Abstract
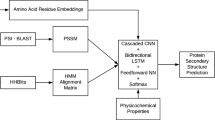
In recent years, deep learning (DL) techniques have been applied in the structural and functional analysis of proteins in bioinformatics, especially in 8-state (Q8) protein secondary structure prediction (PSSP). In this paper, we have explored the performance of various DL architectures for Q8 PSSP, by developing six DL architectures, using convolutional neural networks (CNNs), recurrent neural networks (RNNs), and combinations of them. These architectures are: CNN-SW (CNNs with sliding window); CNN-WP (CNNs with whole protein as input); LSTM+ (Long Short-Term Memory (LSTM) & Bidirectional LSTM (BLSTM)); GRU+ (Gated Recurrent Unit (GRU) & bidirectional GRU (BGRU)); CNN-BGRU (CNNs & BGRUs); and CNN-BLSTM (CNNs & BLSTMs). They include batch normalization, dropout, and fully-connected layers. We have used CB6133 and CB513 datasets for training and testing, respectively. The experiments showed that combining CNN with BLSTM or BGRU overcame overfitting, and achieved better Q8 accuracy, precision, recall and F-score. The experiments on CB513 showed that CNN-SW, CNN-BGRU, and CNN-BLSTM achieved Q8 accuracy comparable with some state-of-the-art models.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Noble, M.E., Endicott, J.A., Johnson, L.N.: Protein kinase inhibitors: insights into drug design from structure. Science 303(5665), 1800–1805 (2004)
Zhou, J., Wang, H., Zhao, Z., Xu, R., Lu, Q.: CNNH_PSS: protein 8-class secondary structure prediction by convolutional neural network with highway. BMC Bioinform. 19(60), 99–119 (2018)
Kabsch, W., Sander, C.: Dictionary of protein secondary structure: pattern recognition of hydrogen-bonded and geometrical features. Biopolymers 22(12), 2577–2637 (1983)
Zhou, J., Troyanskaya, O.G.: Deep supervised and convolutional generative stochastic network for protein secondary structure prediction. In: 31st International Conference on Machine Learning (ICML 2014), pp. 745–53. PMLR, Bejing (2014)
Li, Z., Yu, Y.: Protein secondary structure prediction using cascaded convolutional and recurrent neural networks. In: 25th International Joint Conference on Artificial Intelligence (IJCAI 2016), pp. 2560–2567. AAAI Press, California (2016)
LeCun, Y., Bottou, L., Bengio, Y., Haffner, P.: Gradient-based learning applied to document recognition. Proc. IEEE 86(11), 2278–2324 (1998)
Hochreiter, S., Schmidhuber, J.: Long short-term memory. Neural Comput. 9(8), 1735–1780 (1997)
Cho, K., Merrienboer, B., Gulcehre, C., Bahdanau, D., Bougares, F., Schwenk, H., Bengio, Y.: Learning phrase representations using RNN encoder–decoder for statistical machine translation. In: 2014 Conference on Empirical Methods in Natural Language Processing (EMNLP), pp. 1724–1734. Association for Computational Linguistics, Doha (2014)
Schuster, M., Paliwal, K.K.: Bidirectional recurrent neural networks. IEEE Trans. Signal Process. 45(11), 2673–2681 (1997)
Sφnderby, S.K., Winther, O.: Protein secondary structure prediction with long short term memory networks. arXiv:1412.7828v2 [q-bio.QM] (2014)
Wang, S., Peng, J., Ma, J., Xu, J.: Protein secondary structure prediction using deep convolutional neural fields. Sci. Rep. 6, Article number 18962 (2016)
Busia, A., Jaitly, N.: Next-step conditioned deep convolutional neural networks improve protein secondary structure prediction. In: Conference on Intelligent Systems for Molecular Biology and European Conference on Computational Biology (ISMB/ECCB 2017). International Society of Computational Biology, Leesburg (2017)
Heffernan, R., Yang, Y., Paliwal, K., Zhou, Y.: Capturing non-local interactions by long short-term memory bidirectional recurrent neural networks for improving prediction of protein secondary structure, backbone angles, contact numbers and solvent accessibility. Bioinformatics 33(18), 2842–2849 (2017)
Fang, C., Shang, Y., Xu, D.: MUFOLD-SS: new deep inception-inside-inception networks for protein secondary structure prediction. Proteins 86(5), 592–598 (2018)
Zhang, B., Li, J., Lü, Q.: Prediction of 8-state protein secondary structures by a novel deep learning architecture. BMC Bioinform. 19(293), 1–13 (2018)
Kumar, P., Bankapur, S., Patil, N.: An enhanced protein secondary structure prediction using deep learning framework on hybrid profile based features. Appl. Soft Comput. J. 86(105926), 1–10 (2020)
Brownlee, J.: Better Deep Learning: Train Faster, Reduce Overfitting, and Make Better Predictions. v1.7. edn. Machine Learning Mastery, Vermont (2020)
Ioffe, S., Szegedy, C.: Batch normalization: accelerating deep network training by reducing internal covariate shift. arXiv preprint arXiv:1502.03167 (2015)
Brownlee, J.: How to Reduce Overfitting with Dropout Regularization in Keras. https://machinelearningmastery.com/how-to-reduce-overfitting-with-dropout-regularization-in-keras/. Accessed 3 Oct 2019
Long short-term memory, From Wikipedia, https://en.wikipedia.org/wiki/Long_short-term_memory. Accessed 29 Aug 2019
Brownlee, J.: How to Develop a Bidirectional LSTM for Sequence Classification in Python with Keras. Long Short-Term Memory Networks. Accessed 16 June 2017
Wang, G., Dunbrack, R.L.: PISCES: a protein sequence culling server. Bioinformatics 19(12), 1589–1591 (2003)
Li, W., Godzik, A.: Cd-hit: a fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics 22(13), 1658–1659 (2006)
Altschul, S.F., Gertz, E.M., Agarwala, R., Schaaffer, A.A., Yu, Y.K.: PSI-Blast pseudo counts and the minimum description length principle. Nucleic Acids Res. 37(3), 815–824 (2009)
Chen, H., Zhou, H.X.: Prediction of solvent accessibility and sites of deleterious mutations from protein sequence. Nucl. Acids Res. 33(10), 3193–3199 (2005)
Narkhede, S.: Understanding AUC - ROC Curve. https://towardsdatascience.com/understanding-auc-roc-curve-68b2303cc9c5. Accessed 26 June 2018
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 The Editor(s) (if applicable) and The Author(s), under exclusive license to Springer Nature Switzerland AG
About this paper
Cite this paper
Girgis, M.R., Elgeldawi, E., Gamal, R.M. (2021). A Comparative Study of Various Deep Learning Architectures for 8-state Protein Secondary Structures Prediction. In: Hassanien, A.E., Slowik, A., Snášel, V., El-Deeb, H., Tolba, F.M. (eds) Proceedings of the International Conference on Advanced Intelligent Systems and Informatics 2020. AISI 2020. Advances in Intelligent Systems and Computing, vol 1261. Springer, Cham. https://doi.org/10.1007/978-3-030-58669-0_45
Download citation
DOI: https://doi.org/10.1007/978-3-030-58669-0_45
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-58668-3
Online ISBN: 978-3-030-58669-0
eBook Packages: Intelligent Technologies and RoboticsIntelligent Technologies and Robotics (R0)