Abstract
Drug Repositioning is the use of existing drugs to treat new diseases. Drug molecules exert their actions by binding to specific 3D biological molecules. Working and reasoning with 3D structures is complex, thus researchers prefer working with 1D or text data. Furthermore, drug-repositioning studies often use data sets from various independent sources, which make data processing and analysis time consuming due to different file formats, missing data, and complex cross-referencing. Here, we integrate 12 publicly available data sets on various biological/chemical entities like disease, gene, protein, pathway, drug, and side effect and 5 ontologies to provide an abstraction paradigm. The resulting integrated repository, which we called Sirius (for shedding light on drug-disease relationships) contains 7,321 disease related phenotypes, 47,063 protein functions, 2,226 drugs functions and 72,787 drug side effects having 12, 11, 9 and 4 abstraction levels, respectively. We illustrate the usefulness of our repository by studying the relationships between drugs and diseases, using side effect and pathway data. Our study predicted 117 associations, of which 93 are confirmed by the CTD database. The database is available on request from the authors as an SQL dump file.
Access provided by Autonomous University of Puebla. Download conference paper PDF
Similar content being viewed by others
Keywords
1 Introduction
Drug development process usually takes 10 to 17 years and has a high failure rate [1]. It costs 1,395 million USD per new drug compound [2], has low approval rate [3] and the trend is not in favor of pharmaceutical companies [2, 4, 5]. Drugs exhibit their functions by binding to particular sites on proteins, which alter the functions of the targeted protein. The drug (chemical) and drug target (protein) share a “key and lock” relation. This implies that it is possible to find drugs that match a particular target by knowing their shapes. Furthermore, one drug can bind to multiple targets and multiple drugs can bind to a target, each having different effects. Drug repositioning is defined as “the use of known drug and compound already on the market or in the later stage of development to treat new indications”. It is considered as the best alternative to drug development [1]. Drug repositioning can also be an incentive for pharmaceutical companies to invest in rare disease treatment.
Drug molecules and proteins have complex 3D shapes. Working and reasoning with 3D structures are process intensive and small changes in 3D structures leverage great effects in biological output. Furthermore, drug repositioning often requires analyzing three main data sets: phenotypes (diseases or side effects), biological entities (drug targets, genes, proteins, protein interactions, and pathways) and drugs (chemicals). Integrating these resources in a centralized database is time-consuming task due to different file formats, missing data and complex cross-referencing.
There are much endeavor among researchers to pursue studies in the field of drug repositioning [6]. For example, Yu et al. assigned weights to disease-protein complex and chemical-protein complex using symmetrical conditional probability to map disease to chemical [7]. This resulted in 40 drug-disease predictions, matching with CTD, and of which 9 are also supported by literature. In another study, Lee integrated multiple data sets mapping drug-protein, protein-protein, protein-disease and drug-disease, and applied a stochastic algorithm, which he called Shared Neighborhood Scoring (SNS) to weight edges between the nodes [8]. Lee et al. predicted 28 drugs for lung cancer, one of which is thia-benzthiazide. A study conducted under hypoxic found that thia-benzthiazidea could suppress proliferation of cancerous cells. Daminelli inferred chemical similarities based on targets and disease similarities. The three sets (drug, target and disease) were graph projected. Then edges were added to produce bicliques, which represent new predictions [9]. Twelve drug-target links and 11 drug-disease links were predicted. Nine drug-disease links were confirmed in literature. Re and Valentini integrated multiple chemical information such as structures, targets, and targets scores from the STITCH data set [10] to find chemical similarities [11]. The top ranked drugs L-Tryptophan, Darunavir and Adinazolam were confirmed by literature. Campillos et al. inferred drug similarities based on similar side effects [12]. The predicted similarities showed positive results when evaluated against drug target similarities. All the aforementioned studies use publicly available data sets, and require some pre-processing and cross-referencing. To avoid repeated work, it is necessary to integrate systematically all useful data into a single repository to aid drug repositioning studies.
This paper presents an integrated resource made from 12 publicly available data sets on various biological/chemical entities like disease, gene, protein, pathway, drug and side effect data, and 5 ontologies on disease-related SNP, disease related phenotype, protein function, drug function and side effect, each having varied levels of abstraction. Data sets from OMIM [13], GWAS [14], NCBI Gene [15], UniProt [16], InterPro [17], BioGRID [18], CORUM [19], Reactome [20], STITCH [10], TTD [21], DrugBank [22], SIDER [23] and data from HPO [24], EFO [25], GO [26], FTC [27] and MedDRA [28, 29] ontologies were processed, mapped and stored in a centralized single repository. To illustrate the usefulness of the repository, we identified relationships between drug and diseases, using side effect and pathway data. Our study predicted 117 associations, of which 93 are confirmed by the CTD database. The database is available on request from the authors as an SQL dump file.
The rest of this paper is organized as follows. Section 2 gives the relevant background on this work, Sect. 3 describes the methodology, and Sect. 4 gives the results and analysis. Section 5 concludes the paper.
2 Background
Biological Terms: Human non-blood cell nucleus contain 46 chromosomes. Chromosomes are coded sequence represented by four letters (bases) called the Deoxyribonucleic acid (DNA). DNA sequence are separated into smaller pieces, called genes, which are smallest units of heredity. Genes can be protein coding genes, which code for proteins or non-coding, which are mainly for regulatory purposes. Most genes code for a single protein. Genes go through a process called transcription where they leave the nucleus as mRNA and then undergo another process called Translation where amino acids from the cell cytoplasm are used to create a polypeptide chain of amino acids commonly referred to as the primary structure of the protein. The polypeptide folds into a 3D structure made up of alpha helices and beta strands, known as secondary structures. Proteins are made up of reusable units, called domains. A protein can have one or more protein domains. Once a protein is formed it may undergo other changes due to cellular surrounding known as post-translational modification (PTM). Multiple protein polypeptides form a protein complex. Proteins interact with each other forming a network of protein-protein interaction (PPI). Protein and small molecule interactions lead to a positive or negative outcome of the cell. A biological pathway consists of several of these interactions.
Human DNA consists of single-nucleotide polymorphism (SNP), a change in a single alphabet of the DNA sequence, which is responsible for people having different physical appearances (phenotype). Humans contain around 3 billion DNA base pairs and differ around one in every 1,200 to 1,500 DNA bases. SNP may be related to diseases and since DNA is inherited from parent to child, SNP related diseases are also inherited, known as non-communicable disease. Disease can be result of single SNP known as single-gene inheritance disease, or multiple SNPs known as multifactorial inheritance diseases.
Data Sources: OMIM [13] contains 7,894 phenotypes. OMIM is mapped to Human Phenotype Ontology (HPO) and Unified Medical Language System (UMLS). HPO [24] who classifies phenotype descriptions currently has 13155 entities and 12 levels. GWAS [14] contains over 100000 manually curated SNPs, having links with phenotypic traits. GWAS is mapped to Experimental Factor Ontology (EFO). EFO [25] has 20458 entities and13 levels. NCBI Gene [15] contains over 60167 homo-sapiens genes. UniProt contains over 60 million protein sequences, of which only about 550000 are manually curated. UniProt also provides annotation and mapping to multiple databases [16]. Gene Ontology (GO) [26] has three top-level namespaces namely ‘biological_process’, ‘molecular_function’ and ‘cellular_component’ which do not interconnect. GO has 47063 terms and 11 levels. It is available for download in OBO format. InterPro [17] contains 26238 domain entries curated from multiple sources along with annotations and links to GO. BioGRID [18] contains 1072173 protein interactions. CORUM [19] contains 2837 protein complexes from 3198 different genes and is linked to Functional Catalogue (FunCat) for complex characterization. Reactome [20] contains 6744 reactions and 1481 pathways. It contains mapping to NCBI Gene and UniProt. STITCH [10] contains chemical-protein mappings. It has 430000 compounds mapped to 9600000 proteins including non homo-sapiens proteins. Therapeutic target database (TTD) [21] contains drug targets details. DrugBank [22] contains 10527 drugs. DrugBank includes experimental, withdrawn and investigational drugs. FTC maps DrugBank entries to GO. FTC [27] is available for download at https://github.com/loopasam/ftc/. SIDER [23] contains 140064 drug-adverse-drug effects (ADRs) mapped to MedDRA and UMLS. MedDRA [28, 29] is a side effect ontology having five levels and 95912 terms.
3 Proposed Methodology
Data from different sources were downloaded, explored, and studied carefully. A subset of the data, which could be useful, was kept for further processing. For example, only Homo sapiens related data were retained. Furthermore, high expressivity data such as protein 3D structures and protein sequences were discarded and abstract data such as grouping and ontologies are taken into consideration.
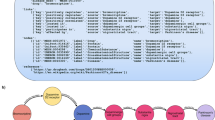
Ontologies are tree structured graphs having nodes and edges. Two nodes can be subset, equivalent, intersection or union of each other. Two edges, called ‘role’ are classified as subset, inverse and/or transitive of each other. Other edges types can be added to meet requirements. For the purpose of abstraction additional role mapping of lower level entities to their corresponding higher-level entities were calculated and stored. This mapping was done using subset edges only. The additional role tag is structured as follows: ‘map_L2’ where 2 represents the level to which the lower level entity was mapped to. The hierarchy level was calculated using the simple path algorithm [30]. This is illustrated in Fig. 1.
Ontologies were downloaded in OBO format and manipulated using R scripts. Data related to graphs are stored as edge-list in conventional RDBMS as it can be expanded quickly. Furthermore edge-list can store multiple roles and easier to retrieve and project using available mining toolkits. The centralized resource was built on PostgreSQL. Figure 2 shows the database Entity-Relationship (ER) diagram of the unified resource. The CTD data set [31] was added to the resource for evaluation purposes. CTD contains 202085 manually curated diseases and 1.8 million inferred chemical-disease.
A simplified ER diagram showing how the 12 data sets and 5 ontologies interlinked. Disease related data set and ontologies include GWAS, OMIM, EFO and HPO. Biological data sets and ontologies include NCBI_Gene, BioGRID, UniProt, InterPro, CORUM, Reactome and GO. Drug related data sets and ontologies include TTD, STITCH, DrugBank, SIDER, MedDRA, CTD and FTC.
4 Results and Analysis
The content of the resulting database, which we call Sirius (for shedding light on drug-disease relationships) is as follows:
-
Diseases
-
7321 phenotypes mapped to 6898 disease terms and to 16740 genes.
-
6108 SNPs mapped to 21327 genes.
-
1358 SNP disease terms mapped to 3685 genes.
-
-
Drugs
-
756 of 10527 drugs mapped to 4291 biological effect terms.
-
4688 of 10527 drugs mapped to 1199 side effects terms.
-
5618 proteins mapped to 2795 drugs.
-
2538 proteins mapped to 2553 drug targets.
-
1493 drugs mapped to 3110 phenotypes having 7139 manually curated relations and 276131 inferred relations.
-
-
Genes, proteins and their interactions:
-
41910 genes mapped to 1470499 PPIs.
-
28098 genes mapped to 33053 proteins.
-
100924 of 160361 proteins mapped to 13197 protein domain families.
-
100625 proteins mapped to 30545 protein functions terms.
-
2537 protein complexes involving a total of 3301 proteins
-
10619 proteins mapped to 2191 biological pathways.
-
10962 proteins mapped to 10984 reactions.
-
We illustrate the usefulness of our repository by studying the relationships between drugs and diseases, using side effect and pathway data. Our hypothesis is as follows: Chemicals affecting similar pathways and having similar side effects cure similar diseases. This implies that entries from DrugBank affecting similar Reactome pathways and having similar MedDRA entries share similar drug-disease relationships. This produces 253728 clusters (Table 1).
For evaluation, a second set of clusters grouping drug according to similar disease are projected. This produces 319 clusters. Three pairs of clusters were compared. The results are illustrated using a bar chart (Fig. 3) (Tables 2 and 3).
Cluster1 shows 28 drugs are predicted and 19 drugs are confirmed in the drug-pathway-side effect cluster1 of which 12 drugs intersect. Cluster2 shows 29 drugs are predicted and 23 drugs are confirmed in the drug-pathway-side effect cluster2, of which 12 drugs intersect. Cluster3 shows 38 drugs are predicted and 21 drugs are confirmed in the drug-pathway-side effect cluster3, of which 12 drugs intersect.
5 Conclusion
The paper presents a useful resource, Sirius, for analyzing drug-disease relationships for drug repositioning. Sirius integrates data from 12 publicly-available data sets and 5 ontologies into a single cleaned, unified repository, which is available for download on request from the authors. In brief, the Sirius contains SNP, phenotype, gene, protein, protein complex and function, PPI, pathway, drug and side effect data and cross-references each other. The resource is expected to avoid repetitive tasks of integrating multiple data sets by researchers doing drug repositioning. Furthermore this is an initiative to exploit the large amount of manually curated annotations publicly available in form of ontology mapping. To our knowledge, to date, there are no repositories cross-referencing these 12 databases and 5 ontologies.
The three cluster pair considered for evaluation of drugs grouping based on pathways and side effects yield at least 31.6% correct predictions. The result of the study showed it is possible to use observable phenomena and ontologies as classification tool for drug repositioning. Furthermore the resource allows multiple drug-disease hypotheses to be tested using statistics. For example, using existing drugs, biological effect terms and protein functions, drugs having similar biological effect may cure disease affecting proteins with similar functions.
References
Ashburn, T., Thor, K.: Drug repositioning: identifying and developing new uses for existing drugs. Nat. Rev. Drug Discov. 3(8), 673 (2004)
DiMasi, J., Grabowski, H., Hansen, R.: Innovation in the pharmaceutical industry: new estimates of R&D costs. J. Health Econ. 47, 20–33 (2016)
Mullard, A.: 2013 FDA drug approvals. Nat. Rev. Drug Discov. 13(2), 85–89 (2014)
DiMasi, J., Hansen, R., Grabowski, H.: The price of innovation: new estimates of drug development costs. J. Health Econ. 22(2), 151–185 (2003)
Pammolli, F., Magazzini, L., Riccaboni, M.: The productivity crisis in pharmaceutical R&D. Nat. Rev. Drug Discovery 10(6), 428–438 (2011)
Brown, A., Patel, C.: A standard database for drug repositioning. Sci. Data 4, 170029 (2017)
Yu, L., Huang, J., Ma, Z., Zhang, J., Zou, Y., Gao, L.: Inferring drug-disease associations based on known protein complexes. BMC Med. Genomics 8(2), 52 (2015)
Lee, H., Bae, T., Lee, J., Kim, D., Oh, Y., Jang, Y., Kim, J., Lee, J., Innocenti, A., Supuran, C., Chen, L.: Rational drug repositioning guided by an integrated pharmacological network of protein, disease and drug. BMC Syst. Biol. 6(2), 80 (2012)
Daminelli, S., Haupt, V., Reimann, M., Schroeder, M.: Drug repositioning through incomplete bi-cliques in an integrated drug–target–disease network. Integr. Biol. 4(7), 778–788 (2012)
Szklarczyk, D., Santos, A., von Mering, C., Jensen, L., Bork, P., Kuhn, M.: STITCH 5: augmenting protein–chemical interaction networks with tissue and affinity data. Nucleic Acids Res. 44(D1), D380–D384 (2015)
Re, M., Valentini, G.: Network-based drug ranking and repositioning with respect to DrugBank therapeutic categories. IEEE/ACM Trans. Comput. Biol. Bioinf. 10(6), 1359–1371 (2013)
Campillos, M., Kuhn, M., Gavin, A., Jensen, L., Bork, P.: Drug target identification using side-effect similarity. Science 5886(263–266), 321 (2008)
Amberger, J., Bocchini, C., Schiettecatte, F., Scott, A., Hamosh, A.: OMIM.org: online Mendelian Inheritance in Man (OMIM®), an online catalog of human genes and genetic disorders. Nucleic Acids Res. 43(D1), D789–D798 (2014)
Welter, D., MacArthur, J., Morales, J., Burdett, T., Hall, P., Junkins, H., Klemm, A., Flicek, P., Manolio, T., Hindorff, L., Parkinson, H.: The NHGRI GWAS Catalog, a curated resource of SNP-trait associations. Nucleic Acids Res. 42(D1), D1001–D1006 (2013)
Brown, G., Hem, V., Katz, K., Ovetsky, M., Wallin, C., Ermolaeva, O., Tolstoy, I., Tatusova, T., Pruitt, K., Maglott, D., Murphy, T.: Gene: a gene-centered information resource at NCBI. Nucleic Acids Res. 43(D1), D36–D42 (2014)
The Uniprot Consortium: UniProt: the universal protein knowledgebase. Nucleic Acids Res. 45(D1), D158–D169 (2017)
Mitchell, A., Chang, H., Daugherty, L., Fraser, M., Hunter, S., Lopez, R., McAnulla, C., McMenamin, C., Nuka, G., Pesseat, S., Sangrador-Vegas, A.: The InterPro protein families database: the classification resource after 15 years. Nucleic Acids Res. 43(D1), D213–D221 (2014)
Chatr-aryamontri, A., Oughtred, R., Boucher, L., Rust, J., Chang, C., Kolas, N., O’Donnell, L., Oster, S., Theesfeld, C., Sellam, A., Stark, C.: The BioGRID interaction database: 2017 update. Nucleic Acids Res. 45(D1), D369–D379 (2017)
Ruepp, A., Waegele, B., Lechner, M., Brauner, B., Dunger-Kaltenbach, I., Fobo, G., Frishman, G., Montrone, C., Mewes, H.: CORUM: the comprehensive resource of mammalian protein complexes—2009. Nucleic Acids Res. 38(suppl_1), D497–D501 (2009)
Croft, D., Mundo, A., Haw, R., Milacic, M., Weiser, J., Wu, G., Caudy, M., Garapati, P., Gillespie, M., Kamdar, M., Jassal, B.: The Reactome pathway knowledgebase. Nucleic Acids Res. 42(D1), D472–D477 (2014)
Li, Y., Yu, C., Li, X., Zhang, P., Tang, J., Yang, Q., Fu, T., Zhang, X., Cui, X., Tu, G., Zhang, Y.: Therapeutic target database update 2018: enriched resource for facilitating bench-to-clinic research of targeted therapeutics. Nucleic Acids Res. 46(D1), D1121–D1127 (2017)
Wishart, D., Feunang, Y., Guo, A., Lo, E., Marcu, A., Grant, J., Sajed, T., Johnson, D., Li, C., Sayeeda, Z., Assempour, N.: DrugBank 5.0: a major update to the DrugBank database for 2018. Nucleic Acids Res. 46(D1), D1074–D1082 (2017)
Kuhn, M., Letunic, I., Jensen, L., Bork, P.: The SIDER database of drugs and side effects. Nucleic Acids Res. 44(D1), D1075–D1079 (2015)
Köhler, S., Doelken, S., Mungall, C., Bauer, S., Firth, H., Bailleul-Forestier, I., Black, G., Brown, D., Brudno, M., Campbell, J., Fitzpatrick, D.: The Human Phenotype Ontology project: linking molecular biology and disease through phenotype data. Nucleic Acids Res. 42(D1), D966–D974 (2013)
Malone, J., Holloway, E., Adamusiak, T., Kapushesky, M., Zheng, J., Kolesnikov, N., Zhukova, A., Brazma, A., Parkinson, H.: Modeling sample variables with an Experimental Factor Ontology. Bioinformatics 26(8), 1112–1118 (2010)
Gene Ontology Consortium: Gene ontology consortium: going forward. Nucleic Acids Res. 43(D1), D1049–D1056 (2014)
Musen, M., Noy, N., Shah, N., Whetzel, P., Chute, C., Story, M., Smith, B., NBCO team: The national center for biomedical ontology. J. Am. Med. Inf. Assoc. 19(2), 190–195 (2011)
Brown, E., Wood, L., Wood, S.: The medical dictionary for regulatory activities (MedDRA). Drug Saf. 20(2), 109–117 (1999)
Merrill G.: The meddra paradox. In: AMIA Annual Symposium Proceedings, vol. 2008, p. 470 (2008)
Csardi, G., Nepusz, T.: The igraph software package for complex network research. InterJ. Complex Syst. 1695(5), 1–9 (2006)
Davis, A., Grondin, C., Johnson, R., Sciaky, D., King, B., McMorran, R., Wiegers, J., Wiegers, T., Mattingly, C.: The comparative toxicogenomics database: update 2017. Nucleic Acids Res. 45(D1), D972–D978 (2017)
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Switzerland AG
About this paper
Cite this paper
Maudarbux, M.M., Ghoorah, A.W., Fowdur, T.P. (2019). Sirius: A Resource for Analyzing Drug-Disease Relationships for Drug Repositioning. In: Fleming, P., Lacquet, B., Sanei, S., Deb, K., Jakobsson, A. (eds) Smart and Sustainable Engineering for Next Generation Applications. ELECOM 2018. Lecture Notes in Electrical Engineering, vol 561. Springer, Cham. https://doi.org/10.1007/978-3-030-18240-3_22
Download citation
DOI: https://doi.org/10.1007/978-3-030-18240-3_22
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-18239-7
Online ISBN: 978-3-030-18240-3
eBook Packages: EngineeringEngineering (R0)