Abstract
Scavenger receptors constitute a group of receptors on the cell surface that attach to various ligands and remove the targets that are non-self or altered. Signaling, transport, endocytosis, phagocytosis, and adhesion resulting in the removal of harmful and degraded substances are some functions of these receptors. Scavenger receptors bind a large repertoire of ligands indicating their involvement in homeostasis and multiple disease pathologies. In this chapter, we describe the role of scavenger receptor group in the pathogenesis of infections and cancer. In addition, we present a variety of ligands with their scavenger receptor binding strategies through different examples of targeted drug delivery systems.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
Scavenge means to clear, accordingly the role played by the scavenger receptors is clearing the body of a variety of moieties, for instance, modified low-density lipoprotein (LDL), bacteria or infected RBCs, apoptotic cells, etc. [1]. The receptor was first identified by Brown and Goldstein in macrophages and they observed that while the receptor internalized and degraded modified and oxidized low-density lipoprotein (OxLDL) or acetylated LDL , native LDL was spared by these receptors. Intracellular internalization of modified LDL may be due to foam cell formation [2]. Such foam cells loaded with cholesterol are integral to the atherosclerotic plaques and are also located in the lesions of blood vessel walls [3, 4]. While the scavenger receptors play a crucial physiological role, they can also perform as mediators in various pathologies. This chapter details the receptor with special emphasis on exploiting the endocytic property of this receptor in the targeted therapy of various diseases.
2 Scavenger Receptors
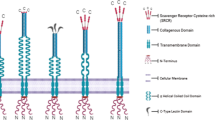
Scavenger receptors encompass a group of membrane proteins along with isoforms and soluble secreted extracellular domain isoforms. Although scavenger receptors are divided into 12 classes A-L (Fig.10.1), a term superfamily is not bestowed, as the receptors reveal no structural homology among the different classes [5]. They are more aptly termed as a supergroup [6]. Though structurally dissimilar, the scavenger receptors show affinity for similar ligands comprising of polyions including lipoproteins, phospholipids, cholesterol ester, apoptotic cells, carbohydrates, proteoglycans, and ferritin. A structural similarity is evident among different members of a class. Owing to their diverse ligand binding ability, the scavenger receptors represent a significant part of the pattern recognition receptors [7, 8]. A schematic representation of scavenger receptor classes and their recognition domains is depicted in Fig.10.1. A major focus of this chapter is on Class A and B scavenger receptors, the receptors that play a role in infections and cancer.
2.1 Scavenger Receptors’ Recognition Domains
2.1.1 Class A
Class A scavenger receptor comprises of type II transmembrane proteins. A cytoplasmic N-terminal domain (40–55 amino acids) is linked to the transmembrane region (26 amino acids). The extracellular domain comprises of three domains, namely, α-helical coiled-coil, C-terminal cysteine-rich (CR), and collagen-like domain and mediates ligand recognition. The unique collagen-like domain has positively charged amino acid residues that bind to polyanions [9,10,11]. The SR-AI/II, MARCO, SCARA5, and SRCL are most widely studied members of this class (Fig.10.1). SR-AI and AII display identical affinity for collagen-rich region [12]. MARCO exhibits an extended collagen-rich domain and expresses cysteine-rich domain as ligand-binding site [13]. SCARA-5 and MARCO receptors reveal a similar ligand binding. The coiled-coil domain is absent in these two receptors [14]. SRCL comprises of a C-terminal lectin-type domain while it lacks cysteine-rich domain [15].
2.1.2 Class B
The members of this class usually contain type III transmembrane proteins of 450–500 amino acid residues. They mainly express 2 transmembrane regions which contain closely placed short intracellular N- and C-terminals with the central extracellular loop comprising N-linked glycosylated domain of 400–450 amino acid residues, involved in ligand recognition [6]. The CD36 and SR-BI are two major members, which are largely glycosylated and fatty acylated [16, 17].
The structural dissimilarity is evident among different classes of scavenger receptors. Class C is not expressed in humans [18]. Class D scavenger receptors contain lysosome-associated membrane glycoprotein (LAMP) domains and mucin-like domains [19], whereas lectin-like LDLR-1, the only member of class E, shows C-type lectin domain. The C-terminal of this domain is connected by transmembrane domain to the cytoplasmic domain of N-terminal [20]. Class F scavenger receptors revealed growth factor domains, while class G receptors exhibit along with a chemokine domain and a mucin-like glycosylated stem as extracellular domain for ligand binding [21]. Class H scavenger receptors comprise of fascillin, epidermal growth factor (EGF) like, and lamin-type EGF-like (FEEL) domain [22]. While class I scavenger receptors consist of multiple group B cysteine-rich domains in their extracellular domain [23], class J contains a single transmembrane domain that connects the amino-terminal ligand recognition and binding ectodomain with a short cytoplasmic domain [24]. The class K scavenger receptor consists of hyaluronan binding domain [25, 26] and class L scavenger receptor consists of ligand-binding repeat, EGF repeat, and β propeller domain [27, 28]. A detailed description of these classes can be accessed from the literature [1, 5, 29].
3 Ligand Binding
Although majority of polyanionic ligands bind to scavenger receptors, their specificity depends on scavenger receptor domains. The broad range of specificity of the scavenger receptors prompted scientists to study the active site of these receptors. The positively charged C-terminal of the collagenous domain is essential for binding of ligands. Binding studies suggest that the collagenous domain is responsible for the broad specificity of the receptor [30]. A sticky surface is provided by the collagenous domain that enables selective binding of polyanions with high affinity. The positively charged residues of this domain are important for binding of polyanions. Presence of few negatively charged residues repels polyanions with low affinity and binds only those with high affinity. A direct or indirect effect on ligand binding is shown by other extracellular domains [31].
Although structurally homologous, SR-A1 and MARCO exhibit ligand uptake by discrete mechanisms. Studies suggest that removal of the cysteine-rich domain of MARCO curbs the internalization, whereas an enhanced uptake was seen following CR domain deletion of SR-A1 [32]. A difference in domain charge may have resulted in this consequence. A negatively charged CR domain is predicted by in silico studies. However, some studies report a mixed positive and negative charge for CR domain in MARCO. Such differences in charges could impact the recognition of pathogens and particulate carriers. Ligand receptor binding of MARCO is dependent on metal ions like calcium. Calcium binding and reduced electrostatic potential at the acidic amino acids enable interaction of MARCO with polyanions [33]. Electrostatic potential changes can also alter the stationing of MARCO domains, in turn affecting ligand binding. A high affinity of CD36 of class B to long-chain fatty acids enables fatty acid transport [29, 34].
4 Intracellular Internalization
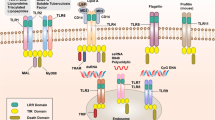
Scavenger receptors based on their class exhibit different endocytic mechanisms. While SR-A receptors follow clathrin-dependent pathways, LOX-1 proceeds via clathrin-independent pathways. Lipid raft-mediated mechanisms are shown by class B scavenger receptors (Fig.10.2). This endocytosis diversity of scavenger receptors is mainly associated with their sequence diversity and a wide variety of endocytic motifs present in cytoplasmic domains of each scavenger receptor.
The ligand binding to scavenger receptor mediates receptor-mediated endocytosis of this scavenger receptor–ligand complex, followed by intracellular trafficking via endosome lysosome system resulting in the metabolism of ligand. Scavenger receptor-mediated endocytosis of ligands stimulates the cascade of intracellular signaling. This leads to apoptosis, lipid peroxidation, and endothelial cell dysfunction. Monocyte infiltration accompanied by differentiation, which leads to foam cell formation, suggest a role in atherosclerotic plaque formation [6].
4.1 Caveolae/Clathrin-Dependent Pathway
A phagocytic cascade is triggered following internalization of modified LDL by SR-A. In the absence of ligands, unlike LDL receptor, the SR-A does not follow continuous cycling through a metabolic pathway. N-terminal cytoplasmic domain of SR-A contains di-leucine motif at amino acid residues 31 and 32, phosphorylation sites have been involved in ligand internalization and adhesion. This internalization of ligands follows classical coated pit pathway (Fig.10.2) [35].
4.2 Clathrin-Independent Pathway
The class E scavenger receptor LOX-1 binds to OxLDL, apoptotic bodies, and phospholipids and endocytoses via clathrin-independent pathway [36].
4.3 Lipid Raft Uptake
Class B scavenger receptor CD36 follows lipid rafts/caveolae-dependent pathway. Lipid rafts are mainly membrane domains containing lipids such as cholesterol, sphingolipids, glycosyl-phosphatidylinositol (GPI)-anchored proteins, and protein-tyrosine kinases of acylated src family. Caveolae present specialized raft subdomain for uptake mechanisms in some cells [37, 38].
5 Scavenger Receptor Location, Expression, and Function
Scavenger receptors are expressed mainly in endothelial cells (EDCs) and myeloid cells, but others are also expressed in epithelial cells. The SR-AI and AII are mostly expressed on macrophages, EDCs, epithelial cells, astrocytes, dendritic cells (DCs), mast cells, smooth muscle cells and mediates lipid metabolism, clearance of modified host components, pathogens, apoptotic cells, B cell–macrophage interactions, antigen presentation, binding of macrophages to extracellular matrix, and intracellular signaling [39,40,41]. MARCO is expressed by macrophages, EDCs, DCs, and astrocytes. Infectious stimuli express MARCO in most tissue macrophages. In DCs, antitumor response induces high-level expression of MARCO. MARCO also regulates the clearance of pathogens, necrotic dead cells, unopsonized particles, and enhances B cell–macrophage interaction [42]. SRCLI/II is mostly expressed by EDCs, stromal cells, astrocytes, and microglia, but not by macrophages. SRCL induces adherence of Lewis X-positive cells to vascular endothelium and elicits clearance of desialylated glycoproteins and β-amyloid [15]. Moreover, SCARA-5, a class A receptor is mostly expressed on epithelial cells of testis, airways, thymus, and adrenal glands. SCARA-5 lacks ability to recognize modified LDL and thus not involved in its endocytosis [14].
CD36 is mostly expressed by myeloid cells, platelets, adipocytes, and EDCs. Monocyte differentiation upregulates CD36 level, a mechanism similar to SR-A. The class B receptors induce lipid transfer activity, clearance of apoptotic cells, and P. falciparum-infected erythrocytes. SR-BI is found on monocytes, DCs, liver cells, and adrenal glands [43]. Class C is not found in humans and expressed on macrophages of the Drosophila and Mosquitoes [44]. The class D CD68 scavenger receptor shows intracellular expression in macrophages, and surface expression on dendritic cells and osteoclasts [45]. Moreover, scavenger receptor class E (LOX-1) is mostly found on EDCs, in various diseased conditions is expressed in smooth muscle cells. Furthermore, LOX-1 is involved in induction of apoptosis of EDCs, monocyte adhesion to EDCs, release of proinflammatory cytokines, and increase in ROS production [46]. The class F receptors are expressed over EDCs, macrophages and are involved in clearance of modified host components, antigen clearance and cross-presentation [47]. The class G scavenger receptors are expressed over macrophages, dendritic cells, and also expressed in multiple organs [48]. The class H scavenger receptors are mostly expressed in EDCs of liver, spleen, and lymphatic system, whereas macrophages only express FEEL-1. The class H facilitates lymphocyte adhesion and transmigration, clearance of modified lipoproteins and apoptotic cells, induces angiogenesis, and is involved in intracellular trafficking [49]. However, Class I CD163 receptor is mainly expressed on myeloid cells and mediates clearance of hemoglobin (Hb):haptoglobin (Hp) complexes, and aids erythroblast adhesion to macrophages [6]. Other classes of scavenger receptors such as class J, K, and L are still in research stage, in which class J is mainly expressed on neurons, class K on macrophages, and class L on kidney proximal tubule cells, lung, thyroid, gallbladder, neuroepithelium, epididymis, prostate, ovaries, uterus, and blood–brain barrier. They are mainly involved in the clearance of extracellular matrix ligands [5]. Although different types of scavenger receptors are expressed at the same site, they show diversity in intracellular trafficking and consequently elicit different responses.
6 Pathophysiological Features
SR-AI/II plays a major role in innate immunity against bacterial infections, where they recognize polyanionic cell wall products of bacteria including lipopolysaccharide (LPS) and lipoteichoic acid (LTA). They mediate unopsonized phagocytosis of Gram-positive bacteria. This innate immune response stimulates scavenger receptor and enhances recognition and rapid internalization of pathogenic materials, thereby playing a role in the host defense mechanism [50].
SR-BI is involved in several processes such as apoptosis, binding and internalization of pathogens, and signaling for induction of anti-inflammatory response. Microorganisms supported by anti-inflammatory activity of SR-BI undergo internalization via multimolecular pathways. This was elucidated based on observations in infectious diseases caused by Gram-positive and -negative bacteria, as also infections caused by dengue virus, hepatitis C virus, Plasmodium species, and many other infectious agents. SR-BI is also involved in the clearance of microbial end products. Binding of SR-BI to lipopolysaccharide is reported [50].
Involvement of scavenger receptors in the regulation of cancer tumor growth and associated immune reactions is reported. Tumor-associated macrophages (TAMs) show elevated levels of SR-A. An overexpression of SR-BI on cancer cell lines is observed. This results in increased lipid uptake in tumor cells, thus promoting growth [51]. Yet another interesting mechanism by which SR-BI increases tumor proliferation is the intracellular signaling cascade involving activation of the PI3K/AKT pathway, thereby causing tumor growth [52, 53].
Scavenger receptors are extensively studied in atherosclerosis, where SR-A and CD36 induce modified LDL uptake which is associated with foam cell formation [54]. On the other hand, SR-BI mediates cholesterol transport which is responsible for its anti-atherogenic role [55]. Although cells present in atherosclerotic lesions expressed LOX-1 and CD68, their exact role in response to atherogenesis is yet to be confirmed.
Interestingly, scavenger receptors are reported to play an important role in the pathophysiology of Alzheimer’s disease, where they exhibited potential endocytosis of β-amyloid fibrils [56].
7 Ligands
Majority of scavenger receptors are able to bind bacteria, virus, and cell surface components. They showed effective binding with polynucleotides, sulfated polysaccharides, and long-chain fatty acids [40]. Almost all scavenger receptors mediate modified LDL uptake. The members of this supergroup such as SR-A, SR-B, and SR-E show efficient binding to both OxLDL and AcLDL, whereas SR-H only mediate AcLDL uptake. Among all, class B receptors bind to unmodified LDL, HDL, and VLDL. Such differential binding dictates their functional diversity in the clearance of modified LDL. The scavenger receptor expresses positively charged amino acids cluster (arginine or lysine) which thereby facilitates ionic interaction with negatively charged polyanionic ligands, lipoprotein particles, and pathogenic materials such as lipopolysaccharide and lipoteichoic acid. The ligand-binding potential of scavenger receptors increases with enhanced oligomer expression [29, 31, 41]. The numerous ligands for different scavenger receptor classes are summarized in Table 10.1.
Class A recognizes lipidic and apolipoprotein functionalities expressed by modified LDL [57]. This class of scavenger receptors exhibits neuronal cytotoxicity by mediating the uptake of β-amyloid fibrils in microglia and thereby contributes to the pathophysiology of Alzheimer’s disease [58]. The affinity of extracellular ligands biglycan and decorin to SR-A induces association of macrophages in the extracellular matrix of smooth muscle cells involved in atherosclerotic plaque formation. Similarly, advanced glycation end products (AGE) efficiently endocytosed by SR-A are released during inflammation. On the other hand, SR-A also mediates uptake of glycated proteins such as glycated collagen IV. Expression of SR-A on adenocarcinoma cells mediates internalization of T-cell tumor antigen and thus plays an important role in cancer pathology. During lung inflammation, MARCO mediates uptake of uteroglobin-related protein-1 (UGRP-1). Besides MARCO another member of class A, SRCL-I recognizes Lewis-X trisaccharides with high affinity and dictates its role in recognizing desialylated glycoproteins. SRCL-I also recognizes β-amyloid peptide in Alzheimer’s patients [5].
Class B receptors show diverse ligand specificity where they bind native lipoprotein particles and hypochlorite modified LDL which are found in atherosclerotic lesions. CD36 present in vascular endothelial cells mediates hexarelin uptake and causes vasoconstriction. Furthermore, this CD36 also binds collagen type I, AGE-modified BSA, and β-amyloid fibrils. It also recognizes oxidized phospholipids expressed on apoptotic cells, thereby mediating macrophage clearance. SRBI receptor of class B, binds AcLDL with greater affinity. It also mediates uptake of native lipoprotein particles and recognizes expressed apolipoprotein components. It also binds AGE-BSA and β-amyloid fibers [5, 29, 57, 58].
Class C receptor binds to AcLDL and pathogens, whereas class D (CD68) binds to OxLDL and negatively charged phosphatidylserine-rich liposomes. The Class E (LOX-1) binds to OxLDL, fibronectin, phosphatidylserine, AGE-modified protein and clears apoptotic cells. LOX-1 mediates Hsp70 internalization in dendritic cells, which is not endocytosed by class A & class B receptors. Both Class F receptors, SREC-I and SREC-II, are involved in the uptake of modified LDL, and mediate recognition of other ligands such as calreticulin, molecular chaperones, gp96, and tumor released heat shock proteins (Hsp70), whereas they lack recognition for AGE-modified proteins.
Class G (SRPSOX) receptor-like LOX1 (Class E) binds to OxLDL but not to AcLDL and it also acts as chemokine ligand for CXC chemokine receptor 16, thereby mediating adhesion of DCs to T cells and NK cells. In class H, FEEL-1 receptor binds to extracellular SPARC (secreted protein acidic and rich in cysteine) glycoprotein and SI-CLP (stabilin-1 interacting chitinase-like protein) sorted in macrophages and Hsp70, while they show poor recognition for AGE-BSA. However, FEEL-2 recognizes hyaluronic acid and AGE-BSA with high affinity [5, 6, 29].
Scavenger receptors bind to foreign ligands including bacteria, fungi, virus, and parasites. In case of pathogens, their extracellular expression containing lipopolysaccharides, lipoteichoic acid, C-reactive protein, endotoxins, and numerous other surface proteins are recognized by these receptors [59]. Researchers have studied various other ligands based on functionality for scavenger receptor targeting, which is summarized in Table 10.2.
8 Receptor-Mediated Targeting Strategies
Endocytic uptake mechanism of scavenger receptor suggests a simplistic way of receptor-mediated targeted drug delivery. The drug gets released intracellularly after efficient internalization of the ligand–receptor complex. This interaction also induces the cascade of inflammatory responses. Although there exist many different scavenger receptor classes, till date, only A and B scavenger receptors have been studied for nanomedicine-mediated response. These scavenger receptors serve as a unique nanomedicine target also for many theranostic applications. Research directed towards site-specific targeted drug delivery through scavenger receptor using various nanoformulations relies on receptor-specific ligands carrier compositions mainly involving polyanions [101].
8.1 Drug–Ligand Conjugates
The chemical coupling of a suitable drug to a scavenger receptor-specific ligand such as maleylated albumin (MBSA) increases recognition by scavenger receptors for high-affinity binding, thereafter it undergoes internalization and metabolically degraded in lysosomes to release free drug for activity. Almost 100-fold enhanced efficacy was observed with such drug–ligand conjugate when studied in leishmaniasis, tuberculosis, and neoplastic conditions. Coupling of methotrexate (MTX) to MBSA exhibited rapid internalization inside leishmania-infected hamster peritoneal macrophages and demonstrated 100-fold antileishmanial effect compared to free drug. It also eliminated intracellular amastigotes of L. donovani and L. mexicana amazonesis. However, in case of M. tuberculosis-infected macrophages the targeting of anti-tubercular drug p-amino salicylic acid (PAS) and MBSA conjugate resulted in only 50% reduction of colony-forming units (CFUs). However, compared to free drug which exhibited CFU reduction of 0.5%, the enhancement in efficacy was nearly 100-fold. In neoplastic condition, the conjugation of drug Daunomycin with MBSA exhibited 100-fold cytotoxicity over free Daunomycin when tested at low concentration of 0.1 μM. Similar cytotoxic results were found with Doxorubicin-MBSA conjugate when tested on human histiocytic lymphoma cells [102].
8.2 Liposomes
Negatively charged phospholipids such as phosphatidylserine (PS) and phosphatidylglycerol (PG) are efficiently recognized by macrophages [103]. The studies confirmed that negatively charged liposomes are efficiently taken up by macrophage scavenger receptors over neutral or cationic liposomes [104,105,106,107]. When macrophage cells which expressed scavenger receptor were treated with negatively charged PS-liposomes and neutral PC-liposomes, the former exhibited 5.3-fold enhanced macrophage uptake over PC liposomes [108]. Incorporation of dicetylphosphate (DCP) also induced net negative surface charge over liposomes [109]. Polyaconitylated-human serum albumin (Aco-HSA) surface anchored liposomes showed effective anti-HIV 1 activity due to high uptake by scavenger receptors expressed on sinusoidal cells. This conjugation of Aco-HSA to liposomes enhanced liver uptake 17-fold, as compared with control liposomes, and the Aco-HSA liposomes were mostly found in liver EDCs and kupffer cells. Further, in this study, reduced liver uptake (24%) of Aco-HSA was found post-injection of polyinosinic acid, which is a known SR ligand [110].
In case of stealth liposomes, endocytic CD163 scavenger receptor enhanced uptake of monoclonal antibody loaded pegylated liposomes in CD163 transfected cells and macrophages [111]. Palmitoyloleoyl-phosphatidylcholine (POPC)-apoE liposomes functionalized using different apoE proteins (apoE2, apoE3, apoE4, apoE165, apoE202, apoE229, and apoE259) enhanced scavenger receptor B binding affinity and were thought to regulate brain cholesterol metabolism [112]. Liposomes carrying fluorescently labeled cholesterol when tested on HepG2 cells (model system for human hepatocytes) showed 20% binding for class B scavenger receptor and only 10% recognition was confined to low-density lipoprotein receptor (LDLR) which provided additional insights for scavenger receptor-mediated uptake of liposomes [113]. It was reported that, in certain cells, liposome uptake is not inhibited by known scavenger receptor ligands suggesting their uptake was not scavenger receptor-mediated. PS-containing liposomes showed enhanced uptake in an African green monkey kidney cell line (CVI) compared to phosphatidylcholine (PC) liposomes, independent of inhibition by known competitors for scavenger receptor [polyinosinic acid or dextran sulfate]. On the other hand, in case of murine macrophage cell line, PS-liposome uptake was inhibited competitively by polyinosinic acid, but not by polycytidylic acid [114]. The liposomes for targeted scavenger receptor delivery in various diseased conditions are described in Table 10.3.
8.3 Nanoparticles
8.3.1 Lipoprotein Particles
In one study, it was found that OxLDL exhibited stronger CD36 binding and HDL has stronger SR-BI binding ability among all lipoproteins [124]. Synthesized HDL nanoparticles also revealed high affinity for SR-BI-rich cancer cells. Furthermore, HDL nanoparticles mediated delivery of siRNA to the cancer tumors overexpressed with SR-BI. Similarly, these HDL nanoparticles exhibited SR-BI-mediated paclitaxel delivery to prostate cancer cells. Such studies proved the potential of SR-BI-mediated targeting of nanoparticles and their subsequent involvement in many disease states [125]. Administration of acylated LDL particles loaded with muramyl tripeptide mediated antitumor efficacy through the scavenger receptors [126]. In one study, the antinociceptive activity of apo lipoprotein functionalized loperamide-loaded albumin nanoparticles was assessed. Three apo lipoproteins E3, A-I, and B-100 exhibited 95%, 65%, and 50% antinociceptive activity, respectively, whereas plain loperamide solution showed no effect, which showed uptake of such particles through SR-BI receptor expressed at BBB [127].
8.3.2 Inorganic Nanoparticles
Dextran sulfate-mediated macrophage uptake of silver nanoparticles (AgNPs) through scavenger receptor is reported. Scavenger receptor-mediated uptake of AgNPs resulted in their intracellular accumulation and thereby apoptosis [128]. Furthermore, protein functionalization of AgNPs reduced its uptake due to decreased surface charge [129, 130]. Inhalation of ZnO nanoparticles induced enhanced expression of both SR-A and SR-B and thereby influenced atherosclerotic disease progression. However, TiO2 nanoparticles did not exhibit the same mechanism [89]. The macrophage phagocytic activity was diminished when subjected to superparamagnetic iron oxide nanoparticles exposure. These iron oxide nanoparticles inhibited macrophage activation for M1 to M2 state and enhanced TNF-α production [33, 88].
8.3.3 Miscellaneous
Gadolinium-containing nanomedicines with anti-CD36 antibodies were efficiently taken up by macrophages in vitro compared to nanomedicines without CD36 antibodies [131]. The scavenger receptor A class member MARCO also exhibited interaction with carbon nanotubes [132] and polystyrene nanoparticles [133]. Furthermore, when scavenger receptor A was overexpressed in human embryonic kidney 293 (HEK293) cells, a cell line which is normally devoid of scavenger receptor expression, elicited enhanced uptake of amorphous silica nanoparticles, demonstrating role of scavenger receptor in uptake of nanomedicines [101,134]. The miscellaneous nanoparticles targeting scavenger receptors are given in Table 10.4.
9 Clinical Trials
Although targeting to scavenger receptor is still in its nascent stage, very few scavenger receptor-mediated delivery systems have reached clinical trials. Herein we discuss such clinical studies with their outcome, demonstrating the role of scavenger receptors and targeting strategies. A study exploring targeting of pegylated interferon α2 plus ribavirin therapy to SR-BI receptor encoded by SCARB1 gene for hepatitis C virus studied the association of single nucleotide polymorphism (SNP) of SCARB1 gene and its response to therapy, where they found SNP may increase antiviral therapy outcome [136, 137]. Interestingly another study was conducted to understand underlying molecular mechanisms causing disruption to HDL regulation through scavenger receptor (SR-BI) in various metabolic diseases including atherosclerosis, where genotype modification affects HDL metabolism and cholesterol homeostasis [138]. One study was conducted to assess the role of scavenger receptor ligands as biomarkers for cardiovascular disease diagnosis, where oxidized phospholipids and apolipoprotein B identification by antibodies can be detected to predict cardiovascular disease state 15 years in advance. The receptor studied here was CD36 [139]. Studies were also conducted for anti-hepatitis C virus efficacy testing of a new molecule ITX 5061 by blocking the virus uptake through scavenger receptor (SR-BI) expressed on hepatocytes to reduce the infection chances in liver transplant patients [140]. No clinical trials are however evident on scavenger receptor-targeted drug delivery.
10 Advantages and Limitations
Targeting scavenger receptors offers great promise for improved therapeutic efficacy. This receptor has recognition specificity for pathogenic materials and plays an important role in various disease conditions. Intracellular delivery of actives can be achieved through scavenger receptor-mediated drug delivery, as the majority of infections are intracellular.
The major limitation of targeting these receptors is their broad ligand binding and recognition including both endogenous and exogenous molecules, which will compete for receptor-mediated endocytosis. Another major challenge is immunogenicity as these receptors are involved in inflammation and expressed on immune cells. Furthermore, they are widely expressed on majority of cell types; hence, specificity is a challenge.
11 Future Perspectives
Recently, newer classes of scavenger receptors were found and many more are still to be discovered, hence targeting to these receptors can provide newer avenues in site-specific drug delivery. Exploitation of scavenger receptor-mediated drug delivery is an option of the future. SR-BI due to its overexpression is reported as biomarker for human nasopharyngeal carcinoma [141]. The role of scavenger receptor as a biomarker for diagnosing various other disease conditions needs to be explored.
12 Conclusion
Scavenger receptors play a multifaceted and dynamic role in various cell-signaling pathways in the human body and are involved in metabolic regulation of macrophages for improved immune response. Scavenger receptors facilitate uptake of a broad spectrum of ligands including endogenous and foreign molecules. However, this receptor poses a challenge in stealth delivery of nanomedicines due to its inherent ability of scavenging numerous components. Targeted drug delivery using scavenger receptor is still in its nascent stage and can be further exploited for the treatment of infections and cancer.
Abbreviations
- AcLDL:
-
Acylated low-density lipoprotein
- Aco-HSA:
-
Polyaconitylated-human serum albumin
- AGE:
-
Advanced glycation end products
- AgNPs:
-
Silver nanoparticles
- BBB:
-
Blood–brain barrier
- BSA:
-
Bovine serum albumin
- CFUs:
-
Colony-forming units
- CR:
-
Cysteine rich
- CXC:
-
Chemokine receptor 16
- DCP:
-
Dicetylphosphate
- DCs:
-
Dendritic cells
- EDCs:
-
Endothelial cells
- EGF:
-
Epidermal growth factor
- Fe2O3:
-
Iron oxide
- FEEL:
-
Fasciclin EGF-like, and lamin-type EGF-like domains
- GPI:
-
Glycosyl-phosphatidylinositol
- HDL:
-
High-density lipoprotein
- Hsp:
-
Heat shock proteins
- LAMP:
-
Lysosome-associated membrane glycoprotein
- LCO:
-
Lithocholic oleate
- LDL:
-
Low-density lipoprotein
- LDLR:
-
Low-density lipoprotein receptor
- LOX-1:
-
Lectin-like oxidized LDL receptor-1
- LPS:
-
Lipopolysaccharide
- LTA:
-
Lipoteichoic acid
- MARCO:
-
Macrophage receptor with collagenous structure
- MBSA:
-
Maleylated albumin
- MTX:
-
Methotrexate
- NMs:
-
Nanomedicines
- NK cells:
-
Natural Killer cells
- OxLDL:
-
Oxidized low-density lipoprotein
- PAS:
-
p-amino salicylic acid
- PC:
-
Phosphatidylcholine
- PG:
-
Phosphatidylglycerol
- POPC:
-
Palmitoyloleoyl-phosphatidylcholine
- PS:
-
Phosphatidylserine
- RBCs:
-
Red blood cells
- ROS:
-
Reactive oxygen species
- S1-CLP:
-
Stabilin-1 interacting chitinase-like protein
- SCARA-5:
-
Scavenger receptor class A member 5
- SiRNA:
-
Small interfering ribonucleic acid
- SNP:
-
Single nucleotide polymorphism
- SPARC:
-
Secreted protein acidic and rich in cysteine
- SR:
-
Scavenger receptor
- SRCL:
-
Scavenger receptors with C-type lectin
- SRPSOX:
-
Scavenger receptor that binds phosphatidylserine and oxidized lipids
- TAMs:
-
Tumor-associated macrophages
- TiO2:
-
Titanium dioxide
- UGPR:
-
Uteroglobin-related protein
- VLDL:
-
Very low-density lipoproteins
- ZnO:
-
Zinc oxide
References
Canton J, Neculai D, Grinstein S. Scavenger receptors in homeostasis and immunity. Nat Rev Immunol. 2013;13(9):621–34.
Brown MS, Goldstein JL. Receptor-mediated endocytosis: insights from the lipoprotein receptor system. Proc Natl Acad Sci. 1979;76(7):3330–7.
Brown MS, Goldstein JL, Krieger M, Ho YK, Anderson RG. Reversible accumulation of cholesteryl esters in macrophages incubated with acetylated lipoproteins. J Cell Biol. 1979;82(3):597–613.
Su T, Zhao L, Ruan X, Zuo G, Gong J. Synergistic effect of scavenger receptor A and low-density lipoprotein receptor on macrophage foam cell formation under inflammatory stress. Mol Med Rep. 2013;7(1):37–42.
PrabhuDas MR, Baldwin CL, Bollyky PL, Bowdish DM, Drickamer K, Febbraio M, Herz J, Kobzik L, Krieger M, Loike J, McVicker B. A consensus definitive classification of scavenger receptors and their roles in health and disease. J Immunol. 2017;198(10):3775–89.
Zani I, Stephen S, Mughal N, Russell D, Homer-Vanniasinkam S, Wheatcroft S, Ponnambalam S. Scavenger receptor structure and function in health and disease. Cell. 2015;4(2):178–201.
Ashraf MZ, Sahu A. Scavenger receptors: a key player in cardiovascular diseases. Biomol Concepts. 2012;3(4):371–80.
de Winther MP, van Dijk KW, Havekes LM, Hofker MH. Macrophage scavenger receptor class A: a multifunctional receptor in atherosclerosis. Arterioscler Thromb Vasc Biol. 2000;20(2):290–7.
Wada Y, Doi T, Matsumoto A, Asaoka H, Honda M, Hatano H, Emi M, Naito M, Mori T, Takahashi K, Nakamura H. Structure and function of macrophage scavenger receptors. Ann N Y Acad Sci. 1994;748(1):226–38.
Bowdish DM, Gordon S. Conserved domains of the class A scavenger receptors: evolution and function. Immunol Rev. 2009;227(1):19–31.
Resnick D, Chatterton JE, Schwartz K, Slayter H, Krieger M. Structures of class A macrophage scavenger receptors electron microscopic study of flexible, multidomain, fibrous proteins and determination of the disulfide bond pattern of the scavenger receptor cysteine-rich domain. J Biol Chem. 1996;271(43):26924–30.
Kodama T, Freeman M, Rohrer L, Zabrecky J, Matsudaira P, Krieger M. Type I macrophage scavenger receptor contains α-helical and collagen-like coiled coils. Nature. 1990;343(6258):531–5.
Elomaa O, Sankala M, Pikkarainen T, Bergmann U, Tuuttila A, Raatikainen-Ahokas A, Sariola H, Tryggvason K. Structure of the human macrophage MARCO receptor and characterization of its bacteria-binding region. J Biol Chem. 1998;273(8):4530–8.
Jiang Y, Oliver P, Davies KE, Platt N. Identification and characterization of murine SCARA5, a novel class A scavenger receptor that is expressed by populations of epithelial cells. J Biol Chem. 2006;281(17):11834–45.
Nakamura K, Funakoshi H, Tokunaga F, Nakamura T. Molecular cloning of a mouse scavenger receptor with C-type lectin (SRCL), a novel member of the scavenger receptor family. Biochim Biophys Acta. 2001;1522(1):53–8.
Thorne RF, Meldrum CJ, Harris SJ, Dorahy DJ, Shafren DR, Berndt MC, Burns GF, Gibson PG. CD36 forms covalently associated dimers and multimers in platelets and transfected COS-7 cells. Biochem Biophys Res Commun. 1997;240(3):812–8.
Reaven E, Cortez Y, Leers-Sucheta S, Nomoto A, Azhar S. Dimerization of the scavenger receptor class B type I formation, function, and localization in diverse cells and tissues. J Lipid Res. 2004;45(3):513–28.
Pearson A, Lux AL, Krieger M. Expression cloning of dSR-CI, a class C macrophage-specific scavenger receptor from Drosophila melanogaster. Proc Natl Acad Sci. 1995;92(9):4056–60.
Jiang Z, Shih DM, Xia YR, Lusis AJ, de Beer FC, de Villiers WJ, van der Westhuyzen DR, de Beer MC. Structure, organization, and chromosomal mapping of the gene encoding macrosialin, a macrophage-restricted protein. Genomics. 1998;50(2):199–205.
Yamanaka S, Zhang XY, Miura K, Kim S, Iwao H. The human gene encoding the lectin-type oxidized LDL receptor (OLR1) is a novel member of the natural killer gene complex with a unique expression profile. Genomics. 1998;54(2):191–9.
Ishii J, Adachi H, Aoki J, Koizumi H, Tomita S, Suzuki T, Tsujimoto M, Inoue K, Arai H. SREC-II, a new member of the scavenger receptor type F family, trans-interacts with SREC-I through its extracellular domain. J Biol Chem. 2002;277(42):39696–702.
Adachi H, Tsujimoto M. FEEL-1, a novel scavenger receptor with in vitro bacteria-binding and angiogenesis-modulating activities. J Biol Chem. 2002;277(37):34264–70.
Rodamilans B, Muñoz IG, Bragado-Nilsson E, Sarrias MR, Padilla O, Blanco FJ, Lozano F, Montoya G. Crystal structure of the third extracellular domain of CD5 reveals the fold of a group B scavenger cysteine-rich receptor domain. J Biol Chem. 2007;282(17):12669–77.
Ibrahim ZA, Armour CL, Phipps S, Sukkar MB. RAGE and TLRs: relatives, friends or neighbours. Mol Immunol. 2013;56(4):739–44.
Naor D, Sionov RV, Ish-Shalom D. CD44: structure, function and association with the malignant process. Adv Cancer Res. 1997;71:241–319. Academic Press.
Dzwonek J, Wilczynski GM. CD44: molecular interactions, signaling and functions in the nervous system. Front Cell Neurosci. 2015;9:175.
Oh SJ, Kim TH, Lim JM, Jeong JW. Progesterone induces expression of Lrp2 in the murine uterus. Biochem Biophys Res Commun. 2013;441(1):175–9.
Bartolome F, Antequera D, Tavares E, Pascual C, Maldonado R, Camins A, Carro E. Obesity and neuroinflammatory phenotype in mice lacking endothelial megalin. J Neuroinflammation. 2017;14(1):26.
Plüddemann A, Neyen C, Gordon S. Macrophage scavenger receptors and host-derived ligands. Methods. 2007;43(3):207–17.
Ashkenas J, Penman M, Vasile E, Acton S, Freeman M, Krieger M. Structures and high and low affinity ligand binding properties of murine type I and type II macrophage scavenger receptors. J Lipid Res. 1993;34(6):983–1000.
Stephen SL, Freestone K, Dunn S, Twigg MW, Homer-Vanniasinkam S, Walker JH, Wheatcroft SB, Ponnambalam S. Scavenger receptors and their potential as therapeutic targets in the treatment of cardiovascular disease. Int J Hypertens. 2010;2010:646929.
Ojala JR, Pikkarainen T, Tuuttila A, Sandalova T, Tryggvason K. Crystal structure of the cysteine-rich domain of scavenger receptor MARCO reveals the presence of a basic and an acidic cluster that both contribute to ligand recognition. J Biol Chem. 2007;282(22):16654–66.
Chao Y, Makale M, Karmali PP, Sharikov Y, Tsigelny I, Merkulov S, Kesari S, Wrasidlo W, Ruoslahti E, Simberg D. Recognition of dextran–superparamagnetic iron oxide nanoparticle conjugates (Feridex) via macrophage scavenger receptor charged domains. Bioconjug Chem. 2012;23(5):1003–9.
Ibrahimi A, Abumrad NA. Role of CD36 in membrane transport of long-chain fatty acids. Curr Opin Clin Nutr Metab Care. 2002;5(2):139–45.
Chen Y, Wang X, Ben J, Yue S, Bai H, Guan X, Bai X, Jiang L, Ji Y, Fan L, Chen Q. The di-leucine motif contributes to class a scavenger receptor-mediated internalization of acetylated lipoproteins. Arterioscler Thromb Vasc Biol. 2006;26(6):1317–22.
Murphy JE, Vohra RS, Dunn S, Holloway ZG, Monaco AP, Homer-Vanniasinkam S, Walker JH, Ponnambalam S. Oxidised LDL internalisation by the LOX-1 scavenger receptor is dependent on a novel cytoplasmic motif and is regulated by dynamin-2. J Cell Sci. 2008;121(13):2136–47.
Zeng Y, Tao N, Chung KN, Heuser JE, Lublin DM. Endocytosis of oxidized low density lipoprotein through scavenger receptor CD36 utilizes a lipid raft pathway that does not require caveolin-1. J Biol Chem. 2003;278(46):45931–6.
Rahaman SO, Lennon DJ, Febbraio M, Podrez EA, Hazen SL, Silverstein RL. A CD36-dependent signaling cascade is necessary for macrophage foam cell formation. Cell Metab. 2006;4(3):211–21.
Hughes DA, Fraser IP, Gordon S. Murine macrophage scavenger receptor: in vivo expression and function as receptor for macrophage adhesion in lymphoid and non-lymphoid organs. Eur J Immunol. 1995;25(2):466–73.
Zingg JM, Ricciarelli R, Azzi A. Scavenger receptors and modified lipoproteins: fatal attractions. IUBMB Life. 2000;49(5):397–403.
Platt N, Gordon S. Scavenger receptors: diverse activities and promiscuous binding of polyanionic ligands. Chem Biol. 1998;5(8):R 193–203.
Elomaa O, Kangas M, Sahlberg C, Tuukkanen J, Sormunen R, Liakka A, Thesleff I, Kraal G, Tryggvason K. Cloning of a novel bacteria-binding receptor structurally related to scavenger receptors and expressed in a subset of macrophages. Cell. 1995;80(4):603–9.
Terpstra V, van Amersfoort ES, van Velzen AG, Kuiper J, van Berkel TJ. Hepatic and extrahepatic scavenger receptors: function in relation to disease. Arterioscler Thromb Vasc Biol. 2000;20(8):1860–72.
Rämet M, Pearson A, Manfruelli P, Li X, Koziel H, Göbel V, Chung E, Krieger M, Ezekowitz RA. Drosophila scavenger receptor CI is a pattern recognition receptor for bacteria. Immunity. 2001;15(6):1027–38.
Ramprasad MP, Terpstra V, Kondratenko N, Quehenberger O, Steinberg D. Cell surface expression of mouse macrosialin and human CD68 and their role as macrophage receptors for oxidized low density lipoprotein. Proc Natl Acad Sci. 1996;93(25):14833–8.
Moriwaki H, Kume N, Sawamura T, Aoyama T, Hoshikawa H, Ochi H, Nishi E, Masaki T, Kita T. Ligand specificity of LOX-1, a novel endothelial receptor for oxidized low density lipoprotein. Arterioscler Thromb Vasc Biol. 1998;18(10):1541–7.
Berwin B, Delneste Y, Lovingood RV, Post SR, Pizzo SV. SREC-I, a type F scavenger receptor, is an endocytic receptor for calreticulin. J Biol Chem. 2004;279(49):51250–7.
Shimaoka T, Kume N, Minami M, Hayashida K, Kataoka H, Kita T, Yonehara S. Molecular cloning of a novel scavenger receptor for oxidized low density lipoprotein, SR-PSOX, on macrophages. J Biol Chem. 2000;275(52):40663–6.
Prevo R, Banerji S, Ni J, Jackson DG. Rapid plasma membrane-endosomal trafficking of the lymph node sinus and high endothelial venule scavenger receptor/homing receptor stabilin-1 (FEEL-1/CLEVER-1). J Biol Chem. 2004;279(50):52580–92.
Vasquez M, Simões I, Consuegra-Fernández M, Aranda F, Lozano F, Berraondo P. Exploiting scavenger receptors in cancer immunotherapy: lessons from CD5 and SR-B1. Eur J Immunol. 2017;47(7):1108–18.
Yu X, Guo C, Fisher PB, Subjeck JR, Wang XY. Scavenger receptors: emerging roles in cancer biology and immunology. Adv Cancer Res. 2015;128:309–64.
Danilo C, Gutierrez-Pajares JL, Mainieri MA, Mercier I, Lisanti MP, Frank PG. Scavenger receptor class B type I regulates cellular cholesterol metabolism and cell signaling associated with breast cancer development. Breast Cancer Res. 2013;15(5):R87.
Twiddy AL, Cox ME, Wasan KM. Knockdown of scavenger receptor class B type I reduces prostate specific antigen secretion and viability of prostate cancer cells. Prostate. 2012;72(9):955–65.
Kamada N, Kodama T, Suzuki H. Macrophage scavenger receptor (SR-A I/II) deficiency reduced diet-induced atherosclerosis in C57BL/6J mice. J Atheroscler Thromb. 2001;8(1):1–6.
Acton S, Rigotti A, Landschulz KT, Xu S, Hobbs HH, Krieger M. Identification of scavenger receptor SR-BI as a high density lipoprotein receptor. Science. 1996;271(5248):518–20.
Husemann J, Loike JD, Anankov R, Febbraio M, Silverstein SC. Scavenger receptors in neurobiology and neuropathology: their role on microglia and other cells of the nervous system. Glia. 2002;40(2):195–205.
Terpstra V, Bird DA, Steinberg D. Evidence that the lipid moiety of oxidized low density lipoprotein plays a role in its interaction with macrophage receptors. Proc Natl Acad Sci. 1998;95(4):1806–11.
El Khoury J, Hickman SE, Thomas CA, Cao L, Silverstein SC, Loike JD. Scavenger receptor-mediated adhesion of microglia to β-amyloid fibrils. Nature. 1996;382(6593):716–9.
Dunne DW, Resnick D, Greenberg J, Krieger M, Joiner KA. The type I macrophage scavenger receptor binds to gram-positive bacteria and recognizes lipoteichoic acid. Proc Natl Acad Sci. 1994;91(5):1863–7.
Fujiwara M, Baldeschwieler JD, Grubbs RH. Receptor-mediated endocytosis of poly (acrylic acid)-conjugated liposomes by macrophages. Biochim Biophys Acta. 1996;1278(1):59–67.
Cordes T, Michelucci A, Hiller K. Itaconic acid: the surprising role of an industrial compound as a mammalian antimicrobial metabolite. Annu Rev Nutr. 2015;35:451–73.
Wang G, Simberg D. Role of scavenger receptors in immune recognition and targeting of nanoparticles. Rev Cell Biol Mol Med. 2006;1(3):166–89.
Minami M, Kume N, Shimaoka T, Kataoka H, Hayashida K, Yonehara S, Kita T. Expression of scavenger receptor for phosphatidylserine and oxidized lipoprotein (SR-PSOX) in human atheroma. Ann N Y Acad Sci. 2001;947(1):373–6.
Gu BJ, Saunders BM, Petrou S, Wiley JS. P2X7 is a scavenger receptor for apoptotic cells in the absence of its ligand, extracellular ATP. J Immunol. 2011;187(5):2365–75.
Podrez EA, Poliakov E, Shen Z, Zhang R, Deng Y, Sun M, Finton PJ, Shan L, Gugiu B, Fox PL, Hoff HF. Identification of a novel family of oxidized phospholipids that serve as ligands for the macrophage scavenger receptor CD36. J Biol Chem. 2002;277(41):38503–16.
Balasubramanian K, Maeda A, Lee JS, Mohammadyani D, Dar HH, Jiang JF, Croix CM, Watkins S, Tyurin VA, Tyurina YY, Klöditz K. Dichotomous roles for externalized cardiolipin in extracellular signaling: promotion of phagocytosis and attenuation of innate immunity. Sci Signal. 2015;8(395):ra95.
Tsubamotoa Y, Yamada N, Watanabe Y, Inaba T, Shiomi M, Shimano H, Gotoda T, Harada K, Shimada M, Ohsuga JI, Kawamura M. Dextran sulfate, a competitive inhibitor for scavenger receptor, prevents the progression of atherosclerosis in Watanabe heritable hyperlipidemic rabbits. Atherosclerosis. 1994;106(1):43–50.
Brown MS, Goldstein JL. Lipoprotein metabolism in the macrophage: implications for cholesterol deposition in atherosclerosis. Annu Rev Biochem. 1983;52(1):223–61.
Urano K, Haba M, Yuasa H, Watanabe J. Internalization of fractionated 3H-heparin by the scavenger-like receptor in rat liver parenchymal cells in primary culture. Drug Deliv. 1997;4(3):181–5.
Harris EN. Heparin clearance by liver scavenger receptors. Biochem Anal Biochem. 2012;1:e114.
Funderburgh JL, Mitschler RR, Funderburgh ML, Roth MR, Chapes SK, Conrad GW. Macrophage receptors for lumican. A corneal keratan sulfate proteoglycan. Invest Ophthalmol Vis Sci. 1997;38(6):1159–67.
Harris EN, Weigel PH. The ligand-binding profile of HARE: hyaluronan and chondroitin sulfates A, C, and D bind to overlapping sites distinct from the sites for heparin, acetylated low-density lipoprotein, dermatan sulfate, and CS-E. Glycobiology. 2008;18(8):638–48.
Sobal G, Sinzinger H. Binding of [99mTc] chondroitin sulfate to scavenger receptors on human chondrocytes as compared to binding of oxidized [125I] LDL on human macrophages. J Recept Sig Transd. 2002;22(1–4):459–70.
Fang W, Bi D, Zheng R, Cai N, Xu H, Zhou R, Lu J, Wan M, Xu X. Identification and activation of TLR4-mediated signaling pathways by alginate-derived guluronate oligosaccharide in RAW264. 7 macrophages. Sci Rep. 2017;7(1):1663.
McCourt PA, Smedsrød BH, Melkko J, Johansson S. Characterization of a hyaluronan receptor on rat sinusoidal liver endothelial cells and its functional relationship to scavenger receptors. Hepatology. 1999;30(5):1276–86.
Kelley JL, Ozment TR, Li C, Schweitzer JB, Williams DL. Scavenger receptor-A (CD204): a two-edged sword in health and disease. Crit Rev Immunol. 2014;34(3):241–61.
Harada M, Imai J, Okuno S, Suzuki T. Macrophage-mediated activation of camptothecin analogue T-2513–carboxymethyl dextran conjugate (T-0128): possible cellular mechanism for antitumor activity. J Control Release. 2000;69(3):389–97.
Tokuda H, Masuda S, Takakura Y, Sezaki H, Hashida M. Specific uptake of succinylated proteins via a scavenger receptor-mediated mechanism in cultured brain microvessel endothelial cells. Biochem Biophys Res Commun. 1993;196(1):18–24.
Hoang B, Ernsting MJ, Roy A, Murakami M, Undzys E, Li SD. Docetaxel-carboxymethylcellulose nanoparticles target cells via a SPARC and albumin dependent mechanism. Biomaterials. 2015;59:66–76.
Martino JV, Van Limbergen J, Cahill LE. The role of carrageenan and carboxymethylcellulose in the development of intestinal inflammation. Front Pediatr. 2017;5:96.
Iesaki T, Takeuchi T, Okano M, Hashimoto R, Kakigi R, Ishii Y, Okada T. Fucoidan, a ligand of scavenger receptor class a, causes vascular relaxation through a nitric oxide/cGMP-mediated pathway in rat aorta. Atherosclerosis. 2014;235(2):e36.
Thelen T, Hao Y, Medeiros AI, Curtis JL, Serezani CH, Kobzik L, Harris LH, Aronoff DM. The class A scavenger receptor, macrophage receptor with collagenous structure, is the major phagocytic receptor for Clostridium sordellii expressed by human decidual macrophages. J Immunol. 2010;185(7):4328–35.
Rost MS, Sumanas S. Hyaluronic acid receptor stabilin-2 regulates Erk phosphorylation and arterial-venous differentiation in zebrafish. PLoS One. 2014;9(2):e88614.
Marshall-Clarke S, Downes JE, Haga IR, Bowie AG, Borrow P, Pennock JL, Grencis RK, Rothwell P. Polyinosinic acid is a ligand for toll-like receptor 3. J Biol Chem. 2007;282(34):24759–66.
Pearson AM, Rich A, Krieger M. Polynucleotide binding to macrophage scavenger receptors depends on the formation of base-quartet-stabilized four-stranded helices. J Biol Chem. 1993;268(5):3546–54.
Zeng J, Zhang Y, Hao J, Sun Y, Liu S, Bernlohr DA, Sauter ER, Cleary MP, Suttles J, Li B. Stearic acid induces CD11c expression in proinflammatory macrophages via epidermal fatty acid binding protein. J Immunol. 2018;200(10):3407–19.
Loison C, Mendy F, Sérougne C, Lutton C. Dietary myristic acid modifies the HDL-cholesterol concentration and liver scavenger receptor BI expression in the hamster. Br J Nutr. 2002;87(3):199–210.
Chao Y, Karmali PP, Mukthavaram R, Kesari S, Kouznetsova VL, Tsigelny IF, Simberg D. Direct recognition of superparamagnetic nanocrystals by macrophage scavenger receptor SR-AI. ACS Nano. 2013;7(5):4289–98.
Suzuki Y, Tada-Oikawa S, Ichihara G, Yabata M, Izuoka K, Suzuki M, Sakai K, Ichihara S. Zinc oxide nanoparticles induce migration and adhesion of monocytes to endothelial cells and accelerate foam cell formation. Toxicol Appl Pharmacol. 2014;278(1):16–25.
Iyer R, Hamilton RF, Li L, Holian A. Silica-induced apoptosis mediated via scavenger receptor in human alveolar macrophages. Toxicol Appl Pharmacol. 1996;141(1):84–92.
Murthy S, Larson-Casey JL, Ryan AJ, He C, Kobzik L, Carter AB. Alternative activation of macrophages and pulmonary fibrosis are modulated by scavenger receptor, macrophage receptor with collagenous structure. FASEB J. 2015;29(8):3527–36.
Resnick D, Freedman NJ, Xu SH, Krieger M. Secreted extracellular domains of macrophage scavenger receptors form elongated trimers which specifically bind crocidolite asbestos. J Biol Chem. 1993;268(5):3538–45.
BEPPU M, HORA M, KIKUGAWA K. A simple method for the assessment of macrophage scavenger receptor-ligand interaction: adherence of erythrocytes coated with oxidized low density lipoprotein and modified albumin to macrophages. Biol Pharm Bull. 1994;17(1):39–46.
Takata K, Horiuchi S, Morino Y. Scavenger receptor-mediated recognition of maleylated albumin and its relation to subsequent endocytic degradation. Biochim Biophys Acta. 1989;984(3):273–80.
Greaves DR, Gordon S. The macrophage scavenger receptor at 30 years of age: current knowledge and future challenges. J Lipid Res. 2009;50(Supplement):S282–6.
Herrmann M, Schäfer C, Heiss A, Gräber S, Kinkeldey A, Büscher A, Schmitt MM, Bornemann J, Nimmerjahn F, Herrmann M, Helming L. Clearance of fetuin-A–containing calciprotein particles is mediated by scavenger receptor -A. Circ Res. 2012;111(5):575–84.
Melkko J, Hellevik T, Risteli L, Risteli J, Smedsrød B. Clearance of NH2-terminal propeptides of types I and III procollagen is a physiological function of the scavenger receptor in liver endothelial cells. J Exp Med. 1994;179(2):405–12.
Facciponte JG, Wang XY, Subjeck JR. Hsp110 and Grp170, members of the Hsp70 superfamily, bind to scavenger receptor-A and scavenger receptor expressed by endothelial cells -I. Eur J Immunol. 2007;37(8):2268–79.
Ben J, Zhang Y, Zhou R, Zhang H, Zhu X, Li X, Zhang H, Li N, Zhou X, Bai H, Yang Q. Major vault protein regulates class A scavenger receptor-mediated tumor necrosis factor-α synthesis and apoptosis in macrophages. J Biol Chem. 2013;288(27):20076–84.
Takahashi T, Suzuki T. Role of sulfatide in normal and pathological cells and tissues. J Lipid Res. 2012;53(8):1437–50.
Shannahan JH, Bai W, Brown JM. Implications of scavenger receptors in the safe development of nanotherapeutics. Recept Clin Invest. 2015;2(3):e811.
Basu SK, Majumdar S, Mukhopadhyay B, Mukhopadhyay A. Receptor-mediated drug delivery to macrophages. Proc Indian Natl Sci Acad Part B. 1994;60:345–56.
Scherphof GL, Kamps JA. Receptor versus non-receptor mediated clearance of liposomes. Adv Drug Deliv Rev. 1998;32(1–2):81–97.
Semple SC, Chonn A, Cullis PR. Interactions of liposomes and lipid-based carrier systems with blood proteins: relation to clearance behaviour in vivo. Adv Drug Deliv Rev. 1998;32(1–2):3–17.
Liu D, Liu F, Song YK. Recognition and clearance of liposomes containing phosphatidylserine are mediated by serum opsonin. Biochim Biophys Acta. 1995;1235(1):140–6.
Devine DV, Wong K, Serrano K, Chonn A, Cullis PR. Liposome complement interactions in rat serum: implications for liposome survival studies. Biochim Biophys Acta. 1994;1191(1):43–51.
Fidler IJ, Raz A, Fogler WE, Kirsh R, Bugelski P, Poste G. Design of liposomes to improve delivery of macrophage-augmenting agents to alveolar macrophages. Cancer Res. 1980;40(12):4460–6.
Chonn A, Semple SC, Cullis PR. Association of blood proteins with large unilamellar liposomes in vivo. Relation to circulation lifetimes. J Biol Chem. 1992;267(26):18759–65.
Vyas SP, Kannan ME, Jain S, Mishra V, Singh P. Design of liposomal aerosols for improved delivery of rifampicin to alveolar macrophages. Int J Pharm. 2004;269(1):37–49.
Kamps JA, Morselt HW, Swart PJ, Meijer DK, Scherphof GL. Massive targeting of liposomes, surface-modified with anionized albumins, to hepatic endothelial cells. Proc Natl Acad Sci. 1997;94(21):11681–5.
Etzerodt A, Maniecki MB, Graversen JH, Møller HJ, Torchilin VP, Moestrup SK. Efficient intracellular drug-targeting of macrophages using stealth liposomes directed to the hemoglobin scavenger receptor CD163. J Control Release. 2012;160(1):72–80.
Li X, Kan HY, Lavrentiadou S, Krieger M, Zannis V. Reconstituted discoidal ApoE-phospholipid particles are ligands for the scavenger receptor BI The amino-terminal 1–165 domain of ApoE suffices for receptor binding. J Biol Chem. 2002;277(24):21149–57.
Sakai-Kato K, Sakurai M, Takechi-Haraya Y, Nanjo K, Goda Y. Involvement of scavenger receptor class B type 1 and low-density lipoprotein receptor in the internalization of liposomes into HepG2 cells. Biochim Biophys Acta. 2017;1859(11):2253–8.
Lee KD, Pitas RE, Papahadjopoulos D. Evidence that the scavenger receptor is not involved in the uptake of negatively charged liposomes by cells. Biochim Biophys Acta. 1992;1111(1):1–6.
Pinheiro M, Lúcio M, Lima JL, Reis S. Liposomes as drug delivery systems for the treatment of TB. Nanomedicine. 2011;6(8):1413–28.
Labana S, Pandey R, Sharma S, Khuller GK. Chemotherapeutic activity against murine tuberculosis of once weekly administered drugs (isoniazid and rifampicin) encapsulated in liposomes. Int J Antimicrob Agents. 2002;20(4):301–4.
Pollock S, Nichita NB, Böhmer A, Radulescu C, Dwek RA, Zitzmann N. Polyunsaturated liposomes are antiviral against hepatitis B and C viruses and HIV by decreasing cholesterol levels in infected cells. Proc Natl Acad Sci. 2010;107(40):17176–81.
Tempone AG, Perez D, Rath S, Vilarinho AL, Mortara RA, de Andrade Jr HF. Targeting Leishmania (L.) chagasi amastigotes through macrophage scavenger receptors: the use of drugs entrapped in liposomes containing phosphatidylserine. J Antimicrob Chemother. 2004;54(1):60–8.
da Costa-Silva TA, Galisteo AJ, Lindoso JA, Barbosa LR, Tempone AG. Nanoliposomal buparvaquone immunomodulates Leishmania infantum-infected macrophages and is highly effective in a murine model. Antimicrob Agents Chemother. 2017;61(4):e02297–16.
Pollock S, Antrobus R, Newton L, Kampa B, Rossa J, Latham S, Nichita NB, Dwek RA, Zitzmann N. Uptake and trafficking of liposomes to the endoplasmic reticulum. FASEB J. 2010;24(6):1866–78.
Di Y, Wasan EK, Cawthray J, Wasan KM. Scavenger receptor class BI (SR-BI) mediates uptake of CPX-351 into K562 leukemia cells. Drug Dev Ind Pharm. 2019;45(1):21–6.
Pont I, Calatayud-Pascual A, López-Castellano A, Albelda EP, García-España E, Martí-Bonmatí L, Frias JC, Albelda MT. Anti-angiogenic drug loaded liposomes: Nanotherapy for early atherosclerotic lesions in mice. PLoS One. 2018;13(1):e0190540.
Rensen PC, Gras JC, Lindfors EK, van Dijk KW, Jukema JW, van Berkel TJ, Biessen EA. Selective targeting of liposomes to macrophages using a ligand with high affinity for the macrophage scavenger receptor class A. Curr Drug Discov Technol. 2006;3(2):135–44.
Gan C, Wang K, Tang Q, Chen Y. Comparative investigation on the sizes and scavenger receptor binding of human native and modified lipoprotein particles with atomic force microscopy. J Nanobiotechnol. 2018;16(1):25.
Zhang X, Huang G. Synthetic lipoprotein as nano-material vehicle in the targeted drug delivery. Drug Deliv. 2017;24(2):16–21.
Shaw JM, Futch WS, Schook LB. Induction of macrophage antitumor activity by acetylated low density lipoprotein containing lipophilic muramyl tripeptide. Proc Natl Acad Sci. 1988;85(16):6112–6.
Kreuter J, Hekmatara T, Dreis S, Vogel T, Gelperina S, Langer K. Covalent attachment of apolipoprotein AI and apolipoprotein B-100 to albumin nanoparticles enables drug transport into the brain. J Control Release. 2007;118(1):54–8.
Singh RP, Ramarao P. Cellular uptake, intracellular trafficking and cytotoxicity of silver nanoparticles. Toxicol Lett. 2012;213(2):249–59.
Shannahan JH, Podila R, Aldossari AA, Emerson H, Powell BA, Ke PC, Rao AM, Brown JM. Formation of a protein corona on silver nanoparticles mediates cellular toxicity via scavenger receptors. Toxicol Sci. 2014;143(1):136–46.
Aldossari AA, Shannahan JH, Podila R, Brown JM. Scavenger receptor B1 facilitates macrophage uptake of silver nanoparticles and cellular activation. J Nanopart Res. 2015;17(7):313.
Lipinski MJ, Frias JC, Amirbekian V, Briley-Saebo KC, Mani V, Samber D, Abbate A, Aguinaldo JG, Massey D, Fuster V, Vetrovec GW. Macrophage-specific lipid-based nanoparticles improve mri detection and characterization of human atherosclerosis. J Am Coll Cardiol Img. 2009;2(5):637–47.
Hirano S, Kanno S, Furuyama A. Multi-walled carbon nanotubes injure the plasma membrane of macrophages. Toxicol Appl Pharmacol. 2008;232(2):244–51.
Kanno S, Furuyama A, Hirano S. A murine scavenger receptor MARCO recognizes polystyrene nanoparticles. Toxicol Sci. 2007;97(2):398–406.
Orr GA, Chrisler WB, Cassens KJ, Tan R, Tarasevich BJ, Markillie LM, Zangar RC, Thrall BD. Cellular recognition and trafficking of amorphous silica nanoparticles by macrophage scavenger receptor A. Nanotoxicology. 2011;5(3):296–311.
Hamilton R, Buckingham S, Holian A. The effect of size on Ag nanosphere toxicity in macrophage cell models and lung epithelial cell lines is dependent on particle dissolution. Int J Mol Sci. 2014;15(4):6815–30.
Hsu CS, Hsu SJ, Liu WL, Chen DS, Kao JH. Association of SCARB1 Gene polymorphisms with virological response in chronic hepatitis C patients receiving pegylated interferon plus ribavirin therapy. Sci Rep. 2016;6:32303.
SR-BI and Antiviral Treatment Response in HCV [Internet]. Available from: https://clinicaltrials.gov/show/NCT02714712. Accessed on 14th Jun 2019.
Genetic and Metabolism of Post-prandial HDL Particles (HDL-PP) [Internet]. Available from: https://clinicaltrials.gov/show/NCT03109067. Accessed on 14th Jun 2019.
Mechanisms of Chronic Kidney Disease (CKD)-Induced Foam Cell Formation [Internet]. Available from: https://clinicaltrials.gov/show/NCT01671605. Accessed on 14th Jun 2019.
A Study of Immunological Biomarkers as Predictors of Cardiovascular Events (BIOKID) [Internet]. Available from: https://clinicaltrials.gov/show/NCT02894060. Accessed on 14th Jun 2019.
Zheng Y, Liu Y, Jin H, Pan S, Qian Y, Huang C, Zeng Y, Luo Q, Zeng M, Zhang Z. Scavenger receptor B1 is a potential biomarker of human nasopharyngeal carcinoma and its growth is inhibited by HDL-mimetic nanoparticles. Theranostics. 2013;3(7):477–86.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 American Association of Pharmaceutical Scientists
About this chapter
Cite this chapter
Lokhande, A.S., Jahagirdar, P., Dandekar, P., Devarajan, P.V. (2019). Scavenger Receptor and Targeting Strategies. In: Devarajan, P., Dandekar, P., D'Souza, A. (eds) Targeted Intracellular Drug Delivery by Receptor Mediated Endocytosis. AAPS Advances in the Pharmaceutical Sciences Series, vol 39. Springer, Cham. https://doi.org/10.1007/978-3-030-29168-6_10
Download citation
DOI: https://doi.org/10.1007/978-3-030-29168-6_10
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-29167-9
Online ISBN: 978-3-030-29168-6
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)