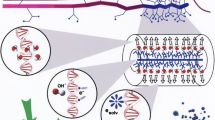
Abstract
In this chapter, we first give an overview of modeling approaches in the literature that are relevant for studying plasma for cancer treatment, including both macro-scale and atomic/molecular scale models. Subsequently, we will present typical results obtained by these models to illustrate the type of information that can be obtained. Finally, we will conclude and discuss some future challenges for modeling plasma for cancer treatment.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Plasma–cell interaction
- Multi-scale modeling
- Molecular dynamic simulations
- Plasma–liquid interaction
- Plasma oxidation
7.1 Introduction
Although plasma is very promising for cancer treatment, as shown in other chapters of this book, the underlying mechanisms are not yet fully understood. Hence, there is a clear need for a better understanding of these mechanisms. This can be obtained from experiments, but measurements cannot always provide detailed information, e.g., on the species concentrations in the plasma, or on the interaction of these species with biomolecules present in or around (cancer) cells. Such information can be obtained from modeling. Different types of models can be applied, depending on the system to be studied and the type of information required. In this chapter, we first give an overview of modeling approaches in literature, relevant for studying plasma for cancer treatment, including both macro-scale and atomic/molecular scale models. Subsequently, we will present typical results obtained by these models, to illustrate the type of information that can be obtained. Finally, we will conclude and discuss some future challenges for modeling plasma for cancer treatment.
7.2 Overview of Modeling Approaches in Literature
7.2.1 Macro-Scale Models for Plasma Sources and Plasma–Liquid Interaction
Several different plasma sources are in use for cancer treatment, as outlined in Chap. XX of this book, but most common are plasma jets (in various designs) and dielectric barrier discharges (DBDs). In addition, besides direct plasma treatment, where the plasma is directly applied on the tissue or cells to be treated, plasma can also be applied to a liquid, so that reactive oxygen and nitrogen species (RONS) produced by the plasma can be captured in the liquid, after which the liquid can be injected into the tissue. These so-called plasma-treated liquids (PTLs), or plasma-activated liquids (PALs) or plasma-activated media (PAM), are also gaining increasing interest, as explained in Chap. 6 of this book.
Various biomedical plasma sources, as well as plasma–liquid interaction, have been modeled by either zero-dimensional (0D) chemical kinetics models or 1D and 2D fluid dynamics models to reveal the behavior of the important RONS in the plasma, the effluent, and/or the liquid.
A 0D chemical kinetics model is based on solving balance equations for the various species (i.e., electrons, various ions, radicals, molecules, and excited species) as a function of time, with production and loss rates determined by the chemical reactions between all species. It can describe a large chemistry without too much computation time, which is its main advantage. On the other hand, it is based on some major approximations. For instance, it needs certain input data (e.g., electric field), and it assumes a uniform plasma (or liquid), and thus, it cannot capture spatial variations, e.g., concentration gradients from the gas–liquid interface towards the bulk liquid, although there exist solutions to overcome this limitation, by applying different 0D models for different subsequent layers in the liquid [1]. In addition, for a plasma jet, a 0D model can provide information on the species density evolution as a function of position in the effluent, by converting the calculated temporal evolution into a spatial evolution, based on the velocity profile of the feed gas, yielding in fact a pseudo-1D model [2].
Fluid dynamics models (in 1D, 2D, or in principle also 3D) are based on solving conservation equations for mass, momentum, and energy. Often, the momentum equation is replaced by a drift–diffusion approximation (transport of species by diffusion due to concentration gradients and drift or migration in the electric field), and the energy equation is often only solved for the electrons. Fluid dynamics models for plasma jets in contact with liquid can be quite complicated, as they need to couple both gas and liquid phase, describing the flow dynamics (often turbulent flow), heat transfer, transport of species, and chemistry, in both gas and liquid phase[3].
Several modeling studies were performed to study plasma bullet propagation in plasma jets, used in plasma medicine. Naidis et al. numerically demonstrated that these wavefronts propagate at very high speeds through the helium channel that develops in the ambient air, by including a simple photo-ionization function [4]. Nevertheless, later simulations (all performed for a pre-developed helium channel in ambient air) suggested that photo-ionization might not be crucial for the propagation of these ionization fronts, as the existence of a pre-ionization channel (created by previous bullets) was sufficient for their propagation [5, 6]. Nevertheless, including the photo-ionization phenomena did affect the velocity of the ionization wave [7], although a higher velocity could also be obtained by using a higher degree of pre-ionization. Anyway, the development of any ionization wave required both an electric field high enough in its front (to be able to ionize the gas) and the presence of seed electrons just ahead of the front of the ionization wave. The latter could be caused both “instantaneously” by photo-ionization or by a pre-ionized channel created by preceding bullets. It was shown by Yousfi et al. that the ionization rate rapidly drops when helium is significantly diluted by air (above 1%) [8]. Therefore, this explained why the mixing of helium with ambient air limits the bullet propagation both in the radial and axial direction. Breden et al. and Liu et al. [9] performed simulations in ambient helium and observed that the plasma (formed within the dielectric tube during the rise time of the voltage pulse) propagated along the surface of the dielectric during the rest of the pulse, without transitioning into a streamer. Finally, Kushner and coworkers simulated the propagation of ionization waves through flexible capillary channels, impinging upon a target, through branched tubes and across different channels, thereby mimicking experiments performed with the plasma gun of GREMI [10,11,12]. In addition, they also studied ionization waves launched from single and multiple plasma jets (pin-electrode setup) after a steady state was obtained for the neutral fluid flow of helium containing 2% oxygen into ambient humid air, taking into account photo-ionization processes. They found that the jets interact through electrostatic, hydrodynamic, and photolytic means [13].
More recently, the effects of various admixtures to a helium feed gas on the evolution of a plasma jet or plasma gun have been investigated. These admixtures include, e.g., nitrogen, oxygen, or air, which are often used in plasma medicine. Naidis et al. [14] focused on the addition of air to a helium-driven plasma jet, whereas Bourdon et al. [15] investigated nitrogen admixtures in a helium-driven plasma gun. Finally, Lazarou et al. [16] employed 2D simulations to study the effect of oxygen admixtures in a helium-driven plasma jet. These studies showed that the dynamics of the plasma devices strongly depend on the composition of the feed gas. Indeed, adding oxygen (1) promoted plasma bullet propagation on the axis of symmetry of the tube while (2) lowering the propagation speed of the plasma bullets. At the same time, much higher electric fields were induced on a dielectric surface treated (in comparison to the pure helium plasma jet) [16]. In the case of nitrogen addition, Bourdon et al. [15] showed that two- and three-body Penning reactions are crucial for the discharge dynamics. Moreover, it was shown that the ionization front velocity increased when increasing the amplitude of the applied voltage. Overall, it was concluded that the ionization front velocity at different levels of nitrogen admixture in the helium gas depends on a complex coupling between the kinetics of the discharge, the photo-ionization, and the 2D structure of the discharge in the tube. In a follow-up study, Viergas et al. [17] showed that the peak electric field is able to propagate outside the plasma jet device. After impact of the ionization front on a metallic target, a rebound front is shown to propagate in the opposite direction, with a velocity 2–3 times higher than the first ionization front.
Recently, 2D models developed by Lietz et al. [18] showed that molecular admixtures (O2 and H2O) can significantly impact the ionization wave propagation speed, the electron density, the electron temperature, and the RONS production in a helium-driven plasma jet. Moreover, they also showed that, in the absence of intentional admixtures, impurities in the feed gas can account for 79–98% of the production of most RONS [18]. Chang et al. [19] investigated the effect of seed electrons on the repeatability of plasma plume propagation in a plasma jet, using helium as feed gas with different concentrations of air added. They found that the minimum seed electron density required for repeatable propagation mode is independent of the air concentration in the feed gas. The main source of the seed electrons, however, depends on the feed gas composition. Indeed, at low air concentrations (0.25%), the main source of seed electrons is due to leftover electrons from previous discharges. At an air concentration of 0.50%, on the other hand, the detachment of O2 − is the main source of the seed electrons [19]. The rate of this detachment was calculated based on Monte Carlo simulations.
A large number of models have also been developed with specific focus on the plasma chemistry in plasma jets (and their effluent) and DBD plasmas, used in plasma medicine applications. Sakiyama et al. [20] developed two coupled 0D models to describe a surface micro-discharge with 50 species and 600 reactions. One of the 0D models described the plasma zone near the electrodes, with a high electric field, thus solving balance equations for both neutral and charged species, whereas the other model only described the neutrals and highlighted the chemistry of the long-lived species. Babaeva et al. [21] developed a combined 0D–2D fluid model to provide more insight into the effect of filaments on the chemical formation of species and the particle fluxes towards wounded skin. The 2D results were used as input for a 0D model calculation and vice versa. The 2D fluid was able to describe streamer development in air between the electrode and the tissue, which acts as a floating electrode, but only for short time scales due to the computational load [22, 23]. The 0D model was used in a complementary way to evaluate which chemical species were formed on the long term from the short-lived chemical species created in the streamers simulated in 2D. Also interesting is that the authors included a cellular structure in the model, mimicking actual skin tissue, and investigated the induced electric fields in the context of electroporation. Later, this work was extended to wounds covered by liquid, possibly containing blood platelets [24]. Similarly, Naidis coupled his 2D fluid model, describing the streamer or “bullet” behavior of a helium plasma jet, with a small 0D chemical kinetics model (He/H2O) for describing the OH radical production mechanism inside the device, thus not in the effluent [25].
An interesting 0D numerical study was reported by Liu et al. [26] with a quite large He/H2O chemistry set, containing 46 species and 577 reactions. In this publication, the authors also suggested several reduced chemistries that can be used to run complex simulations. An example of such a model was the work of Waskoenig et al. [27] for a He/O2 mixture with 16 species and 116 reactions. This code was developed to comprise chemical and physical phenomena across the gap between two parallel plate electrodes (hence 1D) of a micro-scaled radio-frequency driven atmospheric pressure plasma jet device (μAPPJ), designed for plasma medicine applications. Results of a similar model were reported by McKay et al. [28] but with the addition of water to the He–O2 mixture. In this study, a very large reaction chemistry set of 61 species and 878 reactions was implemented in a 1D fluid model. The chemistry of the μAPPJ was further studied by Murakami et al. [29,30,31], by means of a global model. The lower complexity of the latter enabled these authors to do a faster kinetic analysis of a He/O2/N2/H2O/CO2 mixture with 59 species and 1048 reactions.
Another plasma medicine tool that has been studied numerically is the plasma needle, initially by Brok et al. and later by Sakiyama and Graves. Brok was the first one to model in 2D (in the field of plasma medicine at least) and his simulations ran for a fixed He/N2 gas ratio over the entire simulation domain, neglecting the gas flow and using only a limited plasma chemistry [32]. The approach of Sakiyama and Graves was similar, yet already more advanced, as they used an unstructured mesh (and not a Cartesian mesh like in Brok’s work), which enabled a high resolution close to the needle electrode tip. Therefore, these authors were able to accurately model the sheath and they found that the corona discharge at low power exhibits a transition to a glow discharge at a critical power, spreading back along the needle surface [33, 34]. Subsequently, they studied this discharge type with an asymmetric electrode in a more fundamental way in 1D spherical coordinates, e.g., looking at the validity of the local field approximation, the influence of nitrogen impurities, and secondary electron emission [35, 36]. Finally, they published the results of a 2D fluid calculation that considered the neutral gas flow (i.e., ambient air diffusion, here represented by N2) [37].
While the above models focused mainly on helium plasma jets, Van Gaens and Bogaerts developed a 0D model for an argon plasma jet flowing into humid air, considering 84 different species and 1880 reactions [2]. It was first applied to the experimental conditions of a plasma jet geometry developed at Eindhoven University of Technology. In a follow-up study [38], the gas phase production and loss pathways of the most important biomedical species were examined, showing that the pathways of the biomedical species drastically change as a function of position in the effluent, i.e., inside the discharge device, the active plasma jet effluent and the afterglow region far from the nozzle, and are also strongly dependent on the conditions, such as ambient air humidity, total deposited power into the plasma, and gas temperature along the jet. In [39] complementary numerical work and experiments were presented on the ozone generation in the same plasma jet from Eindhoven University of Technology. Another combined experimental-computational study was applied to two cold atmospheric pressure plasma jets [40], operating in Ar + 2% air, with a different electrode geometry but the same power dissipated in the plasma. This combined work allowed to derive the most probable spatial power distribution in an RF-driven plasma jet, by comparing measured and calculated species density profiles, because of the strong effect of the power distribution on the NO and O density profiles. In addition, the dominant reaction pathways for both NO and O were identified. In [41], the kINPen plasma jet was studied, operating in argon with different admixture fractions up to 1% pure N2, O2, and N2 + O2, and a shielding gas of dry air, focusing on the production of O3 and NO2, in comparison with experiments performed in the far effluent by infrared absorption spectroscopy.
Schmidt-Bleker et al. [42] also developed a 0D model for the kINPen plasma jet, with a reduced reaction scheme, to model the chemistry of the effluent without including the electron dynamics. By varying the shield gas humidity and composition (i.e., O2/N2 ratio) they obtained valuable insights regarding the chemistry of the plasma jet effluent. An extension to this work [43] took different time- and length-scales into account by using an approach involving two separate kinetic models. A first model accounts for the electron impact reactions occurring in the vicinity of the streamer head, whereas the second model only accounts for neutral molecules on a much larger time- and length-scale, using the obtained densities of the first model as an input. The major finding was that primary RONS are dominantly generated from exited Ar species.
Next to the kINPen, 0D models are also employed to study the COST reference microplasma jet. Indeed, Gorbanev et al. [44] used such simulations to study the chemistry occurring inside the source, as well as during transport to a liquid surface. Moreover, a 3D fluid dynamics model was developed to assess the humidity induced in the effluent by evaporation of water from the liquid target. In a follow-up study [45], the 0D model was used to help unraveling reaction pathways leading to the formation of aqueous ClO−. Schröter et al. [46] employed 0D simulations to study the effect of surface reaction probabilities of reactive neutral species (H, O, and OH). They found that the densities of low-mass species (e.g., H) can change significantly based on the choice of the surface reaction probabilities. This in turn offers potential for tailoring the composition of these plasmas by choosing different wall materials or plasma geometries.
Besides gas phase simulations, plasma–liquid modeling is also crucial for plasma medicine applications, for two reasons: (1) the growing interest in PTLs and (2) even in direct plasma treatment, biological samples are usually covered by a liquid medium. The chemical cocktail in the gas phase is not necessarily the same after the gas phase species are transferred to the liquid phase.
Plasma–liquid modeling has also been studied by 0D chemical kinetics and (mainly 2D) fluid dynamics models, for medical (as well as environmental) applications of plasmas. Locke et al. [47] wrote a general review paper on modeling of electrical discharges in liquid water, including streamer-like plasma channels, spark and arc discharges. van Gils et al. [48] presented a 0D chemical kinetics model for the liquid phase chemistry upon plasma treatment, assuming certain values for the fluxes of O3, NO, and OH originating from the plasma, using an optimization algorithm, to find the values at which the end concentrations of nitrate and nitrite correspond to the values measured by ion chromatography in the plasma-treated liquid. They used the model to elucidate the bacterial inactivation mechanisms in the liquid phase. Likewise, Lukes et al. [49] performed a kinetic study of the aqueous-phase chemistry, and more specifically the formation of peroxynitrite in plasma-activated water, also to investigate the bactericidal effects of an air discharge in contact with water. Hamaguchi and coworkers [50] developed a 1D reaction–diffusion model, as well as a 0D model, to describe the behavior of reactive species in pure water (with dissolved O2 and N2 in equilibrium with air) exposed to an atmospheric pressure plasma. The model contained 37 species and 111 chemical reactions. They estimated the flux of RONS towards the liquid (i.e., the initial conditions for the 0D model of the liquid), based on their typical gas phase concentrations in a representative plasma medicine device, and on the Henry’s constants, which determine how easily a species dissolves in liquid. Chen et al. [51] evaluated the plasma–liquid chemistry for a He/O2 DBD with a 0D model, for treating biofilms and biological tissues. Later, the same group [52] developed a semi-1D model for plasma–biofilm interaction and for plasma–tissue interaction, based on a reactive penetration model for mass transfer of plasma species across the gas–liquid boundary. The liquid phase model contained 19 species and 84 chemical reactions.
The most comprehensive models for describing (DBD) plasma–liquid interaction, especially for the purpose of plasma for wound treatment, were developed by Kushner and coworkers. As mentioned before, Babaeva et al. developed a combined 0D–2D multi-species fluid model, called nonPDPSIM, for atmospheric pressure DBDs interacting with slightly wounded skin, focusing on a dry wound [21]. This model was later extended by Tian and Kushner [53, 54] to the interaction of DBD plasma with liquid-covered tissue. The water layer (with liquid phase chemistry) was represented by an extremely dense gas and separated from the actual gas phase (with gas phase chemistry) by a boundary layer. Transport of RONS across this boundary was also dictated by Henry’s law. Also water evaporation from the liquid layer and photolysis of water molecules into OHliq (triggered by photons created in the plasma above the liquid layer) was taken into account. The authors also found that ROS are readily consumed by hydrocarbons in the liquid, thus affecting their concentration [53, 54]. The same group also studied the interaction of DBD filaments with wounds and blood plasma, focusing on the electric fields delivered to blood platelets and cells [24], as well as the interaction of a DBD treating liquid-covered tissue [55]. Lietz et al. [55, 56] studied plasma–liquid interactions by coupling two 0D models. In the first model, the gas phase chemistry of a pulsed DBD operating in ambient air is studied. The second model investigates the chemistry occurring in a liquid target. These two models are characterized by a separate set of chemical species (gaseous and aqueous) as well as a separate set of chemical reactions. The species in these two zones (gas and liquid) only interact through the interface between the zones by means of diffusion into or out of the liquid, which is based on Henry’s law. By using two 0D models, elaborate chemistry sets can be investigated on large time scales, which allowed the authors to model 5000 consecutive pulses, followed by an afterglow period of 5 min. The effect of the applied voltage, gas flow rate, pulse repetition frequency, and the presence of organic molecules in the liquid was investigated [55, 56].
To the best of our knowledge, only a limited number of papers reported the interaction between a plasma jet and a liquid. Van Boxem et al. applied a 0D model to a liquid treated by the kINPen plasma jet, to elucidate the reaction mechanisms for the formation of H2O2 and HNO2 in order to explain experimental observations [57]. Du et al. [58] developed a 1D drift–diffusion model for the mass transfer process from a plasma jet into the liquid. A few papers also reported 2D simulations for the interaction between a plasma jet and a liquid layer. Lindsay et al. [59] developed a 2D model to study the transport of reactive species produced by a helium plasma jet to liquid water, using 13 species and 23 reactions. Lietz and Kushner [18] studied the effect of H2O and O2 admixtures in the feed gas of a helium plasma jet, by means of a 2D fluid dynamics model. Norberg et al. [60] investigated the influence of the pulse repetition frequency on the plasma treatment of a reactive liquid layer. Verlackt et al. [61] studied the transport and accumulation of plasma-generated species in aqueous solution by a 2D fluid dynamics model, considering 20 gas phase and 22 liquid species, and 57 and 42 reactions in the gas and liquid phase, respectively. This model was recently extended by Heirman et al. [3], by coupling it to a 0D model, in order to take advantage and avoid the limitations of both models (in terms of calculation time vs approximations). Furthermore, this model also showed simulation results after plasma treatment, to reveal the stability of RONS in the liquid after treatment. The 0D model included 91 gas phase and 32 liquid species, which interact in 1390 gas phase and 89 liquid phase reactions, while the 2D model included 21 gas phase and 25 liquid species, interacting in 56 gas phase and 52 liquid phase reactions. This clearly illustrates the difference between the number of species, and especially number of reactions, in gas phase and liquid between 2D and 0D models, as explained in the beginning of this section.
7.2.2 Atomic Scale Models for the Interaction of RONS with Biomolecules
Besides modeling the plasma devices and plasma–liquid interactions, studying the interaction of RONS with biomolecules, important in the context of plasma for cancer treatment, such as the phospholipid bilayer (PLB), DNA, and proteins, is also needed to obtain a better insight in the underlying mechanisms. The same applies to the effect of these interactions (e.g., oxidation) on structural changes in the biomolecules. For these purposes, atomic (or molecular) scale models must be developed, as macroscopic models will not provide the required information. A wide range of atomic/molecular scale methods can be applied, ranging from quantum mechanics, density functional theory (DFT), density functional tight binding (DFTB), classical reactive molecular dynamics (MD), and non-reactive MD, including all-atom, united-atom, and coarse-grained approaches (see below). These methods vary in accuracy and level of detail they can describe, as well as the time- and length-scales they can handle, as illustrated in Fig. 7.1.
Overview of the computational methods that allow to obtain atomic (or molecular) level insight in the interaction of RONS with biomolecular systems, or the (longer-term) effects of these interactions, illustrating the length- and time-scales that can be handled. QM = quantum mechanics, DFTB = density functional-tight binding, QM/MM = quantum mechanics/molecular mechanics, rMD = reactive molecular dynamics, nrMD = non-reactive molecular dynamics, uaMD = united-atom molecular dynamics, cgMD = coarse-grained molecular dynamics. Adopted from [62] with permission
Quantum mechanical (QM) calculations are the most accurate computational methods, as they are based on first principles. Various QM techniques exist, varying in their approach to solve the Schrödinger equation and in their level of approximations (see more details in [62]). For plasma medicine, DFT calculations are the most suitable approach, keeping in mind the required system sizes. They are more approximate than other QM methods, but still very time-consuming, able to handle system sizes only in the order of 100 atoms. In principle, they are static, but they can be integrated in MD simulations, where all atoms in the system are followed through space and time, by integrating their equations of motion. The forces acting on the atoms are obtained as the derivative of some suitable interatomic potential, which can thus be based on QM data. DFT-based MD calculations, also called “ab initio MD” (AIMD) can handle time scales in the order of picoseconds.
Somewhat larger systems can be handled with DFTB, which is an approximate DFT method, based on a Taylor series expansion of the DFT total energy expression [63]. DFTB can typically handle a few 1000 atoms on time scales of 10s of picoseconds. This method has been applied in the context of plasma medicine to study the interaction of ROS with the head group of the PLB [64], a specific protein (P-glycoprotein) [65] and peptides [66], as well as the behavior of O and OH in water [67].
Classical reactive MD simulations, in which the forces are not based on QM data but on classical fitting parameters, can typically handle much longer time scales and larger systems than DFT or DFTB calculations, ranging from 1 ps to 100 ns, and from 104 to 106 atoms, depending on the complexity of the interatomic potential. This (classical) potential is constructed with many parameters obtained by fitting against DFT calculations. The Brenner potential [68] and the ReaxFF potential [69] are two reactive potentials used for plasma medicine applications. Reactive MD simulations can describe bond breaking and formation, so they allow to study chemical reactions of RONS with biomolecules. This has been illustrated for the interaction of ROS with peptidoglycan [70, 71], lipid A [72], lipids [23, 73, 74], DNA [75, 76], a water layer [77], and simple organic molecules in water [78].
As opposed to reactive classical MD simulations, in non-reactive MD simulations, also called “molecular mechanics” (MM), the molecular connectivity in the system is fixed. Hence, the bond order of each bond does not have to be recalculated in every step, so they can handle system sizes and time scales two orders of magnitude larger than reactive MD simulations, hence, in the order of 106–108 atoms, at time scales of 0.1 ns to 10 μs, for the so-called all-atom force fields. A drawback is that they cannot describe chemical reactions (as they do not consider bond breaking and formation), but as they can follow the system over a longer time scale, they can study conformational changes and stability of biomolecules.
In all-atom force fields, as the name suggests, all atoms in the system are treated separately. In addition, non-reactive MD simulations can also use the so-called united-atom and coarse-grained force fields, which can handle even larger system sizes (typically up to one order of magnitude larger), for the same time scales. United-atom force fields (e.g., [79]) treat all heavy atoms separately, but the H atoms bound to a C atom (e.g., in the apolar tails of phospholipids) are combined and treated as one (methyl or methylene) group. Hence, this yields a lower number of separate particles in the system, allowing to simulate larger systems. Some well-known non-reactive interatomic potentials are AMBER [80], CHARMM [81], and GROMOS [82]. Coarse-grained methods further reduce the number of particles in the system, and thus they allow even larger system sizes (or speed up the calculations), because the atoms comprising entire functional groups, i.e., typically, 3–5 heavy atoms with their H atoms, are represented by the so-called coarse-grained particles. The Martini force field [83] is an example of a coarse-grained potential.
Finally, it is also possible to combine the above methods in the so-called quantum mechanical/molecular mechanics (QM/MM) methods [84]. Indeed, it is clear that the various methods can handle different length- and time-scales (see Fig. 7.1), attributed to the accuracy and type of information they provide. In the QM/MM method, a small (chemically most relevant) part of the system (e.g., the active site of the biological system) will be described at the quantum chemical (electronic) level, while the surrounding embedding atoms and molecules will be treated at a classical (atomic) level.
For more details on these methods and examples of their simulation results for plasma medicine, we refer to [62, 85].
7.3 Typical Modeling Results
7.3.1 Gas Phase Simulations
7.3.1.1 0D Modeling Results
Figure 7.2 illustrates typical results obtained by a 0D model of a plasma jet. Using the pseudo-1D model approach discussed above, the evolution of species along the plasma axis can be obtained. These results illustrate, e.g., the difference in behavior of short- and long-lived RONS. The densities of short-lived RONS, such as OH or exited species, increase rapidly in the effluent when ambient air species enter the effluent. However, during transport to a target, these species recombine again into more stable, long-lived RONS, which are therefore generated gradually throughout the effluent.
Calculated density profiles of different RONS in the effluent, obtained with a 0D model for the kINPen. Adopted from [43] with permission
Due to the low computational cost of such 0D models, the effect of different parameters on the generation of species can easily be investigated, including, e.g., the gas temperature at the nozzle exit, the flow rate of the feed gas, or the humidity of the ambient air. The effect of varying the flow rate of the feed gas in an atmospheric pressure argon plasma jet is illustrated in Fig. 7.3. This shows that altering one operational parameter can have different effects on different important species. Indeed, lowering the feed gas velocity yields an increase in the density of, e.g., H2O2, HNO2, or HNO3, whereas the density of NO, OH, and O2 − decreases.
Calculated relative changes (%) in the density of different biomedically important RONS upon altering the flow rate of the feed gas, compared to a flow rate of 2 slm, as obtained from 0D modeling. In case (a), the flow rate is lowered to 1.5 slm, whereas in case (b), it is enhanced to 2.5 slm. Adopted from [2] with permission
7.3.1.2 2D Modeling Results
Tian et al. [53] employed 2D simulations to study the interaction of DBDs in humid air with a thin water layer covering tissue. Such a model allows studying the spatio-temporal evolution of species in the discharge zone, as well as in the water layer, during consecutive filament discharges onto the liquid surface and afterglow periods. One of the important observations made in this study is the effect of water evaporation from the water layer covering the tissue. Indeed, when accounting for this evaporation, the densities of different important species changed significantly, as is illustrated in Fig. 7.4.
Effect of water evaporation on the density profiles of H2O+ ions and OH radicals during a filament incident on the water surface, as obtained from 2D modeling. Contours for H2O+ and OH are plotted on a log scale over three decades with the maximum density shown in each frame. Adopted from [53] with permission
As mentioned before, Bourdon et al. [15] employed 2D simulations to study the propagation of the ionization front in a plasma gun setup. Using these models, they discovered that Penning reactions not only have an effect on the discharge dynamics, but also on its structure, as illustrated in Fig. 7.5. As Penning reactions produce ions and electrons, excluding these reactions makes the plasma column behind the ionization front less conductive, which leads to a slower propagation of the discharge in the tube.
Cross-sectional views of the magnitude of the electric field and electron density, as obtained from 2D modeling, at 1500 ns, for an applied voltage of 16 kV, 1000 ppm of N2, with a pre-ionization level n init = 104 cm−3 and for a photo-ionization coefficient A ph = 10−2, with the reference kinetic scheme presented in [15], as well as without two- and three-body Penning reactions. Adopted from [15] with permission
7.3.2 Gas–Liquid Interactions and Liquid Phase Simulations
7.3.2.1 0D Modeling Results
Lietz et al. [55] used a coupling of two 0D models (also called global models) to investigate the chemistry in the gas and liquid phase of a DBD device interacting with a water layer. As the computational cost of 0D models is much lower than for 2D models, the authors were able to study the chemistry in 5000 consecutive pulses, followed by an afterglow of several minutes. Figure 7.6 shows the accumulation of species in the water layer during the first 50 pulses (a–c), as well as the decay of some important RONS after treatment (d). After treatment, gas phase species continue to solvate, which serves as a source term of the aqueous species. Liquid reactions, on the other hand, lead to a depletion of these species.
The accumulation of RNS (a) and ROS (b, c) during the first 50 pulses, as well as the decay of some important RONS after plasma treatment (d), as obtained from 0D modeling. The time refers to the time after the last pulse. Adopted from [55] with permission
In the same paper, Lietz et al. investigated the effect of hydrocarbons present in the liquid on the generation and stability of aqueous species. Their results clearly show that these hydrocarbons (e.g., peptidoglycan) have a big impact on the density of certain RONS, due to reactions occurring between plasma-generated RONS and hydrocarbons, which is shown in Fig. 7.7.
Calculated (a) gas phase and (b) liquid phase densities of different RONS after the 5000th discharge pulse, with and without the presence of peptidoglycan in the liquid. O, OH, O3, and H2O2 are able to react directly with peptidoglycan in this mechanism, so their densities are zero in the liquid upon presence of peptidoglycan. Adopted from [55] with permission
7.3.2.2 2D Modeling Results
2D fluid dynamics models also provide information on the flow profiles in both gas and liquid phase. This is illustrated in Fig. 7.8, for the kINPen plasma jet interacting with water, contained in a well of 2 mL, at a flow rate of 3 slm Ar [3]. A maximum velocity of 57 m/s inside the plasma jet was predicted, while the maximum gas velocity outside the jet is 34 m/s. When the gas flow reaches the liquid surface, it flows towards the edge of the well, causing a shear stress on the liquid surface. Upon hitting the edge of the well, the gas flow results in a vortex within the well, so that the gas flows back towards the afterglow. Because of the shear stress on the liquid surface, the upper layer of the liquid starts to move in the same direction as the gas, but with a lower velocity. The maximum velocity in the liquid (2.5 m/s) is reached near the edge of the well. The liquid movement results in another vortex in the liquid phase (see close-up in Fig. 7.8). Note that this vortex behavior is characteristic for the small volume of the liquid. In larger volumes, other vortex patterns can be observed [61].
Calculated gas and liquid flow profiles, obtained with a 2D fluid dynamics model for the kINPen plasma jet interacting with water, contained in a well of 2 mL, at an inlet argon flow rate of 3 slm. The color scale gives the velocity magnitudes, while the arrows show the flow direction. Adopted from [3]—reproduced by permission of the PCCP Owner Societies
The calculated gas phase densities and liquid phase concentrations of the three major RONS, i.e., H2O2, HNO2, and HNO3, obtained with the same model and at the same conditions, are plotted in Fig. 7.9, for 10 s of plasma treatment [3]. The profiles clearly follow the gas and liquid flow profiles of Fig. 7.8. Indeed, convection of species was found to be more important than diffusion. The model predicts that the H2O2 density is at maximum underneath the plasma jet, just above the liquid surface, and its density drops to half its value towards the edge of the well and in the bulk gas. HNO2 is only formed to some extent underneath the plasma jet, just above the liquid surface, but its density reaches its maximum towards the edge of the well. Finally, the HNO3 density profile does not rise at all underneath the plasma jet. The HNO2 and HNO3 species accumulate in the gas phase inside the vortex.
2D plots of the gas phase densities and liquid phase concentrations of H2O2, HNO2, and HNO2, calculated with a 2D fluid dynamics model for the kINPen plasma jet interacting with water, contained in a well of 2 mL, at an inlet Ar flow rate of 3 slm, for 10 s of plasma treatment. Adopted from [3]—reproduced by permission of the PCCP Owner Societies
Transport to the liquid phase is dictated by the Henry’s constants, which are above 1 for H2O2, HNO2, and HNO3. Hence, their equilibrium is towards the liquid phase. This is clear from the difference between gas phase density just above the liquid surface (being small, because the species are transported towards the liquid) and the liquid phase concentration just below the liquid surface (being the highest in the first few nm, because the species enter from the gas phase); see close-ups in Fig. 7.9.
The highest concentration of H2O2 in the liquid was calculated to be ten times higher than that of HNO2, which is ten times higher than that of HNO3 (cf. different values in the color scales of Fig. 7.9).
This model also calculated the behavior of the RONS in the liquid after plasma treatment, to check their stability, which is very important for the application of PTLs. It was predicted that only H2O2, HNO2, HNO3, HO2, O3, and ONOOH can accumulate (to some extent) in the liquid (the so-called long-lived liquid species), while the other RONS, such as OH, NO, and NO2, are only important in the first few nm below the liquid surface and have a lifetime around 10 ms (“short-lived species”).
Figure 7.10 illustrates the volume-integrated liquid phase concentrations of the long-lived RONS, as a function of time, both during and after plasma treatment, for the same conditions as Figs. 7.8 and 7.9. During plasma treatment (10 s), the H2O2, HNO2, and HNO3 concentrations increase linearly with time, which is in agreement with experimental data [57]. The concentrations of O3 and ONOOH rise less than linearly with treatment time, while the HO2 concentration rises significantly in the first milliseconds, followed by a steady state till the end of plasma treatment.
Volume-integrated liquid phase concentrations of the long-lived RONS, as a function of time during and after plasma treatment, calculated with a 2D fluid dynamics model for the kINPen plasma jet interacting with water, contained in a well of 2 mL, at an inlet argon flow rate of 3 slm, for 10 s of plasma treatment. Adopted from [3]—reproduced by permission of the PCCP Owner Societies
After plasma treatment (i.e., after 10 s), obviously H2O2, HNO2, and HNO3 are the only stable species in the liquid, with a constant concentration, at least in the first 10 s after plasma treatment. This could be explained by the model because both the production and loss processes of these species in the liquid become negligible after plasma treatment, as their rates depend on short-lived radicals, which are not present anymore after treatment. This stability also correlates with experiments [57], where the H2O2 and HNO2 concentrations were observed not to change in the plasma-treated liquid for up to 2 h after plasma treatment (HNO3 was not measured). On the other hand, the concentrations of HO2, ONOOH, and NO2 drop quickly after plasma treatment, to virtually zero after 10 s, while the O3 concentration decreases linearly. After 10 s, it reaches a value equal to 80% of its value at the end of plasma treatment. Extrapolating this linear drop to longer times would mean that it takes about 50 s for the O3 concentration to drop to negligible values. This drop in O3 concentration can be explained by transport of O3 back to the gas phase, because its Henry’s constant is lower than 1; in order words, its equilibrium points towards the gas phase.
Note that the calculated liquid phase concentrations of H2O2, HNO2, and HNO3 in this model are significantly underestimated, compared to experiments [3]. This was attributed to the static liquid interface defined in the model. It was argued that a turbulently moving liquid layer would deplete the liquid phase concentrations in the interface, so that the gas-to-liquid equilibrium shifts towards the liquid, which will enhance the liquid phase concentrations, but this hypothesis will need to be investigated further.
Tian et al. [53] also investigated the transport of RONS from the gas phase into a liquid layer upon treatment of a water layer with a DBD device. Figure 7.11 illustrates the time evolution of the OH and H2O2 densities in the (top) gas gap and (bottom) 200 μm water layer. These results again clearly illustrate the dual behavior of (1) short-lived species (such as OH), which are lost in the liquid interface, versus (2) long-lived species (such as H2O2), which accumulate in the bulk of the liquid and can reach targets below the water layer. The generation of OHaq was attributed to both diffusion and solvation processes, as well as photo-dissociation at the surface of the water by VUV produced by the plasma.
Time evolution of the OH and H2O2 densities in the (top) gas gap and (bottom) 200 μm water layer, as obtained from 2D fluid dynamics simulations. The densities of OH and OHaq are shown during the first pulse and interpulse afterglow. H2O2 and H2O2aq densities are shown accumulating after each pulse (0, 10 ms, 20 ms) and during the terminal afterglow. The contours are plotted on a 3-decade log scale with the maximum values noted in each frame. Adopted from [53] with permission
7.3.3 Permeability of RONS and Pore Formation Through Native and Oxidized Phospholipid Bilayers (PLBs)
7.3.3.1 Different Permeability of Hydrophilic and Hydrophobic RONS
When plasma-produced RONS come in contact with cells, either directly or through a liquid layer covering cells, or upon application of PTLs to cells, they first come in contact with the cell membrane. The latter is composed of a PLB with proteins embedded. Hence, it is interesting to investigate the permeability of RONS through the PLB. In addition, as plasma-produced RONS can oxidize the PLB, it is interesting to compare the RONS permeability through both native and oxidized PLBs. Although the cell membrane consists of both lipids and proteins, each contributing for about 50% to the mass of the cell membrane, the PLB plays a crucial role in the structure of the cell membrane, so it makes sense, in first instance, to study the RONS permeability through only the PLB. Figure 7.12 depicts the free energy profiles (FEPs), obtained from umbrella sampling (US) MD simulations, of various ROS and RNS across both native and 50% oxidized PLBs, assuming oxidation of the lipid tails into aldehyde (see details in [86]). The structure of the PLB is drawn in pale color behind the FEPs. The center of the PLB is at z = 0, the head groups are around z = ± 2 nm, and beyond this distance is the water phase surrounding the PLB.
As is clear from Fig. 7.12a, when the hydrophilic ROS (i.e., OH, HO2, H2O2) move from the water phase to the PLB center, their FEP first reaches a minimum around the head groups, in agreement with [87], followed by a steep rise towards the center, thus creating a clear energy barrier when they cross the PLB. In case of the oxidized PLB, this free energy barrier is greatly reduced (see Fig. 7.12b). Indeed, upon oxidation, the hydrophilicity of the PLB rises, resulting in a larger permeability for hydrophilic ROS.
As the hydrophilic ROS prefer to reside close to the head groups, it is interesting to study oxidation of the head groups of the PLB, which was performed by DFTB [64]. While HO2 and H2O2 molecules do not seem to react with the head groups and only show weak attractive non-bonded interactions, OH radicals do react with the head groups, resulting in detachment of some parts in the lipids, which reduces the lipid order and enhances the membrane fluidity, in agreement with experiments [64]. The latter might allow RONS to penetrate more easily through the PLB, causing further lipid tail (per)oxidation, which might result in pore formation (see next section). Note that OH radicals react with virtually all biomolecules and thus, oxidation of the head groups will only occur if the OH radicals are generated nearby the cell membrane, due to their small diffusion distance.
As is obvious from Fig. 7.12c, d, the hydrophobic ROS (such as O2 and O3) and RNS (e.g., NO, NO2, and N2O4) show a completely different behavior, with very low permeation barriers around the PLB head groups and minima in the center (compared to the water phase). Indeed, these species prefer to reside in the lipid tail region, where they can cause lipid (per)oxidation. Upon oxidation of the PLB, the FEPs of these hydrophobic ROS and RNS become a bit smoother, due to the higher membrane fluidity [86], but otherwise they do not change significantly. Note that hydrophilic RNS, like HNO2, HNO3/NO3 −, and ONOOH, might behave the same as the hydrophilic ROS, but their permeability across the PLB could not yet be investigated, as no force fields are available.
These simulation results are in good qualitative correlation with experiments, where the permeability of hydrophobic RONS (NO and O2) was found to be 3–6 orders of magnitude higher than the permeability of hydrophilic ROS (H2O2) [88, 89]. Obviously, pores or aquaporin (AQP) channels will be needed for the active transport of hydrophilic ROS in and out of the cell, as will be illustrated in the next sections, while hydrophobic RONS may permeate through the cell membrane even in the absence of AQP channels and pores.
7.3.3.2 Effect of Oxidation on Pore Formation in the PLB
As hydrophilic RO(N)S cannot easily penetrate through the PLB, caused by their high free energy barriers, it is interesting to study pore formation in the cell membrane after lipid (per)oxidation. For this purpose, some lipid (per)oxidation products, based on data from literature [90], were added to the model systems of a native PLB, with concentrations varying between 0 and 100% (see details in [91]).
Non-reactive MD simulations revealed that, upon oxidation, the thickness of the PLB drops, followed by a rise when the oxidation reaches 100%, and this was attributed to pore formation [91]. Figure 7.13 illustrates such a pore formation for a PLB with 100% lipid oxidation. The initial conformation (after 10 ns) does not show water defects (Fig. 7.13a), while after 40 ns (Fig. 7.13b), there is a significant amount of water in the center of the PLB, and after 80 ns (Fig. 7.13c), a pore is formed with diameter of 15 Å, which might allow RONS to penetrate and reach the cell interior. Figure 7.13 also indicates the thickness of the PLB: as pore formation allows water to enter the PLB, this yields swelling, resulting in a somewhat thicker bilayer (cf. Fig. 7.13c vs Fig. 7.13a, b). Other MD simulations in literature on oxidized PLBs [92,93,94] also reported an overall rise in membrane permeability [92], a change in the lipid mobility [93], as well as pore formation and bilayer disintegration [94] upon lipid oxidation.
Snapshot of MD simulations, at 10 ns (a), 40 ns (b), and 80 ns (c), illustrating pore formation in a model system of a PLB with 100% oxidation. A pore with diameter of ca. 15 Å is formed in (c). Adopted from [91]—Published by The Royal Society of Chemistry
7.3.3.3 Synergistic Effect of Plasma Oxidation and Electric Field on Pore Formation
Besides plasma oxidation, electric fields can also cause pore formation, which is applied in the so-called electroporation [95,96,97]. As some biomedical plasma sources generate electric fields, with values in the order of a few up to 100 kV/cm (see, e.g., [98,99,100]), these electric fields might also contribute to pore formation in the cell membrane, and perhaps there is even synergy with plasma-induced lipid oxidation.
Figure 7.14 indeed illustrates that lipid oxidation yields a drop in the electric field threshold needed for pore formation in the PLB, as well as a shorter average pore formation time. More details can be found in [101]. Note that the pore formation time can fluctuate considerably, resulting in large error bars in the figure, but the latter cannot be reduced by increasing the number of simulations, because pore formation is a stochastic process. Nevertheless, it is clearly visible that oxidation of the lipid tails in the PLB facilitates pore formation, by lowering the threshold electric field, as well as the pore formation time.
Average pore formation time for three different electric field values, as a function of the oxidation degree of the PLB, for lipid oxidation into aldehydes. Adopted from [101] with permission
We need to remark that the applied electric fields in electroporation simulations are typically much stronger than the fields used in electroporation or plasma medicine [98,99,100]. Indeed, the latter typically vary between 0.1 and 100 kV/cm (corresponding to 0.01 and 10 mV/nm). However, it should be noted that the macroscopic field applied in experiments is not at all equivalent to the field experienced by the cell membrane (and thus applied in MD simulations), so the actual electric field values should not be directly compared, as explained in detail in [101]. In addition, it is not possible to directly relate the average pore formation times obtained in MD simulations to the experimental pore formation kinetics, as also explained in [101]. Nevertheless, the trends of pore formation times for different values of electric fields and oxidation degrees shown in Fig. 7.14 qualitatively illustrate the synergistic effect of plasma oxidation and electric fields on the permeability of the cell membrane.
7.3.3.4 Stabilizing Effect of Cholesterol in the PLB: Possible Explanation for the Selectivity of Plasma for Cancer Treatment
It is known that the cholesterol-to-phospholipid ratio in the cell membrane of some cancer cells (e.g., leukemic cells) is lower than in their normal counterparts (such as lymphocytes) [102]. Therefore, it is interesting to study the effect of the presence of cholesterol in the cell membrane, in varying concentrations, on the FEPs of ROS (both hydrophilic and hydrophobic), to reveal whether this can provide an explanation for the selectivity of plasma towards cancer cells vs. normal cells [103].
In Fig. 7.15 (upper part) the FEP of H2O2 is plotted for various cholesterol concentrations. Higher cholesterol concentrations obviously enhance the free energy barrier height and width and also result in local free energy minima in the center of the PLB, which will make the penetration of H2O2 towards the intracellular environment more difficult, even if it would succeed to penetrate into the PLB center. Other hydrophilic ROS, such as OH and HO2, showed the same behavior [103].
FEPs of H2O2 (upper part) and O2 (lower part) across the PLB, for various cholesterol concentrations in the cell membrane. Adopted from [103] with permission
Figure 7.15 (lower part) illustrates the FEP of O2, which exhibits a minimum in the center (cf. also Fig. 7.12). However, upon increasing cholesterol concentrations, some extra free energy barriers are formed at about 1 nm from the center, which is due to the presence of the bulky sterol rings. These extra barriers will limit the probability of lipid (per)oxidation of the lipid tails, and hence pore formation. As some cancer cells have less cholesterol in their cell membrane than normal cells, this might explain why RONS can more easily reach the cell interior of cancer cells, and hence, the selective action of plasma treatment.
As indicated above, the FEPs of the hydrophilic RO(N)S are too high for permeation, even without the presence of cholesterol, indicating the need for pore formation to allow these hydrophilic RO(N)S to enter the cell. Hence, it is also interesting to study the effect of cholesterol on pore formation. It was found that cholesterol fractions above 15% do not give rise to increasing cell membrane fluidity upon oxidation, and even for 100% oxidation degree, no pore formation was observed [91]. Indeed, Fig. 7.16 illustrates the water density in the center of the PLB in case of 100% lipid oxidation, as a function of the cholesterol fraction in the PLB. A cholesterol fraction up to 11% yields a significant water density, resulting from pore formation, while for higher cholesterol fractions, the water density drops to negligible values, indicating that no pores are formed. Hence, RONS should be able to penetrate more easily through the membrane of cells with lower cholesterol fraction, such as cancer cells, resulting in oxidative stress inside the cell, possibly inducing apoptosis of cancer cells, while this effect might not happen in normal cells, because the cholesterol in their cell membrane inhibits pore formation. Hence, this might be one of the explanations of the selectivity of plasma towards cancer cells.
Calculated average water density in the center of the PLB, for model systems with 100% oxidation, as a function of cholesterol concentration in the PLB, indicating that pore formation occurs more easily in cell membranes containing less cholesterol, which is typical for cancer cells. This might be one of the explanations of the selectivity of plasma treatment for cancer cells vs normal cells. Adopted from [91]—Published by The Royal Society of Chemistry
7.3.3.5 Permeation of RONS Through Aquaporins (AQPs) Vs PLB: Possible Explanation for the Selectivity of Plasma for Cancer Treatment
Another possible explanation for the selectivity of plasma towards cancer cells vs. normal cells is the higher expression of AQPs in the cell membrane of cancer cells. Indeed AQPs are known as H2O2 channels [104,105,106,107]. It is thus interesting to compare the FEPs and the diffusion rate profiles of H2O2 across AQP and the PLB, as well as the resulting permeability coefficient, which can all be obtained from MD simulations [108].
As illustrated in Fig. 7.17, the free energy barrier for H2O2 transport through AQP1 (which is one model system of AQP) is ca. three times lower than through the PLB, and the resulting permeability coefficient was found to be more than two orders higher through AQP than through the PLB, i.e., 2.57 cm/s vs. 6.62 × 10−3 cm/s. We can thus conclude that AQP creates a more favorable path for H2O2 permeation, which might also explain the selectivity of plasma towards cancer cells.
In [109] the permeation of other RONS, both hydrophilic (OH) and hydrophobic (NO2 and NO) through AQP was studied, and it was found that these RONS can all penetrate across the pores of AQP, with a lower permeation free energy barrier for OH and NO than for H2O2 and NO2, suggesting that these radicals may have easier access to the pore interior and interact with the amino acid residues of AQP. Furthermore, oxidation of both the phospholipids and AQP (i.e., sulfenylation of Cys191) seemed to slightly enhance the free energy barrier for H2O2 and NO2 permeation, while for OH and NO no clear effect was observed [109].
7.3.4 Interaction of RONS with DNA: Single and Double Strand Breaks
Besides studying the behavior of the PLB, the interaction of RONS with DNA might also be important in the context of cancer treatment. This has been investigated with classical reactive MD simulations [76]. As mentioned in Sect. 7.2.2, this method is less time-consuming than DFT and DFTB, so it can handle somewhat larger model systems, although the latter are still quite limited (order of a few 10,000 atoms). Hence, DNA as a whole cannot be simulated, because the structure is far too large. However, it is possible to select some representative fragments, as a model system. This is illustrated in Fig. 7.18, for a small part of a DNA double helix, containing 12 base pairs, surrounded by water molecules.
Part of a DNA double helix, consisting of 12-base pairs surrounded by water molecules, used as model system for the interaction with OH radicals in reactive MD simulations. Adopted from [76] with permission
When studying the interaction of OH radicals with this DNA model system, two main types of reactions were observed, i.e., (i) H-abstraction from a (primary or secondary) amine in the nucleic bases and (ii) OH-addition on the purine ring of the nucleic bases (see details in [76]).
The H-abstractions result in a radical that may give rise to some more (intermolecular) H-abstractions, which might affect the H-bridge structure between both DNA strands, possibly leading to single strand breaks (SSBs). When combined with a second SSB at the opposite strand in close vicinity, this might result in a double strand break (DSB), cleaving the DNA double helix, which might activate pro-apoptotic factors. It is indeed known from literature [110] that when both strands are broken within 20–57 base pairs by SSBs, DSB will eventually occur when enough H-bridges between both helices are broken. This could, however, not be observed in the MD simulations, because they are limited to a certain time scale, due to the high computational cost.
The OH-addition on the purine ring of the nucleic bases, more specifically at the C-8′ position of dAMP and dGMP, yields 8-hydroxy-purine adduct radicals (8-OH-Ade• or 8-OH-Gua•), as illustrated in Fig. 7.19 for dGMP. This reaction is the first step towards 8-oxo-guanine (8-O-Gua) and 2,6-diamino-4-hydroxy-5-formamidopyrimidine (FapydG) formation [111], which are markers for oxidative stress in cells [110,111,112,113]. This may affect biochemical pathways within the affected cell, e.g., introduction of DNA mutations or inhibition of gene expressions, possibly leading to apoptosis. Again, however, because of the high computational cost of the MD simulations, the simulated time was not long enough to observe these further reactions.
7.3.5 Effect of (Plasma) Oxidation on Proteins
As third type of biomolecules, proteins also play a crucial role in the context of cancer. In [114] non-reactive MD and molecular docking simulations, as well as binding free energy calculations, were performed to investigate the structural conformation and binding affinity of human epidermal growth factor (hEGF) to its receptor (hEGFR) under oxidative stress. hEGF is one of the important signaling proteins playing a role in both wound healing and cancer treatment. The simulations showed that a low amount of oxidation has minor effect on the binding affinity of hEGF with its receptor, while a higher level of oxidation causes less interaction with its receptor. This can be deduced from Fig. 7.20, illustrating the alignments of the oxidized structures with the native hEGF. Indeed, the structure with mild oxidation has a better alignment with the native hEGF, whereas the structures with more enhanced oxidation show significant deviations. Furthermore, the helical structures also start to form at random sites, which is more visible in the strongest oxidized structure. Thus, we can conclude that a higher level of oxidation may lead to completely new conformations in the hEGF protein (see Fig. 7.20).
Alignment of the oxidized hEGF structures (red), at mild (a), intermediate (b), and strong (c) oxidation, with native hEGF (cyan), illustrating more conformational changes upon higher oxidation of hEGF, compared to the native hEGF. Adopted from [114] with permission
Molecular docking and binding free energy simulations furthermore predicted that these structural modifications, and corresponding conformational changes, affect the binding free energies of hEGF with its receptor (hEGFR) [114]. Indeed, the binding free energy of native hEGF with its receptor was calculated to be −105.15 kcal/mol, while the binding free energies for the oxidized hEGF structures were found to be −71.21, −63.85, and −49.49 kcal/mol, for the mildly, intermediate, and strongly oxidized structures, respectively, thus indicating that higher oxidation causes a lower interaction.
The fact that mild oxidation does not significantly affect the binding affinity of hEGF with its receptor means that most likely, it does not strongly influence the signaling pathways in a cell, and thereby the cell proliferation, which might explain why short plasma treatment times are beneficial for chronic wound healing. On the other hand, the reduced alignment and interaction upon a higher level of oxidation of hEGF may cause a disturbance of the cell signaling pathways, ultimately leading to disruption of the cell proliferation. The latter might explain why longer plasma treatment times result in inhibiting cancer cell proliferation and even cancer cell death.
Although not in the context of plasma for cancer treatment, some other studies focused on the effect of oxidation on proteins by means of computer simulations. For instance, in [115] classical MD simulations were carried out to study the structure and dynamics of the oxidized (i.e., carbonylated) headpiece domain of villin. The modification of certain residues in this protein (i.e., seven lysine, arginine, and proline) resulted in a destabilization and unfolding of the protein structure, especially at high levels of carbonylation. This was attributed to the disruption of specific stabilizing elements, i.e., proline kinks and salt bridges, and a change of surface hydrophobicity [115]. The same group also examined the effect of oxidation on the stability and dimerization of superoxide dismutase 1 (SOD1), a key antioxidant enzyme in human cells [116]. The MD simulation results revealed that the oxidative modification of residues on the dimer interface (i.e., carbonylation of lysine, arginine, proline, and threonine, as well as cysteine oxidation) may lead to significant destabilization in the SOD1 homodimer. Moreover, oxidation can also decrease the stability of SOD1 monomer, resulting in a partial local unfolding and subsequent increase of the aggregation propensity [116]. Finally, MD simulations were also applied to investigate the role of post-translational modifications (PTMs), namely side chain hydroxylations, C- and N-methylations, on the stability of β6.3-helical structure of the polytheonamide B (pTB) peptide [117], by reverting the modified residues to their precursors. It was shown that the reversion of the hydroxylations and C-methylations represents insignificant or even no effect on the structure. On the other hand, the reversion of the N-methylations leads to the complete loss of the β-helical conformation [117].
In general, the effects of PTMs (e.g., methylation, phosphorylation, glycosylation, and acetylation) on the protein structure have been widely studied. However, it is also important to examine the effect of oxidation (and nitrosylation induced by RNS) on the protein stability and its binding affinity, especially in the context of plasma for cancer treatment, which is till now only scarcely investigated. Indeed, such studies would help to unravel the molecular level mechanisms of cancer cell death induced by plasma, as well as by other cancer therapies based on oxidation.
7.4 Conclusion and Future Challenges
This chapter gave an overview of the different types of modeling approaches that can be applied to study plasma for cancer treatment. This includes both macro-scale models, i.e., 0D chemical kinetics and 1D/2D fluid dynamics models of the most common plasma sources and their interaction with liquids, and atomic/molecular scale models, to study the interaction of plasma-produced RONS with biomolecules important in the context of plasma for cancer treatment, such as the phospholipid bilayer (PLB), DNA, and proteins, as well as the effect of these interactions (e.g., oxidation) on structural changes in the biomolecules.
Subsequently, typical calculation results of these models were presented, to illustrate the type of information that can be obtained from the different models.
Several different 0D chemical kinetics models and 1D/2D fluid dynamics models have been developed for different types of plasma sources and their interaction with liquids. They typically provide information on the important RONS present inside the plasma, the effluent, and/or the liquid. 0D models focus on a detailed chemistry, as they can handle a large number of species and chemical reactions with limited computational cost, while 1D/2D (or in principle 3D) fluid dynamics simulations typically consider a more limited chemistry (due to calculation time), but can be used to model specific plasma devices, and provide spatial information, e.g., gas and liquid flow profiles, species concentration profiles in the gas and liquid phase (including transport of RONS from the interface to the bulk), and chemical pathways for the various species in gas and liquid phase.
Furthermore, we showed several calculation results of the atomic/molecular scale simulations, for instance, to gain more insight in the permeability of various RONS across both native and oxidized PLBs, as model systems for the cell membrane, as well as in the synergistic effect of plasma oxidation and electric fields on pore formation, the effect of cholesterol present in the cell membrane on the permeability and on the chance for pore formation, and the different permeability of H2O2 (as representative RONS) across AQP channels vs. the PLB, in order to contribute to a better understanding of the selectivity of plasma for cancer treatment.
Hydrophobic RONS are shown to penetrate quite easily across both native and oxidized PLBs, while hydrophilic ROS experience too high energy barriers. Although the latter are reduced upon oxidation of the PLB, they are still too high, indicating the need for specific protein channels (e.g., AQPs) or pores created by plasma oxidation and/or electric fields, to allow their penetration into the cytoplasm, eventually to cause oxidative damage. In addition, the calculations reveal that the lower cholesterol fraction and the higher AQP expression in the cell membrane of various types of cancer cells might explain the selectivity of plasma towards cancer cells vs. normal cells.
Although these simulations give more insight in the formation and reactivity/stability of RONS inside plasma and in plasma-treated liquids, and how these RONS can penetrate through the cell membrane, either by passive transport (for hydrophobic RONS) or through pores or AQP channels (for hydrophilic RONS), more research will be needed to elucidate all the underlying mechanisms of plasma for cancer treatment. For instance, the simulations for RONS penetration through cell membranes are only applied to PLB model systems, while in reality the cell membrane is more complex and contains more proteins besides AQP, such as antiporters, catalase, etc., which can also be important in the context of plasma for cancer treatment. In addition, it would be interesting to study the cell membrane permeability of other RONS, e.g., hydrophilic RNS, like HNO2, HNO3/NO3 −, and ONOOH. They were not investigated up to now, because no accurate force fields are available yet for these simulations. This obviously shows the need to develop such force fields, to obtain a more comprehensive picture of the behavior of all possible RONS in the cell membrane. Furthermore, the various RONS might create a myriad of different lipid oxidation (and nitration) products in the cell membrane, so more knowledge is crucial on these different products formed, and how they affect the biophysical properties of the cell membrane, and thus its function. Up to now, simulations of oxidized membranes were based on rather simplified descriptions of membrane composition, while lipid peroxidation might lead to liquid ordered–liquid disordered phase separation in membranes [118], and this might favor pore formation. Thus, it would be interesting to account for phase-separated membranes.
We also showed examples of the interaction of RONS with DNA, as well as of the effect of plasma-induced oxidation on the structural conformation of proteins, more specifically for hEGF, playing a role in both wound healing and cancer therapy. Of course, many more proteins play a role in cancer therapy and could also be the subject of future investigations.
Specifically, it would be interesting to investigate the interaction between immune cell proteins and cancer cell proteins, under native conditions and oxidative stress (as could be induced by plasma therapy), which would be of great interest for plasma cancer immunotherapy. Protein–protein interactions have indeed been computationally studied in many different fields (e.g., [119, 120]). In particular, some MD simulations have already been reported for the molecular level functions of immune checkpoint proteins. For instance, Du et al. investigated the high affinity of PD-1 mutant by MD simulations, which helps to design engineered PD-1 mutants to modulate the PD-1/PD-L1 interaction [121]. Likewise, the binding of major histocompatibility complex (MHC) with different peptides and their receptors has also been studied by MD simulations [122]. In general, the effect of mutations in proteins and drugs that block the binding of immune checkpoint proteins to receptors has been studied for PD-1 and PD-L1, CD47 and SIRPα, and CD86 and CTLA-4 [123, 124]. However, to the best of our knowledge, the interaction between immune cell and cancer cell proteins under oxidative stress, as induced by plasma, has not yet been investigated by computer simulations, showing the need for such type of modeling, for a better insight in plasma for cancer immunotherapy.
Besides the above atomic/molecular level simulations, it would also be very interesting to study the cell signaling pathways, which might also explain the selectivity of plasma for cancer therapy [125]. Indeed, experiments showed that two apoptosis-inducing signaling pathways that are inhibited by catalase in the extracellular compartment can be reactivated when catalase in the cell membrane of cancer cells is inactivated. As plasma might be able to inactivate catalase, it might induce cancer cell death by reactivation of these signaling pathways. It would be interesting to obtain a better understanding of this apoptosis-inducing mechanism and the role of catalase inactivation. Such mechanisms can of course not be simulated by molecular level simulations, but it requires reaction kinetics modeling, including also spatial dynamics. Such modeling is very challenging and would need to be built up step-by-step, but it would be very rewarding to gain further insight in the underlying mechanisms of the selective action of plasma towards cancer cells.
Finally, to validate the model predictions, especially those at the molecular level, dedicated experiments should be designed, under very controlled conditions, generating, for instance, only a beam of OH radicals, instead of a complex mixture of RONS and other plasma effects, and for well-defined model systems of biomolecules, gradually mimicking the more complex tissues. Such experiments are set up in various labs, for instance, for studying the separate and synergistic effects of plasma-generated radicals and UV/VUV photons at the cellular and molecular level for various kinds of biomolecules, or experiments with simple model systems for the cell membrane, based on synthetic phospholipid membrane vesicles or liposomal model membranes (e.g., [126,127,128,129,130,131,132,133,134,135,136,137]). We believe that the combination of such experiments and modeling is crucial for gaining a deeper understanding of the underlying mechanisms of plasma for cancer treatment.
References
J. Van der Paal, Generation, transport and molecular interactions of reactive species in plasma medicine, PhD-dissertation, University of Antwerp, 2019
W. Van Gaens, A. Bogaerts, Kinetic modelling for an atmospheric pressure argon plasma jet in humid air. J. Phys. D Appl. Phys. 46, 275201 (2013)
P. Heirman, W. Van Boxem, A. Bogaerts, Reactivity and stability of plasma-generated oxygen and nitrogen species in buffered water solution: a computational study. Phys. Chem. Chem. Phys. 21, 12881–12894 (2019)
G.V. Naidis, Modelling of streamer propagation in atmospheric-pressure helium plasma jets. J. Phys. D Appl. Phys. 43, 402001 (2010)
G.V. Naidis, Modelling of plasma bullet propagation along a helium jet in ambient air. J. Phys. D Appl. Phys. 44, 215203 (2011)
J.P. Boeuf, L.L. Yang, L.C. Pitchford, Dynamics of a guided streamer (‘plasma bullet’) in a helium jet in air at atmospheric pressure. J. Phys. D Appl. Phys. 46, 015201 (2013)
D. Breden, K. Miki, L.L. Raja, Self-consistent two-dimensional modeling of cold atmospheric-pressure plasma jets/bullets. Plasma Sources Sci. Technol. 21, 034011 (2012)
M. Yousfi, O. Eichwald, N. Merbahi, N. Jomaa, Analysis of ionization wave dynamics in low-temperature plasma jets from fluid modeling supported by experimental investigations. Plasma Sources Sci. Technol. 21, 045003 (2012)
X.Y. Liu, X.K. Pei, X.P. Lu, D.W. Liu, Numerical and experimental study on a pulsed-dc plasma jet. Plasma Sources Sci. Technol. 23, 035007 (2014)
Z. Xiong, E. Robert, V. Sarron, J.M. Pouvesle, M.J. Kushner, Atmospheric-pressure plasma transfer across dielectric channels and tubes. J. Phys. D Appl. Phys. 46, 155203 (2013)
Z. Xiong, M.J. Kushner, Atmospheric pressure ionization waves propagating through a flexible high aspect ratio capillary channel and impinging upon a target. Plasma Sources Sci. Technol. 21, 034001 (2012)
Z. Xiong, E. Robert, V. Sarron, J.M. Pouvesle, M.J. Kushner, Dynamics of ionization wave splitting and merging of atmospheric-pressure plasmas in branched dielectric tubes and channels. J. Phys. D Appl. Phys. 45, 275201 (2012)
N.Y. Babaeva, M.J. Kushner, Interaction of multiple atmospheric-pressure micro-plasma jets in small arrays: He/O2 into humid air. Plasma Sources Sci. Technol. 23, 015007 (2014)
G.V. Naidis, On the ring-shaped structure of helium plasma jets. IEEE Trans. Plasma Sci. 43, 733–736 (2015)
A. Bourdon, T. Darny, F. Pechereau, et al., Numerical and experimental study of the dynamics of a μs helium plasma gun discharge with various amounts of N2 admixture. Plasma Sources Sci. Technol. 25, 035002 (2016)
C. Lazarou, C. Anastassiou, C. Topala, et al., Numerical simulation of capillary helium and helium−oxygen atmospheric pressure plasma jets: propagation dynamics and interaction with dielectric. Plasma Sources Sci. Technol. 27, 105007 (2018)
P. Viergas, F. Péchereau, A. Bourdon, Numerical study on the time evolutions of the electric field in helium plasma jets with positive and negative polarities. Plasma Sources Sci. Technol. 27, 025007 (2018)
A.M. Lietz, M.J. Kushner, Molecular admixtures and impurities in atmospheric pressure plasma jets. J. Appl. Phys. 124, 153303 (2018)
L. Chang, L. Nie, Y. Xian, X. Lu, The effect of seed electrons on the repeatability of atmospheric pressure plasma plume propagation. II. Modeling. Phys. Plasmas 23, 123513 (2016)
Y. Sakiyama, D.B. Graves, H.W. Chang, T. Shimizu, G.E. Morfill, Plasma chemistry model of surface microdischarge in humid air and dynamics of reactive neutral species. J. Phys. D Appl. Phys. 45, 425201 (2012)
N.Y. Babaeva, M.J. Kushner, Reactive fluxes delivered by dielectric barrier discharge filaments to slightly wounded skin. J. Phys. D Appl. Phys. 46, 025401 (2013)
H.W. Lee, G.Y. Park, Y.S. Seo, Y.H. Im, S.B. Shim, H.J. Lee, Modelling of atmospheric pressure plasmas for biomedical applications. J. Phys. D Appl. Phys. 44, 053001 (2011)
N.Y. Babaeva, N. Ning, D.B. Graves, M.J. Kushner, Ion activation energy delivered to wounds by atmospheric pressure dielectric-barrier discharges: sputtering of lipid-like surfaces. J. Phys. D Appl. Phys. 45, 115203 (2012)
N.Y. Babaeva, W. Tian, M.J. Kushner, The interaction between plasma filaments in dielectric barrier discharges and liquid covered wounds: electric fields delivered to model platelets and cells. J. Phys. D Appl. Phys. 47, 235201 (2014)
G.V. Naidis, Modelling of OH production in cold atmospheric-pressure He–H2O plasma jets. Plasma Sources Sci. Technol. 22, 035015 (2013)
D.X. Liu, P. Bruggeman, F. Iza, M.Z. Rong, M.G. Kong, Global model of low-temperature atmospheric-pressure He + H2O plasmas. Plasma Sources Sci. Technol. 19, 025018 (2010)
J. Waskoenig, K. Niemi, N. Knake, L.M. Graham, S. Reuter, V. Schulz-vonder Gathen, T. Gans, Atomic oxygen formation in a radio-frequency driven micro-atmospheric pressure plasma jet. Plasma Sources Sci. Technol. 19, 045018 (2010)
K. McKay, D.X. Liu, M.Z. Rong, F. Iza, M.G. Kong, Generation and loss of reactive oxygen species in low-temperature atmospheric-pressure RF He + O2 + H2O plasmas. J. Phys. D Appl. Phys. 45, 172001 (2012)
T. Murakami, K. Niemi, T. Gans, D. O’Connell, W.G. Graham, Afterglow chemistry of atmospheric-pressure helium–oxygen plasmas with humid air impurity. Plasma Sources Sci. Technol. 23, 025005 (2014)
T. Murakami, K. Niemi, T. Gans, D. O’Connell, W.G. Graham, Interacting kinetics of neutral and ionic species in an atmospheric-pressure helium–oxygen plasma with humid air impurities. Plasma Sources Sci. Technol. 22, 045010 (2013)
T. Murakami, K. Niemi, T. Gans, D. O’Connell, W.G. Graham, Chemical kinetics and reactive species in atmospheric pressure helium–oxygen plasmas with humid-air impurities. Plasma Sources Sci. Technol. 22, 015003 (2013)
W.J.M. Brok, M.D. Bowden, J. van Dijk, J.J.A.M. van der Mullen, G.M.W. Kroesen, Numerical description of discharge characteristics of the plasma needle. J. Appl. Phys. 98, 013302 (2005)
Y. Sakiyama, D.B. Graves, Finite element analysis of an atmospheric pressure RF-excited plasma needle. J. Phys. D Appl. Phys. 39, 3451 (2006)
Y. Sakiyama, D.B. Graves, Corona-glow transition in the atmospheric pressure RF-excited plasma needle. J. Phys. D Appl. Phys. 39, 3644 (2006)
Y. Sakiyama, D.B. Graves, Non-thermal atmospheric RF plasma in one-dimensional spherical coordinates. IEEE Trans. Plasma Sci. 35, 1279 (2007)
Y. Sakiyama, D.B. Graves, Nonthermal atmospheric rf plasma in one-dimensional spherical coordinates: asymmetric sheath structure and the discharge mechanism. J Appl. Phys. 101, 073306 (2007)
Y. Sakiyama, D.B. Graves, Neutral gas flow and ring-shaped emission profile in non-thermal RF-excited plasma needle discharge at atmospheric pressure. Plasma Sources Sci. Technol. 18, 025022 (2009)
W. Van Gaens, A. Bogaerts, Reaction pathways of biomedically active species in an Ar plasma jet. Plasma Sources Sci. Technol. 23, 035015 (2014)
S. Zhang, W. Van Gaens, B. Van Gessel, S. Hofmann, E. van Veldhuizen, A. Bogaerts, P. Bruggeman, Spatially resolved ozone densities and gas temperatures in a time modulated RF driven atmospheric pressure plasma jet: an analysis of the production and destruction mechanisms. J Phys. D Appl. Phys. 46, 205202 (2013)
W. Van Gaens, P.J. Bruggeman, A. Bogaerts, Numerical analysis of the NO and O generation mechanism in a needle-type plasma jet. New J. Phys. 16, 063054 (2014)
W. Van Gaens, S. Iséni, A. Schmidt-Bleker, K.-D. Weltmann, S. Reuter, A. Bogaerts, Numerical analysis of the effect of nitrogen and oxygen admixtures on the chemistry of an argon plasma jet operating at atmospheric pressure. New J. Phys. 17, 033003 (2015)
A. Schmidt-Bleker, J. Winter, S. Iseni, M. Dünnbier, K.D. Weltmann, S. Reuter, Reactive species output of a plasma jet with a shielding gas device - combination of FTIR absorption spectroscopy and gas phase modelling. J. Phys. D Appl. Phys. 47, 145201 (2014)
A. Schmidt-Bleker, J. Winter, A. Bösel, S. Reuter, K.-D. Weltmann, On the plasma chemistry of a cold atmospheric argon plasma jet with shielding gas device. Plasma Sources Sci. Technol. 25, 015005 (2016)
Y. Gorbanev, C.C.W. Verlackt, S. Tinck, E. Tuenter, K. Foubert, P. Cos, A. Bogaerts, Combining experimental and modelling approaches to study the sources of reactive species induced in water by the COST RF plasma jet. Phys. Chem. Chem. Phys. 20, 2797–2808 (2018)
Y. Gorbanev, J. Van der Paal, W. Van Boxem, S. Dewilde, A. Bogaerts, Reaction of chloride anion with atomic oxygen in aqueous solutions: can cold plasma help in chemistry research? Phys. Chem. Chem. Phys. 21, 4117–4121 (2019)
S. Schröter, R.A. Gibson, M.J. Kushner, T. Gans, D. O’Connell, Numerical study of the influence of surface reaction probabilities on reactive species in an rf atmospheric pressure plasma containing humidity. Plasma Phys. Control. Fusion 60, 014035 (2018)
B.R. Locke, S.M. Thagard, Analysis and review of chemical reactions and transport processes in pulsed electrical discharge plasma formed directly in liquid water. Plasma Chem. Plasma Process. 32, 875 (2012)
C.A.J. van Gils, S. Hofmann, B.K.H.L. Boekema, R. Brandenburg, P.J. Bruggeman, Mechanisms of bacterial inactivation in the liquid phase induced by a remote RF cold atmospheric pressure plasma jet. J Phys. D. Appl. Phys. 46, 175203 (2013)
P. Lukes, E. Dolezalova, I. Sisrova, M. Clupek, Aqueous-phase chemistry and bactericidal effects from an air discharge plasma in contact with water: evidence for the formation of peroxynitrite through a pseudo-second-order post-discharge reaction of H2O2 and HNO2. Plasma Sources Sci. Technol. 23, 015019 (2014)
S. Hamaguchi, K. Ikuse, T. Kanazawa, Generation of free radicals in liquid by atmospheric-pressure plasmas and its application to biology and medicine. JPS Conf. Proc. 1, 015055 (2014)
C. Chen, D.X. Liu, Z.C. Liu, A.J. Yang, H.L. Chen, G. Shama, M.G. Kong, A model of plasma-biofilm and plasma-tissue interactions at ambient pressure. Plasma Chem. Plasma Process. 34, 403 (2014)
D.X. Liu, Z.C. Liu, C. Chen, A.J. Yang, D. Li, M.Z. Rong, H.L. Chen, M.G. Kong, Aqueous reactive species induced by a surface air discharge: heterogeneous mass transfer and liquid chemistry pathways. Sci. Rep. 6, 23737 (2016)
W. Tian, M.J. Kushner, Influence of excitation pulse duration of dielectric barrier discharges on biomedical applications. J. Phys. D Appl. Phys. 47, 165201 (2014)
W. Tian, M.J. Kushner, Long-term effects of multiply pulsed dielectric barrier discharges in air on thin water layers over tissue: stationary and random streamers. J. Phys. D Appl. Phys. 48, 494002 (2015)
A.M. Lietz, M.J. Kushner, Air plasma treatment of liquid covered tissue: long timescale chemistry. J. Phys. D Appl. Phys. 49, 425204 (2016)
A.M. Lietz, M.J. Kushner, Corrigendum: air plasma treatment of liquid covered tissue: long timescale chemistry (2016 J. Phys. D: Appl. Phys. 49 425204). J. Phys. D Appl. Phys. 50, 119501 (2017)
W. Van Boxem, J. Van der Paal, Y. Gorbanev, S. Vanuytsel, E. Smits, S. Dewilde, A. Bogaerts, Anti-cancer capacity of plasma-treated PBS: effect of chemical composition on cancer cell cytotoxicity. Sci. Rep. 7, 16478 (2017)
J. Du, Z. Liu, C. Bai, L. Li, Y. Zhao, L. Wang, J. Pan, Concentration distributions and reaction pathways of species in the mass transfer process from atmospheric pressure plasma jet to water. Eur. Phys. J. D 72, 179 (2018)
A. Lindsay, C. Anderson, E. Slikboer, S. Shannon, D. Graves, Momentum, heat, and neutral mass transport in convective atmospheric pressure plasma-liquid systems and implications for aqueous targets. J. Phys. D Appl. Phys. 48, 424007 (2015)
S.A. Norberg, G.M. Parsey, A.M. Lietz, E. Johnsen, M.J. Kushner, Atmospheric pressure plasma jets onto a reactive water layer over tissue: pulse repetition rate as a control mechanism. J. Phys. D Appl. Phys. 52, 015201 (2019)
C.C.W. Verlackt, W. Van Boxem, A. Bogaerts, Transport and accumulation of plasma generated species in aqueous solution. Phys. Chem. Chem. Phys. 20, 6845–6859 (2018)
A. Bogaerts, N. Khosravian, J. Van der Paal, C.C.W. Verlackt, M. Yusupov, B. Kamaraj, E.C. Neyts, Multi-level molecular modeling for plasma medicine. J. Phys. D Appl. Phys. 49, 054002 (2016)
M. Elstner, D. Porezag, G. Jungnickel, J. Elsner, M. Haugk, T. Frauenheim, S. Suhai, G. Seifert, Self-consistent-charge density-functional tight-binding method for simulations of complex materials properties. Phys. Rev. B 58, 7260–7268 (1998)
M. Yusupov, K. Wende, S. Kupsch, E.C. Neyts, S. Reuter, A. Bogaerts, Effect of head group and lipid tail oxidation in the cell membrane revealed through integrated simulations and experiments. Sci. Rep. 7, 5761 (2017)
N. Khosravian, B. Kamaraj, E.C. Neyts, A. Bogaerts, Structural modification of P-glycoprotein induced by OH radicals: insights from atomistic simulations. Sci. Rep. 6, 19466 (2016)
C.C.W. Verlackt, W. Van Boxem, D. Dewaele, F. Lemière, F. Sobott, J. Benedikt, E.C. Neyts, A. Bogaerts, Mechanisms of peptide oxidation by hydroxyl radicals: insight at the molecular scale. J. Phys. Chem. C 121, 5787–5799 (2017)
C.C.W. Verlackt, E.C. Neyts, A. Bogaerts, Atomic scale behavior of oxygen-based radicals in water. J. Phys. D Appl. Phys. 50, 11LT01 (2017)
D.W. Brenner, Empirical potential for hydrocarbons for use in simulating the chemical vapor deposition of diamond films. Phys. Rev. B 42, 9458–9471 (1990)
A.C.T. van Duin, S. Dasgupta, F. Lorant, W.A. Goddard, ReaxFF: a reactive force field for hydrocarbons. J. Phys. Chem. A 105, 9396–9409 (2001)
M. Yusupov, E.C. Neyts, U. Khalilov, R. Snoeckx, A.C.T. van Duin, A. Bogaerts, Atomic scale simulations of plasma species interacting with bacterial cell walls. New J. Phys. 14, 093043 (2012)
M. Yusupov, A. Bogaerts, S. Huygh, S. Snoeckx, A.C.T. van Duin, E.C. Neyts, Plasma-induced destruction of bacterial cell wall components: a reactive molecular dynamics simulation. J. Phys. Chem. C 117, 5993–5998 (2013)
M. Yusupov, E.C. Neyts, C.C. Verlackt, U. Khalilov, A.C.T. van Duin, A. Bogaerts, Inactivation of the endotoxic biomolecule lipid a by oxygen plasma species: a reactive molecular dynamics study. Plasma Process. Polym. 12, 162–171 (2015)
J. Van der Paal, S. Aernouts, A.C.T. van Duin, E.C. Neyts, A. Bogaerts, Interaction of O and OH radicals with a simple model system for lipids in the skin barrier: a reactive molecular dynamics simulation for plasma medicine. J. Phys. D Appl. Phys. 46, 395201 (2013)
J. Van der Paal, C.C. Verlackt, M. Yusupov, E.C. Neyts, A. Bogaerts, Structural modification of the skin barrier by OH radicals: a reactive molecular dynamics study for plasma medicine. J. Phys. D Appl. Phys. 48, 155202 (2015)
R.M. Abolfath, P.K. Biswas, R. Rajnarayanam, T. Brabec, R. Kodym, L. Papiez, Multiscale QM/MM molecular dynamics study on the first steps of guanine damage by free hydroxyl radicals in solution. J. Phys. Chem. A 116, 3940–3945 (2012)
C.C.W. Verlackt, E.C. Neyts, T. Jacob, D. Fantauzzi, M. Golkaram, Y.-K. Shin, A.C.T. van Duin, A. Bogaerts, Atomic-scale insight in the interactions between hydroxyl radicals and DNA in solution using the ReaxFF reactive force field. New J. Phys. 17, 103005 (2015)
M. Yusupov, E.C. Neyts, P. Simon, G. Bergiyorov, R. Snoeckx, A.C.T. van Duin, A. Bogaerts, Reactive molecular dynamics simulations of oxygen species in a liquid water layer of interest for plasma medicine. J. Phys. D Appl. Phys. 47, 025205 (2014)
N. Khosravian, A. Bogaerts, S. Huygh, M. Yusupov, E.C. Neyts, How do plasma-generated OH radicals react with biofilm components? Insights from atomic scale simulations. Biointerphases 10, 029501 (2015)
O. Berger, O. Edholm, F. Jähnig, Molecular dynamics simulations of a fluid bilayer of dipalmitoylphosphatidylcholine at full hydration, constant pressure, and constant temperature. Biophys. J. 72, 2002–2013 (1997)
W.D. Cornell, P. Cieplak, C.I. Bayly, I.R. Gould, K.M. Merz Jr., D.M. Ferguson, D.C. Spellmeyer, T. Fox, J.W. Caldwell, P.A. Kollman, A second generation force field for the simulation of proteins, nucleic acids, and organic molecules. J. Am. Chem. Soc. 117, 5179–5197 (1995)
W. Yu, X. He, K. Vanommeslaeghe, A.D. MacKerell Jr., Extension of the CHARMM general force field to sulfonyl-containing compounds and its utility in biomolecular simulations. J. Comput. Chem. 33, 2451 (2012)
W.F. van Gunsteren, H.J.C. Berendsen, Groningen Molecular Simulation (GROMOS) Library Manual (Biomos, Groningen, 1987), p. 221. http://www.gromos.net/gromos87/GROMOS87_manual.pdf
S.J. Marrink, H.J. Risselada, S. Yefimov, D.P. Tieleman, A.H. De Vries, The MARTINI force field: coarse grained model for biomolecular simulations. J. Phys. Chem. B 111, 7812–7824 (2007)
J. Åqvist, A. Warshel, Simulation of enzyme reactions using valence bond force fields and other hybrid quantum/classical approaches. Chem. Rev. 93, 2523–2544 (1993)
E.C. Neyts, M. Yusupov, C.C. Verlackt, A. Bogaerts, Computer simulations of plasma-biomolecule and plasma-tissue interactions for a better insight in plasma medicine. J. Phys. D Appl. Phys. 47, 293001 (2014)
J. Razzokov, M. Yusupov, R.M. Cordeiro, A. Bogaerts, Atomic scale understanding of the permeation of plasma species across native and oxidized membranes. J. Phys. D Appl. Phys. 51, 365203 (2018)
R.M. Cordeiro, Reactive oxygen species at phospholipid bilayers: distribution, mobility and permeation. Biochim. Biophys. Acta Biomembr. 1838, 438–444 (2014)
M.N. Möller, Q. Li, J.R. Lancaster, A. Denicola, Acceleration of nitric oxide autoxidation and nitrosation by membranes. IUBMB Life 59, 243–248 (2007)
W.K. Subczynski, M. Lomnicka, J.S. Hyde, Permeability of nitric oxide through lipid bilayer membranes. Free Radic. Res. 24, 343–349 (1996)
A. Reis, M.R.M. Domingues, F.M.L. Amado, A.J.V. Ferrer-Correia, P. Domingues, Separation of peroxidation products of diacyl-phosphatidylcholines by reversed-phase liquid chromatography-mass spectrometry. Biomed. Chromatogr. 19, 129–137 (2005)
J. Van der Paal, E.C. Neyts, C.C.W. Verlackt, A. Bogaerts, Effect of lipid peroxidation on membrane permeability of cancer and normal cells subjected to oxidative stress. Chem. Sci. 7, 489–498 (2016)
J. Wong-Ekkabut, Z. Xu, W. Triampo, I.-M. Tang, D.P. Tieleman, L. Monticelli, Effect of lipid peroxidation on the properties of lipid bilayers: a molecular dynamics study. Biophys. J. 93, 4225–4236 (2007)
L. Beranova, L. Cwiklik, P. Jurkiewicz, M. Hof, P. Jungwirth, Oxidation changes physical properties of phospholipid bilayers: fluorescence spectroscopy and molecular simulations. Langmuir 26, 6140–6144 (2010)
L. Cwiklik, P. Jungwirth, Massive oxidation of phospholipid membranes leads to pore creation and bilayer disintegration. Chem. Phys. Lett. 486, 99–103 (2010)
P.T. Vernier, M.J. Ziegler, Nanosecond field alignment of head group and water dipoles in electroporating phospholipid bilayers. J. Phys. Chem. B 111, 12993–12996 (2007)
M. Casciola, M. Tarek, A molecular insight into the electro-transfer of small molecules through electropores driven by electric fields. Biochim. Biophys. Acta. Biomembr. 1858, 2278–2289 (2016)
S.J. Marrink, A.H. de Vries, D.P. Tieleman, Lipids on the move: simulations of membrane pores, domains, stalks and curves. Biochim. Biophys. Acta Biomembr. 1788, 149–168 (2009)
A.M. Hirst, F.M. Frame, M. Arya, N.J. Maitland, D. O’Connell, Low temperature plasmas as emerging cancer therapeutics: the state of play and thoughts for the future. Tumor Biol. 37, 7021–7031 (2016)
E. Robert, T. Darny, S. Dozias, S. Iseni, J.-M. Pouvesle, New insights on the propagation of pulsed atmospheric plasma streams: from single jet to multi jet arrays. Phys. Plasmas 22, 122007 (2015)
A. Begum, M. Laroussi, M.R. Pervez, Atmospheric pressure He-air plasma jet: breakdown process and propagation phenomenon. AIP Adv. 3, 062117 (2013)
M. Yusupov, J. Van der Paal, E.C. Neyts, A. Bogaerts, Synergistic effect of electric field and lipid oxidation on the permeability of cell membranes. Biochim. Biophys. Acta Gen. 1861, 839–847 (2017)
M. Shinitzky, Membrane fluidity in malignancy adversative and recuperative. Biochim. Biophys. Acta Rev. Cancer 738, 251–261 (1984)
J. Van der Paal, C. Verheyen, E.C. Neyts, A. Bogaerts, Hampering effect of cholesterol on the permeation of reactive oxygen species through phospholipids bilayer: possible explanation for plasma cancer selectivity. Sci. Rep. 7, 39526 (2017)
P.S. Hole, J. Zabkiewicz, C. Munje, Z. Newton, L. Pearn, P. White, N. Marquez, R.K. Hills, A.K. Burnett, A. Tonks, R.L. Darley, Overproduction of NOX-derived ROS in AML promotes proliferation and is associated with defective oxidative stress signaling. Blood 122, 3322–3330 (2013)
M.C. Papadopoulos, S. Saadoun, Key roles of aquaporins in tumor biology. Biochim. Biophys. Acta Biomembr. 1848, 2576–2583 (2015)
R.M. Cordeiro, Molecular dynamics simulations of the transport of reactive oxygen species by mammalian and plant aquaporins. Biochim. Biophys. Acta Gen. 1850, 1786–1794 (2015)
D. Yan, A. Talbot, N. Nourmohammadi, J.H. Sherman, X. Cheng, M. Keidar, Toward understanding the selective anticancer capacity of cold atmospheric plasma - a model based on aquaporins (review). Biointerphases 10, 040801 (2015)
M. Yusupov, D. Yan, R.M. Cordeiro, A. Bogaerts, Atomic scale simulation of H2O2 permeation through aquaporin: toward the understanding of plasma-cancer treatment. J. Phys. D Appl. Phys. 51, 125401 (2018)
M. Yusupov, J. Razzokov, R.M. Cordeiro, A. Bogaerts, Transport of reactive oxygen and nitrogen species across aquaporin: a molecular level picture. Oxidative Med. Cell Longev. 2019, 2930504 (2019)
C. Shao, M. Saito, Z. Yu, Formation of single- and double-strand breaks of pBR322 plasmid irradiated in the presence of scavengers. Radiat. Environ. Biophys. 38, 105 (1999)
M. Dizdaroglu, P. Jaruga, Mechanisms of free radical-induced damage to DNA. Free Radic. Res. 46, 382 (2012)
J. Cadet, T. Douki, J.-L. Ravanat, P. Di Mascio, Sensitized formation of oxidatively generated damage to cellular DNA by UVA radiation. Photochem. Photobiol. Sci. 8, 903 (2009)
N.R. Jena, P.C. Mishra, Mechanisms of formation of 8-oxoguanine due to reactions of one and two OH• radicals and the H2O2 molecule with guanine: a quantum computational study. J. Phys. Chem. B 109, 14205 (2005)
M. Yusupov, J.-W. Lackmann, J. Razzokov, S. Kumar, K. Stapelmann, A. Bogaerts, Impact of plasma oxidation on structural features of human epidermal growth factor. Plasma Process. Polym. 15, e1800022 (2018)
D. Petrov, B. Zagrovic, Microscopic analysis of protein oxidative damage: effect of carbonylation on structure, dynamics, and aggregability of villin headpiece. J. Am. Chem. Soc. 133, 7016–7024 (2011)
D. Petrov, X. Daura, B. Zagrovic, Effect of oxidative damage on the stability and dimerization of superoxide dismutase 1. Biophys. J. 110, 1499–1509 (2016)
A. Renevey, R. Sereina, The importance of N-methylations for the stability of the β6.3-helical conformation of polytheonamide B. Eur. Biophys. J. 46, 363–374 (2017)
C.K. Haluska, M.S. Baptista, A.U. Fernades, A.P. Schroder, C.M. Marques, R. Itri, Photo-activated phase separation in giant vesicles made from different lipid mixture. Biochim. Biophys. Acta Biomembr. 1818, 666–672 (2012)
D.E. Scott et al., Small molecules, big targets: drug discovery faces the protein-protein interaction challenge. Nat. Rev. Drug Discov. 15, 533–550 (2016)
T. Kortemme et al., Computational redesign of protein-protein interaction specificity. Nat. Struct. Mol. Biol. 11, 371–379 (2004)
J. Du et al., The design of high affinity human PD-1 mutants by using molecular dynamics simulations (MD). Cell. Commun. Signal. 16, 25 (2018)
B. Knapp et al., Current status and future challenges in T-cell receptor/peptide/MHC molecular dynamics simulations. Brief. Bioinform. 16, 1035 (2015)
K. Weiskopf et al., Engineered SIRPα variants as immunotherapeutic adjuvants to anticancer antibodies. Science 341, 88–91 (2013)
J.Y. Lee et al., Structural basis of checkpoint blockade by monoclonal antibodies in cancer immunotherapy. Nat. Commun. 7, 13354 (2016)
G. Bauer, D.B. Graves, Mechanisms of selective antitumor action of cold atmospheric plasma-derived reactive oxygen and nitrogen species. Plasma Process. Polym. 13, 1157–1178 (2016)
J.-W. Lackmann, S. Schneider, E. Edengeiser, F. Jarzina, S. Brinckmann, E. Steinborn, M. Havenith, J. Benedikt, J.E. Bandow, Photons and particles emitted from cold atmospheric-pressure plasma inactivate bacteria and biomolecules independently and synergistically. J. R. Soc. Interface 10, 20130591 (2013)
T.-Y. Chung, N. Ning, J.W. Chu, D.B. Graves, E. Bartis, J. Seog, G.S. Oehrlein, Plasma deactivation of endotoxic biomolecules: vacuum ultraviolet photon and radical beam effects on Lipid A. Plasma Process. Polym. 10, 167–180 (2013)
E.A.J. Bartis, D.B. Graves, J. Seog, G.S. Oehrlein, Atmospheric pressure plasma treatment of lipopolysaccharide in a controlled environment. J. Phys. D Appl. Phys. 46, 312002 (2013)
E.A.J. Bartis, C. Barrett, T.Y. Chung, N. Ning, J.W. Chu, D.B. Graves, J. Seog, G.S. Oehrlein, Deactivation of lipopolysaccharide by Ar and H2 inductively coupled low-pressure plasma. J. Phys. D Appl. Phys. 47, 045202 (2014)
J.H. Park, N. Kumar, D.H. Park, M. Yusupov, E.C. Neyts, C.C.W. Verlackt, A. Bogaerts, M.H. Kang, H.S. Uhm, E.H. Choi, P. Attri, A comparative study for the inactivation of multidrug resistance bacteria using dielectric barrier discharge and nano-second pulsed plasma. Sci. Rep. 5, 13849 (2015)
M. Marschewski, J. Hirschberg, T. Omairi, O. Hofft, W. Viol, S. Emmert, W. Maus-Friedrichs, Electron spectroscopic analysis of the human lipid skin barrier: cold atmospheric plasma-induced changes in lipid composition. Exp. Dermatol. 21, 921–925 (2012)
E. Takai, T. Kitamura, J. Kuwabara, S. Ikawa, S. Yoshizawa, K. Shiraki, H. Kawasaki, R. Arakawa, K. Kitano, Chemical modification of amino acids by atmospheric-pressure cold plasma in aqueous solution. J. Phys. D Appl. Phys. 47, 285403 (2014)
G.S. Madugundu, J. Cadet, J.R. Wagner, Hydroxyl-radical-induced oxidation of 5-methylcytosine in isolated and cellular DNA. Nucleic Acids Res. 42, 7450–7460 (2014)
S.-H. Hong, E.J. Szili, A.T.A. Jenkins, R.D. Short, Ionized gas (plasma) delivery of reactive oxygen species (ROS) into artificial cells. J. Phys. D Appl. Phys. 47, 362001 (2014)
E.J. Szili, J.W. Bradley, R.D. Short, A ‘tissue model’ to study the plasma delivery of reactive oxygen species. J. Phys. D Appl. Phys. 47, 152002 (2014)
M.U. Hammer, E. Forbrig, S. Kupsch, K.-D. Weltmann, S. Reuter, Influence of plasma treatment on the structure and function of lipids. Plasma Med. 3, 97–114 (2013)
J. Van der Paal, S.-H. Hong, M. Yusupov, N. Gaur, J.-S. Oh, R.D. Short, E.J. Szili, A. Bogaerts, Cell membrane response to oxidative stress for anticancer therapies: an experimental and computational study. Phys. Chem. Chem. Phys. 21(35), 19327–19341 (2019)
Acknowledgements
We acknowledge the financial support from the Research Foundation—Flanders (FWO; grant numbers: 1200219 N and 11U5416N). The computational work was carried out using the Turing HPC infrastructure at the CalcUA core facility of the Universiteit Antwerpen (UA), a division of the Flemish Supercomputer Center VSC, funded by the Hercules Foundation, the Flemish Government (department EWI), and the UA. We are also very thankful to R. Cordeiro for the very interesting discussions.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Bogaerts, A., Van der Paal, J., Heirman, P., Razzokov, J., Yusupov, M. (2020). Plasma and Plasma–Cell Interaction Simulations. In: Keidar, M. (eds) Plasma Cancer Therapy. Springer Series on Atomic, Optical, and Plasma Physics, vol 115. Springer, Cham. https://doi.org/10.1007/978-3-030-49966-2_7
Download citation
DOI: https://doi.org/10.1007/978-3-030-49966-2_7
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-49965-5
Online ISBN: 978-3-030-49966-2
eBook Packages: Physics and AstronomyPhysics and Astronomy (R0)