Abstract
The application of one-class machine learning is gaining attention in the computational biology community. Many biological cases can be considered as multi one-class classification problem. Examples include the classification of multiple cancer types, protein fold recognition and, molecular classification of multiple tumor types. In all of those cases the real world appropriately characterized negative cases or outliers are impractical to be achieved and the positive cases might be consists from different clusters which in turn might reveal to accuracy degradation. In this paper, we present multi-one-class classifier to deal with this problem. The key point of our classification method is to run a clustering algorithm such as the well-known k-means over the positive cases and then building up a classifier for every cluster separately. For a given new example, we apply all the generated classifiers. If it rejected by all of those classifiers, the given example will be considered as a negative case, otherwise it is a positive case.
Access provided by Autonomous University of Puebla. Download conference paper PDF
Similar content being viewed by others
Keywords
1 Introduction
The one-class approach in machine learning has been receiving more attention particularly for solving problems where the negative class is not well defined [1,2,3,4,5]; moreover, the one class approach has been successfully applied in various fields including text mining [6], functional Magnetic Resonance Imaging (fMRI) [7], signature verification [8] and miRNA gene and target discovery [5, 9].
One and two class classification methods can both give useful classification accuracies. The advantage of one-class methods is that they do not require any additional effort for choosing the best way of generating the negative class. Some data is composed from multiple category or classes. For such a data, special methods are required and the one class gives insufficient results.
Here, we present a new approach that devise the positive class into sub-groups and then builds a classifier (one-class) for each sub-group.
The key point of our classification method is to run the k-means clustering algorithm over the positive cases and then building up a classifier for every cluster separately. For a given new sample the algorithm applies all the generated classifiers. If one of those classifiers classifies the given sample as a positive sample then it will be considered as a positive sample, otherwise it is a negative sample.
The problem of how to implement a multi-class classifier by an ensemble of one-class classifiers has attracted much attention in the machine-learning community [10, 11]. Tax et al. [12] proposed a method for combining different one-class classifiers, which improves dramatically the performance and the robustness of the classification, to the handwritten digit recognition problem. Similar method investigated by Lai et al. [13] to the problem of image retrieval problem. They reported that combining multi SVM based classifiers improves the retrieval precision.
Several methods for handling missing feature values which are based on combining one-class classifiers have been suggested by Juszczak et al. [14]. They indicated that their methods are more flexible and more robust to small sample size problems than the standard methods such as the linear programming dissimilarity-data description. Ban et al. [15] suggested using multiple one-class classifiers which can deal with the feature space and the nonlinear classification problem. They trained the multi one-class classifiers on each class and then extracted a decision function which based on minimum distance rules. Their experiments showed that the proposed method outperforms the OC-SVM.
Lyu and Fraid [16] provided a multi one-class SVMs which combines a beforehand clustering process for detecting hidden (Steganographic) messages in digital images. They show how a multi one-class SVM greatly simplifies the training stage of the classifiers. Their results show that even though the overall detection improves with an increasing number of hyper-spheres (i.e., the clusters), the false-positive rate begins to increase considerably with the increases of the number of the hyper-spheres. Recently, Menahem et al. [17] described a different multiple one-class classification approach called TUPSO which combines multi one-class classifiers via meta-classifier. They showed that TUPSO outperforms existing methods such as the OC-SVM.
Much works on multi one-class classification are existed concerning the computational biology community. Spinosa et al. [18] suggested a multi one-class classification technique to detect novelty in gene expression data. They combined different one-class classifiers such as OC-Kmeans and OC-KNN. For a given example, each classifier votes ‘positive’ or ‘outlier’. The final decision is taken according to the majority votes of all classifiers. Results showed that the robustness of the classification is increased because it takes into account different classifiers such that each one judges the examples in different point of view. Recently, similar approach provided by Zhang et al. [19] for the avian influenza outbreak classification problem.
Our new approach distinguished itself from those predecessors in the way that it first clusters the positive data into groups before the classification process. This pre-processing prevents the drawback of using only a single hyper-sphere (i.e., the one generated by the one-class classifier) which may not provide a particularly compact support for the training data. Our experiments show that our approach significantly improves the accuracy of the classification.
The rest of this paper is organized as follows: Sect. 2 describes necessary preliminaries. Our multi one-class approach is described in Sect. 3 and evaluated in Sect. 4. Our main conclusions and future work can be found in Sect. 5.
2 Methods
2.1 One-Class Methods
In general, a binary learning (two-class) approach to miRNA discovery considers both positive (miRNA) and negative (non-miRNA) classes by providing examples for the two-classes to a learning algorithm in order to build a classifier that will attempt to discriminate between them. The most common term for this kind of learning is supervised learning where the labels of the two-classes are known beforehand.
One-class uses only the information for the target class (positive class) building a classifier which is able to recognize the examples belonging to its target and rejecting others as outliers. Among the many classification algorithms available, we chose four one-class algorithms to compare for miRNA discovery. We give a brief description of each one-class classifier and we refer the references [20, 21] for additional details including a description of parameters and thresholds. The LIBSVM library [22] was used as implementation of the SVM (one-class using the RBF kernel function) and the DDtools [23] for the other one-class methods. The WEKA software [24] was used as implementation of the two-class classifiers.
2.2 One-Class Support Vector Machines (OC-SVM)
Support Vector Machines (SVMs) are a learning machine developed as a two-class approach [25, 26]. The use of one-class SVM was originally suggested by [21]. One-class SVM is an algorithmic method that produces a prediction function trained to “capture” most of the training data. For that purpose a kernel function is used to map the data into a feature space where the SVM is employed to find the hyper-plane with maximum margin from the origin of the feature space. In this use, the margin to be maximized between the two classes (in two-class SVM) becomes the distance between the origin and the support vectors which define the boundaries of the surrounding circle, (or hyper-sphere in high-dimensional space) which encloses the single class.
2.3 One-Class Gaussian (OC-Gaussian)
The Gaussian model is considered as a density estimation model. The assumption is that the target samples form a multivariate normal distribution, therefore for a given test sample z in n-dimensional space, the probability density function can be calculated as:
where \( \mu \) and \( \varSigma \) are the mean and covariance matrix of the target class estimated from the training samples.
2.4 One-Class Kmeans (OC-Kmeans)
Kmeans is a simple and well-known unsupervised machine learning algorithm used in order to partition the data into k clusters. Using the OC-Kmeans we describe the data as k clusters, or more specifically as k centroids, one derived from each cluster. For a new sample, z, the distance d(z) is calculated as the minimum distance to each centroid. Then based on a user specified threshold, the classification decision is made. If d(z) is less than the threshold the new sample belongs to the target class, otherwise it is rejected.
2.5 One-Class K-Nearest Neighbor (OC-KNN)
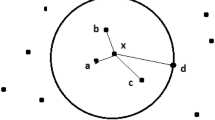
The one-class nearest neighbor classifier (OC-KNN) is a modification of the known two-class nearest neighbor classifier which learns from positive examples only. The algorithm operates by storing all the training examples as its model, then for a given test example z, the distance to its nearest neighbor y (y = NN(z)) is calculated as d(z,y). The new sample belongs to the target class when:
where NN(y) is the nearest neighbor of y, in other words, it is the nearest neighbor of the nearest neighbor of z. The default value of \( \delta \) is 1. The average distance of the k nearest neighbors is considered for the OC-KNN implementation.
3 Multi One-Class Classifiers
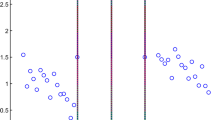
For a given data, the positive class might be consisting of different clusters (See Fig. 1). If we try to find the small hyper-sphere that contains all the data points belongs to the positive class, then the negative data points will be a part of the hyper-sphere yielding in low performance.
To alleviate this type of data we propose the Multi One-Class Classifiers that works with small groups of the positive data. Our approach based on the OC-SVM that builds a hypersphere for a given positive data and we aim that working with compacted groups is better that work with wide group.
As a result, we run clustering to identify the similar data points. The main idea of our approach is to cluster the positive data into several clusters using the k-means clustering algorithm and then build up a classifier from each cluster using the OC-SVM as can be seen in Fig. 2.
For a given new example, we apply all the classifiers. If all of them reject it then it considers as negative example; otherwise it considers as belongs to the one-class data. In this work, without loss of generality, we choose to evaluate our Multi One-Class approach using the OC-SVM as the basis classification algorithm and the standard K-means as the clustering method. We leave other possible combinations for future work.
4 Results
The first experiment is a typical one, which illustrates the performance of Multi-OC-SVM versus that of the normal OC-SVM over a synthetic data set (See Fig. 1). In each experiment, the data set is split into two subsets, one for training and one for testing. Both algorithms were trained using 80% of the positive class and the remaining 20% together with all the negative examples were used for testing. The positive examples are divided into four clusters beforehand. We used the standard k-means algorithm for this purpose.
As can be seen from Fig. 3, executing the OC-SVM method reveals into poor classification. This is to be expected since OC-SVM classifies part of the positive examples as negative ones (or as outliers). Figure 4 depicts the classification done by the Multi-OC-SVMs. As can be seen, obviously Multi-OC-SVM outperforms OC-SVM.
Next we ran both OC-SVM and Multi-OC-SVM twenty times over the data set in order to evaluate their performance and stability using different values of k. The data set used for this experiment is similar to the first one. Additionally, the Matthews Correlation Coefficient (MCC) (see [27] for more details) measurement is used to take into account both over-prediction and under-prediction in imbalanced data sets. Looking at Fig. 4, one can see that the accuracy of Multi-OC-SVM is far better than that of OC-SVM. Furthermore, it shows that Multi-OC-SVM is stable over different number of clusters.
5 Conclusion
The current results show that it is possible to build up a multi one-class classifier with a combined clustering beforehand process based only on positive examples yielding a reasonable performance.
Further research can proceed in several interesting directions. First, the suitability of the framework of our approach to different data types can be investigated. Second, it would be interesting to apply our approach to other types of classifiers and to more robust clustering methods such as Mean-Shift [28]. Lastly, it would be interesting to test our approach on more real data-sets and real problems.
References
Kowalczyk, A., Raskutti, B.: One class SVM for yeast regulation prediction. SIGKDD Explor. 4(2), 99–100 (2002)
Spinosa, E.J., Carvalho, A.C.: Support vector machines for novel class detection in Bioinformatics. Genet. Mol. Res. (GMR) 4(3), 608–615 (2005)
Crammer, K., Chechik, G.: A needle in a haystack: local one-class optimization. In: Proceedings of the Twenty-First International Conference on Machine Learning (ICML) (2004)
Gupta, G., Ghosh, J.: Robust one-class clustering using hybrid global and local search. In: Proceedings of the 22nd International Conference on Machine Learning. ACM Press, Bonn (2005)
Yousef, M., Najami, N., Khalifa, W.: A comparison study between one-class and two-class machine learning for MicroRNA target detection. J. Biomed. Sci. Eng. 3, 247 (2010)
Manevitz, L.M., Yousef, M.: One-class SVMs for document classification. J. Mach. Learn. Res. 2, 139–154 (2001)
Thirion, B., Faugeras, O.: Feature characterization in fMRI data: the information bottleneck approach. Med. Image Anal. 8(4), 403 (2004)
Koppel, M., Schler, J.: Authorship verification as a one-class classification problem. In: Proceedings of the Twenty-First International Conference on Machine Learning. ACM Press, Banff (2004)
Yousef, M., et al.: Learning from positive examples when the negative class is undetermined- microRNA gene identification. Algorithms Mol. Biol. 3(1), 2 (2008)
AbedAllah, L., Shimshoni, I.: k nearest neighbor using ensemble clustering. In: Cuzzocrea, A., Dayal, U. (eds.) DaWaK 2012. LNCS, vol. 7448, pp. 265–278. Springer, Heidelberg (2012). https://doi.org/10.1007/978-3-642-32584-7_22
Landgrebea, T.C., Paclıka, D.M., Andrew, R.P.: One-Class and Multi-Class Classifier Combining for Ill-Defined Problems. Elsevier Science, Amsterdam (2005)
Tax, D.M.J., Duin, R.P.W.: Combining one-class classifiers. In: Kittler, J., Roli, F. (eds.) MCS 2001. LNCS, vol. 2096, pp. 299–308. Springer, Heidelberg (2001). https://doi.org/10.1007/3-540-48219-9_30
Lai, C., Tax, D.M.J., Duin, R.P.W., Pękalska, E., Paclík, P.: On combining one-class classifiers for image database retrieval. In: Roli, F., Kittler, J. (eds.) MCS 2002. LNCS, vol. 2364, pp. 212–221. Springer, Heidelberg (2002). https://doi.org/10.1007/3-540-45428-4_21
Juszczak, P., Duin, R.P.W.: Combining one-class classifiers to classify missing data. In: Roli, F., Kittler, J., Windeatt, T. (eds.) MCS 2004. LNCS, vol. 3077, pp. 92–101. Springer, Heidelberg (2004). https://doi.org/10.1007/978-3-540-25966-4_9
Ban, T., Abe, S.: Implementing multi-class classifiers by one-class classification methods. In: International Joint Conference on Neural Networks, IJCNN 2006 (2006)
Lyu, S., Farid, H.: Steganalysis using color wavelet statistics and one-class support vector machines. In: SPIE Symposium on Electronic Imaging, pp. 35–45 (2004)
Menahem, E., Rokach, L., Elovici, Y.: Combining One-Class Classifiers via Meta-learning. CoRR, abs/1112.5246 (2011)
Spinosa, E.J., de Carvalho, A.C.P.L.F.: Combining one-class classifiers for robust novelty detection in gene expression data. In: Setubal, J.C., Verjovski-Almeida, S. (eds.) BSB 2005. LNCS, vol. 3594, pp. 54–64. Springer, Heidelberg (2005). https://doi.org/10.1007/11532323_7
Zhang, J., Lu, J., Zhang, G.: Combining one class classification models for avian influenza outbreaks. In: Computational Intelligence in Multicriteria Decision-Making (MDCM), pp. 190–196. IEEE (2011)
Tax, D.M.J.: One-class classification; concept-learning in the absence of counter-examples. Delft University of Technology, June 2001
Schölkopf, B., et al.: Estimating the support of a high-dimensional distribution. Neural Comput. 13(7), 1443–1471 (2001)
Chang, C.-C., Lin, C.-J.: LIBSVM: a library for support vector machines. ACM Trans. Intell. Syst. Technol. (TIST) 2, 27 (2001)
Tax, D.M.J.: DDtools, the Data Description Toolbox for Matlab. Delft University of Technology (2005)
Witten, I.H., Frank, E.: Data Mining: Practical Machine Learning Tools and Techniques, 2nd edn. Morgan Kaufmann, San Francisco (2005)
Schölkopf, B., Burges, C.J.C., Smola, A.J.: Advances in Kernel Methods. MIT Press, Cambridge (1999)
Vapnik, V.: The Nature of Statistical Learning Theory. Springer, Heidelberg (1995). https://doi.org/10.1007/978-1-4757-3264-1
Matthews, B.W.: Comparison of the predicted and observed secondary structure of T4 phage lysozyme. Biochim. Biophys. Acta 405(2), 442–451 (1975)
Comaniciu, D., Meer, P.: Mean shift: a robust approach toward feature space analysis. IEEE Trans. Pattern Anal. Mach. Intell. 24(5), 603–619 (2002)
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Switzerland AG
About this paper
Cite this paper
Abedalla, L., Badarna, M., Khalifa, W., Yousef, M. (2019). K – Means Based One-Class SVM Classifier. In: Anderst-Kotsis, G., et al. Database and Expert Systems Applications. DEXA 2019. Communications in Computer and Information Science, vol 1062. Springer, Cham. https://doi.org/10.1007/978-3-030-27684-3_7
Download citation
DOI: https://doi.org/10.1007/978-3-030-27684-3_7
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-27683-6
Online ISBN: 978-3-030-27684-3
eBook Packages: Computer ScienceComputer Science (R0)