Abstract
Fusarium wilt caused by Fusarium oxysporum f. sp. ciceris (FOC) is a widespread disease of chickpea that adversely affects the crop. The pathogen is more diverse in its pathogenicity. The more severe effect of FOC on yield and it varied from 10% to 100% depending upon the environmental conditions and host susceptibility. The main effects were transience in young seedlings, color differentiation (brown to black), and leaf senescence. Characterization and screening of high-yielding varieties for better remediation of the disease are needed. Chemical control is costly and not adequate to manage the crop. Moreover, the use of biocontrol agents, i.e., Burkholderia, Trichoderma, Pseudomonas, and Bacillus, also limits the pathogenetic effect substantial and best supernumerary against chemical control. Thus, plant omics technologies are needed for screening and monitoring of Fusarium sp. growth and improved the efficiency of disease management. Identification of new genes responsible for slow wilting and introducing into chickpea crops using integrated management protocols. The substantial development has ended the extrication of genetics for race-specific confrontation. The roots are directly in contact with these organisms while pathogen determination from the root zone is extremely less. So, there is a dire need for identification of transcription factors that play an important role to limit the pathogen activity participated in Fusarium oxysporum activity in the soil as well as chickpea. Functional genomics studies would play a significant role to help better understanding of pathogen chickpea interaction and play an auspicious role in resistance development against legume wilt.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
4.1 Introduction
Chickpea is an important founder of crops in agriculture, having diploid (2n = 16) chromosome number. It belongs to legumes and papilionoid (subfamily) from its wild Cajanus reticulatus ancestor present in Turkish Kurdistan dating back (8000–9000) years (Lev-Yadun et al. 2000) and considered a major source of human food due to the presence of lysine-rich protein. It is an important legume and pulse crop in the world having 41–50.8% carbohydrates, 3–6% oil, 17–24% protein, and considerable amount of other minerals like phosphorus, magnesium, calcium, potassium, iron, zinc, and manganese. Chickpea also plays an important role as an alternate rotation crop followed by cereals and manages soil fertility and productivity by improving the N fertilization (nitrogen-fixing ability) from the atmosphere (Jiménez Díaz et al. 2015). Over the past few years, it is stated that chickpea productivity has been marginal decreases due to the effect of biotic factors (Fusarium wilt and pod borer) and abiotic factors. Reducing the pressure of these factors (biotic and abiotic) is important to increase production. Chickpea ranked second among the important food legume crops in tropical, subtropical, South, and West Asia. Overall, about 1.35 × 107 ha of chickpea are growing and yield about 1.31 × 107 in more than 50 countries. Chickpea is used not as a valuable crop for export in developed countries but a good source of protein supplement in cereal-based diets in developing countries. Chickpea is generally grown under the rainfed condition and depends on available soil water showing drought tolerance over the year. Fusarium oxysporum f. sp. ciceris (FOC ) affects the chickpea crop by inducing wilt disease, more damaging worldwide for their occurrence, and accounts 90% annual yield losses worldwide. The disease was first reported by Butler in 1918, but etiology was not confirmed until 1940 and later was spread in Americas, Europe, and Africa but not reported in Australia. Fusarium wilt has become a limiting factor for chickpea production in the Mediterranean basin, the Indian subcontinent, and America. The most important symptoms of wilt, i.e., the patch in group form and occurs at any stage and spread across a field (Haware 1990). The main reason for Fusarium wilt is soilborne pathogen and observing signs like delaying crown, leaf anomalies, and rolled brown leaves. The number of strains is unknown to the soilborne pathogen and is difficult to control without solid information and identification of the pathogen (Cha et al. 2016).
The susceptible varieties showed symptoms in 25 days after sowing such as including flabbiness in leaves tailed by a dull green streak, dehydration, and downfall of the plant. Though disease marks are commonly more visible at the initiation of flowering for 6–8 weeks, in some studies, it is reported that it appeared at the podding stage. The leaves dropping has occurred in the upper part of the plant, but within a few days, it ensures on the whole plant. In partial wilt, few branches were affected initially, but later roots of affected material affect the nearby plants. In partial wilting, no color discoloration was recorded visually. In general, symptoms of the disease occur at any stage of plant growth (Jiménez Díaz et al. 2015) while more visible at the early stage of flowering and appears at the podding stage (late wilt). Late wilted plants exhibited falling of petioles, rachis, and leaflets as well as necrosis and discoloration of foliage (Jiménez Díaz et al. 2015). Early Fusarium wilt affects more than late wilting. However, late wilted plants produce lighter, rougher, and duller seeds as compared to normal (Haware and Nene 1980; Navas-Cortés et al. 2000). If the cross-sectional study was done on the affected plant, a dark brown color discoloration was observed in xylem tissues. The discoloration was also recorded in vascular tissues of roots as well as in stems. The symptoms were also recorded as cavity formation among xylem and phloem, medulla and cortical parenchyma, and cell proliferation in vascular cambium.
During the defense mechanism, the plant uses many molecular signals or protein receptors to know the presence of microbes . Two modes of pathogen recognition used by the host, i.e., effector-triggered immunity (ETI) and pathogen-triggered immunity (PTI) . The invariant epitope types are called microbe-associated molecular patterns (MAMPs) and are composed of flagellin, chitin, and lipopolysaccharides that help spread the disease. Moreover, pathogen-induced danger-associated molecular patterns are composed of fructans, callose, and glucans. As a result, host secretes effector R protein domains have nibblers act as PTI. Studies also reported that the sensing of bacteria produce siderophores and fungi serve as MAMPs and hydroxyproline and rapid alkalization factors, but their role was not clear yet in defense mechanism. The current has described the chickpea Fusarium wilt etiology, occurrence, and management practices including the most recent molecular breeding, high-throughput sequencing techniques, as well as identification of transcription factors that could favor the crop and enhance the tolerance mechanism to control the disease.
4.2 Casual Organism and Symptoms
It is caused by Fusarium oxysporum f. sp. ciceris [Fusarium oxysporum Schlecht. f. sp. concerns (Padw.) Matuo & Sato] (Jimenez-Fernandez et al. 2011; Haware 1990). The aerial mycelium in the first appearance was whitish and cotton, on potato sucrose agar, potato dextrose agar and under UV light, but turn into salmon in color and some cases, remain white (Jimenez Diaz et al. 2011). Fusarium wilt of chickpea produces microconidia, macroconidia, and chlamydospores. The microconidia are elliptical or tubular and straight. Macroconidia are thinner than microconidia and typically 3–5 septate or fusoid, while chlamydospores are produced in 15-day-old cultures and infected chickpea tissues, smooth or rough-walled (Castro et al. 2012; Jimenez Diaz et al. 2011). Maximum sporulation was recorded at pH ranges from 7.1 to 7.9 (Jimenez Diaz et al. 2011). Hyphae are septate and split abundantly. Optimum growth was recorded at 25–27 °C and pH 5.1–5.9 and liable on strains.
4.3 Epidemiology
The severity of the chickpea wilt is depending upon the pathogen, genotypes, pathogenic races, inoculum density, environmental condition, and cultivar sensitivity. The activity of the wilting disease was triggered by a combination of pathogen activities. It includes fungus mycelium in the xylem that produced contaminant components that affect host defense response, production of gels, teloses, and vessel crushing by the propagation of linked parenchyma cells (Beckman 1987). The mycelium might survive as a pathogen in seed, soil and toxic residues (crop), roots, and stem tissue concealed in the soil for more than 6 years or even in absence of host (Singh et al. 2008). Dicotyledonous weeds that don’t show the symptoms but have the infection that could enhance the pathogen activity and survived in fallow soils. Moreover, infected soil is an important source of primary inoculum for the development of Fusarium wilt (Al-taae et al. 2013). The transmission can also be done by the seed and can survive in plant debris as well as in the soil. Moreover, it also observed that fungus chlamydospore was present in soil freely (Haware et al. 1996), seed hilum (Haware et al. 1978), and cotyledon axis (Shakir and Mirza 1994). Chlamydospores or mycelia are the main and basic sources of infection, even the conidia of the fungus are short-lived, while chlamydospores can remain feasible up to the next available crop in the field (Chand and Khirbat 2009). Chlamydospore production is contingent on the nutrient availability of the inoculum. Fungal inoculum may be exposed to lower nutrient levels in the field condition as compared to grow under well-fed macroconidia form under agar media (Schippers and Van Eck 1981). The pathogen grows very well in roots and stems in apparently looking good condition but concealing adequate fungus (Trapero Casas and Jimenez Diaz 1985). The pathogen remains dormant until triggered to germinate when carbohydrate is released from decaying tissue or roots, present in the form of chlamydospores (Schippers and Van Eck 1981). The provocation for germination could be the host or non-host plant roots or plant wreckage (Nelson 2012), after the germination of chlamydospores, conidia, hyphae, and new chlamydospores is formed. After conidia and hyphae production, thallus formation took place and leads to chlamydospore production in 2–3 days if suitable condition prevailed (Beckman and Roberts 1995). By penetration of the epidermal cells, attack on the roots occurs on the host or non-host plants (Beckman and Roberts 1995) and caused vascular disease (Stover 1970). The infiltration occurs directly or by wounds (Nelson 2012), the common sites for infiltration are the root tip of both tap and lateral roots (Lucas 1998). The infiltration is stopped by different factors, such as fungal compounds, and inhibits the spore formation (fungal), plant surface structures, and germ tube production (Mendgen et al. 1996). The more adverse form is, mycelium moved through intercellular root cortex and finally reaches to xylem vessels during colonization and remains within the xylem vessels and colonize in the host (Bishopt and Cooper 1983).
4.4 Breeding
The Fusarium wilt activity can be reduced in the host using breeding approaches in chickpea crop. Breeding approaches involved availability of genetic diversity considered the most important step for a breeding program, wild relatives, and selection of desirable plant for trait and disease resistance and evaluate the plant for commercial production (Salimath et al. 2007). As chickpea is a self-pollinated crop, it requires genes to fix the breeding problems by pure lines development. Initial screening was done by mass or pure line selection and later crossing programs and alteration in pedigree and bulk methods were employed for segregating generation (Gaur et al. 2012; Millan et al. 2015). In the intraspecific hybrid program, the single cross method was used in desi and Kabuli chickpea genotypes with variant genetic history (Berrada et al. 2007). Parents from desi varieties have been used for gene transfer in Kabuli varieties against Fusarium wilt resistance, as parents from Kabuli parents are used to improve large size seed and seed quality in desi variety (Gaur et al. 2007). The breeding development efforts were also made for interspecific crosses and enhance genetic diversity and interrogate useful genes from wild cicer into cultivated spp. The FOC resistance has been recognized from desi germplasm as well as in wild Cicer spp. (Kaiser et al. 1994). For genetic gains enhancement, there is a need precise and efficient selection of segregating populations (Gaur et al. 2012). For successful wilt, sick breeding programs hot spot location, field, greenhouse and laboratory methods have been used for the selection of resistance varieties (Gaur et al. 2007). It has been reported about 5174 Kabuli genotypes were screened against Fusarium wilt resistance at ICARDA, and about 110 genotypes were recognized as resistant. Fusarium wilt resistance depends upon monogenic or oligogenic depending upon the resistance resource (Sharma and Muehlbauer 2007; Upadhyaya et al. 1983; Sharma et al. 2005). It is also reported that FOC genetic resistance cultivar contains three independent genes (h1, h2, and h3) (Singh et al. 2014). Moreover, it is also suggested that late wilting was controlled by the presence of any one gene nut combination of two genes confirm the wilt resistance in chickpea (Castro et al. 2012; Jiménez Díaz et al. 2015). Similar results also stated that resistance was confirmed by the presence of these genes in the combine or individual form (Tullu et al. 1999). Some ICARDA lines, i.e., WR-315, CA-1938, and CA2139, contain these genes (Halila et al. 2009; Rubio et al. 2003). However, the genetic of resistance for some chickpea races like 1B/C and 6 is still unknown.
4.5 Genetic and Pathogen
The first name of the fungus was Fusarium orthoceras apple and swollen. var. cicerone by Padwick and modified by Chattopadhyay and Sen Gupta and was renamed as F. oxysporum Schl. f. sp. ciceri (Padwick) Snyder and Hansen. Fusarium oxysporum is among the monophyletic origin in the Fusarium oxysporum complex of the gibberella clade and considered as polyphyletic and currently known as Fusarium oxysporum (Schlechtend. Fr.) f. sp. ciceris (Padwick) Matuo & K. Sato. Fusarium is the only pathogen in Cicer sp. (Kaiser et al. 1994), and oxysporum is an attack on root tissue in faba bean, lentil, and pea and recorded as symptomless carters for the pathogen (Trapero Casas and Jimenez Diaz 1985).Yellow or wilting syndromes along with brown discoloration were recognized based on two pathotypes and induce in sensitive chickpeas. The recorded symptoms are considered slow, foliar yellowing and death of plant at a later stage while wilting is considered reckless, adverse chlorosis, flabbiness, and plant death during an early stage of growth (Trapero Casas and Jimenez Diaz 1985).The susceptibility of the pathogen depends upon the races and efficient use of available resources for the chickpea breeding program. The identification of the races against pathogens is simple but depends upon the cost, available resources, and facilities. So, there is a dire need to develop new methods that are more rapid and effective, and reproducible identification of pathogen and races is used to determine the diversity and resistance among the genotypes. Polymerase chain reaction (PCR)-based molecular markers have been used to determine the Fusarium oxysporum f. sp. ciceris and its related pathogen races identified by the method developed by Jiménez-Gasco et al. (2001).
The screening and legacy of the gene of interest (GOI) and traits are possible now with the development of marker-assisted selection (MAS) and provide beneficial information to exploit the genes useful for agronomic traits (Allahverdipoor et al. 2011). Molecular markers are an important tool for identification, characterizing, and screening and determine the diversity among the pathogens and diseases. Commonly, internal transcribed spacer (ITS) markers are used for classification and screening of the fungi (White et al. 1990), while ITS data is not enough for complex identification and diverse gene information; therefore, it is not suitable for genetic diversity or characterization of fungus. The Fusarium genus is improbable as compared to the genetic study of F. oxysporum f. sp. fragariae has not yet been reported. Among the various available technologies, restriction fragment length polymorphism (RFLP) markers are important rDNA used to determine the genetic diversity of plant pathogenic fungi. It is also used to group the isolated strains with low cost (Kachuei et al. 2015). Based on symptoms, the two pathogens were genetically distinguished by random amplified polymorphic DNA markers (RAPD) and sequence characterized amplified region (SCAR) . The specific Fusarium assays were successfully characterized using RAPD and SCAR molecular markers. Another study stated that evaluation and screening of resistant wilt lines were done against Fusarium by using RAPD and SSR molecular markers. The results represent that about 70% cultivars were resistant to disease while 30% showed susceptibility for wilt response. SSR marker (TA194) recorded an 85% probability locus at wilt resistant among the total primer used, and it was later reconfirmed by the receiver operating characteristic curve (Ahmad et al. 2014). Gowda et al. (2009) former designing the linkage map for FOC 1–5 gene resistance races with SSR and RAPD in recombinant inbred lines (RILS) developed by sensitive and resistant parents. About eight races were recognized as the specific fungus, out of which six are more infectious (Jimenez-Diaz et al. 1993). Introgression of Ascochyta blight resistance with double podding traits in chickpea was confirmed by marker-assisted backcrossing. SSR markers are used in separate backcross generation to assist in selection against the resistance of Fusarium (Varshney et al. 2014). SCAR markers are used for Ascochyta blight resistance to determine the QTLs in chickpea, and respected QTLs were identified, i.e., SCY17590 and SCAE19336, tightly linked with Ascochyta blight resistance gene at QTLAR2 location (Iruela et al. 2006) and later on successfully used for tagging in chickpea resistance lines for germplasm collection (Imtiaz et al. 2008). Combinations of SCAR with a codominant marker (CaETR) linked with QTLAR1 for Ascochyta tagging and help to identify the resistance alleles from a core collection of resistant cultivars (Madrid et al. 2013). Near-isogenic lines (NILs) were developed by using STMS markers that are tightly linked FOC 5 and FOC 01 for the selection of susceptible genotypes and resistant genotypes in LG2 and LG5 (Castro et al. 2010). Moreover, NILs are used as a valuable tool for mapping, refining the target region and selection of the desired gene for resistance to foc0 (Jendoubi et al. 2016). Jendoubi et al. (2016) reported that the results obtained from the population were useful for position refining of the target area involved in resistance mechanisms. Similar results were obtained by Ali et al. (2015) that identify the target regions associated with growth habit and double-podding base morphological position-based markers that are used in chickpea.
4.6 Integrated Genomic Approaches
The identification and construction of the genetic map of the segregating population is the foremost objective of the breeders. Efforts have been made to construct the genetic map using molecular markers for tagging traits and site-specific gene of interest in chickpea (Millan et al. 2010; Millan et al. 2015). The first maps were constructed using the isozymes F2 population from interspecific crosses (Gaur and Slinkard 1990). Many researchers reported identified genes regarding flower color, wilt resistance (Fusarium), double pod, and growth habit (Gaur and Slinkard 1990; Kazan et al. 1993; Cobos et al. 2005), and other agronomic characters and Ascochyta blight resistance linked QTLs were identified on these maps (Lichtenzveig et al. 2006). The larger numbers of maps were derived from crosses with C. reticulatum as well as many markers identification related to specific traits. However, the populations derived from interspecific crosses were made due to microsatellite markers and exploit more genetic polymorphisms among the chickpea genotypes (Cobos et al. 2007). The first transcriptome study for the chickpea genome was done with the advancement of next-generation sequencing (Hiremath et al. 2011). With the advancement of transcriptome information, detail genetic maps were made using large-scale molecular markers (Hiremath et al. 2012; Thudi et al. 2011). The availability of the draft genome sequencing in desi and Kabuli varieties would also facilitate the genetic population used for mapping and positioning of the QTLs in chickpea genome (Ali et al. 2015). Omics approaches gathered genomic information and triggered molecular markers development of tightly linked QTLs (Kumar et al. 2011).
4.7 Transcription Factors
Recent advances in molecular plant sciences boost the knowledge, and transcriptomic emerged as a powerful method to understand differential genic response over specific time-bound fashion. Transcriptomic is the techniques used to study the whole set of RNA transcripts (coding and non-coding) of a cell at a specific time and conditions. Expression analysis of tissue under different growth conditions reveals the regulatory network of the responsive gene for that specific stage or conditions, it could also help to annotate those genes which were previously unannotated due to lack of information. TF has the function to regulate the cell development, differentiation, and growth by tagging specific site with DNA or multiple sites and triggered the activation or repression of the TF through various mechanism and interaction, i.e., DNA-protein, protein-protein, and alteration in chromatin structure (Kusuya et al. 2018). The soilborne fungus is a causal agent of chickpea wilt disease. The infection includes root identification, colonization, penetration, adhesion, and penetration of the root cortex, and hyphal proliferation within the xylem vessels are controlled by transcription factors (TFs) . Transcriptome analysis based on RFLP and RAPD-based cDNA techniques were used and identified many defense-related genes in chickpea (Gurjar et al. 2012). Moreover, next-generation sequencing identified microRNA responsive genes regulating plant development and pathogen growth depending on target genes (Kohli et al. 2014). Fusarium spp. produced about 50 unique types of secondary metabolites, i.e., growth regulators, pigments, and mycotoxins, that are important for feed and food concerns. TFs have been shown to manage the mycotoxin biosynthesis compound that is favorable for other pathogenic Fusarium species (Brown et al. 2014).
Identification of FolCZF1 encoded for (C2H2) transcription factor. It is also known to affect pathogenicity in wheat (F. graminearum) and rice (Magnaporthe oryzae). The critical role of gene FolCZF1 is to produce fusaric acid and regulate the expression of fusaric acid biosynthesis. Fusaric acid (FA) taking part in the severity of Fusarium diseases, i.e., damping off, vascular wilt, and root rot (Ding et al. 2015). Fusaric acid is linked with vascular wilt symptoms caused by F. oxysporum; some transcription factors are involved in the regulation of virulence and FA biosynthesis. FolCZF1 affects the FA and influence the virulence (Yun et al. 2019). Moreover, FolCZF1 is also reported that it requires secondary metabolism and early host infection (Yun et al. 2019). Zinc finger proteins (C2H2) are widely studied in filamentous fungi.
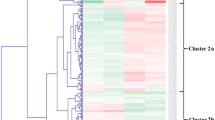
A similar study was conducted to determine the molecular basis of wilt disease in chickpea by comparing the analysis of the transcriptome of resistant and susceptible wilt cultivars under Fusarium oxysporum f. sp. ciceri and controlled condition . Analysis results stated that novel genes with differential or unique expression causative to lignification, hormonal balance, plant defense signaling, and ROS. Moreover, the study also provides information about the functional characterization of the genes involved in resistance mechanism and their use in a breeding program against wilt resistance and tolerance mechanism as well as target pathogen identification for the facilitation of the development of novel control management strategy (Upasani et al. 2017). Microscopic, proteomic, and metabolic approaches are also used to characterize the chickpea cultivars under Fusarium oxysporum interaction. The resulting expression at the microscopic level stated that differential colonization of FOC was present in susceptible and resistant genotypes. It is also reported that resistant host severely restricted the pathogen growth while opposite results were observed in susceptible cultivars. Moreover, proteomics and metabolomics results notified that the upregulation of several metabolic pathways was observed in resistant genotypes (Kumar et al. 2015; Kumar et al. 2016; Upasani et al. 2016).
ROS played an important role in recognized insight and defense signaling, but their redox relation in plant is still unknown for the defensive network. A study was conducted to determine the role of FOC 1 by inducing redox-responsive transcript for regulating defense signaling in chickpea. Microscopic studies emphasized invasion and colonization along with tissue damage and confession of degraded products at the xylem vessels in diseased roots area. Due to confession clogging of the xylem vessels incompatible hosts while resistant plant not. Assays related to lipid peroxidation represent membrane injury, and other remarkable changes were recorded such as cell shrinkage and gradual nuclear depression in fungal ingress. Moreover, qPCR results showed expression of redox regulators, cellular transport, and transcription factors in FOC 1 analysis. Functional analysis results stated that respiratory homolog, vacuolar sorting receptors, and zinc finger domain TF provide deep insight regarding the complex structure of wilt disease defense mechanism in chickpea as well as other legume crops (Gupta et al. 2013). The study also reported that chickpea transcript is used for involvement to regulate the redox state when infection occurs due to FOC 1 races (Gupta et al. 2009; Ashraf et al. 2009; Gupta et al. 2010; Garcia-Limones et al. 2002). Moreover, it is also reported that modification in the RBOH recorded regulatory role during an invasion in resistant plants while sensitive plants do not show similar variation. The other modification in OCP and FSD has a role in ROS signaling and OCP considered as ABA-dependent TF regulator, recorded down regulation in Arabidopsis thaliana (62). Also reported that cationic peroxidase has the function to accumulate in the xylem vessels in rice plants.
Genome-wide analysis of chickpea genotypes against Fusarium oxysporum was done and transcriptome study conducted by illumining technology at conidial germination stage at variant points. The results revealed that; genes linked to fungal developments are transcribed at consecutive ways were discovered. It was also reported that genes related to secret effectors, cell wall degrading, metabolism, peptidases, and transporters-related enzymes were determined at the germination stage of conidial growth. Moreover, metabolism genes are upregulated at germination, while secondary metabolites and transporters genes were upregulated at a later stage (Sharma et al. 2016). The root structure and colonization (hypocotyl) and their expression profiling in infected genotypes and plant response factors were determined using two Fusarium oxysporum. The results revealed that less colonization in xylem vessels was recorded in weekly infected genotypes. After the analysis of virulent genes, the expression profiling results represent that two genes (SIX1, SIX6) include TF (FTF1) were upregulated in root crown and hypocotyl. Both strains performed differently, the virulent strain showed strong transcription in PR1 gene while other strains respond to ethyne factor ERF2 (Niño-Sánchez et al. 2015).
In general plant colonization by fungal vascular wilt pathogens after invasion colonization was done in cortical cells, latterly hyphae intercellularly move toward vascular parenchyma cells and occupied xylem vessels. Once reached to xylem, mycelium is restricted in the vessels; as a result necrosis occurs in host tissue for general colonization (Yadeta and Thomma 2013). Ma et al. (2010) also reported that Fusarium oxysporum-specific sequences present in replaceable chromosomal position are the basis of host specialization and polyphyletic origins of most formae specials.
4.8 Exclusion and Eradication of the Pathogen
The exclusion and eradication of the pathogen is the basic paradigm for crop improvement programs. For this purpose, integrated approaches have been used to exclude and eradicate crop diseases, pests, and weeds. Though disease control by the integrated management approach is no cure for plant disease control, it is considered as an ecology approach by which different disease control measures are adopting such as pathogen-free planting material, avoiding planting in high-risk soil, exclusion and eradication of F. oxysporum inoculum from rhizosphere, and using of biocontrol measures for healthy planting materials. It is transmitted through virulent seeds and plant residues (Jimenez Diaz et al. 2011; Nelson et al. 1981), infected materials than propagating into pathogen-free soils. For this purpose, strict legislation and inspection of the seeds material and planting area and optimize the use of FOC spp. in the non-virulent area (Jimenez Diaz et al. 2011). For quantification, evaluation, inspection, and legislation of the quarantine measurement, Jiménez-Fernández et al. (2011) established a qPCR protocol that permits to measure the DNA quantity in root and stems from infected asymptomatic chickpea. Seed dressing with Benlate could be used to remove seed borne inoculum (Haware et al. 1978).
Soil having problems of Fusarium oxysporum can be reclaimed by reducing or lessening the initial inoculum or reducing the disease potential (Passari et al. 2017; Jimenez Diaz et al. 2011), and this can be achieved by various methods, i.e., biological, physical, and chemical means. A most important method is soil solarization, and Fusarium wilt can be controlled in many crops in this way (Stapleton and de Vay 1986). By solarization, pathogen not only kills but also weakens and reduces the severity and increases the availability of other components in soil microbiota (Strange 2003). Moreover, soil pathogen can also be controlled by flooding (Strange 2003), by removing the plant residue from wilt affected crop, by killing the FOC chlamydospore, and by limiting the severity of the disease for the next crop (Jiménez Díaz et al. 2015). During biological control, use bio-agents to reduce the pathogen activity by making colonization in the rhizosphere while no toxic residue remains in the soil (Dubey et al. 2007). Trichoderma has been used against Fusarium wilt in greenhouse and field condition and gives tremendous result to control the disease (Kaur and Mukhopadhyay 1992).
Moreover, the application of Pseudomonas restricted the FOC in vitro and allowed significant growth in shoot length, dry weight, and yield (Nautiyal 1997). Application of nonpathogenic type strains such as Bacillus sp. and Pseudomonas recorded a significant reduction in the severity of Fusarium oxysporum f. sp. ciceris (Nautiyal 1997). Another practice could also reduce the severity of plant pathogen effect on the chickpea crop. An adequate amount of cultural practices takes the benefit of Fusarium management. A study reported that Fusarium can live about 6 years and 3 years of crop rotation but is not effective to reduce the effect of Fusarium incidence (Haware et al. 1996). Moreover, widespread disease development is due to the sowing date (Navas-Cortés et al. 1998); sowing chickpea from early spring to early winter could slow the Fusarium wilt development and ultimately enhance the yield (Landa et al. 2004). Along with the sowing date, the use of resistant cultivars also appears to be benefitted to control the wilt disease. Resistant varieties played an important role in an integrated disease management program (Landa et al. 2004; Jimenez Diaz et al. 2011; Jiménez Díaz et al. 2015). Resistant desi genotypes have been identified against FOC that reduced the disease incidence in wild and desi chickpea varieties (Jiménez Díaz et al. 2015). The availability of high genetic diversity in pathogenicity reduces the effectiveness and extensive use of present resistance (Bayraktar and Dolar 2012).
4.9 Conclusion
Fusarium oxysporum f. sp. ciceri (FOC) affects the chickpea crop causing wilt disease, more damaging and worldwide in occurrence. The main reason of Fusarium wilt is soilborne pathogen and showed symptoms, i.e., delaying crown, leaf anomalies, and rolled brown leaves. The number of strains is unknown of the soilborne pathogen and is difficult to control without solid information and identification of the pathogen. In general, symptoms of the disease occur at any stage of plant growth while more visible at the early stage of flowering and appears at the podding stage (late wilt). Late wilted plants exhibited falling of petioles, rachis, and leaflets as well as necrosis and discoloration of foliage. Early Fusarium wilt affects more than late wilting. However, late wilted plants produce lighter, rougher, and duller seeds as compared to normal. During the defense mechanisms, the plant uses many molecular signals or protein receptors to know the presence of microbes. Two modes of pathogen recognition are used by the host, i.e., effector-triggered immunity and pathogen-triggered immunity (PTI) . The invariant epitope types are called microbe-associated molecular patterns and are composed of flagellin, chitin, and lipopolysaccharides that help spread the disease. Studies also reported that the sensing of bacteria produce siderophores and fungi serve as MAMPs and hydroxyproline and rapid alkalization factors, but their role was not clear yet in defense mechanism. The severity of the chickpea wilt is depending upon the pathogen, genotypes, pathogenic races, inoculum density, environmental condition, and cultivar sensitivity. The activity of the wilting disease was triggered by a combination of pathogen activities.
Breeding approaches involved genetic diversity the most important step for a breeding program, selection of desirable plants for trait resistance and disease resistance and evaluation of the plant for commercial production. In an intraspecific hybrid program, the single-cross method was used in desi and Kabuli chickpea genotypes with variant genetic history. Molecular markers are an important tool for identification, characterizing, screening, and diversity among the pathogens and diseases. Commonly, internal transcribed spacer (ITS) markers are used for classification and screening of the fungi. While data regarding pathogen diversity is compulsory to comprehend pathology and development for control measures, SSR markers are used in separate backcross generation to assist in the selection against the resistance of Fusarium. Many pathogenic FOC spp. cause alike symptoms in chickpea crop as with FOC. For this purpose, screening, identification, and insight are more important among the pathogen FOC spp. This approach provides a deep understanding of the epidemiology of the disease and triggered the development of elite resistant genotypes by adopting breeding, molecular, and plant omics technology. QTLs linked molecular markers would also facilitate to identify the desired traits is the basic requisite for the application of molecular markers in the breeding program and enhance the selection process. Combinations of SCAR with a codominant marker (CaETR) linked with QTLAR1 for Ascochyta tagging and help to identify the resistance alleles from a core collection of resistant cultivars. Moreover, NILs are used as a value able tool for mapping, refining the target region and selection of the desired gene for resistance to FOC 0. Efforts have been made to construct the genetic map using molecular markers for tagging traits and site-specific gene of interest in chickpea. However, the population derived from interspecific crosses was made due to microsatellite markers exploiting more genetic polymorphisms. Recent advances in molecular plant sciences boost the knowledge, and transcriptomic emerged as a powerful method to understand differential genic response over specific time-bound fashion. TF has the function to regulate the cell development, differentiation, and growth by tagging specific site with DNA or multiple sites and triggered the activation or repression of the TF through various mechanisms and interactions, i.e., DNA-protein, protein-protein, and alteration in chromatin structure. The infection includes root identification, colonization, penetration, adhesion, and penetration of the root cortex and hyphal proliferation within the xylem vessels are controlled by transcription factors (TFs) . The functional characterization of the genes would also facilitate resistance mechanisms and their use in the breeding program against wilt resistance and crop tolerance mechanism along with target pathogen identification for the facilitation of the development of novel control management strategy .
References
Ahmad Z, Mumtaz AS, Ghafoor A, Ali A, Nisar M. Marker Assisted Selection (MAS) for chickpea Fusarium oxysporum wilt resistant genotypes using PCR based molecular markers. Mol Biol Rep. 2014;41:6755–62.
Ali L, Azam S, Rubio J, Kudapa H, Madrid E, Varshney RK, Castro P, Chen W, Gil J, Millan T. Detection of a new QTL/gene for growth habit in chickpea CaLG1 using wide and narrow crosses. Euphytica. 2015;204:473–85.
Allahverdipoor KH, Bahramnejad B, Amini J. Selection of molecular markers associated with resistance to Fusarium wilt disease in chickpea (Cicer arietinum L.) using multivariate statistical techniques. Aust J Crop Sci. 2011;5:1801–9.
Al-taae AK, Hadwan HA, SAE AJ. Physiological Races of Fusarium oxysporum f. sp. ciceris in Iraq. J Life Sci. 2013;7:1070–5.
Ashraf N, Ghai D, Barman P, Basu S, Gangisetty N, Mandal MK, Chakraborty N, Datta A, Chakraborty S. Comparative analyses of genotype dependent expressed sequence tags and stress-responsive transcriptome of chickpea wilt illustrate predicted and unexpected genes and novel regulators of plant immunity. BMC Genomics. 2009;10:415.
Bayraktar H, Dolar FS. Pathogenic variability of Fusarium oxysporum f. sp. ciceris isolates from chickpea in Turkey. Pak J Bot. 2012;44:821–3.
Beckman CH, Roberts EM. On the nature and genetic basis for resistance and tolerance of fungal wilt diseases. Adv Bot Res. 1995;21:35–77.
Beckman CH. The nature of wilt diseases of plants. St. Paul, MN: American Phytopathological Society; 1987.
Berrada AF, Shivakumar BG, Yaburaju NT. Chickpea in cropping systems. In: Yadav SSS, Redden R, Chen W, Sharma B, editors. Chickpea breeding and management. Wallingford: CABI Publishing; 2007. p. 193–212.
Bishopt GD, Cooper RM. An ultrastructural study of root invasion in three vascular wilt diseases. Physiol Plant Pathol. 1983;22:15–27.
Brown DW, Busman M, Proctor RH. Fusarium verticillioides SGE1 is required for full virulence and regulates expression of protein effector and secondary metabolite biosynthetic genes. Mol Plant-Microbe Interact. 2014;27:809–23.
Castro P, Pistón F, Madrid E, Millán T, Gil J, Rubio J. Development of chickpea near-isogenic lines for Fusarium wilt. Theor Appl Genet. 2010;121:1519–26.
Castro P, Rubio J, Millán T, Gil J, Cobos MJ. Fusarium wilt in chickpea: general aspect and molecular breeding. In: Rios TF, Ortega ER, editors. Fusarium: epidemiology, environmental sources and prevention. New York, NY: Nova Science Publishers; 2012. p. 101–22.
Cha JY, Han S, Hong HJ, Cho H, Kim D, Kwon Y, et al. Microbial and biochemical basis of a Fusarium wilt-suppressive soil. ISME J. 2016;10:119–29.
Chand H, Khirbat SK. Chickpea wilt and its management-a review. Agric Rev. 2009;30:1–12.
Cobos MJ, Fernández MJ, Rubio J, Kharrat M, Moreno MT, Gil J, Millán T. A linkage map of chickpea (Cicer arietinum L.) based on populations from Kabuli × Desi crosses: location of genes for resistance to Fusarium wilt race 0. Theor Appl Genet. 2005;110:1347–53.
Cobos MJ, Rubio J, Fernández-Romero MD, Garza R, Moreno MT, Millán T, Gil J. Genetic analysis of seed size, yield and days to flowering in a chickpea recombinant inbred line population derived from a Kabuli × Desi cross. Ann Appl Biol. 2007;151:33–42.
Ding Z, Li M, Sun F, Xi P, Sun L, Zhang L, Jiang Z. Mitogen-activated protein kinases are associated with the regulation of physiological traits and virulence in Fusarium oxysporum f. sp. cubense. PLoS One. 2015;10:e0122634.
Dubey SC, Suresh M, Singh B. Evaluation of Trichoderma species against Fusarium oxysporum f. sp. ciceris for integrated management of chickpea wilt. Biol Control. 2007;40:118–27.
Garcia-Limones C, Hervás A, Navas-Cortés JA, Jiménez-Diaz RM, Tena M. Induction of an antioxidant enzyme system and other oxidative stress markers associated with compatible and incompatible interactions between chickpea (Cicer arietinum L.) and Fusarium oxysporum f.sp. ciceris. Physiol Mol Plant Pathol. 2002;61:325–37.
Gaur PM, Gowda CLL, Knights EJ, Warkentin T, Acikoz N, Yadav SS, Kumar J. Breeding achivements. In: Chickpea breeding. Wallingford: Centre for Agriculture and Bioscience International (CABI); 2007.
Gaur PM, Jukanti AK, Varshney RK. Impact of genomic technologies on chickpea breeding strategies. Agronomy. 2012;2:199–221.
Gaur PM, Slinkard AE. Genetic control and linkage relations of additional isozyme markers in chick pea. Theor Appl Genet. 1990;80:648–56.
Gowda SJM, Radika P, Kadoo NY, Mhase LB, Gupta VS. Molecular mapping of wilt resistance genes in chickpea. Mol Breed. 2009;24:177–83.
Gupta S, Bhar A, Chatterjee M, Das S. Fusarium oxysporum f. sp. ciceri race 1 induced redox state alterations are coupled to downstream defense signaling in root tissues of chickpea (Cicer arietinum L.). PLoS One. 2013;8:e73163.
Gupta S, Chakraborti D, Rangi RK, Basu D, Das S. A molecular insight into the early events of chickpea (Cicer arietinum) and Fusarium oxysporum f. sp. ciceri (Race 1) interaction through cDNA-AFLP analysis. Phytopathology. 2009;99:1245–57.
Gupta S, Chakraborti D, Sengupta A, Basu D, Das S. Primary metabolism of chickpea is the initial target of wound inducing early sensed Fusarium oxysporum f. sp. ciceri race 1. PLoS One. 2010;5:e9030.
Gurjar GS, Giri AP, Gupta VS. Gene expression profiling during wilting in chickpea caused by Fusarium oxysporum f.sp. Ciceri. Am J Plant Sci. 2012;3:190–201.
Halila I, Cobos MJ, Rubio J, Millán T, Kharrat M, Marrakchi M, Gil J. Tagging and mapping a second resistance gene for Fusarium wilt race 0 in chickpea. Eur J Plant Pathol. 2009;124:87–92.
Haware MP, Nene YL, Natarajan M. The survival of Fusarium oxysporum f. sp. ciceri in the soil in the absence of chickpea. Phytopathol Mediterr. 1996;35:9–12.
Haware MP, Nene YL, Rajeshwari R. Eradication of Fusarium oxysporum f. sp. ciceri transmitted in chickpea seed. Phytopathol. 1978;68:1364–7.
Haware MP, Nene YL. Influence of wilt and different growth stages on yield loss in chickpea. Trop Grain Legum Bull. 1980;19:38–40.
Haware MP. Fusarium wilt and other important diseases chickpea in the Mediterranean area. In: Saxena MC, Cubero JI, Wery J, editors. Present status and future prospects of chickpea crop production and improvement in the Mediterranean countries. Zaragoza: CIHEAM; 1990. p. 61–4.
Hiremath PJ, Farmer A, Cannon SB, Woodward J, Kudapa H, Tuteja R, et al. Large scale transcriptome analysis in chickpea (Cicer arietinum L.), an orphan legume crop of the semiarid tropics of Asia and Africa. Plant Biotechnol J. 2011;9:922–31.
Hiremath PJ, Kumar A, Penmetsa RV, Farmer A, Schlueter JA, Chamarthi SK, et al. Large scale development of cost effective SNP marker assays for diversity assessment and genetic mapping in chickpea and comparative mapping in legumes. Plant Biotechnol J. 2012;10:716–32.
Imtiaz M, Materne M, Hobson K, van Ginkel M, Malhotra RS. Molecular genetic diversity and linked resistance to ascochyta blight in Australian chickpea breeding materials and their wild relatives. Aust J Agric Res. 2008;59:554–60.
Iruela M, Rubio J, Barro F, Cubero JI, Millán T, Gil J. Detection of two quantitative trait loci for resistance to ascochyta blight in an intra-specific cross of chickpea (Cicer arietinum L.): development of SCAR markers associated with resistance. Theor Appl Genet. 2006;112:278–87.
Jendoubi W, Bouhadida M, Millan T, Kharrat M, Gil J, Rubio J, Madrid E. Identification of the target region including the Foc0 1 /foc0 1 gene and development of near isogenic lines for resistance to Fusarium Wilt race 0 in chickpea. Euphytica. 2016;210:119–33.
Jimenez Diaz RM, Jimenez Gasco MM, Landa BB, Castillo P, Navas-Cortes JA. Fusarium wilt of chikpea. In: Chen W, Sharma HC, Muehlbauer FJ, editors. Compendium of chikpea and lentil diseases and pests. St. Paul, MN: The American Phytopathological Society; 2011.
Jimenez-Diaz RM, Alcala-Jimenez AR, Hervas A, Trapero-Casas JL. Pathogenic variability and host resistance in the Fusarium oxysporum f. sp. ciceris/Cicer arietinum. Pathosystem. In: Proceedings of the 3rd European Seminar on Fusarium mycotoxins, taxonomy, pathogenicity and host resistance. Plant Breeding and Acclimatization Institute, Radzikov, Poland; 1993, pp. 87–94.
Jiménez Díaz RM, Castillo P, del Mar Jiménez Gasco M, Landa BB, Navas Cortés JA. Fusarium wilt of chickpeas: biology, ecology and management. Crop Prot. 2015;73:16–27.
Jiménez-Fernández D, Montes-Borrego M, Jiménez-Díaz RM, Navas-Cortés JA, Landa BB. In Plant and soil quantification of Fusarium oxysporum f. sp. ciceris and evaluation of Fusarium wilt resistance in chickpea with a newly developed quantitative polymerase chain reaction assay. Phytopathology. 2011;101:250–62.
Jimenez-Fernandez D, Navas-Cortes JA, Montes-Borrego M, Jimenez-Diaz RM, Landa BB. Molecular and pathogenic characterization of Fusarium redolens, a new causal agent of Fusarium yellows in chickpea. Plant Dis. 2011;95:860–70.
Jiménez-Gasco M, Pérez-Artés E, Jiménez-Diaz RM. Identification of Pathogenic Races 0, 1B/C, 5, and 6 of Fusarium oxysporum f. sp. ciceris with Random Amplified Polymorphic DNA (RAPD). Eur J Plant Pathol. 2001;107:237–48.
Kachuei R, Yadegari MH, Safaie N, Ghiasian A, Noorbakhsh F, Piranfar V, Rezaie S. PCR-RFLP patterns for the differentiation of the Fusarium species in virtue of ITS rDNA. Curr Med Mycol. 2015;1:4–11.
Kaiser WJ, Alcala-Jimenez AR, Hervas-Vargas A, Trapero-Casas JL, Jiménez-Díaz RM. Screening of wild Cicer species for resistance to races 0 and 5 of Fusarium oxysporum f. sp. ciceris. Plant Dis. 1994;78:962–7.
Kaur NP, Mukhopadhyay AN. Integrated control of chickpea wilt complex by trichoderma and chemical methods in India. Trop Pest Manag. 1992;38:37–41.
Kazan K, Muehlbauer FJ, Weeden NE, Ladizinsky G. Inheritance and linkage relationships of morphological and isozyme loci in chickpea (Cicer arietinum L.). Theor Appl Genet. 1993;86:417–26.
Kohli D, Joshi G, Deokar AA, Bhardwaj AR, Agarwal M, Agarwal SK, Srinivasan R, Jain PK. Identification and characterization of wilt and salt stress-responsive microRNAs in chickpea through high-throughput sequencing. PLoS One. 2014;9:e108851.
Kumar J, Choudhary AK, Solanki RK, Pratap A. Towards marker-assisted selection in pulses: a review. Plant Breed. 2011;130:297–313.
Kumar Y, Dholakia BB, Panigrahi P, Kadoo NY, Giri AP, Gupta VS. Metabolic profiling of chickpea-Fusarium interaction identifies differential modulation of disease resistance pathways. Phytochemistry. 2015;116:120–9.
Kumar Y, Zhang L, Panigrahi P, Dholakia BB, Dewangan V, Chavan SG, Kunjir SM, Wu X, Li N, Rajmohanan PR, Kadoo NY, Giri AP, Tang H, Gupta VS. Fusarium oxysporum mediates systems metabolic reprogramming of chickpea roots as revealed by a combination of proteomics and metabolomics. Plant Biotechnol J. 2016;14:1589–603.
Kusuya Y, Hagiwara D, Sakai K, Yaguchi T, Gonoi T, Takahashi H. Transcription factor Afmac1 controls copper import machinery in Aspergillus fumigatus. Curr Genet. 2018;63:777–89.
Landa BB, Navas-Cortés JA, Jiménez-Díaz RM. Integrated management of Fusarium wilt of chickpea with sowing date, host resistance, and biological control. Phytopathology. 2004;94:946–60.
Lev-Yadun S, Gopher A, Abbo S. The cradle of agriculture. Science. 2000;288:1602–3.
Lichtenzveig J, Bonfil DJ, Zhang HB, Shtienberg D, Abbo S. Mapping quantitative trait loci in chickpea associated with time to flowering and resistance to Didymella rabiei the causal agent of Ascochyta blight. Theor Appl Genet. 2006;113:1357–69.
Lucas J. Plant diseases. In: Plant pathology and plant pathogens. Malden, MA: Blackwell Publishing; 1998.
Ma LJ, van der Does HC, Borkovich KA, Coleman JJ, Daboussi MJ, Di Pietro A, et al. Comparative genomics reveals mobile pathogenicity chromosomes in Fusarium. Nature. 2010;464:367–73.
Madrid E, Chen W, Rajesh PN, Castro P, Millán T, Gil J. Allele-specific amplification for the detection of ascochyta blight resistance in chickpea. Euphytica. 2013;189:183–90.
Mendgen K, Hahn M, Deising H. Morphogenesis and mechanisms of penetration by plant pathogenic fungi. Annu Rev Phytopathol. 1996;34:367–86.
Millan T, Madrid E, Cubero JI, Amri M, Castro P, Rubio J. Chickpea. In: De Ron Antonio M, editor. Grain legumes. New York, NY: Springer; 2015.
Millan T, Winter P, Jüngling R, Gil J, Rubio J, Cho S, et al. A consensus genetic map of chickpea (Cicer arietinum L.) based on 10 mapping populations. Euphytica. 2010;175:175–89.
Nautiyal CS. Selection of chickpea-rhizosphere-competent Pseudomonas fluorescens NBRI1303 antagonistic to Fusarium oxysporum f. sp. ciceri, Rhizoctonia bataticola and Pythium sp. Curr Microbiol. 1997;35:52–8.
Navas-Cortés JA, Hau B, Jiménez-Díaz RM. Effect of Sowing Date, Host Cultivar, and Race of Fusarium oxysporum f. sp. ciceri on development of Fusarium wilt of chickpea. Phytopathology. 1998;88:1338–46.
Navas-Cortés JA, Hau B, Jiménez-Díaz RM. Yield loss in chickpeas in relation to development of Fusarium wilt epidemics. Phytopathology. 2000;90:1269–78.
Nelson PE, Tousson TA, Cook RJ. Fusarium: diseases, biology and taxonomy. University Park, PA: Pennsylvania State University Press; 1981.
Nelson PE. Life cycle and epidemiology of Fusarium oxysporum. In: Mace M, Bell AA, Beckman C, editors. Fungal wilt diseases of plants. London, UK: Academic Press; 2012. p. 51–78.
Niño-Sánchez J, Tello V, Casado-del Castillo V, Thon M, Benito EP, Díaz-Mínguez JM. Gene expression patterns and dynamics of the colonization of common bean (Phaseolus vulgaris L.) by highly virulent and weakly virulent strains of Fusarium oxysporum. Front Microbiol. 2015;6:1–34.
Rubio J, Hajj-Moussa E, Kharrat M, Moreno MT, Millan T, Gil J. Two genes and linked RAPD markers involved in resistance to Fusarium oxysporum f. sp. ciceri race in chickpea. Plant Breed. 2003;122:188–91.
Salimath PM, Toker C, Sandhu JS, Kumar J, Suma B, Yadav SS, Bahl PN. Conventional breeding methods. In: Yadav SS, Redden RJ, Chen W, Sharma B, editors. Chickpea breeding and management. Wallingford: Centre for Agriculture and Bioscience International (CABI); 2007.
Schippers B, Van Eck WH. Formation and survival of chlamydospores in Fusarium. In: Fusarium: diseases, biology and taxonomy. London, UK: The Pennysylvania State University Press; 1981.
Shakir AS, Mirza JH. Location of seed-borne fungi in chickpea seed. Pak J Phytopathol. 1994;6:87–90.
Sharma KD, Chen W, Muehlbauer FJ. Genetics of chickpea resistance to five races of Fusarium wilt and a concise set of race differentials for Fusarium oxysporum f. sp. ciceri. Plant Dis. 2005;89:385–90.
Sharma KD, Muehlbauer FJ. Fusarium wilt of chickpea: physiological specialization, genetics of resistance and resistance gene tagging. Euphytica. 2007;157:1–14.
Sharma M, Sengupta A, Ghosh R, Agarwal G, Tarafdar A, Nagavardhini A, et al. Genome wide transcriptome profiling of Fusarium oxysporum f. sp. ciceri conidial germination reveals new insights into infection-related genes. Sci Rep. 2016;6:1–11.
Singh R, Sharma P, Varshney RK, Sharma SK, Singh NK. Chickpea improvement: role of wild species and genetic markers. Biotechnol Genet Eng Rev. 2008;25:267–314.
Singh S, Singh I, Kapoor K, Gaur PM, Chaturvedi SK, Singh NP, Sandhu JS. Chickpea. In: Broadening the genetic base of grain legumes. New Delhi, India: National Bureau of Plant Genetic Resources; 2014. p. 51–73.
Stapleton JJ, de Vay JE. Soil solarization: a non chemical approach for management of plant pathogens and pests. Crop Prot. 1986;5:190–8.
Stover RH. Banana root diseases caused by Fusarium oxysporum f. sp. cubense, Pseudomonas solanacearum, and Radopholus similis: A comparative study of life cycles in relation to control. In Root diseases and soil borne pathogens; 1970, pp. 197–200.
Strange RN. Introduction to plant pathology. London, UK: Wiley; 2003.
Thudi M, Bohra A, Nayak SN, Varghese N, Shah TM, Penmetsa RV, et al. Novel SSR markers from BAC end sequences, DArT arrays and a comprehensive genetic map with 1,291 marker loci for chickpea (Cicer arietinum L.). PLoS One. 2011;6:1–12.
Trapero Casas A, Jimenez Diaz RM. Fungal wilt and root rot diseases of chickpea in southern Spain. Phytopathology. 1985;75:1146–51.
Tullu A, Kaiser WJ, Kraft JM, Muehlbauer FJ. A second gene for resistance to race 4 of Fusarium wilt in chickpea and linkage with a RAPD marker. Euphytica. 1999;109:43–50.
Upadhyaya HD, Haware MP, Kumar J, Smithson JB. Resistance to wilt in chickpea. I. Inheritance of late-wilting in response to race 1. Euphytica. 1983;32:447–52.
Upasani ML, Gurjar GS, Kadoo NY, Gupta VS. Dynamics of colonization and expression of pathogenicity related genes in Fusarium oxysporum f.sp. ciceri during chickpea vascular wilt disease progression. PLoS One. 2016;11:1–21.
Upasani ML, Limaye BM, Gurjar GS, Kasibhatla SM, Joshi RR, Kadoo NY, Gupta VS. Chickpea-Fusarium oxysporum interaction transcriptome reveals differential modulation of plant defense strategies. Sci Rep. 2017;7:1–12. https://doi.org/10.1038/s41598-017-07114-x.
Varshney RK, Mohan SM, Gaur PM, Chamarthi SK, Singh VK, Srinivasan S, et al. Marker-assisted backcrossing to introgress resistance to Fusarium wilt race 1 and Ascochyta blight in C 214, an elite cultivar of chickpea. Plant Genom. 2014;7:1–11.
White TJ, Bruns T, Lee S, Taylor J. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ, editors. PCR protocols a guide to methods and applications. San Diego, CA: Academic Press; 1990. p. 315–22.
Yadeta KE, Thomma BPHJ. The xylem as battleground for plant hosts and vascular wilt pathogens. Front Plant Sci. 2013;4:1–13.
Yun Y, Zhou X, Yang S, Wen Y, You H, Zheng Y, et al. Fusarium oxysporum f. sp. lycopersici C2H2 transcription factor FolCzf1 is required for conidiation, fusaric acid production, and early host infection. Curr Genet. 2019;65(3):1–11.
Acknowledgments
The authors would like to extend their sincere appreciation to the Deanship of Scientific Research at King Saud University for funding this research group NO (RG-1435-014).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Hashem, A., Tabassum, B., Abd_Allah, E.F. (2020). Omics Approaches in Chickpea Fusarium Wilt Disease Management. In: Singh, B., Singh, G., Kumar, K., Nayak, S., Srinivasa, N. (eds) Management of Fungal Pathogens in Pulses. Fungal Biology. Springer, Cham. https://doi.org/10.1007/978-3-030-35947-8_4
Download citation
DOI: https://doi.org/10.1007/978-3-030-35947-8_4
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-35946-1
Online ISBN: 978-3-030-35947-8
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)