Abstract
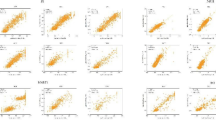
This work presents an approach that uses both genetic algorithm (GA) and many machine learning algorithms (MLA) for feature selection molecular descriptors in a quantitative structure-activity relationships (QSAR) classification and prediction problem. The MLA is used to evaluate an individual population in the GA process. So the fitness function is introduced and defined by the best accuracy classification of the GA and MLA combination. The proposed approach has been implemented and tested using a data set with experimental value antihuman immunodeficiency virus (anti-HIV) molecules. The classification parameters sensibility is equal to 0.99, specificity is equal to 0.91, and accuracy is equal to 0.98. These results reveal the capacity for achieving data subset of molecular descriptors, with high predictive capacity as well as the effectiveness and robustness of the proposed approach.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
UNAIDS, Ending the AIDS epidemic 2020: Global HIV Statistics, (2020)
C.P. Swathik, K.D. Jaspreet, M. Vidhi, R. Navaneethan, J. Mannu, S. Durai, Quantitative structure-activity relationship (QSAR): modeling approaches to biological applications. Encycl. Bioinforma. Comput. Biol. 2, 661–676 (2019)
I. Hdoufane, J. Stoycheva, A. Tadjer, D. Villemin, M. Najdoska-Bogdanov, J. Bogdanov, D. Cherqaoui, QSAR and molecular docking studies of indole-based analogs as HIV-1 attachment inhibitors. J. Mol. Struct. 1193, 429–443 (2019)
R. Todeschini, V. Consonni, Molecular Descriptors for Chemoinformatics (Wiley-VCH, 2009)
M. Eklund, U. Norinder, S. Boyer, L. Carlsson, Choosing feature selection and learning algorithms in QSAR. J. Chem. Inf. Model. 54(3), 837–843 (2014)
F. Grisoni, V. Consonni, R. Todeschini, Impact of molecular descriptors on computational models, in Computational Chemogenomics. Methods in Molecular Biology, ed. by J. Brown, vol. 1825, (Humana Press, New York, 2018)
X.Y. Liu, Y. Liang, S. Wang, Z.Y. Yang, H.S. Ye, Hybrid genetic algorithm with wrapper-embedded approaches for feature selection. IEEE Access. 6, 22863–22874 (2018)
B. Wutzl, K. Leibnitz, F. Rattay, M. Kronbichler, M. Murata, S.M. Golaszewski, Genetic algorithms for feature selection when classifying severe chronic disorders of consciousness. PLoS One 14(7), 1–16 (2019)
K. Nagasubramanian, S. Jones, S. Sarkar, A.K. Singh, A. Singh, B. Ganapathysubramanian, Hyperspectral band selection using genetic algorithm and support vector machines for early identification of charcoal rot disease in soybean stems. Plant Methods 14(86), 1–13 (2018)
H. Labjar, M. Kissi, R. Mouhibi, O. Khadir, H. Chaair, M. Zahouily, QSAR study of 1-(3, 3-diphenylpropyl)-piperidinyl amides and ureas using genetic algorithms and artificial neural networks. Int. J. Bioinforma. Res. Appl. 12(2), 116–128 (2016)
N. Salari, S. Shohaimi, F. Najafi, M. Nallappan, I. Karishnarajah, A novel hybrid classification model of genetic algorithms, modified k-nearest neighbor and developed backpropagation neural network. PLoS One 9(11), 1–50 (2014)
A.K. Srivastava, D. Singh, A.S. Pandey, T. Maini, A novel feature selection and short-term Price forecasting based on a decision tree (J48) model. Energies 12, 1–17 (2019)
C.P. Swathik, K.D. Jaspreet, M. Vidhi, R. Navaneethan, J. Mannu, S. Durai, Quantitative structure-activity relationship (QSAR): modeling approaches to biological applications. Encycl. Bioinform. Comput. Biol. 2, 661–676 (2019)
B. Liu, H. He, H. Luo, T. Zhang, J. Jiang, Artificial intelligence and big data facilitated targeted drug discovery. Stroke Vasc. Neurol. 4, 206–213 (2019)
A. Racz, D. Bajusz, K. Héberger, Intercorrelation limits in molecular descriptor preselection for QSAR/QSPR. Mol. Inf. 38, 1–6 (2019)
J. H. Holland. Adaptation in Natural and Artificial Systems. (Ann Arbor, MI, University of Michigan Press. 1992)
E. Pourbasheer, R. Aalizadeh, M.R. Ganjali, P. Norouzi, J. Shadmanesh, QSAR study of ACK1 inhibitors by genetic algorithm–multiple linear regression (GA–MLR). J. Saudi Chem. Soc. 18, 681–688 (2014)
I.I. Baskin, D. Winkler, I.V. Tetko, A renaissance of neural networks in drug discovery. Expert Opin. Drug Discovery 11, 785–795 (2016)
P. Pradeep, R.J. Povinelli, S. White, S.J. Merrill, An ensemble model of QSAR tools for regulatory risk assessment. J. Chem. 8(48), 1–9 (2016)
T.K. Shameera Ahamed, V.K. Rajan, K. Sabira, K. Muraleedharan, QSAR classification-based virtual screening followed by molecular docking studies for identification of potential inhibitors of 5-lipoxygenase. Comput. Biol. Chem. 77, 154–166 (2018)
K. Lee, M. Lee, D. Kim, Utilizing random Forest QSAR models with optimized parameters for target identification and its application to target-fishing server. BMC Bioinf. 18, 75–86 (2017)
L. Wen, Q. Li, W. Li, Q. Cai, Y.M. Cai, A QSAR study based on SVM for the compound of hydroxyl benzoic esters. Bioinorg. Chem. Appl., 1–10 (2017)
H. Tanaka, H. Takashima, M. Ubasawa, K. Sekiya, I. Nitta, M. Baba, S. Shigata, R.T. Walker, E. De Clercq, T. Miyasaka, Structure-activity relationships of 1-[(2-hydroxyethoxy) methyl]-6-(phenylthio) thymine (HEPT) analogues: effect of substitutions at the C-6 phenyl ring and the C-5 position on anti-HIV-1 activity. J. Med. Chem. 35, 337–345 (1992)
R. Garg, S.P. Gupta, H. Gao, M.S. Babu, A.K. Debnath, Comparative quantitative structure-activity relationships studies on anti-HIV drugs. Chem. Rev. 99, 3525–3601 (1999)
H. Bazoui, M. Zahouily, S. Boulajaaj, S. Sebti, D. Zakarya, QSAR for anti-HIV activity of HEPT derivatives. SAR QSAR Environ. Res. 13(6), 567–577 (2002)
MMP, molecular modelling pro-Demo (TM) Revision 301 demo. ChemSW Software (TM)
S. Anacleto de Souza, L.G. Leonardo Ferreira, S. Aldo de Oliveira, D. Adriano Andricopulo, Quantitative structure–activity relationships for structurally diverse Chemotypes having anti-Trypanosoma cruzi activity. Int. J. Mol. Sci. 20, 1–21 (2019)
L. Wen, Q. Li, W. Li, Q. Cai, Y.M. Cai, A QSAR study based on SVM for the compound of hydroxyl benzoic esters. Bioinorg. Chem. Appl., 1–10 (2017)
S.M. Marunnan, B.P. Pulikkal, A. Jabamalairaj, S. Bandaru, M. Yadav, A. Nayarisseri, V.A. Doss, Development of MLR and SVM aided QSAR models to identify common SAR of GABA uptake herbal inhibitors used in the treatment of schizophrenia. Curr. Neuropharmacol. 15(8), 1085–1092 (2017)
S.K. Chakravarti, S.R.M. Alla, Descriptor free QSAR modeling using deep learning with long short-term memory neural networks. Front. Artif. Intell. 2(17), 1–18 (2019)
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this chapter
Cite this chapter
Labjar, H., Labjar, N., Kissi, M. (2022). QSAR Anti-HIV Feature Selection and Prediction for Drug Discovery Using Genetic Algorithm and Machine Learning Algorithms. In: Ouaissa, M., Boulouard, Z., Ouaissa, M., Guermah, B. (eds) Computational Intelligence in Recent Communication Networks . EAI/Springer Innovations in Communication and Computing. Springer, Cham. https://doi.org/10.1007/978-3-030-77185-0_12
Download citation
DOI: https://doi.org/10.1007/978-3-030-77185-0_12
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-77184-3
Online ISBN: 978-3-030-77185-0
eBook Packages: Intelligent Technologies and RoboticsIntelligent Technologies and Robotics (R0)