Abstract
Entomopathogenic nematodes are parasitic organisms with an exceptional capacity to infect rapidly and efficiently a wide range of insect species. Their distinct pathogenic properties have established entomopathogenic nematodes as supreme biocontrol agents of insects as well as excellent models to simulate and dissect the molecular and physiological bases of conserved strategies employed by parasitic nematodes that cause infectious diseases in humans. The extreme infectivity of entomopathogenic nematodes is due in part to the presence of certain species of Gram-negative bacteria that live in mutualistic symbiosis during the infective juvenile stage, which forms the central part of the nematode life cycle. Both nematodes and their mutualistic bacteria are capable of interfering and undermining several aspects of the insect host innate immune system during the infection process. The mutualistic bacteria are also able to modulate other biological functions in their nematode host including growth, development, and reproduction. In this review, we will focus our attention on the mutualistic relationship between entomopathogenic nematodes and their associated bacteria to discuss the nature and distinct characteristics of the regulatory mechanisms, and their molecular as well as physiological components that control this specific biological partnership.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
Bacteria–host interactions are ubiquitous in nature (Ruby 2008). They form complex relationships that influence critical biological processes such as the nutrition, development, and immunity of plants and animals (Hentschel et al. 2000; Ochman and Moran 2001). Relationships range from being ancient, stable, and beneficial mutualisms, as exemplified by the origin of the mitochondria and chloroplasts from the endosymbiosis, to those which are more recent, dynamic, and highly pathogenic, such as the evolution of Yersinia pestis, the causative agent of plague, from a relatively benign ancestor (Sagan 1967; Parkhill et al. 2001). Although the outcomes of these interactions have very different consequences for their hosts, there is increasing evidence that common mechanisms regulate the ability of bacteria to act as either mutualists or pathogens. Both lifestyles are thought to have evolved from living in close proximity to their hosts and both require the ability to circumvent host immunity and modulate the host environment (Dale and Moran 2006). The diversity of these associations and their importance to medicine and agriculture define them as a key area for research (Ochman and Moran 2001; Maurelli 2007). Recent works have adopted model organisms to determine the overlap between the nature of molecular signaling pathways and their specific genes that are necessary for regulating mutualism and pathogenicity lifestyles in bacteria and their invertebrate hosts. Understanding the genetic mechanisms and dictating the outcomes of bacteria–host interactions will ultimately allow us to determine how microbes switch from one lifestyle to another, thus shedding light on the evolution of complex multi-organism relationships.
Insight into the delicate balance between mutualism and pathogenicity requires a system that allows for the direct study of both interactions (Chaston and Goodrich-Blair 2010). Entomopathogenic nematodes are microscopic worms that target and naturally infect a diverse range of insect hosts, and therefore, they have been implemented in modern agricultural practices as promising biological control agents and alternatives to chemical insecticides for managing destructive insect pests of plants and deleterious vectors of infectious diseases. Due to their remarkable pathogenic properties toward various insect stages, and their unique life cycle that involves mutualistic cooperation with specific bacterial species, entomopathogenic nematodes have been employed in recent years in biomedical research as outstanding and simultaneously environmentally safe tools for unraveling the molecular and physiological basis of mutualistic relationships in animals, and resolving pathogenicity mechanisms in nematode–bacterial complexes, in relation to the host innate immune function. The mutualistic bacteria of entomopathogenic nematodes perform critical biological tasks that are not restricted only to promoting pathogenicity and compromising the insect immune system during infection, but they also protect their nematode host by producing antimicrobial molecules to support the growth of other competitive bacteria. In addition, they promote nematode dispersal and development, growth, and reproduction by supplying nutrients from the bioconverted insect tissues and organs, as well as by acting themselves as a rich food source (Herbert and Goodrich-Blair 2007a).
Application of entomopathogenic nematodes in the field requires careful analysis of various traits of parasites and their associated bacteria acting together as a complex or as a separate one, during the distinct phases of their life cycle. Due to the lack of understanding of the molecular and physiological determinants that control the harmonious coordination between the two mutualistic players, it is considered imperative to invest future efforts and resources on deconstructing the life cycle of different entomopathogenic nematode species to expose the exact elements that enhance or diminish the interaction with the insect host. This approach would, in turn, provide us with the necessary knowledge to validate the attributes that increase the performance of entomopathogenic nematodes, and improve their stability and efficiency in the field. Such information would ultimately enable us to make convenient interventions to refine biocontrol programs that would reduce pesticide use and improve food safety and production.
2 Entomopathogenic Nematode–Bacterial Complexes
Nematode–bacterial complexes with insect pathogenic properties are formed specifically in the soil nematodes of the genera Heterorhabditis and Steinernema, which develop mutualistic relationship with the proteobacteria Photorhabdus spp. and Xenorhabdus spp., respectively. The nematodes together with their associated bacteria undergo a complex life cycle that comprises two stages; a mutualistic stage that takes place in the nematode gut, during which the bacteria are vectored by their cognate nematode and a pathogenic stage that occurs in the insect host during infection, and involves the manipulation of humoral and cellular innate immune defenses by both partners that lead to the accelerated insect death. Although entomopathogenic nematode life cycles exhibit similar characteristics, variation especially in certain features of nematode reproduction and population growth rate, as well as in host range and phase variants of mutualistic bacteria, can be observed among different genera and species (Forst et al. 1997).
Heterorhabditis nematodes from the Heterorhabditidae family act as ‘cruiser’ parasites, a behavior that involves active seeking out of suitable insect hosts by burrowing into the soil. Heterorhabditis parasitic nematodes form a mutually beneficial symbiotic relationship with the entomopathogenic Gram-negative bacteria from the genus Photorhabdus, which belong to the Enterobacteriaceae family (Waterfield et al. 2009). The bacteria are found in the gut of the infective juvenile stage (Ciche et al. 2006; Ciche 2007). The infective juvenile is an obligate stage in the nematode life cycle and is required for the infection of larval stages of mainly lepidopteran insects (Ciche 2007; Kaya and Gaugler 1993) (Fig. 17.1). This stage is analogous to the C. elegans dauer stage and the developmentally arrested infective third-stage larva (L3) of many important parasitic nematodes. Infective juveniles gain entry to the insect through natural openings (anus, spiracles, and mouth) or by abrading the insect cuticle using a dorsal tooth (Ciche 2007). Once inside the insect, the infective juveniles expel a small number of Photorhabdus cells into the hemolymph where the bacteria begin to divide exponentially. After two to three days of bacterial growth, the insect succumbs to the infection due to septicemia with the concomitant conversion of the internal organs and tissues into bacterial biomass. This bioconversion is facilitated by the production of a wide range of toxins and hydrolytic enzymes by the bacteria (Ffrench-Constant et al. 2007; Eleftherianos 2009; Bode 2009). The worms feed on the bacterial biomass, and subsequent nematodes’ growth and development require the presence of high-density Photorhabdus bacteria (Ciche and Ensign 2003). The infective juvenile nematodes mature to first-generation hermaphrodite females, which give rise to the second generation of amphimictic males and females (cross-fertilization) and to the self-fertile hermaphrodite females and infective juveniles. Nematodes reproduce and the progeny develops through four juvenile stages (L1, L2, L3, and L4) to adults. Nematode reproduction continues over two to three generations until the nutrient status of the cadaver deteriorates, whereupon adult development is suppressed, and the infective juvenile stage accumulates. These non-feeding infective juveniles enter the soil where they may survive for several months in the absence of a suitable host. The transmission of mutualistic bacteria by infective juveniles is essential for the nematodes to reproduce (Goodrich-Blair and Clarke 2007).
Life cycle of the entomopathogenic nematode Heterorhabditis bacteriophora. The infective juveniles (IJs) form the only free-living stage of this parasite. Nematode mating, reproduction, and development occur within the hemolymph of the infected insects (e.g. larvae of the greater wax moth, Galleria mellonella) in the presence of high titers of their mutualistic bacteria Photorhabdus luminescens. Depending on the amount of resources in the dead insect, two or three generations may take place within the insect cadavers. L1, L2, L3, and L4: Larval molts. Images are made using Biorender graphic software (https://biorender.com)
Similar to Heterorhabditis, Steinernema nematodes are found free in the soil where they gain access to insect larvae through natural body openings but they lack a dorsal tooth that facilitates penetration through the cuticle. However, in contrast to Heterorhabditis, the Steinernema nematodes exhibit ‘ambushing’ behavior, which involves waiting and attacking sensitive insect hosts in their vicinity. Steinernema nematodes follow a similar life cycle to Heterorhabditis that mainly differs in the initial stage of recovery during which amphimictic reproduction occurs. This means that Steinernema infective juveniles develop into reproductive males and females. Interestingly, this infective juvenile behavior also takes place during the first- and second-generation offspring. In addition, third-generation females produce eggs all of which develop through the ‘endotokia matricida’ process that occurs due to the cessation of egg laying and involves intra-uterine birth causing maternal death, a relatively common phenomenon in entomopathogenic nematodes induced in response to the low food supply. As opposed to Heterorhabditis, the resulting juvenile stages of Steinernema develop into infective juveniles after they exit the mother nematode (Kooliyottil et al. 2013).
3 Mutualism Regulators in Photorhabdus Bacteria
Recent progress in quantitative proteomic techniques has been started to contribute to the identification and preliminary examination of the factors that control symbiotic processes between animal hosts and microbes, including entomopathogenic nematodes and their related bacteria. To determine the identity of bacterial proteins that underlie symbiotic specificity in the entomopathogenic nematodes Heterorhabditis, 2D-gel electrophoresis followed by mass spectrometry were used to analyze and compare the proteomic profiles of two P. luminescens subspecies (P. luminescens ssp. laumondii and P. luminescens ssp. akhurstii), each occupying a distinct Heterorhabditis nematode species (H. bacteriophora and H. indica, respectively) (Kumar et al. 2016). Results from the proteomic and bioinformatic analyses revealed that either bacterial subspecies expresses several unique proteins, a subset of which (e.g. outer membrane proteins, proteins regulating secondary metabolites, and hypothetical proteins) may define nematode specificity. The functional characterization of certain candidate proteins will undoubtedly provide clues on the evolutionary and mechanistic basis of host–symbiont associations (Fig. 17.2).
Photorhabdus molecular regulators of symbiosis. Growth and reproduction of Heterorhabditis bacteriophora nematodes are controlled by genes rpoB, ngrA, and stlA in P. luminescens. Recovery of nematode infective juveniles (IJ) and bacterial metabolism are controlled by P. luminescens gene stlA. The madswitch promoter and gene hexA regulate the two distinct forms (mutualistic and pathogenic) in P. luminescens and P. temperata bacteria, respectively. The fimbrial locus mad in P. luminescens participates in initiating symbiosis through bacterial colonization of the posterior maternal intestinal cells in H. bacteriophora. LPS biosynthesis is modulated by genes galE, galU, and pbgPE in P. luminescens. Biofilm formation in P. luminescens is modulated by genes galU, proQ, and stlA. Images are made using Biorender graphic software (https://biorender.com)
Proteomic analysis coupled with bacterial genetics has further explored the role of the rpoB gene in the symbiosis between P. luminescens LN2 bacteria and their H. bacteriophora H06 nematode vectors (Qiu et al. 2012). Gene rpoB codes for the bacterial beta subunit of RNA polymerase and interestingly rifampicin prevents the initiation of transcription by repressing the rpoB gene. This research showed that certain rifampicin-resistant P. luminescens LN2 mutant strains, which surprisingly also contained mutations in the rpoB gene, were able to support the growth of H. bacteriophora H06 infective juveniles. It was further demonstrated that mutations in the rpoB gene reconstitute the bacteria as the nutrient source for sustaining nematode reproduction; however, without conferring the ability of the bacteria to colonize the nematode intestines during the infective juvenile stage. Rifampicin selection of P. luminescens rpoB mutant strains supporting nematode growth may provide an elegant approach for increasing the production of H. bacteriophora in order to achieve more efficient insect pest control in the field.
Genetic analysis of the nematode–bacterial symbiotic relationship using a transposon mutagenesis and screening approach identified a single mutant strain of P. luminescens that was deficient in providing growth and reproduction to the H. bacteriophora nematode vector (Ciche et al. 2001). Characterization of the mutation localized the transposon insertion into gene ngrA encoding the enzyme Ppant transferase, which is involved in the biosynthesis of the siderophore enterobactin. Although this mutation also conferred an inability of the bacteria to produce antibiotics and siderophores, and probably interrupted the biosynthesis of fatty acids or lipids, these deficiencies were not attributed to the nematode defects. Instead, the assumption is that the inactivation of the ngrA gene possibly affects the biosynthesis of hormones, polyketides, or other secondary metabolites that are produced by P. luminescens when H. bacteriophora is also present, and act as signal molecules to promote the nematode’s growth and development.
A subsequent study continued this work to examine whether gene ngrA encoding a putative phosphopantetheinyl transferase (PPT) that is involved in the biosynthesis of siderophore forms a determining factor for P. luminescens to support the growth and reproduction of H. bacteriohora nematodes (Ciche et al. 2003). Following a mini-Tn5 mutagenesis approach, P. luminescens mutant strain NS414 with a deficiency in producing measurable siderophore activity was first isolated, and then its properties were characterized. The results showed that the mutant bacteria were not able to grow normally in media depleted of iron, but they were capable of promoting the growth and reproduction of their nematode hosts, as well as their transmission by H. bacteriophora infective juveniles. Interestingly, the transposon was found to be inserted into gene photobactin synthetase (phbH) encoding a putative peptidyl carrier protein, which is covalently modified by PPTase for siderophore production. As phbH is not essential for nematode symbiosis, these findings signify that failure of the ngrA mutants to support nematode symbiosis was not due to their inability to produce functional siderophore but rather due to their incapacity to synthesize another currently unknown peptide that performs this function.
An interesting feature of Photorhabdus is that the bacteria can exist in two forms, the primary and secondary, which are morphologically distinct and are associated with the different phases of the pathogen’s lifestyle. Only the primary form bacteria can colonize the intestinal tract of their associated nematode host and promote its growth and development due to the production of extracellular enzymes and antibiotic compounds that support the interaction between the two symbiotic partners during the infection of a suitable insect (Waterfield et al. 2009). Remarkably, it has been previously shown that inactivation through transposon insertion of gene transcriptional regulator LrhA (hexA) in the secondary phase of P. temperata bacteria results in the suppression of nematode colonization, and concomitantly, the mutant bacteria can foster growth and development of the host nematodes H. downesi (Joyce and Clarke 2003). These findings provide proof that hexA in the secondary phase of P. temperata bacteria encodes a molecule that confers direct or indirect repressive effects on symbiotic factors, which are normally expressed in the primary phase variants.
Another library screen of GFP-labeled P. luminescens transposon mutants, involving symbiotic assays to examine the qualitative ability of the mutant bacteria to colonize the gut of the infective juvenile stage of H. bacteriophora nematodes, further aimed at identifying bacterial genes, and their encoded factors responsible for the symbiotic collaboration between the two organisms (Easom et al. 2010). This work showed that mutations in a subset of genetic loci (e.g. pbgPE operon and genes galE and galU) involved specifically in the biosynthesis of lipopolysaccharide (LPS) and assembly and maintenance of LPS structure, as well as of other bacterial cell surface components, conferred substantially reduced transmission frequency of the mutant bacteria to associate with their nematode host. In addition, the P. luminescens mutant for genes proQ (encoding an RNA chaperone) and galU were also defective in biofilm formation as shown through testing the ability of the mutant bacteria to attach to an abiotic surface. This information highlights further the vital role of cell surface molecules in P. luminescens, and probably in other entomopathogenic bacteria, in adjusting the symbiotic outcome of bacterial–nematode partnerships.
Using a similar random transposon mutagenesis screening approach, the transmission ability of GFP-labeled P. luminescens mutants in the intestine of H. bacteriophora nematode parasites was analyzed in detail. The genetic analysis detected that the maternal adhesion defective (mad) fimbrial locus in P. luminescens has an essential role in initiating symbiosis through the bacterial colonization of the posterior maternal intestinal cells in H. bacteriophora. This process facilitates bacterial symbiont transmission from the maternal nematodes to the infective juveniles. Importantly, this is a specialized function because mad is required for symbiosis but not for insect pathogenesis, and the effect is regulated by bacterial phase variation in the wild type bacteria but not in the mad mutants (Somvanshi et al. 2010). These are the findings of particular significance because although fimbriae are known colonization factors that were previously shown to promote animal tissue or cell colonization by various bacterial pathogens through receptor recognition events, this was the first time that these adhesive organelles were assigned a similar function in modulating nematode–bacteria mutualistic symbiosis.
A previous study identified the production of crystalline inclusion proteins containing high levels of essential amino acids by P. luminescens bacteria to assist nematode reproduction (Bintrim and Ensign 1998), and a more recent work linked the two distinct forms of P. luminescens (M, initiating nematode Mutualism and P, initiating insect Pathogenicity) with the expression of the mad fimbrial locus, which occurs after the inversion of the madswitch promoter (Somvanshi et al. 2012). More precisely, it was demonstrated that during the first stage of mutualism in H. bacteriophora, P. luminescens bacteria switch to the M-form in the posterior intestine of the maternal nematodes, while the P-form bacteria are temporarily present in the intestines. The M-form cells then occupy the intestines of the new generation of infective juveniles before turning into the P-form cells to provide nematodes with bacteria possessing properties that promote infection of susceptible insects. Strikingly, the M-form bacteria are smaller than the P-form bacteria; they grow slower and exhibit decreased bioluminescence, virulence, and ability to secrete secondary metabolic compounds. Therefore, the biological implication of these findings underlines the importance of madswitch promoter orientation, which defines not only the phenotypic appearance of P. luminescens cells but also influences the lifestyle of the bacteria.
Additional factors have been associated with the persistence of P. temperata in the intestine of H. bacteriophora nematodes, as demonstrated by the drastic changes in the transcriptional profile of the bacteria during mutualism in the non-feeding infective juvenile stage (An and Grewal 2010). To determine strategies adapted by the bacteria to strengthen their persistence in the parasite through diminishing nutritional reliance on the nematode host, the number and identity of the differentially expressed genes in P. temperata was explored using the selective capture of the transcribed sequences technique. Analysis of the results displayed a large number of differentially regulated genes in P. temperata when the bacteria reside in the nematode intestine compared to the bacteria being cultured in vitro or being present in the insect host hemolymph. The differentially expressed genes denote modifications in physiological functions when residing in the nematodes’ intestine including the activation of the pentose phosphate pathway, alteration in amino acid metabolisms, modification in LPS, induction of intracellular acidification and urea cycle mechanisms, proton transport and biofilm formation, as well as processes involving bacterial replication, transcription, and translation. These findings provide an exciting pool of potential molecular regulators of bacterial symbiosis in parasitic nematodes and future work awaits to dissect the specific mechanistic roles of these symbiosis factors and whether (and how) they are interconnected to enable the close biological link between the two organisms.
P. luminescens is a bacterium that is able to produce the antibiotic 3,5-dihydroxy-4-isopropylstilbene (stilbene, ST) during insect infection. The production of this antibiotic compound eliminates non-symbiotic micro-organisms that colonize insect tissues to compete for resources and nutrients, it prevents decay of the insect carcass and therefore, provides a favorable environment to their H. bacteriophora nematode vectors to grow, replicate, and complete their life cycle (Hu and Webster 2000). ST functions as a signal for the nematodes by stimulating the recovery of infective juveniles to adult hermaphrodites. This was demonstrated by the finding that P. luminescens mutants deficient in ST production were also unable to support H. bacteriophora growth and development (Joyce et al. 2008). The biosynthesis of ST involves the non-oxidative deamination of phenylalanine that leads to the synthesis of cinnamic acid by the enzyme phenylalanine-ammonium lyase, which is encoded by the gene stlA (Williams et al. 2005). Interestingly, it was subsequently shown that stlA expression is temporally controlled during growth, and it can be regulated by nutrient limitation (Lango-Scholey et al. 2013). This gene regulatory mechanism is further controlled by three transcriptional regulators; LysR-type TyrR, which is absolutely essential for stlA expression, as well as Leucine-responsive regulatory protein (Lrp) and RNA polymerase Sigma factor (rpoS), which are also required for normal stlA expression under suitable environmental conditions. These findings signify the molecular players that modulate secondary metabolism, and as a consequence, the mutualism of a potent entomopathogenic bacterium with its nematode host. Recently, additional findings provided evidence for the role of P. luminescens stilbene in the biology of the bacteria and their association with H. bacteriophora nematodes (Hapeshi et al. 2019). Exogenous ectopic addition of stilbene to P. luminescens stlA mutants reduces biofilm formation and downregulates the transcriptional expression of genes participating in the secondary metabolism, and basic cellular processes. These findings illustrate that stilbene cannot be only produced but also be detected by P. luminescens and plays a modulatory role by possibly acting as a signal for the bacteria to regulate the symbiotic phase of their life cycle through promoting the production of other molecules that are important for the recovery of nematode infective juveniles. This is a crucial process because it facilitates bacterial transmission to the next insect host following parasitic nematode infection.
4 Mutualism Regulators in Xenorhabdus Bacteria
Similar to Photorhabdus, Xenorhabdus bacteria are considered to have evolved distinct physiological and metabolic mechanisms that facilitate the close association with their cognate nematode hosts, as revealed by a previous study comparing the genome sequences of the two entomopathogenic bacterial symbionts (Chaston et al. 2011). The current speculation is that the two bacteria most likely share a single progenitor and as a result of multiple selective pressure events following differentiation, they have acquired unique factors to support the close relationship with their related nematode vectors.
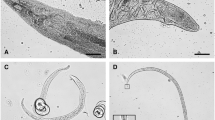
Transposon mutagenesis to detect factors in X. nematophila that promote symbiosis with Steinernema carpocapsae nematodes led to the isolation of a mutant strain that was able to produce certain phospholipases such as lecithinase, but was unable to produce antibiotics and the mutant bacteria failed to grow and emerge normally from their nematode host (Volgyi et al. 2000) (Fig. 17.3). The transposon mutation was detected in the gene var1 encoding a protein that is involved in the formation of the variant cell type. Deficient growth of the mutant bacteria may suggest a negative effect on the survival of this variant type of cells in the nematode intestines or the inability of the bacteria to swarm properly, which may delay their exit from the nematode. Although the specific physiological and biochemical bases of this phenotype in X. nematophila is unclear, further elucidation of the processes and the particular factors modulating the switch from primary to secondary phase cells will enhance our knowledge regarding the type of features that are required to support the symbiotic interactions between entomopathogenic nematodes and their affiliated bacteria.
Xenorhabdus molecular regulators of symbiosis. Filamentation and replication in X. nematophila are regulated by gene cpxRA. Bacterial growth and survival are controlled by gene var1. Colonization of Steinernema carpocapsae infective juveniles by X. nematophila is controlled by the bacterial genes nilD and tdk. Growth of competitor microbes in the dead insect is suppressed by phenazine compounds produced by X. nematophila as well as by gene ngrA. The latter also modulates the development and emergence of S. carpocapsae nematodes. Images are made using Biorender graphic software (https://biorender.com)
During the symbiotic relationship with S. carpocapsae, the X. nematophila uses nutrients from their nematode host; however, the exact identity of these compounds required for bacterial growth during this interaction is not well-determined. Since bacteria can utilize salvaged nucleosides as a supplement to endogenous nucleotides for DNA synthesis, they are able to form nitrogen sources, they can participate in the activation of signal transduction, and they are involved in the construction of cell structures. Therefore, it was previously hypothesized that the regulation of pyrimidine salvage pathways might constitute a process that facilitates the X. nematophila-nematode interplay (Orchard and Goodrich-Blair 2005). Curiously, it has been shown that the X. nematophila mutant for gene tdk encoding the enzyme deoxythymidine kinase, which synthesizes the pyrimidine nucleotide deoxythymidine monophosphate from deoxythymidine, are deficient in nematode colonization in vitro but not when present in the insect host. This defect is also fully restored by the addition of the wild copy tdk allele to the mutant strain. Such a mechanism could represent a broader strategy for entomopathogenic bacteria to associate with their nematode hosts.
An important aspect to consider in host–microbe interactions is the ability of the organisms involved in symbiotic relationships to sense and respond to external environmental changes. The two-component regulatory system CpxRA in X. nematophila consists of a sensor histidine kinase (CpxA) and a cytoplasmic response regulator (CpxR). It forms a signaling pathway, which in other bacteria, such as E. coli, regulates the function of structural components that permit interaction with the host. They might also act as a transducer to transmit internal signals inside the cells for initiating signaling pathways that would generate a cellular response (Herbert and Goodrich-Blair 2007b). Testing the symbiotic competence of X. nematophila cpxR1 mutant bacteria, in which expression of both cpxA and cpxP genes is abolished, revealed that their ability to associate with S. carpocapsae infective juveniles is markedly impaired, and this effect is not due to a reduced survival ability of the mutants. Alternatively, this effect is primarily associated with modifications in cell morphological features in cpxR1 mutants that might alter cell division dynamics or cause filamentation, which in turn could lead to interference in the symbiotic partnership between the bacteria and their nematodes, as well as with the decreased expression of genes that participate in nematode colonization. The regulatory role of CpxRA in X. nematophila is not unique, as other genes including Lrp (Leucine responsive regulatory protein) have also been shown to possess regulatory properties (Cowles et al. 2007). Overall, these findings imply the presence of certain genes in the X. nematophila genome performing multiple activities to promote the symbiotic connection of the bacteria with their affiliated entomopathogenic nematode partners.
It is given that molecules with antimicrobial activity may also modulate molecular and physiological processes by acting as signals in certain bacteria, previous efforts have focused on the role of bacterial secondary metabolites in promoting symbiotic relationships. This is because the P. luminescens ngrA gene, which encodes various secondary metabolites, is important for the growth and reproduction of H. bacteriophora nematodes, the involvement of X. nematophila ngrA in establishing or maintaining symbiosis with S. carpocapsae nematodes was also analyzed (Singh et al. 2015). Results from this research indicated that the number of nematode progenies from Manduca sexta caterpillars inoculated with S. carpocapsae infective juveniles containing the X. nematophila ngrA mutant were remarkably decreased compared to nematode progeny containing X. nematophila wild-type bacteria. These findings support the notion of a dual role for ngrA-derived compounds in not only combating competitor microbes in the insect cadaver to facilitate nematode development but also acting as signals for accelerating nematode development and emergence from the infected insect host.
Other molecules with multiple activities in Xenorhabdus that might serve as crucial factors for providing efficient bacteria–nematode symbiotic cooperation are the phenazines, which are commonly found in Gram-negative bacteria (Shi et al. 2019). It was recently tested whether phenazine compounds derived from X. szentirmaii influence the symbiotic ability between the bacteria and their S. rarum nematode vectors. Although the exact function of these molecules in regulating bacteria–nematode interactions is currently obscure because phenazines possess broad-spectrum as well as specific antibiotic activity, the current working hypothesis is that these compounds target a variety of competitor soil microbes present in the insect carcass. Elimination of these competitor microbes enables the S. rarum parasitic nematodes to complete and maintain their life cycle together with their closely related X. szentirmaii bacteria.
Further efforts to detect bacterial molecular factors promoting nematode colonization have included a signature-tagged mutagenesis screen in X. nematophila, an approach that identified a transposon mutant that lost its ability to colonize infective juveniles of S. carpocapsae nematodes (Veesenmeyer et al. 2014). Of note, the transposon was found to be inserted into gene nilD that codes for a Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR) element. CRISPR associated sequences (Cas) are widely found in bacteria and perform several cellular functions including the regulation of gene expression and DNA repair mechanisms as well as bacterial behavior and resistance to foreign nucleotide sequences (Li and Peng 2019; Hampton et al. 2020). The investigators were able to demonstrate elegantly that the nilD CRISPR sequence is sufficient to support the colonization of S. carpocapsae, as well as S. anatoliense and S. websteri nematodes. The nilD RNA is expressed in a Cas6e-dependent manner under in vitro growth conditions and during nematode symbiosis, but in the latter case only within a specific genetic background of X. nematophila. These exciting new findings open novel avenues of investigation for designing strategies to decode the precise mechanism of CRISPR bacterial systems in modulating symbiotic interdependence with entomopathogenic nematodes.
5 Development of Tools for Identifying Nematode Regulators of Mutualism
Although most information currently available on the number and nature of genes and their products acting as regulators of mutualism with entomopathogenic nematodes has been obtained from bacteria, knowledge of analogous molecules in parasites playing a central or complementary role to the evolution, and stability of this process, is missing so far. This, at least till now, was mostly attributed to the lack of availability of genetic and genomic tools in entomopathogenic nematodes, which has impeded progress with dissecting the molecular basis of the mutualistic interaction between the two partners. To this end, recent efforts have mainly focused on developing whole-genome sequencing approaches and molecular procedures in H. bacteriophora to genetically manipulate the vector nematode in a similar way to the C. elegans model. Earlier work identified species-specific satellite DNA motifs in H. bacteriophora and Steinernema glaseri nematodes, and employed DNA reassociation kinetics to determine the genome size and complexity in Heterorhabditis bacteriophora and Steinernema carpocapsae (Grenier et al. 1996, 1997). At the same time, the expressed sequence tags were generated in H. bacteriophora to elucidate their involvement in various biological functions and more recently RNA-sequencing studies were carried out to understand the molecular basis of nematode parasitism (Sandhu et al. 2006; Bai et al. 2007, 2009; Vadnal et al. 2017). A tremendous breakthrough in entomopathogenic nematode research, including the nematode–bacterial mutualistic symbiosis, came about with the complete sequencing of the H. bacteriophora genome, the annotation which was recently improved (Bai et al. 2013; Vadnal et al. 2018; McLean et al. 2018). In terms of genetic techniques, the development of gene silencing RNAi interference through soaking and microinjection in H. bacteriophora has substantially upgraded the research value of this model organism (Ciche and Sternberg 2007; Ratnappan et al. 2016). Such advances are particularly significant because not only they promise to uncover previously unknown players of the interrelationship between the nematodes and their associated bacteria, but also to reveal the exact contribution of the parasites to the infection process of insects.
6 Concluding Remarks
Entomopathogenic nematodes are spectacular organisms that have received particular attention due to their complex life cycle that involves the mutualistic interdependence with specific bacteria that act as symbionts for the parasites and potent pathogens for the invaded insects. This extremely efficient relationship provides a fascinating model system for studying the interactions between invertebrates and their mutualistic microbes in relation to the host immune system (Ffrench-Constant et al. 2003; Joyce et al. 2006; Clarke 2008). Appreciating the details of the participation of the molecular processes and the specific genes or their products in fine-tuning the relationship between entomopathogenic nematodes and their associated bacteria is essential for understanding mutualistic interactions in other invertebrate organisms including beneficial insects, and devastating vectors of infectious diseases. It is also important for deciphering the conserved mechanisms that regulate similar types of interconnection between microbes and vertebrate animals, including humans. Such knowledge will significantly advance our biological interpretation of a wide range of host–microbial interplays that occur not only in the lab but also in different environmental settings.
References
An R, Grewal PS (2010) Molecular mechanisms of persistence of mutualistic bacteria Photorhabdus in the entomopathogenic nematode host. PLoS One 5:e13154
Bai X, Grewal PS, Hogenhout SA et al (2007) Expressed sequence tag analysis of gene representation in insect parasitic nematode Heterorhabditis bacteriophora. J Parasitol 93:1343–1349
Bai X, Adams BJ, Ciche TA et al (2009) Transcriptomic analysis of the entomopathogenic nematode Heterorhabditis bacteriophora TTO1. BMC Genomics 10:205
Bai X, Adams BJ, Ciche TA et al (2013) A lover and a fighter: the genome sequence of an entomopathogenic nematode Heterorhabditis bacteriophora. PLoS One 8:e69618
Bintrim SB, Ensign JC (1998) Insertional inactivation of genes encoding the crystalline inclusion proteins of Photorhabdus luminescens results in mutants with pleiotropic phenotypes. J Bacteriol 180:1261–1269
Bode HB (2009) Entomopathogenic bacteria as a source of secondary metabolites. Curr Opin Chem Biol 13:224–230
Chaston J, Goodrich-Blair H (2010) Common trends in mutualism revealed by model associations between invertebrates and bacteria. FEMS Microbiol Rev 34:41–58
Chaston JM, Suen G, Tucker SL et al (2011) The entomopathogenic bacterial endosymbionts Xenorhabdus and Photorhabdus: convergent lifestyles from divergent genomes. PLoS One 6:e27909
Ciche T (2007) The biology and genome of Heterorhabditis bacteriophora. WormBook:1–9. https://doi.org/10.1895/wormbook.1.135.1
Ciche TA, Ensign JC (2003) For the insect pathogen Photorhabdus luminescens, which end of a nematode is out? Appl Environ Microbiol 69:1890–1897
Ciche TA, Sternberg PW (2007) Postembryonic RNAi in Heterorhabditis bacteriophora: a nematode insect parasite and host for insect pathogenic symbionts. BMC Dev Biol 7:101
Ciche TA, Bintrim SB, Horswill AR et al (2001) A Phosphopantetheinyl transferase homolog is essential for Photorhabdus luminescens to support growth and reproduction of the entomopathogenic nematode Heterorhabditis bacteriophora. J Bacteriol 183:3117–3126
Ciche TA, Blackburn M, Carney JR et al (2003) Photobactin: a catechol siderophore produced by Photorhabdus luminescens, an entomopathogen mutually associated with Heterorhabditis bacteriophora NC1 nematodes. Appl Environ Microbiol 69:4706–4713
Ciche TA, Darby C, Ehlers RU et al (2006) Dangerous liaisons: the symbiosis of entomopathogenic nematodes and bacteria. Biol Control 38:22–46
Clarke DJ (2008) Photorhabdus: a model for the analysis of pathogenicity and mutualism. Cell Microbiol 10:2159–2167
Cowles KN, Cowles CE, Richards GR et al (2007) The global regulator Lrp contributes to mutualism, pathogenesis and phenotypic variation in the bacterium Xenorhabdus nematophila. Cell Microbiol 9:1311–1323
Dale C, Moran NA (2006) Molecular interactions between bacterial symbionts and their hosts. Cell 126:453–456
Easom CA, Joyce SA, Clarke DJ (2010) Identification of genes involved in the mutualistic colonization of the nematode Heterorhabditis bacteriophora by the bacterium Photorhabdus luminescens. BMC Microbiol 10:45
Eleftherianos IG (2009) Novel antibiotic compounds produced by the insect pathogenic bacterium Photorhabdus. Recent Pat Antiinfect Drug Discov 4:81–89
Ffrench-Constant R, Waterfield N, Daborn P et al (2003) Photorhabdus: towards a functional genomic analysis of a symbiont and a pathogen. FEMS Microbiol Rev 26:433–456
Ffrench-Constant RH, Dowling A, Waterfield NR (2007) Insecticidal toxins from Photorhabdus bacteria and their potential use in agriculture. Toxicon 49:436–451
Forst S, Dowds B, Boemare N et al (1997) Xenorhabdus and Photorhabdus spp.: bugs that kill bugs. Annu Rev Microbiol 51:47–72
Goodrich-Blair H, Clarke DJ (2007) Mutualism and pathogenesis in Xenorhabdus and Photorhabdus: two roads to the same destination. Mol Microbiol 64:260–268
Grenier E, Laumond C, Abad P (1996) Molecular characterization of two species-specific tandemly repeated DNAs from entomopathogenic nematodes Steinernema and Heterorhabditis (Nematoda: Rhabditida). Mol Biochem Parasitol 83:47–56
Grenier E, Catzeflis FM, Abad P (1997) Genome sizes of the entomopathogenic nematodes Steinernema carpocapsae and Heterorhabditis bacteriophora (Nematoda: Rhabditida). Parasitology 114:497–501
Hampton HG, Watson BNJ, Fineran PC (2020) The arms race between bacteria and their phage foes. Nature 577:327–336
Hapeshi A, Benarroch JM, Clarke DJ, Waterfield NR (2019) Iso-propyl stilbene: a life cycle signal? Microbiology 165:516–526
Hentschel U, Steinert M, Hacker J (2000) Common molecular mechanisms of symbiosis and pathogenesis. Trends Microbiol 8:226–231
Herbert EE, Goodrich-Blair H (2007a) Friend and foe: the two faces of Xenorhabdus nematophila. Nat Rev Microbiol 5:634–646
Herbert EE, Goodrich-Blair H (2007b) CpxRA regulates mutualism and pathogenesis in Xenorhabdus nematophila. Appl Environ Microbiol 73:7826–7836
Hu K, Webster JM (2000) Antibiotic production in relation to bacterial growth and nematode development in Photorhabdus-Heterorhabditis infected Galleria mellonella larvae. FEMS Microbiol Lett 189:219–223
Joyce SA, Clarke DJ (2003) A hexA homologue rom Photorhabdus regulates pathogenicity, symbiosis and phenotypic variation. Mol Microbiol 47:1445–1457
Joyce SA, Watson RJ, Clarke DJ (2006) The regulation of pathogenicity and mutualism in Photorhabdus. Curr Opin Microbiol 9:127–132
Joyce SA, Brachmann AO, Glazer I et al (2008) Bacterial biosynthesis of a multipotent stilbene. Angew Chem Int Ed Engl 47:1942–1945
Kaya HK, Gaugler R (1993) Entomopathogenic nematodes. Annu Rev Entomol 38:181–206
Kooliyottil R, Upadhyay D, Inman F III et al (2013) A comparative analysis of entomoparasitic nematodes Heterorhabditis bacteriophora and Steinernema carpocapse. Open J Anim Sci 3:326–333
Kumar R, Kushwah J, Ganguly S et al (2016) Proteomic investigation of Photorhabdus Bacteria for nematode-host specificity. Indian J Microbiol 56:361–367
Lango-Scholey L, Brachmann AO, Bode HB et al (2013) The expression of stlA in Photorhabdus luminescens is controlled by nutrient limitation. PLoS One 8:e82152
Li Y, Peng N (2019) Endogenous CRISPR-Cas system-based genome-editing and antimicrobials: review and prospects. Front Microbiol 10:2471
Maurelli AT (2007) Black holes, antivirulence genes, and gene inactivation in the evolution of bacterial pathogens. FEMS Microbiol Lett 267:1–8
McLean F, Berger D, Laetsch DR, Schwartz HT, Blaxter M (2018) Improving the annotation of the Heterorhabditis bacteriophora genome. Gigascience 7:1–12
Ochman H, Moran NA (2001) Genes lost and genes found: evolution of bacterial pathogenesis and symbiosis. Science 292:1096–1098
Orchard SS, Goodrich-Blair H (2005) Pyrimidine nucleoside salvage confers an advantage to Xenorhabdus nematophila in its host interactions. Appl Environ Microbiol 71:6254–6259
Parkhill J, Wren BW, Thomson NR et al (2001) Genome sequence of Yersinia pestis, the causative agent of plague. Nature 413:523–527
Qiu X, Yan X, Liu M et al (2012) Genetic and proteomic characterization of rpoB mutations and their effect on nematicidal activity in Photorhabdus luminescens LN2. PLoS One 7:e43114
Ratnappan R, Vadnal J, Keaney M et al (2016) RNAi-mediated gene knockdown by microinjection in the model entomopathogenic nematode Heterorhabditis bacteriophora. Parasit Vectors 9:160
Ruby EG (2008) Symbiotic conversations are revealed under genetic interrogation. Nat Rev Microbiol 6:752–762
Sagan L (1967) On the origin of mitosing cells. J Theor Biol 14:255–274
Sandhu SK, Jagdale GB, Hogenhout SA, Grewal PS (2006) Comparative analysis of the expressed genome of the infective juvenile entomopathogenic nematode, Heterorhabditis bacteriophora. Mol Biochem Parasitol 145:239–244
Shi YM, Brachmann AO, Westphalen MA et al (2019) Dual phenazine gene clusters enable diversification during biosynthesis. Nat Chem Biol 15:331–339
Singh S, Orr D, Divinagracia E et al (2015) Role of secondary metabolites in establishment of the mutualistic partnership between Xenorhabdus nematophila and the entomopathogenic nematode Steinernema carpocapsae. Appl Environ Microbiol 81:754–764
Somvanshi VS, Kaufmann-Daszczuk B, Kim KS et al (2010) Photorhabdus phase variants express a novel fimbrial locus, mad, essential for symbiosis. Mol Microbiol 77:1021–1038
Somvanshi VS, Sloup RE, Crawford JM et al (2012) A single promoter inversion switches Photorhabdus between pathogenic and mutualistic states. Science 337:88–93
Vadnal J, Ratnappan R, Keaney M et al (2017) Identification of candidate infection genes from the model entomopathogenic nematode Heterorhabditis bacteriophora. BMC Genomics 18:8
Vadnal J, Granger OG, Ratnappan R et al (2018) Refined ab initio gene predictions of Heterorhabditis bacteriophora using RNA-seq. Int J Parasitol 48:585–590
Veesenmeyer JL, Andersen AW, Lu X et al (2014) NilD CRISPR RNA contributes to Xenorhabdus nematophila colonization of symbiotic host nematodes. Mol Microbiol 93:1026–1042
Volgyi A, Fodor A, Forst S (2000) Inactivation of a novel gene produces a phenotypic variant cell and affects the symbiotic behavior of Xenorhabdus nematophilus. Appl Environ Microbiol 66:1622–1628
Waterfield NR, Ciche T, Clarke D (2009) Photorhabdus and a host of hosts. Annu Rev Microbiol 63:557–574
Williams JS, Thomas M, Clarke DJ (2005) The gene stlA encodes a phenylalanine ammonia-lyase that is involved in the production of a stilbene antibiotic in Photorhabdus luminescens TT01. Microbiology 151:2543–2550
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Eleftherianos, I., Heryanto, C. (2020). Molecular Regulators of Entomopathogenic Nematode–Bacterial Symbiosis. In: Kloc, M. (eds) Symbiosis: Cellular, Molecular, Medical and Evolutionary Aspects. Results and Problems in Cell Differentiation, vol 69. Springer, Cham. https://doi.org/10.1007/978-3-030-51849-3_17
Download citation
DOI: https://doi.org/10.1007/978-3-030-51849-3_17
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-51848-6
Online ISBN: 978-3-030-51849-3
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)