajam 2020).
Metabolomics is a systematic research in which all the metabolites of a crop plant are identified and quantified. A plant’s biochemical response to climate change may be described using metabolomics, which allows us to understand the interconnections between different components in a biological system. Other functional genomics techniques, such as genomes, proteomics, and transcriptomics, can be utilized in combination with metabolomics (Razzaq et al. 2019).
Endogenous plant component proteomics and metabolomics study is becoming increasingly popular, and a strong technique like MS imaging is being used frequently to conduct this research. For protein analysis, mass spectrometry methods like matrix-assisted laser desorption ionization (MALDI) and electrospray ionization (ESI) have been frequently employed (Agarrwal and Nair 2020).
Primary and secondary metabolites make up the majority of the plant’s metabolome. The metabolic profiling of primary and secondary metabolites offers substantial information on the biochemical processes that take place during plant metabolism (Sung et al. 2015). Some plant secondary and primary metabolites are related to metabolic processes that are quite complicated. These metabolites may be effectively identified, measured, and studied using contemporary metabolomics methods such as gas chromatography-mass spectrometry (GC-MS), liquid chromatography-mass spectroscopy (LCMS), and non-destructive nuclear magnetic resonance (NMR) spectroscopy (Che-Othman et al. 2020).
As none of the single technique or instrument can be utilized to examine all of a metabolome’s metabolites, many distinct technologies are necessary for comprehensive coverage. The techniques involved in metabolomics study include mass spectrometry (MS) (Yadav et al. 2019), non-destructive nuclear magnetic resonance (NMR) spectroscopy (Cuperlovic-Culf et al. 2019), high-performance thin-layer chromatography (HPTLC) (D’Amelia et al. 2018), gas chromatography-mass spectrometry (GC-MS) (Chang et al. 2019), liquid chromatography-mass spectrometry (LC-MS) (Zhou et al. 2019), direct infusion mass spectrometry (DIMS), ultra-performance liquid chromatography (UPLC), and high-resolution mass spectrometry (HRMS) (Thomason et al. 2018).
Bernardo et al. (2019) studied the drought-induced changes in wheat secondary metabolism. By employing metabolomics, mycorrhizal inoculation was initiated in two wheat varieties at various phases of drought. According to the findings, arbuscular mycorrhizal fungi (AMF) colonization in a water-stressed environment modulates several molecules, most of which are sugars and lipids. Integrated transcriptome and metabolic profile analysis were used to determine the response of sunflower under drought stress. The findings indicated that increasing the expression level of photosynthesis in sunflower plants delays senescence by affecting many key metabolic pathways and candidate genes (Moschen et al. 2017).
3.4.4 Phenomic Approach
Breeding operations rely heavily on genetic and phenotypic connections. The study of a plant’s phenotypic or genotypic expression under certain environmental conditions is known as plant phenomics (Deery and Jones 2021). Due to the intricate biosynthetic mechanisms that regulate various plants’ abiotic stress resistance, phenotyping remains a difficult challenge under abiotic stress circumstances. The importance of phenotyping has emerged in the postgenomic age since crop improvement techniques such as GWAS, GS, MAS, and QTL mapping largely rely on high-throughput phenotyping (HTP) in crops (Ndlovu 2020). Modern phenomics methods make use of hyperspectral/multispectral cameras to collect hundreds of reflectance data points in a variety of settings and phases of crop development using discrete narrow bands. Agronomic characteristics may now be rapidly and precisely gathered using phenotyping technology (Atkinson et al. 2018). Furthermore, utilizing highly heritable secondary phenotypes closely related to the selection phenotypes, the main goal of a high-throughput phenotype (HTP) is to reduce the cost of data per plot and enhance the accuracy of the crop-growing season forecast (Matias et al. 2020). The proper mapping of genes/QTLs/alleles responsible for the trait of interest requires precise phenotyping of germplasms. Phenotyping using traditional methods is time-consuming and labor expensive (Bohra et al. 2021). Phenomics has proven successful in selecting field crops that can withstand abiotic stressors, such as drought (Jangra et al. 2021).
High-throughput, precise, and automated measurements of phenotypic information such as plant growth, architecture, and composition are all part of the process. This new technology has allowed for the collection and analysis of a huge amount of phenotyping data, which has previously been a barrier to functional genomics research and agricultural breeding (Pazhamala et al. 2021).
3.4.5 Ionomic Approach
Ionomics is the study of a plant’s whole mineral nutrient and traces element makeup. Additionally, the field of ionomics studies the molecular mechanisms that underlie plant elemental composition (Du et al. 2020). There have been several applications of ionomics in plant biology, ranging from studying plant growth and stress biology to meeting human food needs (Shakoor et al. 2016). To find out how metabolic, genetic, developmental, and environmental factors impact the elemental composition of target tissues and organs, ionome research looks at the entire organism’s chemical makeup (Pita-Barbosa et al. 2019).
Ionomics relies on mutant screening and natural variation to uncover genes and alleles that are crucial for elemental accumulation and variations in the ionomes of different genotypes (Hindt et al. 2017). Ionomics-based biomarkers in stress biology can help identify whether or not a plant has achieved a specific biochemical or physiological state under a variety of unfavorable environmental conditions. As a result, they can aid in the screening of plants that are more vulnerable to biotic and abiotic stressors, which is impossible with current high-throughput techniques. Ionomics has also identified genes that govern natural variation in the plant ionome and are important for stress tolerance (Ali et al. 2021).
The physiological and ionomic differences between Cd-tolerant and Cd-sensitive maize genotypes will be investigated using ionomic and physiological studies. Cd-tolerant cultivars had considerably greater proline, phenolics, and antioxidant accumulation, as well as enhanced rhizosphere and root cell sap absorption and translocation of N, P, K, Ca2+, Mg, Zn, and Fe when compared to Cd-sensitive cultivars. These nutrients are crucial in enhancing plant physiological, biochemical, and molecular responses to stress (Abbas et al. 2021). The role of silicon nanoparticles (SiNPs) in rice toxicity reduction was investigated. They observed that molecular priming with SiNPs decreased F accumulation and increased the amount of non-enzymatic antioxidants in rice grains, such as glutathione, flavonoids, anthocyanins, and phenols, leading to improved injuries and yield (Banerjee et al. 2021). A comparison of gluten-containing and gluten-free foods was studied. Ionomic analysis revealed that gluten-free foods contain a high amount of arsenic and mercury than the gluten-containing foods (Punshon and Jackson 2018).
Ionomics study reveals the susceptibility and resistance characteristics of two olive cultivars to discover target genes for the treatment of olive disease (D’Attoma et al. 2019). By increasing the content of non-enzymatic antioxidants in T. aestivum L., 60 mg/kg K substantially decreased the harmful effects of Cd (Yasin et al. 2018). Malus halliana was subjected to saline-alkali stress, which generated an ion imbalance and Na+ toxicity, and was then analyzed at the molecular level. An ionomics study demonstrated the importance of Ca2+-mediated signaling for maintaining Na+/K+ balance and decreasing stress-induced damage (Jia et al. 2020).
3.5 Abiotic Stress Tolerance in Crops: Integrating Omics Methods
Genomics is the study of the complex biological activities of a cell’s genetic material, the genome, using high-throughput technology. Functional genomics is critical for determining and proving the functions and roles of all genes discovered on the genome in cellular metabolism. Genomic analysis is widely used in agriculture to discover single-nucleotide polymorphisms (SNPs) using a comparative genomics method. The ultimate goal is to find SNPs or quantitative trait loci (QTLs) associated with a certain phenotype. SNPs are relatively prevalent in genome, however, and normally do not generate a discernible phenotype because of significant environmental stress on plants. This is particularly essential in the case of abiotic stress studies that need the monitoring of the wanted phenotypic expression, the tolerance of abiotic stress, to prevent changes in the intended trait under various environmental circumstances due to genotype x environment (G x E) interaction. As a result, genome-wide association studies (GWAS) and genotyping-by-sequencing (GBS) techniques are frequently used to find SNPs linked to abiotic stress tolerance in field crops (He et al. 2014; Unamba et al. 2015; Kang et al. 2016). For numerous field crops, both GWAS and GBS have been used to discover SNPs or QTLs that may be linked to tolerance to a variety of abiotic stressors (Table 3.2). More study is needed to enhance field crop tolerance to abiotic stressors, and more research is needed to assure agriculture’s long-term viability.
Transcriptomic is the result that all RNA molecules from the genome are sequenced, profiled, and counted, enabling comprehensive and/or comparative analysis in quantitative terms of gene expression. Gene expression or transcription, on the other hand, varies depending on the stage of development, tissue, and even cells within the same tissue. Transcriptomics, therefore, shows the metabolic activity of the plant at a specific moment in time, development stage, and tissues or on the environmental circumstances. Microarray platforms or next-generation sequencing (NGS) platforms are often used for transcriptomic research, and both provide high-throughput data for all transcripts depending on the available genome sequence in NGS and the sequence depth in microarrays (Unamba et al. 2015). Most work on transcriptome of abiotic stress tolerance reacts to a particular treatment, such as drought, salinity, cold, or heat. However, as previously indicated, the climate change models predict numerous stressors to impact field crop output. Thus, transcriptomic alterations following combination therapies became the principal technique in the study of transcriptomes. The knowledge of changes in gene expression patterns when they are primed is another perspective that may provide insights into the important components to increase the tolerance of the plant toward abiotic stress situations. Both of these approaches are required to get new insights into transcriptome experiments in order to improve field plant abiotic stress tolerance.
Proteomics uses high-performance technologies to identify and investigate proteins, potential post-translational modifications, structures, activities, and interactions with other proteins or cellular components in order to better understand cellular metabolism (Table 3.2). Proteomics research focused on using two-dimensional polyacrylamide gel electrophoresis (2D-PAGE) to examine the reactivity and/or tolerance of agricultural crops to abiotic stress, as well as the identification and characterization of the protein spots produced. In 2D-PAGE, the first dimension separates proteins based on their isoelectric point (pI), whereas the second dimension separates proteins based on their molecular weight (MW). In theory, no two proteins in a cell can have the same pI or MW, allowing for high-resolution separation with each spot on the gel representing a different protein. Interactions with other proteins, DNA, or RNA can be investigated using yeast two-hybrid systems or protein microarrays, as well as structural studies using nuclear magnetic resonance (NMR) or X-ray crystallography and subcellular localization using fluorescent proteins like EGFP or X-ray tomography (Moreno-Risueno et al. 2010). Nonetheless, proteomics has been widely utilized to better understand how many field crops, including rice, wheat, and maize, respond to abiotic stress situations (Table 3.2). Proteomics, rather than genomes and transcriptomics, appears to be more promising in terms of giving relevant background information for understanding and creating abiotic stress-tolerant field crops, and technical advances will definitely be necessary to attain these aims.
Metabolomics is the study of all metabolic byproducts, hormones, signaling molecules, and secondary metabolites present in a biological sample using methods such as mass spectroscopy (MS), gas chromatography-MS (GC-MS), liquid chromatography-MS (LC-MS), capillary electrophoresis-MS (CE-MS), and Fourier transform infrared spectroscopy (FTIR) (Oliver et al. 1998; Hong et al. 2016; Tian et al. 2016; Samota et al. 2017). Understanding complex phenotypic responses of plants to changes in environmental conditions requires metabolomic analysis. Metabolomics is perhaps the most difficult of the omics techniques, because the metabolome varies greatly depending on developmental stage and growth factors (Fuhrer and Zamboni 2015). As a result, it is reasonable to state that metabolomics has not been employed efficiently in investigations of field crop abiotic stress tolerance. Despite the fact that it does not yield definitive results like genomes, transcriptomics, or even proteomics, metabolomics provides a consistent metabolic quantitative trait locus (mQTL) that seems to be used as an indicator for breeding experiments to enhance agricultural crop tolerance to abiotic stress (Mochida and Shinozaki 2011). The mQTL can be used as metabolite markers in response to abiotic stress conditions, in addition to phenotypic studies such as GWAS, if the change in metabolite concentration influences the phenotypic change.
The field of omics technologies has broadened as a result of the introduction of high-performance technologies in many study fields. To integrate data from genomics, transcriptomics, proteomics, and metabolomics studies, systems biology techniques are employed (Fig. 3.2). The primary challenge is the integration of data from diverse omics techniques to abiotic stress with different plants and/or genotypes.
3.6 Conclusion and Future Perspectives
Climate change affects crop growth cycles, phenological development, crop productivities, and food quality. Variations in temperature, CO2 levels, and rainfall patterns have substantial impacts on agricultural crops; thus it could be a threat to future food security. Studying growth and yield responses of crop plants to different abiotic stresses owing to climate change is important in order to develop climate-resilient crops. Development and integration of modern genetics into classical breeding and development of new plant types and resistant cultivars through genetic manipulation are now being focused to meet the future food demands to reduce food security risk. Multi-omics approaches, i.e., genomics, proteomics, transcriptomics, phenomics, and ionomics, are playing a crucial role to identify a genetic basis of crop performance and improvement as well as stress tolerance mechanisms in crop plants under various environmental stresses. In the future, integration and combination of multi-omics approaches would be helpful to identify potential candidate genes and their pathways to develop models to mark agronomically important traits to enhance crop performance under changing climatic scenario through precision breeding.
References
Abbas S, Javed MT, Ali Q, Akram MS, Tanwir K, Ali S, Chaudhary HJ, Iqbal N (2021) Elucidating Cd-mediated distinct rhizospheric and in planta ionomic and physio-biochemical responses of two contrasting Zea mays L. cultivars. Physiol Mol Biol Plants 27(2):297–312
Adhikari L, Makaju SO, Missaoui AM (2019) QTL mapping of flowering time and biomass yield in tetraploid alfalfa (Medicago sativa L.). BMC Plant Biol 19(1):1–15
Agarrwal R, Nair S (2020) Metabolomics-assisted crop improvement. Advancement in crop improvement techniques. Elsevier, pp 263–274
Ahmad M, Ali Q, Hafeez M, Malik A (2021) Improvement for biotic and abiotic stress tolerance in crop plants. Biol & Clinic Sci Res J 2021(1)
Akhunov E, Nicolet C, Dvorak J (2009) Single nucleotide polymorphism genotyping in polyploid wheat with the Illumina GoldenGate assay. Theor Appl Genet 119(3):507–517
Akram NA, Shafiq F, Ashraf M (2017) Ascorbic acid-a potential oxidant scavenger and its role in plant development and abiotic stress tolerance. Front Plant Sci 8:613
Akula R, Ravishankar GA (2011) Influence of abiotic stress signals on secondary metabolites in plants. Plant Signal Behav 6(11):1720–1731
Ali M (2000) Dynamics of vegetable production, distribution and consumption in Asia. AVRDC
Ali S, Tyagi A, Bae H (2021) Ionomic Approaches for Discovery of Novel Stress-Resilient Genes in Plants. Int J Mol Sci 22(13):7182
Alonge M, Wang X, Benoit M, Soyk S, Pereira L, Zhang L, Suresh H, Ramakrishnan S, Maumus F, Ciren D (2020) Major impacts of widespread structural variation on gene expression and crop improvement in tomato. Cell 182(1):145–161
Amrutha V, Shanija S, Beena R, Nithya N, Jaslam M, Soni K, Viji M (2021) Population structure analysis and marker trait association in selected set of Indian tomato (Solanum lycopersicum L.) varieties under high temperature condition. Genet Resour Crop Evol:1–25
Anders S, Cowling W, Pareek A, Gupta KJ, Singla-Pareek SL, Foyer CH (2021) Gaining Acceptance of Novel Plant Breeding Technologies. Trends Plant Sci 26(6):575–587
Anjum SA, Ashraf U, Tanveer M, Khan I, Hussain S, Shahzad B, Zohaib A, Abbas F, Saleem MF, Ali I, Wang LC (2017) Drought induced changes in growth, osmolyte accumulation and antioxidant metabolism of three maize hybrids. Front Plant Sci 8:69. https://doi.org/10.3389/fpls.2017.00069
Ashraf U, Kanu AS, Mo Z, Hussain S, Anjum SA, Khan I, Abbas RN, Tang X (2015) Lead toxicity in rice: effects, mechanisms, and mitigation strategies—a mini review. Environ Sci Pollut Res 22(23):18318–18332
Ashraf U, Hussain S, Anjum SA, Abbas F, Tanveer M, Noor MA, Tang X (2017a) Alterations in growth, oxidative damage, and metal uptake of five aromatic rice cultivars under lead toxicity. Plant Physiol Biochem 115:461–471
Ashraf U, Kanu AS, Deng Q, Mo Z, Pan S, Tian H, Tang X (2017b) Lead (Pb) toxicity; physio-biochemical mechanisms, grain yield, quality, and Pb distribution proportions in scented rice. Front Plant Sci 8:259. https://doi.org/10.3389/fpls.2017.00259
Ashraf U, Mahmood MH, Hussain S, Abbas F, Anjum SA, Tang X (2020) Lead (Pb) distribution and accumulation in different plant parts and its associations with grain Pb contents in fragrant rice. Chemosphere 248:126003
Atkinson JA, Jackson RJ, Bentley AR, Ober E, Wells DM (2018) Field phenotyping for the future. Ann Plant Rev Online:719–736
Azmat M, Ilyas F, Sarwar A, Huggel C, Vaghefi SA, Hui T, Qamar MU, Bilal M, Ahmed Z (2021) Impacts of climate change on wheat phenology and yield in Indus basin, Pakistan. Sci Total Environ 790:148221
Bajwa HK, Khan H (2021) Genomics, proteomics and metabolic approaches against abiotic stress. Environmental Stress Physiology of Plants and Crop Productivity, p 183
Baldos UL, Hertel TW, Moore FC (2019) Understanding the spatial distribution of welfare impacts of global warming on agriculture and its drivers. Am J Agric Econ 101(5):1455–1472
Banerjee A, Singh A, Sudarshan M, Roychoudhury A (2021) Silicon nanoparticle-pulsing mitigates fluoride stress in rice by fine-tuning the ionomic and metabolomic balance and refining agronomic traits. Chemosphere 262:127826
Barbazuk WB, Emrich SJ, Chen HD, Li L, Schnable PS (2007) SNP discovery via 454 transcriptome sequencing. Plant J 51(5):910–918
Basu P, Ali M, Chaturvedi S (2009) Terminal heat stress adversely affects chickpea productivity in northern India—strategies to improve thermo tolerance in the crop under climate change. W3 Workshop Proceedings: Impact of Climate Change on Agriculture, International Society for Photogrammetry and Remote Sensing New Delhi
Batayeva D, Labaco B, Ye C, Li X, Usenbekov B, Rysbekova A, Dyuskalieva G, Vergara G, Reinke R, Leung H (2018) Genome-wide association study of seedling stage salinity tolerance in temperate japonica rice germplasm. BMC Genet 19(1):1–11
Battisti DS, Naylor RL (2009) Historical warnings of future food insecurity with unprecedented seasonal heat. Science 323(5911):240–244
Bernardo L, Carletti P, Badeck FW, Rizza F, Morcia C, Ghizzoni R, Rouphael Y, Colla G, Terzi V, Lucini L (2019) Metabolomic responses triggered by arbuscular mycorrhiza enhance tolerance to water stress in wheat cultivars. Plant Physiol Biochem 137:203–212
Bisbis MB, Gruda N, Blanke M (2018) Potential impacts of climate change on vegetable production and product quality–a review. J Clean Prod 170:1602–1620
Bohra A, Satheesh Naik S, Kumari A, Tiwari A, Joshi R (2021) Integrating Phenomics with Breeding for Climate-Smart Agriculture. In: Omics Technologies for Sustainable Agriculture and Global Food Security (Vol II). Springer, pp 1–24
Bortesi L, Zhu C, Zischewski J, Perez L, Bassié L, Nadi R, Forni G, Lade SB, Soto E, Jin X (2016) Patterns of CRISPR/Cas9 activity in plants, animals and microbes. Plant Biotechnol J 14(12):2203–2216
Bouzroud S, Gasparini K, Hu G, Barbosa MAM, Rosa BL, Fahr M, Bendaou N, Bouzayen M, Zsögön A, Smouni A (2020) Down regulation and loss of auxin response factor 4 function using CRISPR/Cas9 alters plant growth, stomatal function and improves tomato tolerance to salinity and osmotic stress. Genes 11(3):272
Brozynska M, Furtado A, Henry RJ (2016) Genomics of crop wild relatives: expanding the gene pool for crop improvement. Plant Biotechnol J 14(4):1070–1085
Cai C, Wu S, Niu E, Cheng C, Guo W (2017) Identification of genes related to salt stress tolerance using intron-length polymorphic markers, association mapping and virus-induced gene silencing in cotton. Sci Rep 7(1):1–15
Carpena F (2019) How do droughts impact household food consumption and nutritional intake? A study of rural India. World Dev 122:349–369
Chang J, Cheong BE, Natera S, Roessner U (2019) Morphological and metabolic responses to salt stress of rice (Oryza sativa L.) cultivars which differ in salinity tolerance. Plant Physiol Biochem 144:427–435
Chebrolu KK, Fritschi FB, Ye S, Krishnan HB, Smith JR, Gillman JD (2016) Impact of heat stress during seed development on soybean seed metabolome. Metabolomics 12(2):28
Cheema A (2018) Plant breeding its applications and future prospects. Int J Eng Technol Sci Res 5:88–94
Chen X-B, Xie Y-H, Sun X-M (2015) Development and characterization of polymorphic genic-SSR markers in Larix kaempferi. Molecules 20(4):6060–6067
Chen K, Wang Y, Zhang R, Zhang H, Gao C (2019) CRISPR/Cas genome editing and precision plant breeding in agriculture. Annu Rev Plant Biol 70:667–697
Cheng L, Gao X, Li S, Shi M, Javeed H, Jing X, Yang G, He G (2010) Proteomic analysis of soybean [Glycine max (L.) Meer.] seeds during imbibition at chilling temperature. Mol Breed 26(1):1–17
Cheng G, Zhang L, Wang H, Lu J, Wei H, Yu S (2020) Transcriptomic profiling of young Cotyledons response to chilling stress in two contrasting cotton (Gossypium hirsutum L.) genotypes at the seedling stage. Int J Mol Sci 21(14):5095
Che-Othman MH, Jacoby RP, Millar AH, Taylor NL (2020) Wheat mitochondrial respiration shifts from the tricarboxylic acid cycle to the GABA shunt under salt stress. New Phytol 225(3):1166–1180
Chun JA, Li S, Wang Q, Lee W-S, Lee E-J, Horstmann N, Park H, Veasna T, Vanndy L, Pros K (2016) Assessing rice productivity and adaptation strategies for Southeast Asia under climate change through multi-scale crop modeling. Agr Syst 143:14–21
Cloutier M, Chatterjee D, Elango D, Cui J, Bruns MA, Chopra S (2021) Sorghum root flavonoid chemistry, cultivar, and frost stress effects on rhizosphere bacteria and fungi. Phytobiome J 5(1):39–50
Cui Y, Jiang N, Xu Z, Xu Q (2020) Heterotrimeric G protein are involved in the regulation of multiple agronomic traits and stress tolerance in rice. BMC Plant Biol 20(1):1–13
Cuperlovic-Culf M, Vaughan MM, Vermillion K, Surendra A, Teresi J, McCormick SP (2019) Effects of atmospheric CO2 level on the metabolic response of resistant and susceptible wheat to Fusarium graminearum infection. Mol Plant Microbe Interact 32(4):379–391
D’Amelia L, Dell’Aversana E, Woodrow P, Ciarmiello LF, Carillo P (2018) Metabolomics for crop improvement against salinity stress. Salinity responses and tolerance in plants, vol 2. Springer, pp 267–287
D’Attoma G, Morelli M, Saldarelli P, Saponari M, Giampetruzzi A, Boscia D, Savino VN, De La Fuente L, Cobine PA (2019) Ionomic differences between susceptible and resistant olive cultivars infected by Xylella fastidiosa in the outbreak area of Salento, Italy. Pathogens 8(4):272
Daloz AS, Rydsaa JH, Hodnebrog Ø, Sillmann J, van Oort B, Mohr CW, Agrawal M, Emberson L, Stordal F, Zhang T (2021) Direct and indirect impacts of climate change on wheat yield in the Indo-Gangetic plain in India. J Agric Food Res 4:100132
Damaris RN, Li M, Liu Y, Chen X, Murage H, Yang P (2016) A proteomic analysis of salt stress response in seedlings of two African rice cultivars. Biochimica et Biophysica Acta (BBA)-Proteins and Proteomics 1864(11):1570–1578
Deery DM, Jones HG (2021) Field phenomics: will it enable crop improvement? Plant Phenomic 2021
Deng C, Zhang Z, Yan G, Wang F, Zhao L, Liu N, Abudurezike A, Li Y, Wang W, Shi S (2020) Salt-responsive transcriptome analysis of triticale reveals candidate genes involved in the key metabolic pathway in response to salt stress. Sci Rep 10(1):1–9
Diambra LA (2011) Genome sequence and analysis of the tuber crop potato. Nature 475:189–195
Díaz ML, Soresi DS, Basualdo J, Cuppari SJ, Carrera A (2019) Transcriptomic response of durum wheat to cold stress at reproductive stage. Mol Biol Rep 46(2):2427–2445
Ding J, Adiconis X, Simmons SK, Kowalczyk MS, Hession CC, Marjanovic ND, Hughes TK, Wadsworth MH, Burks T, Nguyen LT (2020) Systematic comparison of single-cell and single-nucleus RNA-sequencing methods. Nat Biotechnol 38(6):737–746
Dong X, Guan L, Zhang P, Liu X, Li S, Fu Z, Tang L, Qi Z, Qiu Z, Jin C (2021) Responses of maize with different growth periods to heat stress around flowering and early grain filling. Agric For Meteorol 303:108378
Du F, Liu P, Wang K, Yang Z, Wang L (2020) Ionomic responses of rice plants to the stresses of different arsenic species in hydroponics. Chemosphere 243:125398
Dudziak K, Zapalska M, Börner A, Szczerba H, Kowalczyk K, Nowak M (2019) Analysis of wheat gene expression related to the oxidative stress response and signal transduction under short-term osmotic stress. Sci Rep 9(1):1–14
Efeoğlu B, Ekmekçi Y, Çiçek N (2009) Physiological responses of three maize cultivars to drought stress and recovery. S Afr J Bot 75(1):34–42
ElBasyoni I, Saadalla M, Baenziger S, Bockelman H, Morsy S (2017) Cell membrane stability and association mapping for drought and heat tolerance in a worldwide wheat collection. Sustainability 9(9):1606
Ergin S, Aydogan C, Turhan E (2021) Physiological effects of high temperature stress in some cucurbit plants. Current Trends in Natural Sciences 10(19):235–245
Fiaz S, Khan SA, Younas A, Shahzad K, Ali H, Noor MA, Ashraf U, Nadeem F (2021) Application of CRISPR/Cas system for genome editing in cotton. In: CRISPR and RNAi Systems. Elsevier, pp 277–301
Fuhrer T, Zamboni N (2015) High-throughput discovery metabolomics. Curr Opin Biotechnol 31:73–78
Fumagalli E, Baldoni E, Abbruscato P, Piffanelli P, Genga A, Lamanna R, Consonni R (2009) NMR techniques coupled with multivariate statistical analysis: tools to analyse Oryza sativa metabolic content under stress conditions. J Agron Crop Sci 195(2):77–88
Ganguly M, Datta K, Roychoudhury A, Gayen D, Sengupta DN, Datta SK (2012) Overexpression of Rab16A gene in indica rice variety for generating enhanced salt tolerance. Plant Signal Behav 7(4):502–509
Gao S, Tang G, Hua D, Xiong R, Han J, Jiang S, Zhang Q, Huang C (2019) Stimuli-responsive bio-based polymeric systems and their applications. J Mater Chem B 7(5):709–729
Godoy F, Olivos-Hernández K, Stange C, Handford M (2021) Abiotic stress in crop species: improving tolerance by applying plant metabolites. Plants 10(2):186
Goff SA, Ricke D, Lan T-H, Presting G, Wang R, Dunn M, Glazebrook J, Sessions A, Oeller P, Varma H (2002) A draft sequence of the rice genome (Oryza sativa L. ssp. japonica). Science 296(5565):92–100
Gorim LY, Vandenberg A (2017) Evaluation of wild lentil species as genetic resources to improve drought tolerance in cultivated lentil. Front Plant Sci 8:1129
Guan S, Xu Q, Ma D, Zhang W, Xu Z, Zhao M, Guo Z (2019) Transcriptomics profiling in response to cold stress in cultivated rice and weedy rice. Gene 685:96–105
Guo Z, Yang W, Chang Y, Ma X, Tu H, Xiong F, Jiang N, Feng H, Huang C, Yang P (2018) Genome-wide association studies of image traits reveal genetic architecture of drought resistance in rice. Mol Plant 11(6):789–805
Guo X, Xin Z, Yang T, Ma X, Zhang Y, Wang Z, Ren Y, Lin T (2020) Metabolomics response for drought stress tolerance in chinese wheat genotypes (Triticum aestivum). Plants 9(4):520
Gupta PK (2008) Single-molecule DNA sequencing technologies for future genomics research. Trends Biotechnol 26(11):602–611
Hao W, Wang S, Liu H, Zhou B, Wang X, Jiang T (2015) Development of SSR markers and genetic diversity in white birch (Betula platyphylla). PLoS One 10(4):e0125235
Hasan MM-U, Ma F, Islam F, Sajid M, Prodhan ZH, Li F, Shen H, Chen Y, Wang X (2019) Comparative transcriptomic analysis of biological process and key pathway in three cotton (Gossypium spp.) species under drought stress. Int J Mol Sci 20(9):2076
Hasegawa T, Fujimori S, Havlík P, Valin H, Bodirsky BL, Doelman JC, Fellmann T, Kyle P, Koopman JF, Lotze-Campen H (2018) Risk of increased food insecurity under stringent global climate change mitigation policy. Nat Clim Chang 8(8):699–703
He J, Zhao X, Laroche A, Lu Z-X, Liu H, Li Z (2014) Genotyping-by-sequencing (GBS), an ultimate marker-assisted selection (MAS) tool to accelerate plant breeding. Front Plant Sci 5:484
Hindt MN, Akmakjian GZ, Pivarski KL, Punshon T, Baxter I, Salt DE, Guerinot ML (2017) BRUTUS and its paralogs, BTS LIKE1 and BTS LIKE2, encode important negative regulators of the iron deficiency response in Arabidopsis thaliana. Metallomics 9(7):876–890
Hong J, Yang L, Zhang D, Shi J (2016) Plant metabolomics: an indispensable system biology tool for plant science. Int J Mol Sci 17(6):767
Hossain Z, Komatsu S (2014) Potentiality of soybean proteomics in untying the mechanism of flood and drought stress tolerance. Proteomes 2(1):107–127
Hosseini S, Ivanov D, Dolgui A (2019) Review of quantitative methods for supply chain resilience analysis. Transp Res E: Logist Transp Rev 125:285–307
Hückelhoven R, Panstruga R (2011) Cell biology of the plant–powdery mildew interaction. Curr Opin Plant Biol 14(6):738–746
Hussain S (2003) Mountain ecosystems: emerging challenges and opportunities for agriculture in Northern Pakistan. Mountains of Pakistan: Protection, Potential and Prospects, pp 180–191
Hussain SS, Mudasser M (2007) Prospects for wheat production under changing climate in mountain areas of Pakistan – an econometric analysis. Agr Syst 94(2):494–501
Hussain HA, Hussain S, Khaliq A, Ashraf U, Anjum SA, Men S, Wang L (2018) Chilling and drought stresses in crop plants: implications, cross talk, and potential management opportunities. Front Plant Sci 9:393. https://doi.org/10.3389/fpls.2018.00393
Hussain HA, Men S, Hussain S, Zhang Q, Ashraf U, Anjum SA, Ali I, Wang L (2020) Maize tolerance against drought and chilling stresses varied with root morphology and antioxidative defense system. Plants 9(6):720. https://doi.org/10.3390/plants9060720
Hyten DL, Song Q, Choi I-Y, Yoon M-S, Specht JE, Matukumalli LK, Nelson RL, Shoemaker RC, Young ND, Cregan PB (2008) High-throughput genotyping with the GoldenGate assay in the complex genome of soybean. Theor Appl Genet 116(7):945–952
Hyten DL, Smith JR, Frederick RD, Tucker ML, Song Q, Cregan PB (2009) Bulked segregant analysis using the GoldenGate assay to locate the Rpp3 locus that confers resistance to soybean rust in soybean. Crop Sci 49(1):265–271
Ibrahim MI, Sapuan SM, Zainudin ES, Zuhri MYM (2019) Extraction, chemical composition, and characterization of potential lignocellulosic biomasses and polymers from corn plant parts. BioResources 14(3):6485–6500
Imran M, Shafiq S, Naeem MK, Widemann E, Munir MZ, Jensen KB, Wang RR-C (2020) Histone deacetylase (HDAC) gene family in allotetraploid cotton and its diploid progenitors: in silico identification, molecular characterization, and gene expression analysis under multiple abiotic stresses, DNA damage and phytohormone treatments. Int J Mol Sci 21(1):321
Jain R, Gupta S, Kachhwaha S, Kothari S (2021) Integrated metabolome and transcriptome analysis: a new platform/technology for functional biology and natural products research. Crop Improvement, CRC Press, pp 37–51
Janes T, McGrath F, Macadam I, Jones R (2019) High-resolution climate projections for South Asia to inform climate impacts and adaptation studies in the Ganges-Brahmaputra-Meghna and Mahanadi deltas. Sci Total Environ 650:1499–1520
Jangra S, Chaudhary V, Yadav RC, Yadav NR (2021) High-throughput phenotyping: a platform to accelerate crop improvement. Phenomics:1–23
Jardak R, Riahi J, Dallagi W, Planchon S, Boubakri H, Bouamama B, Bouagila A, Nefissi R, Mejri S, Renaut J (2021) Proteomic analysis of salt-responsive proteins in the leaves of two contrasting Tunisian barley landraces. Plant Growth Regul:1–18
Ji L, Zhou P, Zhu Y, Liu F, Li R, Qiu Y (2017) Proteomic analysis of Rice seedlings under cold stress. Protein J 36(4):299–307
Jia X, Zhu Y, Zhang R, Zhu Z, Zhao T, Cheng L, Gao L, Liu B, Zhang X, Wang Y (2020) Ionomic and metabolomic analyses reveal the resistance response mechanism to saline-alkali stress in Malus halliana seedlings. Plant Physiol Biochem 147:77–90
Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier E (2012) A programmable dual-RNA–guided DNA endonuclease in adaptive bacterial immunity. Science 337(6096):816–821
Jorrin-Novo JV (2020) What is new in (plant) proteomics methods and protocols: the 2015–2019 quinquennium. plant proteomics. Springer, pp 1–10
Julkowska MM, Testerink C (2015) Tuning plant signaling and growth to survive salt. Trends Plant Sci 20(9):586–594
Junior CAS, D'Amico-Damião V, Carvalho RF (2021) Phytochrome type B family: the abiotic stress responses signaller in plants. Ann Appl Biol 178(2):135–148
Kaler AS, Ray JD, Schapaugh WT, King CA, Purcell LC (2017) Genome-wide association mapping of canopy wilting in diverse soybean genotypes. Theor Appl Genet 130(10):2203–2217
Kalisz A, Sękara A, Cebula S, Grabowska A, Kunicki E (2014) Impact of low-temperature transplant treatment on yield and quality of cauliflower curds in late spring production. Sci Hortic 176:134–142
Kamyab S, Alami-Saeid K, Eslahi M, Moradi M (2021) cDNA-AFLP technique discloses differential gene expression in response to salinity in wheat (Triticum aestivum L.). Genet Resour Crop Evol:1–13
Kang YJ, Lee T, Lee J, Shim S, Jeong H, Satyawan D, Kim MY, Lee SH (2016) Translational genomics for plant breeding with the genome sequence explosion. Plant Biotechnol J 14(4):1057–1069
Katam R, Shokri S, Murthy N, Singh SK, Suravajhala P, Khan MN, Bahmani M, Sakata K, Reddy KR (2020) Proteomics, physiological, and biochemical analysis of cross tolerance mechanisms in response to heat and water stresses in soybean. PLoS One 15(6):e0233905
Kaur J (2014) A comprehensive review on metabolic syndrome. Cardiology Research and Practice 2014
Khazaei H, O’Sullivan DM, Stoddard FL, Adhikari KN, Paull JG, Schulman AH, Andersen SU, Vandenberg A (2020) Recent advances in faba bean genetic and genomic tools for crop improvement. Legume Sci
Kim Y-A, Moon H, Park C-J (2019) CRISPR/Cas9-targeted mutagenesis of Os8N3 in rice to confer resistance to Xanthomonas oryzae pv. oryzae. Rice 12(1):1–13
Klap C, Yeshayahou E, Bolger AM, Arazi T, Gupta SK, Shabtai S, Usadel B, Salts Y, Barg R (2017) Tomato facultative parthenocarpy results from Sl AGAMOUS-LIKE 6 loss of function. Plant Biotechnol J 15(5):634–647
Kong L, Ashraf U, Cheng S, Rao G, Mo Z, Tian H, Pan S, Tang X (2017) Short-term water management at early filling stage improves early-season rice performance under high temperature stress in South China. Eur J Agron 90:117–126
Korhonen A, Traum D, Màrquez L (2019) Celebrity profiling. Proceedings of the 57th Annual Meeting of the Association for Computational Linguistics
Kosová K, Vítámvás P, Urban MO, Prášil IT, Renaut J (2018) Plant abiotic stress proteomics: the major factors determining alterations in cellular proteome. Front Plant Sci 9:122
Kumar S, Banks TW, Cloutier S (2012) SNP discovery through next-generation sequencing and its applications. Int J Plant Genomic 2012:831460
Kumar S, Rajendran K, Kumar J, Hamwieh A, Baum M (2015) Current knowledge in lentil genomics and its application for crop improvement. Front Plant Sci 6:78
Kumar R, Sahoo S, Joanni E, Singh RK, Tan WK, Kar KK, Matsuda A (2019) Recent progress in the synthesis of graphene and derived materials for next generation electrodes of high performance lithium ion batteries. Prog Energy Combust Sci 75:100786
Kumari M, Grover A, Yadav PV, Arif M, Ahmed Z (2013) Development of EST-SSR markers through data mining and their use for genetic diversity study in Indian accessions of Jatropha curcas L.: a potential energy crop. Gene & Genomic 35(5):661–670
Le TT (2016) Effects of climate change on rice yield and rice market in Vietnam. J Agric Appl Economic 48(4):366–382
Lee K, Zhang Y, Kleinstiver BP, Guo JA, Aryee MJ, Miller J, Malzahn A, Zarecor S, Lawrence-Dill CJ, Joung JK (2019) Activities and specificities of CRISPR/Cas9 and Cas12a nucleases for targeted mutagenesis in maize. Plant Biotechnol J 17(2):362–372
Lenaerts B, Collard BC, Demont M (2019) Improving global food security through accelerated plant breeding. Plant Sci 287:110207
Li C, Unver T, Zhang B (2017) A high-efficiency CRISPR/Cas9 system for targeted mutagenesis in cotton (Gossypium hirsutum L.). Sci Rep 7(1):1–10
Li J, Manghwar H, Sun L, Wang P, Wang G, Sheng H, Zhang J, Liu H, Qin L, Rui H (2019) Whole genome sequencing reveals rare off-target mutations and considerable inherent genetic or/and somaclonal variations in CRISPR/Cas9-edited cotton plants. Plant Biotechnol J 17(5):858–868
Li J, Nadeem M, Chen L, Wang M, Wan M, Qiu L, Wang X (2020) Differential proteomic analysis of soybean anthers by iTRAQ under high-temperature stress. J Proteomics 229:103968
Lin C, Heng S, Jie K, Haijiang X, Xiyan Y (2021) Integrated transcriptome and proteome analysis reveals complex regulatory mechanism of cotton in response to salt stress. J Cotton Res 4(1):1–13
Liu X, He P, Chen W, Gao J (2019) Improving multi-task deep neural networks via knowledge distillation for natural language understanding. arXiv. preprint arXiv:1904.09482
Liu G, Zha Z, Cai H, Qin D, Jia H, Liu C, Qiu D, Zhang Z, Wan Z, Yang Y (2020) Dynamic transcriptome analysis of anther response to heat stress during anthesis in thermotolerant rice (Oryza sativa L.). Int J Mol Sci 21(3):1155
Liu B, Wang X, Li K, Cai Z (2021a) Spatially resolved metabolomics and lipidomics reveal salinity and drought-tolerant mechanisms of cottonseeds. J Agric Food Chem 69(28):8028–8037
Liu Y, Zhang J, Pan T, Ge Q (2021b) Assessing the adaptability of maize phenology to climate change: the role of anthropogenic-management practices. J Environ Manage 293:112874
Lu Y, Lam H, Pi E, Zhan Q, Tsai S, Wang C, Kwan Y, Ngai S (2013) Comparative metabolomics in Glycine max and Glycine soja under salt stress to reveal the phenotypes of their offspring. J Agric Food Chem 61(36):8711–8721
Ma J, Li R, Wang H, Li D, Wang X, Zhang Y, Zhen W, Duan H, Yan G, Li Y (2017) Transcriptomics analyses reveal wheat responses to drought stress during reproductive stages under field conditions. Front Plant Sci 8:592
Makeen AAMA, Mohamed HA, Omara S (2021) Modification of microclimate for better growth and develop of summer tomato crop. Gezira J Agric Sci 2(1):59–73
Mammadov J, Aggarwal R, Buyyarapu R, Kumpatla S (2012) SNP markers and their impact on plant breeding. Int J Plant Genomics 2012
Mao Y, Zhang H, Xu N, Zhang B, Gou F, Zhu J-K (2013) Application of the CRISPR–Cas system for efficient genome engineering in plants. Mol Plant 6(6):2008
Mardis ER (2008) Next-generation DNA sequencing methods. Annu Rev Genomics Hum Genet 9:387–402
Maruyama A, Haneishi Y, Okello SE, Asea G, Tsuboi T, Takagaki M, Kikuchi M (2014a) Rice green revolution and climatic change in East Africa: an approach from the technical efficiency of rainfed rice farmers in Uganda. Agric Sci:2014
Maruyama K, Urano K, Yoshiwara K, Morishita Y, Sakurai N, Suzuki H, Kojima M, Sakakibara H, Shibata D, Saito K (2014b) Integrated analysis of the effects of cold and dehydration on rice metabolites, phytohormones, and gene transcripts. Plant Physiol 164(4):1759–1771
Mathan J, Singh A, Ranjan A (2021) Sucrose transport in response to drought and salt stress involves ABA-mediated induction of OsSWEET13 and OsSWEET15 in rice. Physiol Plant 171(4):620–637
Matias FI, Caraza-Harter MV, Endelman JB (2020) FIELDimageR: an R package to analyze orthomosaic images from agricultural field trials. The Plant Phenom J 3(1):e20005
Meng Z, Meng Z, Zhang R, Liang C, Wan J, Wang Y, Zhai H, Guo S (2015) Expression of the rice arginase gene OsARG in cotton influences the morphology and nitrogen transition of seedlings. PLoS One 10(11):e0141530
Meng L, Zhang T, Geng S, Scott PB, Li H, Chen S (2019) Comparative proteomics and metabolomics of JAZ7-mediated drought tolerance in Arabidopsis. J Proteomics 196:81–91
Min X, Liu Z, Wang Y, Liu W (2020) Comparative transcriptomic analysis provides insights into the coordinated mechanisms of leaves and roots response to cold stress in Common Vetch. Ind Crop Prod 158:112949
Mo ZW, Pan SG, Ashraf U, Kanu AS, Li W, Wang ZM, Duan MY, Tian H, Kargbo MB, Tang XR (2017) Local climate affects growth and grain productivity of precision hill-direct-seeded rice in South China. Appl Ecol Environ Res 15(1):113–125
Mochida K, Shinozaki K (2011) Advances in omics and bioinformatics tools for systems analyses of plant functions. Plant Cell Physiol 52(12):2017–2038
Moose SP, Mumm RH (2008) Molecular plant breeding as the foundation for 21st century crop improvement. Plant Physiol 147(3):969–977
Moreno-Risueno MA, Busch W, Benfey PN (2010) Omics meet networks—using systems approaches to infer regulatory networks in plants. Curr Opin Plant Biol 13(2):126–131
Moschen S, Di Rienzo JA, Higgins J, Tohge T, Watanabe M, González S, Rivarola M, García-García F, Dopazo J, Hopp HE (2017) Integration of transcriptomic and metabolic data reveals hub transcription factors involved in drought stress response in sunflower (Helianthus annuus L.). Plant Mol Biol 94(4):549–564
Muthamilarasan M, Singh NK, Prasad M (2019) Multi-omics approaches for strategic improvement of stress tolerance in underutilized crop species: a climate change perspective. Adv Genet 103:1–38
Mwadzingeni L, Shimelis H, Rees DJG, Tsilo TJ (2017) Genome-wide association analysis of agronomic traits in wheat under drought-stressed and non-stressed conditions. PLoS One 12(2):e0171692
Naveed SA, Zhang F, Zhang J, Zheng T-Q, Meng L-J, Pang Y-L, Xu J-L, Li Z-K (2018) Identification of QTN and candidate genes for salinity tolerance at the germination and seedling stages in rice by genome-wide association analyses. Sci Rep 8(1):1–11
Nawaz F, Zulfiqar B, Ahmad KS, Majeed S, Shehzad MA, Javeed HMR, Tahir MN, Ahsan M (2021) Pretreatment with selenium and zinc modulates physiological indices and antioxidant machinery to improve drought tolerance in maize (Zea mays L.). S Afr J Bot 138:209–216
Ndlovu N (2020) Application of genomics and phenomics in plant breeding for climate resilience. Asian Plant Res J:53–66
Nekrasov V, Wang C, Win J, Lanz C, Weigel D, Kamoun S (2017) Rapid generation of a transgene-free powdery mildew resistant tomato by genome deletion. Sci Rep 7(1):1–6
Oliver SG, Winson MK, Kell DB, Baganz F (1998) Systematic functional analysis of the yeast genome. Trends Biotechnol 16(9):373–378
Ortigosa A, Gimenez-Ibanez S, Leonhardt N, Solano R (2019) Design of a bacterial speck resistant tomato by CRISPR/Cas9-mediated editing of Sl JAZ 2. Plant Biotechnol J 17(3):665–673
Osnato M, Cereijo U, Sala J, Matías-Hernández L, Aguilar-Jaramillo AE, Rodríguez-Goberna MR, Riechmann JL, Rodríguez-Concepción M, Pelaz S (2021) The floral repressors TEMPRANILLO1 and 2 modulate salt tolerance by regulating hormonal components and photo-protection in Arabidopsis. Plant J 105(1):7–21
Paixão JFR, Gillet F-X, Ribeiro TP, Bournaud C, Lourenço-Tessutti IT, Noriega DD, de Melo BP, de Almeida-Engler J, Grossi-de-Sa MF (2019) Improved drought stress tolerance in Arabidopsis by CRISPR/dCas9 fusion with a histone acetyltransferase. Sci Rep 9(1):1–9
Palazzo A, Vervoort JM, Mason-D’Croz D, Rutting L, Havlík P, Islam S, Bayala J, Valin H, Kadi HAK, Thornton P (2017) Linking regional stakeholder scenarios and shared socioeconomic pathways: quantified West African food and climate futures in a global context. Glob Environ Chang 45:227–242
Pandey P, Irulappan V, Bagavathiannan MV, Senthil-Kumar M (2017) Impact of combined abiotic and biotic stresses on plant growth and avenues for crop improvement by exploiting physio-morphological traits. Front Plant Sci 8:537
Paterson AH, Bowers JE, Bruggmann R, Dubchak I, Grimwood J, Gundlach H, Haberer G, Hellsten U, Mitros T, Poliakov A (2009) The Sorghum bicolor genome and the diversification of grasses. Nature 457(7229):551–556
Pazhamala LT, Kudapa H, Weckwerth W, Millar AH, Varshney RK (2021) Systems biology for crop improvement. The Plant Genome (TSI), pp 1–23
Peng Z, He S, Gong W, Xu F, Pan Z, Jia Y, Geng X, Du X (2018) Integration of proteomic and transcriptomic profiles reveals multiple levels of genetic regulation of salt tolerance in cotton. BMC Plant Biol 18(1):1–19
Peng R, Jones DC, Liu F, Zhang B (2020) From sequencing to genome editing for cotton improvement. Trends Biotechnol 39(3)
Pita-Barbosa A, Ricachenevsky FK, Wilson M, Dottorini T, Salt DE (2019) Transcriptional plasticity buffers genetic variation in zinc homeostasis. Sci Rep 9(1):1–11
Porter G, Grills N (2016) Medication misuse in India: a major public health issue in India. J Public Health 38(2)
Punshon T, Jackson BP (2018) Essential micronutrient and toxic trace element concentrations in gluten containing and gluten-free foods. Food Chem 252:258–264
Qiu Y, Kaiser C, Schmidt C, Broders K, Robertson AE, Jamann TM (2020) Identification of quantitative trait loci associated with maize resistance to bacterial leaf streak. Crop Sci 60(1):226–237
Rajam MV (2020) RNA silencing technology: A boon for crop improvement. J Biosci 45(1):1–5
Ray DK, Gerber JS, MacDonald GK, West PC (2015) Climate variation explains a third of global crop yield variability. Nat Commun 6(1):1–9
Raza A, Ashraf F, Zou X, Zhang X, Tosif H (2020) Plant adaptation and tolerance to environmental stresses: mechanisms and perspectives. In: Plant ecophysiology and adaptation under climate change: mechanisms and perspectives I. Springer, pp 117–145
Razzaq A, Sadia B, Raza A, Khalid Hameed M, Saleem F (2019) Metabolomics: a way forward for crop improvement. Metabolites 9(12):303
Reddy SK, Liu S, Rudd JC, Xue Q, Payton P, Finlayson SA, Mahan J, Akhunova A, Holalu SV, Lu N (2014) Physiology and transcriptomics of water-deficit stress responses in wheat cultivars TAM 111 and TAM 112. J Plant Physiol 171(14):1289–1298
Resop JP, Fleisher DH, Timlin DJ, Mutiibwa D, Reddy V (2016) Climate, water management, and land use: Estimating potential potato and corn production in the US northeastern seaboard region. Trans ASABE 59(6):1539–1553
Rojas C, Pino J, Basnou C, Vivanco M (2013) Assessing land-use and-cover changes in relation to geographic factors and urban planning in the metropolitan area of Concepción (Chile). Implications for biodiversity conservation. Appl Geogr 39:93–103
Rostoks N, Ramsay L, MacKenzie K, Cardle L, Bhat PR, Roose ML, Svensson JT, Stein N, Varshney RK, Marshall DF (2006) Recent history of artificial outcrossing facilitates whole-genome association mapping in elite inbred crop varieties. Proc Natl Acad Sci 103(49):18656–18661
Rounsevell M, Evans S, Bullock P (1999) Climate change and agricultural soils: impacts and adaptation. Clim Change 43(4):683–709
Saia S, Fragasso M, De Vita P, Beleggia R (2019) Metabolomics provides valuable insight for the study of durum wheat: a review. J Agric Food Chem 67(11):3069–3085
Saidi A, Hajibarat Z (2020) Application of Next Generation Sequencing, GWAS, RNA seq, WGRS, for genetic improvement of potato (Solanum tuberosum L.) under drought stress. Biocatalysis and Agricultural. Biotechnology 29:101801
Sammina M, AShraf, Rehman Z, Ikram M, Mehmood S (2021) Plant resilience to abiotic stress mitigated through phytohormones’ production and their transcriptional control. In: Plant growth regulators for climate-smart agriculture. CRC Press, pp 165–186
Samota M, Bhatt L, Yadav D, Garg N, Bajiya R (2017) Metabolomics for functional genomics. Int J Curr Microbiol App Sci 6:2531–2537
Shahbaz M, Mushtaq Z, Andaz F, Masood A (2013) Does proline application ameliorate adverse effects of salt stress on growth, ions and photosynthetic ability of eggplant (Solanum melongena L.)? Sci Hortic 164:507–511
Shakoor N, Ziegler G, Dilkes BP, Brenton Z, Boyles R, Connolly EL, Kresovich S, Baxter I (2016) Integration of experiments across diverse environments identifies the genetic determinants of variation in Sorghum bicolor seed element composition. Plant Physiol 170(4):1989–1998
Shen L, Wang C, Fu Y, Wang J, Liu Q, Zhang X, Yan C, Qian Q, Wang K (2018) QTL editing confers opposing yield performance in different rice varieties. J Integr Plant Biol 60(2):89–93
Shendure J, Ji H (2008) Next-generation DNA sequencing. Nat Biotechnol 26(10):1135–1145
Shi J, Gao H, Wang H, Lafitte HR, Archibald RL, Yang M, Hakimi SM, Mo H, Habben JE (2017) ARGOS 8 variants generated by CRISPR-Cas9 improve maize grain yield under field drought stress conditions. Plant Biotechnol J 15(2):207–216
Shiferaw B, Smale M, Braun H-J, Duveiller E, Reynolds M, Muricho G (2013) Crops that feed the world 10. Past successes and future challenges to the role played by wheat in global food security. Food Secur 5(3):291–317
Silvente S, Sobolev AP, Lara M (2012) Metabolite adjustments in drought tolerant and sensitive soybean genotypes in response to water stress. PLoS One 7(6):e38554
Singh J, Mishra N, Banerjee S, Sharma YC (2011) Comparative studies of physical characteristics of raw and modified sawdust for their use as adsorbents for removal of acid dye. bioresources 6(3):2732–2743
Singh BK, Mishra DC, Yadav S, Ambawat S, Vaidya E, Tribhuvan KU, Kumar A, Kumar S, Kumar S, Chaturvedi K (2016) Identification, characterization, validation and cross-species amplification of genic-SSRs in Indian Mustard (Brassica juncea). J Plant Biochem Biotechnol 25(4):410–420
Singh A, Müller B, Schnürer A (2021) Profiling temporal dynamics of acetogenic communities in anaerobic digesters using next-generation sequencing and T-RFLP. Sci Rep 11(1):1–14
Solankey SS, Kumari M, Akhtar S, Singh HK, Ray PK (2021) Challenges and opportunities in vegetable production in changing climate: mitigation and adaptation strategies. Advances in research on vegetable production under a changing climate, vol 1. Springer, pp 13–59
Soltabayeva A, Ongaltay A, Omondi JO, Srivastava S (2021) Morphological, physiological and molecular markers for salt-stressed plants. Plants 10(2):243
Song A, Li P, Fan F, Li Z, Liang Y (2014) The effect of silicon on photosynthesis and expression of its relevant genes in rice (Oryza sativa L.) under high-zinc stress. PLoS One 9(11):e113782
Soora NK, Aggarwal PK, Saxena R, Rani S, Jain S, Chauhan N (2013) An assessment of regional vulnerability of rice to climate change in India. Clim Change 118(3):683–699
Soriano JM, Colasuonno P, Marcotuli I, Gadaleta A (2021) Meta-QTL analysis and identification of candidate genes for quality, abiotic and biotic stress in durum wheat. Sci Rep 11(1):1–15
Subramanian I, Verma S, Kumar S, Jere A, Anamika K (2020) Multi-omics data integration, interpretation, and its application. Bioinformatic Biol Insight 14:1177932219899051
Sun J, Wang J, Guo W, Yin T, Zhang S, Wang L, Xie D, Zou D (2021) Identification of alkali-tolerant candidate genes using the NGS-assisted BSA strategy in rice. Mol Breed 41(8):1–14
Sung J, Lee S, Lee Y, Ha S, Song B, Kim T, Waters BM, Krishnan HB (2015) Metabolomic profiling from leaves and roots of tomato (Solanum lycopersicum L.) plants grown under nitrogen, phosphorus or potassium-deficient condition. Plant Sci 241:55–64
Swamy BM, Sarla N (2008) Yield-enhancing quantitative trait loci (QTLs) from wild species. Biotechnol Adv 26(1):106–120
Tanaka N, Shenton M, Kawahara Y, Kumagai M, Sakai H, Kanamori H, Yonemaru J, Fukuoka S, Sugimoto K, Ishimoto M (2020) Whole-genome sequencing of the NARO World Rice Core Collection (WRC) as the basis for diversity and association studies. Plant Cell Physiol 61(5):922–932
Tao F, Hayashi Y, Zhang Z, Sakamoto T, Yokozawa M (2008) Global warming, rice production, and water use in China: developing a probabilistic assessment. Agric For Meteorol 148(1):94–110
Tester M, Langridge P (2010) Breeding technologies to increase crop production in a changing world. Science 327(5967):818–822
Thabet SG, Moursi YS, Sallam A, Karam MA, Alqudah AM (2021) Genetic associations uncover candidate SNP markers and genes associated with salt tolerance during seedling developmental phase in barley. Environ Exp Bot 188:104499
Thomason K, Babar MA, Erickson JE, Mulvaney M, Beecher C, MacDonald G (2018) Comparative physiological and metabolomics analysis of wheat (Triticum aestivum L.) following post-anthesis heat stress. PLoS One 13(6):e0197919
Tian H, Lam SM, Shui G (2016) Metabolomics, a powerful tool for agricultural research. Int J Mol Sci 17(11):1871
Ucar B, Kajtez J, Foidl BM, Eigel D, Werner C, Long KR, Emnéus J, Bizeau J, Lomora M, Pandit A (2021) Biomaterial based strategies to reconstruct the nigrostriatal pathway in organotypic slice co-cultures. Acta Biomater 121:250–262
Ul Qamar MT, Faryad A, Bari A, Zahid B, Zhu X, Chen L-L (2020) Effectiveness of conventional crop improvement strategies vs. omics. Environment, climate, plant and vegetation growth. Springer, pp 253–284
Unamba CI, Nag A, Sharma RK (2015) Next generation sequencing technologies: the doorway to the unexplored genomics of non-model plants. Front Plant Sci 6:1074
Varshney RK, Nayak SN, May GD, Jackson SA (2009) Next-generation sequencing technologies and their implications for crop genetics and breeding. Trends Biotechnol 27(9):522–530
Velasco R, Zharkikh A, Affourtit J, Dhingra A, Cestaro A, Kalyanaraman A, Fontana P, Bhatnagar SK, Troggio M, Pruss D (2010) The genome of the domesticated apple (Malus × domestica Borkh.). Nat Genet 42(10):833–839
Vescio R, Abenavoli MR, Sorgonà A (2021) Single and combined abiotic stress in maize root morphology. Plants 10(1):5
Voss-Fels KP, Stahl A, Hickey LT (2019) Q&A: Modern crop breeding for future food security. BMC Biol 17(1):1–7
Wada H, Hatakeyama Y, Nakashima T, Nonami H, Erra-Balsells R, Hakata M, Nakata K, Hiraoka K, Onda Y, Nakano H (2020) On-site single pollen metabolomics reveals varietal differences in phosphatidylinositol synthesis under heat stress conditions in rice. Sci Rep 10(1):1–11
Wang TL, Uauy C, Robson F, Till B (2012) TILLING in extremis. Plant Biotechnol J 10(7):761–772
Wang Y, Cheng X, Shan Q, Zhang Y, Liu J, Gao C, Qiu J-L (2014) Simultaneous editing of three homoeoalleles in hexaploid bread wheat confers heritable resistance to powdery mildew. Nat Biotechnol 32(9):947–951
Wang F, Wang C, Liu P, Lei C, Hao W, Gao Y, Liu Y-G, Zhao K (2016) Enhanced rice blast resistance by CRISPR/Cas9-targeted mutagenesis of the ERF transcription factor gene OsERF922. PLoS One 11(4):e0154027
Wang Y, Meng Z, Liang C, Meng Z, Wang Y, Sun G, Zhu T, Cai Y, Guo S, Zhang R (2017) Increased lateral root formation by CRISPR/Cas9-mediated editing of arginase genes in cotton. Science China. Life Sci 60(5):524
Wang Y, Wang L, Zhou J, Hu S, Chen H, Xiang J, Zhang Y, Zeng Y, Shi Q, Zhu D (2019) Research progress on heat stress of rice at flowering stage. Ric Sci 26(1):1–10
Wong MM, Bhaskara GB, Wen T-N, Lin W-D, Nguyen TT, Chong GL, Verslues PE (2019) Phosphoproteomics of Arabidopsis Highly ABA-Induced1 identifies AT-Hook–Like10 phosphorylation required for stress growth regulation. Proc Natl Acad Sci 116(6):2354–2363
Xiang Z, Pan-pan L, Chen-hua W, Guo-qiang D, Jian W, Sheng P, Yuan C, De-hua C (2018) Mechanism of reduced insecticidal protein expression in Bt cotton under high temperature and drought based on proteomics. Yingyong Shengtai Xuebao 29(8)
Xu X, Liu X, Ge S, Jensen JD, Hu F, Li X, Dong Y, Gutenkunst RN, Fang L, Huang L (2012) Resequencing 50 accessions of cultivated and wild rice yields markers for identifying agronomically important genes. Nat Biotechnol 30(1):105–111
Xu C, Xia C, Xia Z, Zhou X, Huang J, Huang Z, Liu Y, Jiang Y, Casteel S, Zhang C (2018) Physiological and transcriptomic responses of reproductive stage soybean to drought stress. Plant Cell Rep 37(12):1611–1624
Xu X, Crow M, Rice BR, Li F, Harris B, Liu L, Demesa-Arevalo E, Lu Z, Wang L, Fox N (2021) Single-cell RNA sequencing of developing maize ears facilitates functional analysis and trait candidate gene discovery. Dev Cell 56(4):557-568. e556
Yadav AK, Carroll AJ, Estavillo GM, Rebetzke GJ, Pogson BJ (2019) Wheat drought tolerance in the field is predicted by amino acid responses to glasshouse-imposed drought. J Exp Bot 70(18):4931–4948
Yadav S, Modi P, Dave A, Vijapura A, Patel D, Patel M (2020) Effect of abiotic stress on crops. In: Sustainable Crop Production. IntechOpen, Rijeka
Yan M, Zheng L, Li B, Shen R, Lan P (2021) Comparative proteomics reveals new insights into the endosperm responses to drought, salinity and submergence in germinating wheat seeds. Plant Mol Biol 105(3):287–302
Yasin NA, Zaheer MM, Khan WU, Ahmad SR, Ahmad A, Ali A, Akram W (2018) The beneficial role of potassium in Cd-induced stress alleviation and growth improvement in Gladiolus grandiflora L. Int J Phytoremediation 20(3):274–283
Yildirim C, Correia A-P (2015) Exploring the dimensions of nomophobia: Development and validation of a self-reported questionnaire. Comput Hum Behav 49:130–137
You J, Zhang H, Liu N, Gao L, Kong L, Yang Z (2011) Transcriptomic responses to aluminum stress in soybean roots. Genome 54(11):923–933
Young J, Zastrow-Hayes G, Deschamps S, Svitashev S, Zaremba M, Acharya A, Paulraj S, Peterson-Burch B, Schwartz C, Djukanovic V (2019) CRISPR-Cas9 editing in maize: systematic evaluation of off-target activity and its relevance in crop improvement. Sci Rep 9(1):1–11
Younis M, Ali S, Rizvi STR, Tantawy M, Tariq KU, Bekir A (2021) Investigation of solitons and mixed lump wave solutions with (3+ 1)-dimensional potential-YTSF equation. Commun Nonlinear Sci Numer Simul 94:105544
Zafar SA, Zaidi SS e A, Gaba Y, Singla-Pareek SL, Dhankher OP, Li X, Mansoor S, Pareek A (2020) Engineering abiotic stress tolerance via CRISPR/Cas-mediated genome editing. J Exp Bot 71(2):470–479
Zalapa JE, Cuevas H, Zhu H, Steffan S, Senalik D, Zeldin E, McCown B, Harbut R, Simon P (2012) Using next-generation sequencing approaches to isolate simple sequence repeat (SSR) loci in the plant sciences. Am J Bot 99(2):193–208
Zeng A, Chen P, Korth K, Hancock F, Pereira A, Brye K, Wu C, Shi A (2017) Genome-wide association study (GWAS) of salt tolerance in worldwide soybean germplasm lines. Mol Breed 37(3):1–14
Zhang B (2015) MicroRNA: a new target for improving plant tolerance to abiotic stress. J Exp Bot 66(7):1749–1761
Zhang F, Wen Y, Guo X (2014) CRISPR/Cas9 for genome editing: progress, implications and challenges. Hum Mol Genet 23(R1):R40–R46
Zhang Y, Liang Z, Zong Y, Wang Y, Liu J, Chen K, Qiu J-L, Gao C (2016) Efficient and transgene-free genome editing in wheat through transient expression of CRISPR/Cas9 DNA or RNA. Nat Commun 7(1):1–8
Zhang Y, Ma X-M, Wang X-C, Liu J-H, Huang B-Y, Guo X-Y, Xiong S-P, La G-X (2017) UPLC-QTOF analysis reveals metabolomic changes in the flag leaf of wheat (Triticum aestivum L.) under low-nitrogen stress. Plant Physiol Biochem 111:30–38
Zhang X, Zhou X, Lin M, Sun J (2018) Shufflenet: An extremely efficient convolutional neural network for mobile devices. Proceedings of the IEEE conference on computer vision and pattern recognition
Zhang M, Liu Q, Yang X, Xu J, Liu G, Yao X, Ren R, Xu J, Lou L (2020a) CRISPR/Cas9-mediated mutagenesis of Clpsk1 in watermelon to confer resistance to Fusarium oxysporum f. sp. niveum. Plant Cell Rep 39(5):589–595
Zhang Y, Gao S-y, Liu H-h, Zhang X-l, Zeng A-s, Wang J-j, Hou X-l, Li Y (2020b) cDNA-AFLP analysis of differentially expressed genes during microspore embryogenesis in non-heading Chinese cabbage. In Vitro Cell Dev Biol Plant 56(1):18–28
Zhang L, Chu Q-Q, Jiang Y-L, Fu C, Lei Y-D (2021a) Impacts of climate change on drought risk of winter wheat in the North China Plain. J Integr Agric 20(10):2601–2612
Zhang X, Chen J, Feng K, Wang N, Zhang S, Ma H, Ge C, Shen Q, Liu R, Zhao X (2021b) Widely targeted metabolomics reveals the different metabolic changes in leaves and roots of two cotton varieties under drought stress. J Agron Crop Sci
Zhou L, Yarra R (2021) Genome-wide identification and expression analysis of bZIP transcription factors in oil palm (Elaeis guineensis Jacq.) under abiotic stress. Protoplasma:1–15
Zhou Z, Jiang Y, Wang Z, Gou Z, Lyu J, Li W, Yu Y, Shu L, Zhao Y, Ma Y (2015) Resequencing 302 wild and cultivated accessions identifies genes related to domestication and improvement in soybean. Nat Biotechnol 33(4):408–414
Zhou S, Zhang YK, Kremling KA, Ding Y, Bennett JS, Bae JS, Kim DK, Ackerman HH, Kolomiets MV, Schmelz EA (2019) Ethylene signaling regulates natural variation in the abundance of antifungal acetylated diferuloylsucroses and Fusarium graminearum resistance in maize seedling roots. New Phytol 221(4):2096–2111
Zhu D, Luo F, Zou R, Liu J, Yan Y (2021) Integrated physiological and chloroplast proteome analysis of wheat seedling leaves under salt and osmotic stresses. J Proteomics 234:104097
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this chapter
Cite this chapter
Ashraf, U., Mahmood, S., Shahid, N., Imran, M., Siddique, M., Abrar, M. (2022). Multi-omics Approaches for Strategic Improvements of Crops Under Changing Climatic Conditions. In: Prakash, C.S., Fiaz, S., Fahad, S. (eds) Principles and Practices of OMICS and Genome Editing for Crop Improvement. Springer, Cham. https://doi.org/10.1007/978-3-030-96925-7_3
Download citation
DOI: https://doi.org/10.1007/978-3-030-96925-7_3
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-96924-0
Online ISBN: 978-3-030-96925-7
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)