Abstract
The main reasons for the slow progress in improving survival outcomes for ovarian cancer are the ‘one-size-fits-all’ therapy and lack of clinically relevant experimental models that represent the advanced stages of the human disease. The interaction of tumour cells with their surrounding niche, the tumour microenvironment, influences the spread of ovarian cancer cells within the peritoneum and their responses to therapeutics. Scientists are increasingly using 3D cell culture models to dissect the role of the tumour microenvironment in cancer development and progression and the treatment of this disease. In this chapter, we will briefly describe the tumour microenvironment of ovarian cancer. Then, we will review some of the clinically relevant experimental approaches, such as spheroid, organoid and organotypic models, that have been developed for the 3D culture of ovarian cancer cells using different tools, including hydrogels, scaffolds and cancer-on-a-chip devices, to mimic selected components of the tumour microenvironment.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- 3D cell culture
- Tumour microenvironment
- Spheroids
- Organoids
- Organotypic cultures
- Hydrogels
- Scaffolds
- Cancer-on-a-chip
9.1 Introduction
Ovarian cancer is the deadliest gynaecological malignancy in the Western world [1]. The majority (over 75%) of patients have metastatic disease at the time of diagnosis, and survival rates have not changed over the past four decades. Two reasons for the slow progress in improving survival outcomes are (1) the ‘one-size-fits-all’ therapeutic approach and (2) the lack of clinically relevant experimental models that mimic the advanced stages of the human disease to find better therapeutic options [2]. In 2015, the first whole-genome study of chemoresistant ovarian cancer was reported [3]. This worldwide, largest DNA analysis revealed key regulatory mechanisms involving interactions with the tumour microenvironment (TME) that pointed to how cancer cells hijack chemotherapy. Thus, there is an urgent need for more targeted strategies and cell models more representative of the TME, specifically new 3D models [4], for treating and studying the most aggressive form—high-grade serous carcinoma (HGSC).
Since then, advances have been made in our understanding of ovarian cancer and the role of the TME in cancer progression and treatment resistance [5, 6]. Scientists need to consider the cellular and extracellular TME when developing experimental cancer models [7]. The majority of our current knowledge about cellular processes and mechanism has been derived from cancer cells grown attached to flat plastic culture dishes as monolayers, namely two-dimensional (2D) cell cultures [8]. However, 2D cell cultures are limited in terms of their complexity and cellular interactions that govern cell behaviour and drug responses [9]. Dynamic processes, such as epithelial-to-mesenchymal transition, cell invasion, treatment resistance or angiogenesis, cannot be adequately explained [10]. Shifting from 2D to three-dimensional (3D) cell cultures allows for the reconstruction of cell–cell and cell–matrix interactions and other critical TME components and enables studies of physiologically relevant cell behaviours, for example, the dynamics in spatial-temporal oxygen, growth factor gradients and shear stress [11].
Other experimental cancer models include xenograft approaches and murine models of ovarian cancer. These models replicate the human disease in terms of disease development and progression, metastasis and partial immune responses and are useful tools to study responses to treatment [12, 13]. However, murine cell infiltration may lead to mouse-specific tumour evolution, and these models are costly and labour-intensive [14]. In the following sections, we will briefly describe the ovarian TME and discuss some of the clinically relevant experimental approaches that have been used to model selected elements of the TME in 3D.
9.2 Ovarian Cancer
Ovarian cancer is a heterogeneous disease, which makes the design of experimental cancer models even harder. There are different subtypes with distinct biological characteristics and molecular aberrations and treatment strategies are stereotypically applied, in particular aggressive surgical debulking and platinum-based chemotherapy [15].
Epithelial ovarian cancer occurs in over 90% of ovarian malignancies, whereas non-epithelial forms, including germ cell and sex cord-stromal tumours, account for about 5% [16]. Epithelial malignancies develop from the fallopian tubes or other epithelial sites. They are categorised into distinct histo-morphological subtypes: low-grade serous, endometrioid, mucinous and clear cell carcinoma (‘type I ovarian tumours’) and HGSC, mixed Mullerian malignancies and high-grade endometrioid ovarian carcinomas (‘type II ovarian tumours’). Both type I and type II ovarian tumours differ in their point-of-origin, gene mutations, disease progression and clinical outcomes [15]. HGSC is the most frequently diagnosed sub-type, accounting for most ovarian cancer deaths (70–80%), and its point-of-origin is still under debate [6].
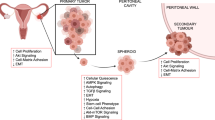
At an early stage, ovarian cancer is asymptomatic and specific biomarkers do not exist. Consequently, patients are diagnosed at an advanced stage, after metastatic spread has already occurred. Dissemination of ovarian cancer is a major clinical problem and results in even further reduced survival rates for patients. Metastatic spread occurs through intraperitoneal dissemination within the tumour fluid (ascites) to secondary sites. Ascites is associated with chemoresistance and disease recurrence and contains single cancer cells as well as multicellular cancer spheroids. These cells and spheroids adhere to mesothelium-lined organs, such as the peritoneum, the small and large bowel serosa, or the omentum and invade into the underlying extracellular matrix (ECM) to form macro-metastases [17, 18]. The cellular composition and cellular states of ascites vary significantly between patients, and within a patient’s primary tumour and metastatic lesions, and need to be considered when developing 3D models using ascites-derived cells or recreating the ascites-specific TME. In a single-cell analysis of ascites-derived cells from patients with advanced HGSC, diverse subpopulations of immunomodulatory fibroblasts, cancer-associated fibroblasts and macrophages were identified [19].
9.3 3D Models That Recreate The Ovarian TME
Signals between malignant and non-malignant cells give rise to a tissue-specific TME that promotes cancer progression, metastasis and chemoresistance. The ovarian TME is highly complex and consists of various non-malignant cell types, including cells of the tumour vasculature, fibroblasts, adipocytes, mesothelial and inflammatory cells, and extracellular components, for example, the ECM, growth factors, cytokines and proteolytic factors [20]. The intrinsically heterogeneous tumour-immune microenvironment contains co-existing regions with immune-cell-excluded and inflammatory areas within the same patient or tumour site. Cytotoxic chemotherapy has an immunogenic effect and induces immune cell infiltration in HGSC as demonstrated, for example, by an enrichment of natural killer cells or an oligoclonal expansion of T cell subsets [21].
Cancer cells recruit and reprogram these non-malignant cells to create tumourigenic niches. The reciprocal crosstalk between cancer cells and the different TME components supports tumour growth and spread. For example, a common matrix response is associated with metastasis and poor survival [20]. Cancer cell dynamics are intricate and challenging to model experimentally [6]. However, 3D cancer models that accurately replicate the different components and the diversity of the TME are urgently needed to study cancer biology and physiology and to identify more effective treatments to improve the clinical outcomes for this disease.
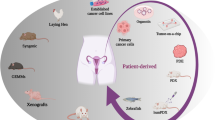
In recent years, great progress in the fields of 3D in vitro cancer models and tumour tissue engineering have been made. A variety of 3D cancer models and bioengineered microenvironments that integrate key elements of the TME in a spatially and biomechanical relevant manner have been developed. These models enable the study of cancer cell behaviour and drug responses under physiological cell culture conditions [7]. Among others, multicellular cancer spheroids, organoid and organotypic models, as well as hydrogel-based and scaffold-based systems are well established. And new technologies, such as cancer-on-a-chip devices, are gaining more attention within the cancer research community.
9.3.1 Cancer Spheroids
Cancer spheroids are an adequate and versatile tool to culture cells in 3D. They are widely used as 3D in vitro cancer model because they closely resemble in vivo tumours in terms of the 3D structure and organisation of tissues or organs [22, 23]. Spheroids are either self-assembling or are forced to grow as cell clusters or aggregates from a single-cell suspension in the absence or presence of exogenous ECM components [24]. Cancer spheroids range in their diameter from 30 to 750 μm and are referred to as bona fide metastatic units with an outermost layer of proliferating cells and a central area of quiescent cells [14]. Larger spheroids (>200 μm) exhibit gradients of oxygen, nutrient, catabolic and soluble factors, including cytokines and growth factors as seen in physiological micro-metastases and avascular tumours, which make them a great 3D model for cancer research [4, 14].
Ovarian cancer spheroids can be isolated from patient’s tumour fluid (ascites) or fabricated by using scaffold-free approaches by which cells produce their own ECM, for example, non-adherent cell culture dishes, hanging drop methods and spinner flask cultures [4]. By using scaffold-based approaches, cells attach and grow on or within polymeric scaffolds, such as matrices and hydrogels [25]. Ovarian cancer spheroids can either be monotypic (malignant cells only) or heterotypic (a mixture of malignant and non-malignant cells).
9.3.2 Hydrogel-Based Models
The ECM provides structural support and triggers biomechanical and biochemical signals that are essential for cancer cell behaviour. ECM properties, including the stiffness, permeability and spatial arrangements, are critical features that influence cancer progression and therapy success and thus need to be considered when developing bioengineered microenvironments. Hydrogels are water-absorbing and water-swollen 3D scaffolds generated from crosslinked biomaterials for the study of cell–cell and cell–matrix interactions. They have physiological properties comparable to native tissues, and their biomechanical and biochemical properties can be widely tailored, which makes them an exciting 3D tool [7].
There are different types of hydrogels , depending on the origin of their main component. Natural hydrogels are generated from naturally derived biomaterials, such as alginate, collagen, hyaluronic acid and Matrigel. Their low cytotoxicity and the presence of cell binding sites make them compatible for 3D cell cultures. However, naturally derived biomaterials have several drawbacks, including a high batch-to-batch variation, undefined and mixed compositions, uncontrolled degradation and poor or low mechanical properties [7]. Synthetic biomaterials, such as polyethylene glycol (PEG) [26] or self-assembling peptide amphiphiles [25], overcome some of these limitations and are applied to increase the experimental reproducibility and to precisely control the biomechanical and biochemical properties. As synthetic biomaterials lack cell adhesion and proteolytic degradation sites, bioactive peptides and molecules are integrated to facilitate cell-stimulatory processes. To combine the biological characteristics of the native ECM with the stable and well-defined properties of synthetic matrices, semi-synthetic hydrogels have been produced [7]. For example, gelatin methacryloyl (GelMA) hydrogels are an alternative 3D model that supports the formation and growth of ovarian cancer spheroids [27]. GelMA hydrogels retain their cell binding and proteolytic degradation sites and have tuneable physical properties, allowing a high degree of experimental control and reproducibility.
The omentum is the primary metastatic site for ovarian cancer. The development of 3D in vitro cancer models that mimic the omental microenvironment may improve the prediction of drug responses. To capture critical omentum-specific ECM protein characteristics [28], omentum-inspired polyethylene glycol-maleimide (PEG-MAL) hydrogels have been developed (Fig. 9.1) [9]. Ovarian cancer cells, allowed to aggregate in microwells, or patient-derived ascites spheroids, were encapsulated into PEG-MAL hydrogels. To support cell viability and proliferation, omentum-specific integrin-binding and ECM-related peptides were added into the hydrogel network. Subsequently, the effects of several anticancer drugs, including the clinically used chemotherapeutic paclitaxel, on cancer spheroids were analysed. While cancer spheroids did not respond to paclitaxel, cell monolayer controls grown on plastic culture dishes were sensitive to paclitaxel and had a reduced cell viability. Moreover, ascites-derived spheroids from patients that had been already treated with paclitaxel did not respond either when grown in PEG-MAL hydrogels. Additionally, cancer spheroids produced their own ECM when cultured in PEG-MAL hydrogels compared to cell monolayers. These findings indicate that the omentum-inspired 3D model may be used as a clinically relevant drug screening platform for ovarian cancer and to identify ECM-related factors that are involved in drug resistance [9].
3D culture of ovarian cancer cells using polyethylene glycol-maleimide (PEG-MAL) hydrogels. Cancer cells seeded in microwells aggregate and form cancer spheroids, which are then collected and grown encapsulated in PEG-MAL hydrogels. The components of the hydrogel network include PEG-MAL, integrin-binding peptides and two different crosslinkers. To recreate the omental microenvironment, different peptide sequences that represent extracellular matrix proteins (e.g. collagen, fibronectin) of the human omentum are crosslinked into the hydrogel network [9]
9.3.3 Organoids
Organoid cultures have been used to study human development and diseases, as well as clinically relevant drug screening platforms and as models of the TME. Patient-derived tumour organoids (or tumouroids) are a great 3D tool for cancer research as they reconstruct the tumour profile in terms of the morphology and gene expression from which they originate [6]. They are established from primary human tumour cells, or murine oviductal (fallopian tube in humans) and ovarian surface epithelium cells that harbour mutagenic modifications [29], and form 3D structures to recapitulate and study tumour heterogeneity and the origin of HGSC [30]. In the presence of growth factors, small molecules and a supporting matrix, mostly Matrigel and collagen gels, organoid cultures are maintained over several months. Organoids grow within days and allow untransformed and precancerous cells to expand [31]. Organoid cultures are cheaper and easier to establish compared to patient-derived xenografts and murine models of ovarian cancer [30].
About 50% of HGSC harbour DNA repair defects, which are targeted by inhibition of a nuclear enzyme, poly-ADP ribose polymerase (PARP). PARP is an important protein that repairs damaged DNA. One of the DNA damage repair mechanism is homologous recombination involving the breast cancer susceptibility genes, BRCA1 and BRCA2. Deficiency, such as mutations, within either gene results in defective homologous recombination, loss of efficient DNA repair and responsiveness to PARP inhibition. For functional profiling of DNA repair and defects in homologous recombination, HGSC patient-derived organoids were established. Regardless of the mutational profile of the DNA repair genes, a functional defect in homologous recombination in the organoids positively correlated with PARP inhibitor sensitivity. In combination with genomic screenings, the functional testing of ovarian tumour organoids is a valid 3D tool for the identification of targetable defects in the repair of DNA damage [30].
In another study, 56 organoids from 32 ovarian cancer patients were established, with a success rate of 65%. For the first time, the generated organoid lines covered all major ovarian cancer subtypes. Moreover, patient-specific genomic features were maintained. These organoid cultures allowed long-term expansion and manipulation, thus offering a platform for drug screening approaches for the different ovarian cancer subtypes [6].
9.3.4 Organotypic Cultures
Organotypic models are composed of multiple cell types found in the cellular TME and an organ-specific ECM to mimic tumour tissues as seen in patients [32]. Ovarian cancer cells preferentially metastasise to the omentum, which is lined by a layer of mesothelial cells. To capture this omental microenvironment, a 3D organotypic model was developed (Fig. 9.2) [18]. Primary omental fibroblasts were mixed with a collagen matrix, followed by addition of mesothelial cells and subsequently co-cultured with fluorescently labelled ovarian cancer cells. This model was used to screen a compound library for their potential to inhibit cell functions in an automated and quantitative high-throughput screen using different HGSC cell lines. Over 44,000 compounds and pharmacologically active small molecules were tested, and only 3 compounds were found to inhibit ovarian cancer cell adhesion, invasion and metastasis, to prolong survival and to reduce omental tumour growth [5].
3D co-culture of ovarian cancer cells with non-malignant cells using an organotypic model. The 3D organotypic model mimics the human omentum through the 3D co-culture of ovarian cancer cells with mesothelial cells and fibroblasts in a collagen matrix. Human primary mesothelial cells and fibroblasts are extracted from human omentum and assessed for the presence of cell type-specific markers (e.g. cytokeratin 8, vimentin and proline-hydroxylase). To mimic the omental basement membrane, patient-derived omental fibroblasts are mixed with a collagen matrix. After cell adhesion, mesothelial cells are seeded on top to reconstruct the mesothelial lining. Subsequently, fluorescently labelled ovarian cancer cells are added and subjected to fluorescence-based assays [18]
Ovarian cancer initiation and progression, including changes in gene expression during early cancer cell dissemination, are poorly understood. To analyse early events in ovarian cancer spread, the aforementioned 3D organotypic model was used. A comprehensive RNA sequencing analysis of healthy fallopian tubes, primary tumours and metastatic lesions was compared with the profiles of their cultures using the 3D organotypic model to identify changes in gene expression. Significant changes in gene expression and key pathways during ovarian cancer initiation, metastasis and early colonisation were identified, which includes the deregulation of ECM proteins and ECM-related factors [33]. The results may help to improve our understanding of critical pathways and their role in ovarian cancer progression in order to develop new and more effective treatment options.
9.3.5 Scaffold-Based Organotypic Cultures
The organ-specific characteristics of the cellular and extracellular TME in disease progression are important, in particular, when designing hydrogel- and scaffold-based 3D models of the omental microenvironment. Tissue engineering approaches that have been used for regenerative medicine can be repurposed and applied to cancer research. Using this interdisciplinary strategy, controllable and reproducible 3D organotypic models have been developed [7]. For example, a 3D TME model was designed to mimic the integral steps in the dissemination of ovarian cancer and its spread to the omentum. Hereby, ovarian cancer spheroids, which formed within PEG hydrogels, were assembled with medical-grade polycaprolactone fibrous scaffolds that were seeded with a layer of mesothelial cells to create a 3D co-culture model (Fig. 9.3) [34]. These 3D co-cultures were carried out for 2 weeks and then, both cell types were mechanically separated for subsequent molecular profiling. Proliferation assays and a high-throughput gene expression and signalling analysis indicated that cancer spheroid growth was enhanced upon 3D co-culture compared to the corresponding 3D monoculture controls and that genes linked to cell growth (e.g. IGFBP7, FGF2, VEGFC, COX2) and proteolytic factors (e.g. KLK5, KLK6, KLK7) were increased in 3D co-cultured cancer spheroids. Upon implantation of 3D co-culture constructs intra-peritoneally into NOD/SCID mice, tumour growth and spread were significantly increased compared to 3D monoculture implants [34]. This tailored and clinically relevant experimental model recreates the organ-specific pattern of early ovarian cancer dissemination within the peritoneal cavity. It represents a quantitative 3D approach to identify the regulatory cellular and molecular mechanisms involved and may help to identify targeted therapies to increase survival rates or to molecular stratify the design of clinical trials for a subset of patients with HGSC.
3D co-culture of ovarian cancer cells with mesothelial cells using a combination of polyethylene glycol (PEG) hydrogels and fibrous scaffolds. To recreate the omental microenvironment, fibrous scaffolds are seeded with mesothelial cells and assembled with ovarian cancer cell-containing PEG hydrogels to form tumour constructs. This combined hydrogel/scaffold model enables the separation of the individual cell types and analysis of low cell yields from 3D co-cultures. Upon cell separation, the effect of 3D co-culture is assessed by confocal and scanning electron microscopy. The presence of cell type-specific markers (e.g. PAX8 and calretinin) confirms the complete separation of the different cell populations. (Modified from [34])
9.3.6 Cancer-on-a-Chip Devices
A variety of experimental models have been developed to study the tumour biology and drug responses of ovarian cancer. Some of the early 3D in vitro cancer models are often oversimplified and unsuited to accurately mimic the complexity of the TME. Consequently, more complex 3D in vitro cancer models, such as multicellular cancer spheroids, organoids and organotypic systems evolved. To overcome their drawbacks, for example, the lack of tissue–tissue interfaces, fluid flow and biochemical cues, new tools, namely organ-on-a-chip devices, have evolved [35]. Organ-on-a-chip models provide cells with fluidic stimuli by perfusing medium in a laminar flow through a porous membrane that separates the individual compartments. This allows the fluidic 3D co-culture of different cell populations to recapitulate complex tissue–tissue interactions. A unique advantage of this technology is its inherent ability to integrate multiple organ functions into a closed microfluidic system, which represents the physiology and metabolism as seen in native tissues, allowing disease modelling and preclinical drug studies [36].
Early and specific biomarkers that allow for HGSC detection at a less-advanced stage do not exist. However, exosomes or extracellular vesicles that are essential for cell–cell communication may be used as a promising biomarker. A cancer-on-chip device has been used for the isolation of intact exosomes from the culture medium of HGSC cells. The proteasome profile was characterised and compared to healthy patient-derived donor cells of the ovarian surface epithelium and the fallopian tube secretory epithelium. Notably, 25 exosomal proteins were differentially expressed in HGSC compared to the controls [37]. These findings may potentially help to detect the disease early and to design targeted therapies.
9.4 Conclusion
Despite intensive research, ovarian cancer remains the leading cause of mortality among gynaecological malignancies for which the treatment options are limited. Given its complexity, experimental models that faithfully mimic the complex microenvironmental stimuli during disease development and progression are urgently needed. In the last decade, it became apparent that traditional cell monolayer models and animal studies are not entirely suited for the modelling of the human disease and treatment response. Hence, new TME models that reconstruct critical elements of the TME in a spatially, physically and chemically relevant manner have been designed. These new 3D platforms have proven more efficient for drug testing and drug discovery and hold enormous potential to improve the treatment options and to screen personalised medicines for patients suffering from ovarian cancer. Personalised medicines aim to move away from the ‘one-size-fits-all’ therapy for patients in order to stratify the variation between individual patients or specific subgroups of patients or subsets of tumours. Personalising cancer treatment includes the molecular profiling of patients to identify biomarkers or genetic profiles that help to select patients for targeted therapies.
Promising new therapies for patients with HGSC, or for patients that have developed resistance to platinum-based chemotherapy, include PARP inhibitors, antiangiogenic therapies and immunotherapies. While PARP inhibitors have shown excellent activity in ovarian cancer, immunotherapies exhibit only modest activity. However, combined PARP and immune checkpoint inhibition has yielded encouraging results for the treatment of ovarian cancer and immunogenomic profiling may identify predictive biomarkers of treatment response [38]. Resistance to PARP inhibitors and platinum-based chemotherapy may be overcome by combining PARP inhibitors with inhibitors of alternative DNA repair pathways, depending on the genetic profile of the individual patient [39, 40]. For ovarian cancer, 3D models that recapitulate physiological aspects and matrix composition of tumour tissues and integrate patient-derived cell populations can be used as patient surrogates to directly test responses to targeted therapies or personalised medicines.
Abbreviations
- 2D:
-
Two-dimensional
- 3D:
-
Three-dimensional
- ECM:
-
Extracellular matrix
- GelMA:
-
Gelatin methacryloyl
- HGSC:
-
High-grade serous carcinoma
- MAL:
-
Maleimide
- PARP:
-
Poly-ADP ribose polymerase
- PEG:
-
Polyethylene glycol
- TME:
-
Tumour microenvironment
References
Siegel, R. L., Miller, K. D., Fuchs, H. E. & Jemal, A. (2021). Cancer statistics, 2021. CA: A Cancer Journal for Clinicians, 71(1), 7–33.
Bowtell, D. D., Bohm, S., Ahmed, A. A., Aspuria, P. J., Bast, R. C., Jr., Beral, V., Berek, J. S., Birrer, M. J., Blagden, S., Bookman, M. A., Brenton, J. D., Chiappinelli, K. B., Martins, F. C., Coukos, G., Drapkin, R., Edmondson, R., Fotopoulou, C., Gabra, H., Galon, J., Gourley, C., Heong, V., Huntsman, D. G., Iwanicki, M., Karlan, B. Y., Kaye, A., Lengyel, E., Levine, D. A., Lu, K. H., McNeish, I. A., Menon, U., Narod, S. A., Nelson, B. H., Nephew, K. P., Pharoah, P., Powell, D. J., Jr., Ramos, P., Romero, I. L., Scott, C. L., Sood, A. K., Stronach, E. A., & Balkwill, F. R. (2015). Rethinking ovarian cancer II: Reducing mortality from high-grade serous ovarian cancer. Nature Reviews. Cancer, 15(11), 668–679.
Patch, A. M., Christie, E. L., Etemadmoghadam, D., Garsed, D. W., George, J., Fereday, S., Nones, K., Cowin, P., Alsop, K., Bailey, P. J., Kassahn, K. S., Newell, F., Quinn, M. C., Kazakoff, S., Quek, K., Wilhelm-Benartzi, C., Curry, E., Leong, H. S., Australian Ovarian Cancer Study Group, Hamilton, A., Mileshkin, L., Au-Yeung, G., Kennedy, C., Hung, J., Chiew, Y. E., Harnett, P., Friedlander, M., Quinn, M., Pyman, J., Cordner, S., O’Brien, P., Leditschke, J., Young, G., Strachan, K., Waring, P., Azar, W., Mitchell, C., Traficante, N., Hendley, J., Thorne, H., Shackleton, M., Miller, D. K., Arnau, G. M., Tothill, R. W., Holloway, T. P., Semple, T., Harliwong, I., Nourse, C., Nourbakhsh, E., Manning, S., Idrisoglu, S., Bruxner, T. J., Christ, A. N., Poudel, B., Holmes, O., Anderson, M., Leonard, C., Lonie, A., Hall, N., Wood, S., Taylor, D. F., Xu, Q., Fink, J. L., Waddell, N., Drapkin, R., Stronach, E., Gabra, H., Brown, R., Jewell, A., Nagaraj, S. H., Markham, E., Wilson, P. J., Ellul, J., McNally, O., Doyle, M. A., Vedururu, R., Stewart, C., Lengyel, E., Pearson, J. V., Waddell, N., de Fazio, A., Grimmond, S. M., & Bowtell, D. D. (2015). Whole-genome characterization of chemoresistant ovarian cancer. Nature, 521(7553), 489–494.
Lengyel, E., Burdette, J. E., Kenny, H. A., Matei, D., Pilrose, J., Haluska, P., Nephew, K. P., Hales, D. B., & Stack, M. S. (2014). Epithelial ovarian cancer experimental models. Oncogene, 33(28), 3619–3633.
Kenny, H. A., Lal-Nag, M., Shen, M., Kara, B., Nahotko, D. A., Wroblewski, K., Fazal, S., Chen, S., Chiang, C. Y., Chen, Y. J., Brimacombe, K. R., Marugan, J., Ferrer, M., & Lengyel, E. (2020). Quantitative high-throughput screening using an organotypic model identifies compounds that inhibit ovarian cancer metastasis. Molecular Cancer Therapeutics, 19(1), 52–62.
Kopper, O., de Witte, C. J., Lohmussaar, K., Valle-Inclan, J. E., Hami, N., Kester, L., Balgobind, A. V., Korving, J., Proost, N., Begthel, H., van Wijk, L. M., Revilla, S. A., Theeuwsen, R., van de Ven, M., van Roosmalen, M. J., Ponsioen, B., Ho, V. W. H., Neel, B. G., Bosse, T., Gaarenstroom, K. N., Vrieling, H., Vreeswijk, M. P. G., van Diest, P. J., Witteveen, P. O., Jonges, T., Bos, J. L., van Oudenaarden, A., Zweemer, R. P., Snippert, H. J. G., Kloosterman, W. P., & Clevers, H. (2019). An organoid platform for ovarian cancer captures intra- and interpatient heterogeneity. Nature Medicine, 25(5), 838–849.
Loessner, D., Holzapfel, B. M., & Clements, J. A. (2014). Engineered microenvironments provide new insights into ovarian and prostate cancer progression and drug responses. Advanced Drug Delivery Reviews, 79–80, 193–213.
Baker, B. M., & Chen, C. S. (2012). Deconstructing the third dimension: How 3D culture microenvironments alter cellular cues. Journal of Cell Science, 125(Pt 13), 3015–3024.
Brooks, E. A., Gencoglu, M. F., Corbett, D. C., Stevens, K. R., & Peyton, S. R. (2019). An omentum-inspired 3D PEG hydrogel for identifying ECM-drivers of drug resistant ovarian cancer. APL Bioengineering, 3(2), 026106.
Al Ameri, W., Ahmed, I., Al-Dasim, F. M., Ali Mohamoud, Y., Al-Azwani, I. K., Malek, J. A., & Karedath, T. (2019). Cell type-specific TGF-beta mediated EMT in 3D and 2D models and its reversal by TGF-beta receptor kinase inhibitor in ovarian cancer cell lines. International Journal of Molecular Sciences, 20(14), 3568.
Masiello, T., Dhall, A., Hemachandra, L. P. M., Tokranova, N., Melendez, J. A., & Castracane, J. (2018). A dynamic culture method to produce ovarian cancer spheroids under physiologically-relevant shear stress. Cell, 7(12), 277.
Maniati, E., Berlato, C., Gopinathan, G., Heath, O., Kotantaki, P., Lakhani, A., McDermott, J., Pegrum, C., Delaine-Smith, R. M., Pearce, O. M. T., Hirani, P., Joy, J. D., Szabova, L., Perets, R., Sansom, O. J., Drapkin, R., Bailey, P., & Balkwill, F. R. (2020). Mouse ovarian cancer models recapitulate the human tumor microenvironment and patient response to treatment. Cell Reports, 30(2), 525–540.e7.
Kim, O., Park, E. Y., Klinkebiel, D. L., Pack, S. D., Shin, Y. H., Abdullaev, Z., Emerson, R. E., Coffey, D. M., Kwon, S. Y., Creighton, C. J., Kwon, S., Chang, E. C., Chiang, T., Yatsenko, A. N., Chien, J., Cheon, D. J., Yang-Hartwich, Y., Nakshatri, H., Nephew, K. P., Behringer, R. R., Fernandez, F. M., Cho, C. H., Vanderhyden, B., Drapkin, R., Bast, R. C., Jr., Miller, K. D., Karpf, A. R., & Kim, J. (2020). In vivo modeling of metastatic human high-grade serous ovarian cancer in mice. PLoS Genetics, 16(6), e1008808.
Loessner, D., Little, J. P., Pettet, G. J., & Hutmacher, D. W. (2013). A multiscale road map of cancer spheroids—Incorporating experimental and mathematical modelling to understand cancer progression. Journal of Cell Science, 126(Pt 13), 2761–2771.
Loessner, D., Goettig, P., Preis, S., Felber, J., Bronger, H., Clements, J. A., Dorn, J., & Magdolen, V. (2018). Kallikrein-related peptidases represent attractive therapeutic targets for ovarian cancer. Expert Opinion on Therapeutic Targets, 22(9), 745–763.
Torre, L. A., Trabert, B., DeSantis, C. E., Miller, K. D., Samimi, G., Runowicz, C. D., Gaudet, M. M., Jemal, A., & Siegel, R. L. (2018). Ovarian cancer statistics, 2018. CA: A Cancer Journal for Clinicians, 68(4), 284–296.
Al Habyan, S., Kalos, C., Szymborski, J., & McCaffrey, L. (2018). Multicellular detachment generates metastatic spheroids during intra-abdominal dissemination in epithelial ovarian cancer. Oncogene, 37(37), 5127–5135.
Kenny, H. A., Kaur, S., Coussens, L. M., & Lengyel, E. (2008). The initial steps of ovarian cancer cell metastasis are mediated by MMP-2 cleavage of vitronectin and fibronectin. The Journal of Clinical Investigation, 118(4), 1367–1379.
Izar, B., Tirosh, I., Stover, E. H., Wakiro, I., Cuoco, M. S., Alter, I., Rodman, C., Leeson, R., Su, M. J., Shah, P., Iwanicki, M., Walker, S. R., Kanodia, A., Melms, J. C., Mei, S., Lin, J. R., Porter, C. B. M., Slyper, M., Waldman, J., Jerby-Arnon, L., Ashenberg, O., Brinker, T. J., Mills, C., Rogava, M., Vigneau, S., Sorger, P. K., Garraway, L. A., Konstantinopoulos, P. A., Liu, J. F., Matulonis, U., Johnson, B. E., Rozenblatt-Rosen, O., Rotem, A., & Regev, A. (2020). A single-cell landscape of high-grade serous ovarian cancer. Nature Medicine, 26(8), 1271–1279.
Pearce, O. M. T., Delaine-Smith, R. M., Maniati, E., Nichols, S., Wang, J., Bohm, S., Rajeeve, V., Ullah, D., Chakravarty, P., Jones, R. R., Montfort, A., Dowe, T., Gribben, J., Jones, J. L., Kocher, H. M., Serody, J. S., Vincent, B. G., Connelly, J., Brenton, J. D., Chelala, C., Cutillas, P. R., Lockley, M., Bessant, C., Knight, M. M., & Balkwill, F. R. (2018). Deconstruction of a metastatic tumor microenvironment reveals a common matrix response in human cancers. Cancer Discovery, 8(3), 304–319.
Jimenez-Sanchez, A., Cybulska, P., Mager, K. L., Koplev, S., Cast, O., Couturier, D. L., Memon, D., Selenica, P., Nikolovski, I., Mazaheri, Y., Bykov, Y., Geyer, F. C., Macintyre, G., Gavarro, L. M., Drews, R. M., Gill, M. B., Papanastasiou, A. D., Sosa, R. E., Soslow, R. A., Walther, T., Shen, R., Chi, D. S., Park, K. J., Hollmann, T., Reis-Filho, J. S., Markowetz, F., Beltrao, P., Vargas, H. A., Zamarin, D., Brenton, J. D., Snyder, A., Weigelt, B., Sala, E., & Miller, M. L. (2020). Unraveling tumor-immune heterogeneity in advanced ovarian cancer uncovers immunogenic effect of chemotherapy. Nature Genetics, 52(6), 582–593.
Dean, M., Jin, V., Bergsten, T. M., Austin, J. R., Lantvit, D. D., Russo, A., & Burdette, J. E. (2019). Loss of PTEN in fallopian tube epithelium results in multicellular tumor spheroid formation and metastasis to the ovary. Cancers (Basel), 11(6), 884.
Singh, M. S., Goldsmith, M., Thakur, K., Chatterjee, S., Landesman-Milo, D., Levy, T., Kunz-Schughart, L. A., Barenholz, Y., & Peer, D. (2020). An ovarian spheroid based tumor model that represents vascularized tumors and enables the investigation of nanomedicine therapeutics. Nanoscale, 12(3), 1894–1903.
Tomas-Bort, E., Kieler, M., Sharma, S., Candido, J. B., & Loessner, D. (2020). 3D approaches to model the tumor microenvironment of pancreatic cancer. Theranostics, 10(11), 5074–5089.
Yang, Z., Xu, H., & Zhao, X. (2020). Designer self-assembling peptide hydrogels to engineer 3D cell microenvironments for cell constructs formation and precise oncology remodeling in ovarian cancer. Advanced Science (Weinh), 7(9), 1903718.
Loessner, D., Stok, K. S., Lutolf, M. P., Hutmacher, D. W., Clements, J. A., & Rizzi, S. C. (2010). Bioengineered 3D platform to explore cell-ECM interactions and drug resistance of epithelial ovarian cancer cells. Biomaterials, 31(32), 8494–8506.
Kaemmerer, E., Melchels, F. P., Holzapfel, B. M., Meckel, T., Hutmacher, D. W., & Loessner, D. (2014). Gelatine methacrylamide-based hydrogels: An alternative three-dimensional cancer cell culture system. Acta Biomaterialia, 10(6), 2551–2562.
Naba, A., Pearce, O. M. T., Del Rosario, A., Ma, D., Ding, H., Rajeeve, V., Cutillas, P. R., Balkwill, F. R., & Hynes, R. O. (2017). Characterization of the extracellular matrix of normal and diseased tissues using proteomics. Journal of Proteome Research, 16(8), 3083–3091.
Lohmussaar, K., Kopper, O., Korving, J., Begthel, H., Vreuls, C. P. H., van Es, J. H., & Clevers, H. (2020). Assessing the origin of high-grade serous ovarian cancer using CRISPR-modification of mouse organoids. Nature Communications, 11(1), 2660.
Hill, S. J., Decker, B., Roberts, E. A., Horowitz, N. S., Muto, M. G., Worley, M. J., Jr., Feltmate, C. M., Nucci, M. R., Swisher, E. M., Nguyen, H., Yang, C., Morizane, R., Kochupurakkal, B. S., Do, K. T., Konstantinopoulos, P. A., Liu, J. F., Bonventre, J. V., Matulonis, U. A., Shapiro, G. I., Berkowitz, R. S., Crum, C. P., & D’Andrea, A. D. (2018). Prediction of DNA repair inhibitor response in short-term patient-derived ovarian cancer organoids. Cancer Discovery, 8(11), 1404–1421.
Maru, Y., & Hippo, Y. (2019). Current status of patient-derived ovarian cancer models. Cell, 8(5), 505.
Lu, M., Henry, C. E., Lai, H., Khine, Y. Y., Ford, C. E., & Stenzel, M. H. (2019). A new 3D organotypic model of ovarian cancer to help evaluate the antimetastatic activity of RAPTA-C conjugated micelles. Biomaterials Science, 7(4), 1652–1660.
Mitra, S., Tiwari, K., Podicheti, R., Pandhiri, T., Rusch, D. B., Bonetto, A., Zhang, C., & Mitra, A. K. (2019). Transcriptome profiling reveals matrisome alteration as a key feature of ovarian cancer progression. Cancers (Basel), 11(10), 1513.
Loessner, D., Rockstroh, A., Shokoohmand, A., Holzapfel, B. M., Wagner, F., Baldwin, J., Boxberg, M., Schmalfeldt, B., Lengyel, E., Clements, J. A., & Hutmacher, D. W. (2019). A 3D tumor microenvironment regulates cell proliferation, peritoneal growth and expression patterns. Biomaterials, 190–191, 63–75.
Sontheimer-Phelps, A., Hassell, B. A., & Ingber, D. E. (2019). Modelling cancer in microfluidic human organs-on-chips. Nature Reviews. Cancer, 19(2), 65–81.
Ronaldson-Bouchard, K., & Vunjak-Novakovic, G. (2018). Organs-on-a-chip: A fast track for engineered human tissues in drug development. Cell Stem Cell, 22(3), 310–324.
Dorayappan, K. D. P., Gardner, M. L., Hisey, C. L., Zingarelli, R. A., Smith, B. Q., Lightfoot, M. D. S., Gogna, R., Flannery, M. M., Hays, J., Hansford, D. J., Freitas, M. A., Yu, L., Cohn, D. E., & Selvendiran, K. (2019). A microfluidic chip enables isolation of exosomes and establishment of their protein profiles and associated signaling pathways in ovarian cancer. Cancer Research, 79(13), 3503–3513.
Farkkila, A., Gulhan, D. C., Casado, J., Jacobson, C. A., Nguyen, H., Kochupurakkal, B., Maliga, Z., Yapp, C., Chen, Y. A., Schapiro, D., Zhou, Y., Graham, J. R., Dezube, B. J., Munster, P., Santagata, S., Garcia, E., Rodig, S., Lako, A., Chowdhury, D., Shapiro, G. I., Matulonis, U. A., Park, P. J., Hautaniemi, S., Sorger, P. K., Swisher, E. M., D’Andrea, A. D., & Konstantinopoulos, P. A. (2020). Immunogenomic profiling determines responses to combined PARP and PD-1 inhibition in ovarian cancer. Nature Communications, 11(1), 1459.
Kim, H., Xu, H., George, E., Hallberg, D., Kumar, S., Jagannathan, V., Medvedev, S., Kinose, Y., Devins, K., Verma, P., Ly, K., Wang, Y., Greenberg, R. A., Schwartz, L., Johnson, N., Scharpf, R. B., Mills, G. B., Zhang, R., Velculescu, V. E., Brown, E. J., & Simpkins, F. (2020). Combining PARP with ATR inhibition overcomes PARP inhibitor and platinum resistance in ovarian cancer models. Nature Communications, 11(1), 3726.
Sanij, E., Hannan, K. M., Xuan, J., Yan, S., Ahern, J. E., Trigos, A. S., Brajanovski, N., Son, J., Chan, K. T., Kondrashova, O., Lieschke, E., Wakefield, M. J., Frank, D., Ellis, S., Cullinane, C., Kang, J., Poortinga, G., Nag, P., Deans, A. J., Khanna, K. K., Mileshkin, L., McArthur, G. A., Soong, J., Berns, E., Hannan, R. D., Scott, C. L., Sheppard, K. E., & Pearson, R. B. (2020). CX-5461 activates the DNA damage response and demonstrates therapeutic efficacy in high-grade serous ovarian cancer. Nature Communications, 11(1), 2641.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Kast, V., Loessner, D. (2021). 3D Models for Ovarian Cancer. In: Schatten, H. (eds) Ovarian Cancer: Molecular & Diagnostic Imaging and Treatment Strategies. Advances in Experimental Medicine and Biology, vol 1330. Springer, Cham. https://doi.org/10.1007/978-3-030-73359-9_9
Download citation
DOI: https://doi.org/10.1007/978-3-030-73359-9_9
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-73358-2
Online ISBN: 978-3-030-73359-9
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)