Abstract
Epidemiological studies provide evidence of a continuous rise in metabolic diseases throughout industrialized countries. Metabolic diseases are commonly associated with different abnormalities that hold a key role in the emergence and progression of frequent disorders including diabetes mellitus (DM), non-alcoholic fatty liver disease (NAFLD), obesity, metabolic syndrome and cardiovascular diseases. The burden of metabolic diseases is believed to arise through complex interaction between genetic and epigenetic factors, lifestyle changes and environmental exposure to triggering stimuli. The diagnosis and treatment of metabolic disorders continue to be an overwhelming challenge. Thus, the development of novel biomarkers may enhance the accuracy of the diagnosis at an early stage of the disease and allow effective intervention. Over the past decade, progress has been made in exploring the potential role of noncoding RNAs (ncRNAs) in the regulation of gene networks involved in metabolic diseases. A growing body of evidence now suggests that aberrant expression of circular RNAs (circRNAs) is relevant to the occurrence and development of metabolic diseases. Accordingly, circRNAs are proposed as predictive biomarkers and potential therapeutic targets for these diseases. As the field of circRNAs is rapidly evolving and knowledge is increasing, the present paper provides current understanding of the regulatory roles of these RNA species mainly in the pathogenesis of DM, NAFLD and obesity. Furthermore, some of the limitations to the promise of circRNAs and perspectives on their future research are discussed.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Circular RNAs (circRNAs)
- Metabolic Diseases
- Diabetes
- NAFLD
- Obesity
- Epigenetics
- Noncoding RNAs (ncRNAs)
10.1 Introduction
Metabolic diseases refer to different disorders including diabetes mellitus (DM), obesity, metabolic syndrome, dyslipidemia, and non-alcoholic fatty liver disease (NAFLD) [1]. These generally occur when metabolism processes fail. The pathogenesis of metabolic diseases and their chronic complications involve multiple molecular processes and pathways. Early studies using different models revealed that metabolic diseases arise through a complex interplay between genetics, epigenetics, environment, and/or lifestyle factors (nutrition, lack of exercise, etc.) and obesity [2,3,4]. However, their exact etiology remains partially elucidated.
Despite intensive research into most aspects of metabolic diseases, their causes are still poorly known and only a few effective drugs are available for accurate treatment. Nonetheless, the effectiveness of the current therapy could be improved if it could be implemented at early stage of the disease and targeted to the right subjects who may actually benefit from it. Such an ideal therapy cannot be achieved unless it is combined with predictive biomarkers to guide the treatment. Hence, the search for additional clinically relevant drugs as well as potential biomarkers with precise prognostic and diagnostic value is becoming increasingly important in the field of metabolic diseases.
Recent years have witnessed increasing interest in studying noncoding RNAs (ncRNAs) including long noncoding RNAs (lncRNAs), microRNAs (miRNAs) and circular RNAs (circRNAs), which are considered as important epigenetic regulators of many physiological processes. Huge efforts have been made to use these RNA molecules as predictive biomarkers for several diseases including metabolic disorders [5,6,7,8,9]. Nowadays, the landscape of miRNAs is by far the most characterized in relation to metabolic diseases whereas the role of circRNAs has not yet been precisely defined.
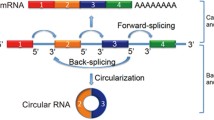
CircRNAs belong to the ever-growing world of ncRNA molecules. They are covalently closed single-stranded molecules generated from precursor mRNA back-splicing [10, 11] and can originate from different genomic regions. The lack of the typical terminal 5′ cap and 3′ polyadenylated tail renders circRNAs more stable and resistant to RNase R digestion compared to the linear RNA counterparts [12,13,14]. With respect to their biogenesis, detailed mechanisms have not been fully elucidated. Several possible models have been proposed including direct back-splicing with ALU and inverted repeats complementation, exon lariat, and RNA binding protein mediated models [15, 16].
Over the past few years, high throughput technologies have enabled a significant breakthrough in discovery of circRNAs. Today, thousands of circRNAs have been identified and annotated. Based on their genomic location, circRNAs can be classified into at least three types with distinct regulatory functions across multiple mammalian cells and species: (1) exonic circular RNAs (ecircRNAs); (2) circular intronic RNAs (ciRNAs); and (3) exon-intron circular RNAs (EIciRNAs) [11]. EcircRNAs appear to be the most abundant RNAs accounting for over 80% of the already known circRNAs. Moreover, the application of highly sophisticated bioinformatics tools has helped create several circRNA databases with searching and browsing functions [17].
CircRNAs are highly represented in the eukaryotic transcriptome, evolutionary conserved across species, and often show tissue or development stage-specific expression patterns [12, 18,19,20,21] suggesting their functional relevance [18, 19, 22, 23]. Interestingly, earlier studies indicated that the expression of a circular RNA does not correlate with the expression of its cognate linear mRNA [24]. In some cases, circRNAs can be more abundantly expressed than their associated linear mRNA isoform [25] while in other situations no circRNA can be detected despite high levels of mRNA expression [22, 23, 26]. The striking expression differences between circRNAs and their mRNA counterparts suggest that the production of circRNAs is a highly orchestrated process [23]. As to their potential functions, research is still limited and challenging. Studies have reported that some circRNAs may act as a sponge for miRNAs via competition with miRNA/mRNA binding or they may interact with RNA-binding proteins (RBPs) or regulate genes at the transcriptional and posttranscriptional levels [11, 27,28,29,30]. With these possible functions, specific circRNAs may control essential biological processes and contribute to the pathogenesis of diverse diseases including metabolic disorders [31,32,33]. However, the exact regulatory mechanisms by which these molecules may carry out these roles are not known. Thus, a more comprehensive understanding of how circRNAs function and what characteristics they should have to interact with other players to orchestrate gene expression in health and diseases states may lay the foundation for the development of RNAs-based diagnostic and therapeutic interventions for complex metabolic diseases. Below I will discuss the most important published studies of circRNAs in DM, NAFLD and obesity. CircRNAs that are most likely to be involved in some of these disorders as well as their putative functions are summarized in Table 10.1.
10.2 CircRNAs and Metabolic Diseases
10.2.1 DM
DM is a multiple-etiology metabolic disorder characterized by chronic hyperglycemia resulting from defects in secretion and/or insulin action [34]. Defects in insulin-mediated uptake of glucose can trigger pathogenic signals including mitochondrial dysfunction, oxidative stress, hypertension, inflammation and dyslipidemia. Additionally, diabetic patients with chronic hyperglycemia are more likely to suffer from many life-limiting and life-threatening complications, such as macrovascular-related stroke, heart disease, peripheral artery disease and/or microvascular-related retinopathy, neuropathy, nephropathy and cancer [35,36,37]. A major concern with these diabetic complications is that the number of DM cases and associated mortality are constantly increasing globally while the effectiveness of current treatments is limited, and this represents a heavy socioeconomic burden. Thus, identification of novel biomarkers that reflect or predict insulin-secretion dysfunction in individuals could transform the way we deal with diabetes, allowing for early prevention and guided therapy as a step toward precision medicine [38].
Over the last decade, efforts have been made to understand the disruption of mRNA-miRNA-lncRNA interaction networks under diabetic conditions [39]. More recently, scientists have shifted their research focus to circRNAs, hoping to develop these molecules as new biomarkers for early detection and management of diabetes. In this respect, the most well-known endogenous circRNA related to diabetes in the literature is CDR1as/ciRS-7 (a natural antisense transcript of CDR1) [19, 28]. Overexpression of this circRNA leads to improved insulin production and secretion in mouse β-cells [40]. By acting as a miR-7 sponge [20], CDR1as promotes islet β-cell proliferation and insulin secretion in diabetes via inhibiting miR-7 and enhancing Myrip and Pax6 expression [40]. These encouraging data suggest that the CDR1as/miR-7axis could serve as a potential therapeutic target for the treatment of diabetes. Similarly, another study reported that CDR1as and circHIPK3 silencing in wild-type animal models causes defective insulin secretion and lower islet cell proliferation [41]. By performing microarray and confirming the data by qRT-PCR, Zhao and colleagues measured the differential expression of circRNAs in the peripheral blood of pre-diabetes and T2DM patients compared to matched control subjects. The most significantly upregulated circRNA was hsa_circ_0054633 (Table 10.1) implying its potential as diagnostic biomarker for prediabetes and type 2 diabetes mellitus (T2DM) in the clinical setting [42]. Circular RNAs have also been investigated in diabetic vascular complications, which are a major cause of mortality among patients with diabetes [43]. In this context, Shan et al. reported that circHIPK3 was significantly induced in the retinas of patients with diabetes [44]. The same study group showed that depletion of circHIPK3 in a mouse model for diabetic retinopathy alleviated the retinal disorder [44]. Mechanistically, circHIPK3 competitively binds different miR-30 isoforms to restore the expression of their target genes including VEGF, FDZ4 and WNT2 which are involved in cell viability, proliferation and migration. In a more recent study, Zhang and colleagues identified circ_0005015 as the most significantly upregulated circRNA in plasma, vitreous samples and fibrovascular membranes of diabetic retinopathy patients [45]. Furthermore, the authors demonstrated that siRNA-mediated silencing of circ_0005015 significantly reduced human retinal vascular endothelial cell proliferation, migration and tube formation. Additional analyses revealed that circ_0005015 acted as an endogenous miR-519d-3p sponge to sequester and inhibit miR-519d-3p, thus facilitating retinal endothelial angiogenic function [45]. Together, these findings suggest that circ_0005015 may be considered as an ideal candidate biomarker for monitoring diabetic retinopathy. CircRNA_000203 is an additional circular transcript linked to diabetes. Tang and colleagues found that circRNA_000203 was upregulated in the diabetic mouse myocardium and in angiotensin (Ang) II-induced mouse cardiac fibroblasts [46]. In fact, circRNA_000203 could specifically increase the expression of fibrosis-associated genes (Col1a2, Col3a1) and α-SMA in cardiac fibroblasts via inhibiting the interaction of miR-26b-5p with the target genes. Therefore, circRNA_000203 might serve as a potential target for prevention and treatment of cardiac fibrosis in diabetic cardiomyopathy [46].
All of the above-mentioned findings suggest that the circRNAs-miRNAs-mRNAs regulatory axis could be a useful therapeutic target for the pathogenesis of diabetes and its complications. However, much more remains to be learned about the biology of circRNAs in diabetes and their beneficial clinical application appears to be a future endeavor.
10.2.2 NAFLD
Non-alcoholic fatty liver disease is emerging as the most common cause of chronic liver disease worldwide. It is a multifaceted disorder that ranges from the simple accumulation of triglycerides in hepatocytes (hepatic steatosis) to steatosis with inflammation, non-alcoholic steatohepatitis (NASH), fibrosis and cirrhosis, which may evolve towards cirrhosis and hepatocellular carcinoma [47,48,49,50]. The prevalence of NAFLD has been estimated to be between 25% and 45% of the general population [51, 52] and 70–90% among patients with obesity, DM or metabolic syndrome [53,54,55].
Although the pathophysiology of NAFLD has not been fully elucidated, recent investigations have brought forward evidence that this disorder may be caused by a plethora of factors including hepatic lipid accumulation, adipose tissue and mitochondrial dysfunction, a high fat diet, obesity, a chronic inflammatory state, insulin resistance, and genetic and epigenetic factors [56,57,58]. NAFLD is clinically important because fatty liver can progress to steatohepatitis in many patients and lead to liver cirrhosis and hepatocellular carcinoma. There is also growing evidence that in patients with NAFLD, hepatic steatosis is closely linked with obesity and the metabolic syndrome [59], which have been well-established as complex metabolic diseases with substantial heterogeneity. It is therefore important to identify biomarkers that may enable earlier prediction and diagnosis of NAFLD and to provide efficient treatment and better management.
An ongoing research effort is attempting to identify biological targets and signals closely associated with NAFLD. Some studies have indicated that miRNAs may have a potential role in this hepatic chronic disease [60]. Indeed, several processes relevant to the development and progression of NAFLD were found to be related to miRNAs [61, 62]. For instance, miR-34 is upregulated in NAFLD and has the potential to be a biomarker for diagnosis of this disorder [63, 64]. Several attempts have been made to translate miRNAs findings to clinical practice. In this sense, Regulus Therapeutics and AstraZeneca have started the development of RG-125, a GalNAc-conjugated anti-miR targeting microRNA-103/107 for the treatment of NASH in pre-diabetes and T2DM patients [65]. Hence, these encouraging findings, combined with the ongoing progress in the field of ncRNAs research, are expected to yield new insights into the pathogenesis of NAFLD.
The circRNA family has also become a key area of focus for research in NAFLD. There is now increasing evidence linking circRNAs to the pathogenesis of NAFLD even though studies in this respect have only just begun. Previous reports have established that the expression of PPARα and associated signaling pathways are inhibited by PPAR1 in patients with NAFLD [66, 67] but the underlying mechanism is not clear. Regarding circRNAs and hepatic steatosis, Guo and colleagues [68] found that circRNA_0046367 was significantly decreased in high-fat-induced hepatic steatosis [68]. Subsequently, the authors demonstrated that the decrease in circRNA_0046367 expression led to miR-34a/PPARα interaction and lipid peroxidative damage, while circRNA_0046367 normalization by intrahepatic overexpression prevented this interaction and therefore reduced steatosis [68]. In a different study, the same authors identified another circRNA, circRNA_0046366, whose expression was also decreased during free fatty acid-induced hepatocellular steatosis and its upregulation abolished the miR-34a-dependent inhibition of PPARα signaling, leading to a marked reduction in triglyceride levels and suppression of hepatocytes steatosis [69]. These findings suggest that the circRNA_0046367/miR-34a/PPARα and circRNA_0046366/miR-34a/PPARα axes play an important role in the pathogenesis of NAFLD [68, 69]. As circRNA_0046367 and circRNA_0046366 have the same target, it would be of interest to examine whether or not these two circRNAs act in synergy and if their transcripts display significant sequence similarities. In another study, the same group used the same model of NAFLD to show that circRNA_021412 is also associated with hepatic steatosis through the circRNA_021412/miR-1972/LPIN1 signaling cascade [70] (Table 10.1). Finally, in a recent study, Li et al. reported that the expression of circScd1 was significantly lower in NAFLD tissues than control groups whereas its over-expression promoted steatosis of NAFLD via JAK2/STAT5 signaling [71].
Together, the above pioneering studies suggest that circRNAs are potentially involved in NAFLD and have the potential to serve as useful tools for the development of diagnostic and interventional pharmacology. However, as mentioned earlier, circRNA data are still lacking functional evidence and their underlaying mechanisms are still awaiting elucidation. Therefore, further carefully designed prospective studies to emphasize and validate the potential use of circRNAs as NAFLD biomarkers are warranted.
10.2.3 Obesity
Obesity is another chronic metabolic disorder affecting adults and children in developed and developing countries [72]. Genetic predisposition, epigenetics, environment, and lifestyle preferences such as diet and low physical activity play crucial roles in excess body fat development and obesity [73, 74]. Obesity is known to be the main risk factor for several disorders including T2DM, cardiovascular disease, hypertension, coronary heart disease, and certain types of cancers [75, 76]. Due to the considerable impact of obesity on human health, it is therefore essential to develop new strategies with potential for early diagnosis and effective treatment.
While the involvement of miRNAs in the physiological processes of obesity has been closely studied [8, 77, 78], the role of circRNAs remains poorly elucidated. To the best of our knowledge, no groundbreaking studies have ever examined the potential link between circRNAs and obesity in humans. However, examination of the potential impact of circRNAs on diverse metabolic processes and a review of examples in the literature, suggest that circRNAs may play a role in the pathogenesis of obesity. For instance, based on the above studies revealing a significant association between circRNA expression and diabetes and NAFLD, and the fact that both are complications of obesity, it is conceivable that circRNAs may also contribute to the development of obesity. In addition, the antisense non-coding RNA in the INK4 locus (ANRIL), a complex gene with many reported linear and circular isoforms (circANRIL), is generated by the 9p21 locus has polymorphisms that have been associated with increased risk of developing cardiometabolic disease, including type 2 (obesity-related) diabetes and manifestations of atherosclerosis such as coronary artery disease [79,80,81]. Furthermore, a previous study by Murray et al. reported that lower level of CpG methylation within the promoter of ANRIL at birth is associated with increased cardiovascular risk [82] and adiposity [83] in later childhood. Carrara and colleagues hypothesized in their recent review that ANRIL could be a genomic site of environmental epigenetic influence on obesity [84]. An additional example that would argue in favor of a possible implication of circRNAs in obesity was shown by Li et al. when they attempted to identify the potential circRNAs associated with adipogenesis and lipid metabolism [85]. The authors analyzed the expression profile of these RNA molecules in subcutaneous adipose tissues of large White pig and Laiwu pig using RNA sequencing technology and bioinformatic methods. Among the differentially expressed circRNAs, they identified circRNA_11897 as the most significantly downregulated while circRNA_26852 was the most significantly upregulated. Subsequent analysis revealed that subcutaneous miR-27a and miR-27b-3p are targets for circRNA_11897 and subcutaneous miR-874 and miR-486 are targets for circRNA_26852 [85]. These target genes are enriched in pathways associated with adipocyte differentiation and lipid metabolism. Since miR-874 and miR-486 were shown to be targets of circRNA_26852, the authors hypothesized that circRNA_26852 may play a role in adipogenic differentiation and lipid metabolism through these miRNAs [85]. On the other hand, since miR-27a is known to promote lipolysis [86] and inhibit adipocyte differentiation by targeting PPARγ [87], it is reasonable to assume that circRNA_11897, which binds miR-27a and miR-27b-3p and consequently provokes upregulation of their target genes, may be implicated in the regulation adipogenic differentiation and lipid metabolism. The fact that several miRNAs have been shown to be involved in the processes of adipogenesis and obesity [8, 88] and lipid metabolism [89, 90], and considering the existing regulatory link and the dynamic interplay between different circRNAs and miRNAs, it is possible to assume that circRNAs may also be part of the complex machinery that orchestrates the regulation of genes associated with obesity. Obesity has been reported to induce a decline in the activity and the amount of PPARγ [91] and an upregulation of miR-130b and miR-138 levels. Considering that miR-130b is known to target 3′-UTR and certain sequences within the coding region of PPARγ [92], while miR-138 indirectly inhibits the expression of PPARγ [93], it is possible that the obesity-associated decline in PPARγ expression may be due to a decline in the expression of yet unknown circRNAs, that normally act as miRNA sponges to target miR-130b. There are reports in the literature that may support this scenario. In a previous study, Deng et al. observed that miR-548 can be regulated by the PPARγ gene, a heart-protective factor shown to be downregulated in acute myocardial infarction (AMI) [94]. Subsequently, when Deng et al. explored the expression profile of circRNAs comparing plasma expression of circRNAs in AMI patients with healthy volunteers, they identified circRNA_081881, which contained seven competitive binding sites for miR-548 as the most significantly downregulated circRNA in AMI. The authors concluded that circRNA_081881 may regulate PPARγ expression by functioning as a competing endogenous RNA (ceRNA) of miR-548 [94].
Collectively, these hypotheses and speculative scenarios are proposed for the purpose of serving as basic framework for further understanding of circRNAs in obesity and providing investigators with potential research directions that may be used for generating new hypotheses for further studies on circRNAs. Finally, as circRNAs research continues, it is expected that new information on the role of these molecules will arise in the field of metabolic diseases. It is hoped that this information will bring evidence for the potential role of circRNAs in metabolic diseases.
10.3 Conclusions and Future Perspectives
Even though circRNAs are increasingly being recognized to play critical regulatory roles in the development of metabolic diseases, the lack of their large exploration and characterization may delay their consideration for clinical settings. In this respect, many concerns are left for their potential future studies. The analytical approaches used in the identification and prediction of circRNAs are still part of a relatively new field of investigation, thus, their sensitivity and specificity require improvement. Furthermore, lack of prospective studies, poor study design and complicated statistical analyses could impede the translation of circRNA results to preclinical and clinical trials, thus, limiting the success of prospective biomarkers. The complex interplay of circRNAs with networks involving transcription factors, mRNA, miRNAs, RBPs and metabolic pathways makes it difficult to evaluate the functions of these RNA molecules under complex metabolic diseases. It should also be noted that circRNAs as transcriptional and posttranscriptional regulators themselves undergo extensive regulation from their biogenesis to the effects that they exert on their target molecules and pathways. Therefore, interpretation of such complex data could be enhanced by deploying systems biology approaches to refine our understanding of circRNAs dynamic and provide insights into their potential regulatory circuits in metabolic disorders. Another limitation that may generate huge incoherencies in circRNA results within a group of patients with metabolic diseases is drug use and other treatment modalities not taken in consideration. Using the example of miRNAs, previous studies reported that statins [95, 96], anticoagulation [97], and antiplatelet drugs [98] can affect quantification of these RNAs in blood samples and therefore should be taken in account. Regarding the patients with metabolic disease, thiazolidinedione drugs are frequently used for patients with impaired fasting glucose tolerance while abdominal obesity can be treated with a variety of lower calorie diets along with regular exercise [99]. Hence, drugs as well as confounding parameters should be also taken in account when examining circRNAs in patients with diabetes, NAFLD, obesity, and metabolic syndrome, as these may impact the disease through these RNA species. Metabolic diseases represent a cluster of disorders such as T2DM, insulin resistance, metabolic syndrome, NAFLD and hypercholesterolemia, which could be linked by numerous metabolic pathways. The interplay between these clinical situations is challenging. Thus, although each of these disorders has different physiological and clinical symptoms, it would be important to identify a signature or set of markers including circRNAs, shared by all disorders constituting metabolic diseases. This idea proses that, rather than relying on a single circRNA biomarker for disease diagnosis, one can use a group of disease-relevant biomarkers which will likely be more accurate and efficient in predicting a complex phenotype. Another type of circRNA that has not been well explored in metabolic diseases is circRNA found within exosomes (exo-circRNAs). The presence of abundant circRNAs within exosomes was firstly reported by Li and colleagues [100] and a web-accessible database (http://www.exoRBase.org), exoRBase, a resource containing all available long RNAs (circRNA, lncRNA and mRNA) derived from RNA-seq data of human blood exosomes, has been recently constructed [101].
With respect to metabolic diseases, many questions remain uncertain. For instance, what is the role of exo-circRNAs in metabolic disorders? What is their origin? Are they horizontally transferred via exosome vehicles to recipient cells as in the case of mRNAs [102]? In studies of cancer, Li et al. [100] observed that the abundance of tumor-derived exo-circRNAs in the serum of patients with colorectal cancer was correlated with tumor mass. They also found that the expression profile of exo-circRNAs in cancer serum was significantly different from that in normal serum. More importantly, a recent study revealed that treatment of lean mice with exosomes isolated from obese mice induced glucose intolerance and insulin resistance in mice [103]. Hence, future studies should aim for answering these questions in order to understand the origin, mode of secretion, target cells and organs of exo-circRNAs. This knowledge may help us to gain more insights into the function of circRNAs in the field of metabolic diseases. With respect to the epigenetic regulation of complex metabolic diseases by ncRNAs, there are only a few published data associating the dysregulation of circRNAs with genes involved diabetes and NAFLD, as mentioned above. Unfortunately, no data are yet available on the potential implication of circRNAs in obesity and metabolic syndrome. Likewise, no reports are available on the potential link between circRNAs and the chronic low-grade inflammation associated with diabetes, obesity and the metabolic syndrome apart from one report indicating an association between circANKRD36 and inflammation in patients with T2DM [104]. All of these pertinent questions represent important issues that must be solved in future investigative attempts to fully understand the role of circRNAs in the pathogenesis of metabolic diseases.
In summary, although the existing studies support a possible association between circRNA molecules and metabolic diseases, it is too early to consider and develop these molecules as sensors and biomarkers for metabolic disorders as claimed by existing reports. Further research in this area is worthwhile and new powerful strategies should be employed to uncover the full biological relevance of circRNAs and their potential therapeutic applications.
References
Stegemann R, Buchner DA (2015) Transgenerational inheritance of metabolic disease. Semin Cell Dev Biol 43:131–140
Baccarelli A, Bollati V (2009) Epigenetics and environmental chemicals. Curr Opin Pediatr 21(2):243–251
Sales VM, Ferguson-Smith AC, Patti ME (2017) Epigenetic mechanisms of transmission of metabolic disease across generations. Cell Metab 25(3):559–571
Barrès R, Zierath JR (2017) The role of diet and exercise in the transgenerational epigenetic landscape of T2DM. Nat Rev Endocrinol 12(8):441–451
Guay C, Regazzi R (2013) Circulating microRNAs as novel biomarkers for diabetes mellitus. Nat Rev Endocrinol 9(9):513–521
Tang Y, Zhou T, Yu X, Xue Z, Shen N (2017) The role of long non-coding RNAs in rheumatic diseases. Nat Rev Rheumatol 13(11):657–669
Gangwar RS, Rajagopalan S, Natarajan R, Deiuliis JA (2018) Noncoding RNAs in cardiovascular disease: pathological relevance and emerging role as biomarkers and therapeutics. Am J Hypertens 31(2):150–165
Zaiou M, El Amri H, Bakillah A (2018) The clinical potential of adipogenesis and obesity-related microRNAs. Nutr Metab Cardiovasc Dis 28(2):91–111
Zaiou M, Bakillah A (2018) Epigenetic regulation of ATP-Binding Cassette Protein A1 (ABCA1) gene expression: a new era to alleviate atherosclerotic cardiovascular disease. Diseases 6(2). pii: E34. doi: https://doi.org/10.3390/diseases6020034
Starke S, Jost I, Rossbach O, Schneider T, Schreiner S, Hung LH et al (2015) Exon circularization requires canonical splice signals. Cell Rep 10(1):103–111
Chen LL (2016) The biogenesis and emerging roles of circular RNAs. Nat Rev Mol Cell Biol 17(4):205–211
Jeck WR, Sorrentino JA, Wang K, Slevin MK, Burd CE, Liu J et al (2013) Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA 19(2):141–157
Chen LL, Yang L (2015) Regulation of circRNA biogenesis. RNA Biol 12(4):381–388
Enuka Y, Lauriola M, Feldman ME, Sas-Chen A, Ulitsky I, Yarden Y (2016) Circular RNAs are long-lived and display only minimal early alterations in response to a growth factor. Nucleic Acids Res 44(3):1370–1383
Dragomir M, Calin GA (2018) Circular RNAs in cancer - lessons learned from microRNAs. Front Oncol 8:179. https://doi.org/10.3389/fonc.2018.00307
Li X, Yang L, Chen LL (2018) The biogenesis, functions, and challenges of circular RNAs. Mol Cell 71(3):428–442
Xu Y (2017) An overview of the main circRNA databases. Non-coding RNA Investig 1:22. https://doi.org/10.21037/ncri.2017.11.05
Salzman J, Gawad C, Wang PL, Lacayo N, Brown PO (2012) Circular RNAs are the predominant transcript isoform from hundreds of human genes in diverse cell types. PLoS One 7(2):e30733. https://doi.org/10.1371/journal.pone.0030733
Memczak S, Jens M, Elefsinioti A, Torti F, Krueger J, Rybak A et al (2013) Circular RNAs are a large class of animal RNAs with regulatory potency. Nature 495(7441):333–338
Guo JU, Agarwal V, Guo H, Bartel DP (2014) Expanded identification and characterization of mammalian circular RNAs. Genome Biol 15(7):409. https://doi.org/10.1186/s13059-014-0409-z
Conn SJ, Pillman KA, Toubia J, Conn VM, Salmanidis M, Phillips CA et al (2015) The RNA binding protein quaking regulates formation of circRNAs. Cell 160(6):1125–1134
Salzman J, Chen RE, Olsen MN, Wang PL, Brown PO (2013) Cell-type specific features of circular RNA expression. PLoS Genet 9(9):e1003777. https://doi.org/10.1371/journal.pone.0030733
You X, Vlatkovic I, Babic A, Will T, Epstein I, Tushev G et al (2015) Neural circular RNAs are derived from synaptic genes and regulated by development and plasticity. Nat Neurosci 18(4):603–610
Nigro JM, Cho KR, Fearon ER, Kern SE, Ruppert JM, Oliner JD et al (1991) Scrambled exons. Cell 64(3):607–613
Maass PG, Glažar P, Memczak, Dittmar G, Hollfinger I, Schreyer L et al (2017) A map of human circular RNAs in clinically relevant tissues. J Mol Med (Berl) 95(11):1179–1189
Rybak-Wolf A, Stottmeister C, Glažar P, Jens M, Pino N, Giusti S et al (2015) Circular RNAs in the mammalian brain are highly abundant, conserved, and dynamically expressed. Mol Cell 58(5):870–885
Li Z, Huang C, Bao C, Chen L, Lin M, Wang X et al (2018) Exon-intron circular RNAs regulate transcription in the nucleus. Nat Struct Mol Biol 22(3):256–264
Hansen TB, Jensen TI, Clausen BH, Bramsen JB, Finsen B, Damgaard CK et al (2013) Natural RNA circles function as efficient microRNA sponges. Nature 495(7441):384–388
Hentze MW, Preiss T (2013) Circular RNAs: splicing’s enigma variations. EMBO J 32(7):923–925
Du WW, Zhang C, Yang W, Yong T, Awan FM, Yang BB (2017) Identifying and characterizing circRNA-protein interaction. Theranostics 7(17):4183–4191
Zhong Y, Du Y, Yang X, Mo Y, Fan C, Xiong F et al (2018) Circular RNAs function as ceRNAs to regulate and control human cancer progression. Mol Cancer 17(1):79. https://doi.org/10.1186/s12943-018-0827-8
Li M, Ding W, Sun T, Tariq MA, Xu T, Li P et al (2018) Biogenesis of circular RNAs and their roles in cardiovascular development and pathology. FEBS J 285(2):220–232
Floris G, Zhang L, Follesa P, Sun T (2017) Regulatory role of circular RNAs and neurological disorders. Mol Neurobiol 54(7):5156–5165
American Diabetes Association (2014) Diagnosis and classification of diabetes mellitus. Diabetes Care 37(Suppl 1):S81–S90
Kahn SE, Cooper ME, Del Prato S (2014) Pathophysiology and treatment of type 2 diabetes: perspectives on the past, present, and future. Lancet 383(9922):1068–1083
Shlomai G, Neel B, LeRoith D, Gallagher EJ (2016) Type 2 diabetes mellitus and cancer: the role of pharmacotherapy. J Clin Oncol 34(35):4261–4269
Rines AK, Sharabi K, Tavares CD, Puigserver P (2016) Targeting hepatic glucose metabolism in the treatment of type 2 diabetes. Nat Rev Drug Discov 15(11):786–804
Zaiou M, El Amri H (2017) Cardiovascular pharmacogenetics: a promise for genomically-guided therapy and personalized medicine. Clin Genet 91(3):355–370
Leung A, Natarajan R (2018) Long noncoding RNAs in diabetes and diabetic complications. Antioxid Redox Signal 29(11):1064–1073
Xu H, Guo S, Li W, Yu P (2015) The circular RNA Cdr1as, via miR-7 and its targets, regulates insulin transcription and secretion in islet cells. Sci Rep 5:12453. https://doi.org/10.1038/srep12453
Stoll L, Sobel J, Rodriguez-Trejo A, Guay C, Lee K, Venø MT et al (2018) Circular RNAs as novel regulators of β-cell functions in normal and disease conditions. Mol Metab 9:69–83
Zhao Z, Li X, Jian D, Hao P, Rao L, Li M (2017) Hsa_circ_0054633 in peripheral blood can be used as a diagnostic biomarker of pre-diabetes and type 2 diabetes mellitus. Acta Diabetol 54(3):237–245
Sena CM, Pereira AM, Seiça R (2013) Endothelial dysfunction - a major mediator of diabetic vascular disease. Biochim Biophys Acta 1832(12):2216–2231
Shan K, Liu C, Liu BH, Chen X, Dong R, Liu X et al (2017) Circular noncoding RNA HIPK3 mediates retinal vascular dysfunction in diabetes mellitus. Circulation 136(17):1629–1642
Zhang SJ, Chen X, Li CP, Li XM, Liu C, Liu BH et al (2017) Identification and characterization of circular RNAs as a new class of putative biomarkers in diabetes retinopathy. Invest Ophthalmol Vis Sci 58(14):6500–6509
Tang CM, Zhang M, Huang L, Hu ZQ, Zhu JN, Xiao Z et al (2017) CircRNA_000203 enhances the expression of fibrosis-associated genes by derepressing targets of miR-26b-5p, Col1a2 and CTGF, in cardiac fibroblasts. Sci Rep 7:40342. https://doi.org/10.1038/srep40342
Chalasani N, Younossi Z, Lavine JE, Diehl AM, Brunt EM, Cusi K et al (2012) The diagnosis and management of non-alcoholic fatty liver disease: practice Guideline by the American Association for the Study of Liver Diseases, American College of Gastroenterology, and the American Gastroenterological Association. Hepatology 55(6):2005–2023
Michelotti GA, Machado MV, Diehl AM (2013) NAFLD, NASH and liver cancer. Nat Rev Gastroenterol Hepatol 10(11):656–665
Adams LA, Angulo P, Lindor KD (2005) Nonalcoholic fatty liver disease. CMAJ 172(7):899–905
Lucas C, Lucas G, Lucas N, Krzowska-Firych J, Tomasiewicz K (2018) A systematic review of the present and future of non-alcoholic fatty liver disease. Clin Exp Hepatol 4(3):165–174
Loomba R, Sanyal AJ (2013) The global NAFLD epidemic. Nat Rev Gastroenterol Hepatol 10(11):686–690
Rinella ME (2015) Nonalcoholic fatty liver disease: a systematic review. JAMA 313(22):2263–2273
Gholam PM, Kotler DP, Flancbaum LJ (2002) Liver pathology in morbidly obese patients undergoing Roux-en-Y gastric bypass surgery. Obes Surg 12(1):49–51
Williamson RM, Price JF, Glancy S, Perry E, Nee LD, Hayes PC et al (2011) Prevalence of and risk factors for hepatic steatosis and nonalcoholic Fatty liver disease in people with type 2 diabetes: the Edinburgh Type 2 Diabetes Study. Diabetes Care 34(5):1139–1144
Chalasani N, Younossi Z, Lavine JE, Diehl AM, Brunt EM, Cusi K et al (2012) The diagnosis and management of non-alcoholic fatty liver disease: practice guideline by the American Association for the Study of Liver Diseases, American College of Gastroenterology, and the American Gastroenterological Association. Am J Gastroenterol 107(6):811–826
Tilg H, Moschen AR (2010) Evolution of inflammation in nonalcoholic fatty liver disease: the multiple parallel hits hypothesis. Hepatology 52(5):1836–1846
Buzzetti E, Pinzani M, Tsochatzis EA (2016) The multiple-hit pathogenesis of non-alcoholic fatty liver disease (NAFLD). Metabolism 65(8):1038–1048
Peverill W, Powell LW, Skoien R (2014) Evolving concepts in the pathogenesis of NASH: beyond steatosis and inflammation. Int J Mol Sci 15(5):8591–8638
Angulo P (2002) Nonalcoholic fatty liver disease. N Engl J Med 346(16):1221–1231
Torres JL, Novo-Veleiro I, Manzanedo L, Alvela-Suárez L, Macías R, Laso FJ et al (2018) Role of microRNAs in alcohol-induced liver disorders and non-alcoholic fatty liver disease. World J Gastroenterol 24(36):4104–4118
Szabo G, Csak T (2016) Role of MicroRNAs in NAFLD/NASH. Dig Dis Sci 61(5):1314–1324
Liu CH, Ampuero J, Gil-Gómez A, Gil-Gómez A, Montero-Vallejo R, Rojas Á et al (2018) miRNAs in patients with non-alcoholic fatty liver disease: a systematic review and meta-analysis. J Hepatol 69(6):1335–1348
Cermelli S, Ruggieri A, Marrero JA, Ioannou GN, Beretta L (2011) Circulating microRNAs in patients with chronic hepatitis C and non-alcoholic fatty liver disease. PLoS One 6(8):e23937. https://doi.org/10.1371/journal.pone.0023937
Salvoza NC, Klinzing DC, Gopez-Cervantes J, Baclig MO (2016) Association of circulating serum miR-34a and miR-122 with dyslipidemia among patients with non-alcoholic fatty liver disease. PLoS One 11(4):e0153497. https://doi.org/10.1371/journal.pone.0153497
Rupaimoole R, Slack FJ (2017) MicroRNA therapeutics: towards a new era for the management of cancer and other diseases. Nat Rev Drug Discov 16(3):203–222
Francque S, Verrijken A, Caron S, Prawitt J, Paumelle R, Derudas B et al (2015) PPARα gene expression correlates with severity and histological treatment response in patients with non-alcoholic steatohepatitis. J Hepatol 63(1):164–173
Huang K, Du M, Tan X, Yang L, Li X, Jiang Y et al (2017) PARP1-mediated PPARα poly(ADP-ribosyl)ation suppresses fatty acid oxidation in non-alcoholic fatty liver disease. J Hepatol 66(5):962–977
Guo XY, Chen JN, Sun F, Wang YQ, Pan Q, Fan JG (2017) circRNA_0046367 prevents hepatoxicity of lipid peroxidation: an inhibitory role against hepatic steatosis. Oxidative Med Cell Longev 2017:3960197. https://doi.org/10.1155/2017/3960197
Guo XY, Sun F, Chen JN, Wang YQ, Pan Q, Fan JG (2018) circRNA_0046366 inhibits hepatocellular steatosis by normalization of PPAR signaling. World J Gastroenterol 24(3):323–337
Guo XY, He CX, Wang YQ, Sun C, Li GM, Su Q et al (2017) Circular RNA profiling and bioinformatic modeling identify its regulatory role in hepatic steatosis. Biomed Res Int 2017:5936171–5936113. https://doi.org/10.1155/2017/5936171
Li P, Shan K, Liu Y, Zhang Y, Xu L, Xu L (2018) CircScd1 promotes fatty liver disease via the Janus Kinase 2/signal transducer and activator of transcription 5 pathway. Dig Dis Sci Sep 27. https://doi.org/10.1007/s10620-018-5290-2. [Epub ahead of print]
James PT, Leach R, Kalamara E, Shayeghi M (2001) The worldwide obesity epidemic. Obes Res 9(Suppl 4):228S–233S
Loos RJ, Bouchard C (2008) FTO: the first gene contributing to common forms of human obesity. Obes Rev 9:246–250
Rojas J, Arraiz N, Aguirre M, Velasco M, Bermúdez V (2011) AMPK as target for intervention in childhood and adolescent obesity. J Obes 2011:252817. https://doi.org/10.1155/2011/252817
Mokdad AH, Ford ES, Bowman BA, Dietz WH, Vinicor F, Bales VS et al (2003) Prevalence of obesity, diabetes, and obesity-related health risk factors, 2001. JAMA 289(1):76–79
Landsberg L, Aronne LJ, Beilin LJ, Burke V, Igel LI, Lloyd-Jones D et al (2013) Obesity-related hypertension: pathogenesis, cardiovascular risk, and treatment: a position paper of The Obesity Society and the American Society of Hypertension. J Clin Hypertens (Greenwich) 15(1):14–33
Hilton C, Neville MJ, Karpe F (2013) MicroRNAs in adipose tissue: their role in adipogenesis and obesity. Int J Obes 37(3):325–332
Wang R, Hong J, Cao Y, Shi J, Gu W, Ning G et al (2015) Elevated circulating microRNA-122 is associated with obesity and insulin resistance in young adults. Eur J Endocrinol 172(3):291–300
Cunnington MS, Santibanez Koref M, Mayosi BM, Burn J, Keavney B (2010) Chromosome 9p21 SNPs associated with multiple disease phenotypes correlate with ANRIL expression. PLoS Genet 6(4):e1000899. https://doi.org/10.1371/journal.pgen.1000899
Kong Y, Sharma RB, Nwosu BU, Alonso LC (2016) Islet biology, the CDKN2A/B locus and type 2 diabetes risk. Diabetologia 59(8):1579–1593
Tajbakhsh A, Khorrami MS, Hassanian SM, Aghasizade M, Pasdar A, Maftouh M et al (2016) The 9p21 locus and its potential role in atherosclerosis susceptibility; molecular mechanisms and clinical implications. Curr Pharm Des 22(37):5730–5737
Murray R, Bryant J, Titcombe P, Barton SJ, Inskip H, Harvey NC et al (2016) DNA methylation at birth within the promoter of ANRIL predicts markers of cardiovascular risk at 9 years. Clin Epigenetics 8:90. https://doi.org/10.1186/s13148-016-0259-5
Lillycrop K, Murray R, Cheong C, Teh AL, Clarke-Harris R, Barton S et al (2017) ANRIL promoter DNA methylation: a perinatal marker for later adiposity. EBioMedicine 19:60–72
Carrara M, Fuschi P, Ivan C, Martelli F (2018) Circular RNAs: methodological challenges and perspectives in cardiovascular diseases. J Cell Mol Med 22(11):5176–5187
Li A, Huang W, Zhang X, Xie L, Miao X (2018) Identification and characterization of CircRNAs of two pig breeds as a new biomarker in metabolism-related diseases. Cell Physiol Biochem 47(6):2458–2470
Wang T, Li M, Guan J, Li P, Wang H, Guo Y et al (2011) MicroRNAs miR-27a and miR-143 regulate porcine adipocyte lipid metabolism. Int J Mol Sci 12(11):7950–7959
Kim SY, Kim AY, Lee HW, Son YH, Lee GY, Lee JW et al (2010) miR-27a is a negative regulator of adipocyte differentiation via suppressing PPARgamma expression. Biochem Biophys Res Commun 392(3):323–328
Sun L, Goff LA, Trapnell C, Alexander R, Lo KA, Hacisuleyman E et al (2013) Long noncoding RNAs regulate adipogenesis. Proc Natl Acad Sci U S A 110(9):3387–3392
Zaiou M, Rihn BH, Bakillah A (2018) Epigenetic regulation of genes involved in the reverse cholesterol transport through interaction with miRNAs. Front Biosci (Landmark Ed) 23:2090–2105
Chen Z (2015) Progress and prospects of long noncoding RNAs in lipid homeostasis. Mol Metab 5(3):164–170
Motawi TK, Shaker OG, Ismail MF, Sayed NH (2017) Peroxisome proliferator-activated receptor gamma in obesity and colorectal cancer: the role of epigenetics. Sci Rep 7(1):10714. https://doi.org/10.1038/s41598-017-11180-6
Lee EK, Lee MJ, Abdelmohsen K, Kim W, Kim MM, Srikantan S et al (2011) miR-130 suppresses adipogenesis by inhibiting peroxisome proliferator-activated receptor gamma expression. Mol Cell Biol 31(4):626–638
Yang Z, Bian C, Zhou H, Huang S, Wang S, Liao L et al (2011) MicroRNA hsa-miR-138 inhibits adipogenic differentiation of human adipose tissue-derived mesenchymal stem cells through adenovirus EID-1. Stem Cells Dev 20(2):259–267
Deng YY, Zhang W, She J, Zhang L, Chen T, Zhou J et al (2016) Circular RNA related to PPARγ function as ceRNA of microRNA in human acute myocardial infarction. J Am Coll cardiol 68(16) Supp S. https://doi.org/10.1016/j.jacc.2016.07.189
Gao W, He HW, Wang ZM, Zhao H, Lian XQ, Wang YS et al (2012) Plasma levels of lipometabolism-related miR-122 and miR-370 are increased in patients with hyperlipidemia and associated with coronary artery disease. Lipids Health Dis 11:55. https://doi.org/10.1186/1476-511X-11-55
Zambrano T, Hirata RDC, Hirata MH, Cerda Á, Salazar LA (2018) Statins differentially modulate microRNAs expression in peripheral cells of hyperlipidemic subjects: a pilot study. Eur J Pharm Sci 117:55–61
Boeckel JN, Thome CE, Leistner D, Zeiher AM, Fichtlscherer S, Dimmeler S (2013) Heparin selectively affects the quantification of micrornas in human blood samples. Clin Chem 59(7):1125–1127
de Boer HC, van Solingen C, Prins J, Duijs JM, Huisman MV, Rabelink TJ et al (2013) Aspirin treatment hampers the use of plasma microRNA-126 as a biomarker for the progression of vascular disease. Eur Heart J 34(44):3451–3457
Wagh A, Stone NJ (2004) Treatment of metabolic syndrome. Expert Rev Cardiovasc Ther 2(2):213–228
Li Y, Zheng Q, Bao C, Li S, Guo W, Zhao J et al (2015) Circular RNA is enriched and stable in exosomes: a promising biomarker for cancer diagnosis. Cell Res 25:981–984
Li S, Li Y, Chen B, Zhao J, Yu S, Tang Y et al (2018) exoRBase: a database of circRNA, lncRNA and mRNA in human blood exosomes. Nucleic Acids Res 46(D1):D106–D112
Valadi H, Ekstrom K, Bossios A, Sjöstrand M, Lee JJ, Lötvall JO (2007) Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat Cell Biol 9:654–659
Castaño C, Kalko S, Novials A, Párrizas M (2018) Obesity-associated exosomal miRNAs modulate glucose and lipid metabolism in mice. Proc Natl Acad Sci USA Nov 14. pii: 201808855. https://doi.org/10.1073/pnas.1808855115. [Epub ahead of print]
Fang Y, Wang X, Li W, Han J, Jin J, Su F et al (2018) Screening of circular RNAs and validation of circANKRD36 associated with inflammation in patients with type 2 diabetes mellitus. Int J Mol Med 42(4):1865–1874
Li X, Zhao Z, Jian D, Li W, Tang H, Li M (2017) Hsa-circRNA11783-2 in peripheral blood is correlated with coronary artery disease and type 2 diabetes mellitus. Diab Vasc Dis Res 14(6):510–515
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Zaiou, M. (2019). Circular RNAs as Potential Biomarkers and Therapeutic Targets for Metabolic Diseases. In: Guest, P. (eds) Reviews on Biomarker Studies of Metabolic and Metabolism-Related Disorders. Advances in Experimental Medicine and Biology(), vol 1134. Springer, Cham. https://doi.org/10.1007/978-3-030-12668-1_10
Download citation
DOI: https://doi.org/10.1007/978-3-030-12668-1_10
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-12667-4
Online ISBN: 978-3-030-12668-1
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)