Abstract
Stem cell antigen-1 (Sca-1) is the first identified member of mouse Ly6 gene family. We discovered that Sca-1 disrupts TGFβ signaling and enhances mammary tumorigenesis in a DMBA-induced mammary tumor model. Sca-1 gene is lost during evolution in humans. Human Ly6 genes Ly6D, LyE, LyH, and LyK on human chromosome 8q24.3 genes are syntenic to the mouse chromosome 15 where Sca-1 is located. We found that Ly6D, E, H, and K are upregulated in human cancer compared to normal tissue and that the increased expression of these genes are associated with poor prognosis of multiple types of human cancer. Several other groups have indicated increased expression of Ly6 genes in human cancer. Here we described the relevance of expression of human Ly6D, LyE, LyH, and LyK in functioning of normal tissues and tumor progression.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
Introduction
Stem cell antigen-1 (Sca-1) was the first identified member of mouse Ly6 gene family [1]. Sca-1 has been described as marker of tissue-resident stem cells and also been recognized as cancer initiating cell population in multiple mouse model of mammary, prostate, and lung cancer among other cancer types [2,3,4]. We described that Sca-1 binds with TGFβ receptor 1 (TβR1) and disrupts the TGFβ receptor complex, leading to tumorigenic progression in mouse model of mammary tumorigenesis [5]. A direct homologue of Sca-1 is missing in humans; however, multiple members of mouse Ly6 genes and syntenic human Ly6 genes have been described. Human Ly6 gene family usually contain LU domain. These molecules are scattered throughout the human chromosome 1, 2, 6, 8, 11, and 19 [6]. We will concern our description to human Ly6D, LyE, LyH, and Ly6K, which are present on human chromosome 8q24.3.
Expression of Ly6D, E, H, and K Genes in Normal Tissues
Ly6D
Ly6D RNA is expressed in highest quantities in esophageal tissue and in skin as shown in HPA dataset, GTEx dataset, and Fantome5 datasets as visualized in Human Protein Atlas available from www.proteinatlas.org [7,8,9]. The RNA expression was undetectable or expressed at a very low level in normal ovary, lung, testis, colon, breast, and prostate (Fig. 1a–c). The protein expression corroborated with RNA expression, showing increased expression of Ly6D in esophageal tissue and in skin (Fig. 2).
RNA expression of Ly6D in normal human tissues. (a) HPA data set, (b) GTEx data set, and (c) FANTOM dataset. The data was visualized on the human proteome webtool with expression filter on. X-axis shows the name of tissue type used. Y-axis shows the units of RNA expression as described below. HPA dataset: RNA-seq tissue data is reported as mean TPM (transcripts per million), corresponding to mean values of the different individual samples from each tissue. Color coding is based on tissue groups, each consisting of tissues with functional features in common. GTEx dataset: RNA-seq data is reported as median RPKM (reads per kilobase per million mapped reads), generated by the Genotype-Tissue Expression (GTEx) project
Ly6D protein expression in normal tissues. Immunohistochemistry was performed using anti-Ly6D antibody-rabbit polyclonal affinity purified antibody HPA024755, HPA064317 from Sigma-Aldrich as described in the human protein atlas webtool: https://www.proteinatlas.org/ENSG00000167656-LY6D/tissue
Ly6E
Ly6E RNA expression was identified in liver tissue, placenta, lung, and spleen by three different databases (Fig. 3a–c) as visualized in Human Protein Atlas available from www.proteinatlas.org [7,8,9]. The corroborative protein data was not yet available at the human protein atlas. Lowest expression of Ly6E RNA in normal tissues was found in pancreas and skeleton tissues (Fig. 3a–c). The Ly6E expression is restricted to syncytiotrophoblast cells of the mouse placenta [10].
RNA expression of Ly6E in normal human tissues. (a) HPA data set, (b) GTEx data set, and (c) FANTOM dataset. The data was visualized on the human proteome webtool with expression filter on. X-axis shows the name of tissue type used. Y-axis shows the units of RNA expression as described below. HPA dataset: RNA-seq tissue data is reported as mean TPM (transcripts per million), corresponding to mean values of the different individual samples from each tissue. Color-coding is based on tissue groups, each consisting of tissues with functional features in common. GTEx dataset: RNA-seq data is reported as median RPKM (reads per kilobase per million mapped reads), generated by the Genotype-Tissue Expression (GTEx) project
Ly6H
Ly6H RNA expression was identified highest in Brain tissue in HPA dataset, GTEx dataset, and Fantome5 (Fig. 4a–c) as visualized in Human Protein Atlas available from www.proteinatlas.org [7,8,9]. Ly6H plays an important role in glutamatergic signaling in brain [11].
RNA expression of Ly6H in normal human tissues. (a) HPA data set, (b) GTEx data set, and (c) FANTOM dataset. The data was visualized on the human proteome webtool with expression filter on. X-axis shows the name of tissue type used. Y-axis shows the units of RNA expression as described below. HPA dataset: RNA-seq tissue data is reported as mean TPM (transcripts per million), corresponding to mean values of the different individual samples from each tissue. Color coding is based on tissue groups, each consisting of tissues with functional features in common. GTEx dataset: RNA-seq data is reported as median RPKM (reads per kilobase per million mapped reads), generated by the Genotype-Tissue Expression (GTEx) project
Ly6K
Ly6K RNA is expressed highest in testis in HPA dataset, GTEx dataset, and Fantome5 datasets (Fig. 5a–c) as visualized in Human Protein Atlas available from www.proteinatlas.org [7,8,9]. The RNA expression was undetectable or expressed at a very low level in normal ovary, lung, testis, colon, breast, and prostate. The protein expression data showed that Ly6K is exclusively expressed in testis (Fig. 6). Male Ly6K homozygous knockout mice showed normal mating habits; however, they were infertile. Female Ly6K homozygous knockout mice were fertile. The male infertility was found to be associated with sperm migration [12].
RNA expression of Ly6K in normal human tissues. (a) HPA data set, (b) GTEx data set, and (c) FANTOM dataset were visualized on the human proteome webtool with expression filter on. X-axis shows the name of tissue type used. Y-axis shows the units of RNA expression as described below. HPA dataset: RNA-seq tissue data is reported as mean TPM (transcripts per million), corresponding to mean values of the different individual samples from each tissue. Color coding is based on tissue groups, each consisting of tissues with functional features in common. GTEx dataset: RNA-seq data is reported as median RPKM (reads per kilobase per million mapped reads), generated by the Genotype-Tissue Expression (GTEx) project
Ly6K protein expression data in normal tissues. Ly6K protein is exclusively expressed on human testis. Immunohistochemistry was performed using anti-Ly6K antibody-rabbit polyclonal affinity-purified antibody HPA017770 and protein A/G purified mouse monoclonal antibody AMAb90986, AMAb90987 from Sigma-Aldrich as described in the human protein atlas webtool: https://www.proteinatlas.org/ENSG00000160886-LY6K/tissue
Expression of Ly6D, E, H, and K Genes in Tumor Tissues
Ly6D
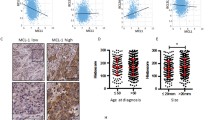
We looked for RNA expression of Ly6D in various publically available dataset and visualized using Oncomine [13] and TCGA [14] as described by Luo et al. [15]. Ly6D was found to be increased in tumors of ovarian, colorectal, gastric, breast, lung, bladder, brain and CNS, cervical, esophageal, head and neck and pancreatic cancer compared to normal tissue in multiple studies. The increased expression of Ly6D was associated with poor survival in ovarian, colorectal, gastric, breast and lung cancer [15] (Table 1). Since our publication new clinical data were available and added to KM plotter tool [16] (http: //kmplot.com/), which showed that increased Ly6D expression is associated with poor prognosis in renal clear cell carcinoma and pancreatic ductal adenocarcinoma (Fig. 7). Recently Ly6D expression in addition to OLFM4, S100A7 was found to be associated with distant metastasis of estrogen receptor-positive breast cancer [17]. Ly6D was shown to be increased in aggressive forms of head and neck cancer [18].
Ly6E
We looked for RNA expression of Ly6E in various publically available dataset and visualized using Oncomine [13] and TCGA [14] as described by Luo et al. [15]. Ly6E was found be increased in tumors of ovarian, colorectal, gastric, breast, lung, bladder, brain and CNS, cervical, esophageal, head and neck and pancreatic cancer compared to normal tissue in multiple studies. The increased expression of Ly6E was associated with poor survival in ovarian, colorectal, gastric, breast, lung, bladder, and brain and CNS cancer patients [15] (Table 1). Recent data added to KM plotter [16] (http://kmplot.com/) show that increased expression of Ly6E is associated with poor overall survival of renal papillary cell carcinoma and good prognostic marker for renal clear cell carcinoma (Fig. 8a, b). These new data indicated that increased expression of Ly6E is associated with poor overall survival of pancreatic ductal adenocarcinoma (Fig. 8c). The use of genome wide data analysis has prompted several new reports showing increased expression of Ly6E in bladder cancer and gastric cancer [19, 20]. The Ly6E gene has been also associated with more aggressive stem-like cells in hepatocellular carcinoma, pancreatic carcinoma, colon, and kidney [21,22,23].
Ly6H
We looked for RNA expression of Ly6H in various publically available dataset and visualized using Oncomine [13] and TCGA [14] as described by Luo et al. [15]. Ly6H was found be increased in tumors of ovarian, colorectal, gastric, breast, lung, bladder, brain and CNS, cervical, esophageal, head and neck and pancreatic cancer compared to normal tissue in multiple studies. The increased expression of Ly6H was associated with poor survival in ovarian, colorectal, gastric, and breast cancer patients [15] (Table 1). Recent data added to KM plotter [16] (http://kmplot.com/) show that increased expression of Ly6H is associated with poor overall survival of renal clear cell carcinoma and pancreatic ductal adenocarcinoma (Fig. 9a, b).
Ly6K
We looked for RNA expression of Ly6K in various publically available dataset and visualized using Oncomine [13] and TCGA [14] as described by Luo et al. [15]. Ly6K was found be increased in tumors of ovarian, colorectal, gastric, breast, lung, bladder, brain and CNS, cervical, esophageal, head and neck and pancreatic cancer compared to normal tissue in multiple studies. The increased expression of Ly6K was associated with poor survival in ovarian, colorectal, gastric, breast, lung, bladder, and brain and CNS cancer patients [15] (Table 1). Increased expression of Ly6K in metastatic ER-positive breast cancer [24, 25], esophageal squamous cancer [26], gingivobuccal cancers [27], bladder cancer [28], and lung cancer [29] was observed. Recent data added to KM plotter [16] (http://kmplot.com/) show that increased expression of Ly6K is associated with poor overall survival of renal clear cell carcinoma, renal papillary cell carcinoma, and uterine corpus endometrial carcinoma (Fig. 10a–c).
Mechanisms Associated Ly6D, E, H, and K Gene Family
Ly6D
The functional role of Ly6D in human or mouse is not very well described. Ly6D was found to be important in selection of CD4+CD8+ subset of T cells in thymus [30]. Ly6D was shown to be involved in cell adhesion using NIH3T3 fibroblast cells [31]. The molecular pathways which may lead to regulation of Ly6D remain to be understood.
Ly6E
We discovered that Ly6E is required for increased TGFβ signaling, IFNγ/PDL1 signaling in breast cancer [32]. Ly6E was also shown to modulate the PTEN/PI3K/Akt/HIF-1 axis in breast cancer cell lines [33]. Ly6E was found to be important in enhancing the infectivity of multiple, enveloped RNA viruses in late steps using influenza A virus model [34]. Ly6E has been shown to be an interferon-inducible gene in lymphoid cells and plays an important role in HIV infection [35, 36]. Mouse Ly6E is a receptor syncytin A and plays important role in syncytiotrophoblast fusion and placental morphogenesis [37, 38].
Ly6H
Ly6H is required for nicotine-induced glutamatergic signaling via synaptic signaling of alpha7 nicotinic acetylcholine receptors [11].
Ly6K
We discovered that Ly6E is required for increased TGFβ signaling, IFNγ/PDL1 signaling in breast cancer [32]. High Ly6K expression is shown to have been linked with low expression of tumor suppression microRNA 500a-3p in non-small cell lung carcinoma [39]. AP1 activation is shown to be important in increased expression of Ly6K in breast cancer cell lines [40]. Ly6K protein expression in testis was shown to be stabilized by TEX101, a glycoprotein important in fertility [41].
Summary
The important role of Ly6 gene family in physiology and disease is emerging. The Ly6 gene family members can emerge as valid therapeutic target in correcting conditions of infection, nicotine addiction, cancer, immunotherapy, and fertility. Our laboratory is focused on understanding the molecular mechanism of Ly6 protein in cancer and validating Ly6 molecules as therapeutic targets in targeted cancer therapy. The network analysis of Ly6 gene family members showed that Ly6 signaling is involved in a broad range of molecules including growth factor, nuclear receptor, and micro RNAs (Fig. 11a). Pathway studio network analysis showed that Ly6 gene family affect multitude of cellular fate and cell–cell interaction with microenvironment ranging from growth, apoptosis, autophagy, and immune response (Fig. 11b).
Network analysis of Ly6 gene family members. (a) Pathway studio network analysis showed that Ly6 signaling is involved in broad range of molecules including growth factor, nuclear receptor, and micro RNAs. The upstream regulators are not highlighted, the downstream effectors are highlighted with blue, and the potential binding partners are highlighted with green. (b) Pathway studio network analysis showed that Ly6 gene family affect multitude of cellular fate and cell–cell interaction with microenvironment ranging from growth, apoptosis, autophagy, and immune response
References
Yutoku, M., Grossberg, A. L., & Pressman, D. (1974). A cell surface antigenic determinant present on mouse plasmacytes and only about half of mouse thymocytes. Journal of Immunology, 112(5), 1774–1781.
Dall, G. V., et al. (2017). SCA-1 labels a subset of estrogen-responsive bipotential repopulating cells within the CD24(+) CD49f(hi) mammary stem cell-enriched compartment. Stem Cell Reports, 8(2), 417–431.
Ceder, J. A., Aalders, T. W., & Schalken, J. A. (2017). Label retention and stem cell marker expression in the developing and adult prostate identifies basal and luminal epithelial stem cell subpopulations. Stem Cell Research & Therapy, 8(1), 95.
Zakaria, N., et al. (2018). Inhibition of NF-kappaB signaling reduces the stemness characteristics of lung cancer stem cells. Frontiers in Oncology, 8, 166.
Upadhyay, G., et al. (2011). Stem cell antigen-1 enhances tumorigenicity by disruption of growth differentiation factor-10 (GDF10)-dependent TGF-beta signaling. Proceedings of the National Academy of Sciences of the United States of America, 108(19), 7820–7825.
Loughner, C. L., et al. (2016). Organization, evolution and functions of the human and mouse Ly6/uPAR family genes. Human Genomics, 10, 10.
Thul, P. J., et al. (2017). A subcellular map of the human proteome. Science, 356(6340), eaal3321.
Uhlen, M., et al. (2015). Proteomics. Tissue-based map of the human proteome. Science, 347(6220), 1260419.
Uhlen, M., et al. (2017). A pathology atlas of the human cancer transcriptome. Science, 357(6352), eaan2507.
Hughes, M., et al. (2013). Ly6e expression is restricted to syncytiotrophoblast cells of the mouse placenta. Placenta, 34(9), 831–835.
Puddifoot, C. A., et al. (2015). Ly6h regulates trafficking of alpha7 nicotinic acetylcholine receptors and nicotine-induced potentiation of glutamatergic signaling. The Journal of Neuroscience, 35(8), 3420–3430.
Fujihara, Y., Okabe, M., & Ikawa, M. (2014). GPI-anchored protein complex, LY6K/TEX101, is required for sperm migration into the oviduct and male fertility in mice. Biology of Reproduction, 90(3), 60.
Rhodes, D. R., et al. (2004). ONCOMINE: A cancer microarray database and integrated data-mining platform. Neoplasia, 6(1), 1–6.
Cancer Genome Atlas Research Network, et al. (2013). The cancer genome atlas pan-cancer analysis project. Nature Genetics, 45(10), 1113–1120.
Luo, L., et al. (2016). Distinct lymphocyte antigens 6 (Ly6) family members Ly6D, Ly6E, Ly6K and Ly6H drive tumorigenesis and clinical outcome. Oncotarget, 7(10), 11165–11193.
Nagy, A., et al. (2018). Validation of miRNA prognostic power in hepatocellular carcinoma using expression data of independent datasets. Scientific Reports, 8(1), 9227.
Mayama, A., et al. (2018). OLFM4, LY6D and S100A7 as potent markers for distant metastasis in estrogen receptor-positive breast carcinoma. Cancer Science, 109(10), 3350–3359.
Colnot, D. R., et al. (2004). Clinical significance of micrometastatic cells detected by E48 (Ly-6D) reverse transcription-polymerase chain reaction in bone marrow of head and neck cancer patients. Clinical Cancer Research, 10(23), 7827–7833.
Dhawan, D., et al. (2018). Naturally-occurring canine invasive urothelial carcinoma harbors luminal and basal transcriptional subtypes found in human muscle invasive bladder cancer. PLoS Genetics, 14(8), e1007571.
Lv, Y., et al. (2018). Overexpression of lymphocyte antigen 6 complex, locus E in gastric cancer promotes cancer cell growth and metastasis. Cellular Physiology and Biochemistry, 45(3), 1219–1229.
Kondoh, N., et al. (1999). Identification and characterization of genes associated with human hepatocellular carcinogenesis. Cancer Research, 59(19), 4990–4996.
Gou, S., et al. (2007). Establishment of clonal colony-forming assay for propagation of pancreatic cancer cells with stem cell properties. Pancreas, 34(4), 429–435.
Bresson-Mazet, C., Gandrillon, O., & Gonin-Giraud, S. (2008). Stem cell antigen 2: A new gene involved in the self-renewal of erythroid progenitors. Cell Proliferation, 41(5), 726–738.
Kim, Y. S., et al. (2016). miRNAs involved in LY6K and estrogen receptor alpha contribute to tamoxifen-susceptibility in breast cancer. Oncotarget, 7(27), 42261–42273.
Kong, H. K., et al. (2016). Epigenetic activation of LY6K predicts the presence of metastasis and poor prognosis in breast carcinoma. Oncotarget, 7(34), 55677–55689.
Zhang, B., et al. (2012). Serological antibodies against LY6K as a diagnostic biomarker in esophageal squamous cell carcinoma. Biomarkers, 17(4), 372–378.
Ambatipudi, S., et al. (2012). Genome-wide expression and copy number analysis identifies driver genes in gingivobuccal cancers. Genes, Chromosomes & Cancer, 51(2), 161–173.
Matsuda, R., et al. (2011). LY6K is a novel molecular target in bladder cancer on basis of integrate genome-wide profiling. British Journal of Cancer, 104(2), 376–386.
Ishikawa, N., et al. (2007). Cancer-testis antigen lymphocyte antigen 6 complex locus K is a serologic biomarker and a therapeutic target for lung and esophageal carcinomas. Cancer Research, 67(24), 11601–11611.
Reese, J. T., et al. (2001). Downregulated expression of Ly-6-ThB on developing T cells marks CD4+CD8+ subset undergoing selection in the thymus. Developmental Immunology, 8(2), 107–121.
Brakenhoff, R. H., et al. (1995). The human E48 antigen, highly homologous to the murine Ly-6 antigen ThB, is a GPI-anchored molecule apparently involved in keratinocyte cell-cell adhesion. The Journal of Cell Biology, 129(6), 1677–1689.
AlHossiny, M., et al. (2016). Ly6E/K signaling to TGFbeta promotes breast cancer progression, immune escape, and drug resistance. Cancer Research, 76(11), 3376–3386.
Yeom, C. J., et al. (2016). LY6E: A conductor of malignant tumor growth through modulation of the PTEN/PI3K/Akt/HIF-1 axis. Oncotarget, 7(40), 65837–65848.
Mar, K. B., et al. (2018). LY6E mediates an evolutionarily conserved enhancement of virus infection by targeting a late entry step. Nature Communications, 9(1), 3603.
Yu, J., Liang, C., & Liu, S. L. (2017). Interferon-inducible LY6E protein promotes HIV-1 infection. The Journal of Biological Chemistry, 292(11), 4674–4685.
Xu, X., et al. (2014). IFN-stimulated gene LY6E in monocytes regulates the CD14/TLR4 pathway but inadequately restrains the hyperactivation of monocytes during chronic HIV-1 infection. Journal of Immunology, 193(8), 4125–4136.
Langford, M. B., et al. (2018). Deletion of the Syncytin A receptor Ly6e impairs syncytiotrophoblast fusion and placental morphogenesis causing embryonic lethality in mice. Scientific Reports, 8(1), 3961.
Bacquin, A., et al. (2017). A cell fusion-based screening method identifies Glycosylphosphatidylinositol-anchored protein Ly6e as the receptor for mouse endogenous retroviral envelope Syncytin-A. Journal of Virology, 91(18), e00832-17.
Liao, X. H., Xie, Z., & Guan, C. N. (2018). MiRNA-500a-3p inhibits cell proliferation and invasion by targeting lymphocyte antigen 6 complex locus K (LY6K) in human non-small cell lung cancer. Neoplasma, 65(5), 673–682.
Kong, H. K., Yoon, S., & Park, J. H. (2012). The regulatory mechanism of the LY6K gene expression in human breast cancer cells. The Journal of Biological Chemistry, 287(46), 38889–38900.
Endo, S., et al. (2016). TEX101, a glycoprotein essential for sperm fertility, is required for stable expression of Ly6k on testicular germ cells. Scientific Reports, 6, 23616.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Upadhyay, G. (2019). Emerging Role of Novel Biomarkers of Ly6 Gene Family in Pan Cancer. In: Rhim, J., Dritschilo, A., Kremer, R. (eds) Human Cell Transformation. Advances in Experimental Medicine and Biology, vol 1164. Springer, Cham. https://doi.org/10.1007/978-3-030-22254-3_4
Download citation
DOI: https://doi.org/10.1007/978-3-030-22254-3_4
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-22253-6
Online ISBN: 978-3-030-22254-3
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)