Abstract
The RECQ family of DNA helicases is a conserved group of enzymes that plays an important role in maintaining genomic stability. Humans possess five RECQ helicase genes, and mutations in three of them – BLM, WRN, and RECQL4 – are associated with the genetic disorders Bloom syndrome, Werner syndrome, and Rothmund-Thomson syndrome (RTS), respectively. These syndromes share overlapping clinical features, and importantly they are all associated with an increased risk of cancer. Patients with RTS have the highest specific risk of developing osteosarcoma compared to all other cancer predisposition syndromes; therefore, RTS serves as a relevant model to study the pathogenesis and molecular genetics of osteosarcoma. The “tumor suppressor” function of the RECQ helicases continues to be an area of active investigation. This chapter will focus primarily on the known cellular functions of RECQL4 and how these may relate to tumorigenesis, as well as ongoing efforts to understand RECQL4’s functions in vivo using animal models. Understanding the RECQ pathways will provide insight into avenues for novel cancer therapies in the future.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- RECQ
- RECQL4
- DNA helicase
- Rothmund-Thomson syndrome
- RTS
- Bloom syndrome
- Werner syndrome
- Osteosarcoma
- Genomic instability
Introduction
The roles of the RECQ helicases in cancer and specifically the role of RECQL4 in osteosarcoma (OS) are areas of active investigation. While it is known that constitutional mutations in the RECQ genes predispose patients to developing cancer, the exact mechanisms of tumorigenesis remain to be fully explored. As basic science research continues to reveal the normal cellular functions of the RECQ helicases, application of this knowledge to OS pathogenesis will provide avenues for future investigation into targeted therapies for this disease. This chapter will primarily focus on what is currently known about the RECQL4 DNA helicase gene, which is mutated in the OS predisposition disorder Rothmund-Thomson syndrome (RTS).
RECQ Family of DNA Helicases and Cancer Predisposition
The RECQ DNA helicases are a family of proteins that are important in maintaining genomic integrity. DNA helicases are ubiquitous molecular motor proteins that harness the chemical free energy of ATP hydrolysis to catalyze the unwinding of duplex DNA and as such play important roles in nearly all aspects of nucleic acid metabolism, including replication, repair, recombination, and transcription [115]. The RECQ helicases belong to the SF2 superfamily of DNA helicases that unwind DNA in a 3′ ↑ 5′ direction in an ATP- and Mg2+-dependent fashion [5, 8]. As such, they contain a conserved region that includes the seven characteristic helicase motifs (I, Ia, II, III, IV, V, and VI) that define this family of helicases and that are important for coupling ATP hydrolysis to the separation of DNA strands. The first RECQ helicase was discovered in Escherichia coli (E. coli) in a screen for resistance to thymineless death [81]. Subsequently, RECQ proteins have been identified in multiple species. These evolutionarily conserved proteins are defined by their common central helicase motif, a highly conserved region of approximately 400 amino acids (Fig. 3.1) [8, 55]. The number of RECQ helicases increases from lower to higher organisms. Bacteria such as E. coli have one (RecQ), as do yeast (Sgs1 in Saccharomyces cerevisiae and Rqh1 in Schizosaccharomyces pombe), while Caenorhabditis elegans has two and Arabidopsis thalianas has seven RECQ helicases [58].
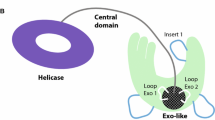
Structural features of RecQ helicases. The RecQ proteins have several structural domains that are conserved from bacteria through humans. All RecQ proteins have a core helicase domain. Most RecQ proteins also contain conserved helicase and RNAse D C-terminal (HRDC) and RecQ C-terminal (RQC) domains that are thought to mediate interactions with nucleic acid and other proteins, respectively. Many RecQ proteins have acidic regions that enable protein-protein interactions, and some of the RecQ proteins have nuclear localization sequences. WRN and FFA-1 protein are unique in that they also contain an exonuclease domain. Sgs1 and Blm are the first characterized members of this family of proteins containing a functional strand exchange domain in their N-terminus. The number of amino acids in each protein is indicated on the right. (Reprinted with permission from Bernstein et al. [8])
In humans, there are five RECQ helicases (Fig. 3.1). Three of these, WRN, BLM, and RECQL4, are associated with human diseases [79]. Mutations in the WRN gene [137] cause Werner syndrome [73], and mutations in the BLM gene [30] are responsible for Bloom syndrome [36]. Mutations in RECQL4 are associated with three overlapping disorders: RTS, RAPADILINO syndrome, and Baller-Gerold syndrome (BGS) [56, 101, 117]. Although RECQL and RECQL5 have not thus far been associated with any human genetic disorders, both have been linked to human tumorigenesis [23, 28, 127]. In one study, rare germ line truncating mutations in the RECQL gene were shown to be associated with an increased risk of breast cancer in two populations of high-risk patients [23]. A few small studies have demonstrated that specific single-nucleotide polymorphisms in RECQL5 are more common in OS patients [28, 139], and decreased expression of RECQL5 in OS tumors may be associated with disease progression [127].
All of the human RECQ disorders are cancer predisposition syndromes, but they have varying cancer profiles (Table 3.1). Patients with Werner syndrome display features of premature aging, such as diabetes, coronary artery disease, cataracts, and osteoporosis. They are susceptible primarily to thyroid cancer, melanoma, meningioma, soft tissue sarcomas, and OS. In a study of the spectrum of cancers in Werner syndrome patients, OS was found to comprise 8% of all neoplasms [62]. In contrast, patients with Bloom syndrome are susceptible to all types of cancers seen in the general population, but at a much higher frequency and at an earlier age. These include leukemia and lymphomas and epithelial cancers of the colon, breast, head and neck, and cervix, as well as OS, which accounted for 2% of the first 100 cases of cancers reported in the Bloom Registry [22, 37]. Among the RECQL4-associated disorders, patients with RTS have a very high and specific risk for OS, in addition to nonmelanoma skin cancers (squamous and basal cell carcinomas). In one clinical cohort study of 41 RTS patients, 30% had a diagnosis of OS [122]. Patients with RAPADILINO syndrome and RECQL4 mutations are also at risk for cancer, most commonly lymphomas as well as OS [102]. These patients share many of the same phenotypes as RTS patients, including small stature, limb deformities, radial ray defects, and absent patellae. Interestingly, these patients do not display poikiloderma, which is a defining feature of RTS. BGS is the least well-characterized of the RECQL4 disorders. These patients are characterized by craniosynostosis and radial ray defects, as well as poikiloderma in some patients. So far only a few cases have been described to have RECQL4 mutations, and cancer has only been described in one patient who developed a midline NK cell lymphoma [26]. Overall there have been over 60 RECQL4 mutations identified among these three disorders [116]. Exact genotype-phenotype correlations with respect to specific mutations and resultant phenotypes, including cancer, remain to be elucidated .
As a group, the RECQ helicases are considered “caretakers” of the genome and as such do not necessarily directly drive tumorigenesis but prevent genomic instability that results in accumulation of structural changes in oncogenes or tumor suppressors that could then lead to cancer [17]. This protection of genome stability is achieved through their various roles in DNA replication, repair, and telomere maintenance. It is also possible that the RECQ helicases could play a more direct role in affecting tumorigenesis. While the exact molecular mechanisms of tumor suppression have yet to be worked out fully, deficiency of the WRN, BLM, and RECQL4 proteins in humans clearly predisposes to the development of cancer.
Structure and Functions of the RECQL4 DNA Helicase
The role of RECQL4 in DNA replication has been extensively studied, and it appears that while RECQL4 may participate in many cellular functions, its primary role is in the initiation of DNA replication [46, 74, 94, 113, 126, 132, 133]. This is achieved primarily through its N-terminal domain (amino acids 1–370) which shares homology to the yeast replication factor Sld2 in S. cerevisiae and Drc1 in S. pombe [72, 74, 94], both of which are important for establishing replication forks during the initiation of DNA replication . After phosphorylation by cyclin-dependent kinases, Sld2 binds Dpb11, a key mediator of the formation of the active replicative helicase complex on replication origins and a crucial factor in the initiation of DNA replication [51, 108, 119]. In Xenopus , it has been shown that xRECQL4 belongs to the replication initiation complex and helps to promote loading of replication factors at the origins, after pre-replication complex formation [94]. The N-terminal amino acid region 1–596 of RECQL4 interacts directly with xCut5 (frog orthologue of Dpb11), which is responsible for recruiting DNA polymerases to the sites of replication [74]. RECQL4 has also been shown to interact with multiple DNA replication factors, such as MCM10, MCM2-7, CTF4, CDC45, GINS, and SLD5 which are essential for initiation of DNA replication [46, 47, 57, 132], as well as TopBP1, the vertebrate orthologue of Dpb11 [87]. The C-terminus of RECQL4 including the helicase domain also appears to play a role in replication under stressed conditions. Human pre-B lymphocyte cells with mutant RECQL4 lacking the C-terminus were shown to have replication defects only after ionizing radiation, perhaps by allowing replication forks to negotiate the radiation-damaged DNA templates [59]. Because RECQL4 is overexpressed in many types of sporadic cancers (see below), the effect of overexpression of RECQL4 on replication has also been studied. Although overexpression of RECQL4 alone did not affect replication, when RECQL4 was fused to a subunit of the origin recognition complex-ORC4 protein, overexpression of this fusion protein induced increased binding of RECQL4 to late replication origins in early S phase and recruitment of replication initiation factors [99]. As a result, early activation of replication was observed in genes with late replication origins, leading to elevated replication stress caused by replication-transcription conflicts [99]. Therefore, the binding of RECQL4 to replication origins needs to be tightly regulated to ensure a normal replication process. In addition to initiation of DNA replication, RECQL4 may also play a role in replication fork restart given its high affinity to Holliday junction substrates demonstrated by in vitro binding assays via N-terminal amino acid residues 320–400 [96].
RECQL4 has been shown to bind additional nucleic acid substrates in vitro, including guanine quadruplex (G4) structures [54]. G4 is a type of secondary structure formed in guanine-rich sequences and is found in replication origins, gene promoter regions, and telomeric DNA sequences [41]. BLM, WRN, and RECQL4 have all been shown to be important for telomere maintenance [20]. Both BLM and WRN helicases bind and unwind G4 DNA substrates [77], while RECQL4 only binds but has no detectable unwinding activity [54]. Gene expression analyses using fibroblasts from both Bloom and Werner syndrome patients showed that BLM and WRN regulate transcription through G4 DNA sequences [85, 109]. The biological function of RECQL4 at G4 sites needs further investigation given the importance of G4 sequences in normal physiological processes as well as in tumorigenesis. In addition to the abovementioned functional domains, the N-terminus of RECQL4 also contains several localization regions, including two nuclear localization domains [9], a region of acetylation by p300 which regulates nuclear to cytoplasmic localization [27], and a predicted mitochondrial localization signal in amino acids 1–84 [25].
Initially, researchers were unable to demonstrate actual DNA unwinding activity by RECQL4 using a variety of DNA substrates [69, 134]. Helicase activity was finally demonstrated for RECQL4 by several groups [11, 91, 107, 131], which was likely masked in previous assays by the strong annealing activity of the enzyme. In vitro biochemical data suggested that RECQL4 possesses another N-terminal region contributing to DNA unwinding besides the well-known conserved helicase domain [131], although known helicase motifs and nucleotide binding sites were not found to be present in this region. The in vivo function of this extra helicase domain requires further investigation. In addition to the helicase domain, other protein domains in RecQ helicases, including the helicase-and-RNase D C-terminal (HRDC) and RecQ-C-terminal (RQC) domains , are also important for helicase unwinding activity. However, RECQL4 lacks the structurally conserved HRDC domain which is felt to be important for interactions with nucleic acids (Fig. 3.1) [8, 80]. Human RECQL4 also appears to lack the structurally conserved RQC domain that is important for zinc and DNA binding and for helicase activity. However, through bioinformatic and biochemical analyses, Mojumdar et al. identified a functional RQC domain in human RECQL4 that is essential for these activities [72, 78]. In addition, the crystal structure of a human RECQL4 fragment (residues 449–1111), including the helicase domain and the majority of the C-terminus, revealed that a RECQL4 zinc binding domain (R4ZBD, residues 836–1045) resides downstream of the helicase domain and is important for DNA unwinding activity in a biochemical DNA helicase activity assay [50]. Interestingly, the last 92 residues of human RECQL4 have also been shown to play an important role in helicase activity by increasing DNA binding [50].
In addition to its role in DNA replication, RECQL4 has also been implicated to function in various aspects of DNA repair, including double-strand break (DSB) repair [61, 66, 67, 90, 97, 103], nucleotide excision repair (NER) [19, 31], and base excision repair (BER) [95]. RECQL4 plays important roles in both homologous recombination (HR)-dependent and nonhomologous end-joining (NHEJ)-mediated repair of DSBs . RECQL4 has been shown to interact physically with the Ku70/Ku80 heterodimer [97], which forms a complex with DNA-PKcs to play a central role in NHEJ-mediated DSB repair. During HR-dependent DSB repair, RECQL4 has been shown to interact physically by co-immunoprecipitation with RAD51, a key protein involved in the HR pathway of DSB repair, and to associate with RAD51 by immunofluorescence in DNA damage foci [61, 90, 103]. Lu et al. reported that RECQL4 participates in 5′ end resection of DSBs, the first step in HR-mediated DSB repair [67]. RECQL4 interacts with the MRE11-RAD50-NBS1 (MRN) complex and increases the recruitment of CtIP which stimulates end resection by the MRN complex [67]. Interestingly, the participation of RECQL4 in both pathways was shown to be cell cycle dependent and was regulated by the phosphorylation of RECQL4 by cyclin-dependent kinases CDK1/CDK2. RECQL4 stimulates NHEJ in G1 phase and promotes HR-mediated DSB repair in S and G2 phases when CDK1/CDK2 activity is high [66]. RECQL4 has also been shown to interact with BLM helicase, which like RECQL4 probably has many functions in the cell, the most important of which is its role in HR. This interaction was strengthened in S-phase and after ionizing radiation treatment in human cells, indicating that RECQL4 coordinates with BLM to function in DNA replication and DNA damage repair [104]. Ribosomal protein S3 (RPS3) , a component of 40S small subunit of the ribosome contributing to protein translation, has also been shown to interact with the N-terminus of RECQL4 and modulates its activity during DNA damage repair [89]. RECQL4 helicase activity appears to be essential for the end resection and HR-dependent repair of DSBs [67]. However, a knock-in mouse model (Recql4K525A) , mimicking human RECQL4K508M, displayed normal development and normal life span compared to wild-type littermates [12]. Cells derived from these mice had no significant difference in growth rate after treatment with genotoxic agents [12]. This discrepancy between human cells and mouse models requires more detailed investigation. Nevertheless, taken together, the data suggest that lack of RECQL4 functional activity in DNA repair can lead to increased DSBs, DNA replication stress, genomic instability, and cancer development.
The NER pathway is a major mediator of repair of UV damage, and RECQL4 has been shown to colocalize with XPA, a key protein involved in NER, and to interact with XPA directly by GST pull-down assay [31]. The BER pathway is the main mechanism for repair of oxidative DNA lesions, and RECQL4 was also found to colocalize and functionally interact with key proteins involved in BER, including APE1, FEN1, and DNA polymerase β, after treatment with H2O2 [95]. Werner et al. showed that after H2O2 treatment, RECQL4 translocates from the cytoplasm to the nucleus and forms nuclear foci in normal human fibroblasts. After recovery from oxidant damage, viable RTS patient fibroblasts underwent irreversible growth arrest and had significantly decreased DNA synthesis [124]. Woo et al. also showed that in response to oxidative stress, RECQL4 had altered cellular localization to the nucleolus and using a T7 phage display screen showed that RECQL4 C-terminus interacts with the single-strand break repair protein, poly(ADP-ribose) polymerase-1 (PARP-1) [125]. PARP-1 is activated in response to a wide variety of DNA-damaging agents and modulates the cellular sensitivity to γ-irradiation [68].
The response of RECQL4 mutant cells to different genotoxic agents has been investigated by several groups; these have included UV and ionizing radiation (IR), hydrogen peroxide, topoisomerases inhibitors, and chemotherapy agents such as doxorubicin and cisplatin [10, 19, 31, 49, 59, 103, 124]. However, the results have been somewhat inconsistent between studies, likely reflecting the use of different primary cells or cell lines (transformed cells vs. untransformed cells, RTS patient cells vs. RECQL4 knockdown cells), different assays to determine sensitivity, and different RECQL4 mutations present in the cells. For example, some studies have demonstrated significant increased sensitivity to UV radiation [88, 100, 105], while others have shown moderate or no increase in sensitivity [49, 59]. Using CRISPR-Cas9 , Kohzaki et al. deleted the C-terminus of RECQL4 after the NLS domain, including the conserved helicase domain, in several human cancer cell lines [60]. These cells displayed hypersensitivity to IR and cisplatin, which primarily introduce DNA DSB and interstrand cross-links, respectively. In vitro cell-based DSB repair reporter assays showed that these cells displayed increased single-strand annealing activity and reduced alternative end-joining mediated pathway. They showed that RAD52 inhibition suppressed the growth of cancer cell lines in vitro and in xenograft mouse models. In addition, cisplatin treatment had an additive inhibitory effect with RAD52 inhibition on tumor cell growth, providing a potential treatment avenue for cancer patients with RECQL4 mutations and increased RAD52 expression [60].
As mentioned earlier, RECQ proteins bind to G4 structures such as those found in telomeric DNA , and RECQL4 has been shown to play a role in telomere maintenance [38]. RTS patient cells and human cells with RECQL4 knockdown exhibit increased fragile telomeric ends. In addition, human RECQL4 localizes to telomeres and interacts with shelterin protein telomeric repeat-binding factor 2 (TRF2) which maintains telomere integrity [38]. RECQL4 also interacts with the WRN protein and stimulates WRN’s activity on telomeric D-loops. Similar to WRN and BLM, RECQL4 also appears to be able to resolve these D-loops, which is necessary for replication to take place at the telomeres, and this resolving activity is stimulated by TRF1 and TRF2 as well as the shelterin protein POT1 [38]. Also similar to WRN and BLM, RECQL4 seems to be more active on telomeric D-loops that contain 8-oxoguanine base lesions, indicative of oxidative damage. Unlike WRN , however, RECQL4 also has a clear preference for unwinding D-loops that contain thymine glycol (Tg) lesions, which are the most common oxidation product of the thymine base, and this activity is stimulated by TRF2 [34]. Thus, mutations in RECQL4 could result in dysfunctional telomeres, which are well known to play a role in both tumor suppression and tumor progression, depending on the cellular milieu, particularly with respect to the checkpoint status of the cells [130].
In addition to these nuclear functions, RECQL4 has also been shown to localize in the cytosol [27, 134] as well as in the mitochondria [16, 21, 25, 120]. Yin et al. showed that RECQL4 interacts with cytosolic ubiquitin ligases UBR1 and UBR2 which function in the N-end rule pathway by ubiquitination and degradation of proteins [134]. Dietschy et al. demonstrated that RECQL4 can be acetylated by histone acetyltransferase p300 resulting in the cytosolic translocation of RECQL4 from the nucleus [27], providing a mechanism to modulate RECQL4 nuclear activities. In the mitochondria , loss of RECQL4 led to abnormalities in mitochondrial DNA (mtDNA) as well as mitochondrial function caused by reduced replication of mtDNA [16, 25] or caused by reduced proofreading and polymerization functions of mitochondrial DNA polymerase-γ (PolγA) [40]. Interestingly, a RECQL4 mutation frequently reported in RAPADILINO patients who are predisposed to lymphoma and osteosarcoma disrupts the interaction between RECQL4 and mitochondrial p32 protein [120] while also enhancing the interaction between RECQL4 and mitochondrial helicase PEO1, leading to increased replication of mtDNA. Both increased or decreased mtDNA content could cause abnormal mitochondrial function demonstrated by abnormal mitochondrial metabolism and glycolysis [40, 120].
In addition to the abovementioned cellular functions, RECQL4 was also recently demonstrated to play a role in mitosis. RECQL4 was shown to be a microtubule-associated protein and to participate in the maintenance of chromosome alignment during mitosis [135]. It was identified among the proteins with a nuclear localization sequence (NLS) that can be pulled down by Taxol-stabilized microtubules in mitotic Xenopus egg extracts. RECQL4-depleted HeLa cells as well as RTS fibroblasts exhibited spindle abnormalities, including misaligned chromosomes and increased micronuclei. Interestingly, using immunoprecipitation with tagged proteins and GST pull-down assays in human cells, RECQL4 was shown to interact with aurora kinase B (AURKB) and to modulate its protein stability by reducing ubiquitination of AURKB [33], an essential protein that modulates mitosis by regulating chromosome alignment and segregation.
Rothmund-Thomson Syndrome (RTS): Nature’s Model of Osteosarcoma
RTS was first described in 1868 by Dr. Auguste Rothmund, who was a German ophthalmologist. He described poikiloderma, the classic skin finding in RTS, along with rapidly developing bilateral juvenile cataracts in several families in an isolated region in the Bavarian Alps [92]. In 1921, Dr. Sydney Thomson, a British dermatologist, described a similar rash in two sisters, but instead of juvenile cataracts, they had bone abnormalities (radial ray defects) [114]. Later, Dr. William Taylor in the United States proposed that the two disorders described by Rothmund and Thomson were the same, and he proposed the eponym Rothmund-Thomson syndrome [112]. Mutations in the RECQL4 gene in RTS were not discovered until 1999 [55, 56], 131 years after the original description by Rothmund. It is now known that approximately two-thirds of patients with RTS have mutations in the RECQL4 gene (designated Type 2 RTS). The other one-third of patients who lack RECQL4 mutations are designated as Type 1 RTS. Mutations in the ANAPC1 gene, which encodes the APC1 protein, a component of the anaphase-promoting complex/cyclosome (APC/C), have recently been identified as causative in a subset of Type 1 RTS patients [3]. Previous studies have shown that the presence of pathogenic mutations in RECQL4 correlates significantly with risk of developing OS (Fig. 3.2) [121]. None of the patients with Type 1 RTS developed OS, while every RTS patient with OS had RECQL4 mutations. These pathogenic mutations included nonsense, frameshift, splice site, and intronic deletions. Unlike other hereditary cancer syndromes known to predispose patients to OS, such as Li-Fraumeni syndrome and hereditary retinoblastoma, where the causative genes, p53 and RB, respectively, are commonly mutated in sporadic OS [14], mutations in RECQL4 have not been detected in sporadic OS tumors [86]. Thus, RECQL4 does not appear to be a direct target for somatic mutations in sporadic OS. However, the extremely high and specific risk for OS in Type 2 RTS patients suggests that the RECQL4 helicase plays a clear role in OS tumor suppression, making RTS a relevant model for the study of human OS pathogenesis.
Estimated probability of osteosarcoma onset in Rothmund-Thomson syndrome, classified by RECQL4 mutation status. The time to OS onset was defined from the date of birth to the first diagnosis of OS. Event-time data were analyzed by Kaplan-Meier method, and the difference between the RECQL4 mutation-positive and RECQL4 mutation-negative patients was compared by the log-rank method
In addition to cancer of the bone, patients with RTS also have prominent developmental defects of the bone. In a study of 28 RTS patients who underwent skeletal surveys, 75% were found to have major skeletal abnormalities, including radial, ulnar, or thumb agenesis/hypoplasia, radioulnar and radiohumeral synostoses, abnormal metaphyseal trabeculation, brachymesophalangy, and osteopenia [75]. This risk correlated with presence of RECQL4 mutations. Understanding the role that RECQL4 plays in normal skeletal development will provide additional insight into the specific risk for OS, since many developmental pathways, such as the Wnt, Hedgehog, and Notch signaling pathways, not only are critical for normal skeletal development [39, 44, 110] but also play important roles in tumorigenesis [7, 18, 52, 111, 118].
Early case reports suggested that OS arising in RTS patients may be different from sporadic OS, i.e., arising in unusual or multiple (multifocal) sites [29]. In addition, because of the implicated role of RECQL4 in DNA damage repair, clinicians may consider decreasing chemotherapy doses up-front for RTS patients diagnosed with OS. However, a study of 12 RTS patients with OS showed that their tumors had features that mirrored OS in the general population with regard to location of primary tumor (distal long bones), histology (conventional OS), histologic response to neoadjuvant chemotherapy, and overall outcomes [42]. The major difference was that the age of onset was younger in the RTS cohort compared to sporadic OS, which is not surprising given the genetic predisposition of RTS patients to OS. Some patients developed mucositis requiring dose modifications, particularly to doxorubicin (no more than 25% decrease), but there is no current method to determine a priori who will experience increased toxicities. Therefore, current recommendations are to treat with standard doses of chemotherapy and to adjust according to the patient’s individual course. The similarities between OS in RTS and sporadic OS support the further study of the contribution of the RECQL4 pathways in the pathogenesis of OS.
Understanding the Role of RECQL4 in Osteosarcoma Development Using Mouse Models
Recql4 Global Knockout Mouse Models
In order to understand the function of RECQL4 in OS tumorigenesis in vivo, three mouse models of global Recql4 disruption have been generated. In the first model, exons 5–8 of Recql4 upstream of the conserved helicase domain (exons 9–15) were replaced with PGKneo and LacZ cassettes [45]. Homozygous mutants died during early embryonic stage E3.5–6.5. Although there was no information about transcripts and protein levels of Recql4 in the paper, presumably this targeting strategy generated a null mutation as a result of nonsense mediated decay. The second mouse model by Hoki et al. targeted exon 13 of the helicase domain of Recql4 with a neomycin cassette [43]. These homozygous mutants were viable at birth, but 95% of them died within 2 weeks. The remaining 5% exhibited growth retardation, skin atrophy, hair abnormalities, and tissue hypoplasia, such as severely reduced bone trabeculae and fewer and smaller villi of the small intestine. The MEFs from these mutants showed reduced proliferation. However, there was no malignancy reported in these mice. The third global mouse model was generated by replacing exons 9–13 in the conserved helicase domain of Recql4 with a PGK-HPRT cassette [71]. Homozygous mutants were born alive with normal Mendelian ratio, but 16% of them died within 24 hours of birth. The remaining mutants exhibited tail pigmentation defects by 12 months, and palatal patterning defects were seen in all examined animals. Furthermore, 6% of these mutants developed limb defects at birth, ranging from preaxial polydactyly of hindlimbs to forelimb aplasia. Interestingly, 5% of these mutants developed OS or lymphoma by 20 months, while heterozygous and wild-type mice had no tumor formation, although this difference was not found to be statistically significant .
Recql4 Conditional (Bone-Specific) Mouse Models
Because the previous global Recql4 knockout mouse models failed to recapitulate the high risk of OS seen in RTS patients with RECQL4 mutations, skeletal-specific conditional knockout mouse models have been developed to assess the effect of Recql4 deficiency in the bone. Lu et al. developed a conditional knockout model of Recql4 in early skeletal progenitor cell system by crossing these Recql4 mice with Prx1-Cre transgenic mice. Resultant mutants developed foreshortened limbs, digit defects, abnormal growth plates and joints, and craniosynostosis, recapitulating the major skeletal defects seen in RTS patients. Mouse tissues lacking Recql4 displayed increased DNA damage and elevated p53 activation, leading to increased cell death, reduced cell proliferation, and increased senescence. These defects were partially rescued by concurrent inactivation of p53, indicating that p53 activation may contribute to the skeletal phenotypes seen in RTS patients. RTS human fibroblasts were also shown to have increased p53 phosphorylation and expression of downstream target genes of p53 [24, 25]. Similarly, depletion of RECQL4 in primary human fibroblasts causes increased DNA damage and cellular senescence as well as p53 activation and increased expression of target genes [65].
Ng et al. developed another conditional knockout model using Osx-Cre to inactivate Recql4 in osteoblast progenitor cells at a later stage of skeletal development, and they observed reduced body weight and decrease in trabecular and cortical bone [83]. Mice lacking Recql4 in the osteocytes and a subset of osteoblasts showed no striking developmental skeletal abnormalities [83], indicating that RECQL4 plays a more important developmental role in the early stages of osteoblast differentiation. Unlike human RTS patients, however, these homozygous Recql4 conditional knockout mice did not develop OS. Interestingly, mice with homozygous loss of both Recql4 and p53 in the osteoblast progenitor cells showed delayed osteosarcoma development and significantly longer survival compared to p53 homozygous loss alone, indicating that Recql4 may actually be necessary for OS development in mice [83]. The mouse models developed to date have not been able to recapitulate the high incidence of OS seen in RTS patients, and further work is in progress to understand these differences and to dissect the molecular mechanisms underlying OS development in RTS patients.
In order to more closely mirror the human disease, induced pluripotent stem cell (iPSC) techniques have been used to model RECQ syndromes using patient-derived somatic cells including peripheral blood mononuclear cells and dermal fibroblasts. Werner syndrome iPSCs have been generated by several groups [15, 35, 98, 123], and they exhibit normal karyotypes and stable chromosomes after long-term culture [98]. In addition, human embryonic stem cells were also used to generate WRN-deficient cells which were further differentiated into human mesenchymal stem cells (MSCs), demonstrating that WRN is essential for maintaining heterochromatin stability and that loss of WRN in human MSCs leads to disorganization of heterochromatin and increased senescence [138]. Thus far, an iPSC line has been generated from dermal fibroblasts derived from a RECQL4 heterozygous carrier [48], and work is ongoing to establish iPSC lines differentiated into osteoblasts from RTS patient fibroblasts with biallelic RECQL4 mutations in order to identify the molecular mechanisms underlying the high risk of OS in RTS patients .
Clinical Implications for Understanding RECQ Gene Defects and Potentially Targeting RECQ-Related Pathways for Cancer Therapy
Based on the roles of the RECQ proteins in normal cellular proliferation, DNA damage response, DNA repair, and telomere maintenance, there is growing interest in exploring inhibition of these functions in susceptible cancer cell types. Small molecule inhibitors of the WRN [1] and BLM [84] proteins have been identified as potential antiproliferative cancer therapies. Both of these molecules were identified through in vitro helicase activity screens. The WRN inhibitor, a small molecule inhibitor identified from the National Cancer Institute Diversity Set, designated NSC 19630 [2], was shown to inhibit cell proliferation and to induce apoptosis in a WRN-dependent manner. It also caused increase in DSBs and accumulation of blocked replication forks in human tumor cells grown in culture. NSC 19630 also had a synergistic effect on inhibiting cell proliferation when cells were co-treated along with telomestatin, a small molecule that binds G4 structures and causes disruption of telomere-associated proteins, as well as a PARP inhibitor KU0058948. It also acted synergistically with the topoisomerase inhibitor topotecan in inducing DSBs. Investigators later characterized a structurally related compound, NSC 617145, which they demonstrated was able to sensitize cancer cells to mitomycin C, resulting in decreased cell proliferation, increased DNA damage, and chromosomal abnormalities [1]. More recently, through high-throughput CRISPR-Cas9-mediated knockout and/or RNA interference screening, the WRN helicase has been shown by several groups to be a promising synthetically lethal target in cancers with high levels of microsatellite instability (MSI), including colorectal, endometrial, ovarian, and gastric cancers [6, 13, 53, 63]. MSI is caused by an impaired DNA mismatch repair pathway leading to small insertions and/or deletions in genomic nucleotide repeats. The helicase function of WRN is essential for this synthetic lethality [13, 63], which was not observed with other RecQ helicases [13, 63]. Similarly, inactivation of WRN leads to increased DNA damage and cell death in MSI high cancer cells, but not in microsatellite stable cancer cells [6, 13, 53, 63]. Therefore, these small molecule inhibitors to the WRN protein may be useful to target cancers with high levels of MSI.
The small molecule inhibitor of BLM, ML216 [84], was found to exert its action by preventing BLM from binding to DNA. Cells treated with ML216 showed decreased proliferation as well as an increase in sister chromatid exchanges, a hallmark of Bloom syndrome. One of the proposed future uses of this BLM-specific inhibitor would be to test its efficacy in treating tumor cells that depend on the ALT (alternative lengthening of telomeres) mechanism for maintenance of telomeres, since previous work showed that the BLM orthologue Sgs1 is required for telomere maintenance in the absence of telomerase[140]. Approximately 5–10% of tumors depend on the ALT pathway for continued proliferation, including OS; therefore, further exploration of this BLM-specific inhibitor could reveal a new therapeutic strategy for targeting susceptible tumors.
Expression of RECQL4 has been found to be upregulated in a variety of cancer types in addition to sporadic OS [70, 93], including soft tissue sarcomas [64], prostate cancer [106], cervical cancer [82], breast cancer [4, 32], gastric cancer [76], and oral cancer [136], suggesting that inactivation of RECQL4, and thus inhibition of its functions in cellular replication/viability, genome stability, DNA repair, and telomere maintenance, may be attractive as a potential adjunct to cancer therapy in susceptible tumor cells. RECQL4 may also work in coordination with other Holliday junction processing proteins, including BLM, to prevent replication fork stalling and reversal in order to maintain cancer cell fitness by resolving increased Holliday junctions in cancer cells with overexpression of RAD51 [128, 129]. Additionally, in gastric cancer cells, overexpression of RECQL4 has been linked to increased resistance to cisplatin by physically interacting with YB1 and AKT, as well as by increasing AKT-dependent YB1 phosphorylation and expression of the downstream drug resistance gene MDR1 [76]. These data suggest that RECQL4 may be required for rapid tumor cell proliferation and chemoresistance, providing a potential therapeutic target for cancer cells with overexpression of RECQL4.
Although sporadic OS tumors have not been found to have somatic RECQL4 mutations, a recent study which examined germ line sequence data from over 5000 sporadic pediatric cancer patients revealed an increase in heterozygous RECQL4 loss-of-function variants in OS patients compared to non-cancer database controls[141]. While presence of a RECQL4 heterozygous mutation does not cause RTS with its associated high risk of OS, it may still confer an elevated OS risk in carriers compared to the general population. This has implications for genetic counseling of these patients and may also offer potential avenues for novel targeted therapies for their specific tumors. Ongoing basic science and clinical research is needed to fully understand the cellular context and molecular mechanisms by which RECQL4 exerts its actions on osteosarcomagenesis, and this will provide useful information on the basic biology of OS and open avenues for potential new therapies for OS .
References
Aggarwal M, Banerjee T, Sommers JA, Brosh RM (2013) Targeting an Achilles’ heel of cancer with a WRN helicase inhibitor. Cell Cycle 12(20):3329–3335
Aggarwal M, Sommers JA, Shoemaker RH, Brosh RM Jr (2011) Inhibition of helicase activity by a small molecule impairs Werner syndrome helicase (WRN) function in the cellular response to DNA damage or replication stress. Proc Natl Acad Sci U S A 108(4):1525–1530
Ajeawung NF, Nguyen TTM, Lu L, Kucharski TJ, Rousseau J, Molidperee S, Atienza J, Gamache I, Jin W, Plon SE, Lee BH, Teodoro JG, Wang LL, Campeau PM (2019) Mutations in ANAPC1, encoding a scaffold subunit of the anaphase-promoting complex, cause Rothmund-Thomson syndrome type 1. Am J Hum Genet 105(3):625–630. https://doi.org/10.1016/j.ajhg.2019.06.011
Arora A, Agarwal D, Abdel-Fatah TM, Lu H, Croteau DL, Moseley P, Aleskandarany MA, Green AR, Ball G, Rakha EA, Chan SY, Ellis IO, Wang LL, Zhao Y, Balajee AS, Bohr VA, Madhusudan S (2016) RECQL4 helicase has oncogenic potential in sporadic breast cancers. J Pathol 238(4):495–501. https://doi.org/10.1002/path.4681
Bachrati CZ, Hickson ID (2003) RecQ helicases: suppressors of tumorigenesis and premature aging. Biochem J 374(Pt 3):577–606. https://doi.org/10.1042/BJ20030491
Behan FM, Iorio F, Picco G, Goncalves E, Beaver CM, Migliardi G, Santos R, Rao Y, Sassi F, Pinnelli M, Ansari R, Harper S, Jackson DA, McRae R, Pooley R, Wilkinson P, van der Meer D, Dow D, Buser-Doepner C, Bertotti A, Trusolino L, Stronach EA, Saez-Rodriguez J, Yusa K, Garnett MJ (2019) Prioritization of cancer therapeutic targets using CRISPR-Cas9 screens. Nature 568(7753):511–516. https://doi.org/10.1038/s41586-019-1103-9
Belyea B, Kephart JG, Blum J, Kirsch DG, Linardic CM (2012) Embryonic signaling pathways and rhabdomyosarcoma: contributions to cancer development and opportunities for therapeutic targeting. Sarcoma 2012:406239
Bernstein KA, Gangloff S, Rothstein R (2010) The RecQ DNA helicases in DNA repair. Annu Rev Genet 44:393–417. https://doi.org/10.1146/annurev-genet-102209-163602
Burks LM, Yin J, Plon SE (2007) Nuclear import and retention domains in the amino terminus of RECQL4. Gene 391(1–2):26–38. https://doi.org/10.1016/j.gene.2006.11.019
Cabral RE, Queille S, Bodemer C, de Prost Y, Neto JB, Sarasin A, ya-Grosjean L (2008) Identification of new RECQL4 mutations in Caucasian Rothmund-Thomson patients and analysis of sensitivity to a wide range of genotoxic agents. Mutat Res 643(1–2):41–47
Capp C, Wu J, Hsieh TS (2009) Drosophila RecQ4 has a 3′-5’ DNA helicase activity that is essential for viability. J Biol Chem 284(45):30845–30852. https://doi.org/10.1074/jbc.M109.008052
Castillo-Tandazo W, Smeets MF, Murphy V, Liu R, Hodson C, Heierhorst J, Deans AJ, Walkley CR (2019) ATP-dependent helicase activity is dispensable for the physiological functions of Recql4. PLoS Genet 15(7):e1008266. https://doi.org/10.1371/journal.pgen.1008266
Chan EM, Shibue T, McFarland JM, Gaeta B, Ghandi M, Dumont N, Gonzalez A, McPartlan JS, Li T, Zhang Y, Bin Liu J, Lazaro JB, Gu P, Piett CG, Apffel A, Ali SO, Deasy R, Keskula P, Ng RWS, Roberts EA, Reznichenko E, Leung L, Alimova M, Schenone M, Islam M, Maruvka YE, Liu Y, Roper J, Raghavan S, Giannakis M, Tseng YY, Nagel ZD, D’Andrea A, Root DE, Boehm JS, Getz G, Chang S, Golub TR, Tsherniak A, Vazquez F, Bass AJ (2019) WRN helicase is a synthetic lethal target in microsatellite unstable cancers. Nature 568(7753):551–556. https://doi.org/10.1038/s41586-019-1102-x
Chen X, Bahrami A, Pappo A, Easton J, Dalton J, Hedlund E, Ellison D, Shurtleff S, Wu G, Wei L, Parker M, Rusch M, Nagahawatte P, Wu J, Mao S, Boggs K, Mulder H, Yergeau D, Lu C, Ding L, Edmonson M, Qu C, Wang J, Li Y, Navid F, Daw NC, Mardis ER, Wilson RK, Downing JR, Zhang J, Dyer MA, St. Jude Children’s Research Hospital-Washington University Pediatric Cancer Genome P (2014) Recurrent somatic structural variations contribute to tumorigenesis in pediatric osteosarcoma. Cell Rep 7(1):104–112. https://doi.org/10.1016/j.celrep.2014.03.003
Cheung HH, Liu X, Canterel-Thouennon L, Li L, Edmonson C, Rennert OM (2014) Telomerase protects werner syndrome lineage-specific stem cells from premature aging. Stem Cell Rep 2(4):534–546. https://doi.org/10.1016/j.stemcr.2014.02.006
Chi Z, Nie L, Peng Z, Yang Q, Yang K, Tao J, Mi Y, Fang X, Balajee AS, Zhao Y (2012) RecQL4 cytoplasmic localization: implications in mitochondrial DNA oxidative damage repair. Int J Biochem Cell Biol 44(11):1942–1951. https://doi.org/10.1016/j.biocel.2012.07.016
Chu WK, Hickson ID (2009) RecQ helicases: multifunctional genome caretakers. Nat Rev Cancer 9(9):644–654. https://doi.org/10.1038/nrc2682
Cleton-Jansen AM, Anninga JK, Briaire-de BI, Romeo S, Oosting J, Egeler RM, Gelderblom H, Taminiau AH, Hogendoorn PC (2009) Profiling of high-grade central osteosarcoma and its putative progenitor cells identifies tumourigenic pathways. Br J Cancer 101(11):1909–1918
Crevel G, Vo N, Crevel I, Hamid S, Hoa L, Miyata S, Cotterill S (2012) Drosophila RecQ4 is directly involved in both DNA replication and the response to UV damage in S2 cells. PLoS One 7(11):e49505. https://doi.org/10.1371/journal.pone.0049505
Croteau DL, Popuri V, Opresko PL, Bohr VA (2014) Human RecQ helicases in DNA repair, recombination, and replication. Annu Rev Biochem 83:519–552. https://doi.org/10.1146/annurev-biochem-060713-035428
Croteau DL, Rossi ML, Canugovi C, Tian J, Sykora P, Ramamoorthy M, Wang ZM, Singh DK, Akbari M, Kasiviswanathan R, Copeland WC, Bohr VA (2012) RECQL4 localizes to mitochondria and preserves mitochondrial DNA integrity. Aging Cell 11(3):456–466. https://doi.org/10.1111/j.1474-9726.2012.00803.x
Cunniff C, Bassetti JA, Ellis NA (2017) Bloom’s syndrome: clinical spectrum, molecular pathogenesis, and cancer predisposition. Mol Syndromol 8(1):4–23. https://doi.org/10.1159/000452082
Cybulski C, Carrot-Zhang J, Kluzniak W, Rivera B, Kashyap A, Wokolorczyk D, Giroux S, Nadaf J, Hamel N, Zhang S, Huzarski T, Gronwald J, Byrski T, Szwiec M, Jakubowska A, Rudnicka H, Lener M, Masojc B, Tonin PN, Rousseau F, Gorski B, Debniak T, Majewski J, Lubinski J, Foulkes WD, Narod SA, Akbari MR (2015) Germline RECQL mutations are associated with breast cancer susceptibility. Nat Genet 47(6):643–646. https://doi.org/10.1038/ng.3284
Davis T, Tivey HS, Brook AJ, Grimstead JW, Rokicki MJ, Kipling D (2013) Activation of p38 MAP kinase and stress signalling in fibroblasts from the progeroid Rothmund-Thomson syndrome. Age (Dordr) 35(5):1767–1783. https://doi.org/10.1007/s11357-012-9476-9
De S, Kumari J, Mudgal R, Modi P, Gupta S, Futami K, Goto H, Lindor NM, Furuichi Y, Mohanty D, Sengupta S (2012) RECQL4 is essential for the transport of p53 to mitochondria in normal human cells in the absence of exogenous stress. J Cell Sci 125(Pt 10):2509–2522. https://doi.org/10.1242/jcs.101501
Debeljak M, Zver A, Jazbec J (2009) A patient with Baller-Gerold syndrome and midline NK/T lymphoma. Am J Med Genet A 149A(4):755–759. https://doi.org/10.1002/ajmg.a.32736
Dietschy T, Shevelev I, Pena-Diaz J, Huhn D, Kuenzle S, Mak R, Miah MF, Hess D, Fey M, Hottiger MO, Janscak P, Stagljar I (2009) p300-mediated acetylation of the Rothmund-Thomson-syndrome gene product RECQL4 regulates its subcellular localization. J Cell Sci 122(Pt 8):1258–1267. https://doi.org/10.1242/jcs.037747
Dong YZ, Huang YX, Lu T (2015) Single nucleotide polymorphism in the RECQL5 gene increased osteosarcoma susceptibility in a Chinese Han population. Genet Mol Res 14(1):1899–1902. https://doi.org/10.4238/2015.March.13.18
el-Khoury JM, Haddad SN, Atallah NG (1997) Osteosarcomatosis with Rothmund-Thomson syndrome. Br J Radiol 70:215–218. https://doi.org/10.1259/bjr.70.830.9135453
Ellis NA, Groden J, Ye TZ, Straughen J, Lennon DJ, Ciocci S, Proytcheva M, German J (1995) The Bloom’s syndrome gene product is homologous to RecQ helicases. Cell 83(4):655–666
Fan W, Luo J (2008) RecQ4 facilitates UV light-induced DNA damage repair through interaction with nucleotide excision repair factor xeroderma pigmentosum group A (XPA). J Biol Chem 283(43):29037–29044. https://doi.org/10.1074/jbc.M801928200
Fang H, Nie L, Chi Z, Liu J, Guo D, Lu X, Hei TK, Balajee AS, Zhao Y (2013) RecQL4 helicase amplification is involved in human breast tumorigenesis. PLoS One 8(7):e69600. https://doi.org/10.1371/journal.pone.0069600
Fang H, Niu K, Mo D, Zhu Y, Tan Q, Wei D, Li Y, Chen Z, Yang S, Balajee AS, Zhao Y (2018) RecQL4-Aurora B kinase axis is essential for cellular proliferation, cell cycle progression, and mitotic integrity. Oncogenesis 7(9):68. https://doi.org/10.1038/s41389-018-0080-4
Ferrarelli LK, Popuri V, Ghosh AK, Tadokoro T, Canugovi C, Hsu JK, Croteau DL, Bohr VA (2013) The RECQL4 protein, deficient in Rothmund-Thomson syndrome is active on telomeric D-loops containing DNA metabolism blocking lesions. DNA Repair 12(7):518–528. https://doi.org/10.1016/j.dnarep.2013.04.005
Gatinois V, Desprat R, Becker F, Pichard L, Bernex F, Corsini C, Pellestor F, Lemaitre JM (2019) Reprogramming of Human Peripheral Blood Mononuclear Cell (PBMC) from a patient suffering of a Werner syndrome resulting in iPSC line (REGUi003-A) maintaining a short telomere length. Stem Cell Res 39:101515. https://doi.org/10.1016/j.scr.2019.101515
German J (1995) Bloom’s syndrome. Dermatol Clin 13(1):7–18
German J (1997) Bloom’s syndrome. XX. The first 100 cancers. Cancer Genet Cytogenet 93(1):100–106
Ghosh AK, Rossi ML, Singh DK, Dunn C, Ramamoorthy M, Croteau DL, Liu Y, Bohr VA (2012) RECQL4, the protein mutated in Rothmund-Thomson syndrome, functions in telomere maintenance. J Biol Chem 287(1):196–209. https://doi.org/10.1074/jbc.M111.295063
Glass DA, Karsenty G (2007) In vivo analysis of Wnt signaling in bone. Endocrinology 148(6):2630–2634
Gupta S, De S, Srivastava V, Hussain M, Kumari J, Muniyappa K, Sengupta S (2014) RECQL4 and p53 potentiate the activity of polymerase gamma and maintain the integrity of the human mitochondrial genome. Carcinogenesis 35(1):34–45. https://doi.org/10.1093/carcin/bgt315
Hansel-Hertsch R, Di Antonio M, Balasubramanian S (2017) DNA G-quadruplexes in the human genome: detection, functions and therapeutic potential. Nat Rev Mol Cell Biol 18(5):279–284. https://doi.org/10.1038/nrm.2017.3
Hicks MJ, Roth JR, Kozinetz CA, Wang LL (2007) Clinicopathologic features of osteosarcoma in patients with Rothmund-Thomson syndrome. J Clin Oncol 25(4):370–375. https://doi.org/10.1200/JCO.2006.08.4558
Hoki Y, Araki R, Fujimori A, Ohhata T, Koseki H, Fukumura R, Nakamura M, Takahashi H, Noda Y, Kito S, Abe M (2003) Growth retardation and skin abnormalities of the Recql4-deficient mouse. Hum Mol Genet 12(18):2293–2299. https://doi.org/10.1093/hmg/ddg254
Hu H, Hilton MJ, Tu X, Yu K, Ornitz DM, Long F (2005) Sequential roles of Hedgehog and Wnt signaling in osteoblast development. Development 132(1):49–60
Ichikawa K, Noda T, Furuichi Y (2002) Preparation of the gene targeted knockout mice for human premature aging diseases, Werner syndrome, and Rothmund-Thomson syndrome caused by the mutation of DNA helicases. Nihon yakurigaku zasshi Folia pharmacologica Japonica 119(4):219–226
Im JS, Ki SH, Farina A, Jung DS, Hurwitz J, Lee JK (2009) Assembly of the Cdc45-Mcm2-7-GINS complex in human cells requires the Ctf4/And-1, RecQL4, and Mcm10 proteins. Proc Natl Acad Sci U S A 106(37):15628–15632. https://doi.org/10.1073/pnas.0908039106
Im JS, Park SY, Cho WH, Bae SH, Hurwitz J, Lee JK (2015) RecQL4 is required for the association of Mcm10 and Ctf4 with replication origins in human cells. Cell Cycle 14(7):1001–1009. https://doi.org/10.1080/15384101.2015.1007001
Jewell BE, Liu M, Lu L, Zhou R, Tu J, Zhu D, Huo Z, Xu A, Wang D, Mata H, Jin W, Xia W, Rao PH, Zhao R, Hung MC, Wang LL, Lee DF (2018) Generation of an induced pluripotent stem cell line from an individual with a heterozygous RECQL4 mutation. Stem Cell Res 33:36–40. https://doi.org/10.1016/j.scr.2018.10.003
Jin W, Liu H, Zhang Y, Otta SK, Plon SE, Wang LL (2008) Sensitivity of RECQL4-deficient fibroblasts from Rothmund-Thomson syndrome patients to genotoxic agents. Hum Genet 123(6):643–653. https://doi.org/10.1007/s00439-008-0518-4
Kaiser S, Sauer F, Kisker C (2017) The structural and functional characterization of human RecQ4 reveals insights into its helicase mechanism. Nat Commun 8:15907. https://doi.org/10.1038/ncomms15907
Kamimura Y, Masumoto H, Sugino A, Araki H (1998) Sld2, which interacts with Dpb11 in Saccharomyces cerevisiae, is required for chromosomal DNA replication. Mol Cell Biol 18(10):6102–6109
Kansara M, Tsang M, Kodjabachian L, Sims NA, Trivett MK, Ehrich M, Dobrovic A, Slavin J, Choong PF, Simmons PJ, Dawid IB, Thomas DM (2009) Wnt inhibitory factor 1 is epigenetically silenced in human osteosarcoma, and targeted disruption accelerates osteosarcomagenesis in mice. J Clin Invest 119(4):837–851
Kategaya L, Perumal SK, Hager JH, Belmont LD (2019) Werner syndrome helicase is required for the survival of cancer cells with microsatellite instability. iScience 13:488–497. https://doi.org/10.1016/j.isci.2019.02.006
Keller H, Kiosze K, Sachsenweger J, Haumann S, Ohlenschlager O, Nuutinen T, Syvaoja JE, Gorlach M, Grosse F, Pospiech H (2014) The intrinsically disordered amino-terminal region of human RecQL4: multiple DNA-binding domains confer annealing, strand exchange and G4 DNA binding. Nucleic Acids Res 42(20):12614–12627. https://doi.org/10.1093/nar/gku993
Kitao S, Ohsugi I, Ichikawa K, Goto M, Furuichi Y, Shimamoto A (1998) Cloning of two new human helicase genes of the RecQ family: biological significance of multiple species in higher eukaryotes. Genomics 54(3):443–452. https://doi.org/10.1006/geno.1998.5595
Kitao S, Shimamoto A, Goto M, Miller RW, Smithson WA, Lindor NM, Furuichi Y (1999) Mutations in RECQL4 cause a subset of cases of Rothmund-Thomson syndrome. Nat Genet 22(1):82–84. https://doi.org/10.1038/8788
Kliszczak M, Sedlackova H, Pitchai GP, Streicher WW, Krejci L, Hickson ID (2015) Interaction of RECQ4 and MCM10 is important for efficient DNA replication origin firing in human cells. Oncotarget 6(38):40464–40479. https://doi.org/10.18632/oncotarget.6342
Kobbe D, Focke M, Puchta H (2010) Purification and characterization of RecQ helicases of plants. Methods Mol Biol 587:195–209
Kohzaki M, Chiourea M, Versini G, Adachi N, Takeda S, Gagos S, Halazonetis TD (2012) The helicase domain and C-terminus of human RecQL4 facilitate replication elongation on DNA templates damaged by ionizing radiation. Carcinogenesis 33(6):1203–1210. https://doi.org/10.1093/carcin/bgs149
Kohzaki M, Ootsuyama A, Sun L, Moritake T, Okazaki R (2019) Human RECQL4 represses the RAD52-mediated single-strand annealing pathway after ionizing radiation or cisplatin treatment. Int J Cancer. https://doi.org/10.1002/ijc.32670
Kumata Y, Tada S, Yamanada Y, Tsuyama T, Kobayashi T, Dong YP, Ikegami K, Murofushi H, Seki M, Enomoto T (2007) Possible involvement of RecQL4 in the repair of double-strand DNA breaks in Xenopus egg extracts. Biochim Biophys Acta 1773(4):556–564. https://doi.org/10.1016/j.bbamcr.2007.01.005
Lauper JM, Krause A, Vaughan TL, Monnat RJ Jr (2013) Spectrum and risk of neoplasia in Werner syndrome: a systematic review. PLoS One 8(4):e59709
Lieb S, Blaha-Ostermann S, Kamper E, Rippka J, Schwarz C, Ehrenhofer-Wolfer K, Schlattl A, Wernitznig A, Lipp JJ, Nagasaka K, van der Lelij P, Bader G, Koi M, Goel A, Neumuller RA, Peters JM, Kraut N, Pearson MA, Petronczki M, Wohrle S (2019) Werner syndrome helicase is a selective vulnerability of microsatellite instability-high tumor cells. Elife 8:e43333. https://doi.org/10.7554/eLife.43333
Linton KM, Hey Y, Saunders E, Jeziorska M, Denton J, Wilson CL, Swindell R, Dibben S, Miller CJ, Pepper SD, Radford JA, Freemont AJ (2008) Acquisition of biologically relevant gene expression data by Affymetrix microarray analysis of archival formalin-fixed paraffin-embedded tumours. Br J Cancer 98(8):1403–1414
Lu H, Fang EF, Sykora P, Kulikowicz T, Zhang Y, Becker KG, Croteau DL, Bohr VA (2014) Senescence induced by RECQL4 dysfunction contributes to Rothmund-Thomson syndrome features in mice. Cell Death Dis 5:e1226. https://doi.org/10.1038/cddis.2014.168
Lu H, Shamanna RA, de Freitas JK, Okur M, Khadka P, Kulikowicz T, Holland PP, Tian J, Croteau DL, Davis AJ, Bohr VA (2017) Cell cycle-dependent phosphorylation regulates RECQL4 pathway choice and ubiquitination in DNA double-strand break repair. Nat Commun 8(1):2039. https://doi.org/10.1038/s41467-017-02146-3
Lu H, Shamanna RA, Keijzers G, Anand R, Rasmussen LJ, Cejka P, Croteau DL, Bohr VA (2016) RECQL4 promotes DNA end resection in repair of DNA double-strand breaks. Cell Rep 16(1):161–173. https://doi.org/10.1016/j.celrep.2016.05.079
Luo X, Kraus WL (2012) On PAR with PARP: cellular stress signaling through poly(ADP-ribose) and PARP-1. Genes Dev 26(5):417–432
Macris MA, Krejci L, Bussen W, Shimamoto A, Sung P (2006) Biochemical characterization of the RECQ4 protein, mutated in Rothmund-Thomson syndrome. DNA Repair 5(2):172–180. https://doi.org/10.1016/j.dnarep.2005.09.005
Maire G, Yoshimoto M, Chilton-MacNeill S, Thorner PS, Zielenska M, Squire JA (2009) Recurrent RECQL4 imbalance and increased gene expression levels are associated with structural chromosomal instability in sporadic osteosarcoma. Neoplasia 11(3):260–268, 263p following 268
Mann MB, Hodges CA, Barnes E, Vogel H, Hassold TJ, Luo G (2005) Defective sister-chromatid cohesion, aneuploidy and cancer predisposition in a mouse model of type II Rothmund-Thomson syndrome. Hum Mol Genet 14(6):813–825. https://doi.org/10.1093/hmg/ddi075
Marino F, Vindigni A, Onesti S (2013) Bioinformatic analysis of RecQ4 helicases reveals the presence of a RQC domain and a Zn knuckle. Biophys Chem 177-178:34–39. https://doi.org/10.1016/j.bpc.2013.02.009
Martin GM, Oshima J (2000) Lessons from human progeroid syndromes. Nature 408(6809):263–266
Matsuno K, Kumano M, Kubota Y, Hashimoto Y, Takisawa H (2006) The N-terminal noncatalytic region of Xenopus RecQ4 is required for chromatin binding of DNA polymerase alpha in the initiation of DNA replication. Mol Cell Biol 26(13):4843–4852. https://doi.org/10.1128/MCB.02267-05
Mehollin-Ray AR, Kozinetz CA, Schlesinger AE, Guillerman RP, Wang LL (2008) Radiographic abnormalities in Rothmund-Thomson syndrome and genotype-phenotype correlation with RECQL4 mutation status. AJR Am J Roentgenol 191(2):W62–W66. https://doi.org/10.2214/AJR.07.3619
Mo D, Fang H, Niu K, Liu J, Wu M, Li S, Zhu T, Aleskandarany MA, Arora A, Lobo DN, Madhusudan S, Balajee AS, Chi Z, Zhao Y (2016) Human helicase RECQL4 drives cisplatin resistance in gastric cancer by activating an AKT-YB1-MDR1 signaling pathway. Cancer Res 76(10):3057–3066. https://doi.org/10.1158/0008-5472.CAN-15-2361
Mohaghegh P, Karow JK, Brosh RM Jr, Bohr VA, Hickson ID (2001) The Bloom’s and Werner’s syndrome proteins are DNA structure-specific helicases. Nucleic Acids Res 29(13):2843–2849. https://doi.org/10.1093/nar/29.13.2843
Mojumdar A, De March M, Marino F, Onesti S (2017) The human RecQ4 helicase contains a functional RecQ C-terminal Region (RQC) that is essential for activity. J Biol Chem 292(10):4176–4184. https://doi.org/10.1074/jbc.M116.767954
Monnat RJ Jr (2010) Human RECQ helicases: roles in DNA metabolism, mutagenesis and cancer biology. Semin Cancer Biol 20(5):329–339. https://doi.org/10.1016/j.semcancer.2010.10.002
Morozov V, Mushegian AR, Koonin EV, Bork P (1997) A putative nucleic acid-binding domain in Bloom’s and Werner’s syndrome helicases. Trends Biochem Sci 22(11):417–418
Nakayama K, Irino N, Nakayama H (1985) The recQ gene of Escherichia coli K12: molecular cloning and isolation of insertion mutants. Mol Gen Genet MGG 200(2):266–271. https://doi.org/10.1007/bf00425434
Narayan G, Bourdon V, Chaganti S, rias-Pulido H, Nandula SV, Rao PH, Gissmann L, Durst M, Schneider A, Pothuri B, Mansukhani M, Basso K, Chaganti RS, Murty VV (2007) Gene dosage alterations revealed by cDNA microarray analysis in cervical cancer: identification of candidate amplified and overexpressed genes. Genes Chromosomes Cancer 46(4):373–384
Ng AJ, Walia MK, Smeets MF, Mutsaers AJ, Sims NA, Purton LE, Walsh NC, Martin TJ, Walkley CR (2015) The DNA helicase recql4 is required for normal osteoblast expansion and osteosarcoma formation. PLoS Genet 11(4):e1005160. https://doi.org/10.1371/journal.pgen.1005160
Nguyen GH, Dexheimer TS, Rosenthal AS, Chu WK, Singh DK, Mosedale G, Bachrati CZ, Schultz L, Sakurai M, Savitsky P, Abu M, McHugh PJ, Bohr VA, Harris CC, Jadhav A, Gileadi O, Maloney DJ, Simeonov A, Hickson ID (2013) A small molecule inhibitor of the BLM helicase modulates chromosome stability in human cells. Chem Biol 20(1):55–62
Nguyen GH, Tang W, Robles AI, Beyer RP, Gray LT, Welsh JA, Schetter AJ, Kumamoto K, Wang XW, Hickson ID, Maizels N, Monnat RJ Jr, Harris CC (2014) Regulation of gene expression by the BLM helicase correlates with the presence of G-quadruplex DNA motifs. Proc Natl Acad Sci U S A 111(27):9905–9910. https://doi.org/10.1073/pnas.1404807111
Nishijo K, Nakayama T, Aoyama T, Okamoto T, Ishibe T, Yasura K, Shima Y, Shibata KR, Tsuboyama T, Nakamura T, Toguchida J (2004) Mutation analysis of the RECQL4 gene in sporadic osteosarcomas. Int J Cancer 111(3):367–372. https://doi.org/10.1002/ijc.20269
Ohlenschlager O, Kuhnert A, Schneider A, Haumann S, Bellstedt P, Keller H, Saluz HP, Hortschansky P, Hanel F, Grosse F, Gorlach M, Pospiech H (2012) The N-terminus of the human RecQL4 helicase is a homeodomain-like DNA interaction motif. Nucleic Acids Res 40(17):8309–8324. https://doi.org/10.1093/nar/gks591
Park SJ, Lee YJ, Beck BD, Lee SH (2006) A positive involvement of RecQL4 in UV-induced S-phase arrest. DNA Cell Biol 25(12):696–703
Patil AV, Hsieh TS (2017) Ribosomal protein S3 negatively regulates unwinding activity of RecQ like helicase 4 through their physical interaction. J Biol Chem. https://doi.org/10.1074/jbc.M116.764324
Petkovic M, Dietschy T, Freire R, Jiao R, Stagljar I (2005) The human Rothmund-Thomson syndrome gene product, RECQL4, localizes to distinct nuclear foci that coincide with proteins involved in the maintenance of genome stability. J Cell Sci 118(Pt 18):4261–4269. https://doi.org/10.1242/jcs.02556
Rossi ML, Ghosh AK, Kulikowicz T, Croteau DL, Bohr VA (2010) Conserved helicase domain of human RecQ4 is required for strand annealing-independent DNA unwinding. DNA Repair 9(7):796–804
Rothmund A (1868) Ueber cataracten in verbindung mit einer eigenthumlichen hautdegeneration. Albrecht von Graefes Arch Fur Ophth 14:159–182
Sadikovic B, Thorner P, Chilton-Macneill S, Martin JW, Cervigne NK, Squire J, Zielenska M (2010) Expression analysis of genes associated with human osteosarcoma tumors shows correlation of RUNX2 overexpression with poor response to chemotherapy. BMC Cancer 10:202. https://doi.org/10.1186/1471-2407-10-202
Sangrithi MN, Bernal JA, Madine M, Philpott A, Lee J, Dunphy WG, Venkitaraman AR (2005) Initiation of DNA replication requires the RECQL4 protein mutated in Rothmund-Thomson syndrome. Cell 121(6):887–898. https://doi.org/10.1016/j.cell.2005.05.015
Schurman SH, Hedayati M, Wang Z, Singh DK, Speina E, Zhang Y, Becker K, Macris M, Sung P, Wilson DM 3rd, Croteau DL, Bohr VA (2009) Direct and indirect roles of RECQL4 in modulating base excision repair capacity. Hum Mol Genet 18(18):3470–3483. https://doi.org/10.1093/hmg/ddp291
Sedlackova H, Cechova B, Mlcouskova J, Krejci L (2015) RECQ4 selectively recognizes Holliday junctions. DNA Repair 30:80–89. https://doi.org/10.1016/j.dnarep.2015.02.020
Shamanna RA, Singh DK, Lu H, Mirey G, Keijzers G, Salles B, Croteau DL, Bohr VA (2014) RECQ helicase RECQL4 participates in non-homologous end joining and interacts with the Ku complex. Carcinogenesis 35(11):2415–2424. https://doi.org/10.1093/carcin/bgu137
Shimamoto A, Kagawa H, Zensho K, Sera Y, Kazuki Y, Osaki M, Oshimura M, Ishigaki Y, Hamasaki K, Kodama Y, Yuasa S, Fukuda K, Hirashima K, Seimiya H, Koyama H, Shimizu T, Takemoto M, Yokote K, Goto M, Tahara H (2014) Reprogramming suppresses premature senescence phenotypes of Werner syndrome cells and maintains chromosomal stability over long-term culture. PLoS One 9(11):e112900. https://doi.org/10.1371/journal.pone.0112900
Shin G, Jeong D, Kim H, Im JS, Lee JK (2019) RecQL4 tethering on the pre-replicative complex induces unscheduled origin activation and replication stress in human cells. J Biol Chem. https://doi.org/10.1074/jbc.RA119.009996
Shinya A, Nishigori C, Moriwaki S, Takebe H, Kubota M, Ogino A, Imamura S (1993) A case of Rothmund-Thomson syndrome with reduced DNA repair capacity. Arch Dermatol 129(3):332–336
Siitonen HA, Kopra O, Kaariainen H, Haravuori H, Winter RM, Saamanen AM, Peltonen L, Kestila M (2003) Molecular defect of RAPADILINO syndrome expands the phenotype spectrum of RECQL diseases. Hum Mol Genet 12(21):2837–2844. https://doi.org/10.1093/hmg/ddg306
Siitonen HA, Sotkasiira J, Biervliet M, Benmansour A, Capri Y, Cormier-Daire V, Crandall B, Hannula-Jouppi K, Hennekam R, Herzog D, Keymolen K, Lipsanen-Nyman M, Miny P, Plon SE, Riedl S, Sarkar A, Vargas FR, Verloes A, Wang LL, Kaariainen H, Kestila M (2008) The mutation spectrum in RECQL4 diseases. Eur J Hum Genet 17(2):151–158
Singh DK, Karmakar P, Aamann M, Schurman SH, May A, Croteau DL, Burks L, Plon SE, Bohr VA (2010) The involvement of human RECQL4 in DNA double-strand break repair. Aging Cell 9(3):358–371. https://doi.org/10.1111/j.1474-9726.2010.00562.x
Singh DK, Popuri V, Kulikowicz T, Shevelev I, Ghosh AK, Ramamoorthy M, Rossi ML, Janscak P, Croteau DL, Bohr VA (2012) The human RecQ helicases BLM and RECQL4 cooperate to preserve genome stability. Nucleic Acids Res 40(14):6632–6648
Smith PJ, Paterson MC (1982) Enhanced radiosensitivity and defective DNA repair in cultured fibroblasts derived from Rothmund Thomson syndrome patients. Mutat Res 94(1):213–228
Su Y, Meador JA, Calaf GM, Proietti De-Santis L, Zhao Y, Bohr VA, Balajee AS (2010) Human RecQL4 helicase plays critical roles in prostate carcinogenesis. Cancer Res 70(22):9207–9217. https://doi.org/10.1158/0008-5472.CAN-10-1743
Suzuki T, Kohno T, Ishimi Y (2009) DNA helicase activity in purified human RECQL4 protein. J Biochem 146(3):327–335
Tanaka S, Komeda Y, Umemori T, Kubota Y, Takisawa H, Araki H (2013) Efficient initiation of DNA replication in eukaryotes requires Dpb11/TopBP1-GINS interaction. Mol Cell Biol 33(13):2614–2622
Tang W, Robles AI, Beyer RP, Gray LT, Nguyen GH, Oshima J, Maizels N, Harris CC, Monnat RJ Jr (2016) The Werner syndrome RECQ helicase targets G4 DNA in human cells to modulate transcription. Hum Mol Genet 25(10):2060–2069. https://doi.org/10.1093/hmg/ddw079
Tao J, Chen S, Lee B (2010) Alteration of Notch signaling in skeletal development and disease. Ann N Y Acad Sci 1192:257–268. https://doi.org/10.1111/j.1749-6632.2009.05307.x
Tao J, Jiang MM, Jiang L, Salvo JS, Zeng HC, Dawson B, Bertin TK, Rao PH, Chen R, Donehower LA, Gannon F, Lee BH (2014) Notch activation as a driver of osteogenic sarcoma. Cancer Cell 26(3):390–401. https://doi.org/10.1016/j.ccr.2014.07.023
Taylor WB (1957) Rothmund’s syndrome; Thomson’s syndrome; congenital poikiloderma with or without juvenile cataracts. AMA Arch Derm 75(2):236–244
Thangavel S, Mendoza-Maldonado R, Tissino E, Sidorova JM, Yin J, Wang W, Monnat RJ Jr, Falaschi A, Vindigni A (2010) Human RECQ1 and RECQ4 helicases play distinct roles in DNA replication initiation. Mol Cell Biol 30(6):1382–1396. https://doi.org/10.1128/MCB.01290-09
Thomson MS (1923) An hitherto undescribed familial disease. Br J Dermatol Syphilis 35:455–462
Tuteja N, Tuteja R (2004) Prokaryotic and eukaryotic DNA helicases. Essential molecular motor proteins for cellular machinery. Eur J Biochem 271(10):1835–1848. https://doi.org/10.1111/j.1432-1033.2004.04093.x
Van Maldergem L, Piard J, Larizza L, Wang LL (2016) RECQL4-related recessive conditions. In: Erickson RP and Winshaw-Boris A (eds) Epstein’s Inborn Errors of Development: The Molecular Basis of Clinical Disorders of Morphogenesis (3 ed.) Oxford University Press, USA.
Van Maldergem L, Siitonen HA, Jalkh N, Chouery E, De Roy M, Delague V, Muenke M, Jabs EW, Cai J, Wang LL, Plon SE, Fourneau C, Kestila M, Gillerot Y, Megarbane A, Verloes A (2006) Revisiting the craniosynostosis-radial ray hypoplasia association: Baller-Gerold syndrome caused by mutations in the RECQL4 gene. J Med Genet 43(2):148–152. https://doi.org/10.1136/jmg.2005.031781
Vijayakumar S, Liu G, Rus IA, Yao S, Chen Y, Akiri G, Grumolato L, Aaronson SA (2011) High-frequency canonical Wnt activation in multiple sarcoma subtypes drives proliferation through a TCF/beta-catenin target gene, CDC25A. Cancer Cell 19(5):601–612
Wang H, Elledge SJ (1999) DRC1, DNA replication and checkpoint protein 1, functions with DPB11 to control DNA replication and the S-phase checkpoint in Saccharomyces cerevisiae. Proc Natl Acad Sci U S A 96(7):3824–3829
Wang JT, Xu X, Alontaga AY, Chen Y, Liu Y (2014) Impaired p32 regulation caused by the lymphoma-prone RECQ4 mutation drives mitochondrial dysfunction. Cell Rep 7(3):848–858. https://doi.org/10.1016/j.celrep.2014.03.037
Wang LL, Gannavarapu A, Kozinetz CA, Levy ML, Lewis RA, Chintagumpala MM, Ruiz-Maldanado R, Contreras-Ruiz J, Cunniff C, Erickson RP, Lev D, Rogers M, Zackai EH, Plon SE (2003) Association between osteosarcoma and deleterious mutations in the RECQL4 gene in Rothmund-Thomson syndrome. J Natl Cancer Inst 95(9):669–674
Wang LL, Levy ML, Lewis RA, Chintagumpala MM, Lev D, Rogers M, Plon SE (2001) Clinical manifestations in a cohort of 41 Rothmund-Thomson syndrome patients. Am J Med Genet 102(1):11–17
Wang S, Liu Z, Ye Y, Li B, Liu T, Zhang W, Liu GH, Zhang YA, Qu J, Xu D, Chen Z (2018) Ectopic hTERT expression facilitates reprograming of fibroblasts derived from patients with Werner syndrome as a WS cellular model. Cell Death Dis 9(9):923. https://doi.org/10.1038/s41419-018-0948-4
Werner SR, Prahalad AK, Yang J, Hock JM (2006) RECQL4-deficient cells are hypersensitive to oxidative stress/damage: insights for osteosarcoma prevalence and heterogeneity in Rothmund-Thomson syndrome. Biochem Biophys Res Commun 345(1):403–409. https://doi.org/10.1016/j.bbrc.2006.04.093
Woo LL, Futami K, Shimamoto A, Furuichi Y, Frank KM (2006) The Rothmund-Thomson gene product RECQL4 localizes to the nucleolus in response to oxidative stress. Exp Cell Res 312(17):3443–3457. https://doi.org/10.1016/j.yexcr.2006.07.023
Wu J, Capp C, Feng L, Hsieh TS (2008) Drosophila homologue of the Rothmund-Thomson syndrome gene: essential function in DNA replication during development. Dev Biol 323(1):130–142. https://doi.org/10.1016/j.ydbio.2008.08.006
Wu J, Zhi L, Dai X, Cai Q, Ma W (2015) Decreased RECQL5 correlated with disease progression of osteosarcoma. Biochem Biophys Res Commun 467(4):617–622. https://doi.org/10.1016/j.bbrc.2015.10.114
Xia J, Chen LT, Mei Q, Ma CH, Halliday JA, Lin HY, Magnan D, Pribis JP, Fitzgerald DM, Hamilton HM, Richters M, Nehring RB, Shen X, Li L, Bates D, Hastings PJ, Herman C, Jayaram M, Rosenberg SM (2016) Holliday junction trap shows how cells use recombination and a junction-guardian role of RecQ helicase. Sci Adv 2(11):e1601605. https://doi.org/10.1126/sciadv.1601605
Xia J, Mei Q, Rosenberg SM (2019) Tools to live by: bacterial DNA structures illuminate Cancer. Trends Genet 35(5):383–395. https://doi.org/10.1016/j.tig.2019.03.001
Xu L, Li S, Stohr BA (2013) The role of telomere biology in cancer. Annu Rev Pathol 8:49–78
Xu X, Liu Y (2009) Dual DNA unwinding activities of the Rothmund-Thomson syndrome protein, RECQ4. EMBO J 28(5):568–577. https://doi.org/10.1038/emboj.2009.13
Xu X, Rochette PJ, Feyissa EA, Su TV, Liu Y (2009a) MCM10 mediates RECQ4 association with MCM2-7 helicase complex during DNA replication. EMBO J 28(19):3005–3014. https://doi.org/10.1038/emboj.2009.235
Xu Y, Lei Z, Huang H, Dui W, Liang X, Ma J, Jiao R (2009b) dRecQ4 is required for DNA synthesis and essential for cell proliferation in Drosophila. PLoS One 4(7):e6107
Yin J, Kwon YT, Varshavsky A, Wang W (2004) RECQL4, mutated in the Rothmund-Thomson and RAPADILINO syndromes, interacts with ubiquitin ligases UBR1 and UBR2 of the N-end rule pathway. Hum Mol Genet 13(20):2421–2430. https://doi.org/10.1093/hmg/ddh269
Yokoyama H, Moreno-Andres D, Astrinidis SA, Hao Y, Weberruss M, Schellhaus AK, Lue H, Haramoto Y, Gruss OJ, Antonin W (2019) Chromosome alignment maintenance requires the MAP RECQL4, mutated in the Rothmund-Thomson syndrome. Life Sci Alliance 2(1). https://doi.org/10.26508/lsa.201800120
Yong ZW, Zaini ZM, Kallarakkal TG, Karen-Ng LP, Rahman ZA, Ismail SM, Sharifah NA, Mustafa WM, Abraham MT, Tay KK, Zain RB (2014) Genetic alterations of chromosome 8 genes in oral cancer. Sci Rep 4:6073. https://doi.org/10.1038/srep06073
Yu CE, Oshima J, Fu YH, Wijsman EM, Hisama F, Alisch R, Matthews S, Nakura J, Miki T, Ouais S, Martin GM, Mulligan J, Schellenberg GD (1996) Positional cloning of the Werner’s syndrome gene. Science 272(5259):258–262. https://doi.org/10.1126/science.272.5259.258
Zhang W, Li J, Suzuki K, Qu J, Wang P, Zhou J, Liu X, Ren R, Xu X, Ocampo A, Yuan T, Yang J, Li Y, Shi L, Guan D, Pan H, Duan S, Ding Z, Li M, Yi F, Bai R, Wang Y, Chen C, Yang F, Li X, Wang Z, Aizawa E, Goebl A, Soligalla RD, Reddy P, Esteban CR, Tang F, Liu GH, Belmonte JC (2015) Aging stem cells. A Werner syndrome stem cell model unveils heterochromatin alterations as a driver of human aging. Science 348(6239):1160–1163. https://doi.org/10.1126/science.aaa1356
Zhi LQ, Ma W, Zhang H, Zeng SX, Chen B (2014) Association of RECQL5 gene polymorphisms and osteosarcoma in a Chinese Han population. Tumour Biol 35(4):3255–3259. https://doi.org/10.1007/s13277-013-1425-4
Huang P, Pryde FE, Lester D, Maddison RL, Borts RH, Hickson ID, Louis EJ (2001) SGS1 is required for telomere elongation in the absence of telomerase. Curr Biol 11 (2):125-129. https://doi.org/10.1016/S0960-9822(01)00021-5
Maciaszek JL, Oak N, Chen W, Hamilton KV, McGee RB, Nuccio R, Mostafavi R, Hines-Dowell S, Harrison L, Taylor L, Gerhardt EL, Ouma A, Edmonson MN, Patel A, Nakitandwe J, Pappo AS, Azzato EM, Shurtleff SA, Ellison DW, Downing JR, Hudson MM, Robison LL, Santana V, Newman S, Zhang J, Wang Z, Wu G, Nichols KE, Kesserwan CA (2019) Enrichment of heterozygous germline RECQL4 loss-of-function variants in pediatric osteosarcoma. Cold Spring Harb Mol Case Stud 5(5):a004218. https://doi.org/10.1101/mcs.a004218
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Lu, L., Jin, W., Wang, L.L. (2020). RECQ DNA Helicases and Osteosarcoma. In: Kleinerman, E., Gorlick, R. (eds) Current Advances in the Science of Osteosarcoma. Advances in Experimental Medicine and Biology, vol 1258. Springer, Cham. https://doi.org/10.1007/978-3-030-43085-6_3
Download citation
DOI: https://doi.org/10.1007/978-3-030-43085-6_3
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-43084-9
Online ISBN: 978-3-030-43085-6
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)