Abstract
We have characterized two highly tumorigenic and metastatic basal B TNBC cell lines, XtMCF and LmMCF, with the additional values of having the normal and early-stage counterparts of them. This model allows the study of the evolution of TNBC, and investigates molecular pathways at different stages of transformation and progression in a relatively constant genetic background. This constitutes an ideal model for developing targeted therapy in two important fields in cancer biology which are the epithelial mesenchymal transition (EMT) and cancer stem cells (CSC).
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- 17ß-estradiol
- Human breast epithelial cells
- Epithelial to mesenchymal transition
- Triple-negative breast cancer
- Chromatin remodeling
- Breast epithelial cell transformation
17ß-Estradiol Induces Transformation and Tumorigenesis in Human Breast Epithelial Cells
Breast cancer is a malignancy whose dependence on estrogen exposure has long been recognized, even though the mechanisms through which estrogens cause cancer are not clearly understood [1,2,3,4,5,6,7,8,9,10,11,12,13,14,15,16,17]. Our work was performed in order to determine whether 17ß-estradiol (E2), the predominant circulating ovarian steroid , is carcinogenic in human breast epithelial cells and whether non-receptor mechanisms are involved in the initiation of breast cancer. For this purpose, the effect of four alternating 24 h treatment periods with 70 nM E2 of the estrogen receptor alpha (ER-α) negative MCF-10F cell line on the in vitro expression of neoplastic transformation was evaluated [7, 8, 11] (Fig. 1).
Schematic representation of the experimental protocol. MCF-10F cells were treated with 70 nM 17ß-estradiol (17ß-E2 70 nM) for 24-h periods, twice a week during 2 weeks. After the last treatment the cells were passaged 7–9 times before being tested for colony efficiency, ductulogenesis in collagen, invasion in Matrigel and tumorigenesis in SCID mice. For the invasion assay, the cells were trypsinized and seeded in the invasion chambers at a concentration of 2.5 × 104 cells/well, incubated for 22 h and then the membranes of the inserts were cut and invasive cells (bsMCF) were cultured in 24-well plates. The invasive cells were expanded and evaluated for the expression of tumorigenesis in SCID mice. Nine out of ten mice injected with bsMCF developed tumors. Tumors from four of the animals were dissected in 0.5–1 mm size fragments, incubated in culture medium until confluent, generating cell lines from each tumor that were subsequently injected to five mice per cell line for the evaluation of their tumorigenic potential. All injected animals developed tumors. All tumors and cell lines were analyzed histopathologically and immunocytochemically as well as for fingerprint and CGH analyses. None of the animals injected with MCF-10F control cells or trMCF developed tumors
E2-treatment induced the expression of anchorage-independent growth, loss of ductulogenesis in collagen, invasiveness in Matrigel, and loss of 9p11-13. Tumorigenesis in SCID mice was expressed only in invasive cells that in addition exhibited a deletion of 4p15.3-16. Tumors formed in SCID mice were poorly differentiated adenocarcinomas that were estrogen receptor α and progesterone receptor negative, expressed keratins, EMA, and e-cadherin. The relationship between cell motility in vitro and the ability of neoplastic cells to invade and metastasize in vivo is well known. This led us to evaluate the migratory behavior of the human breast epithelial cell line MCF-10F after neoplastic transformation with 70 nM of 17ß-estradiol (E2), 4 OH estradiol (4-OH-E2) and 2-OH estradiol (2-OH-E2). Cells thus transformed express colony formation in agar methocel, loss of ductulogenic capacity in collagen matrix, and invasiveness in a Matrigel artificial membrane [7, 8, 11]. We set up a time-lapse video microscopy system to directly observe and capture the cells’ images using a Nikon DXM digital camera attached to an Olympus IMT-2 microscope that was equipped with a Plexiglas incubation chamber. From each cell line, a random number of cells were selected for tracking at 1-hour intervals. Cell motility was evaluated by determining the speed of the tracked cell expressed in mm/min (S), the direction persistence in time (P), and the random motility coefficient (μ) that provides a measure of how fast a cell population will grow to cover a surface. Our findings [1] indicated that the transformation of HBEC by estrogen and its metabolites induces changes in cell motility in vitro, 4-OH-E2 being the one inducing the most significant changes, its effect correlated to the expression of phenotypes indicative of cell transformation .
Epithelial to Mesenchymal Transition in Human Breast Epithelial Cells Transformed by 17-Beta-Estradiol
We have demonstrated that 17ß-estradiol (E2) induces complete neoplastic transformation of the human breast epithelial cells MCF-10F [7, 8, 11]. E2-treatment of MCF-10F cells progressively induced high colony efficiency and loss of ductulogenesis in early transformed (trMCF) cells, invasiveness in a Matrigel invasion chambers. The cells that crossed the chamber membrane were collected and identified as bsMCF, and their subclones designated bcMCF, and the cells harvested from carcinoma formation in SCID mice designated (caMCF) (Fig. 2) [11]. These phenotypes correlated with gene dysregulation during the progression of the transformation. The highest number of dysregulated genes was observed in caMCF cells, being slightly lower in bcMCF cells, and lowest in trMCF cells. This order was consistent with the extent of chromosome aberrations (caMCF > bcMCF >>> trMCF). Chromosomal amplifications were found in 1p36.12-pter, 5q21.1-qter, and 13q21.31-qter. Losses of the complete chromosome 4 and of 8p11.21-23.1 were found only in tumorigenic cells. In tumor-derived cell lines, additional losses were found in 3p12.1-14.1, 9p22.1-pter, and 18q11.21-qter [11].
Transformation of MCF-10F cells by 17ß-estradiol treatment. Experimental protocol: MCF-10F cells treated with 70 nM 17ß-estradiol (E2) that expressed high colony efficiency (CE) and loss of ductulogenic capacity in collagen matrix were classified as transformed (trMCF). Transformed cells that were invasive in a Matrigel Boyden-type invasion chambers were selected (bsMCF) and plated at low density for cloning (bcMCF). MCF-10F, trMCF, bsMCF, and bcMCF were tested for carcinogenicity by injecting them into the mammary fat pad of 45-day-old female SCID mice. MCF-10F and trMCF cells did not induce tumors (canceled arrow); bsMCF and bcMCF formed solid tumors from which four cell lines, identified as caMCF, were derived and proven to be tumorigenic in SCID mice (reprinted from [11])
Functional profiling of deregulated genes revealed progressive changes in the integrin signaling pathway, inhibition of apoptosis, acquisition of tumorigenic cell surface markers, and epithelial to mesenchymal transition. In tumorigenic cells, the levels of E-cadherin, EMA, and various keratins were low and CD44E/CD24 were negative, whereas SNAI2, vimentin, S100A4, FN1, HRAS and TGFβ1, and CD44H were high (Fig. 3) [11].
(a) A list of EMT markers and promoting genes was generated a priory by literature search. Hierarchical clustering of cell lines and genes was performed using dChip software. Two sample clusters (κ and λ) and two gene clusters (α and β) were identified. The red, white, and blue colors represent level above, at, and below mean expression, respectively. (b) Detection of epithelial and mesenchymal markers by immunocytochemistry: (a) Histological sections of MCF-10F cells, reacted with pre-immune mouse serum, were used as the negative control (×100); (b) MCF-10F reacted for EMA (×100); (c) MCF-10F reacted for E-Cadherin (×100); (d) MCF-10F reacted for vimentin (×100); (e) trMCF cells reacted with pre-immune mouse serum used as negative control (×100); (f, g, and h) trMCF cells reacted for EMA, E-cadherin, and vimentin, respectively (×100); (i) bsMCF cells reacted with pre-immune mouse serum as a negative control (×100); (j, k, l) bsMCF cells reacted for EMA, E-cadherin, and vimentin, respectively (×100); (m) caMCF tumor cell line cells reacted with pre-immune mouse serum used as negative control (×100); (n, o, p) caMCF tumor cell lines reacted for EMA, E-cadherin, and vimentin, respectively (×100); (q and r) invasive ductal carcinoma of the breast as positive control and immunoreacted for EMA and E-cadherin, respectively (×100); (s) histological section of an invasive adenocarcinoma immunoreacted for vimentin (×100) (Reprinted from: [11])
The phenotypic and genomic changes triggered by estrogen exposure that lead normal cells to tumorigenesis confirm the role of this steroid hormone in cancer initiation. Our work emphasizes the importance of being able to make a normal cell-like MCF-10F neoplastically transformed by a treatment with a natural hormone. More importantly the cell is estrogen receptor negative, indicating that the traditional pathway of action for estrogen and its receptors is not the main pathway of the neoplastic process. It is known that prolonged exposure to estrogen is a risk factor for human breast cancer, but the role of estrogen in the development of human breast cancer has been difficult to ascertain. There are three mechanisms that have been considered responsible for the carcinogenicity of estrogens: a receptor-mediated hormonal activity, cytochrome P450-mediated metabolic activation, and induction of aneuploidy. The receptor-mediated hormonal activity of estrogen has generally been related to stimulation of cellular proliferation, resulting in more opportunities for accumulation of genetic damages leading to carcinogenesis. Since local synthesis of estrogen in the stromal component can increase the estrogen levels and growth rate of breast carcinoma, a paracrine mechanism is likely to account for interactions between aromatase-containing stromal cells and ER-containing breast tumor epithelial cells. More importantly, estrogen may not need to activate nuclear receptors alpha to initiate or promote breast carcinogenesis. We have evidence that ERP may also be involved in this process and that oxidative catabolism of estrogens mediated by various CYP complexes constitutes a pathway of their metabolic activation and generates reactive free radicals and intermediate metabolites reactive intermediates that can cause oxidative stress and genomic damage directly. Estrogen-induced genotoxic effects include increased mutation rates, MSI, and LOH in chromosomes 3 and 11. Compromised DNA repair system allows accumulation of genomic lesions essential to estrogen-induced tumorigenesis. Metabolic biotransformation of estrogen does occur in human mammary explant culture. Increased formation of catechol estrogens as a result of elevated hydroxylation of 17β-estradiol at C-4 and C-16a positions has been observed in human breast cancer patients and in women at a higher risk of developing this disease. There is also evidence that formation of superoxide and hydrogen peroxide, as a result of the metabolism of estrogen, might also be involved in estrogen-mediated oxidative stress. In fact, a substantial increase in base lesions observed in the DNA of invasive ductal carcinoma of the breast has been postulated to result from the oxidative stress associated with metabolism of 17β-estradiol. Altogether the data thus far accumulated indicate that more than one pathway may be necessary to initiate neoplastic transformation and maintaining of the transformation phenotypes leading to tumorigenesis.
Developing a Unique Model of Triple Negative Breast Cancer
Triple-negative breast cancer (TNBC) represents a heterogeneous group of cancers characterized by a lack of ER, PgR, and HER2 expression. Cluster analysis of human TNBC identified six subtypes displaying unique gene expression and ontologies [18]. Approximately 80% of TNBC show features of basal-like cancers [19]. Transcriptional profile analysis assigned 21 TNBC cell lines into three clusters: luminal, basal A, and basal B [20,21,22]. Basal A contains cell lines such as BT-20, Sum149, and MDA-MB-468, which preferentially express genes such as CK5/6, CK14, and EGFR. Basal B includes cell lines such as MDA-MB-231, Sum159pt, and Hs578t, which preferentially express genes such as CD44, VIM, and SNAI2, and exhibits a stem-cell-like profile [20]. This classification of TNBC cell lines is closely associated with cell morphology and invasive potential. Basal B cells have a more mesenchymal-like appearance and are less differentiated and much more invasive compared to the other two clusters. Analysis of the relationship between TNBC cell lines and tumor subtypes showed that most of basal A and basal B cell lines resemble basal-like tumors [20], indicating that TNBC cell lines are suitable for investigations of subtype-specific cancer cell biology.
Although there are over 20 commercially available TNBC cell lines, MDA-MB-231 is the most widely used in vitro and in vivo. In BALB/CAJCI-nu/nu mice, it took 5 weeks to form a xenograft around 6.5 mm in diameter with the subcutaneous injection of 5 × 106 MDA-MB-231 cells [23]. MDA-MB-468 cells had a growth speed similar to MDA-MB-231 in the same mouse strain [23].
The growth speed of MDA-MB-231 xenograft in CB17/SCID was almost the same as in nude mice, while BT-549 cells grew a little bit slower than MDA-MB-231 cells in CB17/SCID mice [24]. Sum149 and Sum159 are two highly tumorigenic cell lines; it was reported that the injection of 1 × 105 cells in nonobese diabetic SCID mice could produce tumors in 3/4 and 5/6 mice, respectively [25]. But these two cell lines are mainly used for the study of inflammatory breast cancer [26, 27].
We have established [28] a progressive TNBC model (Fig. 4) consisting of normal MCF-10F, transformed cell line trMCF, and tumorigenic cell lines bsMCF, XtMCF, and LmMCF. Compared to the other nine tumorigenic TNBC cell lines, our cell lines XtMCF and LmMCF are the most tumorigenic and metastatic.
The expression of cytokeratin 18 (CK18) confirmed the epithelial origin of this cell model, and we observed that CK18 was down-regulated in bsMCF cell line and its derivatives. Furthermore, CK18 was lost in the lung metastases, whereas still present in the xenografts of both XtMCF and LmMCF cells, suggesting down-regulation of CK18 may be related to breast tumor progression [29]. Our study also showed that CK5-positive cell number was inversely correlated to clinical stage of TNBC [30, 31], suggesting that our cell model reflects features of TNBC progression.
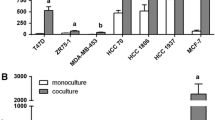
The EMT process is not only closely related to cancer invasion and metastasis but also conferred to the generation of cancer stem cells (CSC) [32,33,34]. As bsMCF-luc, XtMCF, and LmMCF have undergone EMT, we evaluated their CSC properties and the results showed that they could form tumorspheres, and the number of tumorspheres was progressively increasing from bsMCF-luc to XtMCF and LmMCF cells, consistent with in vivo tumorigenic and metastatic potential.
In this work [30], we postulated that the evaluation of CSC markers would give us a rationale for the high tumorigenic and metastatic potential of these two cell lines (Figs. 5 and 6). Our results showed that the bsMCF-luc and XtMCF cells were CD24low/CD44+, whereas LmMCF cells were CD44+ with moderate CD24 expression. CD24−/low/CD44+ has been frequently used as CSC markers of breast cancers [35,36,37]. However, it was shown that the percentage of CD24−/low/CD44+ associates with a basal-like phenotype, not tumorigenicity, but CD24−/low/CD44+/EpCAM+ cells enrich for tumorigenicity [31, 36]. EpCAM induces expressions of reprogramming factors and EMT genes, regulates EMT progression, and tumorigenesis [38]. In addition, EpCAM can be cleaved at several sites, and the nuclear translocation of cytoplasmic domain (EpCID) associates with Wnt pathway and promotes cell proliferation and tumor formation in mice [39]. One of the EpCAM cleavage sites between two arginine residues (AA80 and AA81) was detected and described in the late 1980s, but the functional consequence is still unknown [40]. Interestingly, we observed the expression of EpCAM in the cell lines we examined by immunofluorescence staining and WB, but the EGF-like domain of EpCAM was absent in mesenchymal-like cells, suggesting the EGF-like domain might be cleaved off from the cleavage site between AA80 and AA81. This was supported by other workers [41,42,43,44,45,46,47]. The majority of commercial antibodies for EpCAM react with overlapped or partly overlapped epitope at EGF-like domain [48]. This may result in failing detection of EpCAM in cells which have undergone EMT . Our study indicates that the EGF-like-domain-cleaved-off EpCAM may be associated with the EMT process. Furthermore, although the total level of EpCAM is low in mesenchymal-like cells, the subcellular localization of EpCAM may be more important to the EpCAM nuclear indicating a strong activation of Wnt signaling in these cells.
XtMCF and LmMCF cells display high tumorigenic and metastatic potential. Representative pictures of xenografts and lungs fixed with Bouin’s solution. Magnification: 6.3× for xenografts, 8× for lungs (Reprinted from: [28])
H&E staining of lungs from the injection of 1 × 106 cells into tail vein. LmMCF cells are more metastatic than XtMCF cells. Arrows indicate the metastases. Magnifications are shown in figure (Reprinted from: [28])
Chromatin Remodeling During Human Breast Epithelial Cell Transformation
We have shown that treatment of the human breast epithelial cells MCF-10F with 17β-estradiol (E2) induces transformation and tumorigenesis. DNA amounts and chromatin supraorganization change in E2-transformed MCF cells [49]. Feulgen-DNA content and chromatin supraorganization were involved during E2-induced transformation and tumorigenesis of the MCF-10F cells. Image analysis was performed for non-transformed and E2-transformed MCF cells, highly invasive cells (C5), and for cell lines (C5-A6-T6 and C5-A8-T8) derived from tumors generated by injection of C5 cells in SCID mice (Fig. 1). A decrease in Feulgen-DNA amounts and nuclear sizes induced by E2 treatment was accented with selection of the highly invasive tumorigenesis potential. However, in the tumor-derived cells, a high variability in cellular phenotypes resulted inclusive in near-polyploidy. Significant changes in textural parameters, including nuclear entropy, indicated chromatin structural remodeling with advancing tumorigenesis. An increased variability in the degree of chromatin packing states in the E2-transformed MCF cells was followed by reduction in chromatin condensation and in contrast between condensed and non-condensed chromatin in the highly invasive C5 cells and tumor-derived cell lines. These observations confirmed previous data [11], showing the role of chromatin remodeling and epigenetic control in the transformation of human breast epithelial cells. We found that a network of several signaling pathways affecting the expression and/or function of a complex hierarchical network of transcription factors (TFs) has been partially elaborated. Known signaling pathways include multiple tyrosine kinase receptors leading to Ras-mediated activation of MAPK and PI3K pathways, TGF-β, Notch, and Wnt. Evidence for enhanced TGF-β and Wnt signaling pathways was found in the EMT expressing bcMCF and caMCF cells . TGF-β acting through Smad transcriptional complexes can repress expression of the Id TFs (Id1, Id2, Id3) and activate HMGA2, a DNA binding protein important for chromatin architecture. Expression of HMGA2 is known to regulate several EMT controlling TFs including TWIST1, SNAI1, and SNAI2 (Slug) (Fig. 7). TGF-β and Wnt signaling also affect the expression of several additional EMT-regulating TFs including ZEB1 (TCF8), TCF3 (E2A encoding E12 and E47), and LEF1. Analysis of the EMT expressing bcMCF cell line revealed the absence of expression of the secreted frizzled-related protein 1 (SFRP1) , a repressor of Wnt signaling. One allele of SFRP1 was deleted in these cells, with the remaining apparently silenced by methylation, accounting for the 28-fold reduction of this transcript. Loss and epigenetic inactivation of SFRP1 occurs often in invasive breast cancer and is associated with poor prognosis. Inspection of the SFRP1 expression levels in Basal B cell lines showed absent calls for four of the eight invasive cell lines, and eightfold decreases in another three invasive cell lines relative to the non-invasive MCF-10A cells (data not shown). Inspection of the expression files for bcMCF cells and the eight invasive Basal B cell lines revealed that LEF1 was always absent, while TCF 3 and TCF 8 were expressed.
Heat map showing the EMT (Reprinted from: [11])
As we have described above the TNBC cell consisting of normal like breast epithelial cell line MCF10F, the trMCF cell line that was transformed from MCF10F cells, the tumorigenic bsMCF cell line derived from trMCF, and two highly tumorigenic and metastatic cell lines XtMCF and LmMCF established from bsMCF cells have undergone epithelial mesenchymal transition (EMT), exhibiting a mesenchymal-like feature, providing a good cell model for identifying new treatments for TNBC .
Concluding Remarks
The relevance of this work is the development and characterization of two highly tumorigenic and metastatic basal B TNBC cell lines, XtMCF and LmMCF. To the best of our knowledge, they are the most tumorigenic and metastatic TNBC cell lines compared to all reported cell models used for TNBC studies. In addition, the normal and early-stage counterparts of these two cell lines are also available. Altogether, these cell lines can be used to study the evolution of TNBC, investigate molecular pathways at different stages of transformation and progression in a relatively constant genetic background, and most importantly, identify new treatments for TNBC. In addition, XtMCF and LmMCF cell lines present CSC properties and can be used for developing CSC-targeted therapy. The finding that the EGF-like domain of EpCAM is cleaved off in cancer cells which have undergone EMT also provides new insights in the research of EMT and CSC, two important fields in cancer biology.
References
Russo, J., Lareef, M. H., Balogh, G., Guo, S., & Russo, I. H. (2003). Estrogen and its metabolites are carcinogenic in human breast epithelial cells. The Journal of Steroid Biochemistry and Molecular Biology, 87, 1–25.
Russo, J., & Russo, I. H. (2004). Genotoxicity of steroidal estrogens. Trends in Endocrinology and Metabolism, 15, 211–214.
Fernandez, S. V., Russo, I. H., Lareef, M. H., Balsara, B., & Russo, J. (2005). Comparative genomic hybridization of human breast epithelial cells transformed by estrogen and its metabolites. International Journal of Oncology, 26(3), 691–695.
Chen, J.-Q., Yager, J. D., & Russo, J. (2005). Regulation of mitochondrial respiratory chain structure and function by estrogens/estrogen receptors and potential physiological/pathophysiological implications: Review. Biochimica et Biophysica Acta, 1746, 1–17.
Fernandez, S. V., Lareef, M. H., Russo, I. H., Balsara, B. R., Testa, J. R., & Russo, J. (2006). Estrogen and its metabolites 4-Hydroxy-estradiol induce mutations in TP53 and LOH in chromosome 13q12.3 near BRCA2 in human breast epithelial cells. International Journal of Cancer, 118(8), 1862–1868.
Cavalieri, E., Chakravarti, D., Guttenplan, J., Hart, E., Ingle, J., Jankowiak, R., Muti, P., Rogan, E., Russo, J., Santen, R., & Sutter, T. (2006). Catechol estrogen quinones as initiators of breast and other human cancers. Implications for biomarkers of susceptibility and cancer prevention. Review. Biochimica et Biophysica Acta, 1766, 63–78.
Russo, J., Fernandez, S. V., Russo, P. A., Fernbaugh, R., Sheriff, F. S., Lareef, H. M., Garber, J., & Russo, I. H. (2006). 17 beta estradiol induces transformation and tumorigenesis in human breast epithelial cells. The FASEB Journal, 20, 1622–1634.
Russo, J., & Russo, I. H. (2006). The role of estrogen in the initiation of breast cancer. The Journal of Steroid Biochemistry and Molecular Biology, 102, 89–96.
Mello, M. L., Vidal, B. C., Lareef, M. H., Russo, I. H., & Russo, J. (2007). DNA content and estradiolbchromatin texture of human breast epithelial cells treated with 17- and the estrogen antagonist ICI 182,780 as assessed by image analysis. Mutation Research, 617, 1–7.
Tiezzi, D. G., Fernandez, S. V., & Russo, J. (2007). Epithelial to mesenchymal transition during breast cancer progression. International Journal of Oncology, 31, 823–827.
Huang, Y., Fernandez, S., Goodwin, S., Russo, P. A., Russo, I. H., Sutter, T., & Russo, J. (2007). Epithelial to mesenchymal transition in human breast epithelial cells transformed by 17-beta-estradiol. Cancer Research, 67, 11147–11157.
Harvey, J. A., Santen, R. J., Petroni, G. R., Bovbjerg, V., Smolkin, M. A., Sheriff, F., & Russo, J. (2008). Histology changes in the breast with menopausal hormone therapy use: Correlation with breast density, ER, PgR, and proliferation indices. Menopause, 15(1), 67–73.
Russo, J., & Russo, I. H. (2007). Estradiol. In M. Schwab (Ed.), Encyclopedia of Cancer (2nd ed.). Heidelberg: Springer.
Chen, J.-Q., Russo, P. A., Cooke, C., Russo, I. H., & Russo, J. (2007). ERβ shifts from the mitochondria to the nucleus during 17β estradiol induced neoplastic transformation of human breast epithelial cells and is involved in E2 induced synthesis of mitochondrial chain proteins. Biochimica et Biophysica Acta Molecular Cell Research, 1773, 1732–1746.
Chen, J.-Q., & Russo, J. (2008). Mitochondrial estrogen receptors and their potential implications in estrogen carcinogenesis in human breast cancer. Journal of Nutritional and Environmental Medicine, 17, 76–89.
Mello, M. L., Russo, P. A., Russo, J., Vidal, B. C., & Benedicto, C. (2007). 17-β-estradiol affects nuclear image properties in MCF-10F human breast epithelial cells with tumorigenesis. Oncology Report, 18, 1475–1481.
Saeed, M., Rogan, E., Fernandez, S. V., Sheriff, F., Russo, J., & Cavalieri, E. (2007). Formation of depurinating N3Adenine and N7Guanine adducts by MCF-10F cells cultured in the presence of 4-hydroxyestradiol. International Journal of Cancer, 120, 1821–1824.
Lehmann, B. D., Bauer, J. A., Chen, X., Sanders, M. E., Chakravarthy, A. B., Shyr, Y., et al. (2011). Identification of human triple-negative breast cancer subtypes and preclinical models for selection of targeted therapies. Journal of Clinical Investigation, 121, 2750–2767.
Tan, D. S., Marchió, C., Jones, R. L., Savage, K., Smith, I. E., Dowsett, M., et al. (2008). Triple negative breast cancer: Molecular profiling and prognostic impact in adjuvant anthracycline-treated patients. Breast Cancer Research and Treatment, 111, 27–44.
Kao, J., Salari, K., Bocanegra, M., Choi, Y.-L., Girard, L., Gandhi, J., et al. (2009). Molecular profiling of breast cancer cell lines defines relevant tumor models and provides a resource for cancer gene discovery. PLoS One, 4, e6146.
Neve, R. M., Chin, K., Fridlyand, J., Yeh, J., Baehner, F. L., Fevr, T., et al. (2006). A collection of breast cancer cell lines for the study of functionally distinct cancer subtypes. Cancer Cell, 10, 515–527.
Grigoriadis, A., Mackay, A., Noel, E., Wu, P. J., Natrajan, R., Frankum, J., et al. (2012). Molecular characterisation of cell line models for triple-negative breast cancers. BMC Genomics, 13, 619.
Yunokawa, M., Koizumi, F., Kitamura, Y., Katanasaka, Y., Okamoto, N., Kodaira, M., et al. (2012). Efficacy of everolimus, a novel mTOR inhibitor, against basal-like triple-negative breast cancer cells. Cancer Science, 103, 1665–1671.
Tate, C. R., Rhodes, L. V., Segar, H. C., Driver, J. L., Pounder, F. N., Burow, M. E., et al. (2012). Targeting triple-negative breast cancer cells with the histone deacetylase inhibitor panobinostat. Breast Cancer Research, 14, R79.
Fillmore, C. M., & Kuperwasser, C. (2008). Human breast cancer cell lines contain stem-like cells that self-renew, give rise to phenotypically diverse progeny and survive chemotherapy. Breast Cancer Research, 10, R25.
Flanagan, L., Van Weelden, K., Ammerman, C., Ethier, S. P., & Welsh, J. (1999). SUM-159PT cells: A novel estrogen independent human breast cancer model system. Breast Cancer Research and Treatment, 58, 193–204.
Zhang, D., LaFortune, T. A., Krishnamurthy, S., Esteva, F. J., Cristofanilli, M., Liu, P., et al. (2009). Epidermal growth factor receptor tyrosine kinase inhibitor reverses mesenchymal to epithelial phenotype and inhibits metastasis in inflammatory breast cancer. Clinical Cancer Research, 15, 6639–6648.
Su, Y., Pogash, T. J., Nguyen, T. D., & Russo, J. (2016). Development and characterization of two human triple-negative breast cancer cell lines with highly tumorigenic and metastatic capabilities. Cancer Medicine, 5, 558–573.
Woelfle, U., Sauter, G., Santjer, S., Brakenhoff, R., & Pantel, K. (2004). Down-regulated expression of cytokeratin 18 promotes progression of human breast cancer. Clinical Cancer Research, 10, 2670–2674.
Su, Y., Gutiérrez-Diez, P. J., Santucci-Pereira, J., Russo, I. H., & Russo, J. (2014). In situ methods for identifying the stem cell of the normal and cancerous breast. In J. Russo & I. H. Russo (Eds.), Techniques and methodological approaches in breast cancer research (1st ed., pp. 151–182). New York: Springer.
Aguiar, F. N., Mendes, H. N., Cirqueira, C. S., Bacchi, C. E., & Carvalho, F. M. (2013). Basal cytokeratin as a potential marker of low risk of invasion in ductal carcinoma in situ. Clinics, 68, 638–643.
Morel, A. P., Lièvre, M., Thomas, C., Hinkal, G., Ansieau, S., & Puisieux, A. (2008). Generation of breast cancer stem cells through epithelial-mesenchymal transition. PLoS One, 3, e28882008.
Mani, S. A., Guo, W., Liao, M.-J., Eaton, E. N., Ayyanan, A., Zhou, A. Y., et al. (2008). The epithelial-mesenchymal transition generates cells with properties of stem cells. Cell, 133, 704–715.
Xue, C., Plieth, D., Venkov, C., Xu, C., & Neilson, E. G. (2003). The gatekeeper effect of epithelial-mesenchymal transition regulates the frequency of breast cancer metastasis. Cancer Research, 63, 3386–3394.
Al-Hajj, M., Wicha, M. S., Benito-Hernandez, A., Morrison, S. J., & Clarke, M. F. (2003). Prospective identification of tumorigenic breast cancer cells. Proceedings of the National Academy of Sciences, 100, 3983–3988.
Sheridan, C., Kishimoto, H., Fuchs, R. K., Mehrotra, S., Bhat-Nakshatri, P., Turner, C. H., et al. (2006). CD44+/CD24-breast cancer cells exhibit enhanced invasive properties: An early step necessary for metastasis. Breast Cancer Research, 8, R59.
Wright, M. H., Calcagno, A. M., Salcido, C. D., Carlson, M. D., Ambudkar, S. V., & Varticovski, L. (2008). Brca1 breast tumors contain distinct CD44+/CD24-and CD133+ cells with cancer stem cell characteristics. Breast Cancer Research, 10, R10.
Lin, C.-W., Liao, M.-Y., Lin, W.-W., Wang, Y.-P., Lu, T.-Y., & Wu, H.-C. (2012). Epithelial cell adhesion molecule regulates tumor initiation and tumorigenesis via activating reprogramming factors and epithelial-mesenchymal transition gene expression in colon cancer. The Journal of Biological Chemistry, 287, 39449–39459.
Maetzel, D., Denzel, S., Mack, B., Canis, M., Went, P., Benk, M., et al. (2009). Nuclear signalling by tumour-associated antigen EpCAM. Nature Cell Biology, 11, 162–171.
Thampoe, I. J., Ng, J. S., & Lloyd, K. O. (1988). Biochemical analysis of a human epithelial surface antigen: Differential cell expression and processing. Archives of Biochemistry and Biophysics, 267, 342–352.
Keller, P. J., Lin, A. F., Arendt, L. M., Klebba, I., Jones, A. D., Rudnick, J. A., et al. (2010). Mapping the cellular and molecular heterogeneity of normal and malignant breast tissues and cultured cell lines. Breast Cancer Research, 12, R87.
Gorges, T. M., Tinhofer, I., Drosch, M., Rose, L., Zollner, T. M., Krahn, T., et al. (2012). Circulating tumour cells escape from EpCAM-based detection due to epithelial-to-mesenchymal transition. BMC Cancer, 12, 178.
Mego, M., De Giorgi, U., Dawood, S., Wang, X., Valero, V., Andreopoulou, E., et al. (2011). Characterization of metastatic breast cancer patients with nondetectable circulating tumor cells. International Journal of Cancer, 129, 417–423.
Sieuwerts, A. M., Kraan, J., Bolt, J., van der Spoel, P., Elstrodt, F., Schutte, M., et al. (2009). Anti-epithelial cell adhesion molecule antibodies and the detection of circulating normal-like breast tumor cells. Journal of the National Cancer Institute, 101, 61–66.
Hayes, D. F. C. M. (2009). Anti-epithelial cell adhesion molecule antibodies and the detection of circulating normal-like breast tumor cells. Journal of the National Cancer Institute, 101, 894–895.
Van Laere, S. J., Elst, H., Peeters, D., Benoy, I., Vermeulen, P. B., & Dirix, L. Y. (2009). Re: Anti-epithelial cell adhesion molecule antibodies and the detection of circulating normal-like breast tumor cells. Journal of the National Cancer Institute, 101, 895–896.
Connelly, M., Wang, Y., Doyle, G. V., Terstappen, L., & McCormack, R. (2009). Re: Anti-epithelial cell adhesion molecule antibodies and the detection of circulating normal-like breast tumor cells. Journal of the National Cancer Institute, 101, 895.
Balzar, M., Briaire-de Bruijn, I., Rees-Bakker, H., Prins, F., Helfrich, W., De Leij, L., et al. (2001). Epidermal growth factor-like repeats mediate lateral and reciprocal interactions of Ep-CAM molecules in homophilic adhesions. Molecular and Cellular Biology, 21, 2570–2580.
Mello, M. L., Russo, P. A., Russo, J., & Vidal, B. C. (2007). 17-ß-estradiol affects nuclear image properties in MCF-10Fhuman breast epithelial cells with tumorigenesis. Oncology Reports, 18, 1475–1481.
Acknowledgements
This work was supported by the Pennsylvania Cancer Cure Grant 6914101, the NIH core grant CA06927 to Fox Chase Cancer Center, the Barbara and Joseph Breitman donation, and the Flyers wives donation. The compound SGI-110 was provided by Astex Pharmaceuticals, Inc., Dublin CA.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Russo, J., Su, Y. (2019). An In Vitro Model of Triple-Negative Breast Cancer. In: Rhim, J., Dritschilo, A., Kremer, R. (eds) Human Cell Transformation. Advances in Experimental Medicine and Biology, vol 1164. Springer, Cham. https://doi.org/10.1007/978-3-030-22254-3_3
Download citation
DOI: https://doi.org/10.1007/978-3-030-22254-3_3
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-22253-6
Online ISBN: 978-3-030-22254-3
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)