Abstract
Traditionally, histopathological evaluations of tissue sections are performed for the diagnosis and grading of cancers using subjective appraisal of the tissue and cell phenotypes. In the last decade, a combination of unprecedented advances in imaging and computing technologies and novel machine learning-based algorithms has driven the field of histopathology into a new dimension. Machine learning and deep learning methodologies applied on digitalized hematoxylin and eosin stained tissue sections have out-performed conventional methods to accurately identify cancer cells or classify tumors into prognostic groups. Nevertheless, we believe that a precise, standardized measurement of nuclear morphology and chromatin texture based on a DNA stain can further improve the diagnosis of cancers and identify patients with high-risk of recurrence, alone or in combination with other clinical, pathological or molecular information. Changes in the morphology of cell nuclei and tissue architecture are intrinsic characteristics and hallmarks of cancer. Nuclei from cancerous samples exhibit different morphological and chromatin texture than nuclei from normal cells, thus, reflecting the structural and molecular effects of genetic and epigenetic alterations driving cancer processes. By image analysis, the chromatin texture can be measured in high-resolution breaking down the components of the nuclear changes into multiple quantifiable units that can be studied independently and in combination using advanced machine learning methods. This allows the investigator to examine associations of such changes with cancer progression and clinical outcomes. There is now increasing interest in developing new algorithms and platforms to decipher spatial relationship between cell subpopulations in whole tissue sections.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Digital pathology
- Image analysis
- Feulgen
- Cancer screening
- Cancer diagnosis
- Prognostic biomarker
- Predictive biomarker
- Cell sociology
1 Introduction and Motivation
Traditionally, the histopathological diagnosis and grading of cancers are based on the appraisal of the changes in the morphology of the cells and tissue organization which are the intrinsic hallmarks of cancers. Nuclei from cancerous samples exhibit different shapes and chromatin textures than nuclei from normal samples reflecting the structural and molecular effects of genetic and epigenetic alterations driving cancer processes [1,2,3]. For instance, increased proportion of heterochromatin condensation is a nuclear characteristic for high-risk cancer [4]. In the last two decades, advances in pathology slide scanning technologies and image processing algorithms have enabled the breakdown of the subjective and qualitative nuclear and architectural changes used by the pathologists into objective quantifiable units that can be studied independently and/or in combination using advanced statistical and machine learning methods [5]. In 2011, Beck et al. [6] developed an algorithm to quantify epithelial and stromal changes in H&E slides of more than 500 breast cancer patients and demonstrated that a scoring system based on these measurements was strongly associated with overall survival and was independent of clinical, pathological and molecular factors. After the first FDA approval of a slide scanner device for primary clinical diagnosis [7], the adoption of digital pathology has gained an exponential popularity in the community of pathologists and is expected to unlock a large number of research and development opportunities including digital applications for diagnosis, prognosis and prediction of treatment response in cancer diseases [8, 9]. While most of the digital pathology algorithms have been developed using the traditional H&E or Papanicolaou slides and artificial neural networks to assist pathologists in many tasks, alternative staining techniques and image analysis processing should not be disregarded, in particular if they can capture the changes in nuclear and chromatin features with improved performances [10], strong consistency [11] and have a proven track of diagnostic, prognostic and predictive clinical applications in a variety of cancer types [12,13,14,15]. In fact, the diversity of information (e.g. clinical, imaging and molecular data), high quality of data and integration of multiple machine learning approaches should be further encouraged for the success of artificial intelligence in oncology and other fields of medicine in general [16,17,18]. In this chapter, the authors review the use of Feulgen as the best stoichiometric stain for accurate quantification of DNA and better representation of the nuclear chromatin texture. Examples of successful applications of computer-assisted image analysis and machine learning in both cytology and histopathology specimens using Feulgen staining will be discussed regarding their accuracy and clinical utility.
2 Glossary
DNA Ploidy:
DNA ploidy is a cytogenetic term describing the number of chromosome sets (n) or deviations from the normal number of chromosomes in a cell. In cytometry, the expression is used either to describe the DNA content in a cell or the total DNA distribution in a cell population.
Digital Image Analysis:
Image analysis is the extraction of meaningful information from images using a computer device or electrical device combined to digital image processing techniques. It involves the fields of computer or machine vision, and medical imaging, and makes heavy use of pattern recognition, digital geometry, and signal processing.
Test Performance:
Diagnostic test performance evaluates the ability of a qualitative or quantitative test to discriminate between two subclasses of subjects.
Test Accuracy:
Diagnostic test accuracy measures the ability of a test to detect a condition when it is present and detect the absence of a condition when it is absent.
Screening:
Screening is defined as the presumptive identification of unrecognized disease in an apparently healthy, asymptomatic population by means of tests, examinations or other procedures that can be applied rapidly and easily to the target population.
Diagnostic Biomarker:
A diagnostic biomarker is used to confirm that a patient has a particular health disorder. Diagnostic biomarkers may facilitate earlier detection of a disorder than can be achieved by physical examination of a patient.
Prognostic Biomarker:
A prognostic biomarker is a clinical or biological characteristic that provides information on the likely patient health outcome (e.g. disease recurrence) irrespective of the treatment.
Predictive Biomarker:
A predictive biomarker indicates the likely benefit to the patient from the treatment compared to their condition at baseline.
3 State of the Art
3.1 Feulgen Stain and DNA-Based Image Analysis
The Feulgen technique is generally accepted as a stoichiometric DNA stain that is used to quantify the amount of DNA in cell nuclei in a reproducible and standardized manner [19]. Feulgen reaction allows the precise densitometric measurement of nuclear DNA because the amount of the dye bound per nucleus is proportional to its DNA content. Briefly, the DNA is submitted to mild acid hydrolysis to split off the purine bases from the double-stranded DNA. The result is an apurinic acid presenting aldehyde groups at the C1-position. A Schiff’s base binds to these aldehyde groups and produces a blue-violet color with 545 nm maximum absorption wavelength [20]. The DNA image analysis is performed using a digital camera that captures images of Feulgen-stained individual nuclei in the specimen. The images are divided into image elements (picture elements - pixels). The gray tone value for each pixel represents the intensity of DNA specific staining. The value is saved in the computer which numerates between 0 (black) and 1023 (white). In-house image analysis software is used to measure the relative amount of DNA in each nucleus (DNA ploidy) by summing the optical density of all the pixels in the nucleus (Fig. 1).
In addition, due to the optimal object-to-background contrast of Feulgen stain, the software can measure the morphometric features including the size, shapes and border smoothness of nuclei. Changes in the nuclear chromatin appearance are common in dysplastic and cancer cell. Features describing the chromatin distribution pattern are referred to as chromatin texture features. Using Feulgen stain, they can be assessed by mathematical formulas that describe the distribution of gray levels in groups of pixels [21]. For instance, Markovian texture features characterize gray-level correlation between adjacent pixels in the image. Non-Markovian texture features describe the local maxima and minima of gray-level differences in the object. Fractal texture features compare local differences integrated over the object at multiple resolutions. Run-length texture features measure the length of consecutive pixels with the same compressed gray-level value along different orientations (0°, 45°, 90°, 135°). Several studies investigated the suitability of Papanicolaou and hematoxylin staining, which are routinely used for daily cytopathology and histopathology, for DNA-based image analysis or ploidy measurement. Unfortunately, the coefficient of variation (CV) is broader in Papanicolaou and hematoxylin stains resulting in significant disproportionality and less reproducibility of the optical density and ploidy values [22, 23]. These studies confirmed that Feulgen remains the gold standard stain for the precise densitometric measurement of DNA content in nuclei [10] (Fig. 2).
The tissue sections are analyzed using in-house developed image analysis software (Getafics; BCCA, Vancouver, Canada). This software was specifically designed for semi-automatic analysis of DNA content, nuclear morphology, chromatin texture and tissue architecture. Briefly, after the Feulgen-stained tissue sections ae digitalized by a whole slide scanning system, the operator selects the region of interest by delineating the boundaries. A threshold algorithm is applied to the image, followed by a segmentation algorithm to separate touching and overlapping nuclei. Autofocusing and edge relocation algorithms are applied to the nuclei to locate the edge of the objects precisely and automatically segment the contour of the highest local gray-level gradient. The digital gray-level images of individual segmented nuclei are stored in a gallery and analyzed the nuclear or architecture features are extracted using computer calculations. The calculated values of these features are used as datasets that will be tested by multiple machine learning and classifier algorithms (Figs. 3 and 4).
3.2 Cancer Screening
Population-based cervical cancer screening programs have been effective in reducing the incidence and mortality of cervical cancers [24]. Pap tests (liquid-based cytology or conventional smears) are widely adopted in developed countries as the gold standard screening method where cells are collected from the cervix to generate Papanicolaou-stained cytology slides for examination under the microscope [25]. However, there are many countries in the world where large scale screening programs have not yet been implemented due to several challenges including the shortage of cytopathologists and lack of skilled cytotechnologists who need to review high volumes of Papanicolaou-stained cytology slides. In addition, many cervical screening programs in resource-limited countries have a high false negative rate [26, 27]. Over the last decades, several automated imaging technologies were developed and clinically implemented to assist the cytotechnologists in reviewing the Pap slides review (i.e. BD FocalPoint Imaging System, ThinPrep Imaging System). Overall, the performance of the automated system has been well accepted and the list of benefits includes improved sensitivity in detection of squamous intraepithelial lesions and increased productivity compared to manual review of conventional Pap smears [28]. However, the adoption of the FDA-approved systems comes with increased costs (equipment and maintenance) and they may not be suitable for low-volume cytology laboratories. The British Columbia Cancer Agency group has developed a series of inexpensive fully automated systems combining point-of-care slide scanners with image analysis software that measures the ploidy of the cells to detect aneuploid cancerous cells based. Briefly, barcoded Feulgen-stained slides are placed in a slide loader and the operator initiates scanning on a supervisor computer. Although such machines can scan smears, liquid base cytology (LBC) slides are generally preferred to generate monolayers of cells simplifying the task of automated imaging. Autofocusing, image capture, segmentation of the nuclei, morphometric and ploidy measurements are all performed automatically without operator intervention. Reporting is done at the conclusion of an interactive review of the scan data for each slide which are comprised of stored images of the cell nuclei, counts of various cell types, and histograms and scatter plots of cell DNA index (calculated by the normalization of a cell DNA content to a population of normal diploid cells) and other morphometric features of the cell nuclei (i.e. nuclear size, smoothness of nuclear boundaries, etc.). The reviewer follows a very simple checklist procedure to systematically examine the data, looking: (a) first to check that the DNA scale (normalization) is valid, then; (b) checking the presence of aneuploid cell nuclei (DI > 2.5), then; (c) looking for aneuploid “stemlines”, then; (d) assessing cell proliferation rate (proliferative cells contains between 2 to 4 copies of DNA); and (e) if none of these are present, then the case is negative.
Head-to-head comparison of automated ploidy-based cytometry versus conventional Pap cytology was performed in multiple studies. In a cohort of 1,555 patients seen in MD Anderson Cancer Center [29], the test performance of DNA-based ploidy (59% sensitivity, 93% specificity, 92% NPV, 63% PPV) was found equivalent to the local cytology laboratory (47% sensitivity, 96% specificity, 90% NPV, 70% PPV). In China, the studies found substantially increased sensitivity (up to the double). For instance, in a cohort of 9,950 screened women [30], the test performance of DNA-based ploidy was 54% sensitivity, 97% specificity, 92% NPV and 58% PPV while conventional Pap cytology performed in local hospital showed 25% sensitivity, 99% specificity, 85% NPV and 54% PPV. Since 2005, the use of DNA-based cytometers in China has continuously expanded to over 1 million tests per year. There are over 70 publications comparing the performance of DNA-based ploidy to conventional Pap smears; however, most of the studies published studies are observational and suffer from missing data or lack of follow-up cervical biopsies as the gold standard method to confirm the presence of high-grade dysplasia or malignancy when one or two tests are positive [31,32,33]. Overall, three conclusions can be drawn from these ground studies. First, the ploidy-based cytometry is a simple and reproducible technique. Second, it can be taught much more quickly than cytology. Records in China have demonstrated that it is routinely possible to teach the technology from slide preparation and staining, to operation of the cytometer, review of the DNA ploidy data and report generation in 10 working days. Third, the test performance of ploidy-based cytometry is comparable to conventional or liquid-based cytology performed by experienced and highly trained cytologists. The diffusion of the automated quantitative image cytometry is expected to expand as the various vendors continue to receive regulatory approvals [34, 35] and endorsement by medical societies and expert groups (Fig. 5).
3.3 Early Cancer Diagnostics
Although the importance of early diagnosis in improving the mortality and morbidity of cancer has long been recognized, the disease is still frequently diagnosed late and prognosis has not dramatically changed for the last decades. A major challenge for early diagnosis of epithelial cancers is our ability to recognize precursors or premalignant lesions at risk of progressing into invasive carcinomas. The progression appears to occur through a low-grade dysplasia (low risk of progression) to high-grade dysplasia (high risk of progression) to carcinoma sequence. One of the most significant challenges confronting the diagnosis of premalignant lesions is the poor agreement among pathologists in the histopathological diagnosis and grading of dysplasia. In Barrett’s esophagus for instance, significant intra- and interobserver variability in the interpretation of biopsy specimens has been well documented, even between expert pathologists, especially at the lower end of the dysplasia spectrum (i.e. benign reactive cytologic atypia versus low-grade dysplasia) [36, 37]. In the context of digital pathology, there is an interest in developing tissue imaging biomarkers both to predict which patients may develop carcinoma (and therefore be offered surgical therapy with curative intent) and to aid guiding surveillance intervals following therapy. Using different image analysis and statistical methods, the potential of image analysis to measure the grade of dysplastic lesions has been demonstrated in different tissues types, such as skin, ovary or prostate [38,39,40]. For instance, our group has previously shown that measuring the chromatin texture features alone can detect serial nuclear changes in the sequential progression of Barret’s esophagus from normal epithelium to dysplastic epithelium to invasive carcinoma, and objectively distinguish reactive epithelial changes (indefinite for dysplasia) from low-grade dysplasia [12]. In addition, our results suggest that quantitative measurement of chromatin texture features has a better correlation with the class of dysplasia (low- versus high-grade). As opposed to morphologic features measuring changes in nuclear sizes and shapes, chromatin texture features are less sensitive to sectioning variation and could have a superior contribution in the differential diagnosis of Barrett’s esophagus classification. We previously observed similar findings in multiple human epithelium sites, including the oral cavity [41], lung [42], cervix [43] and breast [44]. We believe that measurement of nuclear chromatin texture is significant because such changes are an indication of genetic or epigenetic changes that lead toward malignant transformation.
Another known problem in early cancer diagnosis is the difficulty to diagnose the well differentiated intraepithelial neoplasia based on traditional morphology. For instance, differentiated vulvar intraepithelial neoplasia (DVIN) possesses a high oncogenic potential but the high degree of differentiation often results in DVINs being mistakenly diagnosed as benign lesions (i.e. lichen simplex chronicus, lichen sclerosus). The p53 immunohistochemistry marker can be used to support the DVIN diagnosis; however, the characteristic suprabasal p53 overexpression can be encountered in any benign condition in which there is increased epithelial proliferation. The lack of p53 specificity encourages the development of alternative aid tools for DVIN diagnosis. We recently investigated the role of chromatin-based image analysis in distinguishing DVIN versus benign mimickers. Sixty-five vulva biopsy specimens with three major diagnosis categories were selected: Lichen simplex chronicus (n = 34); (2) Lichen sclerosis (n = 21); DVIN (n = 20). A total of 44,483 nuclei were individually captured from the squamous epithelium of the 65 study cases and analyzed for over 100 parameters that assess the shapes, DNA content, chromatin texture and overall architecture of the nuclei. The averages of individual parameters in each specimen are included in a stepwise discriminant analysis. Limiting the classifier model to only 2 nuclear texture features, we achieved an overall accuracy of 95.5% in distinguishing DVIN versus lichen simplex chronicus (sensitivity of 80% with 95% CI: 45% to 98%; specificity of 98% with 95% CI: 90% to 99.9%), and overall accuracy of 96.8% in distinguishing DVIN versus lichen sclerosus (sensitivity of 100% with 95% CI: 69% to 100%; specificity of 95% with 95% CI: 76% to 100%). Limiting the classifier to 2-parameters would increase the chances for reproducibility in independent cohorts. Additional test sets are needed to validate or improve the classifier performances (Figs. 6, 7 and 8).
3.4 Cancer Prognostics
Breast cancer is the leading cause of cancer-related deaths among women worldwide. Adjuvant systemic therapy, including hormonal and chemotherapy, has reduced mortality from breast cancer. As chemotherapy is toxic and has a negative impact on quality of life, it should ideally be given only to those patients who gain significant benefit from it. At present, many patients are over-treated [45]. Apart from traditional prognostic markers that include TNM stage, Estrogen receptor (ER), Progesterone Receptor (PR), human epidermal growth factor receptor (HER2) and pathological features (grade), new genomic profiling tests are being developed to aid refinement in treatment recommendations. Recent recommendations from ASCO support the use of several biomarker assays, including OncotypeDX Recurrence Score (RS), EndoPredict, PAM50 and Breast Cancer Index [46]. The most commonly used assay is OncotypeDX, a multigene reverse transcriptase (RT)-PCR assay designed to quantify the 10-year risk of metastatic recurrence. The major obstacles of this assay are the high cost (about $4,000 per test) and the necessity to ship specimens to California for centralized testing which delays patient care. Our group investigated the contribution of quantitative image analysis in the discrimination between survivors and deceased patients with more than 10 years follow-up after surgery [14]. Feulgen-stained tissue sections of 80 breast carcinomas were processed by our in-house image analysis software. A random forest algorithm selected the best five nuclear texture features and generated a survival score. This classifier model could discriminate between survivor and deceased breast cancer patient with a sensitivity of 88% and a specificity of 85%. Using a multivariate Cox proportional hazards analysis, we assessed the added prognostic value of survival score with other clinical and pathological factors, such as age, lymph node status, tumor size and grade. The survival score was significantly associated with 10-year survival, independent of any tumor grade (1, 2 or 3) or other clinical factors (p = 0.005). In earlier studies, our group pioneered the use of imaging analysis to detect changes in early precancerous breast tumors (DCIS) and demonstrated continuous morphometric changes from hyperplasia to invasive cancers [47, 48] (Figs. 9 and 10).
3.5 Cancer Theranostics
Prostate cancer is the most commonly diagnosed form of cancer in men worldwide. PSA screening has led to a steep increase in incidence of indolent PSA-detected cases, which ultimately do not contribute significantly to the overall mortality rates. Although some prostate cancers behave aggressively and will result in death, most of PSA detected cancers are non-aggressive, slow growing, and do not require immediate intervention. Active surveillance is a preferred approach for PSA detected early prostate cancer. Significantly, 5–10% of individuals with low-risk disease treated up-front with prostate brachytherapy or radical prostatectomy will experience poor outcomes. Additionally, >50% of active surveillance patients will progress, and will require treatment 5 years after initial diagnosis [49]. The effectiveness of active surveillance is limited without a tool to accurately provide prognostic information. Current treatment recommendations are based on PSA levels (iPSA), clinical staging, and Gleason score. Molecular assays are being proposed to predict the risk of clinical metastasis within 5 years of radical prostatectomy surgery; however, these molecular assays have financial and logistical limitations similar to those described in breast. We investigated whether image analysis of nuclear features and tissue architecture can distinguish patients with biochemical failure from biochemical non-evidence of disease (BNED) after radical prostatectomy (RP) for prostate cancer [15]. Of the 78 prostate cancer tissue cores collected from patients treated with RP, 16 who developed biochemical relapse (failure group) and 16 who were BNED patients (non-failure group) were included in the analyses (36 cores from 32 patients). A section from this TMA was stained stoichiometrically for DNA using the Feulgen methodology and stained slides were scanned. Prostate TMA core classification as biochemical failure or BNED after RP was conducted (a) based on cell type and cell position within the epithelium (all cells, all epithelial cells, epithelial >2 cell layers away from basement membrane) from all cores, and (b) based on epithelial cells more than two cell layers from the basement membrane using a Classifier trained on Gleason 6, 8, 9 (16 cores) only and applied to a Test set consisting of the Gleason 7 cores (20 cores). Successful core classification as biochemical failure or BNED after RP by a linear classifier was 75% using all cells, 83% using all epithelial cells, and 86% using epithelial >2 layers. Overall success of predicted classification by the linear Classifier of (b) was 87.5% using the Training Set and 80% using the Test Set. The success of predicted progression using traditional morphologic Gleason score alone was 75% for Gleason >7 as failures and 69% for Gleason >6 as failures. Combination of Tissue Architecture score and Gleason score yielded an overall accuracy of 89% suggesting that the combination of image analysis and conventional morphologic assessment can have a synergistic impact (Fig. 11).
4 Open Problems
So far, most of the image analysis algorithms applied to digital pathology are focused on the characterization of the tissue phenotype; however, cell proliferation, immune evasion, hypoxia and tumor heterogeneity are also important hallmarks of cancer. A better understanding of their individual role and of their mutual interactions is needed to unlock all the valuable information in the glass slide. Advances in optical imaging and immunohistochemistry technologies will allow us to decipher with unprecedented details, precision, and depth the individual expression and intensity of multiple markers as well as the spatial interaction of cell subpopulations (“cell sociology”) in large histological images.
5 Future Outlook
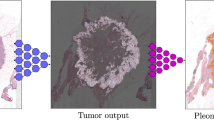
The British Columbia Cancer Agency group developed several hyperspectral microscopy systems to rapidly collect 7–16 wavelength specific images (from 400 to 780 nm) across entire slides. These hyperspectral images can be used to spectrally un-mix the components within a slide that have unique features [50]. As an example, using the absorption spectra of each immunohistochemistry antibody, the stains are computationally unmixed to determine the concentration of each stain for every pixel in the selected area. Our program basically makes the assumption that every pixel in the recorded images (16 wavelengths) are a linear combination of the concentration of the individual stains occurring at that pixel weighted by the absorption characteristics of each of the stains occurring at that pixel. The method we used to separate these linear combinations of absorption stains with different concentration at each pixel was the Multivariate Curve Resolution – Alternating Least Squares algorithm. Different immunohistochemistry stains are available to assess each component of the tumor microenvironment. High expression of Ki-67 protein is known to be associated with aggressive cancers. Ki-67 is a robust marker due to the reproductive and strong signal of his antibody (Mib1). The tumor microenvironment is a complex mixture of tumor epithelium, stroma and immune cells, and the immune component of the tumor microenvironment is highly prognostic for tumour progression and patient outcome. The role of the immune evasion pathways (PD-1/PD-L1-CD8) and T lymphocytes (CD3-CD8) can be studied by immunohistochemistry markers which are currently being used to guide immunotherapy. Cell sociology approach may be critical to examine immune cell function, as anti-tumor immune activation depends on a complex network of interactions between antigen presenting cells, T cells, and target cells. Importantly, quantification via cell sociology has the potential to provide greater prognostic or predictive insight than cell density readings have historically provided. Recently, we showed that the characterization of the spatial tumor-immune cell interactions is associated with lung cancer recurrence [50]. The presence of poorly oxygenated (hypoxic) cells is associated with poor outcome after radiation, chemotherapy, and surgery in a wide range of solid tumors. Hypoxia can be measured by quantifying endogenous expression of hypoxia-induced proteins by immunohistochemistry (e.g., carbonic anhydrase-9; CA9 or glucose transporter-1; Glut-1).
Overall, we believe the application of hyperspectral imaging combined with cell sociology studies will significantly increase our understanding of cancer biological behavior and facilitate the development of robust imaging biomarkers to improve risk stratification and complement clinical prognostic factors. Moreover, hyperspectral imaging platform can also improve the ability of some nuclear morphometric and chromatin texture features to differentiate between cell groups. We discovered that diffraction effects in microscopy images can be readily separated from Feulgen stained material (sharp absorption max at 600 nm). Removal of these diffraction effects simplifies all downstream image analysis including segmentation and differentiation of overlapping cells (Fig. 12).
References
Zink, D., Fischer, A.H., Nickerson, J.A.: Nuclear structure in cancer cells. Nat. Rev. Cancer 4(9), 677–687 (2004)
Singh, H., Sekinger, E.A., Gross, D.S.: Chromatin and cancer: causes and consequences. J. Cell. Biochem. 79(Suppl. 35), 61–68 (2000)
Almassalha, L.M., Tiwari, A., Ruhoff, P.T., et al.: The global relationship between chromatin physical topology, fractal structure, and gene expression. Sci. Rep. 7, 41061 (2017)
Hveem, T.S., Njolstad, T.S., Nielsen, B., et al.: Changes in chromatin structure in curettage specimens identifies high-risk patients in endometrial cancer. Cancer Epidemiol. Biomarkers Prev. 26(1), 61–67 (2017)
O’Connor, J.P., Aboagye, E.O., Adams, J.E., et al.: Imaging biomarker roadmap for cancer studies. Nat. Rev. Clin. Oncol. 14(3), 169–186 (2016)
Beck, A.H., Sangoi, A.R., Leung, S., et al.: Systematic analysis of breast cancer morphology uncovers stromal features associated with survival. Sci. Transl. Med. 3(108), 108ra113 (2011)
Mukhopadhyay, S., Feldman, M.D., Abels, E., et al.: Whole slide imaging versus microscopy for primary diagnosis in surgical pathology: a multicenter blinded randomized noninferiority study of 1992 cases (pivotal study). Am. J. Surg. Pathol. 42(1), 39–52 (2018)
Tizhoosh, H.R., Pantanowitz, L.: Artificial intelligence and digital pathology: challenges and opportunities. J. Pathol. Inform. 9, 38 (2018)
Bychkov, D., Linder, N., Turkki, R., et al.: Deep learning based tissue analysis predicts outcome in colorectal cancer. Sci. Rep. 8(1), 3395 (2018)
Biesterfeld, S., Beckers, S., Del Carmen Villa Cadenas, M., et al.: Feulgen staining remains the gold standard for precise DNA image cytometry. Anticancer Res. 31(1), 53–58 (2011)
Ghizoni, J.S., Sperandio, M., Lock, C., et al.: Image cytometry DNA ploidy analysis: correlation between two semi-automated methods. Oral Dis. 24(7), 1204–1208 (2018)
Guillaud, M., Zhang, L., Poh, C., et al.: Potential use of quantitative tissue phenotype to predict malignant risk for oral premalignant lesions. Cancer Res. 68(9), 3099–3107 (2008). PMID: 18451134
El Hallani, S., Guillaud, M., Korbelik, J., et al.: Evaluation of quantitative digital pathology in the assessment of barrett esophagus-associated dysplasia. Am. J. Clin. Pathol. 144(1), 151–164 (2015)
Guillaud, M., Ye, Q., Leung, S., Carraro, A., et al.: Large-scale DNA organization is a prognostic marker of breast cancer survival. Med. Oncol. 35(1), 9 (2017)
MacAulay, C., Keyes, M., Hayes, M., et al.: Quantification of large scale DNA organization for predicting prostate cancer recurrence. Cytometry A 91(12), 1164–1174 (2017)
Holzinger, A., Haibe-Kains, B., Jurisica, I.: Why imaging data alone is not enough: AI-based integration of imaging, omics, and clinical data. Eur. J. Nucl. Med. Mol. Imaging 46(13), 2722–2730 (2019)
Jean-Quartier, C., Jeanquartier, F., Jurisica, I., Holzinger, A.: In silico cancer research towards 3R. BMC Cancer 18(1), 408 (2018)
O’Sullivan, S., Holzinger, A., Zatloukal, K., et al.: Machine learning enhanced virtual autopsy. Autops Case Rep. 7(4), 3–7 (2017)
Chatelain, R., Willms, A., Biesterfeld, S., et al.: Automated Feulgen staining with a temperature controlled machine. Anal. Quant. Cytol. Histol. 11, 211–217 (1989)
Haroske, G., Baak, J.P., Danielsen, H., et al.: Fourth updated ESACP consensus report on diagnostic DNA image cytometry. Anal. Cell. Pathol. 23(2), 89–95 (2001)
Doudkine, A., Macaulay, C., Poulin, N., et al.: Nuclear texture measurements in image cytometry. Pathologica 87(3), 286–299 (1995)
Bahr, G.F., Bartels, P.H., Bibbo, M., et al.: Evaluation of the Papanicolaou stain for computer-assisted cellular pattern recognition. Acta Cytol. 17, 106–112 (1973)
Schulte, E.K.W., Fink, D.K.: Hematoxylin staining in quantitative DNA cytometry: an image analysis study. Anal. Cell. Pathol. 9, 257–268 (1995)
Liu, S., Semenciw, R., Probert, A., et al.: Cervical cancer in Canada: changing patterns in incidence and mortality. Int. J. Gynecol. Cancer 11, 24–31 (2001)
Guidozzi, F.: Screening for cervical cancer. Obstet. Gynecol. Surv. 51, 247–252 (1996)
McCord, M.L., Stovall, T.G., Meric, J.L., et al.: Cervical cytology: a randomized comparison of four sampling methods. Am. J. Obstet. Cynecol. 166, 1772–1779 (1992)
Stenkvist, B., Soderstrom, J.: Reasons for cervical cancer despite extensive screening. J. Med. Screen. 3, 204–207 (1996)
Wilbur, D.C., Cibas, E.S., Merritt, S., et al.: ThinPrep processor: clinic trials demonstrate an increased detection rate of abnormal cervical cytologic specimens. Am. J. Clin. Pathol. 101, 209–214 (1994)
Guillaud, M., Benedet, J.L., Cantor, S.B., et al.: DNA ploidy compared with human papilloma virus testing (Hybrid Capture II) and conventional cervical cytology as a primary screening test for cervical high-grade lesions and cancer in 1555 patients with biopsy confirmation. Cancer 107(2), 309–318 (2006)
Sun, X.R., Wang, J., Garner, D., et al.: Detection of cervical cancer and high grade neoplastic lesions by a combination of liquid-based sampling preparation and DNA measurements using automated image cytometry. Cell Oncol. 27(1), 33–41 (2005)
Xu, S., Mei, J.H., Han, Y.L., et al.: Quantitative analysis of cervical cell DNA and liquid-based cytology in cervical screening of early lesions comparison. Guangdong Yixue 34, 1387–1390 (2013)
Tian, Y.W., Liu, G., Zhou, J., et al.: Application of DNA image cytometry in the cervical intraepithelial neoplasia and cervical cancer screening. Zhenduan Binglixue Zazhi 20, 425–428 (2013)
Wei, L.F., Lin, H.M., Qin, Y.A.F.: Clinical research of cervical cell DNA quantitative analysis in opportunistic screening of cervical diseases among women in cities of Zhuang national minority regions. Huaxia Yixue 25, 181–184 (2012)
Motic Regulatory. http://www.med.motic.com/Page.aspx?PageCategoryId=7&PageContentId=4&LangCode=zhcn
Landing Med Regulatory. http://landingmed.com/cn/
Reid, B.J., Haggitt, R.C., Rubin, C.E., et al.: Observer variation in the diagnosis of dysplasia in Barrett’s esophagus. Hum. Pathol. 19, 166–178 (1988)
Montgomery, E., Bronner, M.P., Goldblum, J.R., et al.: Reproducibility of the diagnosis of dysplasia in Barrett esophagus: a reaffirmation. Hum. Pathol. 32, 368–378 (2001)
Glazer, E.S., Bartels, P.H., Prasad, A.R., et al.: Nuclear morphometry identifies a distinct aggressive cellular phenotype in cutaneous squamous cell carcinoma. Cancer Prev. Res. (Phila) 4, 1770–1777 (2011)
Nielsen, B., Albregtsen, F., Kildal, W., et al.: The prognostic value of adaptive nuclear texture features from patient gray level entropy matrices in early stage ovarian cancer. Anal. Cell. Pathol. (Amst) 35, 305–314 (2012)
Veltri, R.W., Khan, M.A., Miller, M.C., et al.: Ability to predict metastasis based on pathology findings and alterations in nuclear structure of normal-appearing and cancer peripheral zone epithelium in the prostate. Clin. Cancer Res. 10, 3465–3473 (2004)
Guillaud, M., Zhang, L., Poh, C., et al.: Potential use of quantitative tissue phenotype to predict malignant risk for oral premalignant lesions. Cancer Res. 68, 3099–3107 (2008)
Guillaud, M., le Riche, J.C., Dawe, C., et al.: Nuclear morphometry as a biomarker for bronchial intraepithelial neoplasia: correlation with genetic damage and cancer development. Cytometry A 63, 34–40 (2005)
Swartz, R., West, L., Boiko, I., et al.: Use of nuclear morphometry characteristics to distinguish between normal and abnormal cervical glandular histologies. Anal. Cell. Pathol. 25, 193–200 (2003)
Poulin, N., Frost, A., Carraro, A., et al.: Risk biomarker assessment for breast cancer progression: replication precision of nuclear morphometry. Anal. Cell. Pathol. 25, 129–138 (2003)
Paik, S., Tang, G., Shak, S., et al.: Gene expression and benefit of chemotherapy in women with node-negative, estrogen receptor-positive breast cancer. J. Clin. Oncol. 24(23), 3726–3734 (2006)
Harris, L.N., Ismaila, N., McShane, L.M., et al.: Use of biomarkers to guide decisions on adjuvant systemic therapy for women with early-stage invasive breast cancer: american society of clinical oncology clinical practice guideline. J. Clin. Oncol. 34(10), 1134–1150 (2016)
Poulin, N., Susnik, B., Guillaud, M., et al.: Histometric texture analysis of DNA in thin sections from breast biopsies. Application to the detection of malignancy-associated changes in carcinoma in situ. Anal. Quant. Cytol. Histol. 17(5), 291–299 (1995)
Susnik, B., Worth, A., Palcic, B., et al.: Differences in quantitative nuclear features between ductal carcinoma in situ (DCIS) with and without accompanying invasive carcinoma in the surrounding breast. Anal. Cell. Pathol. 8(1), 39–52 (1995)
Grimm, P., Billiet, I., Bostwick, D., et al.: Comparative analysis of prostate-specific antigen free survival outcomes for patients with low, intermediate and high risk prostate cancer treatment by radical therapy. Results from the Prostate Cancer Results Study Group. BJU Int. 109, 22–29 (2012)
Enfield, K.S.S., Martin, S.D., Marshall, E.A., et al.: Hyperspectral cell sociology reveals spatial tumor immune cell interactions associated with lung cancer recurrence. J. Immunother. Cancer 7(1), 13 (2019)
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Switzerland AG
About this chapter
Cite this chapter
El Hallani, S., MacAulay, C., Guillaud, M. (2020). Digital Image Analysis in Pathology Using DNA Stain: Contributions in Cancer Diagnostics and Development of Prognostic and Theranostic Biomarkers. In: Holzinger, A., Goebel, R., Mengel, M., Müller, H. (eds) Artificial Intelligence and Machine Learning for Digital Pathology. Lecture Notes in Computer Science(), vol 12090. Springer, Cham. https://doi.org/10.1007/978-3-030-50402-1_15
Download citation
DOI: https://doi.org/10.1007/978-3-030-50402-1_15
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-50401-4
Online ISBN: 978-3-030-50402-1
eBook Packages: Computer ScienceComputer Science (R0)