Abstract
Nicotiana benthamiana originates from northern Australia and belongs to the Suaveolentes section. It is used extensively as a model organism for many types of research, including plant–pathogen interactions, RNA interference, and functional genomics. Recent publications that used N. benthamiana as a model for plant–pathogen interactions focused mainly on bacteria, viruses, oomycete, and fungi. Two different N. benthamiana whole genome assemblies were published in 2012. These assemblies have been improved and structurally annotated in later versions but are still incomplete. The lineage most widely used in research originates from a population that has retained a loss-of-function mutation in Rdr1 (RNA-dependent RNA polymerase 1) that makes it highly susceptible to viruses. In this chapter, we review some of the techniques used in N. benthamiana to study plant–pathogen interactions, including virus-induced gene silencing, transient protein expression by agroinfiltration, stable genetic manipulation, and transcriptomics analysis, and discuss some of the results. Descriptions and links to some of the most relevant online resources for N. benthamiana are also provided.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
14.1 Evolutionary History of Nicotiana benthamiana
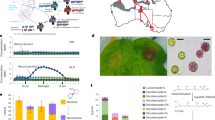
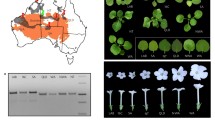
N. benthamiana Domin is among the most popular plants used for plant pathology studies. Despite this, specific details about the origin of this species are still unknown. N. benthamiana is a herbaceous plant with white flowers that is native to Australia where it can be found along the north coast, the Northern Territory, and Queensland (Fig. 14.1) (Global Biodiversity Information Facility (GBIF) 2018).
It can be identified following the dichotomous key described by Burbidge. N. benthamiana is distinguished from N. umbratica specimens by: “Upper cauline leaves sessile and forming leafy bracts. Laminae ovate to broad lanceolate, obtuse. Corolla lobes obtuse compared with the corolla lobes acute of the last one” (Burbidge 1960). The first available record of N. benthamiana can be found in the Herbarium Hookerianum at Kew Royal Botanical Gardens (http://specimens.kew.org/herbarium/K000196107), with a specimen collected by Benjamin Bynoe during the voyage of the Beagle (1837–1843) (Orchard 1999). Like other Australian Nicotiana species, N. benthamiana belongs to the Suaveolentes section, which contains 30 species and represents an important radiation of the Nicotiana genus in Australia (26 species), South Pacific (3 species), and Africa (1 species) (Marks et al. 2011). They are adapted to a wide range of conditions, from the high humidity coastal regions of Cairns (e.g., N. debneyi) to the extremely dry Great Sandy and Tanami deserts (e.g., N. benthamiana). All the Suaveolentes species have been described as allotetraploids with a variable number of chromosomes ranging from 2n = 4 × = 30 (N. suaveolens) to 2n = 4 × = 48 (N. debneyi).
Elucidation of the origin of the Suaveolentes section is still ongoing. An early publication by DeWolf and Goodspeed (1957) proposed that it could have formed after the hybridization of members of the Petunioides, Alatae/Sylvestres, and/or Noctiflorae sections. Later phylogenetic studies using plastid gene sequences such as matK indicated a Noctiflorae species as maternal donor (Aoki and Ito 2000; Clarkson et al. 2004). The use of interspacers ITS1 and ITS2, and 5.8S ribosomal genes indicated that an ancestral member of Alatae could be the possible paternal progenitor (Chase et al. 2003). A study using the GS (glutamine synthetase) gene, with both copies usually retained in polyploid Nicotiana species, indicated Sylvestres as the maternal donor and Trigonophyllae as the paternal donor (Clarkson et al. 2010). Nevertheless, the most accepted hypothesis based on several genes, such as ADH (alcohol dehydrogenase), LFY/FLO (LEAFY/FLORICAULA), GS, and nrITS (nuclear ribosomal interspacer), suggests a complex history where the most probable donors are Sylvestres (paternal) and Noctiflorae/Petunioides (maternal) (Kelly et al. 2013). The divergency ages between the Suaveolentes subgenomes and the corresponding proposed progenitors were estimated as 6.4 million years ago for the maternal contribution from the Noctiflorae section and 5.5 million years ago for the paternal contribution from the Sylvestres section (Clarkson et al. 2017).
14.2 Nicotiana benthamiana Genome Assemblies and Genetic Data
N. benthamiana was one of the first plant models positively affected by next-generation sequencing (NGS) technologies. Although several plant whole genome assemblies were built using NGS before 2012, most of them were for crops such as cucumber (Huang et al. 2009), apple (Velasco et al. 2010), and soybean (Schmutz et al. 2010). Two different N. benthamiana whole genome assemblies were published in 2012 (Bombarely et al. 2012; Naim et al. 2012). N. benthamiana is an allotetraploid plant (2n = 4 × = 38) with a large genome of 3.1 Gb, which made it difficult to assemble using the short reads obtained by NGS. Although both genomes were incomplete, they have been used extensively for plant–pathogen research. Statistics of the two N. benthamiana genome assemblies are summarized in Table 14.1.
Although a chromosome level assembly has still not been achieved (as October 2018), these assemblies have been improved. Statistics of the latest versions are summarized in Table 14.2. Evaluation of these genome assemblies using BUSCO (Simao et al. 2015) indicated they had similar completeness (95.4% for Niben1.0.1 and 94.4% for Nbv0.5), but Niben1.0.1 had a higher proportion of duplicated genes (46.0%) than Nbv0.5 (43.4%). These assemblies were also structurally annotated revealing 59,814 and 49,818 primary transcripts for Niben1.0.1 and Nbv0.5, respectively. Because of the ongoing diploidization process in N. benthamiana, its gene number is higher than the gene numbers for diploid Solanaceae species such as Solanum lycopersicum (Tomato Genome Consortium 2012), Solanum tuberosum (Potato Genome Sequencing Consortium 2011), Capsicum annuum (Kim et al. 2014), N. sylvestris and N. tomentosiformis (Sierro 2013), and Petunia axillaris (Bombarely et al. 2016), but lower than the gene number for allotetraploid N. tabacum (Sierro et al. 2014; Edwards et al. 2017). As expected, the BUSCO evaluation of the completeness of the annotations produced a lower value (88.1% for Niben1.0.1 and 75.2% for the transcriptome Nbv5.1 from the assembly Nbv0.5) than the evaluation of the genome assembly. Kourelis et al. (2018) used the genomes of other Nicotiana species to reanalyze the four genomes (Niben1.0.1, Niben0.4.2, Nbv0.5, and Nbv0.3) improving the quality of the N. benthamiana gene annotations.
To date, dozens of experiments using NGS technologies to investigate Solanaceae species have been published by the National Center for Biotechnology Information (NCBI). Indeed, the NCBI’s Taxonomy Browser has links to 342 SRA datasets, 442 BioSamples, and 74 BioProjects (up to December 2018) for N. benthamiana. These BioProjects not only study plant–pathogen interaction using resistance gene enrichment sequencing (RenSeq) (e.g., BioProject accession PRJNA496490) and RNA sequencing (RNA-Seq) (e.g., PRJNA360110), but also study expression regulatory mechanisms involving small RNAs (e.g., PRJNA481240, PRJNA309389) and circular RNAs (e.g., PRJNA422356), transcriptomic landscapes in several organs such as nectaries (e.g., PRJNA448133), transition and floral meristems (e.g., PRJNA343677), and grafting experiments (e.g., PRJDB3306).
14.3 Nicotiana benthamiana as a Model for the Study of Plant–Pathogen Interactions
The use of N. benthamiana as a model for plant–pathogen interactions has been thoroughly reviewed by Goodin et al. 2008; therefore, we will focus mainly on the information that has been generated since then. In their review, Goodin et al. performed a PubMed search using the term “Nicotiana benthamiana” and found 1,743 publications until 2006. We conducted a similar search, which yielded 3,606 hits (as of October 2018). By curating the publications since 2008, we identified 314 papers in which N. benthamiana was used as a tool to study different pathosystems, not taking into account studies in which N. benthamiana was used merely for transient protein expression and was unrelated to plant immunity. Among the 314 publications, the focus of study included plant interactions with bacterium (Senthil-Kumar and Mysore 2010; Kim et al. 2016), virus (Pavli et al. 2011; Zhu et al. 2014), oomycete (Adachi et al. 2015; King et al. 2014), fungus (De Jonge et al. 2012; Li et al. 2015), nematode (Mantelin et al. 2011; Ali et al. 2015), aphid (Peng et al. 2016; Atamian et al. 2013), insect (Chen et al. 2014), and viroid (Adkar-Purushothama et al. 2015). Among these pathogen types, the first four were the most widely studied using N. benthamiana (Fig. 14.2).
Pathogen types studied in publications involving N. benthamiana (2009 to October 2018). The input for WordSift (https://wordsift.org) was derived from the information from 314 publications in which N. benthamiana was employed in studies of plant–pathogen interaction published after 2008. Word font size indicates frequency of use
N. benthamiana was adopted as a model in the plant–virus research field because of its remarkable susceptibility (Goodin et al. 2008). In particular, the lineage employed at that time, which continues to be widely used in research, has a disruptive insertion in the gene coding RNA-dependent RNA polymerase I (Rdr1), which enhances plant fitness but simultaneously leads to its high susceptibility to viruses (Balli et al. 2015). Other reasons why N. benthamiana is so extensively used include high efficiency of gene silencing, ease of protein transient expression using Agrobacterium tumefaciens, the possibility of stable genetic manipulation (Todesco and De Felippes 2016), and the availability of a draft genome (Bombarely et al. 2012; Naim et al. 2012). In addition, the ongoing development of NGS techniques may further enhance the use of N. benthamiana to study plant–pathogen interactions. In the following sections, we describe and discuss each of the uses and techniques mentioned in this section.
14.4 Virus-Induced Gene Silencing (VIGS)
This technique relies on the silencing machinery of plants in order to be able to target the transcripts of genes of interest (Burch-Smith et al. 2004). Tobacco rattle virus (TRV)-based vectors are commonly used in numerous plant species, particularly those belonging to the Solanaceae family (Senthil-Kumar and Mysore 2014). VIGS technology offers several advantages such as rapid reverse genetic screens, mainly because the time-consuming process of plant transformation is avoided (Velasquez et al. 2009). VIGS also allows the simultaneous knockdown of multiple genes by selecting a single fragment with enough homology to the target genes or by arranging several fragments in a single construct (Miki et al. 2005; Zhou and Zeng 2017). An important general aspect of VIGS is the choice of a control against which the target gene’s silencing performance can be compared. An empty vector (TRV2::00) is not recommended as a control, rather TRV2 carrying an insert is preferred (Hartl et al. 2008; Wu et al. 2011). Inserts derived from green fluorescent protein (GFP) (Ryu et al. 2004), β-glucuronidase (Gonorazky et al. 2014), or an Escherichia coli gene (Ec1) (Rosli et al. 2013) have been used previously. Another key aspect of VIGS is the selection of an insert that will effectively target the gene(s) of interest while avoiding off-targets. The Sol Genomics Network (SGN) VIGS Tool allows the interactive identification of most probable targets and off-targets, thereby assisting in construct design (Fernandez-Pozo et al. 2015a, b) (see the section “Online resources for Nicotiana benthamiana” in this chapter for more details). In N. benthamiana, VIGS has been used to silence some target genes (Kang et al. 2010; Choi et al. 2011; Liebrand et al. 2012 Rosli et al. 2013; Pombo et al. 2014), as well as large sets of candidate genes in a high-throughput fashion (Chakravarthy et al. 2010; Zhu et al. 2010; Mantelin et al. 2011; Rojas et al. 2012; Senthil-Kumar and Mysore 2012; Xu et al. 2012; Du et al. 2013; Lee et al. 2013; Nakano et al. 2013). The silencing step is followed by a readout experiment that depends on the particular process under study and may involve transient protein expression and elicitation of programmed cell death (PCD) (see below), pathogen challenge (Asai et al. 2008; Tanaka et al. 2009; Senthil-Kumar and Mysore 2010; Chaparro-Garcia et al. 2011; Kiba et al. 2012; Du et al. 2013; Ohtsu et al. 2014; Adachi et al. 2015; Bruckner et al. 2017; Turnbull et al. 2017), reactive oxygen species (ROS) production (Shibata et al. 2010; Segonzac et al. 2011; Deng et al. 2016; Pfeilmeier et al. 2016; Saur et al. 2016), nitric oxide production (Zhang et al. 2010), and stomatal aperture measurement (Zhang et al. 2012, 2016). Although N. benthamiana is susceptible to many pathogens, it may not be a host to the pathogen under study. This has been overcome by engineering the pathogen (Wei et al. 2007) or using a related species that causes disease in N. benthamiana (Yu et al. 2012; Yin et al. 2013; Wang et al. 2016). Chakravarthy et al. (2010) developed an assay that can be used to test the effect of silencing a candidate gene on the pattern-triggered immunity (PTI) response. This requires the infiltration of a PTI inducer (Pseudomonas fluorescens) and, a few hours later, a second infiltration performed in an overlapping manner with a challenger (Pseudomonas syringae pv. tomato, Pst). The speed of PCD progression in the overlapping area is related to the functionality of PTI. The authors coupled this assay with VIGS high-throughput screening and identified genes involved in the PTI response.
Recently, Zhou and Zeng 2017 developed a novel VIGS strategy to specifically and efficiently knockdown members of a highly homologous gene family using fragments of approximately 70 base pairs. The authors combined the SGN VIGS Tool (Fernandez-Pozo et al. 2015a, b) with a manual optimization step to select the fragments in order to analyze functional redundancy among members of a gene family.
14.5 Transient Protein Expression by Agroinfiltration
This technique uses Agrobacterium tumefaciens carrying an expression vector system. Usually a suspension is infiltrated into a leaf and within 1–3 days the tissue is ready for downstream analysis or treatment. The most commonly used vector cloning methods rely mainly on Gateway (Life Technologies, Carlsbad, CA, USA) (Karimi et al. 2002; Nakagawa et al. 2007) and type IIS assembly-based technologies (Golden Gate and GreenGate) (Engler et al. 2008; Lampropoulos et al. 2013). Expression vectors allow the targeted proteins to be expressed under different promoters, depending on the final purpose of the experiment. In plant–pathogen studies, transient expression in N. benthamiana has been driven mainly by the constitutive cauliflower mosaic virus 35S (CaMV 35S) promoter (Kang et al. 2010; Anderson et al. 2012; Stirnweis et al. 2014; Song et al. 2015; Saur et al. 2016) and in some cases under the native promoter (Sato et al. 2014; El Kasmi et al. 2017; Ramachandran et al. 2017). Alternatively, when controlled timing of protein expression was required or when prolonged protein expression could lead to detrimental effects, gene expression was modulated by inducible promoters (Stork et al. 2015; Hwang et al. 2017). To study plant immunity, transient expression in N. benthamiana has been followed mainly by subcellular and tissue localization (Thiel et al. 2012; Rodriguez et al. 2014; Su et al. 2015; Zhuang et al. 2016), protein–protein interactions through co-immunoprecipitation (Co-IP) (Zhao et al. 2013; Hurni et al. 2014; Kim and Hwang 2015; El Kasmi et al. 2017), bimolecular fluorescence complementation (Sun et al. 2014; Du et al. 2015; Liu et al. 2016) or luciferase complementation imaging assay (Du et al. 2013), effect on pathogen performance (Bae et al. 2011; Medina-Hernandez et al. 2013; Song et al. 2015), expression of immune-related or marker genes (Nguyen et al. 2010; Li et al. 2014; Rodriguez et al. 2014; Su et al. 2014; Chaparro-Garcia et al. 2015), electrolyte leakage (Yu et al. 2012; Teper et al. 2014; Gupta et al. 2015), and ROS production (Stork et al. 2015). Because N. benthamiana does not respond with a ROS burst when challenged with the microbe-associated molecular pattern (MAMP) flgII-28 from flagellin, transient overexpression of the tomato receptor-like kinase flagellin-sensing FLS3 was used to confer responsiveness to N. benthamiana (Hind et al. 2016). The transient expression of a mutated version of FLS3 showed that its kinase activity was required for downstream signaling associated with the flgII-28 ROS burst.
Transient protein expression in N. benthamiana also was employed to identify the receptor of another MAMP from the bacterial cold shock protein, the csp22 peptide (Saur et al. 2016). Under the hypothesis that this receptor should interact with the coreceptor BAK1 (that is part of several activated receptor complexes) upon csp22 challenge, BAK1 was expressed fused to a GFP tag that was used for immunoprecipitation, followed by liquid chromatography–mass spectrometry (LC-MS/MS). This strategy allowed the identification of the N. benthamiana receptor-like protein required for csp22 responsiveness, NbCSPR.
Attachment of fatty acids as a post-translational modification is important for the regulation of protein location and is of particular interest in the study of plant–pathogen interactions (Boyle and Martin 2015). In N. benthamiana, transient protein expression coupled with click chemistry has been exploited for the detection of modifications such as N-myristoylation and S-acylation of both pathogen and host proteins (Boyle et al. 2016).
Transient overexpression of the transcription factor CabZIP63 from Capsicum annuum (pepper) in N. benthamiana leaves, followed by chromatin immunoprecipitation combined with PCR, was used to study the transcriptional regulation that CabZIP63 exerted on CaWRKY40, a transcription factor involved in the response to the bacterial pathogen Ralstonia solanacearum (Shen et al. 2016).
One of the most frequently used outcomes following transient protein expression is the observation of, usually macroscopic, PCD symptoms (Kang et al. 2010; Cunnac et al. 2011; Chronis et al. 2013; Mafurah et al. 2015). The large N. benthamiana leaves allow testing several elicitors using different concentrations or combinations. Coupled with VIGS, this approach has been used to test if the targeted candidate gene participates in PCD associated with highly divergent types of pathogens, such as bacterium, virus, nematode, and oomycete (Del Pozo et al. 2004; Oh and Martin 2011; Pombo et al. 2014). This approach also has been used to test PCD-suppressing activity of pathogen effectors by co-infiltration with inducers of PCD (Teper et al. 2014; Stork et al. 2015). The importance of using appropriate controls in these types of experiments has been highlighted by Adlung and Bonas 2017 who found that some effectors affected Agrobacterium tumefaciens performance in leaf tissues, which could lead to overall lower amounts of the PCD-eliciting protein than expected.
Protein expression also has been used in high-throughput approaches. A simple toothpick method (Takken et al. 2000) coupled with the observation of PCD development allowed the identification of plant proteins involved in resistance (Nasir et al. 2005; Coemans et al. 2008; Takahashi et al. 2009). High-throughput transient in planta expression assays were performed to study the biological activities of pathogen effector proteins (Caillaud et al. 2012; Stam et al. 2013; Petre et al. 2015). The availability of genome sequences from a variety of pathogens allows the computational prediction of candidate effector genes based on conserved host translocation motifs and their presence in well-defined genome regions (Pais et al. 2013). Cloning and transient Agrobacterium-mediated expression of candidate effectors can give valuable insights into the virulence activities of effector proteins, particularly regarding the suppression of host plant immunity (Pais et al. 2013). Using this approach named “effectoromics,” Petre et al. 2015 selected, cloned, and expressed 20 candidate effectors in N. benthamiana leaf cells to determine their subcellular localizations and to identify the plant proteins they interacted with, through downstream experiments such as Co-IP and mass spectrometry. A similar approach was used for the phenotypic characterization of 84 members of a subclass of Phytophthora capsici effectors, which allowed the identification of one member that, when expressed in planta, enhanced P. capsici virulence in N. benthamiana (Stam et al. 2013). By transiently expressing 49 RxLR effector candidates (HaRxLs) of the filamentous phytopathogen Hyaloperonospora arabidopsidis fused to fluorescent tags in N. benthamiana, two major classes of HaRxLs were defined as those that accumulated in the plant cell nucleus and those that accumulated in the plant membranes. Functional analysis revealed that, in particular, a membrane-localized effector, HaRxL17, enhanced the susceptibility of N. benthamiana to this pathogen (Caillaud et al. 2012).
14.6 Stable Genetic Manipulation
The technique has been used frequently in studies of plant–virus interactions for stably overexpressing virus-derived transcripts by taking advantage of post-transcriptional gene silencing (PTGS) to generate N. benthamiana plants more resistant to a pathogen (Ling et al. 2008; Reyes et al. 2009). Plant PTGS machinery has been exploited to improve resistance by overexpressing transcripts derived from viral DNA fragments (Lin et al. 2012), double-strand RNA from viral replicase (Pavli et al. 2012), artificial microRNA (Ali et al. 2013; Wagaba et al. 2016), and interfering satellite RNA and RNA interference (RNAi) (Montes et al. 2014). Expression of a whitefly GroEL chaperonin, a protein that can bind to several viruses, produced N. benthamiana plants more tolerant to tomato leaf curl virus and cucumber mosaic virus (Edelbaum et al. 2009).
Broad-spectrum resistance was explored using stable expression of artificial transcript activator-like effectors, assembled based on highly conserved regions within begomovirus genomes, that conferred partial resistance to three begomoviruses tested (Cheng et al. 2015). The CRISPR/Cas9 system (clustered regulatory interspaced short palindromic repeat/CRISPR-associated DNA endonuclease 9), which revolutionized plant and animal genome editing (Samanta et al. 2016), has been used successfully in N. benthamiana (Nekrasov et al. 2013; Li et al. 2013). CRISPR/Cas9 N. benthamiana plants with an inactivated Argonaute 2 gene were used to investigate broad range resistance, which showed that the Argonaute 2 protein had antiviral activity against at least three viruses in a virus-specific manner (Ludman et al. 2017). Stable plastid protein expression (transplastomics) in N. benthamiana using a plastid-transformation vector and biolistic was employed to express multiple defense genes (Chen et al. 2014). The results showed that a combination of sweet potato sporamin, taro cystatin, and chitinase from Paecilomyces javanicus conferred broad-spectrum resistance against insects, pathogens, and abiotic stresses.
As mentioned in the “Transient protein expression by agroinfiltration” section, N. benthamiana has been used to study the molecular mechanisms of pathogen effectors. Plasmopara viticola effectors with immune-suppressing activities have been identified by combining transient and stable protein expression. In particular, the overexpression of the effector PvRxLR28 in N. benthamiana and grapevine produced plants with enhanced susceptibility to this oomycete (Xiang et al. 2016). Interestingly, the stable overexpression of two Phytophthora sojae effectors enhanced disease resistance and tolerance to salt and drought stresses in N. benthamiana plants (Rajput et al. 2015; Zhang et al. 2015). These results suggested the possible use of these effectors in crop breeding strategies.
A good degree of conservation of certain molecular pathways, which allows interfamily gene transfer, has been key to the use of N. benthamiana in plant–pathogen studies. For example, stably expressed proteins from Arabidopsis thaliana (Lacombe et al. 2010; Narusaka et al. 2013; Huang et al. 2014; Wang et al. 2016), tomato (Rommens et al. 1995), and cotton (Lu et al. 2013; Li et al. 2014; Xu et al. 2014) were shown to have functional roles in N. benthamiana immunity. A pathogen-induced nucleotide-binding (NB)-leucine-rich repeat (LRR) candidate gene from Vitis amurensis was stably overexpressed in N. benthamiana (Li et al. 2017) and the transgenic plants were more resistant not only to the oomycete Plasmopara viticola, but also to drought and salt stresses, suggesting that the NB-LRR protein may have immune and non-immune roles.
Some stably modified N. benthamiana lines have been employed as tools to study the plant immune response. Line SLJR15 expresses the reporter protein Aequorin (Knight et al. 1993), which allows cytoplasmatic Ca2+ dynamics to be studied through luminescence imaging (Segonzac et al. 2011; Saur et al. 2016). Line 16c, which expresses Aequorea victoria GFP targeted to the endoplasmic reticulum (Ruiz et al. 1998), is the most frequently used N. benthamiana line, in particular, to study small RNAs (Philips et al. 2017). Using NGS, the T-DNA insertion region was identified in line 16c and, surprisingly, a portion of a bacterial transposon was found to have co-integrated with this insertion, raising the concern that such events may occur in lines designed for commercial use (Philips et al. 2017). A Cas9-overexpressing (Cas9-OE) N. benthamiana line was developed as part of a virus-mediated genome editing system (Ali et al. 2015). In these plants, the DNA endonuclease Cas9 is stably overexpressed under the 35S promoter, and the single guide RNA (sgRNA), which determines the target sequence, is systemically delivered via tobacco rattle virus (Ali et al. 2015). This approach was used to rapidly test different sgRNAs to confer better immunity more efficiently against the DNA virus, tomato yellow leaf curl virus.
14.7 Transcriptomics Analysis of Plant–Pathogen Interactions Using N. benthamiana
Before a microarray derived entirely from N. benthamiana expressed sequence tags (EST) was developed, potato cDNA arrays were used to determine changes in gene expression in response to virus infection (Senthil et al. 2005; Dardick 2007). Subsequently, a N. benthamiana microarray was developed and used to comparatively analyze gene expression changes in response to the necrotrophic Pectobacterium carotovorum and hemibiotrophic Pst DC3000 bacteria (Kim et al. 2011). Analysis of the data showed that the transcriptomic expression profiles of N. benthamiana in response to P. carotovorum were similar to those in response to a mutated Pst DC3000 without a type III secretion system.
The development of NGS techniques and the availability of a draft genome (Bombarely et al. 2012; Naim et al. 2012) may further enhance the use of N. benthamiana as a model plant. So far, the Illumina RNA-Seq approach has been used to analyze changes in messenger RNA (mRNA) or small RNA levels. High-throughput small RNA sequencing was used to study the effectiveness of different RNA silencing approaches in the control of virus infections based on the expression of large virus-derived sequences (Montes et al. 2014; Zhao et al. 2015). NGS also was employed to identify and characterize microRNAs involved in the N protein-mediated immune response to tobacco mosaic virus (Yin et al. 2015). RNA-Seq was recently used for the identification of a set of stably expressed genes in N. benthamiana which were validated as reference genes for reverse transcription-quantitative PCR (qPCR) in plant–bacteria interaction experiments (Pombo et al. 2019).
The use of integrated omics, which included RNA-Seq analysis of healthy and Odontoglossum ringspot virus (ORSV)-infected N. benthamiana leaves combined with proteomics, allowed the identification of putative host proteins that interacted with ORSV capsid protein, which is important for viral long-distance movement in N. benthamiana (Lin et al. 2015). Recently, transcriptomic differences detected between mock-treated and Phytophthora parasitica-inoculated N. benthamiana leaves provided broad insights into N. benthamiana defense mechanisms against this oomycete pathogen (Shen et al. 2016).
14.8 Online Resources for Nicotiana benthamiana
Many online resources are available for N. benthamiana ranging from bioinformatics tools and sequence databases to germplasm collections. Some of the most relevant of these resources are summarized in Table 14.3.
The three main genomics and transcriptomics resources for N. benthamiana sequences and annotations are the Queensland University of Technology (QUT) database, the SGN, and NCBI’s GenBank. The data in the QUT database are based mostly on the Nbv0.5 genome assembly and transcriptomes v5.1 and v6.1 from Naim et al. 2012. The main features and tools in this database are BLAST and keyword searches, data downloading, genome browsing, expression visualization, and a transcript lookup tool to find corresponding transcripts among the four genome assemblies (Niben1.0.1, Niben0.4.4, Nbv0.5, and Nbv0.3) of Bombarely et al. (2012) and Naim et al. (2012). The SGN contains resources and tools for Niben0.4.4 and Niben1.0.1 (Bombarely et al. 2012). The sequences, annotations, and proteomics resources from these genome versions are available for downloading and the data also can be queried in BLAST and JBrowse (genome browser) tools. The SGN also hosts the SGN VIGS Tool (see below) and SolCyc, a bioinformatics tool to visualize metabolic pathways based on genes from Solanaceae species. GenBank is a large database that contains sequences, annotations, scientific publications, and much more information for all species. Links to the most useful NCBI resources for N. benthamiana can be found on the N. benthamiana page in NCBI’s Taxonomy Browser (https://www.ncbi.nlm.nih.gov/Taxonomy/Browser/wwwtax.cgi?id=4100). In addition to these three web portals, the Boyce Thompson Institute (BTI) N. benthamiana website (https://btiscience.org/our-research/research-facilities/research-resources/nicotiana-benthamiana/) has a collection of links to bioinformatics tools and experimental protocols and resources for N. benthamiana.
Two bioinformatics tools, CRISPR-P and CCTop, provide support to design targets for genome editing using CRISPR with Niben0.4.4 and Niben1.0.1, respectively, as the reference genomes (see Table 14.3).
VIGS is an important and efficient tool for functional genomics in N. benthamiana (see the “Virus induced gene silencing (VIGS)” section). Several resources for designing and performing VIGS analysis in N. benthamiana are available, including, for example, the SGN VIGS Tool (Fernandez-Pozo et al. 2015), the VIGS database (Senthil-Kumar and Mysore 2014), and a video titled “Virus-induced gene silencing (VIGS) in Nicotiana benthamiana and tomato” (Velasquez et al. 2009). The SGN VIGS Tool assists in the design of VIGS constructs based on Niben1.0.1 or Niben0.4.4 using an interactive and intuitive interface. This tool predicts the best target of a gene of interest, thereby allowing the design of constructs to silence multiple genes and minimizing the silencing of off-target genes. The VIGS database contains phenotypic information for a large number of genes silenced in N. benthamiana. Currently, the database contains about 1,300 descriptions and/or photographs of gene-silenced plants as well as sequence information of about 4,500 ESTs used for VIGS. This database also includes keyword and BLAST searches to explore all the resources.
Another resource of interest for biotechnology is the International Genetically Engineered Machine (iGem) Foundation’s Registry of Standard Biological Parts for N. benthamiana (http://parts.igem.org/Collections/Plants#Nicotiana_benthamiana), which provides a collection of expression constructs, reporters, promoters, and other elements tested or that could be used in N. benthamiana.
More information about N. benthamiana taxonomic resources, populations and natural occurrence, and germplasm collections can be found in the links provided in Table 14.3.
References
Adachi H, Nakano T, Miyagawa N et al (2015) WRKY transcription factors phosphorylated by MAPK regulate a plant immune NADPH oxidase in Nicotiana benthamiana. Plant Cell 27:2645–2663
Adkar-Purushothama CR, Kasai A, Sugawara K et al (2015) RNAi mediated inhibition of viroid infection in transgenic plants expressing viroid-specific small RNAs derived from various functional domains. Sci Rep 5:17949
Adlung N, Bonas U (2017) Dissecting virulence function from recognition: cell death suppression in Nicotiana benthamiana by XopQ/HopQ1-family effectors relies on EDS1-dependent immunity. Plant J 91:430–442
Ali I, Amin I, Briddon RW, Mansoor S (2013) Artificial microRNA-mediated resistance against the monopartite begomovirus Cotton leaf curl Burewala virus. Virol J 10:231
Ali S, Magne M, Chen S et al (2015a) Analysis of Globodera rostochiensis effectors reveals conserved functions of SPRYSEC proteins in suppressing and eliciting plant immune responses. Front Plant Sci 6:623
Ali Z, Abul-Faraj A, Li L et al (2015b) Efficient virus-mediated genome editing in plants using the CRISPR/Cas9 system. Mol Plant 8:1288–1291
Ali Z, Abulfaraj A, Idris A et al (2015c) CRISPR/Cas9-mediated viral interference in plants. Genome Biol 16:238
Anderson RG, Casady MS, Fee RA et al (2012) Homologous RXLR effectors from Hyaloperonospora arabidopsidis and Phytophthora sojae suppress immunity in distantly related plants. Plant J 72:882–893
Asai S, Ohta K, Yoshioka H (2008) MAPK signaling regulates nitric oxide and NADPH oxidase-dependent oxidative bursts in Nicotiana benthamiana. Plant Cell 20:1390–1406
Atamian HS, Chaudhary R, Cin VD et al (2013) In planta expression or delivery of potato aphid Macrosiphum euphorbiae effectors Me10 and Me23 enhances aphid fecundity. Mol Plant Microbe Interact 26:67–74
Aoki S, Ito M (2000) Molecular phylogeny of Nicotiana (Solanaceae) based on the nucleotide sequence of the matK gene. Plant Biol 2:316–324
Bae H, Roberts DP, Lim HS et al (2011) Endophytic Trichoderma isolates from tropical environments delay disease onset and induce resistance against Phytophthora capsici in hot pepper using multiple mechanisms. Mol Plant Microbe Interact 24:336–351
Balli J, Nakasugi K, Jia F et al (2015) The extremophile Nicotiana benthamiana has traded viral defence for early vigour. Nat Plants 1:15165
Benson DA, Karsch-Mizrachi I, Lipman DJ et al (2004) GenBank: update. Nucleic Acids Res 32:D23–D26
Burbidge NT (1960) The Australian species of Nicotiana L. (Solanaceae). Aust J Botany 8(3):342–380
Bombarely A, Rosli HG, Vrebalov J et al (2012) A draft genome sequence of Nicotiana benthamiana to enhance molecular plant-microbe biology research. Mol Plant Microbe Interact 25:1523–1530
Bombarely A, Moser M, Amrad A et al (2016) Insight into the evolution of the Solanaceae from the parental genomes of Petunia hybrida. Nature Plants 2:16074
Boyle PC, Martin GB (2015) Greasy tactics in the plant-pathogen molecular arms race. J Exp Bot 66:1607–1616
Boyle PC, Schwizer S, Hind SR et al (2016) Detecting N-myristoylation and S-acylation of host and pathogen proteins in plants using click chemistry. Plant Methods 12:38
Bruckner FP, Xavier ADS, Cascardo RS et al (2017) Translationally controlled tumour protein (TCTP) from tomato and Nicotiana benthamiana is necessary for successful infection by a potyvirus. Mol Plant Pathol 18:672–683
Burch-Smith TM, Anderson JC, Martin GB et al (2004) Applications and advantages of virus-induced gene silencing for gene function studies in plants. Plant J 39:734–746
Caillaud MC, Piquerez SJ, Fabro G et al (2012) Subcellular localization of the Hpa RxLR effector repertoire identifies a tonoplast-associated protein HaRxL17 that confers enhanced plant susceptibility. Plant J 69:252–265
Chakravarthy S, Velasquez AC, Ekengren SK et al (2010) Identification of Nicotiana benthamiana genes involved in pathogen-associated molecular pattern-triggered immunity. Mol Plant Microbe Interact 23:715–726
Chaparro-Garcia A, Schwizer S, Sklenar J et al (2015) Phytophthora infestans RXLR-WY Effector AVR3a Associates with Dynamin-Related Protein 2 Required for Endocytosis of the Plant Pattern Recognition Receptor FLS2. PLoS ONE 10:e0137071
Chase M, Knapp S, Cox AV et al (2003) Molecular systematics, GISH and the origin of hybrid taxa in Nicotiana (Solanaceae). Ann Bot-London 92:107–127
Chaparro-Garcia A, Wilkinson RC, Gimenez-Ibanez S et al (2011) The receptor-like kinase SERK3/BAK1 is required for basal resistance against the late blight pathogen Phytophthora infestans in Nicotiana benthamiana. PLoS ONE 6:e16608
Chen PJ, Senthilkumar R, Jane WN et al (2014) Transplastomic Nicotiana benthamiana plants expressing multiple defence genes encoding protease inhibitors and chitinase display broad-spectrum resistance against insects, pathogens and abiotic stresses. Plant Biotechnol J 12:503–515
Cheng X, Li F, Cai J et al (2015) Artificial TALE as a convenient protein platform for engineering broad-spectrum resistance to Begomoviruses. Viruses 7:4772–4782
Choi HW, Kim YJ, Hwang BK (2011) The hypersensitive induced reaction and leucine-rich repeat proteins regulate plant cell death associated with disease and plant immunity. Mol Plant Microbe Interact 24:68–78
Chronis D, Chen S, Lu S et al (2013) A ubiquitin carboxyl extension protein secreted from a plant-parasitic nematode Globodera rostochiensis is cleaved in planta to promote plant parasitism. Plant J 74:185–196
Clarkson JJ et al (2004) Phylogenetic relationships in Nicotiana (Solanaceae) inferred from multiple plastid DNA regions. Mol Phylogenet Evol 33:75–90
Clarkson JJ, Kelly LJ, Leitch AR et al (2010) Nuclear glutamine synthetase evolution in Nicotiana: phylogenetics and the origins of allotetraploid and homoploid (diploid) hybrids. Mol Phylogenet Evol 55:99–112
Clarkson JJ, Dodsworth S, Chase MW (2017) Time-calibrated phylogenetic trees establish a lag between polyploidisation and diversification in Nicotiana (Solanaceae). Plant Syst Evol 303:1001–1012
Coemans B, Takahashi Y, Berberich T et al (2008) High-throughput in planta expression screening identifies an ADP-ribosylation factor (ARF1) involved in non-host resistance and R gene-mediated resistance. Mol Plant Pathol 9:25–36
Cunnac S, Chakravarthy S, Kvitko BH et al (2011) Genetic disassembly and combinatorial reassembly identify a minimal functional repertoire of type III effectors in Pseudomonas syringae. Proc Natl Acad Sci USA 108:2975–2980
Dardick C (2007) Comparative expression profiling of Nicotiana benthamiana leaves systemically infected with three fruit tree viruses. Mol Plant Microbe Interact 20:1004–1017
De Jonge R, Van Esse HP, Maruthachalam K et al (2012) Tomato immune receptor Ve1 recognizes effector of multiple fungal pathogens uncovered by genome and RNA sequencing. Proc Natl Acad Sci USA 109:5110–5115
DeWolf GP, Goodspeed TH (1957) The Genus Nicotiana. Origins, Relationships and Evolution of Its Species in the Light of Their Distribution, Morphology and Cytogenetics. The Southwestern Naturalist 2, p 177
Del Pozo O, Pedley KF, Martin GB (2004) MAPKKKalpha is a positive regulator of cell death associated with both plant immunity and disease. EMBO J 23:3072–3082
Deng XG, Zhu T, Zou LJ et al (2016) Orchestration of hydrogen peroxide and nitric oxide in brassinosteroid-mediated systemic virus resistance in Nicotiana benthamiana. Plant J 85:478–493
Du J, Tian Z, Liu J et al (2013a) Functional analysis of potato genes involved in quantitative resistance to Phytophthora infestans. Mol Biol Rep 40:957–967
Du Y, Berg J, Govers F et al (2015) Immune activation mediated by the late blight resistance protein R1 requires nuclear localization of R1 and the effector AVR1. New Phytol 207:735–747
Du Y, Zhao J, Chen T et al (2013b) Type I J-domain NbMIP1 proteins are required for both Tobacco mosaic virus infection and plant innate immunity. PLoS Pathog 9:e1003659
Edelbaum D, Gorovits R, Sasaki S et al (2009) Expressing a whitefly GroEL protein in Nicotiana benthamiana plants confers tolerance to tomato yellow leaf curl virus and cucumber mosaic virus, but not to grapevine virus A or tobacco mosaic virus. Arch Virol 154:399–407
Edwards KD, Fernandez-Pozo N, Drake-Stowe K et al (2017) A reference genome for Nicotiana tabacum enables map-based cloning of homeologous loci implicated in nitrogen utilization efficiency. BMC Genom 18:448
El Kasmi F, Chung EH, Anderson RG et al (2017) Signaling from the plasma-membrane localized plant immune receptor RPM1 requires self-association of the full-length protein. Proc Natl Acad Sci USA 114:E7385–E7394
Engler C, Kandzia R, Marillonnet S (2008) A one pot, one step, precision cloning method with high throughput capability. PLoS One 3:e3647
Fernandez-Pozo N, Rosli HG, Martin GB, Mueller LA (2015a) The SGN VIGS tool: user-friendly software to design virus-induced gene silencing (VIGS) constructs for functional genomics. Mol Plant 8:486–488
Fernandez-Pozo N, Menda N, Edwards JD et al (2015b) The Sol genomics network (SGN)—from genotype to phenotype to breeding. Nucleic Acids Res 43(D1):D1036–D1041
Foerster H, Bombarely A, Battey JND et al (2018) SolCyc: a database hub at the sol genomics network (SGN) for the manual curation of metabolic networks in Solanum and Nicotiana specific databases. Database, Volume 2018, 1 January 2018, bay035
Global Biodiversity Information Facility (GBIF) (2018) The species database. https://www.gbif.org/species/3800423. Accessed 19 Oct 2018
Gonorazky G, Ramirez L, Abd-El-Haliem A et al (2014) The tomato phosphatidylinositol-phospholipase C2 (SlPLC2) is required for defense gene induction by the fungal elicitor xylanase. J Plant Physiol 171:959–965
Goodin MM, Zaitlin D, Naidu RA, Lommel SA (2008) Nicotiana benthamiana: its history and future as a model for plant-pathogen interactions. Mol Plant Microbe Interact 21:1015–1026
Gupta MK, Nathawat R, Sinha D et al (2015) Mutations in the predicted active site of Xanthomonas oryzae pv. oryzae XopQ differentially affect virulence, suppression of host innate immunity, and induction of the HR in a nonhost plant. Mol Plant Microbe Interact 28:195–206
Hartl M, Merker H, Schmidt DD, Baldwin IT (2008) Optimized virus-induced gene silencing in Solanum nigrum reveals the defensive function of leucine aminopeptidase against herbivores and the shortcomings of empty vector controls. New Phytol 179:356–365
Hind SR, Strickler SR, Boyle PC et al (2016) Tomato receptor FLAGELLIN-SENSING 3 binds flgII-28 and activates the plant immune system. Nat Plants 2:16128
Huang PY, Yeh YH, Liu AC et al (2014) The Arabidopsis LecRK-VI.2 associates with the pattern-recognition receptor FLS2 and primes Nicotiana benthamiana pattern-triggered immunity. Plant J 79:243–255
Huang S et al (2009) The genome of the cucumber, Cucumis sativus L. Nat Genet 41:1275–1281
Hurni S, Brunner S, Stirnweis D et al (2014) The powdery mildew resistance gene Pm8 derived from rye is suppressed by its wheat ortholog Pm3. Plant J 79:904–913
Hwang IS, Brady J, Martin GB, Oh CS (2017) Ser360 and Ser364 in the kinase domain of tomato SlMAPKKKalpha are critical for programmed cell death associated with plant immunity. Plant Pathol J 33:163–169
Kang HG, Oh CS, Sato M et al (2010) Endosome-associated CRT1 functions early in resistance gene-mediated defense signaling in Arabidopsis and tobacco. Plant Cell 22:918–936
Karimi M, Inze D, Depicker A (2002) GATEWAY vectors for Agrobacterium-mediated plant transformation. Trends Plant Sci 7:193–195
Kelly LJ, Leitch AR, Clarkson JJ et al (2013) Reconstructing the complex evolutionary origin of wild allopolyploid tobaccos (Nicotiana section Suaveolentes). Evolution 67:80–94
Kiba A, Nakano M, Vincent-Pope P et al (2012) A novel Sec14 phospholipid transfer protein from Nicotiana benthamiana is up-regulated in response to Ralstonia solanacearum infection, pathogen associated molecular patterns and effector molecules and involved in plant immunity. J Plant Physiol 169:1017–1022
Kim HS, Park SC, Ji CY et al (2016) Molecular characterization of biotic and abiotic stress-responsive MAP kinase genes, IbMPK3 and IbMPK6, in sweetpotato. Plant Physiol Biochem 108:37–48
Kim HS, Thammarat P, Lommel SA et al (2011) Pectobacterium carotovorum elicits plant cell death with DspE/F but the P. carotovorum DspE does not suppress callose or induce expression of plant genes early in plant-microbe interactions. Mol Plant Microbe Interact 24:773–786
Kim NH, Hwang BK (2015) Pepper heat shock protein 70a interacts with the type III effector AvrBsT and triggers plant cell death and immunity. Plant Physiol 167:307–322
Kim S et al (2014) Genome sequence of the hot pepper provides insights into the evolution of pungency in Capsicum species. Nat Genet 46:270–278. https://doi.org/10.1038/ng.2877
King SR, Mclellan H, Boevink PC et al (2014) Phytophthora infestans RXLR effector PexRD2 interacts with host MAPKKK epsilon to suppress plant immune signaling. Plant Cell 26:1345–1359
Knight MR, Read ND, Campbell AK, Trewavas AJ (1993) Imaging calcium dynamics in living plants using semi-synthetic recombinant aequorins. J Cell Biol 121:83–90
Kourelis J, Kaschani F, GrossHolz FM et al (2018) Re-annotated Nicotiana benthamiana gene models for enhanced proteomics and reverse genetics. bioRxiv: e373506
Kumar S, Stecher G, Suleski M, Hedges SB (2017) Timetree: a resource for timelines, timetrees, and divergence times. Mol Biol Evol 34(7):1812–1819
Lacombe S, Rougon-Cardoso A, Sherwood E et al (2010) Interfamily transfer of a plant pattern-recognition receptor confers broad-spectrum bacterial resistance. Nat Biotechnol 28:365–369
Lampropoulos A, Sutikovic Z, Wenzl C, Maegele I, Lohmann JU, Forner J (2013) GreenGate—a novel, versatile, and efficient cloning system for plant transgenesis. PLoS One 8:e83043
Lee JH, Kim YC, Choi D, Park JM (2013) Identification of novel pepper genes involved in Bax- or INF1-mediated cell death responses by high-throughput virus-induced gene silencing. Int J Mol Sci 14:22782–22795
Lei Y, Lu L, Liu H, Sen L, Xing F, Chen L (2014) CRISPR-P: a web tool for synthetic single-guide RNA design of CRISPR-system in plants. Mol Plant 7(9):1494–1496
Li D, Zhang H, Song Q et al (2015) Tomato Sl3-MMP, a member of the matrix metalloproteinase family, is required for disease resistance against Botrytis cinerea and Pseudomonas syringae pv. tomato DC3000. BMC Plant Biol 15:143
Li JF, Norville JE, Aach J et al (2013) Multiplex and homologous recombination-mediated genome editing in Arabidopsis and Nicotiana benthamiana using guide RNA and Cas9. Nat Biotechnol 31:688–691
Li X, Zhang Y, Huang L et al (2014a) Tomato SlMKK2 and SlMKK4 contribute to disease resistance against Botrytis cinerea. BMC Plant Biol 14:166
Li X, Zhang Y, Yin L, Lu J (2017) Overexpression of pathogen-induced grapevine TIR-NB-LRR gene VaRGA1 enhances disease resistance and drought and salt tolerance in Nicotiana benthamiana. Protoplasma 254:957–969
Li Y, Zhang L, Lu W, Wang X, Wu CA, Guo X (2014b) Overexpression of cotton GhMKK4 enhances disease susceptibility and affects abscisic acid, gibberellin and hydrogen peroxide signalling in transgenic Nicotiana benthamiana. Mol Plant Pathol 15:94–108
Liebrand TW, Smit P, Abd-El-Haliem A et al (2012) Endoplasmic reticulum-quality control chaperones facilitate the biogenesis of Cf receptor-like proteins involved in pathogen resistance of tomato. Plant Physiol 159:1819–1833
Lin CY, Tsai WS, Ku HM, Jan FJ (2012) Evaluation of DNA fragments covering the entire genome of a monopartite begomovirus for induction of viral resistance in transgenic plants via gene silencing. Transgenic Res 21:231–241
Lin PC, Hu WC, Lee SC et al (2015) Application of an integrated omics approach for identifying host proteins that interact with Odontoglossum ringspot virus capsid protein. Mol Plant Microbe Interact 28:711–726
Ling KS, Zhu HY, Gonsalves D (2008) Resistance to Grapevine leafroll associated virus-2 is conferred by post-transcriptional gene silencing in transgenic Nicotiana benthamiana. Transgenic Res 17:733–740
Liu C, Pedersen C, Schultz-Larsen T et al (2016) The stripe rust fungal effector PEC6 suppresses pattern-triggered immunity in a host species-independent manner and interacts with adenosine kinases. New Phytol. https://doi.org/10.1111/nph.14034
Lu W, Chu X, Li Y, Wang C, Guo X (2013) Cotton GhMKK1 induces the tolerance of salt and drought stress, and mediates defence responses to pathogen infection in transgenic Nicotiana benthamiana. PLoS One 8:e68503
Ludman M, Burgyan J, Fatyol K (2017) Crispr/Cas9 mediated inactivation of Argonaute 2 reveals its differential involvement in antiviral responses. Sci Rep 7:1010
Mafurah JJ, Ma H, Zhang M et al (2015) A virulence essential CRN effector of Phytophthora capsici suppresses host defense and induces cell death in plant nucleus. PLoS One 10:e0127965
Mantelin S, Peng HC, Li B, Atamian HS, Takken FL, Kaloshian I (2011) The receptor-like kinase SlSERK1 is required for Mi-1-mediated resistance to potato aphids in tomato. Plant J 67:459–471
Marks CE, Newbigin E, Ladiges PY (2011) Comparative morphology and phylogeny of Nicotiana section Suaveolentes (Solanaceae) in Australia and the South Pacific. Aust Syst Bot 24:61–86
Medina-Hernandez D, Rivera-Bustamante RF, Tenllado F, Holguin-Pena RJ (2013) Effects and effectiveness of two RNAi constructs for resistance to Pepper golden mosaic virus in Nicotiana benthamiana plants. Viruses 5:2931–2945
Miki D, Itoh R, Shimamoto K (2005) RNA silencing of single and multiple members in a gene family of rice. Plant Physiol 138:1903–1913
Montes C, Castro A, Barba P et al (2014) Differential RNAi responses of Nicotiana benthamiana individuals transformed with a hairpin-inducing construct during Plum pox virus challenge. Virus Genes 49:325–338
Naim F et al (2012) Advanced engineering of lipid metabolism in Nicotiana benthamiana using a draft genome and the V2 viral silencing-suppressor protein. PLoS One 7:e52717
Nakagawa T, Suzuki T, Murata S et al (2007) Improved Gateway binary vectors: high-performance vectors for creation of fusion constructs in transgenic analysis of plants. Biosci Biotechnol Biochem 71:2095–2100
Nakano M, Nishihara M, Yoshioka H et al (2013) Suppression of DS1 phosphatidic acid phosphatase confirms resistance to Ralstonia solanacearum in Nicotiana benthamiana. PLoS One 8:e75124
Nakasugi K, Crowhurst RN, Bally J, Wood C, Hellens RP, Waterhouse PM (2013) De novo transcriptome sequence assembly and analysis of RNA silencing genes of Nicotiana. PLoS One 8(3):e59534. https://doi.org/10.1371/journal.pone.0059534
Narusaka M, Kubo Y, Hatakeyama K et al (2013) Interfamily transfer of dual NB-LRR genes confers resistance to multiple pathogens. PLoS One 8:e55954
Nasir KH, Takahashi Y, Ito A et al (2005) High-throughput in planta expression screening identifies a class II ethylene-responsive element binding factor-like protein that regulates plant cell death and non-host resistance. Plant J 43:491–505
Nekrasov V, Staskawicz B, Weigel D, Jones JD, Kamoun S (2013) Targeted mutagenesis in the model plant Nicotiana benthamiana using Cas9 RNA-guided endonuclease. Nat Biotechnol 31:691–693
Nguyen HP, Chakravarthy S, Velasquez AC et al (2010) Methods to study PAMP-triggered immunity using tomato and Nicotiana benthamiana. Mol Plant Microbe Interact 23:991–999
Oh CS, Martin GB (2011) Tomato 14-3-3 protein TFT7 interacts with a MAP kinase kinase to regulate immunity-associated programmed cell death mediated by diverse disease resistance proteins. J Biol Chem 286:14129–14136
Ohtsu M, Shibata Y, Ojika M et al (2014) Nucleoporin 75 is involved in the ethylene-mediated production of phytoalexin for the resistance of Nicotiana benthamiana to Phytophthora infestans. Mol Plant Microbe Interact 27:1318–1330
Orchard AE (1999) A history of systematic botany in Australia. Australian Biological Resources Study/CSIRO, Canberra, pp 11–103. ISBN: 0643059652
Pais M, Win J, Yoshida K et al (2013) From pathogen genomes to host plant processes: the power of plant parasitic oomycetes. Genome Biol 14:211
Pavli OI, Kelaidi GI, Tampakaki AP, Skaracis GN (2011) The hrpZ gene of Pseudomonas syringae pv. phaseolicola enhances resistance to rhizomania disease in transgenic Nicotiana benthamiana and sugar beet. PLoS One 6:e17306
Pavli OI, Tampakaki AP, Skaracis GN (2012) High level resistance against rhizomania disease by simultaneously integrating two distinct defense mechanisms. PLoS One 7:e51414
Peng HC, Mantelin S, Hicks GR, Takken FL, Kaloshian I (2016) The conformation of a plasma membrane-localized somatic embryogenesis receptor kinase complex is altered by a potato aphid-derived effector. Plant Physiol 171:2211–2222
Petre B, Saunders DG, Sklenar J et al (2015) Candidate effector proteins of the rust pathogen Melampsora larici-populina target diverse plant cell compartments. Mol Plant Microbe Interact 28:689–700
Pfeilmeier S, Saur IM, Rathjen JP, Zipfel C, Malone JG (2016) High levels of cyclic-di-GMP in plant-associated Pseudomonas correlate with evasion of plant immunity. Mol Plant Pathol 17:521–531
Philips JG, Naim F, Lorenc MT, Dudley KJ, Hellens RP, Waterhouse PM (2017) The widely used Nicotiana benthamiana 16c line has an unusual T-DNA integration pattern including a transposon sequence. PLoS ONE 12:e0171311
Pombo MA, Zheng Y, Fernandez-Pozo N, Dunham DM, Fei Z, Martin GB (2014) Transcriptomic analysis reveals tomato genes whose expression is induced specifically during effector-triggered immunity and identifies the Epk1 protein kinase which is required for the host response to three bacterial effector proteins. Genome Biol 15:492
Pombo MA, Ramos RN, Zheng Y, Fei Z, Martin GB, Rosli HG (2019) Transcriptome-based identification and validation of reference genes for plant-bacteria interaction studies using Nicotiana benthamiana. Sci Rep 9(1):1632
Potato Genome Sequencing Consortium et al (2011) Genome sequence and analysis of the tuber crop potato. Nature 475:189–195
Rajput NA, Zhang M, Shen D et al (2015) Overexpression of a Phytophthora cytoplasmic CRN effector confers resistance to disease, salinity and drought in Nicotiana benthamiana. Plant Cell Physiol 56:2423–2435
Ramachandran SR, Yin C, Kud J et al (2017) Effectors from wheat rust fungi suppress multiple plant defense responses. Phytopathol 107:75–83
Reyes CA, Pena EJ, Zanek MC, Sanchez DV, Grau O, Garcia ML (2009) Differential resistance to Citrus psorosis virus in transgenic Nicotiana benthamiana plants expressing hairpin RNA derived from the coat protein and 54 K protein genes. Plant Cell Rep 28:1817–1825
Rodriguez PA, Stam R, Warbroek T, Bos JI (2014) Mp10 and Mp42 from the aphid species Myzus persicae trigger plant defenses in Nicotiana benthamiana through different activities. Mol Plant Microbe Interact 27:30–39
Rojas CM, Senthil-Kumar M, Wang K, Ryu CM, Kaundal A, Mysore KS (2012) Glycolate oxidase modulates reactive oxygen species-mediated signal transduction during nonhost resistance in Nicotiana benthamiana and Arabidopsis. Plant Cell 24:336–352
Rommens CMT, Salmeron JM, Oldroyd GED, Staskawicz BJ (1995) Intergeneric transfer and functional expression of the tomato disease resistance gene Pto. Plant Cell 7:1537–1544
Rosli HG, Zheng Y, Pombo MA et al (2013) Transcriptomics-based screen for genes induced by flagellin and repressed by pathogen effectors identifies a cell wall-associated kinase involved in plant immunity. Genome Biol 14:R139
Ruiz MT, Voinnet O, Baulcombe DC (1998) Initiation and maintenance of virus-induced gene silencing. Plant Cell 10:937–946
Ryu CM, Anand A, Kang L, Mysore KS (2004) Agrodrench: a novel and effective agroinoculation method for virus-induced gene silencing in roots and diverse Solanaceous species. Plant J 40:322–331
Samanta MK, Dey A, Gayen S (2016) CRISPR/Cas9: an advanced tool for editing plant genomes. Transgenic Res 25:561–573
Sato Y, Ando S, Takahashi H (2014) Role of intron-mediated enhancement on accumulation of an Arabidopsis NB-LRR class R-protein that confers resistance to Cucumber mosaic virus. PLoS One 9:e99041
Saur IM, Kadota Y, Sklenar J et al (2016) NbCSPR underlies age-dependent immune responses to bacterial cold shock protein in Nicotiana benthamiana. Proc Natl Acad Sci USA 113:3389–3394
Schmutz J et al (2010) Genome sequence of the palaeopolyploid soybean. Nature 463:178–183
Segonzac C, Feike D, Gimenez-Ibanez S, Hann DR, Zipfel C, Rathjen JP (2011) Hierarchy and roles of pathogen-associated molecular pattern-induced responses in Nicotiana benthamiana. Plant Physiol 156:687–699
Senthil G, Liu H, Puram VG, Clark A, Stromberg A, Goodin MM (2005) Specific and common changes in Nicotiana benthamiana gene expression in response to infection by enveloped viruses. J Gen Virol 86:2615–2625
Senthil-Kumar M, Mysore KS (2010) Assessing functional role of three water deficit stress-induced genes in nonhost disease resistance using virus-induced gene silencing in Nicotiana benthamiana. Plant Signal Behav 5:586–590
Senthil-Kumar M, Mysore KS (2012) Ornithine-delta-aminotransferase and proline dehydrogenase genes play a role in non-host disease resistance by regulating pyrroline-5-carboxylate metabolism-induced hypersensitive response. Plant Cell Environ 35:1329–1343
Senthil-Kumar M, Mysore KS (2014) Tobacco rattle virus-based virus-induced gene silencing in Nicotiana benthamiana. Nat Protoc 9:1549–1562
Shen D, Chai C, Ma L, Zhang M, Dou D (2016a) Comparative RNA-Seq analysis of Nicotiana benthamiana in response to Phytophthora parasitica infection. Plant Growth Regul 80:59–67
Shen L, Liu Z, Yang S et al (2016b) Pepper CabZIP63 acts as a positive regulator during Ralstonia solanacearum or high temperature-high humidity challenge in a positive feedback loop with CaWRKY40. J Exp Bot 67:2439–2451
Sierro N, et al. (2013) Reference genomes and transcriptomes of Nicotiana sylvestris and Nicotiana tomentosiformis. Genome Biol 14, R60 (2013)
Sierro N et al (2014) The tobacco genome sequence and its comparison with those of tomato and potato. Nat Commun 5:3833
Simao F et al. (2015) BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. 31, 3210–3212
Shibata Y, Kawakita K, Takemoto D (2010) Age-related resistance of Nicotiana benthamiana against hemibiotrophic pathogen Phytophthora infestans requires both ethylene- and salicylic acid-mediated signaling pathways. Mol Plant Microbe Interact 23:1130–1142
Song Q, Li D, Dai Y et al (2015) Characterization, expression patterns and functional analysis of the MAPK and MAPKK genes in watermelon (Citrullus lanatus). BMC Plant Biol 15:298
Stam R, Jupe J, Howden AJ et al (2013) Identification and characterisation CRN effectors in Phytophthora capsici shows modularity and functional diversity. PLoS One 8:e59517
Stemmer M, Thumberger T, del Sol Keyer M, Wittbrodt J, Mateo JL (2015) CCTop: an intuitive, flexible and reliable CRISPR/Cas9 target prediction tool. PLoS One. https://doi.org/10.1371/journal.pone.0124633
Stirnweis D, Milani SD, Jordan T, Keller B, Brunner S (2014) Substitutions of two amino acids in the nucleotide-binding site domain of a resistance protein enhance the hypersensitive response and enlarge the PM3F resistance spectrum in wheat. Mol Plant Microbe Interact 27:265–276
Stork W, Kim JG, Mudgett MB (2015) Functional analysis of plant defense suppression and activation by the Xanthomonas core type III effector XopX. Mol Plant Microbe Interact 28:180–194
Su Y, Xu L, Fu Z et al (2014) ScChi, encoding an acidic class III chitinase of sugarcane, confers positive responses to biotic and abiotic stresses in sugarcane. Int J Mol Sci 15:2738–2760
Su Y, Xu L, Wang S et al (2015) Identification, phylogeny, and transcript of chitinase family genes in sugarcane. Sci Rep 5:10708
Sun Y, Wang C, Yang B et al (2014) Identification and functional analysis of mitogen-activated protein kinase kinase kinase (MAPKKK) genes in canola (Brassica napus L.). J Exp Bot 65:2171–2188
Szczesniak MW, Makalowska I (2014) miRNEST 2.0: a database of plant and animal microRNAs. Nucleic Acids Res 42:D74–D77
Takahashi Y, Berberich T, Kanzaki H et al (2009) Serine palmitoyltransferase, the first step enzyme in sphingolipid biosynthesis, is involved in nonhost resistance. Mol Plant Microbe Interact 22:31–38
Takken FL, Luderer R, Gabriels SH et al (2000) A functional cloning strategy, based on a binary PVX-expression vector, to isolate HR-inducing cDNAs of plant pathogens. Plant J 24:275–283
Tanaka S, Ishihama N, Yoshioka H et al (2009) The Colletotrichum orbiculare SSD1 mutant enhances Nicotiana benthamiana basal resistance by activating a mitogen-activated protein kinase pathway. Plant Cell 21:2517–2526
Teper D, Salomon D, Sunitha S, Kim JG, Mudgett MB, Sessa G (2014) Xanthomonas euvesicatoria type III effector XopQ interacts with tomato and pepper 14-3-3 isoforms to suppress effector-triggered immunity. Plant J 77:297–309
Thiel H, Hleibieh K, Gilmer D, Varrelmann M (2012) The P25 pathogenicity factor of Beet necrotic yellow vein virus targets the sugar beet 26S proteasome involved in the induction of a hypersensitive resistance response via interaction with an F-box protein. Mol Plant Microbe Interact 25:1058–1072
Todesco M, De Felippes FF (2016) Why Benthamiana went viral. Trends Plant Sci 21:4–6
Tomato Genome Consortium (2012) The tomato genome sequence provides insights into fleshy fruit evolution. Nature 485:635–641
Turnbull D, Yang L, Naqvi S et al (2017) RXLR effector AVR2 up-regulates a brassinosteroid-responsive bHLH transcription factor to suppress immunity. Plant Physiol 174:356–369
Velasco R et al (2010) The genome of the domesticated apple (Malus x domestica Borkh.). Nat Genet 42:833
Velasquez AC, Chakravarthy S, Martin GB (2009) Virus-induced gene silencing (VIGS) in Nicotiana benthamiana and tomato. J Vis Exp 28:1292
Wagaba H, Patil BL, Mukasa S, Alicai T, Fauquet CM, Taylor NJ (2016) Artificial microRNA-derived resistance to Cassava brown streak disease. J Virol Methods 231:38–43
Wang Y, Nsibo DL, Juhar HM, Govers F, Bouwmeester K (2016) Ectopic expression of Arabidopsis L-type lectin receptor kinase genes LecRK-I.9 and LecRK-IX.1 in Nicotiana benthamiana confers Phytophthora resistance. Plant Cell Rep 35:845–855
Wei CF, Kvitko BH, Shimizu R et al (2007) A Pseudomonas syringae pv. tomato DC3000 mutant lacking the type III effector HopQ1-1 is able to cause disease in the model plant Nicotiana benthamiana. Plant J 51:32–46
Wu C, Jia L, Goggin F (2011) The reliability of virus-induced gene silencing experiments using tobacco rattle virus in tomato is influenced by the size of the vector control. Mol Plant Pathol 12:299–305
Xiang J, Li X, Wu J, Yin L, Zhang Y, Lu J (2016) Studying the mechanism of Plasmopara viticola RxLR effectors on suppressing plant immunity. Front Microbiol 7:709
Xu G, Li S, Xie K et al (2012) Plant ERD2-like proteins function as endoplasmic reticulum luminal protein receptors and participate in programmed cell death during innate immunity. Plant J 72:57–69
Xu L, Zhang W, He X et al (2014) Functional characterization of cotton genes responsive to Verticillium dahliae through bioinformatics and reverse genetics strategies. J Exp Bot 65:6679–6692
Yin K, Tang Y, Zhao J (2015) Genome-wide characterization of miRNAs involved in N gene-mediated immunity in response to Tobacco mosaic virus in Nicotiana benthamiana. Evol Bioinform Online 11:1–11
Yin W, Dong S, Zhai L, Lin Y, Zheng X, Wang Y (2013) The Phytophthora sojae Avr1d gene encodes an RxLR-dEER effector with presence and absence polymorphisms among pathogen strains. Mol Plant Microbe Interact 26:958–968
Yu X, Tang J, Wang Q et al (2012) The RxLR effector Avh241 from Phytophthora sojae requires plasma membrane localization to induce plant cell death. New Phytol 196:247–260
Zhang H, Dong S, Wang M et al (2010) The role of vacuolar processing enzyme (VPE) from Nicotiana benthamiana in the elicitor-triggered hypersensitive response and stomatal closure. J Exp Bot 61:3799–3812
Zhang H, Teng W, Liang J et al (2016) MADS1, a novel MADS-box protein, is involved in the response of Nicotiana benthamiana to bacterial harpinXoo. J Exp Bot 67:131–141
Zhang H, Wang M, Wang W et al (2012) Silencing of G proteins uncovers diversified plant responses when challenged by three elicitors in Nicotiana benthamiana. Plant Cell Environ 35:72–85
Zhang M, Ahmed Rajput N, Shen D et al (2015) A Phytophthora sojae cytoplasmic effector mediates disease resistance and abiotic stress tolerance in Nicotiana benthamiana. Sci Rep 5:10837
Zhao J, Liu Q, Zhang H, Jia Q, Hong Y, Liu Y (2013) The rubisco small subunit is involved in tobamovirus movement and Tm-2(2)-mediated extreme resistance. Plant Physiol 161:374–383
Zhao M, San Leon D, Mesel F, Garcia JA, Simon-Mateo C (2015) Assorted processing of synthetic trans-acting siRNAs and its activity in antiviral resistance. PLoS One 10:e0132281
Zhou B, Zeng L (2017) Elucidating the role of highly homologous Nicotiana benthamiana ubiquitin E2 gene family members in plant immunity through an improved virus-induced gene silencing approach. Plant Methods 13:59
Zhu F, Xi DH, Yuan S, Xu F, Zhang DW, Lin HH (2014) Salicylic acid and jasmonic acid are essential for systemic resistance against Tobacco mosaic virus in Nicotiana benthamiana. Mol Plant Microbe Interact 27:567–577
Zhu X, Caplan J, Mamillapalli P, Czymmek K, Dinesh-Kumar SP (2010) Function of endoplasmic reticulum calcium ATPase in innate immunity-mediated programmed cell death. EMBO J 29:1007–1018
Zhuang J, Coates CJ, Mao Q, Wu Z, Xie L (2016) The antagonistic effect of Banana bunchy top virus multifunctional protein B4 against Fusarium oxysporum. Mol Plant Pathol 17:669–679
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Pombo, M.A., Rosli, H.G., Fernandez-Pozo, N., Bombarely, A. (2020). Nicotiana benthamiana, A Popular Model for Genome Evolution and Plant–Pathogen Interactions. In: Ivanov, N.V., Sierro, N., Peitsch, M.C. (eds) The Tobacco Plant Genome. Compendium of Plant Genomes. Springer, Cham. https://doi.org/10.1007/978-3-030-29493-9_14
Download citation
DOI: https://doi.org/10.1007/978-3-030-29493-9_14
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-29492-2
Online ISBN: 978-3-030-29493-9
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)