Abstract
The white-fly borne begomoviruses (family Geminiviridae) have circular single-stranded (css) DNA genome, which is encapsidated as monopartite (DNA-A) or bipartite (DNA-A and DNA-B) in the twinned icosahedrons. During the course of their evolution and to escape host defense machinery, begomoviruses adopt small cssDNA satellites called alpha-, beta-, and deltasatellites. Alphasatellties are found to be associated with begomovirus–betasatellite complexes and encode their own replication-associated protein (Rep), thus capable of autonomous replication. These satellite-like molecules are not well known to serve any critical function for their helper begomovirus except for few reports about attenuation of helper-virus accumulation and/or occasionally suppression of the host defense. Most of the monopartite begomoviruses in the Old World (OW) are found to be associated with betasatellites; however, none of the New World (NW) begomoviruses are known to be associated with betasatellites. Begomoviruses replicate their genome through rolling circle replication (RCR), which requires the virus-encoded Rep to recognize and bind to the iterated sequences (iterons) in the origin of replication (ori) region. Betasatellites lack such iterated sequences; however, they can be transreplicated by a diverse range of begomoviruses, following a similar pattern for replication. Betasatellites play a significant role in viral pathogenesis by interacting with certain host factors, attenuation of disease symptoms, suppression of host defense, and sometimes inter- or intracellular shuttling of begomovirus genome. Likewise, the noncoding molecules deltasatellites depend upon their helper virus for their replication. However, their precise role in viral pathogenesis still needs to be explored.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
1 Introduction
Many plant viruses coexist with certain nucleic acid, either DNA or RNA molecules, termed as “satellites.” Satellites lack the ability for an independent existence and are entirely dependent on their helper virus for their replication, encapsidation, movement, and proliferation (Briddon and Stanley 2006). Satellite molecules have a very simple genomic organization and usually encode few or no genes and have little or no sequence homology to their helper virus. The word “satellite” coins two major classes of nucleic acid agents: satellite viruses, which are capable of self-encapsidation by producing capsid protein, and the virus-associated satellites, which lack their own capsid protein and therefore utilize helper-virus proteins for their encapsidation (Mayo et al. 2005). Satellite virus was described for the first time in 1962 when scientists discovered some exogenous nucleic acid agents being associated with few strains of Tobacco necrosis virus (TNV), a Necrovirus, laterally recognized as Tobacco necrosis satellite virus (TNSV) (Kassanis 1962). Until now, several satellite molecules have been found to be associated with different classes of plant viruses, particularly RNA viruses such as Rice yellow mottle virus satellite (RYMV-sat) and Cucumber mosaic virus satellites (CMV-sat) (Adams et al. 2017; Mayo et al. 2005). Majority of the satellites have single-stranded (ss) RNA genome; however, double-stranded (ds) RNA genome is also present in few members. RNA genome of few ssRNA satellites encodes some proteins that may or may not assist in replication process (Palukaitis et al. 2008). The first begomovirus-associated DNA satellite [Tomato leaf curl satellite (ToLCV-sat)] was reported in 1997 from tomato plants infected with Tomato leaf curl virus (ToLCV) (Dry et al. 1997). In most cases, these satellites overload the resources of helper virus for their own replication/survival and interfere with viral infectivity (Brown et al. 2012). However, few members of DNA satellites have been identified causing coinfections with helper viruses and result in severe disease symptoms as compared to single viral infection (Nawaz-ul-Rehman and Fauquet 2009).
2 History and Current Status of ssDNA Satellites Associated with Begomoviruses
Most of the begomoviruses (family: Geminiviridae) in the Old World (OW) and few in the New World (NW) are being associated with circular ssDNA satellites (Fig. 1). Till now, DNA satellites associated with majority of the OW monopartite begomoviruses include more frequently found betasatellites and occasionally alphasatellites, whereas newly characterized deltasatellites are most frequently reported in association with the NW begomoviruses (Adams et al. 2017) (Table 1). Recently, betasatellites and deltasatellites have been assigned new genera Betasatellite and Deltasatellite in the sub-viral family, Tolecusatellitidae, respectively (Adams et al. 2017).
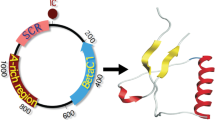
Genome organization of cssDNA satellites. Schematic position and orientation of genes is shown with colored arrows. (a) Alphasatellites encode Rep protein and contain an adenine rich (A-rich) sequence. (b) Betasatellites encode single protein, βC1, have a satellite conserved region (SCR) and an A-rich sequence. The purple and green parts of deltasatellite SCR represent the predicted conserved stem-loop and secondary stem-loop structures, respectively. For all satellites, the intergenic region (IR) contains a predicted hairpin-loop structure, which contains the nonanucleotide sequence (TAA/GTATTAC) as part of the loop
2.1 Alphasatellites
After their first discovery in 1999 from Ageratum conyzoides (Saunders and Stanley 1999), approximately 200 complete alphasatellite (earlier described as DNA-1) sequences have been deposited in the GenBank database. Alphasatellites are circular and ssDNA (css) molecules associated with OW monopartite begomoviruses, begomovirus–betasatellite complexes, and/or NW bipartite begomoviruses (Paprotka et al. 2010). Alphasatellites do not truly represent satellite molecules because of their self-encoded replication-associated protein (Rep) and autonomous replication ability (Briddon et al. 2004) and could survive in permissive hosts (Mansoor et al. 1999). However, for encapsidation, transmission by the insect vector, and in planta movement, they are reliant on helper viruses.
The genome of alphasatellites comprised of ~1380 nucleotides (nt) that encode a single open reading frame (ORF): Rep (36 kDa) subsiding in virion-sense strand (coding strand), a highly conserved A-rich genomic sequence (~200 nt), and an origin of replication (Ori) containing a conserved nonanucleotide sequence (TAGTATT/AC) present in a predicted hairpin structure (Fig. 1a) (Briddon et al. 2004). The nonanucleotide sequence and the Rep-encoding segments of alphasatellite genome resemble the nanoviruses (another family of ssDNA viruses) (Brown et al. 2012), which suggests their possible capture by a begomovirus during mixed infections (Briddon and Stanley 2006). It is presumed that the captured Rep-encoded component (~1000 nt) of nanoviruses was reorganized through embedding A-rich sequences to gain a ~1400 nt size (half the size of begomovirus, i.e., ~2800 nt) in order to encode a structurally stabilized Rep required for self-encapsidation (Briddon and Stanley 2006; Mansoor et al. 2003). Three different subclasses of alphasatellites, DNA-1-type, DNA-2-type, and DNA-3-type, are frequently reported. The most commonly occurring alphasatellites are DNA-1-type that predominantly occur in the Indian subcontinent (Paprotka et al. 2010). The DNA-2-type alphasatellites are far rare, found in Singapore and Oman having low nt sequence identity with DNA-1-type, while DNA-3-type are novel alphasatellites detected from Guatemala, Brazil, and Puerto Rico (Rosario et al. 2016). According to Rosario et al. (2016), the DNA-3-type alphasatellites share 51–55% nt sequence identity with the DNA-1-type. Moreover, they help to increase the symptom severity in the host plants and form a separate monophyletic group when analyzed through phylogenetic studies (Rosario et al. 2016).
Although predominantly alphasatellites are found to be associated with begomoviruses, quite recently, an alphasatellite has been found associated with a mastrevirus, Wheat dwarf India virus (WDIV), in a natural field infection, which shows that alphasatellites have fewer constraints for their helper virus, host plant, or the insect vector. Their enigmatic role in virus pathogenesis has not been clearly answered yet. However, in few studies, the Rep proteins of alphasatellites have been described as the post-transcriptional gene silencing (PTGS) suppressors (Nawaz-ul-Rehman et al. 2010). Alphasatellites still need extensive explorations in plant virology as there are no consolidated reports available that describe their precise function and association mechanism with begomoviruses.
2.2 Betasatellites
Betasatellites (formerly described as DNAβ) are cssDNA-satellite molecules, which have recently been classified into the sub-viral family Tolecusatellitidae genus Betasatellite (Adams et al. 2017) (Fig. 1b). Betasatellites are predominantly found to be associated with monopartite begomoviruses in the OW; however, since the last few years, these molecules have also been found in association with bipartite begomoviruses (Hameed et al. 2017; Jyothsna et al. 2013) and recently with a mastrevirus (Kumar et al. 2014). Unlike alphasatellites, betasatellites are true satellite molecules as they entirely depend on the helper virus for their encapsidation, replication, and systemic dispersal (Briddon and Stanley 2006). Betasatellites variably interact with their helper-virus component and result in multiple types of coinfections. In some cases, the betasatellites synergistically infect the host plants by increasing their helper-virus accumulation and are essential for symptom induction (Chandel et al. 2016). For example, the interactions of Cotton leaf curl Multan virus (CLCuMuV) with Cotton leaf curl Multan betasatellite (CLCuMuB) and Ageratum yellow vein virus (AYVV) with Ageratum yellow vein betasatellite (AYVB) result in severe disease symptoms and enhanced virus titer as compared to their helper begomovirus alone in cotton and A. conyzoides, respectively (Briddon et al. 2001; Saunders et al. 2000). In other cases, a facultative interaction has also been observed in begomovirus:betasatellite complex, where the begomovirus component could infect alone and does not necessarily require betasatellite for symptom induction and/or enhanced viral titer (Chandel et al. 2016). For example, Tobacco curly shoot virus (TbCSV):(TbCSB) could make coinfections but TbCSV could infect alone; however, the presence of betasatellite induces more severe symptoms (Li et al. 2005). Another promising role played by betasatellite is the substitution of DNA-B component. Coinoculation of CLCuMuB and Tomato leaf curl New Delhi virus (ToLCNDV) DNA-A induced leaf curl disease phenotype in the model plant Nicotiana benthamiana in the absence of DNA-B component (Saeed et al. 2007). Since their first description in 2000, genome sequences of more than 1000 betasatellite isolates have been submitted to GenBank, depicting their ongoing diversity and evolution (Adams et al. 2017) (Fig. 2).
The betasatellite genome (~1350 nt) exhibits three conserved features: a complementary-sense single βC1 gene (Briddon et al. 2003), a highly conserved A-rich region (~150-200 nt), and a ~100 nt satellite conserved region (SCR) (Nawaz-ul-Rehman and Fauquet 2009). The betasatellite genome shares no sequence homology with their cognate viruses except for a similar nonanucleotide sequence (TAATATTAC) present in the SCR (Briddon et al. 2003). Betasatellite-encoded βC1 (13-14 kDa) is a multifunctional protein involved in pathogenesis, enhancing viral DNA accumulation in the nucleus and suppression of the host antiviral defense response (Saunders et al. 2004). The other important ability of βC1 is self-interaction and localization at cell periphery, thus presumably having a role in viral movement (Cheng et al. 2011). Additionally, βC1 interacts with numerous host factors like ubiquitin-conjugating enzymes (UBC) (Eini et al. 2009) and asymmetric leaves 1 (AS1) factor, etc. (Yang et al. 2008).
The studies on geographical occurrence and diversity of betasatellites showed that the major center of their diversity lies in the Indian subcontinent and Southeast Asia (Fig. 2). On the basis of phylogeny, betasatellites may be broadly categorized into two major groups: the first group constitutes all the betasatellites isolated from the plant family Malvaceae (hibiscus, cotton, okra, hollyhock, etc.) while the betasatellites belonging to the second group were isolated from non-malvaceous plants (tomato, chillies, ageratum, zinnia, etc.). These findings are suggestive of the important role of host plants in the evolution of betasatellites.
2.3 Deltasatellites
Recently, another class of cssDNA satellites (approx. quarter the size of helper begomoviruses) has been classified as a new genus “Deltasatellite” (family Tolecusatellitidae) (Fiallo-Olivé et al. 2012; Lozano et al. 2016). Deltasatellites have been further categorized into three types of noncoding DNA satellites, i.e., Tomato leaf curl virus–satellite (ToLCV-sat) identified from Australia (Dry et al. 1997), DNA satellites associated with sweepoviruses in Venezuela and Spain (Lozano et al. 2016 ), and those isolated from malvaceous hosts in the Caribbean (Fiallo-Olivé et al. 2012). Besides their structural resemblance (Fig. 1b), phylogenetically deltasatellites are not closely related to each other; however, they entirely depend upon the helper virus for their vital functions. Their genome contains a stem-loop structure with a nonanucleotide (TAATATT/AC) sequence, an A-rich region, and does not encode any putative ORF. These satellites have a second putative stem-loop structure situated close to the iteron-like sequences, and a short region that resembles the SCR of the betasatellites. Contrary to the betasatellites, the emergence of deltasatellites in the NW might be due to agricultural trades of infected plants, like sweet potato, from OW to the NW (Lozano et al. 2016).
To date, the precise function of deltasatellites in context to begomovirus-deltasatellite complex is unclear, although some studies reported the influence of deltasatellite in lowering helper-virus accumulation in host plant that might facilitate the sequential movement of viruses to other plant parts (Fiallo-Olivé et al. 2016; Hassan et al. 2016).
3 Replication Mechanism of DNA Satellites
In the course of successful infection after entering the host plant cell, begomoviral ssDNA genomes along with the genome of the associated DNA satellite(s) access the cell nuclei for replication. The replication of begomovirus genome is achieved through dsDNA intermediates either by a rolling circle replication (RCR) and/or recombination-dependent replication (RDR) mechanism (Hanley-Bowdoin et al. 2013) (Fig. 3). The dsDNA intermediates are transcribed by host RNA-polymerase II to translate the first viral protein Rep, which then initiates RCR of both the begomovirus and the satellite DNA (Hanley-Bowdoin et al. 2013). These circular dsDNAs are assembled into transcriptionally active viral mini-chromosomes with the help of host histone proteins (Pilartz and Jeske 2003). The viral Rep protein creates a conducive environment to commence the replication. The Rep protein of begomoviruses drives the initiation and termination of RCR by nicking the dsDNA intermediates and rejoining the circular DNA at the specific site in the nonanucleotide sequences (TAATATT/AC) (Hanley-Bowdoin et al. 2013; Laufs et al. 1995). The successful commencement of RCR specifically requires high-affinity interactions between the begomovirus Rep protein and ori in the intergenic region (IR). The synthesis of complementary strand is initiated with a nick by Rep protein in the nonanucleotide sequence of the ssDNA, and an RNA primer is synthesized by the host DNA primase to initiate this phenomenon. The host plant replisome machinery (DNA polymerase and associated factors) is hijacked and reprogrammed during the elongation step to accomplish the viral dsDNA synthesis (Bagewadi et al. 2004; Kaliappan et al. 2011). The newly synthesized strand is displaced and released by Rep as a circular ssDNA. The synthesized dsDNA is used as a template to start next replication cycle. At the end of optimum replication cycles, the Rep protein downregulates its own synthesis and ultimately activates the expression of TrAP, which leads to the production of CP to start the virus and DNA satellites assembly (Fig. 3).
Pictorial model of replication mechanism of begomoviruses/satellites. Once the virion DNA is delivered to the nucleus, host machinery initiates the synthesis of the complementary strand. Host-derived polymerases convert the single-stranded virion DNA into a double-stranded intermediate, which performs as a transcriptionally active mini-chromosome. This mini-chromosome mediates the cell-to-cell movement and nuclear trafficking. Virion derived replication-associated protein (Rep) then binds to the iterons, produces a site-specific nick in the origin of replication (ori), and becomes covalently linked to the 5′ end of the nicked DNA via a tyrosine residue. The 3’OH end acts as a primer for synthesis of new virion-sense DNA by host-encoded factors, using the complementary-sense as a template. The nicking-joining activity of Rep releases unit length virion-sense ssDNA molecules. The newly synthesized ssDNA either continues the replication cycle (acting as a template for complementary-strand synthesis), is moved from cell-to-cell (possibly as virions), or is packaged by the coat protein (CP) for onward transmission by insect vectors. Image was reproduced from Briddon and Stanley (2006)
The DNA satellites employ a similar mechanism of DNA replication as their helper begomovirus (Alberter et al. 2005). However, beta- and deltasatellites are devoid of Rep protein (unlike begomovirus and alphasatellites) and hence depend exclusively upon the helper virus to commence their replication (Zhou 2013). Alphasatellites are capable of autonomous replication mechanism through RCR directed by their own protein, alpha-Rep. As betasatellites are devoid of self-replication, the replication strategy of the betasatellites is determined by the helper begomovirus (Alberter et al. 2005). The replication model for begomoviruses suggests the imperative binding of Rep protein to the iterative sequences upstream of the nonanucleotide sequences followed by the recognition of the ori. Apparently, betasatellites frequently lack the iteron sequences, which is suggestive of some other mechanism involved in the ori recognition of betasatellites (Leke et al. 2012). The nicking site for Rep in betasatellites is expected to present in the nonanucleotide stem-loop sequences adjacent to the SCR. The stem sequences and the adjacent hairpin structures of betasatellites are remarkably similar in all betasatellites, and thus, it reaffirms that they participate in the ori recognition by helper Rep (Zhou 2013). The position of the highly conserved SCR present immediately upstream of the stem-loop sequences is also analogous to the relative position of IR of the helper begomoviruses. Furthermore, the conservation of SCRs in the defective forms of betasatellites further supports that the SCR region is involved in the replication of the betasatellites (Zhang et al. 2016). However, the sequence between the downstream of SCR and βC1 is required for efficient replication of betasatellites (Eini and Behjatnia 2016). The betasatellites lack iteron-like Rep-binding motifs (RBM); thus, the presence of G-box motif (CACGTG) may serve the binding of Rep protein (Eini and Behjatnia 2016). Thus, the high-affinity binding of Rep has a critical role in betasatellite replication. However, the exact mechanism of Rep binding is needed to be explored yet.
4 Transreplication and Pseudo-Recombination of DNA Satellites
As discussed earlier, the only sequence homology between the betasatellites and their helper begomoviruses is the presence of stem-loop structure (Briddon and Stanley 2006). In contrast to DNA-B of bipartite begomoviruses, betasatellites do not necessarily require cognate DNA-A of a bipartite or monopartite begomovirus genome. Instead, they are transreplicated by a diverse range of non-cognate begomoviruses for their transreplication. For example, Cotton leaf curl Gezira betasatellite (CLCuGeB) is known to be a cognate associate of Okra yellow crinkle virus (OYCrV) and Cotton leaf curl Gezira virus (CLCuGeV) causing okra leaf curl disease (Leke et al. 2013). However, it can also be transreplicated with three other distinct begomoviruses Tomato yellow leaf curl virus (TYLCV), Tomato leaf curl Mali virus (ToLCMLV), and Tomato yellow leaf crumple virus (TYLCrV), each from a diverse geographic origin (Saunders 2008). Similarly, CLCuMuB is a cognate member of CLCuD-associated begomoviruses (CABs) in Asia (Iqbal et al. 2012; Sattar et al. 2017). The non-cognate associations between CLCuMuB with ToLCNDV and TbCSB with Clerodendrum golden mosaic China virus (CGMCV) are significant examples of its transreplication by a bipartite begomovirus (Li and Zhou 2010; Saeed 2010). Moreover, CLCuMuB can also be transreplicated by other non-cognate viruses such as TYLCV, Tomato leaf curl Karnatka virus (ToLCKnV), and Tomato leaf curl virus (ToLCV) (Kharazmi et al. 2012). Apart from begomoviruses, betasatellites are also known to be transreplicated by the members of other genera of the family Geminiviridae. Beet curly top virus (BCTV; genus Becurtovirus) successfully transreplicates Ageratum yellow vein betasatellite (AYVB) and Tomato yellow leaf curl China betasatellite (TYLCCNB) (Yang et al. 2011a). In another study, a Curtovirus, Beet severe curly top virus (BSCTV), successfully supported the transreplication of CLCuMuB (Kharazmi et al. 2012). Likewise, association of Cotton leaf curl Multan alphasatelliite (CLCuMuA), Guar leaf curl alphasatellite (GLCuA), and Ageratum yellow leaf curl betasatellite (AYLCB) with the WDIV (genus Mastrevirus) highlights the natural transreplication of DNA satellites by the member of a different genus (Kumar et al. 2014). Such associations are quite surprising because the functional betasatellites are mostly known to be associated with monopartite begomoviruses during natural infections. Moreover, the DNA-B component of bipartite begomoviruses has specific interactions with the Rep of the cognate DNA-A only. Such indistinguishable replication of betasatellites depicts that these molecules are quite flexible for their transreplication as compared to the specificity of recognition between Rep protein and DNA-B component of a bipartite begomovirus.
Apart from the fact that betasatellites have quite a promiscuous mode of replication, two distinct betasatellite species rarely coexist with a single helper virus within the same host plant. Apparently, this is because betasatellites are adapted to their cognate helper virus for replication during the course of evolution (Zhou et al. 2003). Thus, the cognate betasatellites are shown to accumulate to higher levels than the non-cognate betasatellites within the same host (Qing and Zhou 2009). For example, the coinoculation of TYLCCNB and TbCSB with one helper virus creates a competition, which causes cognate betasatellite dominant over non-cognate betasatellite. However, switching their sequence elements also switched the preferential replication of the respective cognate helper virus (Zhang et al. 2016).
Under natural environmental conditions, although betasatllites may coexist with the alphasatellites, the binding of alpha-Rep with the Rep protein of the helper virus may obstruct betasatellite replication. The alphasatellites can ameliorate begomovirus symptoms and hinder high accumulation of betasatellites during coinfections (Idris et al. 2011).
The deltasatellites contain a stem-loop with nonanucleotide, TATA box, and a second predicted stem-loop with iteron-like sequences. Moreover, their A-rich region and a short region also share high homology with betasatellite SCRs (Lozano et al. 2016). However, further investigations are needed to decipher their mode of replication and roles in viral pathogenesis. Most probably, these molecules are also transreplicated by the helper begomovirus Rep due to the presence of begomovirus iteron-like sequences upstream of the second stem loop (Fiallo-Olivé et al. 2016). The deltasatellite, Tomato leaf curl virus-satellite (ToLCV-sat), has been shown to be transreplicated by ToLCV as well as geographically distinct geminiviruses like Tomato yellow leaf curl Sardinia virus (TYLCSV), African cassava mosaic virus (ACMV), and a becurtovirus BCTV. In another study, the deltasatellites sat-177 and sat-603 could only be transreplicated by the cognate begomoviruses, Sida golden yellow vein virus (SiGYVV), and a monopartite begomovirus Tomato leaf deformation virus (ToLDV) from the NW. However, the OW TYLCV, TYLCSV, and ACMV could not support their transreplication (Fiallo-Olivé et al. 2016).
5 Deciphering the Role of DNA Satellites in the Begomovirus Pathogenesis
All the functions of betasatellites are accredited to their single gene product βC1. This protein, when expressed transiently through Potato virus X (PVX) or through a stable transformation in model host plants (N. benthamiana and N. tabacum) cells, induces typical begomovirus disease symptoms of leaf curling, vein thickening, and enations (Kon et al. 2007; Qazi et al. 2007). The βC1 protein regulates the expression of several different miRNAs involved in the developmental processes when expressed through PVX in N. benthamiana plants. The accumulation of miR159 and miR160 was significantly enhanced, while the accumulation of miR164, miR165/166, miR169, and miR170 was reduced when the βC1 gene was transiently expressed in the inoculated plants (Amin et al. 2011). The βC1 accumulates primarily in the nucleus, localizes at the periphery of the infected host cells, and colocalizes along the endoplasmic reticulum. These localization patterns and presence of both nuclear import and export signals point toward the putative role of βC1 in intracellular transport and movement (Cheng et al. 2011). Moreover, βC1 forms punctate bodies, both in vivo and in vitro, by self-interaction, which presumably has a role in symptom induction. A deletion mutagenesis study shows that amino acids spanning two α-helices at C-terminal are important in self-interaction.
The βC1 interacts with a variety of host-encoded factors such as with AS1 and AS2 factors. Self-interaction of these two factors is required for leaf development; βC1 mimics the function of AS2 by interacting with AS1 and thus affects the leaf development (Yang et al. 2008). The CLCuMuB-encoded βC1 interacts with a UBC to induce betasatellite-specific symptoms in the host plant (Eini et al. 2009). It is speculated that βC1 interaction with UBC perturbs the ubiquitin-proteasome pathway to enhance βC1 accumulation, which ultimately led to the development of viral symptoms.
Plants have developed a fine-tuned defense mechanism, which is operated through PTGS and TGS, against invading pathogens. To counter the host defense response, βC1 has the ability to suppress the PTGS-mediated host defense by interacting with one of the important host defense components, Argonaute-1 (AGO-1), which binds to the siRNAs and represses the target RNAs (Eini 2017). The βC1 protein can bind to ss- as well as dsDNA, dsRNA, and both long and short RNAs in a sequence-independent manner to suppress the host defense. This binding activity is mediated by the nuclear localization signals (NLS) present in the βC1. TYLCCNB-encoded βC1 has the ability to suppress PTGS by upregulating the N. benthamiana calmodulin-related protein (Nbrgs-CaM), which can repress the expression of N. benthamiana RNA-dependent RNA polymerase 6 (Zhou 2013). Besides PTGS, the βC1 also has the ability to reduce the TGS or can even reverse the established TGS (Yang et al. 2011b) in the host plants. This suppression of TGS is mediated by interacting with S-adenosyl homocysteine hydrolase (SAHH), an enzyme generally required for the generation of S-adenosyl methionine (SAM), through a NLS (49KKK51) present in βC1 (Yang et al. 2011b). The CLCuMuB-βC1 can suppress the host defense by downregulating the jasmonic acid (JA)-responsive genes such as CORI3, PR4, NbPHAN, and PDF1 (Yang et al. 2008) and can interact with certain host-encoded factors involved in metabolic and defense pathways (Tiwari et al. 2013). The expression of βC1 can differentially regulate the genes involved in electron carrier for photosynthesis, respiration, and ATP synthesis (Andleeb et al. 2010). The CLCuMuB-βC1 also interacts with ATG8 protein, a ubiquitin-like protein having a role in the biogenesis of autophagosomes (Shelly et al. 2009). It is thus speculated that the interaction of βC1 with ATG8 may likely be an antiviral defense mechanism. Besides, βC1 can interact with tomato UBC, an enzyme required for ubiquitination and ultimately the degradation of the target protein (Eini et al. 2009). This interaction interferes with ubiquitin-proteasome pathway that could enhance the βC1 accumulation.
To counter the βC1 pathogenesis, host plants have developed a sophisticated counterattack mechanism. Tomato plants employed Sucrosenonfermenting1-related kinase (SISnRK1)-mediated defense against the betasatellites. Hyperexpression of SISnRK1 leads to the reduction in betasatellite accumulation and delayed onset of the symptoms. It has been showed that SISnRK1 phosphorylates TYLCCNB-βC1 at the amino acid positions 33 (serine residue) and 78 (threonine), thus negatively regulating the βC1 functions (Cui et al. 2004; Yang et al. 2008).
6 Role of Rep-A of Alphasatellite in Viral Pathogenesis
To date, the interactions of the alphasatellite-encoded Rep protein with the host-encoded factors and its role in successful begomovirus infection have not been fully explored. Only a few studies are available, which reported that the Rep protein encoded by few alphasatellites have PTGS suppressor activity (Nawaz-ul-Rehman et al. 2010), suggesting the role of alphasatellite in overcoming RNAi-mediated host defense. The type-2 alphasatellites are known to have a role in the symptom attenuation by reducing the accumulation of begomovirus (Nawaz-ul-Rehman et al. 2010) and/or betasatellites (Idris et al. 2011; Wu and Zhou 2005) in the begomovirus–betasatellite complexes. This attenuation in symptoms may likely increase the chance of host survival and virus transmission.
7 Conclusion
Host–pathogen interactions are like arms race with typical zero-sum game, which ultimately leads to the disease development or the host recovery. In this subtle type of intimate relationship, both counterparts continuously deploy different strategies to take advantage over each other. The acquisition of DNA satellites by begomoviruses is the continuity of this process. DNA satellites have equipped their helper begomoviruses to suppress the host defense (both TGS and PTGS) and/or help in the symptom attenuation, which ultimately helps the virus to evade the host defense. During the acquisition process, begomoviruses resized these DNA satellites precisely in a mathematical way, alpha- and betasatellites are almost half, while deltasatellites are one-fourth of the helper-virus genome, to support their replication.
The maintenance of these DNA satellites by the helper-virus replication machinery is dependent upon dynamic, mainly undefined, interactions between begomovirus, DNA satellite, and host-encoded factors. However, ori is the only common feature between DNA satellites and the helper viruses, so interaction between the geminivirus Rep and DNA satellites is principally dependent on this region. Likewise, ori region (particularly nonanucleotide) determines the successful commencement of RCR. The importance of ori in replication has been proven experimentally where switching of the ori sequence has switched the preferential transreplication of betasatellite by helper begomovirus (Zhang et al. 2016). Although, no strong selection mechanism is present between DNA satellites and their helper virus, the interaction between DNA satellites and their helper virus is not merely a transreplication but stacking of a multilayer interaction (Iqbal et al. 2012, 2017).
References
Adams MJ, Lefkowitz EJ, King AMQ, Harrach B et al (2017) Changes to taxonomy and the international code of virus classification and nomenclature ratified by the International Committee on Taxonomy of Viruses. Arch Virol 162:2505–2538
Akhtar S, Khan AJ, Singh AS, Briddon RW (2014) Identification of a disease complex involving a novel monopartite begomovirus with beta-and alphasatellites associated with okra leaf curl disease in Oman. Arch Virol 159:1199–1205
Alberter B, Rezaian MA, Jeske H (2005) Replicative intermediates of Tomato leaf curl virus and its satellite DNAs. Virology 331:441–448
Amin I, Patil BL, Briddon RW, Mansoor S et al (2011) A common set of developmental miRNAs are upregulated in Nicotiana benthamiana by diverse begomoviruses. Virol J 8:143
Andleeb S, Amin I, Bashir A, Briddon RW et al (2010) Transient expression of βC1 protein differentially regulates host genes related to stress response, chloroplast and mitochondrial functions. Virol J 7:373
Bagewadi B, Chen S, Lal SK, Choudhury NR et al (2004) PCNA interacts with Indian mung bean yellow mosaic virus rep and downregulates rep activity. J Virol 78:11890–11903
Briddon RW, Stanley J (2006) Sub-viral agents associated with plant single-stranded DNA viruses. Virology 344:198–210
Briddon RW, Mansoor S, Bedford ID, Pinner MS et al (2001) Identification of DNA components required for induction of cotton leaf curl disease. Virology 285:234–243
Briddon RW, Bull SE, Amin I, Idris AM et al (2003) Diversity of DNA β, a satellite molecule associated with some monopartite begomoviruses. Virology 312:106–121
Briddon RW, Bull SE, Amin I, Mansoor S et al (2004) Diversity of DNA 1; a satellite-like molecule associated with monopartite begomovirus-DNA β complexes. Virology 324:462–474
Brown JK, Fauquet CM, Briddon RW, Zerbini M et al (2012) Geminiviridae. Virus taxonomy – Ninth report of the International Committee on Taxonomy of Viruses. Associated Press, Elsevier Inc., London, Waltham, San Diego, pp 351–373
Bull SE, Tsai W-S, Briddon RW, Markham PG et al (2004) Diversity of begomovirus DNA β satellites of non-malvaceous plants in east and south East Asia. Arch Virol 149:1193–1200
Chandel V, Singh MK, Jangid A, Dhatwalia S (2016) Emerging satellites associated with begomoviruses: world scenario. In: Gaur RK, Petrov NM, Patil BL, Stoyanova MI (eds) Plant viruses: evolution and management. Springer, Singapore, pp 145–169
Cheng X, Wang X, Wu J, Briddon RW et al (2011) βC1 encoded by tomato yellow leaf curl China betasatellite forms multimeric complexes in vitro and in vivo. Virology 409:156–162
Cui XF, Tao XR, Xie Y, Fauquet CM et al (2004) A DNAβ associated with tomato yellow leaf curl China virus is required for symptom induction. J Virol 78:13966–13974
Dong JH, Luo YQ, Ding M, Zhang ZK et al (2007) First report of tomato yellow leaf curl China virus infecting kidney bean in China. Plant Pathol 56:342
Dry I, Krake LR, Rigden JE, Rezaian MA (1997) A novel subviral agent associated with a geminivirus: the first report of a DNA satellite. Proc Natl Acad Sci USA 94:7088–7093
Eini O (2017) A betasatellite-encoded protein regulates key components of gene silencing system in plants. Mol Biol 51:579–585
Eini O, Behjatnia SAA (2016) The minimal sequence essential for replication and movement of Cotton leaf curl multan betasatellite DNA by a helper virus in plant cells. Virus Genes 52:679–687
Eini O, Dogra S, Selth LA, Dry IB et al (2009) Interaction with a host ubiquitin-conjugating enzyme is required for the pathogenicity of a geminiviral DNA b satellite. Mol Plant-Microbe Interact 22:737–746
Fauquet CM, Sawyer S, Idris AM, Brown JK (2005) Sequence analysis and classification of apparent recombinant begomoviruses infecting tomato in the Nile and Mediterranean basins. Phytopathology 95:549–555
Fiallo-Olivé E, Martínez-Zubiaur Y, Moriones E, Navas-Castillo J (2012) A novel class of DNA satellites associated with New World begomoviruses. Virology 426:1–6
Fiallo-Olivé E, Hamed A, Navas-Castillo J, Moriones E (2013) Cotton leaf curl Gezira alphasatellite associated with tomato leaf curl Sudan virus approaches the expected upper size limit in the evolution of alphasatellites. Virus Res 178:506–510
Fiallo-Olivé E, Tovar R, Navas-Castillo J (2016) Deciphering the biology of deltasatellites from the New World: maintenance by New World begomoviruses and whitefly transmission. New Phytol 212:680–692
Geetanjali SA, Shilpi S, Mandal B (2013) Natural association of two different betasatellites with sweet potato leaf curl virus in wild morning glory (Ipomoea purpurea) in India. Virus Genes 47:1–5
Guo XJ, Zhou XP (2006) Molecular characterization of a new begomovirus infecting Sida cordifolia and its associated satellite DNA molecules. Virus Genes 33:279–285
Ha C, Coombs S, Revill P, Harding R et al (2008a) Molecular characterization of begomoviruses and DNA satellites from Vietnam: additional evidence that the New World geminiviruses were present in the Old World prior to continental separation. J Gen Virol 89:312–326
Ha C, Revill P, Harding RM, Vu M et al (2008b) Identification and sequence analysis of potyviruses infecting crops in Vietnam. Arch Virol 153:45–60
Hameed A, Tahir MN, Amin I, Mansoor S (2017) First report of tomato leaf curl New Delhi virus and a tomato yellow leaf curl Thailand betasatellite causing severe leaf curl disease of potato in Pakistan. Plant Dis 101:1065
Hanley-Bowdoin L, Bejarano ER, Robertson D, Mansoor S (2013) Geminiviruses: masters at redirecting and reprogramming plant processes. Nat Rev Microbiol 11:777–788
Harimalala M, Bruyn A, Hoareau M, Andrianjaka A et al (2013) Molecular characterization of a new alphasatellite associated with a cassava mosaic geminivirus in Madagascar. Arch Virol 158(8):1–4
Hassan I, Orílio AF, Fiallo-Olivé E, Briddon RW et al (2016) Infectivity, effects on helper viruses and whitefly transmission of the deltasatellites associated with sweepoviruses (genus Begomovirus, family Geminiviridae). Sci Rep 6:30204
Huang JF, Zhou XP (2006) Molecular characterization of two distinct begomoviruses from Ageratum conyzoides and Malvastrum coromandelianum in China. J Phytopathol 154:648–653
Idris AM, Shahid MS, Briddon RW, Khan AJ et al (2011) An unusual alphasatellite associated with monopartite begomoviruses attenuates symptoms and reduces betasatellite accumulation. J Gen Virol 92:706–717
Iqbal Z, Sattar MN, Kvarnheden A, Mansoor S et al (2012) Effects of the mutation of selected genes of Cotton leaf curl Kokhran virus on infectivity, symptoms and the maintenance of Cotton leaf curl Multan betasatellite. Virus Res 169:107–116
Iqbal Z, Shafiq M, Ali I, Mansoor S et al (2017) Maintenance of Cotton leaf curl Multan betasatellite by tomato leaf curl New Delhi virus—analysis by mutation. Front Plant Sci 8:2208
Jose J, Usha R (2003) Bhendi yellow vein mosaic disease in India is caused by association of a DNA b satellite with a begomovirus. Virology 305:310–317
Jyothsna P, Haq QMI, Singh P, Sumiya KV et al (2013) Infection of tomato leaf curl New Delhi virus (ToLCNDV), a bipartite begomovirus with betasatellites, results in enhanced level of helper virus components and antagonistic interaction between DNA B and betasatellites. Appl Microbiol Biotechnol 97:5457–5471
Kaliappan K, Choudhury NR, Suyal G, Mukherjee SK (2011) A novel role for RAD54: this host protein modulates geminiviral DNA replication. FASEB J 26(3):1142–1160
Kassanis B (1962) Properties and behaviour of a virus depending for its multiplication on another. Microbiology 27:477–488
Kharazmi S, Behjatnia SAA, Hamzehzarghani H, Niazi A (2012) Cotton leaf curl Multan betasatellite as a plant gene delivery vector trans-activated by taxonomically diverse geminiviruses. Arch Virol 157:1269–1279
Kon T, Sharma P, Ikegami M (2007) Suppressor of RNA silencing encoded by the monopartite tomato leaf curl Java begomovirus. Arch Virol 152:1273–1282
Kumar J, Kumar A, Roy J, Tuli R et al (2010) Identification and molecular characterization of begomovirus and associated satellite DNA molecules infecting Cyamopsis tetragonoloba. Virus Genes 41:118–125
Kumar J, Singh S, Kumar A, Khan J et al (2013) Detection and characterization of a new betasatellite: variation in disease symptoms of tomato leaf curl Pakistan virus-India due to associated betasatellite. Arch Virol 158:257–261
Kumar J, Kumar J, Singh SP, Tuli R (2014) Association of satellites with a mastrevirus in natural infection: complexity of Wheat dwarf India virus disease. J Virol 88:7093–7104
Laufs J, Schumacher S, Geisler N, Jupin I et al (1995) Identification of the nicking tyrosine of geminivirus rep protein. FEBS Lett 377:258–262
Leke W, Kvarnheden A, Ngane E, Titanji V et al (2011) Molecular characterization of a new begomovirus and divergent alphasatellite from tomato in Cameroon. Arch Virol 156:925–928
Leke WN, Brown JK, Ligthart ME, Sattar N et al (2012) Ageratum conyzoides: a host to a unique begomovirus disease complex in Cameroon. Virus Res 163:229–237
Leke WN, Sattar MN, Ngane EB, Ngeve JM et al (2013) Molecular characterization of begomoviruses and DNA satellites associated with okra leaf curl disease in Cameroon. Virus Res 174:116–125
Li J, Zhou X (2010) Molecular characterization and experimental host-range of two begomoviruses infecting Clerodendrum cyrtophyllum in China. Virus Genes 41:1–10
Li ZH, Xie Y, Zhou XP (2005) Tobacco curly shoot virus DNA b is not necessary for infection but intensifies symptoms in a host-dependent manner. Phytopathology 95:902–908
Lozano G, Trenado HP, Fiallo-Olivé E, Chirinos D et al (2016) Characterization of non-coding DNA satellites associated with sweepoviruses (genus Begomovirus, Geminiviridae) – definition of a distinct class of begomovirus-associated satellites. Front Microbiol 7:162
Mansoor S, Khan SH, Bashir A, Saeed M et al (1999) Identification of a novel circular single-stranded DNA associated with cotton leaf curl disease in Pakistan. Virology 259:190–199
Mansoor S, Briddon RW, Zafar Y, Stanley J (2003) Geminivirus disease complexes: an emerging threat. Trends Plant Sci 8:128–134
Marwal A, Kumar Sahu A, Gaur RK (2013a) Molecular characterization of begomoviruses and DNA satellites associated with a new host Spanish flag (Lantana camara) in India. ISRN Virol 2013:5
Marwal A, Sahu A, Choudhary D, Gaur RK (2013b) Complete nucleotide sequence of a begomovirus associated with satellites molecules infecting a new host Tagetes patula in India. Virus Genes 47:1–5
Mayo MA, Leibowitz MJ, Palukaitis P, Scholthof K-BG et al (2005) Satellites. In: Fauquet CM, Mayo MA, Maniloff J, Desselberger U, Ball LA (eds) VIIIth report of the International Committee on Taxonomy of Viruses. Virus taxonomy. Elsevier/Academic Press, London, pp 1163–1169
Mubin M, Briddon RW, Mansoor S (2009) Complete nucleotide sequence of chili leaf curl virus and its associated satellites naturally infecting potato in Pakistan. Arch Virol 154:365–368
Mubin M, Shahid MS, Tahir MN, Briddon RW et al (2010) Characterization of begomovirus components from a weed suggests that begomoviruses may associate with multiple distinct DNA satellites. Virus Genes 40:452–457
Nawaz-ul-Rehman MS, Fauquet CM (2009) Evolution of geminiviruses and their satellites. FEBS Lett 583:1825–1832
Nawaz-ul-Rehman MS, Nahid N, Mansoor S, Briddon RW et al (2010) Post-transcriptional gene silencing suppressor activity of the alpha-rep of non-pathogenic alphasatellites associated with begomoviruses. Virology 405:300–308
Nehra C, Gaur RK (2014) Molecular characterization of Chilli leaf curl viruses infecting new host plant Petunia hybrida in India. Virus Genes 50:1–5
Ogawa T, Sharma P, Ikegami M (2008) The begomoviruses Honeysuckle yellow vein mosaic virus and tobacco leaf curl Japan virus with DNAb satellites cause yellow dwarf disease of tomato. Virus Res 137:235–244
Packialakshmi R, Srivastava N, Girish K, Usha R (2010) Molecular characterization of a distinct begomovirus species from Vernonia cinerea and its associated DNA-β using the bacteriophage Φ29 DNA polymerase. Virus Genes 41:135–143
Palukaitis P, Rezaian A, García-Arenal F (2008) Satellite nucleic acids and viruses. In: Mahy BWJ, van Regenmortel MHV (eds) Encyclopedia of virology. Academic Press, Oxford, pp 526–535
Paprotka T, Metzler V, Jeske H (2010) The first DNA 1-like a satellites in association with New World begomoviruses in natural infections. Virology 404:148–157
Pilartz M, Jeske H (2003) Mapping of abutilon mosaic geminivirus minichromosomes. J Virol 77:10808–10818
Qazi J, Amin I, Mansoor S, Iqbal J et al (2007) Contribution of the satellite encoded gene βC1 to cotton leaf curl disease symptoms. Virus Res 128:135–139
Qing L, Zhou X (2009) Trans-replication of, and competition between, DNA β satellites in plants inoculated with tomato yellow leaf curl China virus and tobacco curly shoot virus. Phytopathology 99:716–720
Rathore S, Bhatt B, Yadav B, Kale R et al (2014) A new begomovirus species in association with betasatellite causing tomato leaf curl disease in Gandhinagar, India. Plant Dis 98:428–428
Romay G, Lecoq H, Desbiez C (2014) Melon chlorotic mosaic virus and associated alphasatellite from Venezuela: genetic variation and sap transmission of a begomovirus-satellite complex. Plant Pathol 64(5):1224–1234
Rosario K, Marr C, Varsani A, Kraberger S et al (2016) Begomovirus-associated satellite DNA diversity captured through vector-enabled metagenomic (VEM) surveys using whiteflies (Aleyrodidae). Viruses 8:36
Saeed M (2010) Tomato leaf curl New Delhi virus DNA a component and Cotton leaf curl Multan betasatellite can cause mild transient symptoms in cotton. Acta Virol 54:317–318
Saeed M, Zafar Y, Randles JW, Rezaian MA (2007) A monopartite begomovirus-associated DNA b satellite substitutes for the DNA B of a bipartite begomovirus to permit systemic infection. J Gen Virol 88:2881–2889
Saeed ST, Khan A, Kumar B, Ajayakumar PV et al (2014) First report of Chilli leaf curl India virus infecting Mentha spicata (Neera) in India. Plant Dis 98:164–164
Sattar MN, Iqbal Z, Tahir MN, Ullah S (2017) The prediction of a new CLCuD epidemic in the Old World. Front Microbiol 8:631
Saunders K (2008) Analysis of geminivirus DNA replication by 2-D gel. In: Gary D, Foster IEJ, Hong Y, Nagy PD (eds) Plant virology protocols, pp 135–143
Saunders K, Stanley J (1999) A nanovirus-like component associated with yellow vein disease of Ageratum conyzoides: evidence for interfamilial recombination between plant DNA viruses. Virology 264:142–152
Saunders K, Bedford ID, Briddon RW, Markham PG et al (2000) A unique virus complex causes Ageratum yellow vein disease. Proc Natl Acad Sci USA 97:6890–6895
Saunders K, Bedford ID, Stanley J (2002) Adaptation from whitefly to leafhopper transmission of an autonomously-replicating nanovirus-like DNA component associated with ageratum yellow vein disease. J Gen Virol 83:909–915
Saunders K, Norman A, Gucciardo S, Stanley J (2004) The DNA β satellite component associated with ageratum yellow vein disease encodes an essential pathogenicity protein (βC1). Virology 324:37–47
Senanayake DMJB, Jayasinghe JEARM, Shilpi S, Wasala SK et al (2013) A new begomovirus–betasatellite complex is associated with chilli leaf curl disease in Sri Lanka. Virus Genes 46:128–139
Shahid M, Ikegami M, Waheed A, Briddon R et al (2014) Association of an alphasatellite with tomato yellow leaf curl virus and ageratum yellow vein virus in Japan is suggestive of a recent introduction. Viruses 6:189–200
Sharma P, Matsuda N, Bajet NB, Ikegami M (2011) Molecular analysis of new isolates of tomato leaf curl Philippines virus and an associated betasatellite occurring in the Philippines. Arch Virol 156:305–312
She X, He Z, Yin G, Du Z et al (2015) A new alphasatellite molecule associated with Ageratum yellow vein China virus in the Philippines. J Phytopathol 163:54–57
Shelly S, Lukinova N, Bambina S, Berman A et al (2009) Autophagy plays an essential anti-viral role in Drosophila against vesicular stomatitis virus. Immunity 30:588–598
Shih S, Kumar S, Tsai W, Lee L et al (2009) Complete nucleotide sequences of okra isolates of Cotton leaf curl Gezira virus and their associated DNA-β from Niger. Arch Virol 154:369–372
Singh M, Singh K, Haq Q, Mandal B et al (2011) Molecular characterization of tobacco leaf curl Pusa virus, a new monopartite Begomovirus associated with tobacco leaf curl disease in India. Virus Genes 43:1–11
Singh M, Haq QMR, Mandal B, Varma A (2012) Evidence of the association of radish leaf curl virus with tobacco yellow leaf curl disease in Bihar, India. Indian J Virol 23:64–69
Srivastava A, Raj S, Kumar S, Snehi S (2013) New record of Papaya leaf curl virus and Ageratum leaf curl beta satellite associated with yellow vein disease of aster in India. New Dis Rep 28(6). https://doi.org/10.5197/j.2044-0588.2013.028.006
Srivastava A, Jaidi M, Kumar S, Raj SK et al (2015) Association of Papaya leaf curl virus with the leaf curl disease of grain amaranth (Amaranthus cruentus L.) in India. Phytoparasitica 43:97–101
Tahir M, Haider MS, Briddon RW (2010) Chili leaf curl betasatellite is associated with a distinct recombinant begomovirus, pepper leaf curl Lahore virus, in Capsicum in Pakistan. Virus Res 149:109–114
Tahir M, Amin I, Haider S, Mansoor S et al (2015) Ageratum enation virus – a begomovirus of weeds with the potential to infect crops. Viruses 7:647–665
Tao X, Zhou X (2008) Pathogenicity of a naturally occurring recombinant DNA satellite associated with tomato yellow leaf curl China virus. J Gen Virol 89:306–311
Tiwari N, Padmalatha K, Singh V, Haq Q et al (2010) Tomato leaf curl Bangalore virus (ToLCBV): infectivity and enhanced pathogenicity with diverse betasatellites. Arch Virol 155:1343–1347
Tiwari N, Sharma PK, Malathi VG (2013) Functional characterization of βC1 gene of Cotton leaf curl Multan betasatellite. Virus Genes 46:111–119
Wu P-J, Zhou X-P (2005) Interaction between a nanovirus-like component and the Tobacco curly shoot virus/satellite complex. Acta Biochim Biophys Sin 37:25–31
Xie Y, Wu P, Tao X, Zhou X (2004) Identification of a nanovirus-like DNA molecule associated with Tobacco curly shoot virus isolates containing satellite DNA. Prog Nat Sci 14:689–693
Xiong Q, Guo XJ, Che HY, Zhou XP (2005) Molecular characterization of a distinct begomovirus species and its associated satellite DNA molecule infecting Sida acuta. J Phytopathol 153:264–268
Yang J-Y, Iwasaki M, Machida C, Machida Y et al (2008) βC1, the pathogenicity factor of TYLCCNV, interacts with AS1 to alter leaf development and suppress selective jasmonic acid responses. Genes Dev 22:2564–2577
Yang X, Guo W, Ma X, An Q et al (2011a) Molecular characterization of Tomato leaf curl China virus, infecting tomato plants in China, and functional analyses of its associated betasatellite. Appl Environ Microbiol 77:3092–3101
Yang X, Xie Y, Raja P, Li S et al (2011b) Suppression of methylation-mediated transcriptional gene silencing by βC1-SAHH protein interaction during geminivirus-betasatellite infection. PLoS Pathog 7:e1002329
Zhang T, Xu X, Huang C, Qian Y et al (2016) A novel DNA motif contributes to selective replication of a geminivirus-associated betasatellite by a helper virus-encoded replication-related protein. J Virol 90:2077–2089
Zhou X (2013) Advances in understanding begomovirus satellites. Annu Rev Phytopathol 51:357–381
Zhou X, Xie Y, Tao X, Zhang Z et al (2003) Characterization of DNA b associated with begomoviruses in China and evidence for co-evolution with their cognate viral DNA-A. J Gen Virol 84:237–247
Zulfiqar A, Zhang J, Cui X, Qian Y et al (2012) A new begomovirus associated with alpha- and betasatellite molecules isolated from Vernonia cinerea in China. Arch Virol 157:189–191
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Sattar, M.N., Iqbal, Z., Hameed, A. (2019). Replication of DNA Satellites and Their Role in Viral Pathogenesis. In: Kumar, R. (eds) Geminiviruses. Springer, Cham. https://doi.org/10.1007/978-3-030-18248-9_9
Download citation
DOI: https://doi.org/10.1007/978-3-030-18248-9_9
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-18247-2
Online ISBN: 978-3-030-18248-9
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)