Abstract
Autism Spectrum Disorder (ASD) is a complex neurological condition characterized by a triad of signs: stereotyped behaviors, verbal and non-verbal communication problems and troubles in social interaction. The scientific community has been interested on quantifying anatomical brain alterations of this disorder to correlate the clinical signs with brain tissue changes. This work presents a fully automatic method to find out brain differences between patients diagnosed with autism and control subjects. After pre-processing, a template (MNI152) is registered to each evaluated brain, obtaining a set of segmented regions. Each region is mapped into a 2D collage image which is decomposed by the Zernike Moments, obtaining magnitude and phase. These features are then used to train, region per region, a binary SVM classifier. The method was evaluated in a children population, aged from 6 to 12 years, from the public database Autism Brain Imaging Data Exchange. The AUC values for the most representative brain region were 77% for ABIDE I and 76% for ABIDE II, demonstrating the robustness of the method.
You have full access to this open access chapter, Download conference paper PDF
Similar content being viewed by others
Keywords
1 Introduction
ASD constitutes a group of neurological alterations that represents a wide variety of clinical expressions. About 1 in 59 children has been identified with ASD according to CDC’s Autism and Developmental Disabilities Monitoring (ADDM) Network. The prevalence in children is higher in boys than in girls, in a proportion of 4 to 1 [2].
Although there exist a large number of syndromes related to autism, the diagnosis remains until now strictly clinical. A reliable diagnosis requires availability of therapists or physicians, resulting in a bottleneck that difficult early detection of this disorder [14]. In addition, most of early signs of brain function alteration are not specific and autistic signs may be observed in patients with no disorder, a source of confusion for many clinicians [31].
Currently, ASD diagnosis is performed using neuro-psychological tests that evaluate the patient-environment interaction and high cerebral functions. These tests register several clinician observations, making diagnosis subjective. The gold standard [12] in terms of diagnosis is the Autism Diagnostic Observation Schedule (ADOS) [24] and/or the Autism Diagnostic Interview (ADI-R) [25]. However the probability of misdiagnosis is high [15] since the clinician may mislead descriptive labels and inevitably introduces bias by her/his judgment [29]. Clinicians’ experience may facilitate ASD diagnosis before the second year, yet the average diagnosis age is above 3 years, which suggests many children may not be diagnosed at all. Early ASD diagnosis is critical because earlier treatments can reduce the degree of deterioration and improve the function of both patients and carers [1]. Curiously, even if modern medical images are at the very base of many decisions, in these kind of pathologies their role is still marginal [36]. Neuro-imaging could be useful to evaluate relationship between the different areas, regions or set of cerebral regions and the cognitive and functional signs that patients present, that is to say, analysis of the structure offers new possibilities of correlating brain changes or alterations at the functional level with ASD signs.
Different researches have been done with the purpose of correlating functional alterations presented in ASD and the anatomical structures. The first approaches to this theory date back in 1991, when Kemper and Bauman analyzed brains of six autistic patients, finding main alterations at the level of the limbic system, cerebellum and inferior olive. These brains showed no major morphological changes, yet it was reported a decrease in the neuronal cell size and an increase of the neuronal density at the level of the amygdala and other limbic structures when comparing to controls [20]. Recent studies have used brain MRI to classify patients with ASD, resulting that main changes were in regions like the basal ganglia, corpus callosum, hippocampus, amygdala and thalamus [3, 11, 22]. Participants in this study aged 6–15 years, volunteer ASD and control subjects [18].
There is a strong ADS relationship with brain areas responsible for normal language development such as Broca’s area and Wernicke’s area at a level of verbal and non-verbal communication. There is evidence of increase in the volume of the right and left temporal gyrus in T1-MRI studies [5, 16]. Other research has used more than three different classification techniques (RF, SVM and GBM) using as main feature the size of the cortical and sub-cortical regions, reporting a sensitivity of 57% and 64% of specificity for the binary classification task [19].
ASD characterization is difficult by the high variability between different medical studies and children development. An automatic morphometry approach has the advantage of including additional information to support early diagnosis, but some approaches based on voxel size, shape, or volume [30] ignore local and regional dependencies. A main contribution of the present research is a fully automatic morphometric method that establishes region differences by using local shape information. The approach starts by a brain segmentation using a known template [27]. Once the brain is segmented, each region is characterized by the magnitude and phase of the region Zernike Moments which inputs a standard classifier. Classification results using such feature space provide representative regions that differentiate between ASD patients and control subjects.
2 Methods
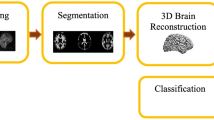
The proposed method is divided in five phases, as illustrated in Fig. 1, starting by firstly pre-processing each volume to eliminate differences coming from acquisition protocols and devices. Afterwards, an atlas [6] is elastically registered to each of the cases with the aim of segmenting brain regions. Each segmented region is then arranged in a two-dimensional collage of images, constructed by sequentially copying each slice into a 2D frame, from the top to the bottom of the region. The obtained 2D image is used as input to calculate the Zernike moments per region. The resultant magnitude and phase are used as features that are challenged to a classic classification task by means of a conventional Support Vector Machine (SVM).
2.1 Pre-processing
Each volume undergoes a pre-processing phase composed by two steps, first an intensity correction to reduce intra-site variability since images were not obtained by the same device and, second a brain extraction to remove skull, spinal cord and eye holes. Intensity correction was performed by the FSLMATH tool provided by the Oxford University [28], which corrects the bias field and normalizes each volume. Afterward, brain extraction was done by using BET (Brain Extraction Tool), which removes non-related brain tissues by using the histogram of the image and triangular tessellations [34].
2.2 Registration and Segmentation
The registration phase was carried out to obtain the corresponding regions from the Harvard-Oxford Atlas [26]. For so doing, the MNI152 template [27] was elastically registered to each brain. The process begins with an affine registration with 12 freedom degrees and a correction for spatial errors computed by means of the FLIRT Tool (FMRIB’s Linear Image Registration Tool) [17]. Then, a finer result is obtained by performing elastic registration using the FNIRT [28] tool with a quadratic spline which optimizes the processing time and ensures that transformation is as accurate as possible.
Pipeline of proposed strategy (1) pre-processing each volume, (2) a particular atlas is elastically registered to each case and the resultant volume is segmented to 117 regions, (3) each 3D region is mapped into a 2D-collage image, (4) the Zernike moments are computed per region, taking the magnitude and phase per moment as features, and (5) finally a classification task between ASD patients and control subjects to establish discriminative regions is performed.
2.3 Anatomic Region Representation and Collage
In the present work each segmented region, a volume structure, is represented as a 2D image that contains each axial plane. This collage is built by placing each slice in an squared resolution image, as seen in Fig. 2. However, note that each volume slice is fit to the size of the region, that is to say, while the volume resolution is \(512 \times 512\), the resulting image resolution for a particular region slice could be \(45 \times 57\). This representation is convenient because it allows to describe structural shape changes on each slice without any loss of information from the 3D original volume.
2.4 Region-Based Characterization: Zernike Moments
Zernike moments are considered as shape descriptors by performing a multiscale frequency analysis which is usually represented as a pyramid, where the scales are the different pyramid levels and frequency analysis (repetitions) is performed through each of the scales. The complex 2D Zernike moments of order m and n repetitions are defined in the unitary circle [23] as:
where \(f(r,\theta )\) stands for the image intensity function, \(V^{*}_{mn} (r,\theta )\) corresponds to the complex conjugate of Zernike polynomial \(V_{mn} (r,\theta )\), and m and n are both integers related as:
For processing a 2D image, Zernike moments are computed by using the discretized form, as illustrated in Eq. 3:
In this work, a morphometric analysis was performed by transforming the 2D image into the Zernike space, a representation that has demonstrated describe complex shapes [23]. This representation allows to characterize each brain region based on shape differences between ASD patients and control subjects. Each brain region was described by using 9 scales (55 Zernike moments), where each moment consists of magnitude and phase components, obtaining at the end a descriptor of 110 features.
2.5 Classifier
Support Vector Machines (SVM) are a set of related methods for supervised learning, applicable to both classification and regression problems. A SVM classifier sets a maximum-margin hyperplane that lies in a transformed input space and splits the space, while maximizing the distance to the nearest sample examples. The parameters of the hyperplane solution are derived from a quadratic programming optimization problem [33]. For this investigation, the SVM algorithm was trained with the feature vectors obtained from previous phase.
3 Experiments and Results
The proposed strategy performance was evaluated by using a open access database [8]. The most important result is the identification of a set of brain regions that better differentiate the two classes, namely ASD patients and control subjects.
3.1 Database
For this study, brain T1-MRI cases were used, available in the Autism Brain Imaging Data Exchange ABIDE I and ABIDE II (first and second version) [7, 8]. ABIDE databases contain 2226 cases (ASD individuals and typical controls aged 5–64 years), scanned across 17 medical centers. For this investigation, two subsamples were used, including only male cases with voxel size of \(1 \times 1 \times 1\) and ages between 6 and 12 years, based on the criterion of growth and development of the cephalic mass in children up to 12.1 years [13], aiming to get an homogeneous population.
The evaluation was then carried out with 196 subjects (98 individuals diagnosed with ASD and 98 controls), 68 from ABIDE I database (Table 1) and 128 from ABIDE II database (Table 2).
3.2 Registration and Segmentation
The Harvard - Oxford atlas [26] was used as the reference space to segment each brain of the experimental group into 96 cortical (48 per hemisphere) and 17 sub-cortical regions. The lateralized template was rigidly and elastically registered to each brain, and the resultant transformation matrix was applied to the atlas parcellation, obtaining the set of brain regions per case. Registration was assessed by measuring the overlapping percentage (Dice Score coefficient [9] metrics) between both complete brain volumes, the registered MNI-152 and each case:
where: X is the MNI152 template and Y the evaluated brain. Once the elastic registration is performed, each brain is compared with the deformed template to verify that there is a high correspondence between the complete brain volumes. Registration results are shown in Tables 3 and 4 respectively.
3.3 Region-Based Characterization: Zernike Moments
In this phase of the proposed approach, there were computed 9 scales of the Zernike transformation, providing the first 55 moments of such representation space. This information corresponds to the shape orientations for each used region, and then magnitude and phase per calculated moment are concatenated on a matrix. Zernike moments were calculated the for the \(n \times n\) 2D region mosaic-image described in the Sect. 2.3, using Matlab [32, 35]. Figure 3 illustrates the complex parameters (magnitude and phase) provided by the Zernike moments for a particular region in the polar space.
Representation of complex parameters (magnitude and phase) for a cortical region (Right Frontal Orbital Cortex) provided by Zernike Moments. In (a) is presented the distribution of the complex values in a control subject and in (b) the distribution of the complex values in a autistic patient evaluated on the same region.
3.4 Classifier
For evaluating the performance of the proposed approach by using a Support Vector Machine (SVM) classifier, a Radial Basis Function (RBF) was selected as kernel because of the high dimensionality of the feature vector per region. A 10-fold cross-validation scheme was used to train and test the constructed model, and four metrics are reported, namely Area Under the Curve (AUC), sensitivity, specificity and F-Score. It is noteworthy that the regions that showed greater accuracy were cortical regions, especially in those that are related to the normal language development, which play an important role in brain morphological changes on autistic patients such as reported in previous researches [10]. Table 5 presents the classification results using the first 55 Zernike Moments for cortical brain regions.
Figure 4 shows the Area Under Curve (AUC) graph for the most significant brain region for ABIDE I (Right Frontal Orbital Cortex) and for ABIDE II (Right Occipital Fusiform Gyrus), found with Zernike Moments characterization.
4 Discussion
In the present study, we examined the orientations of the shape from each brain region using the Zernike Moments, in a large sample of children with ASD (Autism Spectrum Disorder), relative to TC (Typical Control). This representation does not find differences at the cellular level, but on the surface and how they are related to the abnormal growth of each region. Frontal Orbital Cortex is highly related to affective functions, decision-making and sensory integration [21]. It is possible to associate alterations in this structure with one of the signs present in autism related to communication and social interaction. [10].
Finally, we found associations with severity symptom within a thalamic surface area cluster. These findings suggest that there are subtle differences in subcortical morphology in ASD. Although this study was cross-sectional, our findings also suggest that there may be atypical developmental of intellectual function and performance deficits in ASD due to this atypical growth [4].
5 Conclusions and Future Work
This work presents a method for classifying patients diagnosed with ASD and how the anatomy of their brains differs from control subjects in particular regions. The variability of the disorder and the methods used by physicians for diagnosis is not completely reliable. The method used in this research works with high level features in MRI in order to represent the information as orientations in brain shape. The obtained results correspond to regions reported in state-of-the-art methods focused on image analysis based on other high level features. Cortical regions remain relevant in the study of autism due to anatomical variability of the brain, especially those related to the social interaction and communication. As a future work, we would like to make an inter-class classification to automatically determine the existing classes in autism spectrum disorder as Asperger, Classical Autism, Rett (present only in women) and other non-invasive developmental disorders described by the DSM-V guide.
References
Alexeeff, S.E., et al.: Medical conditions in the first years of life associated with future diagnosis of ASD in children. J. Autism Dev. Disord. 47(7), 2067–2079 (2017)
Baio, J., et al.: Prevalence of autism spectrum disorder among children aged 8 years-autism and developmental disabilities monitoring network, 11 sites, United States, 2014. MMWR Surveill Summ. 67(6), 1 (2018)
Brambilla, P., Hardan, A., di Nemi, S.U., Perez, J., Soares, J.C., Barale, F.: Brain anatomy and development in autism: review of structural mri studies. Brain Res. Bull. 61(6), 557–569 (2003)
Dager, S.R., et al.: Shape mapping of the hippocampus in young children with autism spectrum disorder. Am. J. Neuroradiol. 28(4), 672–677 (2007)
De Fossé, L., et al.: Language-association cortex asymmetry in autism and specific language impairment. Ann. Neurol. 56(6), 757–766 (2004)
Desikan, R.S., et al.: An automated labeling system for subdividing the human cerebral cortex on MRI scans into gyral based regions of interest. Neuroimage 31(3), 968–980 (2006)
Di Martino, A., et al.: Enhancing studies of the connectome in autism using the autism brain imaging data exchange II. Sci. Data 4, 170010 (2017)
Di Martino, A., et al.: The autism brain imaging data exchange: towards a large-scale evaluation of the intrinsic brain architecture in autism. Mol. Psychiatry 19(6), 659–667 (2014)
Dice, L.R.: Measures of the amount of ecologic association between species. Ecology 26(3), 297–302 (1945)
Dichter, G.S., Felder, J.N., Bodfish, J.W.: Autism is characterized by dorsal anterior cingulate hyperactivation during social target detection. Soc. Cogn. Affect. Neurosci. 4(3), 215–226 (2009)
Ecker, C., et al.: Describing the brain in autism in five dimensions-magnetic resonance imaging-assisted diagnosis of autism spectrum disorder using a multiparameter classification approach. J. Neurosci. 30(32), 10612–10623 (2010)
Falkmer T., Anderson K., F.M., C., H.: Diagnostic procedures in autism spectrum disorders: a systematic literature review. Eur. Child Adolesc. Psychiatry 22(10), 329–340 (2013)
Giedd, J.N.: Structural magnetic resonance imaging of the adolescent brain. Ann. N. Y. Acad. Sci. 1021(1), 77–85 (2004)
Ginn, N.C., Clionsky, L.N., Eyberg, S.M., Warner-Metzger, C., Abner, J.P.: Child-directed interaction training for young children with autism spectrum disorders: parent and child outcomes. J. Clin. Child Adolesc. Psychol. 46(1), 101–109 (2017)
Hedley, D., Brewer, N., Nevill, R., Uljarević, M., Butter, E., Mulick, J.A.: The relationship between clinicians’ confidence and accuracy, and the influence of child characteristics, in the screening of autism spectrum disorder. J. Autism Dev. Disord. 46(7), 2340–2348 (2016)
Herbert, M.R., et al.: Abnormal asymmetry in language association cortex in autism. Ann. Neurol. 52(5), 588–596 (2002)
Jenkinson, M., Bannister, P., Brady, M., Smith, S.: Improved optimisation for the robust and accurate linear registration and motion correction of brain images. NeuroImage 17(2), 825–841 (2002)
Jiao, Y., Chen, R., Ke, X., Chu, K., Lu, Z., Herskovits, E.H.: Predictive models of autism spectrum disorder based on brain regional cortical thickness. Neuroimage. 50(2), 589–599 (2010)
Katuwal, G.J., Cahill, N.D., Baum, S.A., Michael, A.M.: The predictive power of structural MRI in autism diagnosis. In: 2015 37th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), pp. 4270–4273, August 2015. https://doi.org/10.1109/EMBC.2015.7319338
Kemper, T.L., Bauman, M.L.: The contribution of neuropathologic studies to the understanding of autism. Neurol. Clin. 11(1), 175–187 (1993)
Kringelbach, M.L.: The human orbitofrontal cortex: linking reward to hedonic experience. Nat. Rev. Neurosci. 6(9), 691 (2005)
Lauvin, M.A., et al.: Functional morphological imaging of autism spectrum disorders: current position and theories proposed. Diagn. Interv. Imaging 93(3), 139–147 (2012). http://www.sciencedirect.com/science/article/pii/S2211568412000587
Liu, M., He, Y., Ye, B.: Image Zernike moments shape feature evaluation based on image reconstruction. Geo-Spat. Inf. Sci. 10(3), 191–195 (2007)
Lord, C., et al.: The autism diagnostic observation schedule-generic: a standard measure of social and communication deficits associated with the spectrum of autism. J. Autism Dev. Disord. 30(3), 205–223 (2000)
Lord, C., Rutter, M., Le Couteur, A.: Autism diagnostic interview-revised: a revised version of a diagnostic interview for caregivers of individuals with possible pervasive developmental disorders. J. Autism Dev. Disord. 24(5), 659–685 (1994)
Makris, N., et al.: Decreased volume of left and total anterior insular lobule in schizophrenia. Schizophr. Res. 83(2), 155–171 (2006)
Mazziotta, J.C., Toga, A.W., Evans, A., Fox, P., Lancaster, J.: A probabilistic atlas of the human brain: theory and rationale for its development: the international consortium for brain mapping (ICBM). Neuroimage 2(2), 89–101 (1995)
Woolrich, M.W.: Bayesian analysis of neuroimaging data in FSL. Neuroimaging 45, S173–S186 (2009)
de la Paz, M.P., Arroyo, M.J.F., Aguilera, E.T., Muñoz, L.B.: Investigación epidemiológica en el autismo: una visión integradora. Revista de Neurología 40(Suppl. 1), S191–S198 (2005)
Riddle, K., Cascio, C.J., Woodward, N.D.: Brain structure in autism: a voxel-based morphometry analysis of the autism brain imaging database exchange (ABIDE). Brain Imaging Behav. 11, 1–11 (2016)
Rodríguez-Barrionuevo, A., Rodríguez-Vives, M.: Diagnóstico clínico del autismo. Revista de Neurología 34(1), 72–77 (2002)
Saki, F., Tahmasbi, A., Soltanian-Zadeh, H., Shokouhi, S.B.: Fast opposite weight learning rules with application in breast cancer diagnosis. Comput. Biol. Med. 43(1), 32–41 (2013)
Shmilovici, A.: Support vector machines. In: Maimon, O., Rokach, L. (eds.) Data Mining and Knowledge Discovery Handbook, pp. 231–247. Springer, Boston (2009). https://doi.org/10.1007/978-0-387-09823-4_12
Smith, S.M.: Fast robust automated brain extraction. Hum. Brain Mapp. 17(3), 143–155 (2002)
Tahmasbi, A., Saki, F., Shokouhi, S.B.: Classification of benign and malignant masses based on Zernike moments. Comput. Biol. Med. 41(8), 726–735 (2011)
Tomás-Vila, M.: Rendimiento del estudio diagnóstico del autismo. la aportación de la neuroimagen, las pruebas metabólicas y los estudios genéticos. Revista de neurología 38(1), 15–20 (2004)
Acknowledgments
This work is supported by INV-ING-2639 project, titled “Caracterización y cuantificación de anomalías estructurales en trastornos del espectro autista a través de imágenes de RM cerebrales (Fase II)”, funded by the research vicerrectorate of Universidad Militar Nueva Granada at Bogotá, Colombia.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Switzerland AG
About this paper
Cite this paper
Múnera, N., Almeida, J., Álvarez, C., Velasco, N., Romero, E. (2019). Autism Spectrum Disorders (ASD) Characterization in Children by Decomposing MRI Brain Regions with Zernike Moments. In: Lepore, N., Brieva, J., Romero, E., Racoceanu, D., Joskowicz, L. (eds) Processing and Analysis of Biomedical Information. SaMBa 2018. Lecture Notes in Computer Science(), vol 11379. Springer, Cham. https://doi.org/10.1007/978-3-030-13835-6_6
Download citation
DOI: https://doi.org/10.1007/978-3-030-13835-6_6
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-13834-9
Online ISBN: 978-3-030-13835-6
eBook Packages: Computer ScienceComputer Science (R0)