Abstract
Epithelial cells and functions of the epithelium are critical to the health of the oral cavity. We used a nonhuman primate model to profile the transcriptome of gingival tissues in health across the lifespan and hypothesized that in older animals, epithelial-related transcriptome patterns would reflect epithelial cells that are aggressively responsive to the surrounding environment and less able to modulate and resolve the noxious challenge from the bacteria. Rhesus monkeys (n = 34) with a healthy periodontium were distributed into four groups: ≤3 years (young), 3–7 years (adolescent), 12–16 years (adult), and 18–23 years (aged), and a buccal gingival sample from the premolar/molar region of each animal was obtained. RNA was subjected to a microarray analysis (GeneChip® Rhesus Macaque Genome Array, Affymetrix), and 336 genes examined that are linked to epithelium and epithelial cell functions categorized into 9 broad functional groups: extracellular matrix and cell structure; extracellular matrix remodeling enzymes; cell adhesion molecules, cytoskeleton regulation; inflammatory response; growth factors; kinases/cell signaling; cell surface receptors; junction associated molecules; autophagy/apoptosis; antimicrobial peptides; and transcription factors. Total of 255 genes displayed a normalized signal >100, and differences across the age groups were observed primarily in extracellular matrix and cell structure, cell adhesion molecules, and cell surface receptor gene categories with elevations in the aged tissues. Keratins 2, 5, 6B, 13, 16, 17 were all significantly increased in healthy-aged tissues versus adults, and keratins 1 and 2 were significantly decreased in young animals. Approximately 15 integrins are highly expressed in the gingival tissues across the age groups with only ITGA8, ITGAM (CD11b), and ITGB2 significantly increased in the aged tissues. Little impact of aging on desmosomal/hemidesmosomal genes was noted. These results suggest that healthy gingival aging has a relatively limited impact on the broader functions of the epithelium and epithelial cells, with some effects on genes for extracellular matrix and cell adhesion molecules (e.g., integrins). Thus, while there is a substantial impact of aging on immune system targets even in healthy gingiva, it appears that the epithelial barrier remains reasonably molecularly intact in this model system.
Access provided by Autonomous University of Puebla. Download conference paper PDF
Similar content being viewed by others
Introduction
The majority of agents that cause infections in humans gain access through the mucosal surfaces of the body. As such, the epithelium and epithelial cells have evolved to provide an array of features to protect from pathogenic challenge. These include barrier functions in which the epithelial cells rapidly mature and are sloughed from the surface while maintaining tight junctions enhancing exclusion of deleterious agents at luminal surfaces (Parrish 2017; Yu et al. 2012). In addition to these mechanical barriers, recent evidence has supported the capacity of epithelial cells to constitutively synthesize an array of innate immune protective molecules, as well as a range of cell communication factors providing an “early warning system” to the host inflammatory and immune armamentarium (Pardo-Camacho et al. 2018; Partida-Rodriguez et al. 2017; Ahluwalia et al. 2017). Moreover, a number of these protective signaling molecules are induced through engagement of microbial-associated molecular patterns (e.g., MAMPs, PAMPs) and danger-associated molecular patterns (DAMPs) (Rajaee et al. 2018; Patel 2018; De Lorenzo et al. 2018; Stocks et al. 2018; Walsh et al. 2013; Olive 2012). The functions of the epithelial cells continue to emerge as critical determinants of maintaining host integrity from challenge with pathogenic bacteria, viruses, and fungi, which includes receptor recognition and engagement resulting in specific intracellular signaling pathways leading to antimicrobial activities in the local mucosal environment (Jin and Weinberg 2018; Guncu et al. 2015; Sukhithasri et al. 2013; Ho et al. 2013; McCormick and Weinberg 2010).
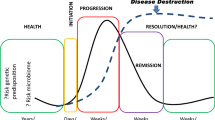
The oral cavity is somewhat unique in the properties of its epithelium. While other mucosal sites in the body consider it a substantial benefit to maintain the integrity of the barrier function, in the oral cavity, the epithelium is routinely deliberately breached from about 6 months to 21 years of age with eruption of the deciduous and permanent dentition. This developmental anomaly of innate immune protection has fostered the development of a unique junctional epithelium that covers connective tissue cells and a collagen matrix that attaches the erupted teeth to the underlying alveolar bone. This junctional epithelial lining of the subgingival sulcus, in health, is attached to the cementoenamel junction of the teeth (Tsukamoto et al. 2012; Hatakeyama et al. 2006; Bosshardt and Lang 2005). Interestingly, in health, this junctional epithelium is somewhat leaky and allows that passage of a low protein fluid transudate in the gingival crevice that mechanically aids in rinsing colonizing bacteria into the saliva, which is swallowed approximating 1 L/day. Accompanying accumulation of bacterial deposits supra- and subgingivally, the gingival tissue reacts with an inflammatory response with the classic signs of acute inflammation. This inflammation, termed gingivitis, is considered a reversible process that responds rapidly to removal of the bacterial insult (Tonetti et al. 2015; Chapple et al. 2015). An inability to clear this stimulus can lead to a persistent immunoinflammatory lesion, i.e., periodontitis, with ulceration of the epithelium, influx of an array of inflammatory cells, breakdown of connective tissue and collagen, vasculitis, and net resorption of alveolar bone at the localized site of the microbial challenge (Tonetti et al. 2015). While substantial strides are being made in the area of tissue regeneration to reestablish normal function for the periodontium following disease, periodontitis remains considered as irreversible once tissue destruction has occurred.
Age-dependent variations in epithelial barrier function have been previously described in different tissues (e.g., skin, lung, intestine, and kidney) of humans and animal models. A common finding is an impaired cell–cell adhesion mediated by tight junctions consistent with aging-increased permeability (Parrish 2017). Additionally, this decline in epithelial barrier function and repair seems to be associated with an alteration in epithelium stem cells niches (Doles et al. 2012; Moorefield et al. 2017). Nevertheless, the molecular mechanisms associated to these observations remain unclear. Thus, as the epithelial cells and functions of the epithelium are critical to the health of the oral cavity, we used a nonhuman primate model to profile the transcriptome of gingival tissues in health across the lifespan. It was hypothesized that in younger animals, epithelial genes related to functions of a more rigid, less developmentally flexible tissue would be decreased, enabling these young animals to respond to the microbial burden by enhanced signaling pathways associated with rapid wound healing, anti-inflammatory/inflammation resolution, maintaining an effective barrier. In contrast, in older animals, these patterns would differ creating epithelial cells highly responsive to the surrounding environment and less able to modulate and resolve the noxious challenge from the bacteria in the absence of some collateral damage of the periodontal tissues and enhancing the long-term risk for initiation and progression of periodontitis.
Methods
Nonhuman Primate Model and Oral Clinical Evaluation
Rhesus monkeys (Macaca mulatta) (n = 23; 10 females and 13 males) housed at the Caribbean Primate Research Center (CPRC) at Sabana Seca, Puerto Rico, were used in these studies. Healthy animals (5–7/group) were distributed by age into four groups: ≤3 years (young), 3–7 years (adolescent), 12–16 years (adult), and 18–23 years (aged). The nonhuman primates are typically fed a 20% protein, 5% fat, and 10% fiber commercial monkey diet (diet 8773, Teklad NIB primate diet modified: Harlan Teklad). The diet is supplemented with fruits and vegetables, and water is provided ad libitum in an enclosed corral setting.
A protocol approved by the Institutional Animal Care and Use Committee (IACUC) of the University of Puerto Rico enabled anesthetized animals to be examined for clinical measures of periodontal including probing pocket depth (PD) and bleeding on probing (BOP) as we have described previously (Ebersole et al. 2008).
Tissue Sampling and Gene Expression Microarray Analysis
A buccal gingival sample from healthy tissues from the premolar/molar maxillary region of each animal was taken using a standard gingivectomy technique and maintained frozen in RNAlater solution. Total RNA was isolated from each gingival tissue using a standard procedure as we have described, and tissue RNA samples submitted to the microarray core to assess RNA quality analyze the transcriptome using the GeneChip® Rhesus Macaque Genome Array (Affymetrix) (Meka et al. 2010; Gonzalez et al. 2011). Individual samples were used for gene expression analyses.
Data Analysis
Normalization of values across the chips was accomplished through signal intensity standardization across each chip using Affymetrix PLIER algorithm. The GeneChip® Rhesus Macaque Genome Array contained matched and mismatched pairs allowing the MAS 5 algorithm to be used. For each gene, we first determined differences in expression across the groups using ANOVA (version 9.3, SAS Inc., Cary, NC). The healthy-aged tissues were then compared among the age groups using a t-test and accepting a p-value ≤0.05 for significance. Because of the cost of these types of nonhuman primate experiments and availability of primates of the various ages, we did not have sufficient samples to identify if the relationship between age and gene expression could be treated using a linear model; thus, the subjects were classified and ANOVA was used for analysis. Correlations with aging and clinical parameters in healthy tissues were determined using a Spearman Rank correlation analysis. A p-value ≤0.05 was used to evaluate the significance of the correlation. The data have been uploaded to http://www.ncbi.nlm.nih.gov/geo/info/submission.html.
Results
Epithelium Gene Transcriptome in Healthy Gingival Tissues
Using the microarray results, we examined 336 genes that are linked to epithelium and epithelial cells functions (Table 1). The set of genes were categorized into 9 broad functional groups: extracellular matrix and cell structure; extracellular matrix remodeling enzymes; cell adhesion molecules, cytoskeleton regulation; inflammatory response; growth factors; kinases/cell signaling; cell surface receptors; junction associated molecules; autophagy/apoptosis; antimicrobial peptides; and transcription factors.
Figure 1a–d summarizes the level of expression of genes in which the normalized signal was >100 in gingival tissues from any of the 4 groups of animals that included 255 genes. From these data, we identified a group of genes that were altered in younger and aged animals when compared to expression levels in the adult tissues, which included selected extracellular matrix components (e.g., KRT2, KRT4, MMP1, MMP9, TIMP1, F13A1, SERPINF1, CTSK, FBN1, LAD1, CHI3L1), cytoskeleton regulators (e.g., ACTN1, TAGLN, ZYX, DES), cell surface receptors and adhesion molecules (e.g., SELL, ICAM2, ITGAL, SPP1, ITGB2, ITGA8, SELP, ITGAM, ICAM1, ITGAX), and host response genes (e.g., PPBP, CAMP, DEFB4, CXCL11).
(a–d): Gene expression levels in gingival tissues reflecting epithelium/epithelial cell functions. The lines represent the mean normalized signal level for each age group on animals. The genes are stratified into general functional categories and grouped in the graphs based upon the magnitude of signal (1: extracellular matrix components; 2: extracellular matrix enzymes; 3: cell adhesion molecules; 4: cytoskeleton regulators; 5: inflammatory cytokines/chemokines; 6: growth factors; 7: kinases/cell signaling; 8: cell surface receptors; 9: junction associated proteins; 10: autophagy/apoptosis; 11: antimicrobial molecules; 12: transcription factors)
Aging Effects on Epithelium Gene Transcriptome
Within the subset of 255 genes, Fig. 2a–c provides volcano plot visualization of the distribution of altered responses and significant differences in the young, adolescent, and aged animals versus healthy adult levels that were considered to be the normal expression level. From these analyses, it appeared that a lower number of genes were significantly different in the young animals versus the other groups, where 10–20% of the genes varied from healthy adult tissues.
Interrogating this dataset more specifically, Fig. 3 provides a heatmap representation of the fold increase or decrease in gene expression in healthy young, adolescent, and aged tissues compared to adults. Additionally, the genes were classified into 9 categories across their range of functions for the epithelium and epithelial cells. The results showed that the extracellular matrix and cell structure, cell adhesion molecules, and cell surface receptor categories appeared to be most frequently different. Several extracellular matrix structural and cell adhesion molecules were elevated in the aged tissues, with generally decreased levels in the young tissue samples. The cell surface receptors were, generally, increased across the different age groups versus the adult levels. Also of interest was the lack of effect on the array of molecules related to epithelium junctions, transcription factors, kinases, and genes linked to autophagy/apoptosis. Of note, specific genes such as SPP1 and PPBP showed and aging-related increase. Table 2 provides a pathway analysis to assess biologic processes that were enriched in the set of genes that were significantly and/or >1.25-fold-regulated. As was seen in the heatmap, cell–matrix, cell–cell adhesion, and differentiation were enriched. While the heatmap did not provide a clear visualization of alterations in MAPK signaling pathway genes, these were enriched in the pathway analysis evaluation.
Figure 4a and b focuses on the details of altered expression of the array of keratins that are critical for epithelial cell functions. The results showed that approximately 20 of the keratins were expressed at high levels in the gingival tissues. Keratins 2, 5, 6B, 13, 16, 17 were all significantly increased in healthy-aged tissues versus adults. In contrast, keratins 1 and 2 were significantly decreased and keratin 17 increased in tissue from young animals compared to healthy adults. An additional set of molecules critical for communication of the epithelial cells are the array of integrin surface receptors. Figure 5 provides an overview of these response profiles across the age groups. Approximately 15 of these integrins are highly expressed in the gingival tissues across the age groups. Only ITGA8, ITGAM (CD11b), and ITGB2 were significantly increased in the aged tissues compared to adults, with no difference in the younger animals. ITGB2 is a component portion of integrins that bind ICAMs, VCAM, and even complement components. ITGAM/ITGB2 is particularly implicated in interactions of monocytes, macrophages, and granulocytes and the uptake of complement-coated particles. Thus, while these integrins can be related to epithelial cell biology, their role in these complex oral tissues may be more related to the physiologic inflammation of the gingiva and reflect tissue maintenance by inflammatory cell responses in these tissues. Lastly, we focused on the array of biomolecules related to epithelial junctions including desmosomal and hemidesmosomal proteins (Fig. 6a, b). As was noted from the heatmap, few of these proteins were significantly altered across the age groups, with only CDSN (corneodesmosin) being increased in younger animals versus adults, and COL7A1 (collagen) and LAMA5 (laminin) decreased in the aged animal tissues.
The data were also analyzed beyond an age categorization (young, adolescent, adult, aged) by evaluating correlations of the gene expression profiles with age as a continuous variable (Fig. 7a). The results demonstrated about 10% of the genes demonstrated significant correlations (p < 0.01) with similar numbers positively and negatively correlated. While those positively correlated genes represented a range of functions, of interest was the number of collagen and integrin genes that were significantly decreased with aging even in healthy tissues. Figure 7b, c provides a similar type of assessment, relating gene profiles to clinical features of the periodontium in the healthy animals (bleeding on probing—BOP; mean probing pocket depth—PPD). In contrast to the correlations with age, fewer relationships were observed with either of the clinical parameters, with only PLAU, SMURF1, and MAP 3 K5 genes positively correlated and KRT17 and BMP2 negatively correlated with both BOP and PPD.
Discussion
Within the paradigms of gingivitis and periodontitis that affect the global population, there remain some observations that have yet to be understood at the molecular level. First, while gingivitis is generally considered to presage to periodontal lesions, identified populations have long-standing, florid gingival inflammation and never progress to periodontitis (Loe et al. 1986; Lang et al. 2009). Second, many cases of localized aggressive periodontitis that tend to occur in younger individuals associated with infection with Aggregatibacter actinomycetemcomitans demonstrate substantial rapid localized bone loss in the absence of gross inflammatory changes in the gingival tissues (Kinane and Hodge 2001; Jenkins and Papapanou 2001; Bimstein et al. 2002). Third, in children and adolescents, there is a high incidence of gingivitis that increases in prevalence and severity through puberty, in the absence of progressing to periodontitis (Albandar and Tinoco 2002; Modeer and Wondimu 2000; Bimstein et al. 2013; Bimstein and Ebersole 1989). Fourth, during pregnancy, subsets of women can develop rather severe pregnancy-associated gingivitis that has been suggested to be linked to hormonal changes that could influence the oral microbial ecology, although there remains sparse data documenting the molecular features of this unique gingivitis that does not progress to periodontitis (Gumus et al. 2016; Gursoy et al. 2014; Barak et al. 2003). Finally, periodontitis has long been described as a disease of aging with substantial increases in incidence and severity in aging populations, and thought to be related to a lifetime accumulation of noxious challenge to the gingival tissues (Papapanou and Susin 2017; Wu et al. 2016; Lamster et al. 2016; Hajishengallis 2014; Huttner et al. 2009). Thus, there remains a need to better understand the underlying molecular biology of the range of cells in gingival tissues and how their functions can dictate variation in disease expression across the lifespan.
This study used a nonhuman primate model to focus in the biology of the epithelium and epithelial cells in gingival tissues to test a hypothesis that alterations in the transcriptome representing a range of functions of these cells/tissues would be altered with aging even in clinically healthy sites. We had previously reported on rather dramatic changes in various immune and inflammatory cells in gingival tissues in this model. There were clear alterations even in healthy-aged tissues with regard to lymphocyte classes (Ebersole et al. 2014, 2016a), apoptosis (Gonzalez et al. 2011, 2013), macrophage function and antigen recognition and presentation (Gonzalez et al. 2014, 2015, 2018), hypoxia (Ebersole et al. 2018), and inflammasome characteristics (Ebersole et al. 2016b). However, while there were some differences in the epithelial-related gene expression profiles in periodontal health with aging, the number of genes affected with a fold-change >1.25 was only about 30% and only 8% >1.5-fold. These alterations were also focused on a more limited functional activity of the epithelium/epithelial cells with extracellular matrix structural components, cell adhesion molecules, and cell surface receptors appearing to be most greatly affected.
Drilling down into these categories, multiple collagen and keratin gene levels were lower in young versus aged tissues, which were confirmed with correlation analysis related to aging. These findings suggested that these altered structural components in healthy aging could either reflect a physiological adaptation with aging that helps to maintain healthy tissues, or potentially these changes reflect altered epithelium characteristics that could increase the risk for initiation of periodontitis. Clear histopathological results demonstrate a breakdown in epithelium integrity accompanying the chronic inflammation of periodontitis (Bosshardt and Lang 2005; Dale 2002; Van der Velden 1984). It is accepted that these microulcerations enhance access of the microbiome components (e.g., bacteria, bacterial structures) into deeper tissues contributing to activating the local inflammatory response responsible for tissue destruction. Additionally, this process is considered as part of the feature allowing bacteria to traverse the gingival tissues and enter into the systemic circulation (Cardoso et al. 2018; Abbayya et al. 2015; Maddi and Scannapieco 2013; Kumar 2013). However, examination of the genes related to cell–cell interactions and cell–matrix interactions (desmosomes, hemidesmosomes) did not show a substantial impact of aging on the expression of these molecules. Thus, how these patterns reflect aging processes in health and risk for disease remains ill-defined, and further studies will be required to discriminate between these options.
An array of genes for cell adhesion molecules including cadherins, integrins, caveolins, and selectins were increased with aging. These molecules are critical for maintaining homeostasis of the epithelium in the septic environment of the oral cavity. Thus, since the tissue samples were from clinically healthy sites in the aged animals, this type of response profile may signify an effective healthy aging process in the tissues from these animals. Of note, remarkably elevated gingival expression levels of SPP (osteopontin) and PPBP (pro-platelet basic protein:CXCL7:NAP-2) were observed with aging. SPP has been shown to play important roles in wound healing seemingly through inhibiting apoptosis and modulating the expression of MMPs (Icer and Gezmen-Karadag 2018). From an epithelial cell function viewpoint, PPBP as a heterodimer with other chemokines is involved in glycosaminoglycan interactions with cells via the CXCR2 receptor (Brown et al. 2017). It is a chemoattractant for neutrophils and has some antimicrobials activities. This chemokine has been associated with the pathogenesis of chronic diseases, such as cancer and arthritis (Yeo et al. 2016; Desurmont et al. 2015). It is also identified as one of a group of platelet-associated chemokines that were systemically elevated in patients with antiphospholipid syndrome (Patsouras et al. 2015), which has also been linked to the microbiome in periodontitis (Schenkein et al. 2003). Finally, a recent study by Shusterman et al. (2017) combining data from murine studies and an existing human dataset identified a gene cluster of platelet factor 4 (PF4:CXCL4)/PPBP/CXCL5 (neutrophil activating peptide 78: ENA-78) being significantly associated with aggressive periodontitis. These variations are consistent with previous reports demonstrating the persistence of inflammatory cells in diseased gingiva that may results from decreased apoptotic responses and/or enhanced transmigration of neutrophils into the inflammatory lesion with aging (Gonzalez et al. 2013; Wael Youssef 2018; Xia et al. 2017; Zhang et al. 2016; Jang et al. 2015; Sakai et al. 1999). Since this study showed elevations in “clinically healthy” aging tissues, there is a potential that this profile describes an enhanced risk of exhibiting disease initiation in the aged individuals.
While considerable effort has been delivered in attempting to delineate the microbiome and host response parameters that drive the disease process, there remains much less information defining, at the molecular level, what tissue responses are required to help maintain health. Recently, understanding the characteristics of the bacteria that constitute a healthy microbiome and the metabolic functions for these commensal bacteria has come under increasing scrutiny as both an explanatory variable in determining the population variation in disease and as a potential therapeutic target for more biologically oriented treatment strategies (Nassar et al. 2017; Ebersole et al. 2017; Hajishengallis and Lamont 2016; Lamont and Hajishengallis 2015; Wade 2013). However, much less is known regarding the host features controlling the periodontal microbiome in health. As an example, there is limited literature that the expression of various epithelial genes/proteins can be regulated by microbial biofilms and that members of the “red complex” can alter components of the epithelial junctions, particularly desmosomal components (Belibasakis et al. 2015). However, if age-associated alterations in these epithelial functions can affect the characteristics of the subgingival microbiome in moving from health to disease related remains unknown.
As noted, our previous examination of the gingival transcriptome in healthy nonhuman primates with aging, as well as with naturally occurring periodontitis demonstrated significant differences in gene profiles that supported innate and adaptive immune responses, inflammation, and cellular senescence changes occur in aging gingival tissues even when clinically healthy. These findings suggested that a basis for increased periodontitis in the human population with age may be linked to inherent changes in the biology of the gingival tissues during aging decreasing the capacity of the tissues to respond to local environmental changes, including alterations in the pathogenic capacity of the microbiome (Belibasakis 2018). The findings from this study suggested some changes in the functional activities of the epithelium and epithelial cells with aging; however, these differences were considerably less that noted with aging effects on immune system components. These more marginal changes in healthy aging will need to be evaluated in the context of the changes taking place in naturally occurring periodontitis, as well as the dynamics of epithelial responses in the gingival during ligature-induced periodontitis using this human-like disease model. Therefore, a more clear understanding of the fundamental biologic responses of the epithelium should provide insight into disease variation related to increased susceptibility or resistance to periodontitis across the population.
References
Abbayya, K., Puthanakar, N. Y., Naduwinmani, S., & Chidambar, Y. S. (2015). Association between periodontitis and Alzheimer’s disease. North American Journal of Medical Sciences, 7, 241–246.
Ahluwalia, B., Magnusson, M. K., & Ohman, L. (2017). Mucosal immune system of the gastrointestinal tract: Maintaining balance between the good and the bad. Scandinavian Journal of Gastroenterology, 52, 1185–1193.
Albandar, J. M., & Tinoco, E. M. (2002). Global epidemiology of periodontal diseases in children and young persons. Periodontology 2000, 29, 153–176.
Barak, S., Oettinger-Barak, O., Oettinger, M., Machtei, E. E., Peled, M., & Ohel, G. (2003). Common oral manifestations during pregnancy: A review. Obstetrical & Gynecological Survey, 58, 624–628.
Belibasakis, G. N. (2018). Microbiological changes of the ageing oral cavity. Archives of Oral Biology, 96, 230–232.
Belibasakis, G. N., Kast, J. I., Thurnheer, T., Akdis, C. A., & Bostanci, N. (2015). The expression of gingival epithelial junctions in response to subgingival biofilms. Virulence, 6, 704–709.
Bimstein, E., & Ebersole, J. L. (1989). The age-dependent reaction of the periodontal tissues to dental plaque. ASDC Journal of Dentistry for Children, 56, 358–362.
Bimstein, E., Ram, D., Irshied, J., Naor, R., & Sela, M. N. (2002). Periodontal diseases, caries, and microbial composition of the subgingival plaque in children: A longitudinal study. ASDC Journal of Dentistry for Children, 69, 133–137. 123.
Bimstein, E., Huja, P. E., & Ebersole, J. L. (2013). The potential lifespan impact of gingivitis and periodontitis in children. The Journal of Clinical Pediatric Dentistry, 38, 95–99.
Bosshardt, D. D., & Lang, N. P. (2005). The junctional epithelium: From health to disease. Journal of Dental Research, 84, 9–20.
Brown, A. J., et al. (2017). “Chemokine CXCL7 Heterodimers: Structural Insights, CXCR2 Receptor Function, and Glycosaminoglycan Interactions.” Int J Mol Sci 18(4).
Cardoso, E. M., Reis, C., & Manzanares-Cespedes, M. C. (2018). Chronic periodontitis, inflammatory cytokines, and interrelationship with other chronic diseases. Postgraduate Medicine, 130, 98–104.
Chapple, I. L., Van der Weijden, F., Dorfer, C., et al. (2015). Primary prevention of periodontitis: Managing gingivitis. Journal of Clinical Periodontology, 42, S71.
Dale, B. A. (2002). Periodontal epithelium: A newly recognized role in health and disease. Periodontology 2000, 30, 70–78.
De Lorenzo, G., Ferrari, S., Cervone, F., & Okun, E. (2018). Extracellular DAMPs in plants and mammals: Immunity, tissue damage and repair. Trends in Immunology, 39, 937–950.
Desurmont, T., Skrypek, N., Duhamel, A., et al. (2015). Overexpression of chemokine receptor CXCR2 and ligand CXCL7 in liver metastases from colon cancer is correlated to shorter disease-free and overall survival. Cancer Science, 106, 262–269.
Doles, J., Storer, M., Cozzuto, L., Roma, G., & Keyes, W. M. (2012). Age-associated inflammation inhibits epidermal stem cell function. Genes & Development, 26, 2144–2153.
Ebersole, J. L., Steffen, M. J., Gonzalez-Martinez, J., & Novak, M. J. (2008). Effects of age and oral disease on systemic inflammatory and immune parameters in nonhuman primates. Clinical and Vaccine Immunology, 15, 1067–1075.
Ebersole, J. L., Kirakodu, S., Novak, M. J., et al. (2014). Cytokine gene expression profiles during initiation, progression and resolution of periodontitis. Journal of Clinical Periodontology, 41, 853.
Ebersole, J. L., Kirakodu, S. S., Novak, M. J., et al. (2016a). Transcriptome analysis of B cell immune functions in periodontitis: Mucosal tissue responses to the oral microbiome in aging. Frontiers in Immunology, 7, 272.
Ebersole, J. L., Kirakodu, S., Novak, M. J., et al. (2016b). Effects of aging in the expression of NOD-like receptors and inflammasome-related genes in oral mucosa. Molecular Oral Microbiology, 31, 18–32.
Ebersole, J. L., Dawson, D., 3rd, Emecen-Huja, P., et al. (2017). The periodontal war: Microbes and immunity. Periodontology 2000, 75, 52–115.
Ebersole, J. L., Novak, M. J., Orraca, L., et al. (2018). Hypoxia-inducible transcription factors, HIF1A and HIF2A, increase in aging mucosal tissues. Immunology, 154, 452–464.
Gonzalez, O. A., Stromberg, A. J., Huggins, P. M., Gonzalez-Martinez, J., Novak, M. J., & Ebersole, J. L. (2011). Apoptotic genes are differentially expressed in aged gingival tissue. Journal of Dental Research, 90, 880–886.
Gonzalez, O. A., John Novak, M., Kirakodu, S., et al. (2013). Effects of aging on apoptosis gene expression in oral mucosal tissues. Apoptosis, 18, 249–259.
Gonzalez, O. A., Novak, M. J., Kirakodu, S., et al. (2014). Comparative analysis of gingival tissue antigen presentation pathways in ageing and periodontitis. Journal of Clinical Periodontology, 41, 327–339.
Gonzalez, O. A., Novak, M. J., Kirakodu, S., et al. (2015). Differential gene expression profiles reflecting macrophage polarization in aging and periodontitis gingival tissues. Immunological Investigations, 44, 643–664.
Gonzalez, O. A., Kirakodu, S., Novak, M. J., et al. (2018). Comparative analysis of microbial sensing molecules in mucosal tissues with aging. Immunobiology, 223, 279–287.
Gumus, P., Ozturk, V. O., Bozkurt, E., & Emingil, G. (2016). Evaluation of the gingival inflammation in pregnancy and postpartum via 25-hydroxy-vitamin D3, prostaglandin E2 and TNF-alpha levels in saliva. Archives of Oral Biology, 63(1–6), 1.
Guncu, G. N., Yilmaz, D., Kononen, E., & Gursoy, U. K. (2015). Salivary antimicrobial peptides in early detection of periodontitis. Frontiers in Cellular and Infection Microbiology, 5, 99.
Gursoy, M., Zeidan-Chulia, F., Kononen, E., et al. (2014). Pregnancy-induced gingivitis and OMICS in dentistry: In silico modeling and in vivo prospective validation of estradiol-modulated inflammatory biomarkers. Omics: A Journal of Integrative Biology, 18, 582–590.
Hajishengallis, G. (2014). Aging and its impact on innate immunity and inflammation: Implications for periodontitis. Journal of Oral Biosciences/JAOB, Japanese Association for Oral Biology, 56, 30–37.
Hajishengallis, G., & Lamont, R. J. (2016). Dancing with the stars: How choreographed bacterial interactions dictate nososymbiocity and give rise to keystone pathogens, accessory pathogens, and pathobionts. Trends in Microbiology, 24, 477–489.
Hatakeyama, S., Yaegashi, T., Oikawa, Y., et al. (2006). Expression pattern of adhesion molecules in junctional epithelium differs from that in other gingival epithelia. Journal of Periodontal Research, 41, 322–328.
Ho, S., Pothoulakis, C., & Koon, H. W. (2013). Antimicrobial peptides and colitis. Current Pharmaceutical Design, 19, 40–47.
Huttner, E. A., Machado, D. C., de Oliveira, R. B., Antunes, A. G., & Hebling, E. (2009). Effects of human aging on periodontal tissues. Special Care in Dentistry, 29, 149–155.
Icer, M. A., & Gezmen-Karadag, M. (2018). The multiple functions and mechanisms of osteopontin. Clinical Biochemistry, 59, 17–24.
Jang, D. H., Bhawal, U. K., Min, H. K., Kang, H. K., Abiko, Y., & Min, B. M. (2015). A transcriptional roadmap to the senescence and differentiation of human oral keratinocytes. Journals of Gerontology, Series A: Biological Sciences and Medical Sciences, 70, 20–32.
Jenkins, W. M., & Papapanou, P. N. (2001). Epidemiology of periodontal disease in children and adolescents. Periodontology 2000, 26, 16–32.
Jin, G., & Weinberg, A. (2018). Human antimicrobial peptides and cancer. Seminars in Cell & Developmental Biology, 88, 156–162.
Kinane, D. F., & Hodge, P. J. (2001). Periodontal disease in children and adolescents: Introduction and classification. Periodontology 2000, 26, 7–15.
Kumar, P. S. (2013). Oral microbiota and systemic disease. Anaerobe, 24, 90–93.
Lamont, R. J., & Hajishengallis, G. (2015). Polymicrobial synergy and dysbiosis in inflammatory disease. Trends in Molecular Medicine, 21, 172–183.
Lamster, I. B., Asadourian, L., Del Carmen, T., & Friedman, P. K. (2016). The aging mouth: Differentiating normal aging from disease. Periodontology 2000, 72, 96–107.
Lang, N. P., Schatzle, M. A., & Loe, H. (2009). Gingivitis as a risk factor in periodontal disease. Journal of Clinical Periodontology, 36(Suppl 10), 3–8.
Loe, H., Anerud, A., Boysen, H., & Morrison, E. (1986). Natural history of periodontal disease in man. Rapid, moderate and no loss of attachment in Sri Lankan laborers 14 to 46 years of age. Journal of Clinical Periodontology, 13, 431–445.
Maddi, A., & Scannapieco, F. A. (2013). Oral biofilms, oral and periodontal infections, and systemic disease. American Journal of Dentistry, 26, 249–254.
McCormick, T. S., & Weinberg, A. (2010). Epithelial cell-derived antimicrobial peptides are multifunctional agents that bridge innate and adaptive immunity. Periodontology 2000, 54, 195–206.
Meka, A., Bakthavatchalu, V., Sathishkumar, S., et al. (2010). Porphyromonas gingivalis infection-induced tissue and bone transcriptional profiles. Molecular Oral Microbiology, 25, 61–74.
Modeer, T., & Wondimu, B. (2000). Periodontal diseases in children and adolescents. Dental Clinics of North America, 44, 633–658.
Moorefield, E. C., Andres, S. F., Blue, R. E., et al. (2017). Aging effects on intestinal homeostasis associated with expansion and dysfunction of intestinal epithelial stem cells. Aging, 9, 1898–1915.
Nassar, M., Tabib, Y., Capucha, T., et al. (2017). GAS6 is a key homeostatic immunological regulator of host-commensal interactions in the oral mucosa. Proceedings of the National Academy of Sciences of the United States of America, 114, E337–E346.
Olive, C. (2012). Pattern recognition receptors: Sentinels in innate immunity and targets of new vaccine adjuvants. Expert Review of Vaccines, 11, 237–256.
Papapanou, P. N., & Susin, C. (2017). Periodontitis epidemiology: Is periodontitis under-recognized, over-diagnosed, or both? Periodontology 2000, 75, 45–51.
Pardo-Camacho, C., Gonzalez-Castro, A. M., Rodino-Janeiro, B. K., Pigrau, M., & Vicario, M. (2018). Epithelial immunity: Priming defensive responses in the intestinal mucosa. American Journal of Physiology. Gastrointestinal and Liver Physiology, 314, G247–G255.
Parrish, A. R. (2017). The impact of aging on epithelial barriers. Tissue Barriers, 5, e1343172.
Partida-Rodriguez, O., Serrano-Vazquez, A., Nieves-Ramirez, M. E., et al. (2017). Human intestinal microbiota: Interaction between parasites and the host immune response. Archives of Medical Research, 48, 690–700.
Patel, S. (2018). Danger-associated molecular patterns (DAMPs): The derivatives and triggers of inflammation. Current Allergy and Asthma Reports, 18, 63.
Patsouras, M. D., Sikara, M. P., Grika, E. P., Moutsopoulos, H. M., Tzioufas, A. G., & Vlachoyiannopoulos, P. G. (2015). Elevated expression of platelet-derived chemokines in patients with antiphospholipid syndrome. Journal of Autoimmunity, 65, 30–37.
Rajaee, A., Barnett, R., & Cheadle, W. G. (2018). Pathogen- and danger-associated molecular patterns and the cytokine response in sepsis. Surgical Infections, 19, 107–116.
Sakai, T., Kiyoshima, T., Kobayashi, I., et al. (1999). Age-dependent changes in the distribution of BrdU- and TUNEL-positive cells in the murine gingival tissue. Journal of Periodontology, 70, 973–981.
Schenkein, H. A., Berry, C. R., Burmeister, J. A., et al. (2003). Anti-cardiolipin antibodies in sera from patients with periodontitis. Journal of Dental Research, 82, 919–922.
Shusterman, A., Munz, M., Richter, G., et al. (2017). The PF4/PPBP/CXCL5 gene cluster is associated with periodontitis. Journal of Dental Research, 96, 945–952.
Stocks, C. J., Schembri, M. A., Sweet, M. J., & Kapetanovic, R. (2018). For when bacterial infections persist: Toll-like receptor-inducible direct antimicrobial pathways in macrophages. Journal of Leukocyte Biology, 103, 35–51.
Sukhithasri, V., Nisha, N., Biswas, L., Anil Kumar, V., & Biswas, R. (2013). Innate immune recognition of microbial cell wall components and microbial strategies to evade such recognitions. Microbiological Research, 168, 396–406.
Tonetti, M. S., Chapple, I. L., Jepsen, S., & Sanz, M. (2015). Primary and secondary prevention of periodontal and peri-implant diseases introduction to, and objectives of the consensus from the 11 European workshop on periodontology. Journal of Clinical Periodontology, 42(Suppl 16), S1–S4.
Tsukamoto, Y., Usui, M., Yamamoto, G., et al. (2012). Role of the junctional epithelium in periodontal innate defense and homeostasis. Journal of Periodontal Research, 47, 750–757.
Van der Velden, U. (1984). Effect of age on the periodontium. Journal of Clinical Periodontology, 11, 281–294.
Wade, W. G. (2013). The oral microbiome in health and disease. Pharmacological Research, 69, 137–143.
Wael Youssef, E. (2018). Age-dependent differential expression of apoptotic markers in rat oral mucosa. Asian Pacific Journal of Cancer Prevention, 19, 3245–3250.
Walsh, D., McCarthy, J., O’Driscoll, C., & Melgar, S. (2013). Pattern recognition receptors--molecular orchestrators of inflammation in inflammatory bowel disease. Cytokine & Growth Factor Reviews, 24, 91–104.
Wu, Y., Dong, G., Xiao, W., et al. (2016). Effect of aging on periodontal inflammation, microbial colonization, and disease susceptibility. Journal of Dental Research, 95, 460–466.
Xia, Y., Sun, M., Xie, Y., & Shu, R. (2017). mTOR inhibition rejuvenates the aging gingival fibroblasts through alleviating oxidative stress. Oxidative Medicine and Cellular Longevity, 2017, 6292630.
Yeo, L., Adlard, N., Biehl, M., et al. (2016). Expression of chemokines CXCL4 and CXCL7 by synovial macrophages defines an early stage of rheumatoid arthritis. Annals of the Rheumatic Diseases, 75, 763–771.
Yu, L. C., Wang, J. T., Wei, S. C., & Ni, Y. H. (2012). Host-microbial interactions and regulation of intestinal epithelial barrier function: From physiology to pathology. World Journal of Gastrointestinal Pathophysiology, 3, 27–43.
Zhang, J., Wang, C. M., Zhang, P., et al. (2016). Expression of programmed death 1 ligand 1 on periodontal tissue cells as a possible protective feedback mechanism against periodontal tissue destruction. Molecular Medicine Reports, 13, 2423–2430.
Acknowledgements
This work was supported by National Institute of Health grants P20GM103538 and UL1TR000117. We express our gratitude to the Caribbean Primate Research Center (CPRC) supported by grant P40RR03640, and the Microarray Core of University Kentucky for their invaluable technical assistance. We thank M. Kirakodu for data management support.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Switzerland AG
About this paper
Cite this paper
Ebersole, J.L., Orraca, L., Novak, M.J., Kirakodu, S., Gonzalez-Martinez, J., Gonzalez, O.A. (2019). Comparative Analysis of Gene Expression Patterns for Oral Epithelium-Related Functions with Aging. In: Belibasakis, G.N., Hajishengallis, G., Bostanci, N., Curtis, M.A. (eds) Oral Mucosal Immunity and Microbiome. Advances in Experimental Medicine and Biology, vol 1197. Springer, Cham. https://doi.org/10.1007/978-3-030-28524-1_11
Download citation
DOI: https://doi.org/10.1007/978-3-030-28524-1_11
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-28523-4
Online ISBN: 978-3-030-28524-1
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)