Abstract
The environment changes faster than the ability of genetic mutation and recombination to generate natural genetic diversity. In this context, epigenetic regulation of gene expression has the potential to provide organisms with an alternative mechanism for phenotypic variation by controlling the extent of plasticity that can be achieved in response to environmental changes. There is now substantial evidence suggesting roles for epigenetic regulation of several different aspects of the plant response to biotic stress. At the basic level of gene expression, posttranscriptional gene silencing mediated by small RNAs and chromatin remodelling controlling transcriptional gene silencing are essential for the induced resistance responses activated during pest and pathogen attack. Beyond this, there is also evidence that histone modifications and DNA methylation are associated with immune memory, or defence priming, such as systemic acquired resistance (SAR). In addition, recent evidence indicates that epigenetic modifications can also generate longer-term defence priming responses that can be inherited across generations. In this chapter, we will discuss the roles of epigenetics in these different modes of biotic stress resistance, and suggest ways in which we may in the future be able to exploit epigenetic systems for crop protection.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
2.1 Introduction
Like all living organisms, plants need to adapt to environmental changes in order to persist. Until the last decades, neo-Darwinian evolutionary theories assigned the origin of phenotypic variability to a set of characteristics determined by the genetic information. In the case of facing a change in the environment, different individuals within a population will have differential survival, depending of their characteristics, that will determine the genetic, and therefore phenotypic features of the new generations (Pigllucci 1996). Surprisingly, some recent observations lead different experts to claim for an implementation of this now classical new Darwinian perspective (Rando and Verstrepen 2007). For example, mutation rates are usually slower than environmental changes, and the phenotypic plasticity observed in natural populations wider than genetic variability. Therefore, an extra source of phenotypic plasticity is expected (Grativol et al. 2012). On the other hand, some environmentally induced adapted states seem to be relatively stable or even inherited for few generations without involving a change in the genetic information. This extra layer of relatively stable phenotypic plasticity has also been called epigenetic buffering (understood as ‘something’ beyond genetics). Epigenetic buffering could have a special importance in the case of plants, which because of their sessile nature, face threats to their survival and fitness from biotic stresses (O’Dea et al. 2016).
Against a pathogen attack, plants counter with a broad range of defence mechanisms. Plants possess some constitutive barriers to protect themselves against potential pathogens that are usually effective against a variety of microbes (Malinovsky et al. 2014). Along their evolutionary arms race for survival, pathogens have developed several strategies to overcome those defences and produce infections. In response to those, plants are able to actively induce defences when they identify a microbe or herbivore as a threat (Jones and Dangl 2006). This is associated with a reprogramming of gene expression. In some cases, once they have suffered a stress that induced their defences, plants are able to remember this first stress encounter. Then, in the case of recurrent stresses, the induced responses are faster and stronger (usually more effective). This is the concept of priming of defence responses, which involves a different control of gene expression and it is inevitably associated with a memory of the stress (Prime-A-Plant Group et al. 2006). Waddington in 1942 coined the term epigenetics to describe the study of phenomena in which the phenotypes observed in nature cannot be explained just by the understanding of their genotypes (Waddington 2012). He was studying development. At that time, the scientific community understood the genome as packages of information encoding specific features, but the control of that information (gene expression control) was an unknown. Of course, they could not imagine that part of the information stored in the genome works, in fact, by controlling gene expression (regulatory regions, transcriptional factors, etc.) and it is therefore genetic. Along the last decades, epigenetics has been redefined as phenotypic changes that can be transmitted through mitotic or even meiotic divisions in the absence of changes in the DNA sequence. Thus, epigenetics is associated with the control of gene expression and certain memory. Importantly, some of the initially considered epigenetic mechanisms have been found to be encoded in the genome (miRNAs, chromatin modellers, histone variants, etc.). However, at all stages of induced immunity, the considered epigenetic mechanisms have been demonstrated to be of a key importance.
In this chapter, we will first introduce the different epigenetic mechanisms. We will focus on their role in controlling gene expression at the transcriptional and posttranscriptional level, contextualizing them by the use of some examples of their involvement in plant response to pest and diseases. Then, we will dedicate a section to assess the role of epigenetics in the memory of the stress and priming of defence responses. We will present and discuss publications supporting the role of epigenetic mechanisms in priming at different timescales, from short to long periods of time or even trans-generationally. Finally, we will summarize and discuss the potential application of epigenetics in the development of alternative programs for plant protection. We believe, this integrated view of epigenetics in plant defence and priming could be inspiring for a new generation of plant scientists aiming to understand plant defence mechanisms, as well as to develop alternative, hopefully more effective and sustainable, crop protection strategies.
2.2 Epigenetic Mechanisms Involved in Plant Defence
As originally defined, epigenetics allows plants to show different phenotypes with the same genotype. The underlying question for years was how? Nowadays, we know that the majority of what are considered epigenetic mechanisms are centred on the control of gene expression. From this perspective, the so-called epigenetic buffering , which is used to explain the extra source of phenotypic plasticity, does not involve a change in the information itself, but the different observed phenotypes are a consequence of modifying the speed and intensity at which genes are expressed (Grativol et al. 2012). Although epigenetics has traditionally been associated with repression of gene expression (reason why they are called ‘silencing’ mechanisms), we now also know of mechanisms considered as epigenetic that are able to promote or facilitate gene expression (Eamens et al. 2008; Matzke and Matzke 2000). This gene expression control can be imposed at either the transcriptional or posttranscriptional level. For this reason, epigenetic mechanisms are typically classified in two groups: those controlling gene expression at transcriptional level, known as ‘transcriptional gene silencing mechanisms’ (TGS), and those controlling gene expression at posttranscriptional level, ‘posttranscriptional gene silencing mechanisms’ (PTGS).
2.2.1 Posttranscriptional Control (PTGS): The Role of Small RNAs
In general terms, the main part of posttranscriptional epigenetic control is associated with the action of small RNAs (sRNA). Despite the fact that sRNAs pathways play important roles in developmental processes, they seem to have evolved originally from a defence mechanism against viruses and transposable elements to later start silencing endogenous genes (Borges and Martienssen 2015; Matzke and Matzke 2000). The first reports of epigenetic mechanisms in plant defence were examples of posttranscriptional gene silencing. Actually, reports of plant defence and priming processes as part of the cross-protection observed against viruses in the 1920s could be considered the very first recognized examples of epigenetic posttranscriptional gene silencing (Ross 1961; Waterhouse et al. 2001). They described how infections with relatively avirulent virus strains can induce protection against related virulent viruses. Unfortunately, at that time, the mechanisms were far from being discovered, and remained unknown for decades. It was not until the late 1980s that the first examples of posttranscriptional gene silencing mechanisms emerged, during experiments by plant biotechnologists trying to alter the colour of petunia flowers (Eamens et al. 2008). They observed how the expression of a transgene could trigger the silencing of the transgene and also homologous endogenous genes, by mechanisms involving small RNAs. Today we know both observations were related. It is generally accepted that all the epigenetic mechanisms involving RNA intermediates were originally part of the plant defence mechanisms against viruses. Virus transcription and replication is carried out inside the host cell, so the plants evolved the ability to detect exogenous nucleic acids as a potential threat (Waterhouse et al. 2001). Thus, plants first developed a system to defend themselves against viruses (exogenous RNAs). Then, the system was adapted to an endogenous control for gene expression and genome stability (controlling the movement of transposable elements—TEs—that are similar to viruses). That is the reason why advances in sRNA-mediated silencing mechanisms and plant defence have been feeding from each other during decades.
Derived from this original antiviral function, all small RNAs possess common features in their biogenesis. For example, most of the small RNAs required a longer double-stranded RNA (dsRNA) precursor molecule. This could be due to the fact that the 90% of plant viruses depend on a dsRNA molecule for their replication (Waterhouse et al. 2001). The dsRNAs are processed by DICER-LIKE proteins (DCLs) to generate fragments of 21–24 nt length. These molecules are then 2′-O-methylated by the protein HUA ENHANCER 1 (HEN1) at the 3′ end, which is a specific aspect of plant sRNAs (Yang et al. 2006). Finally, mature sRNAs are loaded into ARGONAUTE (AGO) proteins which can interact with other proteins to form the RNA-induced silencing complexes (RISCs; Fang and Qi 2016). In posttranscriptional gene silencing mechanisms, RISC complexes find the target mRNA by base pairing and affect its stability through mRNA cleavage (degrading the target mRNA), or repress its translation.
2.2.1.1 Silencing of Exogenous RNAs. The Special Case of Viruses
Due to its evolutionary origin, the majority of antiviral defences are triggered by exogenous RNAs (viral RNAs). Viral infection activates epigenetic mechanisms in order to destroy or silence the invading viral genome (Waterhouse et al. 2001). In Fig. 2.1 we represent a summary of the different silencing mechanisms against viruses as discovered in Arabidopsis (adapted from Ruiz-Ferrer and Voinnet 2009). These biological roles of the silencing components were mainly deciphered by analysing plant defective mutants (reviewed in Katiyar-Agarwal and Jin 2010; Seo et al. 2013). For RNA viruses (the majority of plant viruses), the viral genome is replicated by a viral replicase to generate a dsRNA molecule (Fig. 2.1a). This dsRNA would be processed by plant DCL proteins to trigger the production of sRNAs that would lead to viral silencing by degrading the dsRNA replication intermediates, the viral transcripts or impeding their translation. Viral transcripts (single-stranded RNAs—ssRNAs) can also trigger silencing by being copied to a dsRNA molecule by plant RNA-dependent RNA polymerases (RDR proteins), feeding the system. Those RDR proteins are also involved in the silencing of the viruses which have ssRNA genomes. In the case of the DNA viruses (Geminivirus—family Geminiviridae—for instance), the viral transcripts are copied to dsRNA by the plant RDR2 protein which in this case would mediate the production of sRNAs similar to the heterochromatic sRNAs, triggering the silencing of the virus at transcriptional level (Fig. 2.1b, and further discussed in Sect. 2.2; Raja et al. 2008). Moreover, viruses spread through the plant using the vasculature (Hipper et al. 2013). In this respect, a striking property of the sRNAs is their cell-to-cell mobility, increasing the efficiency of the silencing mechanism. Once the silencing is locally triggered in the infected tissue, the sRNAs can travel through the plant xylem and phloem system and provide systemic resistance by silencing targets in distal parts of the plant (Chitwood and Timmermans 2010; Kalantidis et al. 2008). This systemic silencing has been broadly used by biotechnologists for decades, but more importantly, it has positioned the sRNAs as good candidates to be the mobile signal involved in some systemic resistance processes (Voinnet 2005).
Epigenetic mechanisms controlling viral infection. Adapted from Ruiz-Ferrer and Voinnet (2009). Epigenetic mechanisms against RNA (a) and DNA (b) virus. Pathogen components are highlighted in blue. Plant proteins are represented in green. Plant defence as a process is represented with red and orange colours
2.2.1.2 Endogenous RNAs: From an Antiviral Defence to the Control of Endogenous Sequences
From these early mechanisms designed as antiviral defences, plants (and other organisms) evolved the capacity to use the silencing machinery in controlling other sequences (Waterhouse et al. 2001). This evolution leads to the sRNA-mediated regulation of endogenous genes and the suppression of transposable element (TE) movement. Nowadays there is an increasing number of different sRNAs recognized (Borges and Martienssen 2015). Several classifications have been proposed. On the basis of their biogenesis, we can consider two major classes: small interfering RNAs (siRNAs) and micro RNAs (miRNAs). Maybe the better-known of the two are the miRNAs, which are typically 20–22 nt length, transcribed by RNA polymerase II (Pol II) and processed by DCL1. Among the endogenous siRNAs there are many different subclasses. On the one hand, there are the hairpin-derived siRNAs (hp-siRNAs) and natural antisense siRNAs (natsiRNAs), both 21–24 nt length. On the other hand, the ‘secondary siRNAs’, including trans-acting siRNAs (tasiRNAs), phased siRNAs (phasiRNAs), epigenetically activated siRNAs (easiRNAs) and the long siRNAs (lsiRNAs). The lsiRNAs represent a class of endogenous siRNAs identified specifically in plant–pathogen interactions with a length of 30–40 nt. Finally, plants also produce heterochromatic siRNAs (hetsiRNAs), which from a classical view of the pathway are 24 nt length and generated by the plant-specific RNA-dependent DNA polymerase IV (Pol IV). hetsiRNAs are involved in transcriptional gene silencing, so they will be addressed later (see Sect. 2.2). Both miRNA and non-heterochromatic siRNAs bind the RISCs complexes and find the target mRNA (in this case a plant gene transcript), by base pairing. In the same way that the viral sRNAs are eliminated, the control of the target gene is achieved by mRNA cleavage (degrading the target mRNA) or repression of its translation.
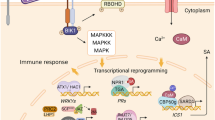
The majority of the sRNA classes have been demonstrated to play a role in the control of plant defence at almost all stages. In general terms, once the pathogen is recognized by the plant, the induced defence process includes some conserved elements (Tsuda et al. 2008) such as the production of reactive oxygen species (ROS), which on specific occasions can lead to a hypersensitive response (HR) and apoptosis, an intracellular cascade mediated by MAP kinase proteins (MAPK) and hormonal signalling which generally involves a transcriptional reprogramming. Salicylic and jasmonic acid (SA and JA, respectively) are considered the two main hormonal pathways involved in plant defence. Plants adjust their immune system depending on the lifestyle of the attacker they encounter. Generally, defence against attack by biotrophic pathogens, which feed from living cells, is mediated by the SA pathway. Conversely, necrotrophic pathogen infections, which kill the tissues to feed from them, or herbivores are resisted by the JA/ethylene (ET) pathway (Pieterse et al. 2012). An effective focus of the resources in defence is achieved by the prioritization of one of the pathways at the expense of the other, once the pathogen is recognized (Glazebrook 2005). As a consequence, it is common to find opposite phenotypes for different lifestyle pathogens when one of the pathways is active (Vos et al. 2015). In addition to SA, JA and ET, other hormones such as auxins, abscisic acid (ABA), cytokinins, gibberellins and brassinosteroids play a secondary, but nevertheless important, role in plant defence (Pieterse et al. 2012). These are general defence mechanisms triggered by the plant independently of the recognition of the pathogen.
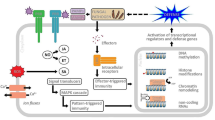
It is generally accepted that there are two main routes by which the plant can activate these various defences, depending on the recognition of the pathogen. These are ‘PAMP triggered immunity (PTI)’ and ‘effector-triggered immunity (ETI)’. In Fig. 2.2, we contextualize some of the most important sRNA examples controlling PTGS described to date in both branches of plant defence.
Posttranscriptional epigenetic mechanisms involved in plant defence. The diagram represents some of the identified small RNAs controlling defence responses at the level of PAMP triggered immunity (a) and effectors triggered immunity (b). Pathogen components are highlighted in blue. Plant elements are represented in green. Plant defence as a process is represented with red and orange colours
2.2.1.2.1 sRNAs in PAMP Triggered Immunity (PTI)
The first branch in plant defence is triggered when the plant recognizes general microbial- or pathogen-associated molecular patterns (PAMPs), such as flagellin (common to different bacteria) or chitin (common to many fungi) by the use of transmembrane pattern recognition receptors (PRRs). This recognition leads into the PAMP triggered immunity (PTI) by the activation of a defence signalling cascade (Fig. 2.2a). miRNAs modulate different stages of the plant defences but they have a special contribution in PTI (Voinnet 2008). As a reflection, dcl1 mutants (strongly impeded in the production of miRNAs) are effectively infected by the usually avirulent hrcC strain of Pseudomonas syringae DC3000 (Navarro et al. 2008). One of the first observations in this respect was a remarkable change in miRNAs populations during infection, and in response to exogenous PAMP applications. During PTI some miRNA species are repressed (represented as ↓ miRNAs in Fig. 2.2a). This is the case of miRNAs controlling positive elements in the defence response, like the miRNA398 (Fig. 2.2a; Jagadeeswaran et al. 2009). The repression of miR398 releases its targets (superoxide dismutases CSD1 and CSD2), and as a consequence, enhances callose deposition, reinforcing the cell wall and impeding the pathogen infection (Li et al. 2010). In contrast, other miRNAs are induced during PTI (↑RNA), which are usually the ones that control negative regulators. Maybe the one that could be considered the most relevant example until date is the case of the miR393. Transcription of miRNA393 is induced during PTI (by both flagellin treatments and Pseudomonas syringae—Pst—infections) and it is accompanied by a repression of its targets, which in turn causes repression of the auxin signalling pathway (Fig. 2.2a). Repression of the auxin pathway in this way seems to lead in resistance against Pst by hormonal crosstalk (Navarro et al. 2006). Apparently, by this hormonal crosstalk, the plant would prioritize the expense of the resources in defence over growth by repressing auxin pathway during a defence response. In fact, there are other miRNAs involved in the repression of the auxin pathway during PTI, like miR160 and miR167 (Fig. 2.2a; Zhang et al. 2011). The fine-tuning of the defence responses through a hormonal control is not exclusive of the auxin pathway. Actually, as introduced before, it is accepted in the field that plants tailor their defence responses in accordance with the attacker’s lifestyle by the negative crosstalk of SA and JA pathways, the two main hormonal pathways involved in immune responses. This crosstalk is also under the control of miRNAs. This is the case of the miR319 which is induced in response to a/virulent hemibiotrophic bacteria (Pst DC3000, Pst DC3000 hrcC, and Pst DC3000 avrRpt2). miR319 represses JA pathway components, and its induction during defence responses against biotrophic pathogens could therefore have the objective of prioritizing SA-related defences at the expense of JA responses (Fig. 2.2a; Zhang et al. 2011). This kind of immunity has been demonstrated to stop the colonization of many different pathogens. However, some plant pathogens evolved to somehow interfere with these PTI mechanisms by the use of specific molecules called as effectors.
Host-adapted pathogens use effectors to suppress PTI and thus successfully colonize their host. There are few identified cases of pathogens manipulating the plant silencing mechanisms as part of the immune system. One of the most relevant cases of pathogen effectors suppressing silencing comes once again from viruses. As effectors, viruses produce viral suppressors of RNA silencing (VSRs) to counteract their silencing (Fig. 2.1). Those are the best-studied examples of suppressors of silencing and are able to interfere with silencing at different stages (Fig. 2.1; Csorba et al. 2015). However, there are also a few identified cases of other pathogens manipulating the plant silencing mechanisms. This is the case of the miR393, and probably miR159, which are repressed by AvrPto effectors of Pst (Navarro et al. 2008; Zhang et al. 2011). One of the most interesting examples of this co-evolution comes from the discovering siRNAs produced by the pathogen Botrytis cinerea. The fungus produces siRNAs as effectors in order to hijack the plant silencing machinery, facilitating its infection (Weiberg et al. 2013). Nonetheless, along their shared evolutionary path, some plants have also acquired the ability to detect pathogen effectors, triggering the second branch of plant immunity.
2.2.1.2.2 sRNAs in Effector-Triggered Immunity (ETI)
Plant recognition of the presence of effectors triggers the second branch of plant defence, which is known as effector-triggered immunity (ETI). Effector detection is carried out by what are sometimes referred to as R proteins (from ‘resistance proteins’). R proteins, also called guard proteins, can be intra- or extracellular and detect the presence of effectors either directly (e.g. by direct protein–protein interaction) or indirectly, via the outcome of the effector’s interference with PTI. During ETI , induced defences are typically much stronger than PTI (Fig. 2.2b). One remarkable example of epigenetic control of ETI concerns the small interfering RNAs nat-siRNAATGB2 and AtlsiRNA-1 (Fig. 2.2). They were both found to take part in the ETI response against the strain avrRpt2 of Pseudomonas syringae pv tomato. In an avirulent interaction, the plant R protein RPS2 is able to detect the action of the effector avrRpt2 degrading the plant defence protein RIN4, and triggers an ETI response. As part of that system nat-siRNAATGB2 and AtlsiRNA-1 are induced and contribute to the releasing of the defences by the inhibition of PPRL (Katiyar-Agarwal et al. 2006) and AtRAP, both negative regulators of RPS2-related defences (Katiyar-Agarwal et al. 2007). Those are just some examples of the roles of silencing mechanisms along the co-evolutionary history between plants and their pathogens. Strikingly, some recent discoveries show how, as a last counteracting measure, some plant sRNAs can be transferred by extracellular vesicles to the pathogen in order to trigger silencing of virulence factors as part of ETI response (Fig. 2.2, Cai et al. 2018). This demonstrates once again the important role of the small RNAs in the plant–pathogen arm race.
2.2.1.3 The State of the Field, from Arabidopsis to Other Species
Unfortunately, even though the first studies started many years ago using tobacco as a model species, in the rise of epigenetics as a hot field during the last 20 past years, the majority of the research has been done in Arabidopsis plants. Infections of Pst in Arabidopsis have been consolidated as a model pathosystem. However, as translational science strategies are rapidly building on this fundamental knowledge, nowadays there is evidence coming from many different pathosystems, including crop species. For instance, the role of the silencing machinery against viruses has been investigated in rice against the rice stripe virus (Jiang et al. 2012). In Brassica, there has been identified a miRNA (bra-miR1885) which appears to be specifically targeted by viral effectors from TuMV virus. In addition, TuMV infections (but not TMV or CMV) induce bra-miR1885 levels which represses a defence-related protein, facilitating virus infection (He et al. 2008). It has also been reported that miR393 is conserved across different plant species such as rice and cucumber (Bian et al. 2012; Xu et al. 2017), and there is even evidence in soybean pointing to a conserved role of its function in PTI (soybean—Phytophthora sojae infections; Wong et al. 2014). Another notable example is the tomato miR482 family, which represses components of the plant basal defences and is down-regulated during bacterial and viral infections (Shivaprasad et al. 2012). Last decade there has also been an increment in the range of pathogen interactions analysed, including bacteria (Alizadeh et al. 2018; Zhao et al. 2013), fungi (Ellendorff et al. 2009; Li et al. 2014, 2016; Shen et al. 2014), oomycetes (Li et al. 2012), viruses (Li et al. 2012), cyst nematodes (Hewezi et al. 2008; Zhao et al. 2015) and other herbivores. Importantly, the majority of the crop studies started with genomic and transcriptomic analysis, with a strong in silico component (Guo et al. 2011; He et al. 2014; Jeyaraj et al. 2017; Kapoor et al. 2008; Lu et al. 2007; Pandey et al. 2008; Pérez-Quintero et al. 2012; Qiu et al. 2009; Radwan et al. 2011; Warren and Covert 2004; Xin et al. 2010; Yin et al. 2012). Much more work needs to be done in vivo to be able to include these epigenetic mechanisms as agronomical tools for the development of crop protection strategies.
2.3 Transcriptional Control (TGS): Chromatin Remodelling
Epigenetics can also modify gene expression at the transcriptional level. The DNA is compacted in the nucleus by association with proteins in what we know as chromatin (Kornberg 1974). The basic units of chromatin are called nucleosomes and are formed by an octamer of histone proteins (two of each H2A, H2B, H3, and H4; Van Holde et al. 1974) and approximately 146 bp of DNA, or 1.7 turns, wrapping the histone octamer. The DNA between nucleosomes is called linker DNA and for higher levels of compaction can be associated with histone H1 (Fig. 2.3a). Depending on the physicochemical interaction of the DNA and histone proteins in the nucleosome, the chromatin can have different levels of compaction (Fig. 2.3b). This compaction is essential as, for the main part of the functions of DNA such as gene expression and replication, the DNA should be accessible to large protein complexes. However, the entire genome does not fit into the nucleus in the uncompacted state (Li et al. 2007). Thus, during interphase, the transcribed regions are uncompacted constituting what is known as open chromatin, while other regions are preserved (silenced) in very compacted chromatin regions. At the same time, the compaction of the DNA at some regions should be maintained, as this preserves the genetic information from damage and the jumping of TEs. In response to an environmental stimulus, the chromatin has been proposed to interpret the signal and facilitate the gene reprogramming (Fig. 2.3b; Badeaux and Shi 2013). As the chromatin compaction has been directly related to the transcriptional control of gene expression, all mechanisms modifying chromatin compaction are considered epigenetic mechanisms controlling transcriptional gene silencing (Fransz and de Jong 2011). Among such mechanisms, the most important are: chromatin remodellers, deposition of histone variants, histone posttranslational modifications and DNA methylation. Similar to the control of PTGS, chromatin compaction has been demonstrated to play a crucial role in the fine-tuning of defence responses. Importantly, the chromatin state has been proposed to be the mechanism underlying priming processes and memory of the stress, which will be addressed in Sect. 2.3.
Transcriptional epigenetic mechanisms involved in plant defence. (a) Representation of a nucleosome unit. The DNA wraps around an octamer of histone proteins (2× -H2A, H2B, H3 and H4-). H1 is located in the linker DNA (between nucleosomes). (b) The main part of the epigenetic mechanisms controlling defence at transcriptional level influences the chromatin compaction. In response to pathogen attack, plants activate epigenetic mechanisms to open the chromatin at the level of genes involved in defence responses, facilitating their expression and/or compacting chromatin regions containing defence repressors.  DNA methylation,
DNA methylation,  histone acetylation,
histone acetylation,  histone methylation as a negative mark (for instance, H3K9me2),
histone methylation as a negative mark (for instance, H3K9me2),  histone methylation as a positive mark (for instance, H3K4me3 or H3K36me),
histone methylation as a positive mark (for instance, H3K4me3 or H3K36me),  histone ubiquitination. (c) Some of the chromatin remodellers involved in plant defence. (d) Deposition of histone variants in plant defence processes. Fig. 2.3 (continued) Specifically, the case of the H2AZ is represented in the diagram. (e) Some histone acetylation examples in response to pathogen attack. (f) Histone methylation examples in response to pathogen attack. (g) Histone ubiquitination associated with plant defence. (h) DNA methylation changes mediated by the RdDM pathway and ROS1 in response to Pst infections
histone ubiquitination. (c) Some of the chromatin remodellers involved in plant defence. (d) Deposition of histone variants in plant defence processes. Fig. 2.3 (continued) Specifically, the case of the H2AZ is represented in the diagram. (e) Some histone acetylation examples in response to pathogen attack. (f) Histone methylation examples in response to pathogen attack. (g) Histone ubiquitination associated with plant defence. (h) DNA methylation changes mediated by the RdDM pathway and ROS1 in response to Pst infections
2.3.1 ATP-Dependent Chromatin Remodellers
The ATP-dependent chromatin remodellers are multiprotein complexes that are able to disrupt the interaction between the DNA and histone proteins using energy provided by the hydrolysis of ATP molecules (Fig. 2.3c; Han et al. 2015). They are conserved cross-kingdom in eukaryotes. In plants, the ATPase function resides in proteins from the Snf2 superfamily. Members of the Snf2 superfamily involved in plant defence identified to date are: BRAHAMA (BRM), SPLAYED (SYD), DECREASE IN DNA METHYLATION 1 (DDM1), PHOTOPERIOD-INDEPENDENT EARLY FLOWERING 1 (PIE1), BIT-RESPONSIVE HISTONE-INTERACTING SNF2 ATPASE 1 (BRHIS1) and CHROMATIN-REMODELLING FACTOR 5 (CHR5). BRM is maybe the most canonical and well-studied chromatin remodeller in plants, and it has been widely studied in responses against abiotic stresses, where it controls abscisic acid (ABA)-related genes (Han et al. 2012; Peirats-Llobet et al. 2016). However, some defence-related genes have been observed as misregulated in the brm101 mutant, pointing to a defence role of BRM, probably by crosstalk between ABA and SA hormonal pathways (Bezhani et al. 2007; Ramirez-Prado et al. 2018). SYD has been reported to play a role in the activation of JA/ET related defences. On the one hand, SYD seems to bind some promoter regions for JA/ET-related genes, probably promoting the opening of the chromatin. On the other hand, mutants defective in SYD are consistently more susceptible to the necrotrophic pathogen B. cinerea and unable to properly induce appropriate target genes (Walley et al. 2008). In addition, some SYD mutants are more resistant to biotrophic pathogens such as Hyaloperonospora arabidopsidis (Hpa) and Pseudomonas syringae pv. maculicola (P.s.m.; Johnson et al. 2015). DDM1 plays a critical role in the maintenance of the DNA and histone H3 methylation pattern. In this case, DDM1-mediated chromatin opening does not recruit gene activators, but it seems to allow the recruitment of methyltransferases to its target regions, inducing gene silencing (Gendrel et al. 2002; Jeddeloh et al. 1998; Zemach et al. 2013). It therefore links the direct opening of the chromatin with changes in the histone protein modifications and DNA methylation. DDM1 has been related with the control of the SA-related defences and the silenced basal state of RPP5, a six-defence gene cluster (Yi and Richards 2007, 2009). Accordingly, the ddm1 mutant plants are more resistant to Hpa (López Sánchez et al. 2016). Recently it has been reported how the SNC1 gene from the RPP5 cluster is also regulated by another ATP-dependent chromatin remodeller, CHR5. In this case, CHR5 acts as a positive regulator (classical chromatin remodeller function) opening the chromatin at the level of SNC1 gene. The mutant chr5 shows an inability to open the chromatin at SNC1 level, exhibits increased nucleosome deposition along the whole genome, and hyper-susceptibility to virulent and avirulent strains of Pst (Zou et al. 2017). Finally, the chromatin remodellers PIE and BRHIS1, like DDM1, act as intermediaries for other chromatin modifications such as the deposition of histone variants and histone monoubiquitination, respectively (discussed later).
2.3.2 Deposition of Histone Variants: Histone Replacement
One of the mechanisms involved in chromatin remodelling is the replacement of the canonical histone by specific histone variants. Due to different physicochemical properties, the interaction of the histone variants with the DNA in the nucleosome can modify chromatin compaction and thus, gene expression at transcriptional level (Coleman-Derr and Zilberman 2012). That is the reason why the deposition of different histone variants has been proposed as a mechanism mediating responses to environmental changes (Talbert and Henikoff 2014). One of the most important examples for this mechanism is the case of the H2A.Z (Fig. 2.3d). First, March-Diaz et al. (2008) described how mutants defective in the H2A.Z coding genes, as well as the SWR1 chromatin remodelling complex, which facilitates its deposition (including the above-mentioned PIE protein), constitutively express SA-related genes and are more resistant against biotrophic pathogens. The authors suggested that the deposition of H2A.Z has a role in controlling the silencing of SA-related genes in a basal state (March-Díaz et al. 2008). Recent works have confirmed the role of this histone variant in repressing SA-related genes, but they report hyper-susceptibility for both biotrophic and necrotrophic pathogens (Berriri et al. 2016). However, the deposition of the histone variant H2A.Z seems to be key for the fine-tuning of the defence responses.
2.3.3 Modification of the Histone Proteins
Histone proteins can be posttranscriptionally modified at different residues, having an important impact in the chromatin state (Kouzarides 2007). More than 60 positions have been detected in the tail of the core histone proteins that can potentially carry posttranscriptional modifications (PTMs) of a variable nature. It is becoming clear that PTM of histone proteins imparts a dynamic and complicated regulation of gene expression, particularly in plant defence processes. This regulation is a consequence not just of the appearance of one PTM, as they can have an individually positive or negative contribution for chromatin compaction, but the consensus of several interacting PTMs, in what is called ‘the histone code’. The PTMs that have been demonstrated to play a key role in plant defence are related with acetylation, methylation and ubiquitination of different residues (Fig. 2.3e–g). There have been very recent and comprehensive reviews in this field (Chen et al. 2017; Ding and Wang 2015; Ramirez-Prado et al. 2018). The majority of cases have been described once again in Arabidopsis, so here we will just outline some of the most notable examples to offer a general view.
2.3.3.1 Acetylation
Generally, the acetylation of residues in histone H3 and H4 proteins is associated with a relaxation of the chromatin (openness). As DNA is negatively charged, the addition of acetyl groups (also negative), loosen the nucleosome association, facilitating gene expression. Histone acetylation is carried out by histone acetyltransferases and the deacetylation by histone deacetylases (HATs and HDACs, respectively, Fig. 2.3e; Berger 2007). In Arabidopsis, two HDACs in particular, HDA19 and HDA6, have been linked with plant defence. Both of them are induced by necrotrophs and/or JA-related signals (for example, wounding; Zhou et al. 2005), suggesting some overlapping functions. hdc19 mutants show increased susceptibility to necrotrophic pathogens and an inability to induce JA-related genes. This phenotype does not seem to be related with direct changes in the PTMs at the level of the JA-related genes, but by the hormonal crosstalk with the SA hormonal pathway (Choi et al. 2012; Koornneef et al. 2008). HDA19 seems to play a key role in the maintenance of the silent basal state of the SA-related genes. At the basal state, HDA19 seems to inhibit the acetylation of histone proteins at the PATHOGENESIS-RELATED1 and 2 (PR1 and PR2) defence gene loci (genes considered marker genes for the SA hormonal pathway). In fact, hda19 mutants show enhanced expression of those genes and hyper-resistance against some biotrophic pathogens (even when originally there were contradictory results at this respect; Choi et al. 2012; Kim et al. 2008). Other histone deacetylases involved in plant defence belong to the SIR2 protein family. In Arabidopsis, AtSRT2 controls the basal repression of the SA-related defences (Wang et al. 2010). Thus, histone deacetylations seems to cause a general repression at the level of different SA-associated defence genes. However, some recent studies have described how during PTI responses, the phosphorylation cascade induced by MAPKs proteins can lead to the specific activation of the HDAC, HD2B. HD2B would deacetylate genic regions, inhibiting their expression and therefore contributing to the gene expression reprogramming during defence processes (Latrasse et al. 2017). Lastly, nitric oxide has been proposed as a repressor of the histone deacetylation during plant defence processes, mediating the hyperacetylation and contributing to the induction of defence-related genes (Mengel et al. 2017; Ramirez-Prado et al. 2018).
2.3.3.2 Methylation
As for acetylation, methylation of different residues of the histone proteins has been proven to play a crucial role in plant defence (De-La-Peña et al. 2012; Ramirez-Prado et al. 2018). It mainly occurs in lysine and arginine residues of histones H3 and H4. For each residue, from one to three methyl groups can be added (Bannister and Kouzarides 2005; Kouzarides 2007). These forms of histone methylation can differentially impact on the chromatin structure. Unlike the case of acetylation that usually involves chromatin relaxation, methylation as a PTM of histone proteins has been linked with both inhibition and priming of gene expression (Fig. 2.3f). The enzymes involved in histone de/methylation are very specific. The most notable examples related to plant defence come from the analysis of methyltransferases. This is the case, for instance, of ARABIDOPSIS HOMOLOG OF TRITHORAX (ATX1), which trimethylates the lysine 4 of histone 3 (H3K4me3) and positively regulates the expression of WRKY70, a transcriptional factor involved in SA hormonal pathway and postulated to be crucial to the SA-JA hormonal crosstalk (Alvarez-Venegas et al. 2007). LAZARUS2 (LAZ2) is another histone methyltransferase involved in trimethylation of lysine 36 of histone 3 (H3K36me3), which is required to activate an R gene involved in ETI responses against Pst (Palma et al. 2010). Recently, roles for demethylases in defence have also been reported. This is the case of Jumonji C demethylases. For example, the H3K9 demethylase JMJ27 is induced during bacterial infection, required for the resistance to the pathogen and the correct expression of defence-related genes (Dutta et al. 2017). A peculiar case is the demethylation of lysine 9 of histone 3 (H3K9me2). H3K9me2 has been traditionally considered as a repressive chromatin mark of TEs. Surprisingly, it has been reported that the levels of H3K9me2 at the defence gene RPP7 can affect the selection of the polyadenylation site, and thus, the production of a different transcript (Tsuchiya and Eulgem 2013). As pointed out previously, the chromatin environment is determined not by just individual marks, but the appearance of different ones at the same time. Therefore, it is easy to imagine that there is co-regulation of the different PTM pathways. This was evident in the study of the methyltransferases SDG8 and SDG25. Mutants in those proteins are altered in plant defence against necrotrophs, biotrophs, and show differences in methylation of several residues of the histone proteins at the level of some defence-related genes (Berr et al. 2010; Lee et al. 2016). They also display an altered pattern in H2B ubiquitination (Lee et al. 2016), which will be the subject of the next section.
2.3.3.3 Ubiquitination
Ubiquitination of proteins typically refers to the addition of the 76-residue peptide known as ubiquitin by the action of three consecutives enzymes, E1, E2 and E3 (Weake and Workman 2008). In general, those enzymes can add one or more residues of ubiquitin to the target proteins. Many different proteins can be ubiquitinated, but in the specific case of histone proteins, ubiquitination has only been found in the form of a single residue in H2A or H2B, and usually acts as a positive mark for gene expression marking open chromatin (Fig. 2.3g). As with the other PTMs, ubiquitination is reversible. Arabidopsis HUB1 and HUB2, the two RING E3 enzymes, have been reported to be required for plant defence against necrotrophs (Dhawan et al. 2009; Hu et al. 2014) and to play a certain role against biotrophic pathogens (Zou et al. 2014), probably through modifications of the cuticle (Ménard et al. 2014).
2.3.4 DNA Methylation
DNA methylation usually refers to the addition of a methyl group at the fifth carbon of the cytosine residues of the DNA. In addition to cytosine methylation, adenine methylation has also been reported in many different organisms, being a key factor for protecting prokaryotic DNA. However, adenine methylation has so far received little attention in plants (Liang et al. 2018). Cytosine DNA methylation in plants can be found in every sequence context and in different extents, depending on the species (Niederhuth et al. 2016; Takuno et al. 2016). Attending to the nature of the context, both symmetrical and asymmetrical contexts can be considered (Cokus et al. 2008). Symmetrical contexts are CG and CHG (H refers to A, T or C), where both strands of the DNA are methylated. Asymmetrical context is CHH. The maintenance of the DNA methylation pattern is carried out by the METHYLTRANSFERASE1 (MET1) in CG context and CHROMOMETHYLASE2 and 3 (CMT2 and CMT3) in CHG contexts. The maintenance of CHH methylation is mainly performed by an RNA-directed DNA methylation pathway (RdDM), and requires the constant production of siRNAs (Fig. 2.3h; Law and Jacobsen 2010). Probably as a consequence of its origin as defence systems against exogenous nucleic acids, the establishment of a de novo pattern in DNA methylation is primarily controlled by siRNAs and has some homologies with the PTGS mechanisms. The current view of the process in Arabidopsis includes an initiation phase (not included in the figure for simplicity reasons) in which it is likely that over-accumulated RNAs are copied to a double-stranded molecule by RDR6, and then processed by DCL2 and DCL4 into 21–22 nt siRNAs. Probably these siRNAs trigger PTGS by binding to AGO1 or AGO2 proteins. However, in subsequent stages, they are loaded into AGO6, which directs the plant-specific DNA-DEPENDENT RNA POLYMERASE V (Pol V) and the DNA methyltransferase, DOMAINS REARRANGED METHYLTRANSFERASE 2 (DRM2), to the target regions in the genome to induce low levels of DNA methylation (Nuthikattu et al. 2013). After this initiation phase, the second branch of the RdDM pathway is activated (Fig. 2.3h). In this, the other plant-specific DNA-DEPENDENT RNA POLYMERASE IV (Pol IV) is involved in the production of RNA molecules from the targets that after being processed by RDR2, DCL3 and HEN1 will be loaded onto AGO4. The base pairing between the siRNA with Pol V-produced RNA transcripts enables the recruitment of DRM2 for establishment of DNA methylation (Matzke and Mosher 2014). DRM2-dependent CHH methylation requires the constant production of siRNAs, and on-going activity by the Pol IV-RDR2-dependent RdDM pathway. Both the overall level of genome-wide DNA methylation and the pattern of methylation are controlled by a balance between DNA methylation and demethylation processes. Removal of DNA methylation can happen passively during replication, or actively by the action of DNA glycosylase/lyases, of which four have been identified to date in Arabidopsis (Zhu 2009). Among these, REPRESSOR OF SILENCING 1 (ROS1) is predominantly responsible for DNA demethylation in vegetative tissues. The primary functions of DNA methylation are controlling genome stability and gene expression. In general, the epigenetics community tend to associate DNA methylation at the level of promoter regions with repression of gene expression, while the consequences of methylation within gene bodies remain uncertain (Bewick and Schmitz 2017).
The first characterized roles for DNA methylation in plant defence came again from the defence against viruses, with several examples showing DNA methylation of the viral genome for the Geminivirus family (Blevins et al. 2006; Raja et al. 2008). The majority of plant viruses possess RNA genomes, but the Geminiviridae family genome is a single-stranded DNA. The silencing of the viral genome is carried out by the TGS mechanisms of the cell as a defence mechanism against exogenous nucleic acids. Therefore, Geminivirus genomes are silenced by part of the RdDM pathway (Fig. 2.1b). A beautiful example of plant–DNA virus interaction is seen between tomato and the TOMATO YELLOW LEAF CURL CHINA VIRUS. During their co-evolution, tomato plants first developed the ability of defend themselves from viral infection by methylating the viral DNA. Some virulent strains of the virus carry what is known as the betasatellite encoding βC1, which is a repressor of silencing used as an effector (Yang et al. 2011). However, resistant strains of tomato plants have developed the ability to polyubiquitinate βC1 to mediate its degradation via the proteasome. Nowadays, a range of different viral effectors acting as repressors of transcriptional silencing are known (Rodríguez-Negrete et al. 2013; Wang et al. 2014). Although the TGS assigned to the defence against Geminiviruses was thought to act at the level of DNA methylation of the viral genome, recent studies found that the role of the RdDM pathway in silencing the viral genome is performed by triggering H3K9 methylation (Jackel et al. 2016). This reflects once again the crosstalk between different TGS mechanisms.
As in the case of the PTGS mechanisms, plants took advantage of the TGS defence system to control different endogenous sequences. In fact, the activation of antiviral defences has direct consequences in endogenous sequences (Castillo-González et al. 2015; Coursey et al. 2018), as the most important role of DNA methylation is controlling TE repression (considered invasive DNAs). The changes in chromatin caused by TE silencing can also modify the expression of some plant genes, and in our case of interest, defence-related genes. There has been increasing evidence for a role of the DNA methylation and demethylation machineries in controlling plant defence. On the one hand, plants trigger changes in DNA methylation during pathogen attack (Dowen et al. 2012; Pavet et al. 2006; Yu et al. 2013). In general terms, an active DNA demethylation process in response to infections of Pst and the application of PAMPs such as flagellin has been observed. This demethylation seems to be a consequence of the repression of some RdDM components and the active removal of methyl-cytosines by the protein ROS1 (Fig. 2.3h; Dowen et al. 2012; Yu et al. 2013). On the other hand, Arabidopsis mutants impeded in DNA methylation (such as met1, drd1, cmt3 and mutants defective in Pol V) have been reported to show increased resistance to biotrophic pathogens like Pst and Hpa (Dowen et al. 2012; López et al. 2011; Yu et al. 2013). Correspondingly, with the SA-JA hormonal crosstalk, those same mutants show hyper-susceptibility against necrotrophic pathogens like Plectosphaerella cucumerina and B. cinerea (López et al. 2011; López Sánchez et al. 2016). Also in accordance, mutants in ROS1 protein, which cannot actively demethylate the DNA, show the opposite phenotype (hyper-susceptibility against biotrophs and enhanced resistance against necrotrophs). The exact mechanisms by which DNA methylation controls plant defences are not known and even when transcriptomic analysis of the mutants during infection point to changes in several genes, only in very limited cases, cis-regulation has been demonstrated (Le et al. 2014; Yu et al. 2013). In just a few specific cases, differentially methylated regions have been directly associated with the transcriptional control defence-related genes, for instance, being TEs located at the promoter regions of defence genes (Yu et al. 2013). Alternative trans-regulatory mechanisms have been proposed, which will need further research in the future. Importantly, even when mutants defective in DNA methylation are more resistant against biotrophic pathogens and they show a better induction of the SA-related defence genes, those mutants do not display constitutive expression of those genes (López et al. 2011; López Sánchez et al. 2016). These and other evidences involving chromatin states and the requirement of DNA methylation machinery in transgenerational phenomena have been crucial in determining their role in priming of induced defences and memory of stress, which will be assessed below.
2.3.5 The State of the Field, from Arabidopsis to Other Species
In line with the case of the PTGS mechanisms, there are not many studies carried out in non-model organisms. Here, we will introduce some of the works reported until now. The BRHIS1 chromatin remodeller from rice is a nice example. It has been demonstrated to repress defence-related genes and maintain them in the basal state. As part of the plant response to fungal pathogens/priming agents, plants repress BRHIS1, which would in turn favour the induction of the defence genes (Li et al. 2015). A few more examples have been reported of posttranslational modification of histone proteins. Also in rice, there has been nice work in the rice interaction with the pathogen Magnaporthe oryzae. The over-expression of the histone deacetylase HDT701 confers to the rice plants hyper-susceptibility to M. oryzae and Xanthomonas oryzae pv. oryzae (Xoo). Thus, HDT701 has been proposed as defence repressor, probably targeted by effectors, as its induction has been detected during infections (Ding et al. 2012a). There is also an elegant work in rice studying the Jumonji C histone demethylases. During Xoo infection, rice plants have been reported to induce the expression of 15 JmjC proteins (Hou et al. 2015). Among these, JMJ704 and JMJ705 have been demonstrated to play a key role for plant defence. Accordingly, a jmj704 mutant is more susceptible, and JMJ705 over-expression lines more resistant, to Xoo (Hou et al. 2015; Li et al. 2013). Both seem to play a role in repressing defence genes during basal conditions and releasing their expression during defence. On the one hand, JMJ704 seems to be important in maintaining low levels of the positive mark H3K4me2/3 in the defence genes during basal conditions (Hou et al. 2015). On the other hand, JMJ705 seems to induce the removal of negative marks such as H3K27me2/3 during infections (Li et al. 2013). Apart from rice, following a similar strategy to the genome-wide analysis of sRNAs, a whole-genome analysis during rust infection in common bean reported global changes in histone methylation and acetylation (Ayyappan et al. 2015). In addition, and built on the basis of the work in Arabidopsis, the histone ubiquitination pathway has also been studied in other species such as tomato (Zhang et al. 2015). In this study, the tomato homologs of the HUB1/2 proteins: SIHUB1 and SIHUB2 were identified. The authors demonstrated that SIHUB1/2 are required for defence against B. cinerea and seem to play a role in the crosstalk of the SA-JA hormonal pathways, probably by a combination of the cuticle properties and the priming of defence genes (Zhang et al. 2015). Lastly, consistent with the role of DNA methylation in response to biotrophic pathogens, treatments with 5-azadeoxycytidine in rice seedlings have been reported to show resistance to Xoo, a phenotype which correlated with the demethylation of specific regions, including a defence gene Xa21G (Akimoto et al. 2007). Again, even though the role of the different TGS mechanisms has been proved to be crucial for plant defence, more work is needed in order to apply the fundamental knowledge generated in models to crop protection programs.
2.4 Epigenetics Is Involved in the Memory of the Stress: Priming
In addition to the roles of small RNAs and chromatin remodelling in the regulation of immediate defence responses and defence-related gene expression, epigenetic processes also play key roles in longer-term defence priming. Priming refers to the immunological memory that can develop following stress exposure, such that responses to future stresses are more effective. The best-studied example of priming in relation to biotic stress is systemic acquired resistance (SAR). As well as transient up-regulation of defence genes in systemic leaves (which have not suffered the infection), SAR typically includes a priming element, such that defence responses are stronger and more rapidly induced in response to a secondary infection for up to several weeks following an initial pathogen infection (Klessig et al. 2018). More recently, evidence has accumulated that under some circumstances, longer-lasting priming memory can be established, which is then inherited by one or more future generations.
2.4.1 Short-Term Priming Memory
2.4.1.1 Changes in Defence Response Signalling Components
Several mechanisms have been proposed that could potentially generate the memory of stress that is required for priming. Short-term memory could be generated relatively easily by changes in the quantity or activity of signalling components required for the regulation of ETI and PTI (Conrath et al. 2015). For example, systemic leaves of Arabidopsis undergoing SAR display elevated levels of unphosphorylated (therefore inactive) mitogen-activated protein kinases MPK3 and MPK6. When inoculated with P. syringae, these primed leaves exhibit higher levels of MPK3/6 activity than non-primed leaves, due to their faster activation, as they are already synthetized (Beckers et al. 2009). The accumulation of transcription factors necessary for defence gene expression may be another similar mechanism for priming. Van der Ent et al. (2009) identified panels of Arabidopsis transcription factors that were up-regulated upon priming by either induced systemic resistance (ISR) triggered by Pseudomonas fluorescens or by root drenching with β-aminobutyric acid (BABA), a well-known chemical inducer of priming. They suggested that the different groups of transcription factors responsive to each of these priming treatments not only provide a mechanism for priming, but can also act as markers for different priming responses. The up-regulation of pattern recognition receptors (PRRs) involved in recognition of biotic attackers has also been suggested as a mechanism for priming (Tateda et al. 2014).
2.4.1.2 Chromatin Remodelling
Aside from the production of additional signalling molecules, the other main area that has received attention is epigenetic mechanisms for encoding stress memories.
As well as being essential for immediate, short-term transcriptional responses, chromatin remodelling via histone and DNA modifications also has the potential for conferring stable patterns of gene expression. Indeed, during development, epigenetic mechanisms are central to changes in gene expression associated with cell-type specialization (Heard and Martienssen 2014). As described above, regulation of defence gene expression involves various histone and DNA modifications which ultimately make genes more accessible to the transcriptional machinery. After a transient burst of stress-induced transcription, the chromatin of a defence-related gene may revert to the basal state, in which case it would be expected to show the same response characteristics to any subsequent experience of stress. Alternatively, if the reversion was only partial, then it might be possible that subsequent access of the transcriptional machinery would be less restricted. This would represent a primed state, in which chromatin modifications, brought about by previous transcriptional activation, leave a memory imprint. Primed genes might therefore be expected to reside on more open chromatin associated with altered levels of key histone and DNA modifications. To date, evidence from several different biotic and abiotic stress response systems has identified a range of histone marks and DNA methylation patterns, but in particular, reduced nucleosome occupancy and increased H3K4me3 are emerging as especially common features of primed stress-related genes.
2.4.1.2.1 Histone Modifications
Several authors have identified H3K4 hypermethylation associated with transcriptional memory/priming, most notably the trimethylation state. Key work linking histone modifications and defence priming came from Jaskiewicz et al. (2011), who demonstrated that local inoculation with P. syringae or treatment with benzothiadiazole (BTH) led to increases in H3K4me2 and H3K4me3, along with increased acetylation at a number of histone H3 and H4 lysine positions in the promoters of several WRKY genes. Histone H3K4me2/3 methylation is generally associated with a permissive transcriptional chromatin state (Berger 2007). Importantly, the increase in H3K4 methylation following BTH treatment was not associated with any immediate change in gene expression, but the affected WRKY genes exhibited augmented expression in response to a secondary stimulus. Furthermore, H3K4me3 hypermethylation was not observed in the SAR-deficient npr1-1 mutant, whereas constitutively primed cpr1 and sni1 mutants showed constitutive high levels of H3K4me3 in the WRKY gene promoters. Similarly, mutants defective RNA polymerase V, which is required for initiation of DNA methylation in the RdDM pathway, also demonstrate constitutive priming of SA-dependent defence genes, and exhibit enhanced H3K4me3 at defence genes (López et al. 2011). Priming responses following application of BABA in common bean were also linked with elevated H3K4me3 states at defence gene promoters (Martínez-Aguilar et al. 2016). Importantly, BABA treatment resulted in enhanced H3K4 trimethylation without any change in gene expression. The subsequent transcriptional activation of genes with this modification was primed, being higher in BABA-treated plants than in control plants, in response to bacterial infection (Martínez-Aguilar et al. 2016).
Other histone modifications may act alongside H3K4 methylation to establish priming. A role for histone acetylation in priming of PTI was identified by Singh et al. (2014a), who showed that priming for bacterial disease resistance was eliminated by mutation of histone acetyltransferase1 (HAC1). Interestingly, the Arabidopsis FLD gene, which encodes a protein homologous to a human lysine-specific demethylase and also associated with histone deacetylase complexes, was identified in a genetic screen for mutants defective in the ability to express SAR (Singh et al. 2013). Mutants in FLD display wild-type levels of basal resistance to Pst, but do not exhibit priming of PR1, WRKY6 and WRKY29 gene expression following secondary inoculation of systemic leaves (Singh et al. 2013, 2014b). H3K4me2 appears to be the major substrate for FLD (Liu et al. 2007), but H3K4me2 methylation was decreased overall rather than increased in the promoters of WRKY6 and WRKY29 following challenge inoculation of an fld mutant (Singh et al. 2014b). Although the data do not identify a clear role for FLD in regulating histone modifications during priming of these genes, the authors suggested it may function as a negative regulator of an alternative histone demethylase that represses H3K4me2 methylation, and therefore priming, of defence genes.
As well as posttranslational modifications of histones, nucleosomes are also regulated by inclusion of variant histone proteins. In particular, H2A.Z has been linked with plant–pathogen resistance responses, although its precise function remains unclear. As noted in Sect. 2.2, March-Díaz et al. (2008) found that mutants defective in the SWR1 chromatin remodelling complex, which is responsible for substitution of canonical H2A with H2A.Z, are resistant to P. syringae and over-express a suite of SAR-regulated genes. More recently, different roles were identified for genes in the SWR1 complex and H2A.Z in SA and JA-mediated basal and effector-triggered immunity, indicating a complex interaction between H2A.Z substitution and immune regulation (Berriri et al. 2016). Overall, while loss of H2A.Z increases basal immunity, it reduces inducible responses. Whether it plays any role in priming of defence genes following induced resistance remains to be tested.
2.4.1.2.2 Nucleosome Occupancy
A second, related, common feature that has been identified in the promoters of primed genes is a more open chromatin configuration. Chromatin assembly factor CAF-1 is required for assembly of nucleosomes on newly replicated DNA, and mutants in CAF-1 subunits display pleiotropic developmental phenotypes (Ramirez-Parra and Gutierrez 2007). One of these phenotypes is constitutive priming of many defence genes. Under normal growth conditions, the fas2-4 mutant (a null allele for one of the CAF-1 subunits) exhibited constitutive expression of SAR-responsive genes. When grown under sterile conditions, these genes were no longer constitutively expressed, but displayed primed expression in response to SA treatment (Mozgová et al. 2015). CAF-1 therefore appears to be involved in repression of the primed state in wild-type plants. In the CAF-1 mutant, chromatin assays identified reduced nucleosome occupancy but increased abundance of H3K4me3 around transcription start sites (TSS) of several SAR genes, and similar chromatin states were observed in the same genes when priming was induced by either SA or BABA treatment (Mozgová et al. 2015). Interestingly, similar profiles were also detected in the promoters of genes exhibiting priming memory following drought stress. In this work, promoters of primed genes possessed stalled RNA polymerase II (PolII) and H3K4me3 hypermethylation (Ding et al. 2012b). Finally, genome-wide profiling of nucleosome positioning in response to SA treatment also identified reduced nucleosome occupancy at the TSS of SA-responsive genes (especially those regulated by NPR1), while SA-repressed genes showed nucleosomal enrichment (Singh et al. 2015).
2.4.1.2.3 DNA Methylation
Changes in DNA methylation are another obvious candidate mechanism for long-term stress memory. As discussed above, DNA hypomethylation has been detected following infection of Arabidopsis with P. syringae in several studies (Dowen et al. 2012; Pavet et al. 2006; Yu et al. 2013). This may play a functional role in immediate induced resistance responses, since hypomethylated loci centred around transposable elements are enriched in defence genes (Dowen et al. 2012). Yu et al. (2013) also identified demethylation of TEs near defence genes in response to treatment with the PTI-eliciting FLG22 peptide. This response was dependent on the DNA glycosylase ROS1. These stress responsive changes in methylation are consistent with the phenotypes of hypomethylated DNA methylation mutants, which are typically more resistant to infection (Dowen et al. 2012; Le et al. 2014; López et al. 2011; López Sánchez et al. 2016; Luna et al. 2012; Luna and Ton 2012). Whether mutants that suffer from extensive genome-wide hypomethylation genuinely reflect what happens following infection is open to question, but plants with more restricted regions of hypomethylation, such as in the so-called epiRIL (epigenetic recombinant inbred) lines generated by back-crossing hypomethylated mutants to wild-type Arabidopsis, can also exhibit altered responses to defence hormones and variations in resistance to pathogens (Latzel et al. 2012). Importantly, increased resistance in some DNA methylation mutants has been attributed to priming rather than constitutive SA-responsive gene expression (López et al. 2011; López Sánchez et al. 2016). Since DNA methylation is readily inherited during mitotic cell divisions, it provides a potential mechanism for long-term priming memory that can extend even into cells and tissues not present during the initiating stress.
2.4.2 Long-Term Priming Memory
Responses such as SAR are well documented to persist over several weeks, but in recent years, examples of defence priming that persist for much longer periods of plant development have also emerged. For example, seed treatments with elicitors including JA, chitosan and BABA resulted in defence priming that persists for many weeks (Haas et al. 2018; Strapasson et al. 2014; Worrall et al. 2012), while seedling root drenches with BABA prime long-lasting disease resistance in Arabidopsis and tomato (Luna et al. 2014; Wilkinson et al. 2018). Not only that, priming can extend from one generation to the next. One early report hinting at what is now referred to as transgenerational acquired resistance (TAR) in the case of SA-dependent disease resistance, or transgenerational immune priming (TGIP) more generally, suggested that tobacco mosaic virus (TMV) infection enhanced resistance in progeny of tobacco (Nicotiana tabacum; Roberts 1983). Other studies found that Brassica spp. suffering biotic stress produced seeds containing higher concentrations of glucosinolates compared with non-infested control plants (Lammerink et al. 1984; Shattuck 1993). Another series of papers demonstrated that insect herbivory on wild radish (Raphanus raphanistrum) enhanced resistance in seedlings of progeny plants (Agrawal 2001, 2002; Agrawal et al. 1999). The increase in resistance observed in these studies was transient and no mechanism was identified. Although intriguing in the context of the potential ecological benefits of TGIP, the examples described above could also be explained by simple maternal effects—the provisioning of seeds with altered resources from the mother plant in response to stress. While widely recognized as ecologically important, maternal effects are distinct from bone fide transgenerational inheritance, which enables the offspring generation to express phenotypes independently of any non-genetic contribution from the parental plants. Such transgenerational inheritance is most likely to be epigenetically encoded.
2.4.2.1 Transgenerational Immune Priming
Several examples have emerged over recent years which provide much stronger evidence for true transgenerational inheritance of biotic stress priming. As well as following pest and pathogen attack, priming can be induced by various chemical agents. One of the best studied among these is β-aminobutyric acid (BABA; Cohen et al. 2016). As well as within-generation priming, it was recently found that the progeny of plants primed either by BABA treatment or infection with avirulent P. syringae bacteria exhibited TAR. Offspring of treated plants were more resistant to infection by both Pst and Hpa because of primed SA-dependent gene expression (Slaughter et al. 2012). Intriguingly, offspring of BABA-primed parents were more responsive to BABA treatment than offspring of control plants, suggesting a ‘primed-to-be-primed’ phenotype. Similarly, treatment of barley (Hordeum vulgare) with the commercial resistance-inducing agent acibenzolar-S-methyl, or with saccharin, resulted in TAR against leaf blotch disease caused by the fungal pathogen, Rhynchosporium commune (Walters and Paterson 2012).
Luna et al. (2012) also reported TAR in Arabidopsis following repeated inoculations with virulent P. syringae. A key aspect of this work was that the authors were not only able to identify priming in the immediate offspring generation, but it could also be detected in the grandchildren of the infected plants. This demonstrates that priming memory can be inherited over at least one stress-free generation and eliminates the possibility that maternal effects alone are responsible for the increased resistance. Subsequent work from the same group now shows that priming can be detected, albeit weakly, even after two stress-free generations (Stassen et al. 2018), indicating that the phenomenon must be epigenetically regulated. In evolutionary terms, the gradual loss of priming after the initial stress episode would be expected in order to avoid excessive costs of priming and could readily be achieved through reversible epigenetic changes. In parallel with this work on transgenerational disease resistance, similar responses to herbivory were reported. terHorst and Lau (2012) found that both herbivore resistance and reproductive fitness were affected by parental exposure to insect herbivory in a field experiment with Lotus wrangelianus. Moreover, herbivory also resulted in JA-dependent transgenerational priming of defence against insects in both Arabidopsis and tomato (Rasmann et al. 2012).
2.4.2.2 Mechanisms for TGIP
The most likely mechanisms for true transgenerational priming effects are epigenetic. Similar to within-generation priming responses such as SAR, histone modifications could be detected in the promoters of the SA-regulated genes, PR1, WRKY6 and WRKY53 in plants derived from P. syringae-infected parents (Luna et al. 2012). Such histone modifications may well contribute to primed defence gene expression in TAR, but there is still wide debate over the roles that histone modifications might play in epigenetic inheritance between generations (Heard and Martienssen 2014). Much better understood is the ability of DNA methylation to be meiotically inherited, and DNA methylation has therefore been suggested as the more likely mechanism for encoding transgenerational epigenetic memory (Quadrana and Colot 2016). It has become increasingly clear over recent years that modifications to the DNA methylome can be maintained through the plant’s lifespan and into subsequent generation(s), and can generate heritable phenotypic changes (Bossdorf et al. 2010; Johannes et al. 2009; Mathieu et al. 2007; Verhoeven et al. 2010). Accordingly, several studies have found that mutants affected in various regulatory mechanisms controlling DNA methylation show altered biotic stress resistance, and/or fail to establish transgenerational priming (López Sánchez et al. 2016; Luna et al. 2012; Luna and Ton 2012; Rasmann et al. 2012). These two modes of epigenetic regulation are not mutually exclusive, since histone modifications and DNA methylation are somewhat inter-dependent (Heard and Martienssen 2014).
The strongest current candidate for generating methylation-dependent stress memory is RNA-dependent DNA methylation (RdDM). As noted above, several studies have shown that mutations in genes involved in the RdDM pathway can have a direct impact on defence responses (López et al. 2011; Le et al. 2014; López Sánchez et al. 2016; Luna and Ton 2012). RdDM-mediated methylation is initiated by the generation of 21–24 nt siRNAs by DICER-like proteins, and transgenerational priming responses to both biotic and abiotic stress have been found to require the production of siRNAs. Furthermore, offspring priming phenotypes triggered by different abiotic stresses in Arabidopsis required DCL2 and/or DCL3, which are responsible for the production of 21/22 and 24 nt siRNAs, respectively (Boyko et al. 2010). Similarly, for biotic stress, the dcl3–1 mutant failed to establish TAR against biotrophic pathogens (Luna and Ton 2012) and a dcl2/dcl3/dcl4 triple mutant failed to establish transgenerational priming against herbivores (Rasmann et al. 2012). siRNAs are able to move systemically throughout the plant (Dunoyer et al. 2010; Lewsey et al. 2016; Molnar et al. 2010), and can therefore be viewed as candidates for long range priming signals both within and between generations, since reprogramming of methylation in germ line cells by siRNAs would allow inheritance by offspring tissues. Interestingly, sRNA populations have been reported to be significantly influenced by parental/grandparental environmental stress in both Brassica rapa and the dandelion, Taraxacum officinale (Bilichak et al. 2015; Morgado et al. 2017). Since pathogen infection triggers genome-wide methylation changes associated with reactivation of transposon sequences, generation of mobile siRNAs and subsequent RdDM presents a feasible mechanism for maintaining stress memories.
Active DNA demethylation also appears to be required for TAR. The DNA glycosylase ROS1 plays a major role in global demethylation, and was found to be essential for transgenerational memory in offspring of Pst-infected Arabidopsis (López Sánchez et al. 2016). Interestingly, the same authors found that ROS1 was not required for within-generation SAR. The nature of the changes imposed by ROS1-dependent demethylation and RdDM in response to biotic stress and the mechanisms by which they impact on defence phenotypes remain to be elucidated. Although methylome profiling of different generations of plants expressing TAR identified differentially methylated sites that correlate with ancestral stress experiences (Stassen et al. 2018), the analysis of these sites does not yet provide a clear insight into the mechanism of transgenerational priming.
2.4.2.3 Re-Setting Epigenetic Priming Memory
Epigenetically mediated transgenerational defence priming provides a novel system by which plants can display phenotypic plasticity in response to environmental stress, providing enhanced evolutionary fitness when parental environments are good predictors of offspring environments. To have evolved, the benefits of transgenerational phenotypic plasticity must outweigh any costs. Because plasticity is epigenetically mediated, it is readily reversible, such that when stress is not present, defence reverts to basal levels, thus minimizing costs. Costs would also be minimized when the signals initiating priming are good predictors of future stress. In other words, long-lasting priming would be expected only under strong, consistent stress, and should decay in the absence of stress. These predictions appear to hold true in the limited instances where these ideas have been tested. Singh et al. (2014a) found that repeated, but not single, mild stress exposures provided short-lived priming of biotic stress resistance but was not sufficient for long-term priming. TAR induced by P. syringae infection of Arabidopsis was maintained at high levels when successive generations of plants were inoculated, but was gradually lost when only a single generation suffered disease (Stassen et al. 2018). The costs of transgenerational priming remain relatively poorly explored, but one clear cost is seen in the antagonism between the major SA- and JA-dependent defence pathways. TAR against biotrophic pathogens was associated with transgenerational increased susceptibility to necrotrophic pathogens (Luna et al. 2012), while transgenerational priming of herbivore resistance caused increased susceptibility to P. syringae infection (Singh et al. 2017). The storage of epigenetic marks reflecting stress memories of previous generations could therefore have deleterious impacts when offspring experience different stresses than their parents (Crisp et al. 2016; Iwasaki and Paszkowski 2014). Avoidance of such maladaptive memories requires a system for re-setting of epigenetic stress memories that can balance the forces imposing them. As an example to keep in mind, the Arabidopsis FLC locus, involved in vernalization, is epigenetically silenced in response to low temperature. The low temperature memory is re-set each generation, meaning that it is the experience of individual plants, not their ancestors, that determines flowering time. Mutation of the ELF3 gene, a histone H3K27me3 demethylase, causes a failure of the ability to re-set the low temperature signal, meaning that offspring flower early regardless of environmental conditions (Crevillén et al. 2014). Other re-setting systems have also been identified. Two chromatin regulators, DECREASE IN DNA METHYLATION1 (DDM1) and MORPHEUS’ MOLECULE1 (MOM1), prevent the transmission of memories of heat stress exposure from parents to their progeny. Genome-wide transcriptional signatures induced by stress were found in the subsequent generation in ddm1/mom1 double mutants, but not wild-type plants (Iwasaki and Paszkowski 2014). More recently, MOM1 has also been identified as an epigenetic regulator of the expression of pattern recognition receptor (PRR) and nucleotide-binding leucine-rich repeat (NLR) genes in Arabidopsis. PRRs and NLRs act as receptors for activation of PTI and ETI, respectively, and therefore changes in their expression could potentially be a mechanism for defence priming. MOM1 indirectly regulates PRR/NLR gene expression via an RdDM-dependent pathway for transposon silencing. Mutants deficient in MOM1 displayed higher expression of PRRs/NLRs and were more resistant to bacterial infection (Cambiagno et al. 2018), consistent with the idea that MOM1 might function antagonistically with defence priming. Such mechanisms of chromatin re-setting could prevent or constrain transgenerational priming of defence to optimize trade-offs between the costs and benefits of priming.
2.5 Potential Application of Epigenetics
World population is growing at a speed that places food security at risk at a global level (FAO 2009a). In this context, increasing crop yields by reducing losses caused by pests and diseases would appear to be essential. At the same time, the use of pesticides, which have had a major influence on protecting yields in the past, has been demonstrated to considerably contribute to soil degradation, ecosystem disturbance, climate change and even to have a negative impact on human health. Given this scenario, global entities are taking action to promote the development and implementation of alternative crop protection technologies and strategies (European Academies Science Advisory Council and Deutsche Akademie der Naturforscher Leopoldina 2014). Although when its impacts were first recognized, epigenetics represented an inconvenience in biotechnology, for example, because of unintended gene silencing effects (Napoli et al. 1990; van der Krol et al. 1990), today, the field of epigenetics presents a clear example of the importance of fundamental research in future applications (Connor 2002). On the one hand, advances in the understanding of epigenetic mechanisms have already brought novel tools for biotechnologists to analyse, control and tailor gene expression. On the other hand, the possibility to produce new stable (even heritable) phenotypes in the absence of genetic changes opens doors to new approaches avoiding transgenesis, which has important implications in view of the current regulatory framework around genetically modified organisms (GMOs) and their public perception. Moreover, boosting the plants’ natural immune system has been suggested as the safest approach to improve crop yields while minimizing environmental impacts (Rapicavoli 2015; Dewen et al. 2017; Kothari and Patel 2004; Quintana-Rodriguez et al. 2018).
Bearing in mind that the origin of the epigenetic machinery seems to be linked to ancestral defence mechanisms, we believe the application of epigenetics to crop protection could provide significant breakthroughs in the development of future integrated pest management programs (Stenberg 2017). Here, we discuss some possible applications to translate the new knowledge of epigenetics to first, keep broadening our understanding of natural phenomena, and second, as a tool to a new generation of plant biotechnologists and breeders in the field of crop protection.
2.5.1 As a Tool in Research
Advances in the understanding of epigenetic mechanisms have significant potential to impact the future research of plant–pathogen interactions. Up to now, such understanding has enabled the development of various techniques to characterize and even manipulate the epigenome. Nowadays, we can detect small RNAs by northern blotting, hybridization with probes for detection and subtraction, PCR/qPCR, etc. (Boccara et al. 2017; Li and Zamore 2018; Ro and Yan 2010; Urbanek et al. 2015). Using the new advances in bioinformatics we can also sequence and map sRNAs in different genomes, compare them in different species or even design specific systems to knock them down. Chromatin changes can be analysed by different techniques such as chromatin immunoprecipitation (ChIP; Furey 2012), DNase I treatments (Cockerill 2011), FAIRE (Simon et al. 2012) and chromatin conformation capture (3C; Dekker et al. 2002). There have also been enormous advances in the analysis of DNA methylation as a result of the development of bisulphite sequencing, which enables the detection of changes in DNA methylation at the level of individual cytosine residues across the whole genome. Other techniques, like ChIP analysis using antibodies against methyl-cytosines (MeDIP-ChIP), allow for the detection of regional differences in DNA methylation (Cortijo et al. 2014b). MeDIP-ChIP has lower resolution than bisulphite sequencing, but it can be used as an affordable method to detect regional differences in the whole genome, or, coupled to PCR, for the detection of differential methylation in discrete regions, which can be helpful in species for which the full genome cannot be sequenced. Another important set of tools is based on the differential activity of restriction enzymes in methylated and non-methylated DNA. This property of the endonucleases can be used for the detection of the whole-genome methylation level by southern blotting, in which no or minimum information about the DNA sequence is required. It can also be used in the design of chop PCR markers, which allow us to detect changes in DNA methylation at specific positions quickly and economically.
Currently, we can not only detect, but also modify the levels of DNA methylation. For example, following the identification of the proteins of the DNA methylation machinery, mutant plants are available with different and relatively stable patterns of DNA methylation that can be used for research and applied purposes. There are also several chemical reagents known that alter the epigenome. For example, 5-azacytidine (5-azaC), zebularine and sulfamethazine reduce the general levels of DNA methylation (Jones 1985; Zhang et al. 2012; Zhou et al. 2002). More targeted manipulation of the epigenome—epigenome editing—is now becoming possible as a result of new advances in molecular biology. By fusing proteins with DNA methylation/demethylation activities to sequence-specific DNA binding proteins such as zinc finger nucleases or the CRISPR/dCas 9 system, it is possible to target methylation/demethylation to specific target sites (Gallego-Bartolomé et al. 2018). Hence, our knowledge of epigenetics made it possible to develop these techniques, and we are now in a position to use them to speed up progress in understanding mechanisms of plant defence.
Now, we can also understand phenotypes that were baffling for decades, as they involved changes in gene expression in the absence of any change in DNA sequence. The use of the aforementioned techniques is accelerating the discovery of the epigenetic mechanisms controlling plant defence. It has opened the door to the identification of new natural epialleles, as well as the understanding of part of the observed natural variation found between species and ecotypes (Cubas et al. 1999; Niederhuth et al. 2016; Richards 2011; Turck and Coupland 2014; Vaughn et al. 2007; Zhai et al. 2008). It has also contributed to unravelling the evolution of the plant immune system. Since the plant immune system has been demonstrated to be controlled by epigenetic mechanisms at almost all levels, the study of the epigenotype will be essential for the future understanding of plant defences and the translation of this new knowledge into crop protection strategies.
2.5.2 As a Tool in Biotechnology
Crop losses due to pests and diseases must be reduced to a minimum to ensure food security and sustainability in the coming decades. World population is expected to grow up to 9.7 billion by 2050. Thus, improving crop yields is a priority action for the world’s public entities in order to produce more food on less land, avoiding the over-exploitation of natural ecosystems and the decline in biodiversity (FAO 2009b). At the same time, global climate change is introducing abiotic stress variables that promote outbreaks of different plant pathogens. To date, the use of pesticides has been the most successful agronomic strategy to protect crops from infections. Unfortunately, pests tend to easily acquire resistance to chemical pesticides. Moreover, the use of pesticides contributes to the disruption of natural ecosystems, degrading soils and contaminating water sources. It has also been demonstrated to be potentially toxic or carcinogenic, at the very least for workers who manipulate the products (Aktar et al. 2009). This provides additional impetus in the search for alternative strategies (European Commission 2009). Moreover, while biotechnologists focused their efforts on genetic engineering of plants to introduce new traits, there has been a strong public pressure against transgenesis and other forms of genetically modified organisms, especially in Europe (Collinge et al. 2010). With this in mind, some experts in the field agree that plant vaccination and priming (or plant immunization) is one of the most promising approaches in pursuit of designing safer and more sustainable crop protection programs (Kothari and Patel 2004; Rapicavoli 2015; Dewen et al. 2017; Quintana-Rodriguez et al. 2018).
As this chapter has described, the plant immune system and priming processes are epigenetically controlled at different levels. Given that it facilitates phenotypic changes while avoiding transgenesis, epigenetics appears to be an attractive tool/target for breeders and agronomical biotechnologists hoping for a new and more sustainable green revolution.
2.5.2.1 PDR: Pathogen-Derived Resistance
The first examples of the exploitation of epigenetics for plant protection were seen in the form of pathogen-derived resistance (PDR) or ‘plant vaccination’. It had been known for some time that infections with some viruses triggered cross-protection against similar viruses (Hamilton 1980). The scientific basis for this observation was unknown for a long time, but resistance was assumed to be triggered by the production of viral components. Thus, PDR was proposed as a biotechnology strategy for plants in which pathogen-specific components could be altered and over-expressed in a non-functional form, or at the wrong developmental stage, to somehow compete with the functional proteins produced by the virus (Sanford and Johnston 1985). Following the suggestion of this concept, there were a few successful cases of protein-mediated resistance (based on viral coat protein expression) that were probably unrelated to epigenetic mechanisms (Abel et al. 1986; Powell et al. 1990). In all cases, the mechanism involved the production of transgenic plants over-expressing genes (or modified versions of them) from the target pathogen. Surprisingly, in many subsequent cases, this approach was discovered to not be always associated with the levels of proteins of the transgene, but rather, RNA expression. In fact, instead of protein-mediated resistance, RNA-mediated resistance appeared to provide near complete immunity against, at least, the target virus. The basis of such RNA-mediated resistance remained obscure until it was linked with the co-suppression phenomenon observed in transgenic petunia plants (Lindbo et al. 1993). It then became clear that in most viral PDR cases, protection was due to the activation of gene silencing mechanisms against the pathogenic virus. With the discovery of the epigenetic mechanisms, PDR mediated by RNA expression has been improved (Prins 2003), optimizing the silencing of the pathogen. RNA silencing-mediated PDR is also now known as host-induced gene silencing (HIGS), which involves plant transformation with constructs expressing efficient siRNAs targeting different pathogen elements.
This strategy has been demonstrated to be effective not just against viruses in model organisms, but also other groups of pathogens and herbivores and in different crop species (Huang et al. 2016). Since its first uses in tobacco and Arabidopsis, PDR has been successfully applied to introduce virus resistance to crops such as papaya (against the papaya ringspot virus, PSRV; Gonsalves 1998; Jia et al. 2017), squash (against the Squash leaf curl virus; Taha et al. 2016), tomato (Schwind et al. 2009), cotton, rice, maize, corn, wheat and barley (Duan et al. 2012; Huang et al. 2016; Koch and Kogel 2014). Beyond its uses in plant-viral protection, it has been adapted to protect plants against many other different pest and diseases such as bacteria, fungi (Nowara et al. 2010; Nunes and Dean 2012), parasitic plants, nematodes or insects (Koch and Kogel 2014). Perhaps most remarkable is the work performed in plant–insect interactions, where HIGS-mediated resistance has been demonstrated following the ingestion of plant-produced siRNAs during insect feeding (Baum et al. 2007). Undoubtedly, the latest discoveries involving cross-kingdom sRNA trafficking will help us to better engineer PDR and/or HIGS, optimizing this kind of plant vaccination (Wang et al. 2016, 2017).
In some cases, PDR confers almost complete resistance. However, it also poses some problems. The first issue is that it involves transgenesis, which faces strong social opposition and is under the control of very restrictive legislation (Lucht 2015). On the other hand, alongside the benefits of resistance, there could be some deleterious consequences in the plant, due to the insertion and over-expression of the constructs. For example, the genomic insertions could disturb genic regions required for specific plant functions in response to environmental changes, which is difficult to predict or identify in field trials. A strong activation of silencing mechanisms could also have consequences for the natural roles of these pathways during plant development or responses. Finally, the expression of the transgene can be modified by environmental changes, or even silenced due to its high expression (Martelli 2001; Zhao et al. 2014). Therefore, even though its high potential has been demonstrated for several decades, its success usually requires an important investment of time and optimization effort, which has restricted its successful commercial application to a relatively small number of specific cases.
2.5.2.2 Epigenetics Mediated Resistance
Since the role of DNA methylation in plant defence has only emerged relatively recently, commercial approaches that exploit epigenetics have not yet reached the market. However, the possibility to select or engineer plants at the epigenetic level to improve resistance without transgenesis makes this one of the most fashionable current strategies that is being actively pursued. In the same way that breeders have been searching for disease resistance genes in crop wild relatives and land races to develop introgression systems, we now know that a proportion of natural phenotypic variability is generated by the epigenome (Zhang et al. 2013). Given that some DNA methylation patterns appear to be stably inherited through meiosis, an attractive idea is the use of natural epigenetic alleles (epialleles) to provide traits for classical selective plant breeding (Gallusci et al. 2017; Hofmeister et al. 2017; Zhang and Hsieh 2013). In this respect, epialleles impacting on defence phenotypes from different backgrounds could be identified and introgressed (Ji et al. 2015).
There are a number of methods by which such epialleles can be identified. The first is through simple screening of populations of a single genotype for phenotypic variation. Repeated selection of an isogenic population of Brassica napus for individuals with elevated energy use efficiency (EUE) resulted in the isolation of ‘epilines’ with improved EUE and which gave increased yield (Hauben et al. 2009). The improvement in EUE appeared to come from altered DNA methylation profiles rather than genetic changes was inherited for eight generations and was stable over 3-year field trials. Later work used a similar strategy to isolate stable epilines with increased EUE and drought tolerance (Verkest et al. 2015). These lines showed increased expression of abiotic stress-related genes, and elevated H3K4me3 methylation at those genes. As well as simply screening for naturally occurring epigenetic variation, it is possible to create recombinant populations in which individual lines carry substantial portions of the genome with altered methylation patterns. Such lines are known as epigenetic recombinant inbred lines (epiRILs). EpiRILs can be generated from two varieties or ecotypes with similar genetic sequences but different epigenetic landscapes (Kawakatsu et al. 2016; Schmitz et al. 2013). The approach for isolating useful epialleles that provide novel traits assumes that novel methylation patterns carried by an epiRIL can be stably inherited and used for crop breeding. One method for generating novel epigenetic patterns is the generation of epiRILs by crossing mutants defective in DNA methylation machinery. Homozygous mutants develop a different epigenetic landscape (e.g. hyper- or hypomethylation), and following removal of the original genetic mutation by back-crossing, the resulting recombinants would carry a mosaic of different epigenetic marks at different locations across the genome. This has already been applied successfully in Arabidopsis (Cortijo et al. 2014a; Johannes et al. 2009; Reinders et al. 2009), and several groups are currently working in epiRILs in crops like tomato, wheat and rice. Genome-wide epigenetic variability can also be induced by the application of DNA methylation inhibitors like sulfamethazine, 5-azacytidine (5-azaC) and zebularine (Jones 1985; Zhang et al. 2012; Zhou et al. 2002), or by introduction of novel DNA methylase/demethylase enzymes (Hollwey et al. 2017). An interesting alternative is to use natural instances of epigenome reprogramming. For example, plants regenerated from tissue culture often exhibit significant phenotypic variation, despite being clonal and therefore, in principle, identical at the DNA sequence level. This is known as somaclonal variation. Wibowo et al. (2018) recently found that Arabidopsis plants regenerated from root (but not leaf) cells were more susceptible to Pst and Hpa infection than non-regenerated controls. Genome-wide DNA methylation profiles also differed with the origin of regenerated plants and altered methylomes were inherited for at least three generations. Finally, where a priori information exists about desirable epigenetic traits, modifications at specific target loci can be achieved through the use of epigenome editing (Gallego-Bartolomé et al. 2018; Kungulovski and Jeltsch 2016) as described above. Despite the novelty of breeding with epialleles, it should not be forgotten that as for other forms of genetically encoded resistance, systems that result in constitutively elevated defences are likely to bring with them costs in yield in the absence of stress, due to the expense of allocating resources to defence. In nature, such costs are minimized by priming of defence, which could therefore be an attractive trait to target.
2.5.2.3 Epigenetic Priming
With the exception of the epigenome editing at specific sites, the approaches described above rely on natural or introduced random epigenetic variation as a source of natural variation from which to select desired traits, such as pest and disease resistance. However, the phenomenon of transgenerational immune priming described above, suggests that epigenetic resistance can be induced by exposure of plants to biotic stress. The primed state involves sensitization of defence responses following initial triggering of priming, but primed plants do not express higher levels of defence prior to attack by pests or pathogens, avoiding costs (van Hulten et al. 2006). Priming is therefore probably more similar conceptually to vaccination or immunization in vertebrates (Hilker et al. 2016; Mak et al. 2014) than PDR, as it involves a certain memory of the first stress-indicating signal. Priming falls within the concept of natural plant immunization, which has been suggested as one of the new targets for the next green revolution (Dewen et al. 2017; Quintana-Rodriguez et al. 2018).
The long-lasting nature of priming is one of its strengths for crop protection . Importantly, across shorter to longer periods, priming memory has been correlated with epigenetic changes (Jaskiewicz et al. 2011; López et al. 2011; Luna and Ton 2012). In fact, the majority of Arabidopsis mutants impeded in de novo DNA methylation show resistance against biotrophic pathogens, but do not constitutively express defences. Rather, they show a constitutive priming phenotype. The lower cost of priming as a defence strategy is evident as such mutants do not usually show alterations in growth or development under control conditions. Thus, the already mentioned induced changes in the epigenetic landscape by the use of mutants, chemical compounds or epigenetic editing could be applied to trigger the epigenetic changes that induce the priming state. One successfully commercialized example is the use of seed treatments with elicitors of defence. Seed treatment with JA or BABA provides long-term priming of pest and disease resistance with either no or minimal costs in terms of growth and development (Paudel et al. 2014; Worrall et al. 2012). Obviously, the most natural priming induction would be to moderately stress plants in order to induce long-lasting priming, or even transgenerational priming. This approach does not involve the use of transgenesis, chemical treatments or any artificial systems, does not involve the constitutive induction of defences and is long-lasting. Priming should also be a sustainable approach, since it is not expected to artificially impact other species or the physicochemical properties of natural ecosystems. Moreover, it is expected to be more resilient to environmental changes as it is naturally responsive to them. Unfortunately, the phenotype induced by these methods is very variable and our poor understanding of the underlying mechanisms prevents the design of molecular markers to track resistance at the current time. Undoubtedly, more research is needed in this promising field to make it feasible and widely applicable. Advances in fundamental understanding, including the interactions between multiple stresses, durability of priming and the development of markers to trace the epigenetic changes are all needed to optimize its application in future crop protection programs.
References
Abel PP, Nelson RS, De B, Hoffmann N, Rogers SG, Fraley RT, Beachy RN (1986) Delay of disease development in transgenic plants that express the tobacco mosaic virus coat protein gene. Science 232:738–743
Agrawal AA (2001) Transgenerational consequences of plant responses to herbivory: an adaptive maternal effect? Am Nat 157:555–569. https://doi.org/10.1086/319932
Agrawal AA (2002) Herbivory and maternal effects: mechanisms and consequences of transgenerational induced plant resistance. Ecology 83:3408–3415. https://doi.org/10.2307/3072089
Agrawal AA, Laforsch C, Tollrian R (1999) Transgenerational induction of defences in animals and plants. Nature 401:60–63. https://doi.org/10.1038/43425
Akimoto K, Katakami H, Kim H-J, Ogawa E, Sano CM, Wada Y, Sano H (2007) Epigenetic inheritance in rice plants. Ann Bot 100:205–217. https://doi.org/10.1093/aob/mcm110
Aktar MW, Sengupta D, Chowdhury A (2009) Impact of pesticides use in agriculture: their benefits and hazards. Interdiscip Toxicol 2:1–12. https://doi.org/10.2478/v10102-009-0001-7
Alizadeh M, Askari H, Najafabadi MS (2018) Temporal expression of three conserved putative microRNAs in response of Citrus × Limon to Xanthomonas citri subsp. citri and Xanthomonas fuscans subsp. Aurantifolii. Biotechnologia 98:257–264. https://doi.org/10.5114/bta.2017.70803
Alvarez-Venegas R, Abdallat AA, Guo M, Alfano JR, Avramova Z (2007) Epigenetic control of a transcription factor at the cross section of two antagonistic pathways. Epigenetics 2:106–113
Ayyappan V, Kalavacharla V, Thimmapuram J, Bhide KP, Sripathi VR, Smolinski TG, Manoharan M, Thurston Y, Todd A, Kingham B (2015) Genome-wide profiling of histone modifications (H3K9me2 and H4K12ac) and gene expression in rust (Uromyces appendiculatus) inoculated common bean (Phaseolus vulgaris L.). PLoS One 10(7):e0132176. https://doi.org/10.1371/journal.pone.0132176
Badeaux AI, Shi Y (2013) Emerging roles for chromatin as a signal integration and storage platform. Nat Rev Mol Cell Biol 14:211–224. https://doi.org/10.1038/nrm3545
Bannister AJ, Kouzarides T (2005) Reversing histone methylation. Nature 436:1103–1106. https://doi.org/10.1038/nature04048
Baum JA, Bogaert T, Clinton W, Heck GR, Feldmann P, Ilagan O, Johnson S, Plaetinck G, Munyikwa T, Pleau M, Vaughn T, Roberts J (2007) Control of coleopteran insect pests through RNA interference. Nat Biotechnol 25:1322–1326. https://doi.org/10.1038/nbt1359
Beckers GJM, Jaskiewicz M, Liu Y, Underwood WR, He SY, Zhang S, Conrath U (2009) Mitogen-activated protein kinases 3 and 6 are required for full priming of stress responses in Arabidopsis thaliana. Plant Cell 21:944–953. https://doi.org/10.1105/tpc.108.062158
Berger SL (2007) The complex language of chromatin regulation during transcription. Nature 447(7143):407–412. https://doi.org/10.1038/nature05915
Berr A, McCallum EJ, Alioua A, Heintz D, Heitz T, Shen W-H (2010) Arabidopsis histone methyltransferase SET DOMAIN GROUP8 mediates induction of the jasmonate/ethylene pathway genes in plant defense response to necrotrophic fungi. Plant Physiol 154:1403–1414. https://doi.org/10.1104/pp.110.161497
Berriri S, Gangappa SN, Kumar SV (2016) SWR1 chromatin-remodeling complex subunits and H2A.Z have non-overlapping functions in immunity and gene regulation in Arabidopsis. Mol Plant 9:1051–1065. https://doi.org/10.1016/j.molp.2016.04.003
Bewick AJ, Schmitz RJ (2017) Gene body DNA methylation in plants. Curr Opin Plant Biol 36:103–110. https://doi.org/10.1016/j.pbi.2016.12.007
Bezhani S, Winter C, Hershman S, Wagner JD, Kennedy JF, Kwon CS, Pfluger J, Su Y, Wagner D (2007) Unique, shared, and redundant roles for the Arabidopsis SWI/SNF chromatin remodeling ATPases BRAHMA and SPLAYED. Plant Cell 19:403–416. https://doi.org/10.1105/tpc.106.048272
Bian H, Xie Y, Guo F, Han N, Ma S, Zeng Z, Wang J, Yang Y, Zhu M (2012) Distinctive expression patterns and roles of the miRNA393/TIR1 homolog module in regulating flag leaf inclination and primary and crown root growth in rice (Oryza sativa). New Phytol 196:149–161. https://doi.org/10.1111/j.1469-8137.2012.04248.x
Bilichak A, Ilnytskyy Y, Wóycicki R, Kepeshchuk N, Fogen D, Kovalchuk I (2015) The elucidation of stress memory inheritance in Brassica rapa plants. Front Plant Sci 6:5. https://doi.org/10.3389/fpls.2015.00005
Blevins T, Rajeswaran R, Shivaprasad PV, Beknazariants D, Si-Ammour A, Park H-S, Vazquez F, Robertson D, Meins F, Hohn T, Pooggin MM (2006) Four plant Dicers mediate viral small RNA biogenesis and DNA virus induced silencing. Nucleic Acids Res 34:6233–6246. https://doi.org/10.1093/nar/gkl886
Boccara M, Sarazin A, Billoud B, Bulski A, Chapell L, Baulcombe D, Colot V (2017) Analysis of small RNA populations using hybridization to DNA tiling arrays. Methods Mol Biol 1456:127–139. https://doi.org/10.1007/978-1-4899-7708-3_11
Borges F, Martienssen RA (2015) The expanding world of small RNAs in plants. Nat Rev Mol Cell Biol 16:727–741. https://doi.org/10.1038/nrm4085
Bossdorf O, Arcuri D, Richards CL, Pigliucci M (2010) Experimental alteration of DNA methylation affects the phenotypic plasticity of ecologically relevant traits in Arabidopsis thaliana. Evol Ecol 24:541–553. https://doi.org/10.1007/s10682-010-9372-7
Boyko A, Blevins T, Yao Y, Golubov A, Bilichak A, Ilnytskyy Y, Hollander J, Jr FM, Kovalchuk I (2010) Transgenerational adaptation of Arabidopsis to stress requires DNA methylation and the function of Dicer-like proteins. PLoS One 5:e9514. https://doi.org/10.1371/journal.pone.0009514
Cai Q, Qiao L, Wang M, He B, Lin F-M, Palmquist J, Huang H-D, Jin H (2018) Plants send small RNAs in extracellular vesicles to fungal pathogen to silence virulence genes. Science 360(6393):1126–1129. https://doi.org/10.1126/science.aar4142
Cambiagno DA, Nota F, Zavallo D, Rius S, Casati P, Asurmendi S, Alvarez ME (2018) Immune receptor genes and pericentromeric transposons as targets of common epigenetic regulatory elements. Plant J 96(6):1178–1190. https://doi.org/10.1111/tpj.14098
Castillo-González C, Liu X, Huang C, Zhao C, Ma Z, Hu T, Sun F, Zhou Y, Zhou X, Wang X-J, Zhang X (2015) Geminivirus-encoded TrAP suppressor inhibits the histone methyltransferase SUVH4/KYP to counter host defense. elife 4:e06671. https://doi.org/10.7554/eLife.06671
Chen W, Zhu Q, Liu Y, Zhang Q (2017) Chapter Nine—Chromatin remodeling and plant immunity. In: Donev R (ed) Advances in protein chemistry and structural biology, chromatin remodelling and immunity. Academic Press, Cambridge, pp 243–260. https://doi.org/10.1016/bs.apcsb.2016.08.006
Chitwood DH, Timmermans MCP (2010) Small RNAs are on the move. Nature 467:415–419. https://doi.org/10.1038/nature09351
Choi S-M, Song H-R, Han S-K, Han M, Kim C-Y, Park J, Lee Y-H, Jeon J-S, Noh Y-S, Noh B (2012) HDA19 is required for the repression of salicylic acid biosynthesis and salicylic acid-mediated defense responses in Arabidopsis. Plant J 71:135–146. https://doi.org/10.1111/j.1365-313X.2012.04977.x
Cockerill PN (2011) Structure and function of active chromatin and DNase I hypersensitive sites. FEBS J 278:2182–2210. https://doi.org/10.1111/j.1742-4658.2011.08128.x
Cohen Y, Vaknin M, Mauch-Mani B (2016) BABA-induced resistance: milestones along a 55-year journey. Phytoparasitica 44:513–538. https://doi.org/10.1007/s12600-016-0546-x
Cokus SJ, Feng S, Zhang X, Chen Z, Merriman B, Haudenschild CD, Pradhan S, Nelson SF, Pellegrini M, Jacobsen SE (2008) Shotgun bisulphite sequencing of the Arabidopsis genome reveals DNA methylation patterning. Nature 452:215–219. https://doi.org/10.1038/nature06745
Coleman-Derr D, Zilberman D (2012) Deposition of histone variant H2A.Z within gene bodies regulates responsive genes. PLoS Genet 8:e1002988. https://doi.org/10.1371/journal.pgen.1002988
Collinge DB, Jørgensen HJL, Lund OS, Lyngkjær MF (2010) Engineering pathogen resistance in crop plants: current trends and future prospects. Annu Rev Phytopathol 48:269–291. https://doi.org/10.1146/annurev-phyto-073009-114430
Connor S (2002) How an experiment to change the colour of a petunia led to a breakthrough in the treatments of cancer and Aids. The Independent. http://www.independent.co.uk/news/science/how-an-experiment-to-change-the-colour-of-a-petunia-led-to-a-breakthrough-in-the-treatment-of-cancer-5547175.html
Conrath U, Beckers GJM, Langenbach CJG, Jaskiewicz MR (2015) Priming for enhanced defense. Annu Rev Phytopathol 53:97–119. https://doi.org/10.1146/annurev-phyto-080614-120132
Cortijo S, Wardenaar R, Colomé-Tatché M, Gilly A, Etcheverry M, Labadie K, Caillieux E, Hospital F, Aury J-M, Wincker P, Roudier F, Jansen RC, Colot V, Johannes F (2014a) Mapping the epigenetic basis of complex traits. Science 343:1145–1148. https://doi.org/10.1126/science.1248127
Cortijo S, Wardenaar R, Colomé-Tatché M, Johannes F, Colot V (2014b) Genome-wide analysis of DNA methylation in Arabidopsis using MeDIP-chip. Methods Mol Biol 1112:125–149. https://doi.org/10.1007/978-1-62703-773-0_9
Coursey T, Regedanz E, Bisaro DM (2018) Arabidopsis RNA polymerase V mediates enhanced compaction and silencing of geminivirus and transposon chromatin during host recovery from infection. J Virol 92:e01320-17. https://doi.org/10.1128/JVI.01320-17
Crevillén P, Yang H, Cui X, Greeff C, Trick M, Qiu Q, Cao X, Dean C (2014) Epigenetic reprogramming that prevents transgenerational inheritance of the vernalized state. Nature 515:587–590. https://doi.org/10.1038/nature13722
Crisp PA, Ganguly D, Eichten SR, Borevitz JO, Pogson BJ (2016) Reconsidering plant memory: intersections between stress recovery, RNA turnover, and epigenetics. Sci Adv 2:e1501340. https://doi.org/10.1126/sciadv.1501340
Csorba T, Kontra L, Burgyán J (2015) Viral silencing suppressors: tools forged to fine-tune host-pathogen coexistence. Virology 479–480:85–103. https://doi.org/10.1016/j.virol.2015.02.028
Cubas P, Vincent C, Coen E (1999) An epigenetic mutation responsible for natural variation in floral symmetry. Nature 401:157–161. https://doi.org/10.1038/43657
Dekker J, Rippe K, Dekker M, Kleckner N (2002) Capturing chromosome conformation. Science 295:1306–1311. https://doi.org/10.1126/science.1067799
De-La-Peña C, Rangel-Cano A, Alvarez-Venegas R (2012) Regulation of disease-responsive genes mediated by epigenetic factors: interaction of Arabidopsis-Pseudomonas. Mol Plant Pathol 13:388–398. https://doi.org/10.1111/j.1364-3703.2011.00757.x
Dewen Q, Yijie D, Yi Z, Shupeng L, Fachao S (2017) Plant immunity inducer development and application. Mol Plant-Microbe Interact 30:355–360. https://doi.org/10.1094/MPMI-11-16-0231-CR
Dhawan R, Luo H, Foerster AM, AbuQamar S, Du H-N, Briggs SD, Scheid OM, Mengiste T (2009) HISTONE MONOUBIQUITINATION1 interacts with a subunit of the mediator complex and regulates defense against necrotrophic fungal pathogens in Arabidopsis. Plant Cell 21:1000–1019. https://doi.org/10.1105/tpc.108.062364
Ding B, Wang G-L (2015) Chromatin versus pathogens: the function of epigenetics in plant immunity. Front Plant Sci 6:675. https://doi.org/10.3389/fpls.2015.00675
Ding B, Bellizzi MR, Ning Y, Meyers BC, Wang G-L (2012a) HDT701, a histone H4 deacetylase, negatively regulates plant innate immunity by modulating histone H4 acetylation of defense-related genes in rice. Plant Cell 24:3783–3794. https://doi.org/10.1105/tpc.112.101972
Ding Y, Fromm M, Avramova Z (2012b) Multiple exposures to drought “train” transcriptional responses in Arabidopsis. Nat Commun 3:740. https://doi.org/10.1038/ncomms1732
Dowen RH, Pelizzola M, Schmitz RJ, Lister R, Dowen JM, Nery JR, Dixon JE, Ecker JR (2012) Widespread dynamic DNA methylation in response to biotic stress. Proc Natl Acad Sci 109:12858–12859. https://doi.org/10.1073/pnas.1209329109
Duan C-G, Wang C-H, Guo H-S (2012) Application of RNA silencing to plant disease resistance. Silence 3:5. https://doi.org/10.1186/1758-907X-3-5
Dunoyer P, Brosnan CA, Schott G, Wang Y, Jay F, Alioua A, Himber C, Voinnet O (2010) Retracted: an endogenous, systemic RNAi pathway in plants. EMBO J 29:1699–1712. https://doi.org/10.1038/emboj.2010.65
Dutta A, Choudhary P, Caruana J, Raina R (2017) JMJ27, an Arabidopsis H3K9 histone demethylase, modulates defense against Pseudomonas syringae and flowering time. Plant J 91:1015–1028. https://doi.org/10.1111/tpj.13623
Eamens A, Wang M-B, Smith NA, Waterhouse PM (2008) RNA silencing in plants: yesterday, today, and tomorrow. Plant Physiol 147:456–468. https://doi.org/10.1104/pp.108.117275
Ellendorff U, Fradin EF, de Jonge R, Thomma BPHJ (2009) RNA silencing is required for Arabidopsis defence against Verticillium wilt disease. J Exp Bot 60:591–602. https://doi.org/10.1093/jxb/ern306
European Academies Science Advisory Council, Deutsche Akademie der Naturforscher Leopoldina (Eds) (2014) Risks to plant health: European Union priorities for tackling emerging plant pests and diseases, EASAC policy report. EASAC Secretariat, Deutsche Akademie der Naturforscher Leopoldina, Halle (Saale)
European Commission (2009) Sustainable use of pesticides - Food Safety - European Commission. https://ec.europa.eu/food/plant/pesticides/sustainable_use_pesticides_en
Fang X, Qi Y (2016) RNAi in plants: an argonaute-centered view. Plant Cell 28:272–285. https://doi.org/10.1105/tpc.15.00920
FAO (2009a) How to feed the world in 2050. Feed World 2050. http://www.fao.org/fileadmin/templates/wsfs/docs/expert_paper/How_to_Feed_the_World_in_2050.pdf
FAO (2009b) 2050 high-level experts forum: the forum. http://www.fao.org/wsfs/forum2050/wsfs-forum/en/
Fransz P, de Jong H (2011) From nucleosome to chromosome: a dynamic organization of genetic information. Plant J 66:4–17. https://doi.org/10.1111/j.1365-313X.2011.04526.x
Furey TS (2012) ChIP-seq and beyond: new and improved methodologies to detect and characterize protein-DNA interactions. Nat Rev Genet 13:840–852. https://doi.org/10.1038/nrg3306
Gallego-Bartolomé J, Gardiner J, Liu W, Papikian A, Ghoshal B, Kuo HY, Zhao JM-C, Segal DJ, Jacobsen SE (2018) Targeted DNA demethylation of the Arabidopsis genome using the human TET1 catalytic domain. Proc Natl Acad Sci 115(9):E2125–E2134. https://doi.org/10.1073/pnas.1716945115
Gallusci P, Dai Z, Génard M, Gauffretau A, Leblanc-Fournier N, Richard-Molard C, Vile D, Brunel-Muguet S (2017) Epigenetics for plant improvement: current knowledge and modeling avenues. Trends Plant Sci 22:610–623. https://doi.org/10.1016/j.tplants.2017.04.009
Gendrel A-V, Lippman Z, Yordan C, Colot V, Martienssen RA (2002) Dependence of heterochromatic histone H3 methylation patterns on the Arabidopsis gene DDM1. Science 297:1871–1873. https://doi.org/10.1126/science.1074950
Glazebrook J (2005) Contrasting mechanisms of defense against biotrophic and necrotrophic pathogens. Annu Rev Phytopathol 43:205–227. https://doi.org/10.1146/annurev.phyto.43.040204.135923
Gonsalves D (1998) Control of papaya ringspot virus in papaya: a case study. Annu Rev Phytopathol 36:415–437. https://doi.org/10.1146/annurev.phyto.36.1.415
Grativol C, Hemerly AS, Ferreira PCG (2012) Genetic and epigenetic regulation of stress responses in natural plant populations. Biochim Biophys Acta 1819:176–185. https://doi.org/10.1016/j.bbagrm.2011.08.010
Guo N, Ye W-W, Wu X-L, Shen D-Y, Wang Y-C, Xing H, Dou D-L (2011) Microarray profiling reveals microRNAs involving soybean resistance to Phytophthora sojae. Genome 54:954–958. https://doi.org/10.1139/g11-050
Haas J, Lozano ER, Haida KS, Mazaro SM, Vismara E d S, Poppy GM (2018) Getting ready for battle: do cabbage seeds treated with jasmonic acid and chitosan affect chewing and sap-feeding insects? Entomol Exp Appl 166:412–419. https://doi.org/10.1111/eea.12678
Hamilton RI (1980) Defenses triggered by previous invaders: viruses. In: Horsfall JG, Cowling EB (eds) Plant disease, an advanced treatise. Academic Press, New York, NY, pp 279–302
Han S-K, Sang Y, Rodrigues A, BIOL425 F2010, Wu M-F, Rodriguez PL, Wagner D (2012) The SWI2/SNF2 chromatin remodeling ATPase BRAHMA represses abscisic acid responses in the absence of the stress stimulus in Arabidopsis. Plant Cell 24:4892–4906. https://doi.org/10.1105/tpc.112.105114
Han S-K, Wu M-F, Cui S, Wagner D (2015) Roles and activities of chromatin remodeling ATPases in plants. Plant J 83:62–77. https://doi.org/10.1111/tpj.12877
Hauben M, Haesendonckx B, Standaert E, Van Der Kelen K, Azmi A, Akpo H, Van Breusegem F, Guisez Y, Bots M, Lambert B, Laga B, De Block M (2009) Energy use efficiency is characterized by an epigenetic component that can be directed through artificial selection to increase yield. Proc Natl Acad Sci U S A 106:20109–20114. https://doi.org/10.1073/pnas.0908755106
He X, Sun Q, Jiang H, Zhu X, Mo J, Long L, Xiang L, Xie Y, Shi Y, Yuan Y, Cai Y (2014) Identification of novel microRNAs in the Verticillium wilt-resistant upland cotton variety KV-1 by high-throughput sequencing. Springerplus 3:564. https://doi.org/10.1186/2193-1801-3-564
He X-F, Fang Y-Y, Feng L, Guo H-S (2008) Characterization of conserved and novel microRNAs and their targets, including a TuMV-induced TIR–NBS–LRR class R gene-derived novel miRNA in Brassica. FEBS Lett 582:2445–2452. https://doi.org/10.1016/j.febslet.2008.06.011
Heard E, Martienssen RA (2014) Transgenerational epigenetic inheritance: myths and mechanisms. Cell 157:95–109. https://doi.org/10.1016/j.cell.2014.02.045
Hewezi T, Howe P, Maier TR, Baum TJ (2008) Arabidopsis small RNAs and their targets during cyst nematode parasitism. Mol Plant-Microbe Interact 21:1622–1634. https://doi.org/10.1094/MPMI-21-12-1622
Hilker M, Schwachtje J, Baier M, Balazadeh S, Bäurle I, Geiselhardt S, Hincha DK, Kunze R, Mueller-Roeber B, Rillig MC, Rolff J, Romeis T, Schmülling T, Steppuhn A, van Dongen J, Whitcomb SJ, Wurst S, Zuther E, Kopka J (2016) Priming and memory of stress responses in organisms lacking a nervous system. Biol Rev 91:1118–1133. https://doi.org/10.1111/brv.12215
Hipper C, Brault V, Ziegler-Graff V, Revers F (2013) Viral and cellular factors involved in phloem transport of plant viruses. Front Plant Sci 4:154. https://doi.org/10.3389/fpls.2013.00154
Hofmeister BT, Lee K, Rohr NA, Hall DW, Schmitz RJ (2017) Stable inheritance of DNA methylation allows creation of epigenotype maps and the study of epiallele inheritance patterns in the absence of genetic variation. Genome Biol 18:155. https://doi.org/10.1186/s13059-017-1288-x
Hollwey E, Out S, Watson MR, Heidmann I, Meyer P (2017) TET3-mediated demethylation in tomato activates expression of a CETS gene that stimulates vegetative growth. Plant Direct 1:e00022. https://doi.org/10.1002/pld3.22
Hou Y, Wang L, Wang L, Liu L, Li L, Sun L, Rao Q, Zhang J, Huang S (2015) JMJ704 positively regulates rice defense response against Xanthomonas oryzae pv. oryzae infection via reducing H3K4me2/3 associated with negative disease resistance regulators. BMC Plant Biol 15:286. https://doi.org/10.1186/s12870-015-0674-3
Hu M, Pei B-L, Zhang L-F, Li Y-Z (2014) Histone H2B monoubiquitination is involved in regulating the dynamics of microtubules during the defense response to Verticillium dahliae toxins in Arabidopsis. Plant Physiol 164:1857–1865. https://doi.org/10.1104/pp.113.234567
Huang J, Yang M, Zhang X (2016) The function of small RNAs in plant biotic stress response. J Integr Plant Biol 58:312–327. https://doi.org/10.1111/jipb.12463
Iwasaki M, Paszkowski J (2014) Epigenetic memory in plants. EMBO J 33:1987–1998. https://doi.org/10.15252/embj.201488883
Jackel JN, Storer JM, Coursey T, Bisaro DM (2016) Arabidopsis RNA polymerases IV and V are required to establish H3K9 methylation, but not cytosine methylation, on geminivirus chromatin. J Virol 90:7529–7540. https://doi.org/10.1128/JVI.00656-16
Jagadeeswaran G, Saini A, Sunkar R (2009) Biotic and abiotic stress down-regulate miR398 expression in Arabidopsis. Planta 229:1009–1014. https://doi.org/10.1007/s00425-009-0889-3
Jaskiewicz M, Conrath U, Peterhänsel C (2011) Chromatin modification acts as a memory for systemic acquired resistance in the plant stress response. EMBO Rep 12:50–55. https://doi.org/10.1038/embor.2010.186
Jeddeloh JA, Bender J, Richards EJ (1998) The DNA methylation locus DDM1 is required for maintenance of gene silencing in Arabidopsis. Genes Dev 12:1714–1725
Jeyaraj A, Liu S, Zhang X, Zhang R, Shangguan M, Wei C (2017) Genome-wide identification of microRNAs responsive to Ectropis oblique feeding in tea plant (Camellia sinensis L.). Sci Rep 7:13634. https://doi.org/10.1038/s41598-017-13692-7
Ji L, Neumann DA, Schmitz RJ (2015) Crop epigenomics: identifying, unlocking, and harnessing cryptic variation in crop genomes. Mol Plant 8:860–870. https://doi.org/10.1016/j.molp.2015.01.021
Jia R, Zhao H, Huang J, Kong H, Zhang Y, Guo J, Huang Q, Guo Y, Wei Q, Zuo J, Zhu YJ, Peng M, Guo A (2017) Use of RNAi technology to develop a PRSV-resistant transgenic papaya. Sci Rep 7:12636. https://doi.org/10.1038/s41598-017-13049-0
Jiang L, Qian D, Zheng H, Meng L-Y, Chen J, Le W-J, Zhou T, Zhou Y-J, Wei C-H, Li Y (2012) RNA-dependent RNA polymerase 6 of rice (Oryza sativa) plays role in host defense against negative-strand RNA virus, Rice stripe virus. Virus Res 163:512–519. https://doi.org/10.1016/j.virusres.2011.11.016
Johannes F, Porcher E, Teixeira FK, Saliba-Colombani V, Simon M, Agier N, Bulski A, Albuisson J, Heredia F, Audigier P, Bouchez D, Dillmann C, Guerche P, Hospital F, Colot V (2009) Assessing the impact of transgenerational epigenetic variation on complex traits. PLoS Genet 5:e1000530. https://doi.org/10.1371/journal.pgen.1000530
Johnson KCM, Xia S, Feng X, Li X (2015) The chromatin remodeler SPLAYED negatively regulates SNC1-mediated immunity. Plant Cell Physiol 56:1616–1623. https://doi.org/10.1093/pcp/pcv087
Jones JDG, Dangl JL (2006) The plant immune system. Nature 444:323–329. https://doi.org/10.1038/nature05286
Jones PA (1985) Altering gene expression with 5-azacytidine. Cell 40:485–486. https://doi.org/10.1016/0092-8674(85)90192-8
Kalantidis K, Schumacher HT, Alexiadis T, Helm JM (2008) RNA silencing movement in plants. Biol Cell 100:13–26. https://doi.org/10.1042/BC20070079
Kapoor M, Arora R, Lama T, Nijhawan A, Khurana JP, Tyagi AK, Kapoor S (2008) Genome-wide identification, organization and phylogenetic analysis of Dicer-like, Argonaute and RNA-dependent RNA polymerase gene families and their expression analysis during reproductive development and stress in rice. BMC Genomics 9:451. https://doi.org/10.1186/1471-2164-9-451
Katiyar-Agarwal S, Jin H (2010) Role of small RNAs in host-microbe interactions. Annu Rev Phytopathol 48:225–246. https://doi.org/10.1146/annurev-phyto-073009-114457
Katiyar-Agarwal S, Morgan R, Dahlbeck D, Borsani O, Villegas A, Zhu J-K, Staskawicz BJ, Jin H (2006) A pathogen-inducible endogenous siRNA in plant immunity. Proc Natl Acad Sci U S A 103:18002–18007. https://doi.org/10.1073/pnas.0608258103
Katiyar-Agarwal S, Gao S, Vivian-Smith A, Jin H (2007) A novel class of bacteria-induced small RNAs in Arabidopsis. Genes Dev 21:3123–3134. https://doi.org/10.1101/gad.1595107
Kawakatsu T, Huang S-SC, Jupe F, Sasaki E, Schmitz RJ, Urich MA, Castanon R, Nery JR, Barragan C, He Y, Chen H, Dubin M, Lee C-R, Wang C, Bemm F, Becker C, O’Neil R, O’Malley RC, Quarless DX, 1001 Genomes Consortium, Schork NJ, Weigel D, Nordborg M, Ecker JR (2016) Epigenomic diversity in a global collection of Arabidopsis thaliana accessions. Cell 166:492–505. https://doi.org/10.1016/j.cell.2016.06.044
Kim K-C, Lai Z, Fan B, Chen Z (2008) Arabidopsis WRKY38 and WRKY62 transcription factors interact with histone deacetylase 19 in basal defense. Plant Cell 20:2357–2371. https://doi.org/10.1105/tpc.107.055566
Klessig DF, Choi HW, Dempsey DA (2018) Systemic acquired resistance and salicylic acid: past, present, and future. Mol Plant-Microbe Interact 31:871–888. https://doi.org/10.1094/MPMI-03-18-0067-CR
Koch A, Kogel K-H (2014) New wind in the sails: improving the agronomic value of crop plants through RNAi-mediated gene silencing. Plant Biotechnol J 12:821–831. https://doi.org/10.1111/pbi.12226
Koornneef A, Rindermann K, Gatz C, Pieterse CM (2008) Histone modifications do not play a major role in salicylate-mediated suppression of jasmonate-induced PDF1.2 gene expression. Commun Integr Biol 1:143
Kornberg RD (1974) Chromatin structure: a repeating unit of histones and DNA. Science 184:868–871
Kothari IL, Patel M (2004) Plant immunization. Indian J Exp Biol 42:244–252
Kouzarides T (2007) Chromatin modifications and their function. Cell 128:693–705. https://doi.org/10.1016/j.cell.2007.02.005
Kungulovski G, Jeltsch A (2016) Epigenome editing: state of the art, concepts, and perspectives. Trends Genet 32:101–113. https://doi.org/10.1016/j.tig.2015.12.001
Lammerink J, MacGibbon DB, Wallace AR (1984) Effect of the cabbage aphid (Brevicoryne brassicae) on total glucosinolate in the seed of oilseed rape (Brassica napus). N Z J Agric Res 27:89–92. https://doi.org/10.1080/00288233.1984.10425735
Latrasse D, Jégu T, Li H, de Zelicourt A, Raynaud C, Legras S, Gust A, Samajova O, Veluchamy A, Rayapuram N, Ramirez-Prado JS, Kulikova O, Colcombet J, Bigeard J, Genot B, Bisseling T, Benhamed M, Hirt H (2017) MAPK-triggered chromatin reprogramming by histone deacetylase in plant innate immunity. Genome Biol 18:131. https://doi.org/10.1186/s13059-017-1261-8
Latzel V, Zhang Y, Karlsson Moritz K, Fischer M, Bossdorf O (2012) Epigenetic variation in plant responses to defence hormones. Ann Bot 110:1423–1428. https://doi.org/10.1093/aob/mcs088
Law JA, Jacobsen SE (2010) Establishing, maintaining and modifying DNA methylation patterns in plants and animals. Nat Rev Genet 11:204–220. https://doi.org/10.1038/nrg2719
Le T-N, Schumann U, Smith NA, Tiwari S, Au PC, Zhu Q-H, Taylor JM, Kazan K, Llewellyn DJ, Zhang R, Dennis ES, Wang M-B (2014) DNA demethylases target promoter transposable elements to positively regulate stress responsive genes in Arabidopsis. Genome Biol 15:458. https://doi.org/10.1186/s13059-014-0458-3
Lee S, Fu F, Xu S, Lee SY, Yun D-J, Mengiste T (2016) Global regulation of plant immunity by histone lysine methyl transferases. Plant Cell 28:1640–1661. https://doi.org/10.1105/tpc.16.00012
Lewsey MG, Hardcastle TJ, Melnyk CW, Molnar A, Valli A, Urich MA, Nery JR, Baulcombe DC, Ecker JR (2016) Mobile small RNAs regulate genome-wide DNA methylation. Proc Natl Acad Sci 113:E801–E810. https://doi.org/10.1073/pnas.1515072113
Li B, Carey M, Workman JL (2007) The role of chromatin during transcription. Cell 128:707–719. https://doi.org/10.1016/j.cell.2007.01.015
Li C, Zamore PD (2018) Analysis of small RNAs by northern hybridization. Cold Spring Harb Protoc 2018:pdb.prot097493. https://doi.org/10.1101/pdb.prot097493
Li D, Wang F, Wang C, Zou L, Wang Z, Chen Q, Niu C, Zhang R, Ling Y, Wang B (2016) MicroRNA-mediated susceptible poplar gene expression regulation associated with the infection of virulent Melampsora larici-populina. BMC Genomics 17:59. https://doi.org/10.1186/s12864-015-2286-6
Li F, Pignatta D, Bendix C, Brunkard JO, Cohn MM, Tung J, Sun H, Kumar P, Baker B (2012) MicroRNA regulation of plant innate immune receptors. Proc Natl Acad Sci 109:1790–1795. https://doi.org/10.1073/pnas.1118282109
Li T, Chen X, Zhong X, Zhao Y, Liu X, Zhou S, Cheng S, Zhou D-X (2013) Jumonji C domain protein JMJ705-mediated removal of histone H3 lysine 27 trimethylation is involved in defense-related gene activation in rice. Plant Cell 25:4725–4736. https://doi.org/10.1105/tpc.113.118802
Li X, Jiang Y, Ji Z, Liu Y, Zhang Q (2015) BRHIS1 suppresses rice innate immunity through binding to monoubiquitinated H2A and H2B variants. EMBO Rep 16:1192–1202. https://doi.org/10.15252/embr.201440000
Li Y, Zhang Q, Zhang J, Wu L, Qi Y, Zhou J-M (2010) Identification of microRNAs involved in pathogen-associated molecular pattern-triggered plant innate immunity. Plant Physiol 152:2222–2231. https://doi.org/10.1104/pp.109.151803
Li Y, Lu Y-G, Shi Y, Wu L, Xu Y-J, Huang F, Guo X-Y, Zhang Y, Fan J, Zhao J-Q, Zhang H-Y, Xu P-Z, Zhou J-M, Wu X-J, Wang P-R, Wang W-M (2014) Multiple rice microRNAs are involved in immunity against the blast fungus Magnaporthe oryzae. Plant Physiol 164:1077–1092. https://doi.org/10.1104/pp.113.230052
Liang Z, Shen L, Cui X, Bao S, Geng Y, Yu G, Liang F, Xie S, Lu T, Gu X, Yu H (2018) DNA N6-adenine methylation in Arabidopsis thaliana. Dev Cell 45:406–416.e3. https://doi.org/10.1016/j.devcel.2018.03.012
Lindbo JA, Silva-Rosales L, Proebsting WM, Dougherty WG (1993) Induction of a highly specific antiviral state in transgenic plants: implications for regulation of gene expression and virus resistance. Plant Cell 5:1749–1759. https://doi.org/10.1105/tpc.5.12.1749
Liu F, Quesada V, Crevillén P, Bäurle I, Swiezewski S, Dean C (2007) The Arabidopsis RNA-binding protein FCA requires a lysine-specific demethylase 1 homolog to downregulate FLC. Mol Cell 28:398–407. https://doi.org/10.1016/j.molcel.2007.10.018
López A, Ramírez V, García-Andrade J, Flors V, Vera P (2011) The RNA silencing enzyme RNA polymerase V is required for plant immunity. PLoS Genet 7:e1002434. https://doi.org/10.1371/journal.pgen.1002434
López Sánchez A, Stassen JHM, Furci L, Smith LM, Ton J (2016) The role of DNA (de)methylation in immune responsiveness of Arabidopsis. Plant J 88:361–374. https://doi.org/10.1111/tpj.13252
Lu S, Sun Y-H, Amerson H, Chiang VL (2007) MicroRNAs in loblolly pine (Pinus taeda L.) and their association with fusiform rust gall development. Plant J 51:1077–1098. https://doi.org/10.1111/j.1365-313X.2007.03208.x
Lucht JM (2015) Public acceptance of plant biotechnology and GM crops. Viruses 7:4254–4281. https://doi.org/10.3390/v7082819
Luna E, Ton J (2012) The epigenetic machinery controlling transgenerational systemic acquired resistance. Plant Signal Behav 7:615–618. https://doi.org/10.4161/psb.20155
Luna E, Bruce TJA, Roberts MR, Flors V, Ton J (2012) Next-generation systemic acquired resistance. Plant Physiol 158:844–853. https://doi.org/10.1104/pp.111.187468
Luna E, López A, Kooiman J, Ton J (2014) Role of NPR1 and KYP in long-lasting induced resistance by β-aminobutyric acid. Front Plant Sci 5:184. https://doi.org/10.3389/fpls.2014.00184
Mak T, Saunders M, Jett B (2014) Primer to the immune response. Academic Cell. Elsevier, Amsterdam. https://doi.org/10.1016/C2009-0-62217-0
Malinovsky FG, Fangel JU, Willats WGT (2014) The role of the cell wall in plant immunity. Front Plant Sci 5:178. https://doi.org/10.3389/fpls.2014.00178
March-Díaz R, García-Domínguez M, Lozano-Juste J, León J, Florencio FJ, Reyes JC (2008) Histone H2A.Z and homologues of components of the SWR1 complex are required to control immunity in Arabidopsis. Plant J 53:475–487. https://doi.org/10.1111/j.1365-313X.2007.03361.x
Martelli GP (2001) Transgenic resistance to plant pathogens: benefits and risks. J Plant Pathol 83:37–46
Martínez-Aguilar K, Ramírez-Carrasco G, Hernández-Chávez JL, Barraza A, Alvarez-Venegas R (2016) Use of BABA and INA as activators of a primed state in the common bean (Phaseolus vulgaris L.). Front Plant Sci 7:653. https://doi.org/10.3389/fpls.2016.00653
Mathieu O, Reinders J, Čaikovski M, Smathajitt C, Paszkowski J (2007) Transgenerational stability of the Arabidopsis epigenome is coordinated by CG methylation. Cell 130:851–862. https://doi.org/10.1016/j.cell.2007.07.007
Matzke MA, Matzke AJM (eds) (2000) Plant gene silencing. Springer, Dordrecht
Matzke MA, Mosher RA (2014) RNA-directed DNA methylation: an epigenetic pathway of increasing complexity. Nat Rev Genet 15:394–408. https://doi.org/10.1038/nrg3683
Ménard R, Verdier G, Ors M, Erhardt M, Beisson F, Shen W-H (2014) Histone H2B monoubiquitination is involved in the regulation of cutin and wax composition in Arabidopsis thaliana. Plant Cell Physiol 55:455–466. https://doi.org/10.1093/pcp/pct182
Mengel A, Ageeva A, Georgii E, Bernhardt J, Wu K, Durner J, Lindermayr C (2017) Nitric oxide modulates histone acetylation at stress genes by inhibition of histone deacetylases. Plant Physiol 173:1434–1452. https://doi.org/10.1104/pp.16.01734
Molnar A, Melnyk CW, Bassett A, Hardcastle TJ, Dunn R, Baulcombe DC (2010) Small silencing RNAs in plants are mobile and direct epigenetic modification in recipient cells. Science 328:872–875. https://doi.org/10.1126/science.1187959
Morgado L, Preite V, Oplaat C, Anava S, Ferreira de Carvalho J, Rechavi O, Johannes F, Verhoeven KJF (2017) Small RNAs reflect grandparental environments in apomictic dandelion. Mol Biol Evol 34:2035–2040. https://doi.org/10.1093/molbev/msx150
Mozgová I, Wildhaber T, Liu Q, Abou-Mansour E, L’Haridon F, Métraux J-P, Gruissem W, Hofius D, Hennig L (2015) Chromatin assembly factor CAF-1 represses priming of plant defence response genes. Nat Plants 1:15127. https://doi.org/10.1038/nplants.2015.127
Napoli C, Lemieux C, Jorgensen R (1990) Introduction of a chimeric chalcone synthase gene into petunia results in reversible co-suppression of homologous genes in trans. Plant Cell 2:279–289. https://doi.org/10.1105/tpc.2.4.279
Navarro L, Dunoyer P, Jay F, Arnold B, Dharmasiri N, Estelle M, Voinnet O, Jones JDG (2006) A plant miRNA contributes to antibacterial resistance by repressing auxin signaling. Science 312:436–439. https://doi.org/10.1126/science.1126088
Navarro L, Jay F, Nomura K, He SY, Voinnet O (2008) Suppression of the microRNA pathway by bacterial effector proteins. Science 321:964–967. https://doi.org/10.1126/science.1159505
Niederhuth CE, Bewick AJ, Ji L, Alabady MS, Kim KD, Li Q, Rohr NA, Rambani A, Burke JM, Udall JA, Egesi C, Schmutz J, Grimwood J, Jackson SA, Springer NM, Schmitz RJ (2016) Widespread natural variation of DNA methylation within angiosperms. Genome Biol 17:194. https://doi.org/10.1186/s13059-016-1059-0
Nowara D, Gay A, Lacomme C, Shaw J, Ridout C, Douchkov D, Hensel G, Kumlehn J, Schweizer P (2010) HIGS: host-induced gene silencing in the obligate biotrophic fungal pathogen Blumeria graminis. Plant Cell 22:3130–3141. https://doi.org/10.1105/tpc.110.077040
Nunes CC, Dean RA (2012) Host-induced gene silencing: a tool for understanding fungal host interaction and for developing novel disease control strategies. Mol Plant Pathol 13:519–529. https://doi.org/10.1111/j.1364-3703.2011.00766.x
Nuthikattu S, McCue AD, Panda K, Fultz D, DeFraia C, Thomas EN, Slotkin RK (2013) The initiation of epigenetic silencing of active transposable elements is triggered by RDR6 and 21-22 nucleotide small interfering RNAs. Plant Physiol 162:116–131. https://doi.org/10.1104/pp.113.216481
O’Dea RE, Noble DWA, Johnson SL, Hesselson D, Nakagawa S (2016) The role of non-genetic inheritance in evolutionary rescue: epigenetic buffering, heritable bet hedging and epigenetic traps. Environ Epigenet 2:dvv014. https://doi.org/10.1093/eep/dvv014
Palma K, Thorgrimsen S, Malinovsky FG, Fiil BK, Nielsen HB, Brodersen P, Hofius D, Petersen M, Mundy J (2010) Autoimmunity in Arabidopsis acd11 is mediated by epigenetic regulation of an immune receptor. PLoS Pathog 6:e1001137. https://doi.org/10.1371/journal.ppat.1001137
Pandey SP, Shahi P, Gase K, Baldwin IT (2008) Herbivory-induced changes in the small-RNA transcriptome and phytohormone signaling in Nicotiana attenuata. Proc Natl Acad Sci U S A 105:4559–4564. https://doi.org/10.1073/pnas.0711363105
Paudel S, Rajotte EG, Felton GW (2014) Benefits and costs of tomato seed treatment with plant defense elicitors for insect resistance. Arthropod Plant Interact 8(6):539–545. https://doi.org/10.1007/s11829-014-9335-y
Pavet V, Quintero C, Cecchini NM, Rosa AL, Alvarez ME (2006) Arabidopsis displays centromeric DNA hypomethylation and cytological alterations of heterochromatin upon attack by pseudomonas syringae. Mol Plant-Microbe Interact 19:577–587. https://doi.org/10.1094/MPMI-19-0577
Peirats-Llobet M, Han S-K, Gonzalez-Guzman M, Jeong CW, Rodriguez L, Belda-Palazon B, Wagner D, Rodriguez PL (2016) A direct link between abscisic acid sensing and the chromatin-remodeling ATPase BRAHMA via core ABA signaling pathway components. Mol Plant 9:136–147. https://doi.org/10.1016/j.molp.2015.10.003
Pérez-Quintero ÁL, Quintero A, Urrego O, Vanegas P, López C (2012) Bioinformatic identification of cassava miRNAs differentially expressed in response to infection by Xanthomonas axonopodis pv. Manihotis. BMC Plant Biol 12:29. https://doi.org/10.1186/1471-2229-12-29
Pieterse CMJ, Van der Does D, Zamioudis C, Leon-Reyes A, Van Wees SCM (2012) Hormonal modulation of plant immunity. Annu Rev Cell Dev Biol 28:489–521. https://doi.org/10.1146/annurev-cellbio-092910-154055
Pigllucci M (1996) How organisms respond to environmental changes: from phenotypes to molecules (and vice versa). Trends Ecol Evol 11:168–173
Powell PA, Sanders PR, Tumer N, Fraley RT, Beachy RN (1990) Protection against tobacco mosaic virus infection in transgenic plants requires accumulation of coat protein rather than coat protein RNA sequences. Virology 175:124–130
Prime-A-Plant Group, Conrath U, Beckers GJM, Flors V, García-Agustín P, Jakab G, Mauch F, Newman M-A, Pieterse CMJ, Poinssot B, Pozo MJ, Pugin A, Schaffrath U, Ton J, Wendehenne D, Zimmerli L, Mauch-Mani B (2006) Priming: getting ready for battle. Mol Plant Microbe Interact 19:1062–1071. https://doi.org/10.1094/MPMI-19-1062
Prins M (2003) Broad virus resistance in transgenic plants. Trends Biotechnol 21:373–375. https://doi.org/10.1016/S0167-7799(03)00183-5
Qiu D, Pan X, Wilson IW, Li F, Liu M, Teng W, Zhang B (2009) High throughput sequencing technology reveals that the taxoid elicitor methyl jasmonate regulates microRNA expression in Chinese yew (Taxus chinensis). Gene 436:37–44. https://doi.org/10.1016/j.gene.2009.01.006
Quadrana L, Colot V (2016) Plant transgenerational epigenetics. Annu Rev Genet 50:467–491. https://doi.org/10.1146/annurev-genet-120215-035254
Quintana-Rodriguez E, Duran-Flores D, Heil M, Camacho-Coronel X (2018) Damage-associated molecular patterns (DAMPs) as future plant vaccines that protect crops from pests. Sci Hortic 237:207–220. https://doi.org/10.1016/j.scienta.2018.03.026
Radwan O, Liu Y, Clough SJ (2011) Transcriptional analysis of soybean root response to Fusarium virguliforme, the causal agent of sudden death syndrome. Mol Plant-Microbe Interact 24:958–972. https://doi.org/10.1094/MPMI-11-10-0271
Raja P, Sanville BC, Buchmann RC, Bisaro DM (2008) Viral genome methylation as an epigenetic defense against geminiviruses. J Virol 82:8997–9007. https://doi.org/10.1128/JVI.00719-08
Ramirez-Parra E, Gutierrez C (2007) The many faces of chromatin assembly factor 1. Trends Plant Sci 12:570–576. https://doi.org/10.1016/j.tplants.2007.10.002
Ramirez-Prado JS, Piquerez SJM, Bendahmane A, Hirt H, Raynaud C, Benhamed M (2018) Modify the histone to win the battle: chromatin dynamics in plant–pathogen interactions. Front Plant Sci 9:355. https://doi.org/10.3389/fpls.2018.00355
Rando OJ, Verstrepen KJ (2007) Timescales of genetic and epigenetic inheritance. Cell 128:655–668. https://doi.org/10.1016/j.cell.2007.01.023
Rapicavoli J (2015) Primed for battle: helping plants fight off pathogens by enhancing their immune systems. The Conversation. https://theconversation.com/primed-for-battle-helping-plants-fight-off-pathogens-by-enhancing-their-immune-systems-43689
Rasmann S, Vos MD, Casteel CL, Tian D, Halitschke R, Sun JY, Agrawal AA, Felton GW, Jander G (2012) Herbivory in the previous generation primes plants for enhanced insect resistance. Plant Physiol 158:854–863. https://doi.org/10.1104/pp.111.187831
Reinders J, Wulff BBH, Mirouze M, Marí-Ordóñez A, Dapp M, Rozhon W, Bucher E, Theiler G, Paszkowski J (2009) Compromised stability of DNA methylation and transposon immobilization in mosaic Arabidopsis epigenomes. Genes Dev 23:939–950. https://doi.org/10.1101/gad.524609
Richards EJ (2011) Natural epigenetic variation in plant species: a view from the field. Curr Opin Plant Biol 14:204–209. https://doi.org/10.1016/j.pbi.2011.03.009
Ro S, Yan W (2010) Detection and quantitative analysis of small RNAs by PCR. In: Sioud M (ed) RNA therapeutics: function, design, and delivery, methods in molecular biology. Humana Press, Totowa, NJ, pp 293–303. https://doi.org/10.1007/978-1-60761-657-3_19
Roberts DA (1983) Acquired resistance to Tobacco mosaic virus transmitted to the progeny of hypersensitive Tobacco. Virology 124:161–163. https://doi.org/10.1016/0042-6822(83)90299-4
Rodríguez-Negrete E, Lozano-Durán R, Piedra-Aguilera A, Cruzado L, Bejarano ER, Castillo AG (2013) Geminivirus Rep protein interferes with the plant DNA methylation machinery and suppresses transcriptional gene silencing. New Phytol 199:464–475. https://doi.org/10.1111/nph.12286
Ross AF (1961) Systemic acquired resistance induced by localized virus infections in plants. Virology 14:340–358. https://doi.org/10.1016/0042-6822(61)90319-1
Ruiz-Ferrer V, Voinnet O (2009) Roles of plant small RNAs in biotic stress responses. Annu Rev Plant Biol 60:485–510. https://doi.org/10.1146/annurev.arplant.043008.092111
Sanford JC, Johnston SA (1985) The concept of parasite-derived resistance-deriving resistance genes from the parasite’s own genome. J Theor Biol 113:395–405. https://doi.org/10.1016/S0022-5193(85)80234-4
Schmitz RJ, He Y, Valdés-López O, Khan SM, Joshi T, Urich MA, Nery JR, Diers B, Xu D, Stacey G, Ecker JR (2013) Epigenome-wide inheritance of cytosine methylation variants in a recombinant inbred population. Genome Res 23:1663–1674. https://doi.org/10.1101/gr.152538.112
Schwind N, Zwiebel M, Itaya A, Ding B, Wang M-B, Krczal G, Wassenegger M (2009) RNAi-mediated resistance to Potato spindle tuber viroid in transgenic tomato expressing a viroid hairpin RNA construct. Mol Plant Pathol 10:459–469. https://doi.org/10.1111/j.1364-3703.2009.00546.x
Seo J-K, Wu J, Lii Y, Li Y, Jin H (2013) Contribution of small RNA pathway components in plant immunity. Mol Plant-Microbe Interact 26:617–625. https://doi.org/10.1094/MPMI-10-12-0255-IA
Shattuck VI (1993) Glucosinolates and glucosinolate degradation in seeds from turnip mosaic virus-infected rapid cycle Brassica campestris L. plants. J Exp Bot 44:963–970. https://doi.org/10.1093/jxb/44.5.963
Shen D, Suhrkamp I, Wang Y, Liu S, Menkhaus J, Verreet J-A, Fan L, Cai D (2014) Identification and characterization of microRNAs in oilseed rape (Brassica napus) responsive to infection with the pathogenic fungus Verticillium longisporum using Brassica AA (Brassica rapa) and CC (Brassica oleracea) as reference genomes. New Phytol 204:577–594. https://doi.org/10.1111/nph.12934
Shivaprasad PV, Chen H-M, Patel K, Bond DM, Santos BACM, Baulcombe DC (2012) A microRNA superfamily regulates nucleotide binding site-leucine-rich repeats and other mRNAs. Plant Cell 24:859–874. https://doi.org/10.1105/tpc.111.095380
Simon JM, Giresi PG, Davis IJ, Lieb JD (2012) Using FAIRE (formaldehyde-assisted isolation of regulatory elements) to isolate active regulatory DNA. Nat Protoc 7:256–267. https://doi.org/10.1038/nprot.2011.444
Singh M, Bag SK, Bhardwaj A, Ranjan A, Mantri S, Nigam D, Sharma YK, Sawant SV (2015) Global nucleosome positioning regulates salicylic acid mediated transcription in Arabidopsis thaliana. BMC Plant Biol 15:13. https://doi.org/10.1186/s12870-014-0404-2
Singh P, Yekondi S, Chen P-W, Tsai C-H, Yu C-W, Wu K, Zimmerli L (2014a) Environmental history modulates Arabidopsis pattern-triggered immunity in a HISTONE ACETYLTRANSFERASE1–dependent manner[C][W]. Plant Cell 26:2676–2688. https://doi.org/10.1105/tpc.114.123356
Singh V, Roy S, Singh D, Nandi AK (2014b) Arabidopsis FLOWERING LOCUS D influences systemic-acquired-resistance-induced expression and histone modifications of WRKY genes. J Biosci 39:119–126. https://doi.org/10.1007/s12038-013-9407-7
Singh P, Dave A, Vaistij FE, Worrall D, Holroyd GH, Wells JG, Kaminski F, Graham IA, Roberts MR (2017) Jasmonic acid-dependent regulation of seed dormancy following maternal herbivory in Arabidopsis. New Phytol 214:1702–1711. https://doi.org/10.1111/nph.14525
Singh V, Roy S, Giri MK, Chaturvedi R, Chowdhury Z, Shah J, Nandi AK (2013) Arabidopsis thaliana FLOWERING LOCUS D is required for systemic acquired resistance. Mol Plant-Microbe Interact 26:1079–1088. https://doi.org/10.1094/MPMI-04-13-0096-R
Slaughter A, Daniel X, Flors V, Luna E, Hohn B, Mauch-Mani B (2012) Descendants of primed Arabidopsis plants exhibit resistance to biotic stress. Plant Physiol 158:835–843. https://doi.org/10.1104/pp.111.191593
Stassen JHM, López A, Jain R, Pascual-Pardo D, Luna E, Smith LM, Ton J (2018) The relationship between transgenerational acquired resistance and global DNA methylation in Arabidopsis. Sci Rep 8:14761. https://doi.org/10.1038/s41598-018-32448-5
Stenberg JA (2017) A conceptual framework for integrated pest management. Trends Plant Sci 22:759–769. https://doi.org/10.1016/j.tplants.2017.06.010
Strapasson P, Pinto-Zevallos DM, Paudel S, Rajotte EG, Felton GW, Zarbin PHG (2014) Enhancing plant resistance at the seed stage: low concentrations of methyl jasmonate reduce the performance of the leaf miner Tuta absoluta but do not alter the behavior of its predator Chrysoperla externa. J Chem Ecol 40:1090–1098. https://doi.org/10.1007/s10886-014-0503-4
Taha O, Farouk I, Abdallah A, Abdallah NA (2016) Use of posttranscription gene silencing in squash to induce resistance against the Egyptian isolate of the squash leaf curl virus. Int J Genom 2016:6053147. https://doi.org/10.1155/2016/6053147
Takuno S, Ran J-H, Gaut BS (2016) Evolutionary patterns of genic DNA methylation vary across land plants. Nat Plants 2:15222. https://doi.org/10.1038/nplants.2015.222
Talbert PB, Henikoff S (2014) Environmental responses mediated by histone variants. Trends Cell Biol 24:642–650. https://doi.org/10.1016/j.tcb.2014.07.006
Tateda C, Zhang Z, Shrestha J, Jelenska J, Chinchilla D, Greenberg JT (2014) Salicylic acid regulates Arabidopsis microbial pattern receptor kinase levels and signaling. Plant Cell 26:4171–4187. https://doi.org/10.1105/tpc.114.131938
terHorst CP, Lau JA (2012) Direct and indirect transgenerational effects alter plant-herbivore interactions. Evol Ecol 26:1469–1480. https://doi.org/10.1007/s10682-012-9560-8
Tsuchiya T, Eulgem T (2013) An alternative polyadenylation mechanism coopted to the Arabidopsis RPP7 gene through intronic retrotransposon domestication. Proc Natl Acad Sci 110:E3535–E3543. https://doi.org/10.1073/pnas.1312545110
Tsuda K, Sato M, Glazebrook J, Cohen JD, Katagiri F (2008) Interplay between MAMP-triggered and SA-mediated defense responses. Plant J 53:763–775. https://doi.org/10.1111/j.1365-313X.2007.03369.x
Turck F, Coupland G (2014) Natural variation in epigenetic gene regulation and its effects on plant developmental traits. Evolution 68:620–631. https://doi.org/10.1111/evo.12286
Urbanek M, Nawrocka A, Krzyzosiak W, Urbanek MO, Nawrocka AU, Krzyzosiak WJ (2015) Small RNA detection by in situ hybridization methods. Int J Mol Sci 16:13259–13286. https://doi.org/10.3390/ijms160613259
Van der Ent S, Hulten MV, Pozo MJ, Czechowski T, Udvardi MK, Pieterse CMJ, Ton J (2009) Priming of plant innate immunity by rhizobacteria and β-aminobutyric acid: differences and similarities in regulation. New Phytol 183:419–431. https://doi.org/10.1111/j.1469-8137.2009.02851.x
van der Krol AR, Mur LA, Beld M, Mol JN, Stuitje AR (1990) Flavonoid genes in petunia: addition of a limited number of gene copies may lead to a suppression of gene expression. Plant Cell 2:291–299
Van Holde KE, Sahasrabuddhe CG, Shaw BR (1974) A model for particulate structure in chromatin. Nucleic Acids Res 1:1579–1586
van Hulten M, Pelser M, van Loon LC, Pieterse CMJ, Ton J (2006) Costs and benefits of priming for defense in Arabidopsis. Proc Natl Acad Sci U S A 103:5602–5607. https://doi.org/10.1073/pnas.0510213103
Vaughn MW, Tanurdžić M, Lippman Z, Jiang H, Carrasquillo R, Rabinowicz PD, Dedhia N, McCombie WR, Agier N, Bulski A, Colot V, Doerge RW, Martienssen RA (2007) Epigenetic natural variation in Arabidopsis thaliana. PLoS Biol 5:e174. https://doi.org/10.1371/journal.pbio.0050174
Verhoeven KJF, Jansen JJ, van Dijk PJ, Biere A (2010) Stress-induced DNA methylation changes and their heritability in asexual dandelions. New Phytol 185:1108–1118. https://doi.org/10.1111/j.1469-8137.2009.03121.x
Verkest A, Byzova M, Martens C, Willems P, Verwulgen T, Slabbinck B, Rombaut D, de Velde JV, Vandepoele K, Standaert E, Peeters M, Lijsebettens MV, Breusegem FV, Block MD (2015) Selection for improved energy use efficiency and drought tolerance in canola results in distinct transcriptome and epigenome changes. Plant Physiol 168:1338–1350. https://doi.org/10.1104/pp.15.00155
Voinnet O (2005) Non-cell autonomous RNA silencing. FEBS Lett 579:5858–5871. https://doi.org/10.1016/j.febslet.2005.09.039
Voinnet O (2008) Post-transcriptional RNA silencing in plant–microbe interactions: a touch of robustness and versatility. Curr Opin Plant Biol 11:464–470. https://doi.org/10.1016/j.pbi.2008.04.006
Vos IA, Moritz L, Pieterse CMJ, Van Wees SCM (2015) Impact of hormonal crosstalk on plant resistance and fitness under multi-attacker conditions. Front Plant Sci 6:639. https://doi.org/10.3389/fpls.2015.00639
Waddington CH (2012) The epigenotype. Int J Epidemiol 41:10–13. https://doi.org/10.1093/ije/dyr184
Walley JW, Rowe HC, Xiao Y, Chehab EW, Kliebenstein DJ, Wagner D, Dehesh K (2008) The chromatin remodeler SPLAYED regulates specific stress signaling pathways. PLoS Pathog 4:e1000237. https://doi.org/10.1371/journal.ppat.1000237
Walters DR, Paterson L (2012) Parents lend a helping hand to their offspring in plant defence. Biol Lett 8:871–873. https://doi.org/10.1098/rsbl.2012.0416
Wang B, Li F, Huang C, Yang X, Qian Y, Xie Y, Zhou X (2014) V2 of tomato yellow leaf curl virus can suppress methylation-mediated transcriptional gene silencing in plants. J Gen Virol 95:225–230. https://doi.org/10.1099/vir.0.055798-0
Wang C, Gao F, Wu J, Dai J, Wei C, Li Y (2010) Arabidopsis putative deacetylase AtSRT2 regulates basal defense by suppressing PAD4, EDS5 and SID2 expression. Plant Cell Physiol 51:1291–1299. https://doi.org/10.1093/pcp/pcq087
Wang M, Weiberg A, Lin F-M, Thomma BPHJ, Huang H-D, Jin H (2016) Bidirectional cross-kingdom RNAi and fungal uptake of external RNAs confer plant protection. Nat Plants 2:16151. https://doi.org/10.1038/nplants.2016.151
Wang M, Thomas N, Jin H (2017) Cross-kingdom RNA trafficking and environmental RNAi for powerful innovative pre- and post-harvest plant protection. Curr Opin Plant Biol 38:133–141. https://doi.org/10.1016/j.pbi.2017.05.003
Warren JM, Covert SF (2004) Differential expression of pine and Cronartium quercuum f. sp. fusiforme genes in fusiform rust galls. Appl Environ Microbiol 70:441–451. https://doi.org/10.1128/AEM.70.1.441-451.2004
Waterhouse PM, Wang M-B, Lough T (2001) Gene silencing as an adaptive defence against viruses [WWW Document]. Nature 411(6839):834–842. https://doi.org/10.1038/35081168
Weake VM, Workman JL (2008) Histone ubiquitination: triggering gene activity. Mol Cell 29:653–663. https://doi.org/10.1016/j.molcel.2008.02.014
Weiberg A, Wang M, Lin F-M, Zhao H, Zhang Z, Kaloshian I, Huang H-D, Jin H (2013) Fungal small RNAs suppress plant immunity by hijacking host RNA interference pathways. Science 342:118–123. https://doi.org/10.1126/science.1239705
Wibowo A, Becker C, Durr J, Price J, Spaepen S, Hilton S, Putra H, Papareddy R, Saintain Q, Harvey S, Bending GD, Schulze-Lefert P, Weigel D, Gutierrez-Marcos J (2018) Partial maintenance of organ-specific epigenetic marks during plant asexual reproduction leads to heritable phenotypic variation. Proc Natl Acad Sci 115:E9145–E9152. https://doi.org/10.1073/pnas.1805371115
Wilkinson SW, Pastor V, Paplauskas S, Pétriacq P, Luna E (2018) Long-lasting β-aminobutyric acid-induced resistance protects tomato fruit against Botrytis cinerea. Plant Pathol 67:30–41. https://doi.org/10.1111/ppa.12725
Wong J, Gao L, Yang Y, Zhai J, Arikit S, Yu Y, Duan S, Chan V, Xiong Q, Yan J, Li S, Liu R, Wang Y, Tang G, Meyers BC, Chen X, Ma W (2014) Roles of small RNAs in soybean defense against Phytophthora sojae infection. Plant J 79:928–940. https://doi.org/10.1111/tpj.12590
Worrall D, Holroyd GH, Moore JP, Glowacz M, Croft P, Taylor JE, Paul ND, Roberts MR (2012) Treating seeds with activators of plant defence generates long-lasting priming of resistance to pests and pathogens. New Phytol 193:770–778. https://doi.org/10.1111/j.1469-8137.2011.03987.x
Xin M, Wang Y, Yao Y, Xie C, Peng H, Ni Z, Sun Q (2010) Diverse set of microRNAs are responsive to powdery mildew infection and heat stress in wheat (Triticum aestivum L.). BMC Plant Biol 10:123. https://doi.org/10.1186/1471-2229-10-123
Xu J, Li J, Cui L, Zhang T, Wu Z, Zhu P-Y, Meng Y-J, Zhang K-J, Yu X-Q, Lou Q-F, Chen J-F (2017) New insights into the roles of cucumber TIR1 homologs and miR393 in regulating fruit/seed set development and leaf morphogenesis. BMC Plant Biol 17:130. https://doi.org/10.1186/s12870-017-1075-6
Yang X, Xie Y, Raja P, Li S, Wolf JN, Shen Q, Bisaro DM, Zhou X (2011) Suppression of methylation-mediated transcriptional gene silencing by βC1-SAHH protein interaction during geminivirus-betasatellite infection. PLoS Pathog 7:e1002329. https://doi.org/10.1371/journal.ppat.1002329
Yang Z, Ebright YW, Yu B, Chen X (2006) HEN1 recognizes 21–24 nt small RNA duplexes and deposits a methyl group onto the 2′ OH of the 3′ terminal nucleotide. Nucleic Acids Res 34:667–675. https://doi.org/10.1093/nar/gkj474
Yi H, Richards EJ (2007) A cluster of disease resistance genes in Arabidopsis is coordinately regulated by transcriptional activation and RNA silencing. Plant Cell 19:2929–2939. https://doi.org/10.1105/tpc.107.051821
Yi H, Richards EJ (2009) Gene duplication and hypermutation of the pathogen resistance gene SNC1 in the Arabidopsis bal variant. Genetics 183:1227–1234. https://doi.org/10.1534/genetics.109.105569
Yin Z, Li Y, Han X, Shen F (2012) Genome-wide profiling of miRNAs and other small non-coding RNAs in the Verticillium dahliae–inoculated cotton roots. PLoS One 7:e35765. https://doi.org/10.1371/journal.pone.0035765
Yu A, Lepère G, Jay F, Wang J, Bapaume L, Wang Y, Abraham A-L, Penterman J, Fischer RL, Voinnet O, Navarro L (2013) Dynamics and biological relevance of DNA demethylation in Arabidopsis antibacterial defense. Proc Natl Acad Sci 110:2389–2394. https://doi.org/10.1073/pnas.1211757110
Zemach A, Kim MY, Hsieh P-H, Coleman-Derr D, Eshed-Williams L, Thao K, Harmer SL, Zilberman D (2013) The Arabidopsis nucleosome remodeler DDM1 allows DNA methyltransferases to access H1-containing heterochromatin. Cell 153:193–205. https://doi.org/10.1016/j.cell.2013.02.033
Zhai J, Liu J, Liu B, Li P, Meyers BC, Chen X, Cao X (2008) Small RNA-directed epigenetic natural variation in Arabidopsis thaliana. PLoS Genet 4:e1000056. https://doi.org/10.1371/journal.pgen.1000056
Zhang C, Hsieh T-F (2013) Heritable epigenetic variation and its potential applications for crop improvement. Plant Breed Biotechnol 1:307–319. https://doi.org/10.9787/PBB.2013.1.4.307
Zhang H, Deng X, Miki D, Cutler S, La H, Hou Y-J, Oh J, Zhu J-K (2012) Sulfamethazine suppresses epigenetic silencing in Arabidopsis by impairing folate synthesis[W]. Plant Cell 24:1230–1241. https://doi.org/10.1105/tpc.112.096149
Zhang W, Gao S, Zhou X, Chellappan P, Chen Z, Zhou X, Zhang X, Fromuth N, Coutino G, Coffey M, Jin H (2011) Bacteria-responsive microRNAs regulate plant innate immunity by modulating plant hormone networks. Plant Mol Biol 75:93–105. https://doi.org/10.1007/s11103-010-9710-8
Zhang Y, Li D, Zhang H, Hong Y, Huang L, Liu S, Li X, Ouyang Z, Song F (2015) Tomato histone H2B monoubiquitination enzymes SlHUB1 and SlHUB2 contribute to disease resistance against Botrytis cinerea through modulating the balance between SA- and JA/ET-mediated signaling pathways. BMC Plant Biol 15:252. https://doi.org/10.1186/s12870-015-0614-2
Zhang Y-Y, Fischer M, Colot V, Bossdorf O (2013) Epigenetic variation creates potential for evolution of plant phenotypic plasticity. New Phytol 197:314–322. https://doi.org/10.1111/nph.12010
Zhao H, Sun R, Albrecht U, Padmanabhan C, Wang A, Coffey MD, Girke T, Wang Z, Close TJ, Roose M, Yokomi RK, Folimonova S, Vidalakis G, Rouse R, Bowman KD, Jin H (2013) Small RNA profiling reveals phosphorus deficiency as a contributing factor in symptom expression for citrus huanglongbing disease. Mol Plant 6:301–310. https://doi.org/10.1093/mp/sst002
Zhao M, San León D, Delgadillo MO, García JA, Simón-Mateo C (2014) Virus-induced gene silencing in transgenic plants: transgene silencing and reactivation associate with two patterns of transgene body methylation. Plant J 79:440–452. https://doi.org/10.1111/tpj.12579
Zhao W, Li Z, Fan J, Hu C, Yang R, Qi X, Chen H, Zhao F, Wang S (2015) Identification of jasmonic acid-associated microRNAs and characterization of the regulatory roles of the miR319/TCP4 module under root-knot nematode stress in tomato. J Exp Bot 66:4653–4667. https://doi.org/10.1093/jxb/erv238
Zhou C, Zhang L, Duan J, Miki B, Wu K (2005) HISTONE DEACETYLASE19 is involved in jasmonic acid and ethylene signaling of pathogen response in Arabidopsis. Plant Cell 17:1196–1204. https://doi.org/10.1105/tpc.104.028514
Zhou L, Cheng X, Connolly BA, Dickman MJ, Hurd PJ, Hornby DP (2002) Zebularine: a novel DNA methylation inhibitor that forms a covalent complex with DNA methyltransferases. J Mol Biol 321:591–599
Zhu J-K (2009) Active DNA demethylation mediated by DNA glycosylases. Annu Rev Genet 43:143–166. https://doi.org/10.1146/annurev-genet-102108-134205
Zou B, Yang D-L, Shi Z, Dong H, Hua J (2014) Monoubiquitination of histone 2B at the disease resistance gene locus regulates its expression and impacts immune responses in Arabidopsis. Plant Physiol 165:309–318. https://doi.org/10.1104/pp.113.227801
Zou B, Sun Q, Zhang W, Ding Y, Yang D-L, Shi Z, Hua J (2017) The Arabidopsis chromatin-remodeling factor CHR5 regulates plant immune responses and nucleosome occupancy. Plant Cell Physiol 58:2202–2216. https://doi.org/10.1093/pcp/pcx155
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Roberts, M.R., López Sánchez, A. (2019). Plant Epigenetic Mechanisms in Response to Biotic Stress. In: Alvarez-Venegas, R., De-la-Peña, C., Casas-Mollano, J. (eds) Epigenetics in Plants of Agronomic Importance: Fundamentals and Applications. Springer, Cham. https://doi.org/10.1007/978-3-030-14760-0_2
Download citation
DOI: https://doi.org/10.1007/978-3-030-14760-0_2
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-14759-4
Online ISBN: 978-3-030-14760-0
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)