Abstract
There has been an increasing effort to incorporate the inner workings of soil microbial communities into conceptual and quantitative models of processes at the ecosystem or global scale. Many studies show that the characteristics of microbial species and their interactions with each other and with plants strongly influence larger-scale processes and that explicitly including microbes can improve the performance of ecosystem models. We review the current understanding of how the physiology and community structure of soil microbial communities can impact cycling of carbon (C), nutrients, and greenhouse gases and recent progress in integrating this knowledge into quantitative models of ecosystems and climate change. Microbes can be characterized by ecological strategies that influence carbon use efficiency, stress physiology, elemental ratios (stoichiometry), production of extracellular enzymes, and responses to temperature. Competitive, synergistic, and trophic interactions within soil microbial communities influence process rates and responses to climate change. Plant-microbe interactions are central in climate change responses of ecosystems and can operate by changes in nutrient cycling or through alterations in the balance of mutualists and parasites. There are trends that connect broad-scale community structure with functioning and evidence that ecological roles of microbes can be mapped to phylogeny at the genus or species level. Models that explicitly simulate microbes have included their physiological limits, growth kinetics, interactions with plants, stoichiometry, dormancy, community structure, and community interactions. Given recent advances in conceptual frameworks for microbial ecology and in techniques for describing microbial communities and computing power, further progress will depend on increased interactions between microbiologists and modelers.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
3.1 Introduction
It has long been known that microbial communities play a central role in biogeochemical cycles and potential biological feedbacks to climate change (Alexander 1964; Baes et al. 1977). Despite this recognition it has been a long-standing challenge in microbial ecology to incorporate the inner workings of microbial communities into conceptual and quantitative ecosystem models (Schimel 1995). Environmental microbial communities are notoriously diverse and dominated by species that resist attempts at cultivation (DeLong and Pace 2001; Rappe and Giovannoni 2003; Yarza et al. 2014; Youssef et al. 2015). The past two decades have seen great technical advances in describing the diversity and metabolism of uncultured microbes, and many previously uncultured bacterial phyla have recently been isolated in pure culture (George et al. 2011; Stewart 2012). Interdisciplinary studies that combine a wide range of techniques to study microbial communities and their role in the environment have become increasingly common. These studies highlight the relevance of microbial ecology and physiology for ecosystem models, but incorporating complex microbial processes and community structure into already complex models is a daunting task. Large uncertainties in coupled climate models arise from biological feedbacks; however, microbes are not yet explicitly included in models used by the Intergovernmental Panel on Climate Change (IPCC) (Hararuk et al. 2015; Wieder et al. 2015; Luo et al. 2016). Given the computational costs of adding more model components and the qualitative nature of much microbial research, it is not obvious how ecosystem models should be modified to fit the emerging understanding of how microbial communities function (Chapin et al. 2009; Lawrence et al. 2009; Todd-Brown et al. 2011).
So, when is a detailed biological understanding of soil microbial communities necessary for predicting ecosystem processes? Models assume microbial activity can be predicted from a set of environmental drivers, like temperature and soil moisture (Schimel 2001; Bloom et al. 2010). This is insufficient when:
-
1.
Microbial activities are driven by variables that are not included in the model, such as soil texture, nutrients, trace metal availability, and specific interactions with plant species. These extraneous variables could control microbial function either directly or indirectly, through effects on the community composition.
-
2.
The microbial community composition is unpredictable due to dispersal limitation and disturbances such as pulse dynamics or local soil properties.
-
3.
There are community interactions such as competition, synergism, and predation that alter rates in complex ways that are hard to predict based on the standard environmental variables.
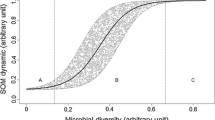
Figure 3.1 outlines the various ways that the properties of soil microbial communities and the species that comprise them can scale up to have impacts on ecosystem functions ranging from plot, regional, and global scales. In this chapter, we summarize the current understanding of how the physiology and community structure of soil microbial communities can scale up to impact cycling of C, nutrients, and greenhouse gases and the recent progress in integrating this knowledge into quantitative models of ecosystems and climate change. A great deal of thought has been given to this subject in recent years, and numerous useful reviews have been published on how soil microbes impact ecosystem processes (Schimel and Schaeffer 2012; Wallenstein and Hall 2012; Nemergut et al. 2014; Sinsabaugh et al. 2014; Bier et al. 2015; Ferris and Tuomisto 2015; García-Palacios et al. 2015; Nielsen et al. 2015; Zechmeister-Boltenstern et al. 2015; van der Putten et al. 2016), incorporating microbes into models (Todd-Brown et al. 2011; Treseder et al. 2012; Nazaries et al. 2013; Wieder et al. 2015; Luo et al. 2016), and the impacts of climate change and other disturbances on microbes (Bradford 2013; de Vries and Shade 2013; Griffiths and Philippot 2013; Classen et al. 2015) and their extracellular enzymes (Burns et al. 2013; Henry 2013).
Microbial traits and community interactions that might have significant impacts at the ecosystem and global scale. Effects are shown as arrows. Through their ecological strategies and interactions within the community and with plants, microbes impact trace gas fluxes and the C balance of ecosystems, influencing climate change, which in turn feeds back on all these processes and characteristics
3.2 Physiological Traits that Scale to Ecosystem Processes
3.2.1 Carbon Use Efficiency
Respiration by soil microbes accounts for about half of the CO2 efflux from terrestrial ecosystems (Ciais et al. 2013). Therefore, the rate and efficiency of soil microbial respiration are key factors regulating the C balance of ecosystems. Carbon use efficiency (CUE), the fraction of C consumed by microbes that is converted to biomass as opposed to CO2, is influenced by factors such as temperature, substrate quality, and the ecological strategies of the microbes that comprise the community. Consequently, CUE is emerging as an important characteristic of microbial communities that can strongly influence the C balance of ecosystems and their response to climate change (Lipson et al. 2009; Allison et al. 2010; Manzoni et al. 2012; Cotrufo et al. 2013; Sinsabaugh et al. 2013; Allison 2014). There is often a trade-off observed between maximum growth rate and growth yield, as thermodynamic constraints generally prevent microorganisms from being both fast-growing and efficient (Lipson 2015). This can lead to an axis representing two contrasting life history strategies: slow-growing efficient microbes that thrive under resource scarce conditions versus fast-growing, inefficient microbes that dominate under resource-rich conditions (Kreft and Bonhoeffer 2005). These trade-offs and ecological strategies that they produce have direct impacts on C cycling. A fast-growing, inefficient microbial community produces much more CO2 per unit biomass than a slow-growing community (Lipson et al. 2009). Studies in a high-elevation forest found that seasonal changes in the soil microbial community and its predominant growth strategy had major repercussions for ecosystem respiration, in particular in the late winter, when a cold-adapted, fast-growing community developed under the thick, protective snow pack, where high concentrations of sugars accumulated due to frost damage of roots (Monson et al. 2006; Scott-Denton et al. 2006; Schmidt et al. 2009). A simple model that included microbial growth kinetics and temperature response parameters performed well in predicting ecosystem respiration (Lipson et al. 2009).
Furthermore, CUE may play a major role in microbial and ecosystem responses to climate change. A modeling study found that warming led to decreased CUE, which in turn led to reduced microbial biomass, limiting C loss from soils under warmer temperature regimes (Allison et al. 2010). A follow-up modeling study found that a trade-off between growth rate and yield would prevent adaptation of higher-yield microbes under this warming scenario, thereby maintaining this limit to C loss from soils (Allison 2014). These studies could explain earlier experimental results showing “acclimatization” of soils to warming treatments, in which the temperature sensitivity of soil respiration decreased in warmed plots (Luo et al. 2001). This so-called acclimatization has often been attributed to changes in substrate availability in warmed soils, but there are also changes in microbial community structure and function that can produce these effects (Wei et al. 2014; Osanai et al. 2015). However, it is uncertain whether global warming will result in a sustained decrease in CUE or whether microbes will adapt their CUE over time (Dijkstra et al. 2011; Tucker et al. 2013). Reductions in CUE with warming have been reported (Steinweg et al. 2008; Manzoni et al. 2012), but there is evidence that the CUE of microbial communities may adapt in the long term (Bradford et al. 2008; Bradford 2013; Frey et al. 2013; Tucker et al. 2013). Increased temperature can also increase rates of microbial turnover in soil, which can have the opposite effect of decreased CUE, increasing the stability of soil organic matter (Hagerty et al. 2014). Interpretation of these studies is further complicated in that microbial temperature responses are sensitive to the chemistry of soil organic matter (Wagai et al. 2013), which can change in response to warming as the decomposition rates and inputs of different forms of soil organic matter shift. Similarly, mineral nutrient availability can also have a strong influence on CUE (Keiblinger et al. 2010; Manzoni et al. 2012; Sinsabaugh et al. 2016). In summary, CUE represents a strong potential link between microbial communities and the global C cycle, and so resolving the uncertainties in this area and incorporating this knowledge into coupled climate change models should be a priority.
3.2.2 Microbial Temperature Responses
Microbial responses to temperature can shape ecosystem processes in unexpected ways. The temperature sensitivity of microbial activities, such as respiration or enzyme activity, is often described as an apparent Q10, the proportional increase in activity with a 10 °C increase in temperature. The Q10 concept was initially developed for simple chemical reactions, and so its application to complex microbial processes in the natural environment presents some difficulties. First of all, biological processes depend on enzymes, which are only functional within a certain temperature range, and so microbial growth is best described by a square root function rather than an exponential Q10 (Arrhenius) function. Furthermore, biological processes such as microbial respiration are the result of many different enzymes and transport processes, both within the cell and in the environment. In soils very high apparent Q10s are observed around the freezing point of water, resulting from changes in the thickness of unfrozen water films in subzero soils, changes in the physiological state of microbes, and growth of microbes over time (when the Q10 is generated from respiration data collected in the field over a period of several weeks) (Mikan et al. 2002; Monson et al. 2006; Panikov et al. 2006; Schmidt et al. 2009). The optimal temperature ranges for microbial growth vary greatly from psychrophiles [less than 15 °C by definition, some at least as low as 5 °C (Morita 1975)] to hyperthermophiles (optimal temperatures over 80 °C), and so clearly the composition of the microbial community could affect how an environment responds to climate change in the short term. However, microbial communities appear to be well-adapted to the ambient temperatures they experience in their environment, and so changes in temperature can have a smaller impact on microbial processes than would be predicted from a model that assumes a simple Q10 relationship for all environments (Giardina and Ryan 2000). Maximum growth rates of bacteria are not well correlated to their optimum temperature (Hanus and Morita 1968; Ratkowsky et al. 1982, 1983). The fastest-growing bacterium currently known, Vibrio natriegens, which doubles in about 9 min at 37 °C, is a mesophile (Maida et al. 2013), and one of the fastest-growing psychrophiles, Pseudoalteromonas haloplanktis, can give many thermophiles a run for their money (doubles in 4 h at 4 °C) (Piette et al. 2010). In the case of permafrost (permanently frozen soil layers found in polar regions), the communities in these frozen soils, despite being capable of growth at surprisingly low temperatures, are currently at suboptimal temperatures for growth (Bakermans et al. 2003; Steven et al. 2006), and melting will certainly lead to substantial losses of C (Schaefer et al. 2014; Schuur et al. 2015). However, current models do not capture the nuances of microbial activity at low temperatures. It is clear that microbial activity continues at low temperatures, for example, in permafrost and during the winter, and these activities are not yet fully incorporated into annual C cycling budgets and models (Clein and Schimel 1995; Brooks et al. 1996, 1997; Lipson et al. 1999; Monson et al. 2006; Panikov 2009; Zona et al. 2016). Despite adaptations for extreme cold, it is likely that winter microbial communities generally experience suboptimal temperatures and should be more sensitive to warming than the communities that are active in the summer. A model that incorporated seasonal microbial dynamics found large C losses in response to winter warming, when microbes were allowed to acclimate their temperature responses to changing seasons (Sistla et al. 2014). The Q10 values of respiration can vary by season and across altitudinal gradients, and these variations are correlated to changes in microbial community structure (Lipson 2007). These results highlight the importance of microbial characteristics and their seasonal dynamics for ecosystem processes.
3.2.3 Stoichiometry of Microbial Cells in Relation to Soil Organic Matter
The stoichiometric ratios of carbon, nitrogen, and phosphorus typically are expressed as C:N:P in plant litter, and soil organic matter in relation to that of microbial biomass are major determinants of C and nutrient cycling (Zechmeister-Boltenstern et al. 2015). While marine microbial primary producers have a relatively narrow range of atomic C:N:P ratios around 106:16:1, known as the Redfield ratio (Redfield 1958), land plants vary more in their nutrient concentrations due to structural materials like lignin and cellulose and variations in nutrient concentrations between lifeforms such as trees vs. herbs, deciduous vs. evergreen, etc. Microbes generally have higher N and P concentrations (lower C:N and C:P ratios) than do macroscopic organisms, and since microbes are the primary decomposers in soils, stoichiometric ratios in soil organic matter are driven toward that of the microbes over time. This occurs because N and P tend to be retained in microbial biomass but C is respired away as CO2. When the C:N and C:P of decomposing litter drop below a threshold value, which depends on microbial CUE, N and P from the litter are in excess of microbial demand and are mineralized and released back into the soil. While the range of element ratios is limited by the nature of cellular life on Earth, there is also considerable variation in C:N:P among microbial groups, particularly between bacteria and fungi. Microbes are also capable of storing C (e.g., as neutral lipids, glycogen, and starch) and also P (typically stored as polyphosphate) when these nutrients are in excess of immediate needs. Therefore different plant litter types lead to variations in stoichiometry of soil organic matter and microbial biomass, especially in terms of P concentrations (Cleveland and Liptzin 2007; Fanin et al. 2013; Hartman and Richardson 2013; Xu et al. 2013). While these variations in microbial element ratios are presumably driven by plants, there is the possibility that changes in microbial community structure could reinforce and accelerate changes in the plant community through their stoichiometry, as discussed in the Plant-Microbe Interactions section below.
There is a strong but complex relationship among stoichiometric ratios, ecological strategies, and CUE in microbes. Faster-growing microbes tend toward higher N and P concentrations and lower CUE (Keiblinger et al. 2010; Zechmeister-Boltenstern et al. 2015; Sinsabaugh et al. 2016). Faster growth rates require more enzymes, ribosomes, and RNA synthesis, increasing the demand for N and P (Gillooly et al. 2005; Hartman and Richardson 2013). Therefore, CUE might decrease with increasing P availability, as C:P ratios decline (Sinsabaugh et al. 2016). On the other hand, under conditions of nutrient limitation (which would occur with high C:N and C:P ratios of soil organic matter), microbial respiration can become decoupled from growth, leading to low CUE (Schimel and Weintraub 2003; Manzoni et al. 2012). In marine and aquatic ecosystems, P limitation commonly reduces bacterial growth efficiency (Del Giorgio and Cole 1998). Soil microbial growth is generally considered to be limited by labile C, even in relatively nutrient-poor soils (Heuck et al. 2015). However, P can limit microbial biomass in some tropical soils (Cleveland et al. 2002), and nutrient additions can stimulate microbial nutrient uptake and activity without necessarily causing an immediate increase in biomass (Jonasson et al. 1996; Schimel and Weintraub 2003; Allen and Schlesinger 2004; Sistla et al. 2012). The effects of high C:N and C:P ratios of plant litter can sometimes have negative effects on CUE (Spohn and Chodak 2015). It has also been shown that social interactions within microbial communities, such as when a subset of microbes “cheat” by not contributing to extracellular enzyme production, lead to different outcomes than those predicted by stoichiometric theory (Kaiser et al. 2014). In summary, ecological stoichiometry is a promising approach for scaling from microbes to ecosystems, but the relationships are complex due to potentially opposing forces of nutrient limitation and ecological strategies.
3.2.4 Extracellular Enzymes
Soil extracellular enzymes are a major link between the microbial community and its impact on the ecosystem, particularly through litter decomposition. The majority of organic inputs to soil are polymers like cellulose, lignin, and proteins that require extracellular enzymes for their degradation, and so extracellular enzymes usually catalyze the rate-limiting steps in the C and N cycles. For example, it was shown that litter decomposition was limited by the activity of phenol oxidases in anaerobic peatlands due to a lack of oxygen required by these enzymes (Freeman et al. 2001). The production and turnover of enzymes along with their substrate specificity, turnover rate, and temperature responses shape ecosystem processes. Shifts in the microbial community can change the compliment of enzymes in the soil, altering patterns of decomposition and nutrient cycling. Soil feedbacks in plant invasions are an example where this seems to be important (Henry 2013) (also discussed in the following section). Similarly, the role of the microbial community in stabilizing soil C could depend on the enzyme profile produced by the community, for example, through the relative dominance of fungi or bacteria (Waring et al. 2013). Because extracellular enzymes represent a cost for cells in terms of both C and N, and because they control rates of decomposition, extracellular enzymes tightly integrate C and N cycling with microbial physiology (Schimel and Weintraub 2003; Burns et al. 2013). Enzymes that degrade cell walls are often regulated by N availability, and it is thought that the main microbial motivation for producing them is increasing their access to N bound in this litter. For example, N deposition can alter soil C storage by reducing production of extracellular enzymes in oak forests with low litter quality (Waldrop et al. 2004). However, this relationship between N availability and lignin degradation varies by biome and microbial community (Sinsabaugh 2010). Another argument for explicitly considering extracellular enzymes in ecosystem-scale processes is that they represent a semiautonomous entity in themselves: they can be functional in the soil matrix after the cells that released them have already died, adding to the complexity of pulse dynamics in soils, such as drying-rewetting or freeze-thaw events (Burns et al. 2013). Furthermore, climate change could bring about changes in the stability of extracellular enzymes in the soil.
One potential advantage to explicitly including extracellular enzymes in ecosystem models might be to more realistically model temperature responses and acclimation to climate change. Enzymes and the whole microorganisms that produce them do not share the same temperature envelopes: psychrophilic enzymes have higher temperature optima than the optimal growth temperature for the whole cell (Huston et al. 2000), and thermophilic enzymes can function at higher temperatures than thermophilic microbes can tolerate (Cowan 2004). This disparity between temperature optima of microbes and their enzymes is one factor in decoupling of respiration and growth that can occur at supraoptimal temperatures in soils (Pietikäinen et al. 2005). Not only is the maximum catalytic rate (V max) of enzymes controlled by temperature, but the substrate affinity (K m) is also sensitive to temperature (German et al. 2012). This cross-latitudinal study found that soil extracellular enzymes had higher affinity for their substrates at low temperatures, partly offsetting decreases in V max, and in the case of β-glucosidase, the K m of enzymes from an Alaskan soil were more sensitive to temperature than those from Costa Rica. This demonstrates that adaptations for high activity at low temperatures can come with a cost in affinity as temperatures are raised.
One final consideration that could earn enzymes (both intracellular and extracellular) some respect in global models is the requirement for trace elements. Enzymes involved in the production and consumption of greenhouse gases, CH4 and N2O, require metals such as Fe, Ni, Cu, Zn, Mo, and W (Glass and Orphan 2012; Wang et al. 2013b), and several phenol oxidases involved in lignin degradation require Cu or Mn (Sinsabaugh 2010). It is established that trace element limitation is a major factor in oceans (Morel and Price 2003) and that alternative nitrogenase enzymes (the key enzyme in N fixation) use V in soils that are poor in Mo (Bellenger et al. 2014). Local limitations in micronutrients could provide surprises in how the soil environment constrains microbial activity.
3.2.5 Stress Tolerance
The capacity for soil microbes to deal with stresses, such as drought or extreme temperatures, will influence ecosystem responses to disturbance and climate change (Schimel et al. 2007). Evolutionary trade-offs, such as the one between rate and yield, can constrain how microbial communities respond to climate change (Wallenstein and Hall 2012). For example, the ecological strategies of soil microbes may also predict the resistance and resilience of the communities to global change, with communities dominated by K-strategists hypothesized to be more resistant to disturbance and those dominated by r-selected microbes to be more resilient (de Vries and Shade 2013).
Local adaptations of microbial communities can lead to unpredictable responses to climate. Microbial responses to climate change experiments tend to vary by ecosystem (Castro et al. 2010; Weber et al. 2011; A’Bear et al. 2014b; Lipson et al. 2014; Classen et al. 2015), showing that microbial communities vary widely in their resistance to disturbance and stress. In arid and semiarid ecosystems, the drought resistance of soil microbes (especially fungi and their extracellular enzymes) and of biological soil crusts can lead to surprising levels of biological activity under very dry conditions (Collins et al. 2008; Austin 2011).
Dry ecosystems are subject to pulse dynamics from drying-rewetting events that cause the soil microbial community and biomass fluctuate rapidly, leading to ecosystem losses of C and N that are not captured by standard models (Collins et al. 2008; Inglima et al. 2009; Dijkstra et al. 2012). During these windows of disequilibrium, the local characteristics of microbial communities can have large impacts on ecosystem function (Lawrence et al. 2009; Placella et al. 2012; Kuzyakov and Blagodatskaya 2015). For example, different phylogenetic groups responded at different rates to rain pulses in California grasslands (Placella et al. 2012). In this study, the set of bacterial groups implicated in immediate, intermediate, and delayed responses was surprising, as the rapid responders included Actinobacteria and Verrucomicrobia, groups generally considered slow-growing and K-selected, while the delayed responders included Proteobacteria associated with rapid growth rates. The authors found that the rapid responders already had high ribosome levels in the pre-wet soils, indicating that the stress resistance of these microbes contributed to their ability to rapidly exploit these pulses, and speculated that the extracellular enzymes these bacteria are known to produce may have been stabilized in the soil matrix and instantly available for activity upon rewetting. Conversely the fast-growing, less stress-tolerant microbes represented a smaller, dormant pool before wetting and therefore required time to become active and grow. Spore formers (Bacillus spp.) showed an intermediate response, as they also required time to germinate but are adapted to rapidly respond to such pulses.
In arid and semiarid ecosystems, biological soil crusts can have major impacts on ecosystem functioning and responses to pulses (Austin et al. 2004; Delgado-Baquerizo et al. 2013). Collins et al. (2008) proposed a “fungal-loop” model for arid ecosystems in which a network of fungi connect plants and biological soil crusts, allowing interchange of fixed N and C. Biological soil crusts are sensitive to disturbance, including N deposition and climate change (Johnson et al. 2012). The continued loss of soil crusts could drastically alter processes such as N fixation, C storage, and plant community dynamics in these ecosystems (Bowker et al. 2014). Incorporating the spatial heterogeneity of biological soil crusts and the “resource islands” produced by patchy plant distribution in dry ecosystems would pose a challenge for ecosystem models but might be worth the effort in terms of improving understanding of processes in arid ecosystems (Austin 2011).
3.3 Microbial Community Interactions that Impact Ecosystem Processes
3.3.1 Microbial Food Webs
Protozoal grazers, soil animals, and viruses can alter outcomes of climate change experiments and alter temperature responses of microbial communities (A’Bear et al. 2014a; Crowther et al. 2015; Pelini et al. 2015). Top-down control of microbial biomass would have a big impact on modeling respiration and other microbial activities, as it is generally assumed that microbes are limited by supply of C or nutrients. For example, climate change models have assumed that elevated CO2 will stimulate soil respiration and methanogenesis in the Arctic by increasing the flux of labile C to soil microbes (Melton et al. 2013), but in C-rich peat soils like those in the Arctic Coastal Plain of Alaska, C additions do not result in stimulation of methanogenesis (von Fischer et al. 2010) or respiration (Allen et al. 2009), and bacteriophage may represent an important top-down control that limits microbial responses to increased substrate (Allen et al. 2009).
3.3.2 Competition and Synergism in Microbial Communities
There are cases where positive and negative interactions within microbial communities have impacts on larger-scale processes, and it may sometimes improve the predictive capabilities of ecosystem models to include these processes. Microbes compete for energy sources, mineral nutrients, and electron acceptors. In an individual-based model of social interactions within microbial communities, it was found that “cheaters” that benefit from extracellular enzymes produced by other microbes can lead to retention of N and accumulation of soil organic matter (Kaiser et al. 2015). Similarly, modeled changes in enzyme production activities depend on interactions among consortia of complimentary microbes that produce extracellular enzymes for acquiring different nutrients (C, N, or P) (Folse and Allison 2012). In anoxic environments, the presence of alternative acceptors can inhibit less thermodynamically favorable pathways. For example, ferric iron [Fe(III)] and humic acids can inhibit sulfate reduction and methanogenesis, though these processes can also coexist in environments where energy is plentiful (Lovley and Phillips 1987; Keller et al. 2009; Miller et al. 2015). Synergistic relationships also contribute to the coexistence of otherwise competing functional groups. For example, the Fe(III)-reducer, Geobacter metallireducens, can donate electrons from the fermentation of ethanol to the methanogen, Methanosarcina barkeri, in a process known as direct interspecies electron transfer (DIET) (Rotaru et al. 2014). There is a plethora of potentially competitive and synergistic interactions that influence the relative and absolute production rates of CO2 and CH4 in soils. Given that CH4 has about 34 times the greenhouse warming potential of CO2, when considered over a 100-year time span (Myhre et al. 2013), the relative fluxes of CH4 and CO2 will have a big impact on the dynamics of climate change over the next century. Hydrogenotrophic methanogens, which use H2 gas, must compete with many other groups for this prized substrate. Acetate can sometimes accumulate in soils as the end product of fermentation and acetogenesis reactions, eventually leading to the establishment of acetoclastic methanogens that can exploit this pool (Hines et al. 2008). Why does it matter if a model includes these two different methanogenic pathways? The two functional groups probably respond differently to environmental factors such as temperature and pH and also to biological factors like competition for substrate or synergistic relationships with fermenters. Therefore, hydrogen- and acetate-dependent methanogens could respond differently to changes in C flux through the soil community driven by elevated CO2 or warming. In a permafrost thaw gradient in Sweden, increasing methane fluxes were associated with a switch to more acetoclastic production, and these changes coincided with the abundance of Candidatus Methanoflorens stordalenmirensis (McCalley et al. 2014). Another warming study found changes in the functional groups responsible for pathways leading to methane production (polysaccharide breakdown, fermentation, and methanogenesis), with different steps limiting the overall process at different temperatures (Tveit et al. 2015).
The majority of CH4 that is produced in soils is thought to be oxidized to CO2 by methanotrophs before it leaves the soil (Le Mer and Roger 2001). The existence of two rapid, nearly balanced processes that are subject to different controls could produce complex behavior, such as rapid spikes or crashes in net fluxes as the production and consumption become uncoupled. Until recently, all methanotrophic activity was mainly attributed to groups of bacteria within the Proteobacteria phylum (Methylocystaceae, Methylococcales; Table 3.1). However, acidophilic methane oxidizers within the Verrucomicrobia phylum have recently been described (Pol et al. 2007), and anaerobic oxidation of methane (AOM), originally discovered in marine environments, also plays a role in soils (Smemo and Yavitt 2011). The presence of alternative electron acceptors in methanogenic environments can lead to a reduction of CH4 flux to the atmosphere due to the activity of AOM species or consortia. The presence of AOM could have the same impact as aerobic methanotrophs, except their activity would be harder to predict. Dissolved oxygen can be modeled relatively simply based on the water table height, but AOM relies on the presence of a variety of other alternative electron acceptors [e.g., nitrite, sulfate, Fe(III)] that are not as easily modeled.
Like CH4, nitrous oxide (N2O) is a powerful greenhouse gas that is subject to complex transformations in soils by diverse groups of microbes. By our current understanding, N2O is produced by (1) facultative anaerobes (including bacteria, archaea, and fungi) that use either nitrite or nitrate as terminal electron acceptors in anaerobic respiration (including denitrification and dissimilatory nitrate reduction to ammonium, DNRA), (2) nitrifiers (chemoautotrophic bacteria and archaea that oxidize \( {\mathrm{NH}}_4^{+} \) for energy using oxygen), (3) anammox (anaerobic oxidation of ammonium using nitrite as the terminal electron acceptor, carried out by chemoautotrophic bacteria within the Planctomyces phylum), and (4) codenitrification (a process carried out by fungi and bacteria in which reduced N compounds, such as ammonium, hydroxylamine, or amino acids, react with oxidized forms of N such as nitrite or nitric oxide) (Giles et al. 2012; Long et al. 2012; Mothapo et al. 2015; Stein and Klotz 2016). The full denitrification pathway leads to the production of N2, with intermediate products NO and N2O emitted depending on the level of oxygen limitation in the soil. Some denitrifying microbes lack N2O reductase, including most denitrifying fungi, and so have a truncated pathway that strictly leads to N2O production rather than N2 (Mothapo et al. 2015; Roco et al. 2016). Recent studies showed that N2O can be consumed by a wider diversity of soil microbes than recently thought (Jones et al. 2012; Sanford et al. 2012). The diversity of microbes that produce and consume N2O leads to complex controls over N2O emissions from soils, as these groups may have different production efficiencies and respond differently to environmental controls such as pH, temperature, oxygen, and energy availability (Pan et al. 2013; Stieglmeier et al. 2014; Jiang et al. 2015; Mothapo et al. 2015).
Microbial community interactions that influence soil processes depend on small spatial scales, such as redox gradients across soil horizons or soil aggregates, diffusion rates between extracellular enzymes and microbial colonies, and differing processes in the rhizosphere, bulk soil, and litter layer compartments. Incorporating this small-scale spatial heterogeneity into the understanding of processes at larger scales is a challenge but is probably worth the effort, at least in some cases (Faust and Raes 2012; Folse and Allison 2012; Giles et al. 2012; Schimel and Schaeffer 2012; Kaiser et al. 2015; Kuzyakov and Blagodatskaya 2015).
3.4 The Impact of Plant-Microbe Interactions on Ecosystem Processes and Global Change
3.4.1 Global Change and Nutrient Cycling
Soil microbes represent an important feedback in the growth responses of plants to global change by modulating the availability of nutrients (Lipson and Kelley 2014). Elevated atmospheric CO2 stimulates photosynthesis rates in the short term, but plants in natural environments generally experience limited growth benefits from elevated CO2 due to nutrient limitations (notably N in most temperate ecosystems). As a result of increased photosynthesis while growth is constrained by nutrient limitations, N concentrations generally decrease in plant tissues when grown under elevated CO2. This lower quality litter can slow down N mineralization in soils, leading to progressive N limitation. Because of this effect, the stimulation of the terrestrial C sink under elevated CO2 is expected to be finite, as plants run out of limiting mineral nutrients from the soil (Ciais et al. 2013). However, increased root growth and microbial activity can partly counteract progressive N limitation (Finzi et al. 2007). Elevated CO2 generally increases plant allocation to roots and to the soil community. The increased roots help plants mine for nutrients more effectively, and increased “rhizodeposition” can stimulate N fixers and mycorrhizae, leading to increased nutrient acquisition. Increased flow of labile exudates from roots to soil can also stimulate N cycling rates through “priming” effects, in which heightened activity in the rhizosphere leads to increased mineralization of N from soil organic matter. And to make matters even more complicated, increased soil temperature generally speeds up N mineralization, partly compensating for progressive N limitation (Dieleman et al. 2012). In summary, the responses of plant growth to climate change depend on changes to the N cycle, which in turn are driven in opposing directions by microbial responses to elevated CO2 and temperature. These biological feedbacks lead to large model uncertainties and are probably best resolved by studies that explicitly examine soil microbes and their interactions with plants and multifactorial climate change.
3.4.2 Soil Feedbacks and Plant Community Change
In climate change experiments, changes in plant communities tend to overwhelm effects of elevated CO2 and temperature (Classen et al. 2015; Steinauer et al. 2015), and so when plant communities change as a result of direct human disturbance or climate change, the rules change completely for the soil microbes. Changes of plant communities to an alternative stable state are often facilitated by feedbacks through the soil microbial community (van der Putten et al. 2016). These can be manifested as changes in nutrient cycling or in a shift of the microbial community in terms of the mutualism-parasitism axis. For example, initial disturbance can allow the establishment of plants with higher litter N content, leading to a faster mineralization rate, further encouraging the invasion by weedy species (Liao et al. 2008; Castro-Díez et al. 2014). Or initial introduction of an exotic plant can favor the presence of microbes that are beneficial or neutral to the invading species and harmful to the natives (Sigüenza et al. 2006; Callaway et al. 2008). The microbial community, if left undisturbed and intact, could also function to limit the invasion of a new community by preferentially benefitting native species (Bozzolo and Lipson 2013; Abbott et al. 2015). Similarly, changes in plant ranges can be limited by the presence of suitable symbiotic microbes. Ectomycorrhizal fungi seem to be more subject to dispersal limitation than arbuscular mycorrhizae (Peay et al. 2010; Davison et al. 2015). For example, invasion by ectomycorrhizal trees into a heathland dominated by ericaceous shrubs is reportedly limited by the influx of mycorrhizal spores (Collier and Bidartondo 2009). Conversely, the maintenance of ectomycorrhizal fungi can help plant species maintain the trailing edge of their range as the climate changes (Lankau et al. 2015).
3.5 Relating Soil Microbial Community Structure to Ecosystem Function
The previous sections dealt with how the individualistic properties of soil microbial species and their interactions might influence ecosystems. But how are these microbial traits and interactions expressed when aggregated into complex communities with tens of thousands of species? Because of their complexity, soil microbial communities are generally described in broad terms, such as the relative proportion of major taxonomic groups (like phyla or classes), or by other coarse metrics like bacterial/fungal ratio. Given the premise that microbial species composition matters at larger scales, how can we know which species are important and how the general structure of communities influences ecosystem processes? One also needs to keep in mind that the relative abundance of microbes is not necessarily proportional to their importance in ecosystem functioning. For example, rare microbes can have very small but active populations due to rapid turnover from predation (Lynch and Neufeld 2015; Neuenschwander et al. 2015). And it might be expected that the microbes that contribute most to CO2 flux might be fast-growing r-selected types that respond quickly to resource pulses but otherwise have low populations most of the time. However, many studies have shown relationships between broad-scale microbial community structure and function, as detailed below.
3.5.1 Predicting Ecological Strategies from Taxonomy
The taxonomic structure of microbial communities varies greatly among soils of the world and is strongly influenced by soil properties (such as pH, texture, and organic and matter content) and by the plant community (Högberg et al. 2007a; Fierer et al. 2009; Caporaso et al. 2011; Chau et al. 2011; Legay et al. 2014; Docherty et al. 2015). Most descriptions of soil microbial communities are focused on bacteria and are based on the sequences of 16S rRNA genes, though a growing number of studies describe entire microbial communities using shotgun sequencing of the soil metagenome (Fierer et al. 2012). While there can be remarkable physiological diversity among closely related bacterial species (Jaspers and Overmann 2004; Hahn and Pöckl 2005), some general trends have emerged that link broad taxonomic groups with ecological strategies (Fierer et al. 2007; Philippot et al. 2010; Goldfarb et al. 2011; Evans and Wallenstein 2014). For example, the Acidobacteria phylum is very common in soils but has few cultured representatives, all of which grow quite slowly (Ward et al. 2009). This group appears to represent a K-selected (or oligotrophic) strategy, growing slowly on complex substrates derived from plant tissues and tolerating stresses. Some species within the Acidobacteria have also been implicated as rhizosphere dwellers (da Rocha et al. 2013). On the other extreme are groups such as the Betaproteobacteria, containing many cultured representatives and representing an r-selected (or copiotrophic) strategy, growing rapidly on labile substrates like amino acids and taking advantage of disturbances that increase resource availability, but with higher sensitivity to stress.
The ratio of bacteria to fungi is another broad index that is linked to soil processes. Fungi are generally associated with improving C sequestration in soils (Six et al. 2006; Fontaine et al. 2011; Waring et al. 2013). Filamentous fungi generally have an advantage over bacteria when growing in complex, high C:N substrates, especially those rich in lignin, and in low pH soils (Högberg et al. 2007a). Their nutrient requirements are lower, having a more flexible C:N ratio, and their extensive hyphal network allows them to exploit sporadic hotspots of resource availability in an otherwise resource-poor environment. The dominance of fungi is associated with nutrient retention in soils and slower mineralization rates (Allen and Zink 1998; Högberg et al. 2007b; Waring et al. 2013). High fungal/bacterial ratios are linked with lower biomass-specific respiration rates (Sakamoto and Oba 1994; Lipson et al. 2005; Six et al. 2006; Lipson et al. 2009). Fungi tend to have lower growth rates and turnover rates than do bacteria in soil (Rousk and Bååth 2011), which would generally lead to lower CO2 production per unit biomass. However, this may not be universally true (Thiet et al. 2006). Fungi are physiologically diverse and include a variety of ecological strategies, growth rates, and growth yields. There is increasing evidence that mycorrhizal fungi have direct and indirect effects on soil C storage. Mycorrhizal fungi, in addition to receiving C from their host plant, can also degrade soil organic matter, either in search of mineral nutrients or for supplemental energy (Talbot et al. 2008). There is also evidence that ectomycorrhizae stimulate C sequestration in soils by competing with saprotrophs for soil nutrients (Averill et al. 2014).
Soil bacteria and fungi generally respond differently to climatic variables. In a study of two soils in Sweden (an agricultural soil and a forest soil), fungal growth was more sensitive to warming than was bacterial growth (Pietikäinen et al. 2005). Additionally, fungal species have been noted as having individualistic phenological responses to past climatic variation, with saprotrophic and mycorrhizal groups associated with deciduous and evergreen trees responding differently to patterns in temperature and rainfall (Diez et al. 2013). These observations indicate that warming could lead to functional changes in fungal communities, with some species increasing vegetative growth and respiration due to delayed fruiting body formation. Several studies have shown differential effects of elevated CO2 on bacteria and fungi, though the results vary by ecosystem (He et al. 2010; Anderson et al. 2011; Lipson et al. 2014).
3.5.2 Assigning Ecological Roles Based on DNA Sequence Data
Numerous techniques are now available to study the functional roles of uncultured environmental microbes, such as shotgun sequencing of soil metagenomes, PCR-based surveys of functional genes, stable isotope probing, fluorescent in situ hybridization and other advanced imaging techniques, single cell genomics, and other innovative approaches such as epicPCR, in which functional genes and 16S rRNA genes from the same cell can be linked (Spencer et al. 2015). However, many descriptions of soil microbial communities are based on 16S rRNA genes, and so it is convenient if conclusions can be drawn from these taxonomic data regarding the functional capabilities of the community. While many prokaryotic taxa are extremely physiologically diverse (e.g., most of the Proteobacteria classes), there are some taxonomic groups that share a reasonably coherent lifestyle (Table 3.1). While it is harder to make generalizations about the relationship between phylogeny and function at finer taxonomic scales, recent studies support the idea that there are consistent relationships between the phylogenetic placement of a bacterial operational taxonomic unit’s (OTU) 16S rRNA gene and its ecological role (Langille et al. 2013).
3.5.3 Temperature Responses Versus Taxonomy
Because of the strong influence of soil chemistry on microbial community structure, no differences are detected between tropical, temperate, and arctic biomes when communities are compared at a broad (e.g., phylum-level) phylogenetic scale (Fierer and Jackson 2006; Fierer et al. 2012). However, temperature is clearly an important selective factor given the tight adaptations to ambient conditions reported in many studies (Bennett and Lenski 1993; Giardina and Ryan 2000; Bárcenas-Moreno et al. 2009; Salvadó et al. 2011; Rousk et al. 2012). In fact, the effect of temperature is so fundamental that cold-adapted species occur in nearly every major bacterial phyla (Margesin and Miteva 2011). This could explain the difficulty in detecting a clear temperature signature in overall community comparisons that are driven by phylum-level differences. However, when focusing on a narrow group of microbes, latitudinal patterns have been observed at a finer taxonomic scale (Rodrigues et al. 2009; Robador et al. 2015). Similarly, a correlation was observed between the diversity of Betaproteobacteria and temperature responses with changes in season and altitude (Lipson 2007).
Adaptations to temperature occur over the entire genome. For example, each enzyme must be adapted for ambient conditions by having the optimal flexibility for functioning at low temperatures or, conversely, high stability for functioning at higher temperatures (Feller and Gerday 2003). Although cold-tolerance genes have been found in plasmids (Dziewit and Bartosik 2014), it is unlikely that a single mobile element could transform a mesophile into a high functioning psychrophile capable of competing within a diverse community. Therefore temperature adaptation should leave an evolutionary signature on a taxonomic marker gene like 16S rRNA, as observed for other complex traits (Langille et al. 2013).
Bacterial genomes from extremely cold environments such as the Arctic, Antarctic, and permafrost zones show very clear signatures of cold adaptation, such as alterations in amino acid composition of the proteins, changes in membrane composition, and enhanced expression of cold shock proteins (Bakermans et al. 2012; Kuhn 2012). However, there is considerable variability among different species and environments (Grzymski et al. 2006; Ayala-del-Río et al. 2010). We are still years away from being able to predict the temperature responses of complex soil processes from metagenomic data, but in principle, all the information is there.
3.5.4 Emergent Properties of Microbial Communities: The Importance of Diversity in Ecosystem Functioning
To this point, we have only considered the constituent microbes that make up microbial communities and how their relative abundance might impact ecosystem processes. But biodiversity (including species richness and evenness) is a property of biological communities with important ecological consequences (McCann 2000), and this appears to be true for microbial communities as well (Ferris and Tuomisto 2015). Diversity-function relationships are often nonlinear, with decreasing impacts of diversity at high levels of species richness. Given the high diversity of soil microbial communities, it would be expected that species-function relationships in soil communities would be fairly flat or require drastic reductions in diversity to see an effect in manipulative experiments. Consistent with this logic, relationships between soil microbial diversity and C cycling are found more frequently in experiments with low diversity levels, and it is often found that because of the high degree of functional redundancy in these communities, species composition matters more than species richness (Nielsen et al. 2011). Nonetheless it still appears that microbial biodiversity is important for the overall functioning of ecosystems and that even given the high functional redundancy within microbial communities, increased diversity can increase process rates (Nielsen et al. 2015). More diverse microbial communities are also more stable and resistant to invasion by pathogens (van Elsas et al. 2012).
The impacts of climate change on microbial diversity vary by ecosystem and microbial group (Nielsen et al. 2015). Elevated CO2 increased fungal diversity in a semiarid shrubland (Lipson et al. 2014), but a similar effect was only seen in two of seven ecosystems in a different study (Weber et al. 2011). In an analysis of several long-term studies, drought stress and pressure on pinyon pine from competitors, herbivory, and parasites decreased the diversity of ectomycorrhizal fungi but did not tend to decrease their mutualistic benefit to the plant host (Gehring et al. 2014). Warming led to increased species evenness in the bacterial community (DeAngelis et al. 2015). Theoretically, more diverse communities should be more resistant to disturbances. Therefore, disturbances or land management practices that reduce soil microbial diversity could lead to increased vulnerability of microbial communities and ecosystems.
3.6 Integrating Microbial Diversity and Physiology into Ecosystem Models
It has been well recognized that microbial mechanisms dominate the biological aspects of soil biogeochemistry (Jenkinson and Ladd 1981; Staley et al. 1997; Schimel and Gulledge 1998; Falkowski et al. 2008). However, soil models do not always simulate the microbial roles on biogeochemistry in an explicit way (Schimel 2001). For example, first-order differential equations have been broadly used to describe transformation rates of soil carbon and nitrogen pools, with rate constants for these equations controlled by a variety of environmental factors such as soil temperature, moisture, soil pH, texture, etc. (Manzoni and Porporato 2009). It has been argued that these traditional soil models do simulate microbial mechanisms implicitly, as the microbial impacts are embedded in the decomposition rate constant, k (turnover rate of the pools) (Schimel 2001; Manzoni and Porporato 2009). However, compared to microbial kinetics, the first-order differential equations lack feedbacks because they are developed based on a strategy of donor-controlled flow. The mechanistic controls (primarily physiological and structural feedback) from microbes are ignored.
3.6.1 Emergence of Microbial Models
Modeling microbial processes started as early as the development of the first soil organic matter model (Veen and Paul 1981; Van Veen et al. 1984; Jenkinson et al. 1987; Parton et al. 1987). When the early soil organic matter models were built, soil microbes were ignored. Later model evolution wherein the microbial pool was treated as a small labile pool without any feedbacks to either upstream litter or soil organic matter decomposition did not result in significant improvements of microbial representation (Jenkinson et al. 1987; Schimel 2001). For example, the VVV model (Veen and Paul 1981; Van Veen et al. 1984), Century model (Parton et al. 1987) and RothC model (Coleman and Jenkinson 1996) separate microbial biomass as an independent labile carbon pool, while none of the three models explicitly simulate the impact of microbial regulation on litter and soil organic matter decomposition. This lack of feedbacks has been identified as a potential uncertainty for projecting soil carbon dynamics with Earth system models (Wieder et al. 2015; Luo et al. 2016). Therefore, there is a strong call for developing a microbial modeling framework for use in Earth system models (DeLong et al. 2011; Treseder et al. 2012; Xu et al. 2014).
3.6.2 Classification of Microbial Physiology and Diversity Simulated in Selected Models
A number of microbial models have been developed (Allison et al. 2010; Allison 2012; Wieder et al. 2013; Sulman et al. 2014). Some are individual-based microbial models, emphasizing the societal and interactions between microbes (Kaiser et al. 2014, 2015); some are functional group-based microbial models to simulate trade-offs among different microbial functions (Wieder et al. 2013, 2015; Xu et al. 2015); some are enzyme-kinetics microbial models, emphasizing the dynamics of microbial processes in response to different substrate quality and environmental conditions (Allison et al. 2010; Wang et al. 2013a). In this section, we will review the state of the art of microbial models simulating microbial physiology and diversity and show the gaps which evidence need for future modeling efforts. Microbial models consider various microbial traits or functions. Categorized below are the primary aspects of microbial physiology and diversity that have been modeled (Table 3.2, Fig. 3.2). This section reviews a few of the typical microbial models developed over the past decade (it is not intended to be a comprehensive review).
Conceptual diagram showing microbial physiology and diversity in models (A) physiological limits, (B) microbial growth, (C) plant-microbe interaction, (D) stoichiometry, (E) microbial interaction, (F) microbial dormancy, and (G) microbial community structure; solid lines indicate flows and dashed lines indicate controls; the red dash lines represent microbial regulation of soil organic matter decomposition; the dashed rectangle represents the entire microbial population in soils, which is composed of different states and functional groups of microbes
3.6.2.1 Physiological Limits
All living organisms have limits for physiological functioning. The physiological limits could be soil temperature, soil moisture, soil pH, oxygen, substrate avail ability, etc. For example, the lowest currently reported temperature for soil microbial respiration is −39 °C, although it has been suggested that activity at lower temperatures is possible (Panikov et al. 2006). Models normally simulate microbial activities over a limited range of temperature and soil moisture. For example, −2 °C has been considered as a threshold for microbial activities in the microbial community land surface model, CLM-Microbe (Xu et al. 2014), which simulates the seasonality of microbial activities in response to changes in soil temperature and moisture. In addition, soil physical conditions provide limits for microbial physiology. For example, MIMIC model, CORPS (Sulman et al. 2014), and the model of Tang and Riley (Tang and Riley 2015) simulate the effects of physical protection on microbial carbon cycling. Manzoni’s model simulates water diffusion and its impacts on microbial activity (Manzoni et al. 2014). All these physiological limits control the microbial activity to maintain microbes functioning well under favorable conditions while avoiding detrimental conditions. A good simulation of microbial physiological limits is fundamental in order for models to accurately capture the real dynamics of ecosystem functions.
3.6.2.2 Microbial Growth
Microbial growth is the fundamental component in microbial models designed to simulate microbial mechanisms and their controls on ecosystem functions. Normally microbial growth is a function of substrate and microbial uptake under the control of environmental factors, simulated by Monod and Michaelis-Menten functions. Most microbial models simulate microbial assimilation of carbon. The CLM-Microbe model simulates microbial assimilation of soil organic carbon under the control of litter quality and microbial physiology and microbial biomass as a net balance between growth and respiration (Xu et al. 2014). The CUE parameter has been considered as an important factor controlling microbial carbon assimilation and carbon sequestration in global soils because it determines how much carbon is released as CO2 versus how much carbon is used for biomass buildup (Manzoni and Porporato 2009; Wieder et al. 2013; Xu et al. 2014; Wang et al. 2013a, b). The environmental controls on microbial growth are another important aspect of microbial modeling, for example, warming impacts on microbial growth efficiency (similar to CUE) (Wieder et al. 2013; Xu et al. 2014) and moisture impacts on microbial activities (Manzoni et al. 2014). Microbial metabolic quotient (the biomass-specific microbial respiration rate) is another important parameter for simulating microbial activities that can benefit the performance of microbial models (Xu et al. 2017).
3.6.2.3 Plant-Microbe Interactions
Plant roots have strong impacts on microbial growth and uptake of nutrients. Of the developed microbial models, the CORPS model simulates plant impacts on microbial cycling of soil carbon (Sulman et al. 2014), and the CLM-Microbe model explicitly simulates root exudation. However, most microbial models are based upon a theoretical framework and have not been incorporated into real ecosystem models. Therefore, microbial models typically lack the important aspects of interactions with plants and plant roots. Plant-microbe interactions have a variety of ecological consequences (Kuzyakov and Xu 2013). For example, plant-microbe competition for nitrogen affects carbon sequestration of the ecosystem (de Vries and Bardgett 2012), and plant-microbe interactions facilitate plant diversity and production (Van Der Heijden et al. 2008). Considering the centrality of plant-microbe interactions to the biology of both plants and soil microbes, a larger investment in these phenomena is warranted in future microbial modeling development and application. The impact of roots on microbial activities is a particularly important mechanism models should represent.
3.6.2.4 Stoichiometry
Substrate quality (primarily expressed as C:N stoichiometry or lignin content) controls microbial activity. The C:N ratio has been explicitly simulated in SCAMPS (Sistla et al. 2014) and implicitly in CLM-Microbe (Xu et al. 2014). The GDM model uses a lignocellulose index (LCI) to simulate substrate quality impacts on litter decomposition. The LCI is well correlated with litter stoichiometry. The individual-based microbial model, such as Kaiser’s model, also explicitly simulates microbial community dynamics and stoichiometry during litter decomposition (Kaiser et al. 2014). Strong microbial homeostatic regulation has also been found for nitrogen, phosphorus, and sulfur (Zechmeister-Boltenstern et al. 2015; Sinsabaugh et al. 2016). We need to further advance our understanding with a framework that explicitly simulates both microbial dynamics in assimilating substrates with varying stoichiometry and how microbes respond to different qualities of substrates. The interplay of stoichiometry in litter and the microbial community under changing environmental conditions is additional critical information needed for better simulations of microbial physiology.
3.6.2.5 Microbial Community Interactions
Interactions within microbial communities are widely recognized as having important controlling effects upon microbial activity (Faust and Raes 2012). For example, microbial interactions lead to evolutional separation of generalists and specialism for enzyme production (Nam et al. 2012), cheaters and producers of enzyme production (Travisano and Velicer 2004). Yet, these interactions have not been well simulated in most microbial models. There are a few approaches used in microbial models to simulate different groups of microbes. For example, the MIMIC model uses the r- and K-strategies for separating the microbial groups, as suggested by Fierer in a previous concept paper (Fierer et al. 2007). The GDM model simulates interactions among three guilds of microbes (groups of microbes that exploit the same resources, see Moorhead and Sinsabaugh 2006). Kaiser’s model simulates societal interaction among individual microbes (Kaiser et al. 2015). The Decomposition Model of Enzymatic Traits (DEMENT) model simulates trade-offs between different functional groups (Allison 2012). Although multiple microbial groups or traits have been simulated to a certain degree in some microbial models, the representation of microbial community interactions in models is far from complete. Given the importance of microbial interactions in ecosystem functions and microbial evolution (Faust and Raes 2012), more effort should be invested to modeling microbial interactions and their impacts on ecosystem functions.
3.6.2.6 Microbial Dormancy
The majority of microbial biomass in soils is in an inactive state during most of the year. Because only a small portion of the microbial biomass is active, the ecosystem functions are carried out by this active microbial biomass. The active microbial biomass and dormant biomass should be differentiated, and this separation has proved to be important for better simulating microbial processes (Wang et al. 2015). Over the past years, few microbial models have been developed to simulate the dormant microbial biomass either as an independent pool or as a season over certain time period. Both the MEND model and Manzoni’s model have separated active versus dormant microbial biomass from the total microbial biomass pool (Manzoni et al. 2014; Wang et al. 2015). In a different approach, the CLM-Microbe model simulates temporally separated active microbial biomass and its impacts on litter mineralization as a seasonality component of microbial functioning (Xu et al. 2014). Both approaches have been proved to be robust in simulating microbial biomass and their contribution to carbon cycling.
3.6.2.7 Community Structure
The Guild Decomposition Model (GDM) is among the first to simulate the dynamic of microbial community shift during litter decay (Moorhead and Sinsabaugh 2006). The MIMIC model simulated two microbial functional groups (MICr and MICk) to represent r- and K-strategists (Wieder et al. 2014). The SCAMPS model is a mechanistic microbial model explicitly simulating the separation of bacteria- and fungi-like microbes and the interplay dynamics of these two groups of microbes (Sistla et al. 2014). Moorhead’s model is based on guilds, which represent the microbial groups with different traits (Moorhead and Sinsabaugh 2006). Xu’s functional group-based methane model (incorporated in CLM-Microbe) also simulates different microbial functional groups and their dynamics in response to substrate and environmental conditions (Xu et al. 2015). These models consider the dynamics of different microbial functional groups, representing the microbial community structure.
In summary, microbial physiology and community structure have been simulated to a certain degree, and some convincing results have been obtained. While much knowledge has accumulated, it is important to note that modeling microbial physiology and diversity is still in its infancy. More effort is particularly needed in the areas of microbial interactions, community structure shifts and their associated changes in microbial functions, ecological stoichiometry of phosphorus beyond carbon and nitrogen, and microbial interactions with plants. All these aspects are beneficial for model improvement in simulating terrestrial microbial biogeochemistry in the context of climate change. We anticipate that the investment of modeling microbial processes in theoretical and applicable ways will pay off with significant contributions to the robustness of Earth system models in one or two decades.
3.7 Conclusion
The broad range of topics relevant to connecting soil microbial communities to ecosystem function underscores the need for interdisciplinary studies. In particular, it is profitable for soil microbiologists and ecosystem modelers to work together, as this can inform microbiologists how to tailor their studies to make them immediately helpful to modelers, while helping modelers become aware of the importance of mechanisms they may not have considered. The field of environmental microbiology has been revolutionized by modern molecular and isotopic techniques. As computers become more powerful, modelers will be less hesitant to build increasing complexity into their models. It is likely that this field is already, or will soon become, limited only by the level of communication among scientists studying the same processes at multiple scales and the imagination of these researchers.
References
A’Bear AD, Jones TH, Boddy L (2014a) Potential impacts of climate change on interactions among saprotrophic cord-forming fungal mycelia and grazing soil invertebrates. Fungal Ecol 10:34–43
A’Bear AD, Jones TH, Kandeler E, Boddy L (2014b) Interactive effects of temperature and soil moisture on fungal-mediated wood decomposition and extracellular enzyme activity. Soil Biol Biochem 70:151–158
Abbott KC, Karst J, Biederman LA, Borrett SR, Hastings A, Walsh V, Bever JD (2015) Spatial heterogeneity in soil microbes alters outcomes of plant competition. PLoS One 10:e0125788
Alexander M (1964) Biochemical ecology of soil microorganisms. Annu Rev Microbiol 18:217–250
Allen A, Schlesinger W (2004) Nutrient limitations to soil microbial biomass and activity in loblolly pine forests. Soil Biol Biochem 36:581–589
Allen MF, Zink TA (1998) The effects of organic amendments on the restoration of a disturbed coastal sage scrub habitat. Restor Ecol 6:52–58
Allen B, Willner D, Oechel WC, Lipson DA (2009) Topdown control of microbial activity and biomass in an Arctic soil ecosystem. Environ Microbiol 12:642–648
Allison SD (2012) A trait-based approach for modeling microbial litter decomposition. Ecol Lett 15:1058–1070
Allison SD (2014) Modeling adaptation of carbon use efficiency in microbial communities. Front Microbiol 5:571
Allison SD, Wallenstein MD, Bradford MA (2010) Soil-carbon response to warming dependent on microbial physiology. Nat Geosci 3:336–340
Anderson T-H, Heinemeyer O, Weigel H-J (2011) Changes in the fungal-to-bacterial respiratory ratio and microbial biomass in agriculturally managed soils under free-air CO2 enrichment (FACE)—a six-year survey of a field study. Soil Biol Biochem 43:895–904
Austin AT (2011) Has water limited our imagination for aridland biogeochemistry? Trends Ecol Evol 26:229–235
Austin AT, Yahdjian L, Stark JM, Belnap J, Porporato A, Norton U, Ravetta DA, Schaeffer SM (2004) Water pulses and biogeochemical cycles in arid and semiarid ecosystems. Oecologia 141:221–235
Averill C, Turner BL, Finzi AC (2014) Mycorrhiza-mediated competition between plants and decomposers drives soil carbon storage. Nature 505:543–545
Ayala-del-Río HL, Chain PS, Grzymski JJ, Ponder MA, Ivanova N, Bergholz PW, Di Bartolo G, Hauser L, Land M, Bakermans C (2010) The genome sequence of Psychrobacter arcticus 273-4, a psychroactive Siberian permafrost bacterium, reveals mechanisms for adaptation to low-temperature growth. Appl Environ Microbiol 76:2304–2312
Baes C, Goeller H, Olson J, Rotty R (1977) Carbon dioxide and climate: the uncontrolled experiment: possibly severe consequences of growing CO 2 release from fossil fuels require a much better understanding of the carbon cycle, climate change, and the resulting impacts on the atmosphere. Am Sci 65:310–320
Bakermans C, Tsapin AI, Souza-Egipsy V, Gilichinsky DA, Nealson KH (2003) Reproduction and metabolism at −10 degrees C of bacteria isolated from Siberian permafrost. Environ Microbiol 5:321–326
Bakermans C, Bergholz PW, Rodrigues DF, Vishnivetskaya TA, Ayala-del-Río HL, Tiedje JM (2012) Genomic and expression analyses of cold-adapted microorganisms. In: Miller RV, Whyte LG (eds) Polar microbiology: life in a deep freeze. American Society of Microbiology, Washington, DC, pp 126–155
Bárcenas-Moreno G, Gómez-Brandón M, Rousk J, Bääth E (2009) Adaptation of soil microbial communities to temperature: comparison of fungi and bacteria in a laboratory experiment. Glob Chang Biol 15:2950–2957
Bellenger J, Xu Y, Zhang X, Morel F, Kraepiel A (2014) Possible contribution of alternative nitrogenases to nitrogen fixation by asymbiotic N 2-fixing bacteria in soils. Soil Biol Biochem 69:413–420
Bennett AF, Lenski RE (1993) Evolutionary adaptation to temperature II. Thermal niches of experimental lines of Escherichia coli. Evolution 47(1):12
Bier RL, Bernhardt ES, Boot CM, Graham EB, Hall EK, Lennon JT, Nemergut DR, Osborne BB, Ruiz-González C, Schimel JP, Waldrop MP, Wallenstein MD, Muyzer G (2015) Linking microbial community structure and microbial processes: an empirical and conceptual overview. FEMS Microbiol Ecol 91:fiv113
Bloom AA, Palmer PI, Fraser A, Reay DS, Frankenberg C (2010) Large-scale controls of methanogenesis inferred from methane and gravity spaceborne data. Science 327:322–325
Bowker MA, Maestre FT, Eldridge D, Belnap J, Castillo-Monroy A, Escolar C, Soliveres S (2014) Biological soil crusts (biocrusts) as a model system in community, landscape and ecosystem ecology. Biodivers Conserv 23:1619–1637
Bozzolo FH, Lipson DA (2013) Differential responses of native and exotic coastal sage scrub plant species to N additions and the soil microbial community. Plant Soil 371:37–51
Bradford MA (2013) Thermal adaptation of decomposer communities in warming soils. Front Microbiol 4:333
Bradford MA, Davies CA, Frey SD, Maddox TR, Melillo JM, Mohan JE, Reynolds JF, Treseder KK, Wallenstein MD (2008) Thermal adaptation of soil microbial respiration to elevated temperature. Ecol Lett 11:1316–1327
Brooks P, Williams MW, Schmidt SK (1996) Microbial activity under alpine snowpacks, Niwot Ridge, Colorado. Biogeochemistry 32:93–113
Brooks PD, Schmidt SK, Williams MW (1997) Winter production of CO2 and N2O from alpine tundra: environmental controls and relationship to inter-system C and N fluxes. Oecologia 110:403–413
Burns RG, DeForest JL, Marxsen J, Sinsabaugh RL, Stromberger ME, Wallenstein MD, Weintraub MN, Zoppini A (2013) Soil enzymes in a changing environment: current knowledge and future directions. Soil Biol Biochem 58:216–234
Callaway RM, Cipollini D, Barto K, Thelen GC, Hallett SG, Prati D, Stinson K, Klironomos J (2008) Novel weapons: invasive plant suppresses fungal mutualists in America but not in its native Europe. Ecology 89:1043–1055
Caporaso JG, Lauber CL, Walters WA, Berg-Lyons D, Lozupone CA, Turnbaugh PJ, Fierer N, Knight R (2011) Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc Natl Acad Sci USA 108:4516–4522
Castro HF, Classen AT, Austin EE, Norby RJ, Schadt CW (2010) Soil microbial community responses to multiple experimental climate change drivers. Appl Environ Microbiol 76:999–1007
Castro-Díez P, Godoy O, Alonso A, Gallardo A, Saldaña A (2014) What explains variation in the impacts of exotic plant invasions on the nitrogen cycle? A meta-analysis. Ecol Lett 17:1–12
Chapin FS, McFarland J, McGuire AD, Euskirchen ES, Ruess RW, Kielland K (2009) The changing global carbon cycle: linking plant–soil carbon dynamics to global consequences. J Ecol 97:840–850
Chau JF, Bagtzoglou AC, Willig MR (2011) The effect of soil texture on richness and diversity of bacterial communities. Environ Forensic 12:333–341
Ciais P, Sabine C, Bala G, Bopp L, Brovkin V, Canadell J, Chhabra A, DeFries R, Galloway J, Heimann M, Jones C, Le Quéré C, Myneni RB, Piao S, Thornton P (2013) Carbon and other biogeochemical cycles. In: Stocker TF, Qin D, Plattner G-K, Tignor M, Allen SK, Boschung J, Nauels A, Xia Y, Bex V, Midgley PM (eds) Climate change 2013: the physical science basis. contribution of Working Group I to the fifth assessment report of the Intergovernmental Panel on Climate Change. Cambridge University Press, Cambridge
Classen AT, Sundqvist MK, Henning JA, Newman GS, Moore JAM, Cregger MA, Moorhead LC, Patterson CM (2015) Direct and indirect effects of climate change on soil microbial and soil microbial-plant interactions: what lies ahead? Ecosphere 6:art130
Clein JS, Schimel JP (1995) Microbial activity of tundra and taiga soils at sub-zero temperatures. Soil Biol Biochem 27:1231–1234
Cleveland CC, Liptzin D (2007) C:N:P stoichiometry in soil: is there a “Redfield ratio” for the microbial biomass? Biogeochemistry 85:235–252
Cleveland CC, Townsend AR, Schmidt SK (2002) Phosphorus limitation of microbial processes in moist tropical forests: evidence from short-term laboratory incubations and field studies. Ecosystems 5:0680–0691
Coleman K, Jenkinson D (1996) RothC-26.3-A model for the turnover of carbon in soil. In: Evaluation of soil organic matter models. Springer, Heidelberg, pp 237–246
Collier FA, Bidartondo MI (2009) Waiting for fungi: the ectomycorrhizal invasion of lowland heathlands. J Ecol 97:950–963
Collins SL, Sinsabaugh RL, Crenshaw C, Green L, Porras-Alfaro A, Stursova M, Zeglin LH (2008) Pulse dynamics and microbial processes in aridland ecosystems. J Ecol 96:413–420
Cotrufo MF, Wallenstein MD, Boot CM, Denef K, Paul E (2013) The Microbial Efficiency-Matrix Stabilization (MEMS) framework integrates plant litter decomposition with soil organic matter stabilization: do labile plant inputs form stable soil organic matter? Glob Chang Biol 19:988–995
Cowan DA (2004) The upper temperature for life–where do we draw the line? Trends Microbiol 12:58–60
Crowther TW, Thomas SM, Maynard DS, Baldrian P, Covey K, Frey SD, van Diepen LTA, Bradford MA (2015) Biotic interactions mediate soil microbial feedbacks to climate change. Proc Natl Acad Sci USA 112:7033–7038
da Rocha UN, Plugge CM, George I, van Elsas JD, van Overbeek LS (2013) The rhizosphere selects for particular groups of acidobacteria and verrucomicrobia. PLoS One 8:e82443
Davison J, Moora M, Öpik M, Adholeya A, Ainsaar L, Ba A, Burla S, Diedhiou A, Hiiesalu I, Jairus T (2015) Global assessment of arbuscular mycorrhizal fungus diversity reveals very low endemism. Science 349:970–973
de Vries FT, Bardgett RD (2012) Plant–microbial linkages and ecosystem nitrogen retention: lessons for sustainable agriculture. Front Ecol Environ 10:425–432
de Vries FT, Shade A (2013) Controls on soil microbial community stability under climate change. Front Microbiol 4:265
DeAngelis KM, Pold G, Topçuoğlu BD, van Diepen LT, Varney RM, Blanchard JL, Melillo J, Frey SD (2015) Long-term forest soil warming alters microbial communities in temperate forest soils. Front Microbiol 6:104
Del Giorgio PA, Cole JJ (1998) Bacterial growth efficiency in natural aquatic systems. Annu Rev Ecol Syst 29:503–541
Delgado-Baquerizo M, Maestre FT, Rodríguez JG, Gallardo A (2013) Biological soil crusts promote N accumulation in response to dew events in dryland soils. Soil Biol Biochem 62:22–27
DeLong EF, Pace NR (2001) Environmental diversity of bacteria and archaea. Syst Biol 50:470–478
DeLong EF, Harwood CS, Chisholm PW, Karl DM, Moran MA, Schmidt TM, Tiedje JM, Treseder KK, Worden AZ (2011) Incorporating microbial processes into climate models. The American Academy of Microbiology, Washington, DC
Dieleman WIJ, Vicca S, Dijkstra FA, Hagedorn F, Hovenden MJ, Larsen KS, Morgan JA, Volder A, Beierk C, Dukes JS, King J, Leuzinger S, Linder S, Luo YQ, Oren R, Angelis PD, Tingey D, Hoosbeek MR, Janssens IA (2012) Simple additive effects are rare: a quantitative review of plant biomass and soil process responses to combined manipulations of CO2 and temperature. Glob Chang Biol 18:2681–2693
Diez JM, James TY, McMunn M, Ibáñez I (2013) Predicting species-specific responses of fungi to climatic variation using historical records. Glob Chang Biol 19:3145–3154
Dijkstra P, Thomas SC, Heinrich PL, Koch GW, Schwartz E, Hungate BA (2011) Effect of temperature on metabolic activity of intact microbial communities: evidence for altered metabolic pathway activity but not for increased maintenance respiration and reduced carbon use efficiency. Soil Biol Biochem 43:2023–2031
Dijkstra FA, Augustine DJ, Brewer P, von Fischer JC (2012) Nitrogen cycling and water pulses in semiarid grasslands: are microbial and plant processes temporally asynchronous? Oecologia 170:799–808
Docherty KM, Borton HM, Espinosa N, Gebhardt M, Gil-Loaiza J, Gutknecht JLM, Maes PW, Mott BM, Parnell JJ, Purdy G, Rodrigues PAP, Stanish LF, Walser ON, Gallery RE (2015) Key edaphic properties largely explain temporal and geographic variation in soil microbial communities across four biomes. PLoS One 10:e0135352
Dziewit L, Bartosik D (2014) Plasmids of psychrophilic and psychrotolerant bacteria and their role in adaptation to cold environments. Front Microbiol 5:596
Evans SE, Wallenstein MD (2014) Climate change alters ecological strategies of soil bacteria. Ecol Lett 17:155–164
Falkowski PG, Fenchel T, Delong EF (2008) The microbial engines that drive Earth’s biogeochemical cycles. Science 320:1034–1039
Fanin N, Fromin N, Buatois B, Hättenschwiler S (2013) An experimental test of the hypothesis of non-homeostatic consumer stoichiometry in a plant litter–microbe system. Ecol Lett 16:764–772
Faust K, Raes J (2012) Microbial interactions: from networks to models. Nat Rev Microbiol 10:538–550
Feller G, Gerday C (2003) Psychrophilic enzymes: hot topics in cold adaptation. Nat Rev Microbiol 1:200–208
Ferris H, Tuomisto H (2015) Unearthing the role of biological diversity in soil health. Soil Biol Biochem 85:101–109
Fierer N, Jackson RB (2006) The diversity and biogeography of soil bacterial communities. Proc Natl Acad Sci USA 103:626–631
Fierer N, Bradford MA, Jackson RB (2007) Toward an ecological classification of soil bacteria. Ecology 88:1354–1364
Fierer N, Strickland MS, Liptzin D, Bradford MA, Cleveland CC (2009) Global patterns in belowground communities. Ecol Lett 12:1238–1249
Fierer N, Leff JW, Adams BJ, Nielsen UN, Bates ST, Lauber CL, Owens S, Gilbert JA, Wall DH, Caporaso JG (2012) Cross-biome metagenomic analyses of soil microbial communities and their functional attributes. Proc Natl Acad Sci 109:21390–21395
Finzi AC, Norby RJ, Calfapietra C, Gallet-Budynek A, Gielen B, Holmes WE, Hoosbeek MR, Iversen CM, Jackson RB, Kubiske ME (2007) Increases in nitrogen uptake rather than nitrogen-use efficiency support higher rates of temperate forest productivity under elevated CO2. Proc Natl Acad Sci USA 104:14014–14019
Folse HJ, Allison SD (2012) Cooperation, competition, and coalitions in enzyme-producing microbes: social evolution and nutrient depolymerization rates. Front Microbiol 3:338
Fontaine S, Henault C, Aamor A, Bdioui N, Bloor JMG, Maire V, Mary B, Revaillot S, Maron PA (2011) Fungi mediate long term sequestration of carbon and nitrogen in soil through their priming effect. Soil Biol Biochem 43:86–96
Freeman C, Ostle N, Kang H (2001) An enzymic ‘latch’ on a global carbon store. Nature 409:149–149
Frey SD, Lee J, Melillo JM, Six J (2013) The temperature response of soil microbial efficiency and its feedback to climate. Nat Clim Chang 3:395–398
García-Palacios P, Vandegehuchte ML, Shaw EA, Dam M, Post KH, Ramirez KS, Sylvain ZA, de Tomasel CM, Wall DH (2015) Are there links between responses of soil microbes and ecosystem functioning to elevated CO2, N deposition and warming? A global perspective. Glob Chang Biol 21:1590–1600
Gehring CA, Mueller RC, Haskins KE, Rubow TK, Whitham TG (2014) Convergence in mycorrhizal fungal communities due to drought, plant competition, parasitism, and susceptibility to herbivory: consequences for fungi and host plants. Front Microbiol 5:306
George IF, Hartmann M, Liles MR, Agathos SN (2011) Recovery of as-yet-uncultured soil Acidobacteria on dilute solid media. Appl Environ Microbiol 77:8184–8188
German DP, Marcelo KRB, Stone MM, Allison SD (2012) The Michaelis-Menten kinetics of soil extracellular enzymes in response to temperature: a cross-latitudinal study. Glob Chang Biol 18:1468–1479
Giardina CP, Ryan MG (2000) Evidence that decomposition rates of organic carbon in mineral soil do not vary with temperature. Nature 404:858–861
Giles M, Morley N, Baggs EM, Daniell TJ (2012) Soil nitrate reducing processes—drivers, mechanisms for spatial variation, and significance for nitrous oxide production. Front Microbiol 3:407
Gillooly JF, Allen AP, Brown JH, Elser JJ, del Rio CM, Savage VM, West GB, Woodruff WH, Woods HA (2005) The metabolic basis of whole-organism RNA and phosphorus content. Proc Natl Acad Sci USA 102:11923–11927
Glass JB, Orphan VJ (2012) Trace metal requirements for microbial enzymes involved in the production and consumption of methane and nitrous oxide. Front Microbiol 3:61
Goldfarb KC, Karaoz U, Hanson CA, Santee CA, Bradford MA, Treseder KK, Wallenstein MD, Brodie EL (2011) Differential growth responses of soil bacterial taxa to carbon substrates of varying chemical recalcitrance. Front Microbiol 2:94
Griffiths BS, Philippot L (2013) Insights into the resistance and resilience of the soil microbial community. FEMS Microbiol Rev 37:112–129
Grzymski JJ, Carter BJ, DeLong EF, Feldman RA, Ghadiri A, Murray AE (2006) Comparative genomics of DNA fragments from six Antarctic marine planktonic bacteria. Appl Environ Microbiol 72:1532–1541
Hagerty SB, van Groenigen KJ, Allison SD, Hungate BA, Schwartz E, Koch GW, Kolka RK, Dijkstra P (2014) Accelerated microbial turnover but constant growth efficiency with warming in soil. Nat Clim Chang 4:903–906
Hahn MW, Pöckl M (2005) Ecotypes of planktonic Actinobacteria with identical 16S rRNA genes adapted to thermal niches in temperate, subtropical, and tropical freshwater habitats. Appl Environ Microbiol 71:766–773
Hanus F, Morita RY (1968) Significance of the temperature characteristic of growth. J Bacteriol 95:736
Hararuk O, Smith MJ, Luo Y (2015) Microbial models with data-driven parameters predict stronger soil carbon responses to climate change. Glob Chang Biol 21:2439–2453
Hartman WH, Richardson CJ (2013) Differential nutrient limitation of soil microbial biomass and metabolic quotients (qCO2): is there a biological stoichiometry of soil microbes? PLoS One 8:e57127
He Z, Xu M, Deng Y, Kang S, Kellogg L, Wu L, Van Nostrand JD, Hobbie SE, Reich PB, Zhou J (2010) Metagenomic analysis reveals a marked divergence in the structure of belowground microbial communities at elevated CO2. Ecol Lett 13:564–575
Henry HAL (2013) Soil extracellular enzyme dynamics in a changing climate. Soil Biol Biochem 56:53–59
Heuck C, Weig A, Spohn M (2015) Soil microbial biomass C:N:P stoichiometry and microbial use of organic phosphorus. Soil Biol Biochem 85:119–129
Hines ME, Duddleston KN, Rooney-Varga JN, Fields D, Chanton JP (2008) Uncoupling of acetate degradation from methane formation in Alaskan wetlands: connections to vegetation distribution. Glob Biogeochem Cycles 22:GB2017. https://doi.org/10.1029/2006GB002903
Högberg M, Högberg P, Myrold D (2007a) Is microbial community composition in boreal forest soils determined by pH, C-to-N ratio, the trees, or all three. Oecologia 150:590–601
Högberg MN, Chen Y, Högberg P (2007b) Gross nitrogen mineralisation and fungi-to-bacteria ratios are negatively correlated in boreal forests. Biol Fertil Soils 44:363–366
Huston AL, Krieger-Brockett BB, Deming JW (2000) Remarkably low temperature optima for extracellular enzyme activity from Arctic bacteria and sea ice. Environ Microbiol 2:383–388
Inglima I, Alberti G, Bertolini T, Vaccari F, Gioli B, Miglietta F, Cotrufo M, Peressotti A (2009) Precipitation pulses enhance respiration of Mediterranean ecosystems: the balance between organic and inorganic components of increased soil CO2 efflux. Glob Chang Biol 15:1289–1301
Jaspers E, Overmann J (2004) Ecological significance of microdiversity: identical 16S rRNA gene sequences can be found in bacteria with highly divergent genomes and ecophysiologies. Appl Environ Microbiol 70:4831–4839
Jenkinson DS, Ladd JN (1981) Microbial biomass in soil: measurement and turnover. In: Paul EA, Ladd JN (eds) Soil biochemistry. Academic Press, Dekker, New York, pp 415–472
Jenkinson DS, Hart PBS, Rayner JH, Parry LC (1987) Modelling the turnover of organic matter in long-term experiments at Rothamstedt. INTECOL Bulletin 15:1–8
Jiang X, Hou X, Zhou X, Xin X, Wright A, Jia Z (2015) pH regulates key players of nitrification in paddy soils. Soil Biol Biochem 81:9–16
Johnson SL, Kuske CR, Carney TD, Housman DC, Gallegos-Graves LV, Belnap J (2012) Increased temperature and altered summer precipitation have differential effects on biological soil crusts in a dryland ecosystem. Glob Chang Biol 18:2583–2593
Jonasson S, Michelsen A, Schmidt IK, Nielsen EV, Callaghan TV (1996) Microbial biomass C, N and P in two arctic soils and responses to addition of NPK fertilizer and sugar: implications for plant nutrient uptake. Oecologia 106:507–515
Jones CM, Graf DRH, Bru D, Philippot L, Hallin S (2012) The unaccounted yet abundant nitrous oxide-reducing microbial community: a potential nitrous oxide sink. ISME J 7:417–426
Kaiser C, Franklin O, Dieckmann U, Richter A (2014) Microbial community dynamics alleviate stoichiometric constraints during litter decay. Ecol Lett 17:680–690
Kaiser C, Franklin O, Richter A, Dieckmann U (2015) Social dynamics within decomposer communities lead to nitrogen retention and organic matter build-up in soils. Nat Commun 6:8960
Keiblinger KM, Hall EK, Wanek W, Szukics U, Hämmerle I, Ellersdorfer G, Böck S, Strauss J, Sterflinger K, Richter A, Zechmeister-Boltenstern S (2010) The effect of resource quantity and resource stoichiometry on microbial carbon-use-efficiency. FEMS Microbiol Ecol 73:430–440
Keller JK, Weisenhorn PB, Megonigal JP (2009) Humic acids as electron acceptors in wetland decomposition. Soil Biol Biochem 41:1518–1522
Kreft JU, Bonhoeffer S (2005) The evolution of groups of cooperating bacteria and the growth rate versus yield trade-off. Microbiology 151:637–641
Kuhn E (2012) Toward understanding life under subzero conditions: the significance of exploring psychrophilic “cold-shock” proteins. Astrobiology 12:1078–1086
Kuzyakov Y, Blagodatskaya E (2015) Microbial hotspots and hot moments in soil: concept & review. Soil Biol Biochem 83:184–199
Kuzyakov Y, Xu X (2013) Competition between roots and microorganisms for nitrogen: mechanisms and ecological relevance. New Phytol 198:656–669
Langille MGI, Zaneveld J, Caporaso JG, McDonald D, Knights D, Reyes JA, Clemente JC, Burkepile DE, Vega Thurber RL, Knight R, Beiko RG, Huttenhower C (2013) Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat Biotechnol 31:814–821
Lankau RA, Zhu K, Ordonez A (2015) Mycorrhizal strategies of tree species correlate with trailing range edge responses to current and past climate change. Ecology 96:1451–1458
Lawrence CR, Neff JC, Schimel JP (2009) Does adding microbial mechanisms of decomposition improve soil organic matter models? A comparison of four models using data from a pulsed rewetting experiment. Soil Biol Biochem 41:1923–1934
Le Mer J, Roger P (2001) Production, oxidation, emission and consumption of methane by soils: a review. Eur J Soil Biol 37:25–50
Legay N, Baxendale C, Grigulis K, Krainer U, Kastl E, Schloter M, Bardgett RD, Arnoldi C, Bahn M, Dumont M, Poly F, Pommier T, Clément JC, Lavorel S (2014) Contribution of above- and below-ground plant traits to the structure and function of grassland soil microbial communities. Ann Bot 114:1011–1021
Liao C, Peng R, Luo Y, Zhou X, Wu X, Fang C, Chen J, Li B (2008) Altered ecosystem carbon and nitrogen cycles by plant invasion: a meta-analysis. New Phytol 177:706–714
Lipson DA (2007) Relationships between temperature responses and bacterial community structure along seasonal and altitudinal gradients. FEMS Microbiol Ecol 59:418–427
Lipson DA (2015) The complex relationship between microbial growth rate and yield and its implications for ecosystem processes. Front Microbiol 6:615
Lipson DA, Kelley ST (2014) Plant-microbe interactions. In: Monson RK (ed) Ecology and the environment. Springer, New York, pp 177–204
Lipson DA, Schmidt SK, Monson RK (1999) Links between microbial population dynamics and N availability in an alpine ecosystem. Ecology 80:1623–1631
Lipson DA, Wilson RF, Oechel WC (2005) Effects of elevated atmospheric CO2 on soil microbial biomass, activity and diversity in a chaparral ecosystem. Appl Environ Microbiol 71:8573–8580
Lipson DA, Monson RK, Schmidt SK, Weintraub MN (2009) The trade-off between growth rate and yield in microbial communities and the consequences for under-snow soil respiration in a high elevation coniferous forest. Biogeochemistry 95:23–35
Lipson DA, Kuske CR, Gallegos-Graves LV, Oechel WC (2014) Elevated atmospheric CO2 stimulates soil fungal diversity through increased fine root production in a semiarid shrubland ecosystem. Glob Chang Biol 20:2555–2565
Long A, Heitman J, Tobias C, Philips R, Song B (2012) Co-occurring anammox, denitrification, and codenitrification in agricultural soils. Appl Environ Microbiol 79:168–176
Lovley DR, Phillips EJP (1987) Competitive mechanisms for inhibition of sulfate reduction and methane production in the zone of ferric iron reduction in sediments. Appl Environ Microbiol 53:2636–2641
Luo YQ, Wan SQ, Hui DF, Wallace LL (2001) Acclimatization of soil respiration to warming in a tall grass prairie. Nature 413:622–625
Luo Y, Ahlström A, Allison SD, Batjes NH, Brovkin V, Carvalhais N, Chappell A, Ciais P, Davidson EA, Finzi A, Georgiou K, Guenet B, Hararuk O, Harden JW, He Y, Hopkins FM, Jiang L, Koven CD, Jackson RB, Jones CD, Lara MJ, Liang J, McGuire AD, Parton W, Peng C, Randerson JT, Salazar A, sierra CA, Smith M, Tian H, Todd-Brown KE, Torn M, van Groenendael J, Wang Y-P, West TO, Wei Y, Wieder WR, Xia J, Xu X, Xu X, Zhou T (2016) Towards more realistic projections of soil carbon dynamics by earth system models. Glob Biogeochem Cycles 30:40–56. https://doi.org/10.1002/2015GB005239
Lynch MD, Neufeld JD (2015) Ecology and exploration of the rare biosphere. Nat Rev Microbiol 13:217–229
Maida I, Bosi E, Perrin E, Papaleo MC, Orlandini V, Fondi M, Fani R, Wiegel J, Bianconi G, Canganella F (2013) Draft genome sequence of the fast-growing bacterium Vibrio natriegens strain DSMZ 759. Genome Announc 1:e00648–e00613
Manzoni S, Porporato A (2009) Soil carbon and nitrogen mineralization: theory and models across scales. Soil Biol Biochem 41(7):1355–1379
Manzoni S, Taylor P, Richter A, Porporato A, Ågren GI (2012) Environmental and stoichiometric controls on microbial carbon-use efficiency in soils. New Phytol 196:79–91
Manzoni S, Schaeffe SM, Katul G, Porporato A, Schimel J (2014) A theoretical analysis of microbial eco-physiological and diffusion limitations to carbon cycling in drying soils. Soil Biol Biochem 73:69–83
Margesin R, Miteva V (2011) Diversity and ecology of psychrophilic microorganisms. Res Microbiol 162:346–361
McCalley CK, Woodcroft BJ, Hodgkins SB, Wehr RA, Kim E-H, Mondav R, Crill PM, Chanton JP, Rich VI, Tyson GW (2014) Methane dynamics regulated by microbial community response to permafrost thaw. Nature 514:478–481
McCann KS (2000) The diversity–stability debate. Nature 405:228–233
Melton J, Wania R, Hodson E, Poulter B, Ringeval B, Spahni R, Bohn T, Avis C, Beerling D, Chen G (2013) Present state of global wetland extent and wetland methane modelling: conclusions from a model intercomparison project (WETCHIMP). Biogeosciences 10:753–788
Mikan CJ, Schimel JP, Doyle AP (2002) Temperature controls of microbial respiration in arctic tundra soils above and below freezing. Soil Biol Biochem 34:1785–1795
Miller KE, Lai C-T, Friedman ES, Angenent LT, Lipson DA (2015) Methane suppression by iron and humic acids in soils of the arctic coastal plain. Soil Biol Biochem 83:176–183
Monson RK, Lipson DA, Burns SP, Turnipseed AA, Delany AC, Williams MW, Schmidt SK (2006) Forest soil respiration controlled by winter climate variation and microbial community composition. Nature 439:711–714
Moorhead DL, Sinsabaugh RL (2006) A theoretical model of litter decay and microbial interaction. Ecol Monogr 76:151–174
Moorhead DL, Lashermes G, Sinsabaugh RL (2012) A theoretical model of C- and N-acquiring exoenzyme activities, which balances microbial demands during decomposition. Soil Biol Biochem 53:133–141
Moorhead D, Lashermes G, Recous S, Bertrand I (2014) Interacting microbe and litter quality controls on litter decomposition: a modeling analysis. PloS One 9:e108769
Morel F, Price N (2003) The biogeochemical cycles of trace metals in the oceans. Science 300:944–947
Morita RY (1975) Psychrophilic bacteria. Bacteriol Rev 39:144
Mothapo N, Chen H, Cubeta MA, Grossman JM, Fuller F, Shi W (2015) Phylogenetic, taxonomic and functional diversity of fungal denitrifiers and associated N2O production efficacy. Soil Biol Biochem 83:160–175
Myhre G, Shindell D, Bréon F, Collins W, Fuglestvedt J, Huang J, Koch D, Lamarque J, Lee D, Mendoza B (2013) Anthropogenic and natural radiative forcing. In: Climate change 2013: the physical science basis. Contribution of Working Group 1 to the Fifth Assessment Report of the Intergovernmental Panel on Climate Change. Table 8, p 714
Nam H, Lewis NE, Lerman JA, Lee D-H, Chang RL, Kim D, Palsson BO (2012) Network context and selection in the evolution to enzyme specificity. Science 337:1101–1104
Nazaries L, Murrell JC, Millard P, Baggs L, Singh BK (2013) Methane, microbes and models: fundamental understanding of the soil methane cycle for future predictions. Environ Microbiol 15:2395–2417
Nemergut DR, Shade A, Violle C (2014) When, where and how does microbial community composition matter? Front Microbiol 5:424
Neuenschwander SM, Pernthaler J, Posch T, Salcher MM (2015) Seasonal growth potential of rare lake water bacteria suggest their disproportional contribution to carbon fluxes. Environ Microbiol 17:781–795
Nielsen U, Ayres E, Wall D, Bardgett R (2011) Soil biodiversity and carbon cycling: a review and synthesis of studies examining diversity–function relationships. Eur J Soil Sci 62:105–116
Nielsen UN, Wall DH, Six J (2015) Soil biodiversity and the environment. Annu Rev Environ Resour 40:63–90
Osanai Y, Janes JK, Newton PCD, Hovenden MJ (2015) Warming and elevated CO2 combine to increase microbial mineralisation of soil organic matter. Soil Biol Biochem 85:110–118
Pan Y, Ni B-J, Bond PL, Ye L, Yuan Z (2013) Electron competition among nitrogen oxides reduction during methanol-utilizing denitrification in wastewater treatment. Water Res 47:3273–3281
Panikov N (2009) Microbial activity in frozen soils. In: Margesin R (ed) Permafrost soils. Springer, Berlin, pp 119–147
Panikov NS, Flanagan PW, Oechel WC, Mastepanov MA, Christensen TR (2006) Microbial activity in soils frozen to below −39°C. Soil Biol Biochem 38:785–794
Parton WJ, Schimel DS, Cole CV, Ojima DS (1987) Analysis of factors controlling soil organic matter levels in Great Plains grasslands. Soil Sci Soc Am J 51:1173–1179
Peay KG, Garbelotto M, Bruns TD (2010) Evidence of dispersal limitation in soil microorganisms: isolation reduces species richness on mycorrhizal tree islands. Ecology 91:3631–3640
Pelini SL, Maran AM, Chen AR, Kaseman J, Crowther TW (2015) Higher trophic levels overwhelm climate change impacts on terrestrial ecosystem functioning. PLoS One 10:e0136344
Philippot L, Andersson SGE, Battin TJ, Prosser JI, Schimel JP, Whitman WB, Hallin S (2010) The ecological coherence of high bacterial taxonomic ranks. Nat Rev Microbiol 8:523–529
Pietikäinen J, Pettersson M, Baath E (2005) Comparison of temperature effects on soil respiration and bacterial and fungal growth rates. FEMS Microbiol Ecol 52:49–58
Piette F, D’Amico S, Struvay C, Mazzucchelli G, Renaut J, Tutino ML, Danchin A, Leprince P, Feller G (2010) Proteomics of life at low temperatures: trigger factor is the primary chaperone in the Antarctic bacterium Pseudoalteromonas haloplanktis TAC125. Mol Microbiol 76:120–132
Placella SA, Brodie EL, Firestone MK (2012) Rainfall-induced carbon dioxide pulses result from sequential resuscitation of phylogenetically clustered microbial groups. Proc Natl Acad Sci USA 109:10931–10936
Pol A, Heijmans K, Harhangi HR, Tedesco D, Jetten MSM, Op den Camp HJM (2007) Methanotrophy below pH1 by a new Verrucomicrobia species. Nature 450:874–878
Rappe MS, Giovannoni SJ (2003) The uncultured microbial majority. Annu Rev Microbiol 57:369–394
Ratkowsky D, Olley J, McMeekin T, Ball A (1982) Relationship between temperature and growth rate of bacterial cultures. J Bacteriol 149:1–5
Ratkowsky D, Lowry R, McMeekin T, Stokes A, Chandler R (1983) Model for bacterial culture growth rate throughout the entire biokinetic temperature range. J Bacteriol 154:1222–1226
Redfield AC (1958) The biological control of chemical factors in the environment. Am Sci 46:205–221
Robador A, Müller AL, Sawicka JE, Berry D, Hubert CRJ, Loy A, Jørgensen BB, Brüchert V (2015) Activity and community structures of sulfate-reducing microorganisms in polar, temperate and tropical marine sediments. ISME J 10:796–809
Roco CA, Bergaust LL, Bakken LR, Yavitt JB, Shapleigh JP (2016) Modularity of nitrogen-oxide reducing soil bacteria: linking phenotype to genotype. Environ Microbiol 19(6):2507–2519
Rodrigues DF, da C Jesus E, Ayala-del-Río HL, Pellizari VH, Gilichinsky D, Sepulveda-Torres L, Tiedje JM (2009) Biogeography of two cold-adapted genera: psychrobacter and Exiguobacterium. ISME J 3:658–665
Rotaru A-E, Shrestha PM, Liu F, Markovaite B, Chen S, Nevin K, Lovley D (2014) Direct interspecies electron transfer between Geobacter metallireducens and Methanosarcina barkeri. Appl Environ Microbiol 80(15):4599–4605
Rousk J, Bååth E (2011) Growth of saprotrophic fungi and bacteria in soil. FEMS Microbiol Ecol 78:17–30
Rousk J, Frey SD, Bååth E (2012) Temperature adaptation of bacterial communities in experimentally warmed forest soils. Glob Chang Biol 18:3252–3258
Sakamoto K, Oba Y (1994) Effect of fungal to bacterial biomass ratio on the relationship between CO2 evolution and total soil microbial biomass. Biol Fertil Soils 17:39–44
Salvadó Z, Arroyo-López F, Guillamón JM, Salazar G, Querol A, Barrio E (2011) Temperature adaptation markedly determines evolution within the genus Saccharomyces. Appl Environ Microbiol 77:2292–2302
Sanford RA, Wagner DD, Wu Q, Chee-Sanford JC, Thomas SH, Cruz-Garcia C, Rodriguez G, Massol-Deya A, Krishnani KK, Ritalahti KM, Nissen S, Konstantinidis KT, Loffler FE (2012) Unexpected nondenitrifier nitrous oxide reductase gene diversity and abundance in soils. Proc Natl Acad Sci USA 109:19709–19714
Schaefer K, Lantuit H, Romanovsky VE, Schuur EAG, Witt R (2014) The impact of the permafrost carbon feedback on global climate. Environ Res Lett 9:085003
Schimel J (1995) Ecosystem consequences of microbial diversity and community structure. In: Arctic and alpine biodiversity: patterns, causes and ecosystem consequences. Springer, Berlin, pp 239–254
Schimel JP (2001) Biogeochemical models: implicit vs. explicit microbiology. In: Schulze E-D (ed) Global biogeochemical cycles in the climate systems. Academic Press, New York, pp 177–183
Schimel JP, Gulledge J (1998) Microbial community structure and global trace gases. Glob Chang Biol 4:745–758
Schimel JP, Schaeffer SM (2012) Microbial control over carbon cycling in soil. Front Microbiol 3:348
Schimel JP, Weintraub MN (2003) The implications of exoenzyme activity on microbial carbon and nitrogen limitation in soil: a theoretical model. Soil Biol Biochem 35:549–563
Schimel J, Balser TC, Wallenstein M (2007) Microbial stress-response physiology and its implications for ecosystem function. Ecology 88:1386–1394
Schmidt SK, Wilson KL, Monson RK, Lipson DA (2009) Exponential growth of “snow molds” at sub-zero temperatures: an explanation for high beneath-snow respiration rates and Q 10 values. Biogeochemistry 95:13–21
Schuur EAG, McGuire AD, Schädel C, Grosse G, Harden JW, Hayes DJ, Hugelius G, Koven CD, Kuhry P, Lawrence DM, Natali SM, Olefeldt D, Romanovsky VE, Schaefer K, Turetsky MR, Treat CC, Vonk JE (2015) Climate change and the permafrost carbon feedback. Nature 520:171–179
Scott-Denton LE, Rosenstiel T, Monson RK (2006) Differential controls by climate and substrate over the heterotrophic and rhizospheric components of soil respiration. Glob Chang Biol 12:205–216
Sigüenza C, Corkidi L, Allen EB (2006) Feedbacks of soil inoculum of mycorrhizal fungi altered by N deposition on the growth of a native shrub and an invasive annual grass. Plant Soil 286:153–165
Sinsabaugh RL (2010) Phenol oxidase, peroxidase and organic matter dynamics of soil. Soil Biol Biochem 42:391–404
Sinsabaugh RL, Manzoni S, Moorhead DL, Richter A (2013) Carbon use efficiency of microbial communities: stoichiometry, methodology and modelling. Ecol Lett 16:930–939
Sinsabaugh RL, Shah JJF, Findlay SG, Kuehn KA, Moorhead DL (2014) Scaling microbial biomass, metabolism and resource supply. Biogeochemistry 122:175–190
Sinsabaugh RL, Turner BL, Talbot JM, Waring BG, Powers JS, Kuske CR, Moorhead DL, Follstad Shah JJ (2016) Stoichiometry of microbial carbon use efficiency in soils. Ecol Monogr 86(2):172–189
Sistla SA, Asao S, Schimel JP (2012) Detecting microbial N-limitation in tussock tundra soil: implications for Arctic soil organic carbon cycling. Soil Biol Biochem 55:78–84
Sistla SA, Rastetter EB, Schimel JP (2014) Responses of a tundra system to warming using SCAMPS: a stoichiometrically coupled, acclimating microbe–plant–soil model. Ecol Monogr 84(1):151–170
Six J, Frey SD, Thiet RK, Batten KM (2006) Bacterial and fungal contributions to carbon sequestration in agroecosystems. Soil Sci Soc Am J 70:555–569
Smemo K, Yavitt J (2011) Anaerobic oxidation of methane: an underappreciated aspect of methane cycling in peatland ecosystems? Biogeosciences 8:779–793
Spencer SJ, Tamminen MV, Preheim SP, Guo MT, Briggs AW, Brito IL, Weitz DA, Pitkänen LK, Vigneault F, Virta MP (2015) Massively parallel sequencing of single cells by epicPCR links functional genes with phylogenetic markers. ISME J 0(2):427–436
Spohn M, Chodak M (2015) Microbial respiration per unit biomass increases with carbon-to-nutrient ratios in forest soils. Soil Biol Biochem 81:128–133
Staley JT, Castenholz RW, Colwell RR, Holt JG, Kane MD, Pace NR, Salyers AA, Tiedje JMT (1997) The microbial world. American Society for Microbiology, Washington, DC
Stein LY, Klotz MG (2016) The nitrogen cycle. Curr Biol 26:R94–R98
Steinauer K, Tilman D, Wragg PD, Cesarz S, Cowles JM, Pritsch K, Reich PB, Weisser WW, Eisenhauer N (2015) Plant diversity effects on soil microbial functions and enzymes are stronger than warming in a grassland experiment. Ecology 96:99–112
Steinweg JM, Plante AF, Conant RT, Paul EA, Tanaka DL (2008) Patterns of substrate utilization during long-term incubations at different temperatures. Soil Biol Biochem 40:2722–2728
Steven B, Leveille R, Pollard WH, Whyte LG (2006) Microbial ecology and biodiversity in permafrost. Extremophiles 10:259–267
Stewart EJ (2012) Growing unculturable bacteria. J Bacteriol 194:4151–4160
Stieglmeier M, Mooshammer M, Kitzler B, Wanek W, Zechmeister-Boltenstern S, Richter A, Schleper C (2014) Aerobic nitrous oxide production through N-nitrosating hybrid formation in ammonia-oxidizing archaea. ISME J 8:1135–1146
Sulman BN, Phillips RP, Oishi AC, Shevliakova E, Pacala SW (2014) Microbe-driven turnover offsets mineral-mediated storage of soil carbon under elevated CO2. Nat Clim Chang 4:1099–1102
Talbot JM, Allison SD, Treseder KK (2008) Decomposers in disguise: mycorrhizal fungi as regulators of soil C dynamics in ecosystems under global change. Funct Ecol 22:955–963
Tang J, Riley WJ (2015) Weaker soil carbon-climate feedbacks resulting from microbial and abiotic interactions. Nat Clim Chang 5:56–60
Thiet RK, Frey SD, Six J (2006) Do growth yield efficiencies differ between soil microbial communities differing in fungal: bacterial ratios? Reality check and methodological issues. Soil Biol Biochem 38:837–844
Todd-Brown KEO, Hopkins FM, Kivlin SN, Talbot JM, Allison SD (2011) A framework for representing microbial decomposition in coupled climate models. Biogeochemistry 109:19–33
Travisano M, Velicer GJ (2004) Strategies of microbial cheater control. Trends Microbiol 12:72–78
Treseder KK, Balser TC, Bradford MA, Brodie EL, Dubinsky EA, Eviner VT, Hofmockel KS, Lennon JT, Levine UY, MacGregor BJ (2012) Integrating microbial ecology into ecosystem models: challenges and priorities. Biogeochemistry 109:7–18
Tucker CL, Bell J, Pendall E, Ogle K (2013) Does declining carbon-use efficiency explain thermal acclimation of soil respiration with warming? Glob Chang Biol 19:252–263
Tveit AT, Urich T, Frenzel P, Svenning MM (2015) Metabolic and trophic interactions modulate methane production by Arctic peat microbiota in response to warming. Proc Natl Acad Sci USA 112:E2507–E2516
Van Der Heijden MGA, Bardgett RD, Van Straalen NM (2008) The unseen majority: soil microbes as drivers of plant diversity and productivity in terrestrial ecosystems. Ecol Lett 11:296–310
van der Putten WH, Bradford MA, Brinkman EP, van de Voorde TF, Veen G (2016) Where, when and how plant-soil feedback matters in a changing world. Funct Ecol 30:1109–1121
van Elsas JD, Chiurazzi M, Mallon CA, Elhottovā D, Krištůfek V, Salles JF (2012) Microbial diversity determines the invasion of soil by a bacterial pathogen. Proc Natl Acad Sci USA 109:1159–1164
Van Veen J, Ladd J, Frissel M (1984) Modelling C and N turnover through the microbial biomass in soil. Plant Soil 76:257–274
Veen JV, Paul E (1981) Organic carbon dynamics in grassland soils. 1. Background information and computer simulation. Can J Soil Sci 61:185–201
von Fischer JC, Rhew RC, Ames GM, Fosdick BK, von Fischer PE (2010) Vegetation height and other controls of spatial variability in methane emissions from the Arctic coastal tundra at Barrow, Alaska. J Geophys Res 115:G00I03. https://doi.org/10.1029/2009JG001283
Wagai R, Kishimoto-Mo AW, Yonemura S, Shirato Y, Hiradate S, Yagasaki Y (2013) Linking temperature sensitivity of soil organic matter decomposition to its molecular structure, accessibility, and microbial physiology. Glob Chang Biol 19:1114–1125
Waldrop MP, Zak DR, Sinsabaugh RL, Gallo M, Lauber C (2004) Nitrogen deposition modifies soil carbon storage through changes in microbial enzymatic activity. Ecol Appl 14:1172–1177
Wallenstein MD, Hall EK (2012) A trait-based framework for predicting when and where microbial adaptation to climate change will affect ecosystem functioning. Biogeochemistry 109:35–47
Wang G, Post WM, Mayes MA (2013a) Development of microbial-enzyme-mediated decomposition model parameters through steady-state and dynamic analyses. Ecol Appl 23:255–272
Wang Q, Burger M, Doane T, Horwath W, Castillo A, Mitloehner F (2013b) Effects of inorganic v. organic copper on denitrification in agricultural soil. Adv Anim Biosci 4:42–49
Wang G, Jagadamma S, Mayes MA, Schadt CW, Steinweg JM, Gu L, Post WM (2015) Microbial dormancy improves development and experimental validation of ecosystem model. ISME J 9:226–237
Ward NL, Challacombe JF, Janssen PH, Henrissat B, Coutinho PM, Wu M, Xie G, Haft DH, Sait M, Badger J (2009) Three genomes from the phylum Acidobacteria provide insight into the lifestyles of these microorganisms in soils. Appl Environ Microbiol 75:2046–2056
Waring BG, Averill C, Hawkes CV (2013) Differences in fungal and bacterial physiology alter soil carbon and nitrogen cycling: insights from meta-analysis and theoretical models. Ecol Lett 16:887–894
Weber CF, Zak DR, Hungate BA, Jackson RB, Vilgalys R, Evans RD, Schadt CW, Megonigal JP, Kuske CR (2011) Responses of soil cellulolytic fungal communities to elevated atmospheric CO2 are complex and variable across five ecosystems. Environ Microbiol 13:2778–2793
Wei H, Guenet B, Vicca S, Nunan N, AbdElgawad H, Pouteau V, Shen W, Janssens IA (2014) Thermal acclimation of organic matter decomposition in an artificial forest soil is related to shifts in microbial community structure. Soil Biol Biochem 71:1–12
Wieder WR, Bonan GB, Allison SD (2013) Global soil carbon projections are improved by modelling microbial processes. Nat Clim Chang 3:909–912
Wieder W, Grandy A, Kallenbach C, Bonan G (2014) Integrating microbial physiology and physio-chemical principles in soils with the MIcrobial-MIneral Carbon Stabilization (MIMICS) model. Biogeosciences 11:3899–3917
Wieder WR, Allison SD, Davidson EA, Georgiou K, Hararuk O, He Y, Hopkins F, Luo Y, Smith MJ, Sulman B (2015) Explicitly representing soil microbial processes in Earth system models. Glob Biogeochem Cycles 29:1782–1800
Xu X, Thornton PE, Post WM (2013) A global analysis of soil microbial biomass carbon, nitrogen and phosphorus in terrestrial ecosystems. Glob Ecol Biogeogr 22:737–749
Xu X, Schimel JP, Thornton PE, Song X, Yuan F, Goswami S (2014) Substrate and environmental controls on microbial assimilation of soil organic carbon: a framework for Earth system models. Ecol Lett 17:547–555
Xu X, Elias DA, Graham DE, Phelps TJ, Carrol SL, Wullschleger SD, Thornton PE (2015) A microbial functional group based module for simulating methane production and consumption: application to an incubation permafrost soil. J Geophys Res Biogeosci 120:1315–1333
Xu X, Schimel JP, Janssens IA, Song X, Song C, Yu G, Sinsabaugh RL, Tang D, Zhang X, Thornton PE (2017) Global pattern and controls of soil microbial metabolic quotient. Ecol Monogr 87(3):429–441
Yarza P, Yilmaz P, Pruesse E, Glöckner FO, Ludwig W, Schleifer K-H, Whitman WB, Euzéby J, Amann R, Rosselló-Móra R (2014) Uniting the classification of cultured and uncultured bacteria and archaea using 16S rRNA gene sequences. Nat Rev Microbiol 12:635–645
Youssef NH, Couger M, McCully AL, Criado AEG, Elshahed MS (2015) Assessing the global phylum level diversity within the bacterial domain: a review. J Adv Res 6:269–282
Zechmeister-Boltenstern S, Keiblinger KM, Mooshammer M, Peñuelas J, Richter A, Sardans J, Wanek W (2015) The application of ecological stoichiometry to plant–microbial–soil organic matter transformations. Ecol Monogr 85:133–155
Zona D, Gioli B, Commane R, Lindaas J, Wofsy SC, Miller CE, Dinardo SJ, Dengel S, Sweeney C, Karion A (2016) Cold season emissions dominate the Arctic tundra methane budget. Proc Natl Acad Sci USA 113:40–45
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Ethics declarations
Conflict of Interest
David A. Lipson declares that he has no conflict of interest. Xiaofeng Xu declares that he has no conflict of interest.
Ethical Approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Rights and permissions
Copyright information
© 2019 Springer Nature Switzerland AG
About this chapter
Cite this chapter
Lipson, D.A., Xu, X. (2019). Integrating Soil Microbiology into Ecosystem Science. In: Hurst, C. (eds) Understanding Terrestrial Microbial Communities. Advances in Environmental Microbiology, vol 6. Springer, Cham. https://doi.org/10.1007/978-3-030-10777-2_3
Download citation
DOI: https://doi.org/10.1007/978-3-030-10777-2_3
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-10775-8
Online ISBN: 978-3-030-10777-2
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)