Abstract
mTOR is a key regulator of a number of critical cellular processes including growth, proliferation, cytoskeletal organization, and differentiation. mTOR mediates its effects on these processes by regulating mRNA translation initiation via phosphorylation of its major downstream targets: the 4E binding proteins (4E-BPs) and the ribosomal protein S6 kinases. Dysregulation of mTOR signalling leads to increased cellular growth and proliferation and is implicated in a number of human cancers. In particular, increased mTOR signalling is associated with human cancers that are characterized by loss or mutations in tumour suppressors such as LKB1, PTEN, and TSC1/2, which are responsible for suppressing the PI3K/AKT pathway. The regulation of mRNA translation by mTOR will be the focus of this chapter. In particular, the role of the translational machinery downstream of mTOR in oncogenesis will be discussed.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
The PI3K/AKT/mTOR signalling pathway regulates a number of critical cellular processes including growth, proliferation, differentiation, and survival. The serine/threonine kinase mTOR integrates a wide range of extra- and intra-cellular signals to control these processes through the regulation of mRNA translation. mTOR mediates its effects on mRNA translation through phosphorylation of its major downstream targets: the 4E binding proteins (4E-BPs) and the ribosomal protein S6 kinases (S6K1 and S6K2). The 4E-BPs suppress translation initiation and are inhibited via mTOR-mediated phosphorylation, while the S6Ks enhance translation upon activation by mTOR by targeting components of the translational machinery. Inappropriate activation of mTOR signalling can promote tumorigenesis through an increase in the translation of mRNAs encoding growth factors, pro-survival proteins, cell cycle regulators, and angiogenic factors. Consequently, dysregulation of the mTOR pathway is implicated in a number of human cancers. The role of mTOR and its downstream targets in mRNA translation initiation will be the focus of this chapter. In particular, the regulation of the translational machinery via mTOR-mediated phosphorylation will be described and the importance of the downstream targets of mTOR in the development of human cancer will be addressed.
2 Translation Initiation
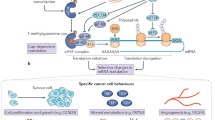
The process of translation is divided into three stages: initiation, elongation, and termination. Initiation is rate limiting under most circumstances and as a result is a primary target of translational control. Translation initiation is a complex, highly ordered process that culminates in the assembly of the 80S ribosome at the initiation codon of an mRNA. The rate-limiting step is thought to be the formation of the eIF4F (eukaryotic initiation factor 4F) complex, which mediates the recruitment of ribosomal subunits to the mRNA [1]. eIF4F is composed of eIF4E, which binds the 7-methylguanosine “cap” (m7GpppX, where X is any nucleotide and m is a methyl group) found on the 5′-end of all nuclear-transcribed cellular mRNAs, the helicase, eIF4A, and the large scaffolding protein, eIF4G (Fig. 1) [2, 3]. While eIF4E is required for recognition and binding of the 5′-cap, eIF4G bridges the ribosome and the mRNA by interacting with eIF3, which binds the 40S ribosome. eIF4A unwinds any 5′-mRNA secondary structure to facilitate binding of the 40S subunit and scanning of the mRNA 5′-untranslated region (UTR). The ATPase and helicase activities of eIF4A are stimulated by eIF4B, a small RNA-binding protein that is involved in ribosomal recruitment to mRNA [2, 4]. The 40S ribosome with its associated initiation factors scans the 5′-UTR until it encounters an initiation codon (AUG or a cognate thereof). Once the initiation codon is encountered, the 60S ribosome joins to form the active 80S ribosome.
eIF4F complex formation: The eIF4F complex mediates the recruitment of ribosomal subunits to the mRNA and is composed of the cap-binding protein eIF4E, the large scaffolding protein eIF4G, and the helicase eIF4A. eIF4G is involved in circularization of the mRNA through an interaction with PABP and bridges the 40S ribosome and the mRNA by interacting with eIF3. eIF4B is an RNA-binding protein that stimulates the ATPase and helicase activities of eIF4A
3 TOR Complex Formation
Two mTOR containing complexes exist in mammalian cells, namely mTORC1 and mTORC2. mTORC1 is rapamycin sensitive, regulated by nutrients and growth factors, and composed of mTOR, raptor (regulatory-associated protein of TOR), and mLst8 (also known as GβL) [5, 6]. Raptor functions as an adaptor protein that is responsible for recruiting the mTORC1 substrates 4E-BP1, S6K, and PRAS40 (proline-rich AKT substrate 40 kDa) [7, 8]. Raptor is required for mTOR-mediated phosphorylation of S6K and 4E-BP1 and interacts with mTOR targets through a TOR signalling (TOS) motif. The TOS motif is located in the NH2 terminus of S6K (FDIDL for S6K1 and FDLDL for S6K2) and the COOH terminus of 4E-BP1 (FEMDI) [9, 10]. Mutation of this motif greatly reduces mTOR-mediated phosphorylation of S6K and 4E-BP1 [9–12]. mLst8 interacts with the kinase domain of mTOR and stabilizes the mTOR–raptor interaction [13]. Recently, it was reported that PRAS40 associates with mTORC1 in cells and negatively regulates mTOR activity [14, 15]. The inhibitory effects of PRAS40 on mTORC1 signalling are due to competition with S6K and 4E-BP1 for binding to raptor, as PRAS40 contains a TOS motif (FVMDE) [16–18]. The activity of PRAS40 is regulated by phosphorylation such that AKT-mediated phosphorylation of PRAS40 on Thr246 reduces its inhibitory effects on mTOR signalling [14, 15]. In addition, PRAS40 was also recently identified as an mTORC1 substrate [8, 17].
In contrast, mTORC2 is composed of mTOR, mLst8, rictor (rapamycin-insensitive component of TOR), and mSIN1 (mammalian stress-activated protein kinase (SAPK)-interacting protein). mTORC2 is rapamycin insensitive, regulated by growth factors, and is implicated in cytoskeletal organization. The only substrate of mTORC2 is believed to be AKT (PKB); however, mTORC2 also regulates the phosphorylation of PKC-α (protein kinase C) and potentially other AGC-family kinases [19–21].
4 Regulation of Translation Initiation by mTOR Signalling
mTOR integrates signals from nutrients, growth factors, cellular energy stores, and other cues to control a variety of important cellular processes including growth, proliferation, differentiation, transcription, and mRNA translation [4, 22, 23]. mTOR is a serine/threonine kinase that co-ordinates these processes via phosphorylation of its downstream targets. The major targets of mTORC1 are integral components of the translational machinery, particularly those involved in translation initiation. Consequently, this chapter will focus on mTORC1 signalling, unless otherwise noted. The eIF4F components eIF4G and eIF4E are regulated by mTORC1, with the activity of eIF4E being controlled by mTORC1-mediated phosphorylation and inactivation of its repressors, the 4E-BPs. In addition, S6K1 and S6K2 are direct targets of mTORC1. S6K1 plays an important role in translational control via regulation of the ribosomal protein S6 (rpS6), eukaryotic elongation factor 2 (eEF2) kinase, and eIF4B.
4.1 The 4E-BPs
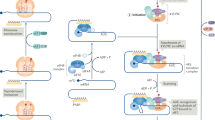
The assembly of the eIF4F complex is controlled by a family of small translational repressor proteins known as the 4E-BPs. Three 4E-BPs exist in mammals (4E-BP1, 4E-BP2, and 4E-BP3), each encoded by a separate gene, while Drosophila only express one 4E-BP [24–26]. The 4E-BPs negatively regulate eIF4F assembly by competing with eIF4G for binding to a shared site on eIF4E. The binding of the 4E-BPs to eIF4E is controlled by mTORC1-mediated phosphorylation. The majority of studies on 4E-BP regulation have focussed on 4E-BP1. Consequently, 4E-BP1 is the best characterized member of the 4E-BP family. In its hypophosphorylated form, 4E-BP1 binds to eIF4E with high affinity, preventing eIF4F complex formation and suppressing cap-dependent translation. However, upon stimulation by growth factors, nutrients, or hormones, mTORC1 phosphorylates and inhibits 4E-BP1 leading to the release of eIF4E and an increase in translation initiation (Fig. 2) [24, 27]. Seven phosphorylation sites have been identified in 4E-BP1 (Thr 37, Thr 46, Ser 65, Thr 70, Ser 83, Ser 101, and Ser 112) and four of which (Thr 37, Thr 46, Ser 65, and Thr 70) are linked to mTORC1. The importance of these individual phosphorylation sites in the regulation of 4E-BP1–eIF4E binding is not completely clear. However, the four mTORC1-specific sites are known to be involved in the release of eIF4E from 4E-BP1 [4, 27–29]. The phosphorylation of 4E-BP1 is a complex process that proceeds in a hierarchical manner (Fig. 2). Upon stimulation, mTOR phosphorylates 4E-BP1 on Thr 37 and Thr 46. The phosphorylation of these sites is believed to act as a priming event that is required for the subsequent phosphorylation of residues Thr 70 and Ser 65 [27, 28]. Phosphorylation of 4E-BP1 on residues Thr 37 and Thr 46 does not disrupt binding of eIF4E by 4E-BP1, and phosphorylation of Ser 65 and Thr 70 alone is insufficient to block eIF4E binding [27, 28]. Therefore, it is likely that in combination, these phosphorylation events co-operate to cause the dissociation of eIF4E from 4E-BP1.
Regulation of eIF4E-BP1 by phosphorylation: Hypophosphorylated 4E-BP1 binds to eIF4E with high affinity and inhibits cap-dependent translation. Upon stimulation, mTORC1 phosphorylates 4E-BP1 on Thr 37 and Thr 46, which acts as a priming event that is required for the subsequent phosphorylation of residues Ser 65 and Thr 70. The hyperphosphorylated form of 4E-BP1 is released from eIF4E, allowing formation of the active eIF4F complex and an increase in cap-dependent translation
4.1.1 S6 Kinase
The ribosomal protein S6 kinases are direct targets of mTORC1 and play important roles in the regulation of mRNA translation. Two S6 kinase proteins (S6K1 and S6K2) are expressed in mammalian cells, and both proteins are phosphorylated and activated by mTORC1 [30]. Despite being encoded by separate genes, the phosphorylation sites on S6K1 and S6K2 are conserved. S6K1, the most characterized of the two kinases, is involved in the regulation of cell growth in Drosophila and mammalian cells [31, 32]. Complete activation of S6K1 requires two phosphorylation events. Phosphorylation at Thr 389 is mediated by mTORC1 and is required for subsequent phosphorylation of Thr 229 by PDK1 (phosphoinositide-dependent kinase 1) [33–35]. S6K1 is believed to promote cell growth by increasing mRNA translation via phosphorylation of its downstream targets rpS6, eIF4B, and eEF2 kinase (Fig. 3). The best characterized substrate of S6K1 is rpS6; however, the functional significance of rpS6 phosphorylation is somewhat unclear. In the past, the phosphorylation of rpS6 correlated well with the translational activation of mRNAs that contain a 5′-terminal oligopyrimidine tract (TOP). However, studies in cells lacking S6K1 and S6K2 have demonstrated that rpS6 phosphorylation is not required for translation of 5′-TOP mRNAs [36–38]. These studies indicate that rpS6 may not be the major target through which the S6 kinases mediate their effects on mRNA translation and cell growth. Despite its unknown function, the phosphorylation of rpS6 is routinely used as an indicator of S6K and mTORC1 activity.
S6 kinase 1 targets and their role in mRNA translation. Phosphorylation of S6K1 by mTORC1 enhances its kinase activity towards its downstream targets (ribosomal protein S6; rpS6, eukaryotic initiation factor 4B; eIF4B, programmed cell death protein 4; PDCD4, insulin receptor substrate-1; IRS-1, S6K1 Aly/REF-like target; SKAR, and eukaryotic elongation factor-2 kinase; eEF2K), which are involved directly or indirectly in the regulation of mRNA translation. eIF4B, rpS6, and SKAR are activated by S6K1-mediated phosphorylation, whereas PDCD4, eEF2K, and IRS-1 are inhibited by phosphorylation
While the physiological relevance of rpS6 phosphorylation remains unclear, S6K1-mediated phosphorylation of eIF4B plays an important role in eIF4F activity and the recruitment of ribosomes to mRNA. eIF4B is an RNA-binding protein that stimulates the ATPase and helicase activities of eIF4A. S6K1 phosphorylates eIF4B on Ser 422, leading to an increase in mRNA translation [39]. This increase in translation is likely due to an enhanced interaction between phosphorylated eIF4B and eukaryotic translation initiation factor 3 (eIF3) [40, 41]. Phosphorylation may also increase the activity of eIF4B towards the helicase eIF4A, leading to more efficient translation of mRNAs containing large amounts of 5′-UTR secondary structure. For example, footprinting assays have demonstrated that eIF4B is required for the binding of ribosomes to an mRNA-containing secondary structure, and knock-down of eIF4B by RNA interference results in reduced translation of highly structured mRNAs [4, 42, 43]. The phosphorylation of eIF4B is increased by a variety of extracellular stimuli that induce cell growth including serum, insulin, and phorbol esters [4, 44]. In addition to S6K1, eIF4B is also phosphorylated on Ser 422 by the p90 ribosomal protein S6 kinase (RSK) in response to stimulation of the ERK1/2 MAPK pathway [40]. S6K1 may also indirectly affect eIF4F activity via regulation of the eIF4A inhibitor PDCD4 (programmed cell death protein 4) [19]. PDCD4 is a tumour suppressor that binds to and inhibits the helicase activity of eIF4A, thus negatively affecting the unwinding of 5′-mRNA secondary structure and translation initiation. Phosphorylation of PDCD4 by S6K1 targets it for ubiquitination and degradation, relieving its inhibitory effect on eIF4A function and protein synthesis [45].
S6K1 also plays an important role in the regulation of translation elongation via phosphorylation of eEF2 kinase. eEF2 kinase is a negative regulator of protein synthesis as it phosphorylates and inactivates eEF2, leading to inhibition of the translocation step in translation elongation [46]. Upon activation, S6K1 phosphorylates eEF2 kinase on Ser 366, which inhibits its kinase activity towards eEF2 and relieves the suppression of translation elongation [47].
An additional target of S6K1 is SKAR (S6K1 Aly/REF-like target). SKAR is phosphorylated on residues Ser 383 and 385 specifically by S6K1, not S6K2 [48]. SKAR is involved in the control of cell growth and, in co-operation with S6K1, is believed to enhance the translation of spliced mRNAs [49].
In addition to being downstream of mTORC1, S6K1 is also involved in the regulation of mTOR activity by controlling insulin signalling. Insulin or IGF (insulin-like growth factor) binds to their receptor, leading to phosphorylation and activation of the insulin receptor substrate-1 (IRS-1). Activated IRS-1 recruits and activates PI3K, which in turn activates AKT and consequently, mTOR. Upon stimulation by mTORC1, S6K1 phosphorylates IRS-1, marking it for degradation and causing a suppression of PI3K/AKT signalling [50–53]. This negative feedback loop is particularly active in cells exhibiting enhanced mTOR activity due to genetic mutations or prolonged nutrient exposure. The regulation of insulin signalling by S6K1 poses a challenge for anti-cancer therapies that target mTORC1 (such as rapamycin and its derivatives) since mTORC1 inhibition reduces S6K1 activity, which results in increased IRS-1 activation and PI3K-AKT signalling that itself plays a role in the control of cell proliferation. As a result, a great deal of research has focussed on the use of mTORC1 inhibitors in combination with inhibitors of insulin signalling for the treatment of cancer [54].
4.1.2 eIF4G
The large scaffolding protein eIF4G is an integral component of the eIF4F translation initiation complex. eIF4G interacts with the other eIF4F components, eIF4E and eIF4A, and is responsible for bridging the 40S ribosomal subunit to the mRNA through an interaction with the ribosome-associated initiation factor eIF3. eIF4G also contains binding sites for PABP (poly-A-binding protein) and the Mnk kinases (mitogen-activated protein kinase signal-integrating kinase; Mnk1, 2) [2]. The interaction between eIF4G and PABP is important for circularization of the mRNA, while the Mnks regulate the phosphorylation of eIF4E on Ser 209 [55–57]. All eukaryotes express two related eIF4G proteins, eIF4GI and eIF4GII, which are encoded by separate genes [2]. Both eIF4GI and eIF4GII are phosphorylated on multiple sites; however, the phosphorylation of eIF4GI is better characterized. eIF4GI is phosphorylated in response to extracellular stimuli that promote cell growth including serum, insulin, and growth factors [58, 59]. The phosphorylation of eIF4GI is mediated by mTORC1 as a number of rapamycin-sensitive phosphorylation sites including Ser 1108, Ser 1148, Ser 1192 have been identified [58]. However, the functional significance of eIF4G phosphorylation remains unclear. Phosphorylation does not affect the activity of eIF4G or its ability to associate with other initiation factors, but may change its structural conformation and alter the translation of specific mRNAs.
4.1.3 Other Targets of mTORC1
No other direct targets of mTORC1 have been identified; however, a number of additional cellular proteins contain putative TOS motifs, indicating that other mTORC1 substrates may exist. For example, HIF1-α, STAT3, and some PKC isoforms contain potential TOS motifs that may target them for raptor recruitment and mTORC1-mediated phosphorylation [60]. In fact, some reports indicate that STAT3 phosphorylation is sensitive to rapamycin treatment, supporting a role for mTORC1 in the regulation of STAT3 activity [61].
4.1.4 mTORC2 Regulation of AKT
While the mTORC1 phosphorylation targets are well characterized, very few mTORC2 substrates have been identified. The mTORC2 complex is rapamycin insensitive and involved in cytoskeletal regulation. Recently, mTORC2 was identified as the kinase responsible for phosphorylating Ser 473 in AKT. AKT is a positive regulator of mTOR signalling that phosphorylates and inhibits the TSC1/2 (tuberous sclerosis complex 1 and 2) complex, which negatively controls mTOR activity. Complete activation of AKT requires phosphorylation on Thr 308 within the activation loop and Ser 473, which lies in the hydrophobic motif [5]. PDK1 is responsible for AKT Thr 308 phosphorylation, and recent studies demonstrate that mTORC2 targets Ser 473. mTORC2 phosphorylates AKT on Ser 473 in vitro and knock-down of rictor in cells reduces Ser 473 phosphorylation, thus establishing mTORC2 as the PDK2 kinase involved in AKT regulation [20, 21]. In addition to AKT, mTORC2 may be involved in the regulation of PKC-α phosphorylation. PKC-α is involved in a number of cellular processes including growth, apoptosis, cellular structure, and motility. Knock-down of rictor reduces PKC-α phosphorylation and stability; however, this effect may not be direct [62].
5 Translation and Cancer
The translation of mRNA is a process that is important for a number of critical cellular processes including growth, survival, proliferation, and differentiation. Therefore, it is not surprising that mRNA translation is implicated in the development of cancer. Increased levels of translation are believed to promote transformation and tumorigenesis through an increase in the expression of proteins involved in growth, proliferation, and survival. Many of these proteins are encoded by a subset of mRNAs that contain long, highly structured 5′-UTRs. For example, the mRNAs encoding survivin, cyclin D1, TGF-β (transforming growth factor), CDK4 (cyclin-dependent kinase), and IGF-II (insulin-like growth factor) contain structured 5′-UTRs [83, 86]. Efficient translation of these mRNAs requires high levels of the eIF4F complex, which through the actions of its helicase eIF4A unwinds the extensive mRNA secondary structure and facilitates translation initiation. Consequently, increased expression or activation of translation initiation factors, particularly components of the eIF4F complex, can promote synthesis of oncogenic proteins and tumorigenesis. As mentioned above, a number of key translation initiation factors, as well as other important regulators of translation, are targets of the mTOR signalling pathway. Increased mTOR signalling can contribute to higher levels of translation and the development of cancer. A number of mechanisms can lead to over-activation of mTOR. Mutational loss or inactivation of tumour suppressors that negatively regulate mTOR can cause hereditary hamartomatous diseases and human cancers. For example, mutation or loss of TSC2, LKB1, and PTEN (phosphatase and tensin homologue deleted on chromosome 10) is associated with cancer-like syndromes in which patients exhibit hamartomatous polyps in multiple organs and an increased risk of cancer development [63–66]. Loss of these tumour suppressors leads to over-active mTOR signalling and an increase in the phosphorylation of S6K and 4E-BP1 [67–70]. In addition, genetic amplification of positive regulators of mTOR, such as AKT and PI3K, has been reported in breast, ovarian, and head and neck cancers [71, 72]. Regardless of the mechanism, enhanced mTOR signalling contributes to tumorigenesis via increased phosphorylation of its downstream targets and elevated mRNA translation.
6 Downstream Targets of mTOR and Their Role in Cancer
6.1 The 4E-BPs
A key mechanism by which mTORC1 signalling contributes to tumorigenesis is increased phosphorylation of the 4E-BPs. The 4E-BPs are believed to act as tumour suppressors by negatively regulating formation of the eIF4F complex that is required for mRNA translation initiation. In support of their role as tumour suppressors, over-expression of 4E-BP1 or 4E-BP2 causes a reversion of the transformed phenotype in cells transformed by eIF4E, ras, or src [73]. As mentioned above, mTORC1-mediated phosphorylation of 4E-BP1 leads to its release from eIF4E and a subsequent increase in eIF4F formation and mRNA translation. Consequently, inactivation of the 4E-BPs by phosphorylation is likely a key step in the development of cancer. For instance, ectopic expression of a non-phosphorylatable 4E-BP1 mutant, which remains capable of binding eIF4E regardless of mTOR activation, suppressed the tumorigenicity of human breast cancer cells [74]. Increased levels of 4E-BP1 phosphorylation have been observed in human breast, prostate, and ovarian tumours, supporting a role for elevated mTORC1 signalling and subsequent eIF4F complex formation in the development of these cancers [75–79]. In fact, 4E-BP1 may be useful as a prognostic indicator in human cancer as increased 4E-BP1 phosphorylation correlated with malignant progression and poor prognosis.
While the 4E-BPs are considered tumour suppressors, their binding partner, eIF4E, acts as an oncoprotein. The cap-binding protein eIF4E is the least abundant component of the eIF4F complex, and its over-expression can lead to oncogenesis. Over-expression of eIF4E transforms rodent fibroblasts and causes the malignant transformation of primary embryo fibroblasts in co-operation with the immortalizing proteins myc and E1A [80, 81]. eIF4E expression also transforms human mammary epithelial cells [74]. Consistent with these data, mice over-expressing eIF4E develop lymphomas, angiosarcomas, lung adenocarcinomas, and hepatocellular adenomas [82, 83]. In humans, elevated eIF4E levels have been reported in colon, breast, bladder, lung, and prostate tumours highlighting its importance in tumour development and progression [84–86]. In fact, eIF4E has emerged as a major target for anti-cancer therapies. Reduction of eIF4E levels by administration of anti-sense oligonucleotides reduced the growth of tumours in mice and use of these anti-sense oligonucleotides is currently under investigation in phase I clinical trials (Eli Lilly Co.) [87, 88].
6.2 S6 Kinase
The S6 kinases are important regulators of protein synthesis as well as cell growth [32]. Phosphorylation of S6K1 by mTORC1 enhances its kinase activity towards its downstream targets, rpS6, eIF4B, eEF2 kinase, PDCD4, and SKAR, leading to increased mRNA translation. The role of S6K1 in oncogenesis is not as well characterized as eIF4E or the 4E-BPs. However, the involvement of S6K1 in the regulation of cell growth strongly suggests that it may be involved in cancer. Over-expression of S6K1 causes increased cell growth and expression of a constitutively active form of S6K1 induces invasive and migratory phenotypes in ovarian cancer cells [89, 90]. Furthermore, it has been reported that S6K1 is important for the epithelial to mesenchymal transition (EMT) of ovarian cancer cells, a key process involved in tumour invasion, metastasis, and progression [91]. S6K1 is believed to promote migration and EMT in ovarian cancer cells via up-regulation of proteins involved in migration and invasion, such as matrix metalloproteinases and Snail [89, 91]. Increased S6K1 activity has been observed in breast cancer cells as well as a number of other cancer cell lines [92–95]. S6K1 is also involved in glial transformation as knock-down of S6K1 by siRNA or shRNA reduces anchorage-independent growth of human astrocytes in culture as well as the growth of intracranial tumours in mice [96]. In humans, S6K1 levels are elevated in 16% of primary breast tumours and the phosphorylation of S6K1 correlates with higher tumour grade in ovarian cancer patients [91, 93, 95, 97–99].
6.3 eIF4G
The functional significance of mTORC1-mediated eIF4G phosphorylation is somewhat unclear; however, due to its importance in eIF4F complex formation and mRNA translation initiation, eIF4G is likely implicated in the development of cancer. Over-expression of eIF4GI causes malignant transformation of rodent fibroblasts and a number of human breast cancer cells contain elevated levels of eIF4G [74, 100]. Recent work by Braunstein and colleagues has demonstrated that together with 4E-BP1, eIF4G plays an important role in causing a hypoxia-activated switch to cap-independent translation, which favours the translation of mRNAs encoding angiogenic factors, and pro-survival proteins [101]. eIF4GI levels are up-regulated in squamous cell lung carcinomas as well as advanced breast tumours [101–103]. Elevated levels of eIF4G likely contribute to tumorigenesis via increased formation of the eIF4F complex and subsequent translation of highly structured mRNAs encoding growth factors and other oncogenic proteins.
6.4 AKT Regulation by mTORC2 in Cancer
mTORC2 is the kinase responsible for phosphorylating and activating AKT. Rapamycin specifically inhibits mTORC1 by disrupting the interaction between mTOR and raptor; however, mTORC2 is not affected by rapamycin. Consequently, mTORC2-AKT signalling represents a significant challenge to the treatment of cancers with rapamycin and its derivatives since AKT is involved in the regulation of cell growth and survival. However, some research indicates that mTORC2 signalling is inhibited by prolonged treatment with rapamycin [104]. Further research is required to elucidate the role of mTORC2 in tumorigenesis, and the development of mTORC2 inhibitors as anti-cancer therapies represents an area of research that requires further examination.
7 Conclusions
Due to its importance in the regulation of cellular growth and proliferation, mTOR has emerged as a key factor in the process of tumorigenesis. Over-active mTOR signalling due to mutation or loss of its negative regulators is implicated in a number of hereditary diseases and human cancers. Consequently, a great deal of research has focussed on the regulation of mTOR and its downstream targets. The majority of mTOR targets are components of the translational machinery, highlighting its importance in the regulation of mRNA translation. The major targets of mTORC1, the 4E-BPs and the S6Ks, have been extensively characterized and their roles in protein synthesis are well described. However, their involvement in the development of cancer is an area of research that requires further attention. In addition, a great deal of work will be required to identify additional substrates of mTORC1 and to elucidate the potential role of the mTORC2 complex in transformation and cancer. Future studies focussing on the function of mTOR and its downstream targets in cancer will not only aid in the understanding of the biochemical regulation and function of mTOR but also provide an opportunity for the identification and development of novel anti-cancer therapies.
References
Pestova TV, Lorsch JR, Hellen CUT (2007) The mechanism of translation initiation in eukaryotes. In: Mathews MB, Sonenberg N, Hershey JWB (eds) Translational control in biology and medicine. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY, pp 87–128
Gingras AC, Raught B, Sonenberg N (1999) eIF4 initiation factors: effectors of mRNA recruitment to ribosomes and regulators of translation. Annu Rev Biochem 68:913–963
Holcik M, Sonenberg N (2005) Translational control in stress and apoptosis. Nat Rev Mol Cell Biol 6:318–327
Hay N, Sonenberg N (2004) Upstream and downstream of mTOR. Genes Dev 18:1926–1945
Guertin DA, Sabatini DM (2005) An expanding role for mTOR in cancer. Trends Mol Med 11:353–361
Sabatini DM (2006) mTOR and cancer: insights into a complex relationship. Nat Rev Cancer 6:729–734
Hara K, Maruki Y, Long X et al (2002) Raptor, a binding partner of target of rapamycin (TOR), mediates TOR action. Cell 110:177–189
Wang L, Harris TE, Lawrence JC Jr (2008) Regulation of proline-rich Akt substrate of 40 kDa (PRAS40) function by mammalian target of rapamycin complex 1 (mTORC1)-mediated phosphorylation. J Biol Chem 283:15619–15627
Schalm SS, Blenis J (2002) Identification of a conserved motif required for mTOR signaling. Curr Biol 12:632–639
Schalm SS, Fingar DC, Sabatini DM, Blenis J (2003) TOS motif-mediated raptor binding regulates 4E-BP1 multisite phosphorylation and function. Curr Biol 13:797–806
Nojima H, Tokunaga C, Eguchi S et al (2003) The mammalian target of rapamycin (mTOR) partner, raptor, binds the mTOR substrates p70 S6 kinase and 4E-BP1 through their TOR signaling (TOS) motif. J Biol Chem 278:15461–15464
Choi KM, McMahon LP, Lawrence JC Jr (2003) Two motifs in the translational repressor PHAS-I required for efficient phosphorylation by mammalian target of rapamycin and for recognition by raptor. J Biol Chem 278:19667–19673
Kim DH, Sarbassov DD, Ali SM et al (2003) GbetaL, a positive regulator of the rapamycin-sensitive pathway required for the nutrient-sensitive interaction between raptor and mTOR. Mol Cell 11:895–904
Sancak Y, Thoreen CC, Peterson TR et al (2007) PRAS40 is an insulin-regulated inhibitor of the mTORC1 protein kinase. Mol Cell 25:903–915
Vander Haar E, Lee SI, Bandhakavi S, Griffin TJ, Kim DH (2007) Insulin signalling to mTOR mediated by the Akt/PKB substrate PRAS40. Nat Cell Biol 9:316–323
Wang L, Harris TE, Roth RA, Lawrence JC Jr (2007) PRAS40 regulates mTORC1 kinase activity by functioning as a direct inhibitor of substrate binding. J Biol Chem 282: 20036–20044
Oshiro N, Takahashi R, Yoshino K et al (2007) The proline-rich Akt substrate of 40 kDa (PRAS40) is a physiological substrate of mammalian target of rapamycin complex 1. J Biol Chem 282:20329–20339
Fonseca BD, Smith EM, Lee VH, MacKintosh C, Proud CG (2007) PRAS40 is a target for mammalian target of rapamycin complex 1 and is required for signaling downstream of this complex. J Biol Chem 282:24514–24524
Guertin DA, Sabatini DM (2007) Defining the role of mTOR in cancer. Cancer Cell 12:9–22
Sarbassov DD, Guertin DA, Ali SM, Sabatini DM (2005) Phosphorylation and regulation of Akt/PKB by the rictor-mTOR complex. Science 307:1098–1101
Bayascas JR, Alessi DR (2005) Regulation of Akt/PKB Ser473 phosphorylation. Mol Cell 18:143–145
Wullschleger S, Loewith R, Hall MN (2006) TOR signaling in growth and metabolism. Cell 124:471–484
Yang Q, Guan KL (2007) Expanding mTOR signaling. Cell Res 17:666–681
Pause A, Belsham GJ, Gingras AC et al (1994) Insulin-dependent stimulation of protein synthesis by phosphorylation of a regulator of 5′-cap function. Nature 371:762–767
Poulin F, Gingras AC, Olsen H, Chevalier S, Sonenberg N (1998) 4E-BP3, a new member of the eukaryotic initiation factor 4E-binding protein family. J Biol Chem 273:14002–14007
Miron M, Verdu J, Lachance PE, Birnbaum MJ, Lasko PF, Sonenberg N (2001) The translational inhibitor 4E-BP is an effector of PI(3)K/Akt signalling and cell growth in Drosophila. Nat Cell Biol 3:596–601
Gingras AC, Gygi SP, Raught B et al (1999) Regulation of 4E-BP1 phosphorylation: a novel two-step mechanism. Genes Dev 13:1422–1437
Gingras AC, Raught B, Gygi SP et al (2001) Hierarchical phosphorylation of the translation inhibitor 4E-BP1. Genes Dev 15:2852–2864
Averous J, Proud CG (2006) When translation meets transformation: the mTOR story. Oncogene 25:6423–6435
Shima H, Pende M, Chen Y, Fumagalli S, Thomas G, Kozma SC (1998) Disruption of the p70(s6k)/p85(s6k) gene reveals a small mouse phenotype and a new functional S6 kinase. EMBO J 17:6649–6659
Montagne J, Stewart MJ, Stocker H, Hafen E, Kozma SC, Thomas G (1999) Drosophila S6 kinase: a regulator of cell size. Science 285:2126–2129
Fumagalli S, Thomas G (2000) S6 Phosphorylation and signal transduction. In: Sonenberg N, Hershey JWB, Mathews MB (eds) Translational control of gene expression. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY, pp 695–717
Pullen N, Dennis PB, Andjelkovic M et al (1998) Phosphorylation and activation of p70s6k by PDK1. Science 279:707–710
Alessi DR, Kozlowski MT, Weng QP, Morrice N, Avruch J (1998) 3-Phosphoinositide-dependent protein kinase 1 (PDK1) phosphorylates and activates the p70 S6 kinase in vivo and in vitro. Curr Biol 8:69–81
Avruch J, Belham C, Weng Q, Hara K, Yonezawa K (2001) The p70 S6 kinase integrates nutrient and growth signals to control translational capacity. Prog Mol Subcell Biol 26: 115–154
Stolovich M, Tang H, Hornstein E et al (2002) Transduction of growth or mitogenic signals into translational activation of TOP mRNAs is fully reliant on the phosphatidylinositol 3-kinase-mediated pathway but requires neither S6K1 nor rpS6 phosphorylation. Mol Cell Biol 22:8101–8113
Tang H, Hornstein E, Stolovich M et al (2001) Amino acid-induced translation of TOP mRNAs is fully dependent on phosphatidylinositol 3-kinase-mediated signaling, is partially inhibited by rapamycin, and is independent of S6K1 and rpS6 phosphorylation. Mol Cell Biol 21:8671–8683
Pende M, Um SH, Mieulet V et al (2004) S6K1(–/–)/S6K2(–/–) mice exhibit perinatal lethality and rapamycin-sensitive 5′-terminal oligopyrimidine mRNA translation and reveal a mitogen-activated protein kinase-dependent S6 kinase pathway. Mol Cell Biol 24:3112–3124
Raught B, Peiretti F, Gingras AC et al (2004) Phosphorylation of eucaryotic translation initiation factor 4B Ser422 is modulated by S6 kinases. EMBO J 23:1761–1769
Shahbazian D, Roux PP, Mieulet V et al (2006) The mTOR/PI3K and MAPK pathways converge on eIF4B to control its phosphorylation and activity. EMBO J 25:2781–2791
Holz MK, Ballif BA, Gygi SP, Blenis J (2005) mTOR and S6K1 mediate assembly of the translation preinitiation complex through dynamic protein interchange and ordered phosphorylation events. Cell 123:569–580
Shahbazian et al, (2010, Mar) Molecular and Cellular Biology. Control of cell survival and proliferation by mammalian eukaryotic initiation factor 4B. 30(6):1478–1485
Dmitriev SE, Terenin IM, Dunaevsky YE, Merrick WC, Shatsky IN (2003) Assembly of 48S translation initiation complexes from purified components with mRNAs that have some base pairing within their 5′ untranslated regions. Mol Cell Biol 23:8925–8933
Duncan R, Hershey JW (1985) Regulation of initiation factors during translational repression caused by serum depletion. Covalent modification. J Biol Chem 260:5493–5497
Dorrello NV, Peschiaroli A, Guardavaccaro D, Colburn NH, Sherman NE, Pagano M (2006) S6K1- and betaTRCP-mediated degradation of PDCD4 promotes protein translation and cell growth. Science 314:467–471
Browne GJ, Proud CG (2002) Regulation of peptide-chain elongation in mammalian cells. Eur J Biochem 269:5360–5368
Wang X, Li W, Williams M, Terada N, Alessi DR, Proud CG (2001) Regulation of elongation factor 2 kinase by p90(RSK1) and p70 S6 kinase. EMBO J 20:4370–4379
Richardson CJ, Broenstrup M, Fingar DC et al (2004) SKAR is a specific target of S6 kinase 1 in cell growth control. Curr Biol 14:1540–1549
Ma XM, Yoon SO, Richardson CJ, Julich K, Blenis J (2008) SKAR links pre-mRNA splicing to mTOR/S6K1-mediated enhanced translation efficiency of spliced mRNAs. Cell 133: 303–313
Manning BD (2004) Balancing Akt with S6K: implications for both metabolic diseases and tumorigenesis. J Cell Biol 167:399–403
Greene MW, Sakaue H, Wang L, Alessi DR, Roth RA (2003) Modulation of insulin-stimulated degradation of human insulin receptor substrate-1 by Serine 312 phosphorylation. J Biol Chem 278:8199–8211
Shah OJ, Wang Z, Hunter T (2004) Inappropriate activation of the TSC/Rheb/mTOR/S6K cassette induces IRS1/2 depletion, insulin resistance, and cell survival deficiencies. Curr Biol 14:1650–1656
Harrington LS, Findlay GM, Gray A et al (2004) The TSC1-2 tumor suppressor controls insulin-PI3K signaling via regulation of IRS proteins. J Cell Biol 166:213–223
O’Reilly KE, Rojo F, She QB et al (2006) mTOR inhibition induces upstream receptor tyrosine kinase signaling and activates Akt. Cancer Res 66:1500–1508
Pyronnet S, Imataka H, Gingras AC, Fukunaga R, Hunter T, Sonenberg N (1999) Human eukaryotic translation initiation factor 4G (eIF4G) recruits mnk1 to phosphorylate eIF4E. EMBO J 18:270–279
Wakiyama M, Imataka H, Sonenberg N (2000) Interaction of eIF4G with poly(A)-binding protein stimulates translation and is critical for Xenopus oocyte maturation. Curr Biol 10:1147–1150
Wells SE, Hillner PE, Vale RD, Sachs AB (1998) Circularization of mRNA by eukaryotic translation initiation factors. Mol Cell 2:135–140
Raught B, Gingras AC, Gygi SP et al (2000) Serum-stimulated, rapamycin-sensitive phosphorylation sites in the eukaryotic translation initiation factor 4GI. EMBO J 19:434–444
Tuazon PT, Merrick WC, Traugh JA (1989) Comparative analysis of phosphorylation of translational initiation and elongation factors by seven protein kinases. J Biol Chem 264:2773–2777
Lee VH, Healy T, Fonseca BD, Hayashi A, Proud CG (2008) Analysis of the regulatory motifs in eukaryotic initiation factor 4E-binding protein 1. FEBS J 275:2185–2199
Yokogami K, Wakisaka S, Avruch J, Reeves SA (2000) Serine phosphorylation and maximal activation of STAT3 during CNTF signaling is mediated by the rapamycin target mTOR. Curr Biol 10:47–50
Sarbassov DD, Ali SM, Kim DH et al (2004) Rictor, a novel binding partner of mTOR, defines a rapamycin-insensitive and raptor-independent pathway that regulates the cytoskeleton. Curr Biol 14:1296–1302
Bjornsti MA, Houghton PJ (2004) The TOR pathway: a target for cancer therapy. Nat Rev Cancer 4:335–348
Hemminki A, Markie D, Tomlinson I et al (1998) A serine/threonine kinase gene defective in Peutz-Jeghers syndrome. Nature 391:184–187
Inoki K, Zhu T, Guan KL (2003) TSC2 mediates cellular energy response to control cell growth and survival. Cell 115:577–590
Di Cristofano A, Pandolfi PP (2000) The multiple roles of PTEN in tumor suppression. Cell 100:387–390
Jaeschke A, Hartkamp J, Saitoh M et al (2002) Tuberous sclerosis complex tumor suppressor-mediated S6 kinase inhibition by phosphatidylinositide-3-OH kinase is mTOR independent. J Cell Biol 159:217–224
Podsypanina K, Ellenson LH, Nemes A et al (1999) Mutation of Pten/Mmac1 in mice causes neoplasia in multiple organ systems. Proc Natl Acad Sci USA 96:1563–1568
Neshat MS, Mellinghoff IK, Tran C et al (2001) Enhanced sensitivity of PTEN-deficient tumors to inhibition of FRAP/mTOR. Proc Natl Acad Sci USA 98:10314–10319
Shaw RJ, Bardeesy N, Manning BD et al (2004) The LKB1 tumor suppressor negatively regulates mTOR signaling. Cancer Cell 6:91–99
Faivre S, Kroemer G, Raymond E (2006) Current development of mTOR inhibitors as anticancer agents. Nat Rev Drug Discov 5:671–688
Fry MJ (2001) Phosphoinositide 3-kinase signalling in breast cancer: how big a role might it play? Breast Cancer Res 3:304–312
Rousseau D, Gingras AC, Pause A, Sonenberg N (1996) The eIF4E-binding proteins 1 and 2 are negative regulators of cell growth. Oncogene 13:2415–2420
Avdulov S, Li S, Michalek V et al (2004) Activation of translation complex eIF4F is essential for the genesis and maintenance of the malignant phenotype in human mammary epithelial cells. Cancer Cell 5:553–563
Armengol G, Rojo F, Castellvi J et al (2007) 4E-binding protein 1: a key molecular “funnel factor” in human cancer with clinical implications. Cancer Res 67:7551–7555
Castellvi J, Garcia A, Rojo F et al (2006) Phosphorylated 4E binding protein 1: a hallmark of cell signaling that correlates with survival in ovarian cancer. Cancer 107:1801–1811
Rojo F, Najera L, Lirola J et al (2007) 4E-binding protein 1, a cell signaling hallmark in breast cancer that correlates with pathologic grade and prognosis. Clin Cancer Res 13: 81–89
Akcakanat A, Sahin A, Shaye AN, Velasco MA, Meric-Bernstam F (2008) Comparison of Akt/mTOR signaling in primary breast tumors and matched distant metastases. Cancer 112:2352–2358
Zhou X, Tan M, Stone Hawthorne V et al (2004) Activation of the Akt/mammalian target of rapamycin/4E-BP1 pathway by ErbB2 overexpression predicts tumor progression in breast cancers. Clin Cancer Res 10:6779–6788
Lazaris-Karatzas A, Sonenberg N (1992) The mRNA 5′ cap-binding protein, eIF-4E, cooperates with v-myc or E1A in the transformation of primary rodent fibroblasts. Mol Cell Biol 12:1234–1238
Lazaris-Karatzas A, Montine KS, Sonenberg N (1990) Malignant transformation by a eukaryotic initiation factor subunit that binds to mRNA 5′ cap. Nature 345:544–547
Ruggero D, Montanaro L, Ma L et al (2004) The translation factor eIF-4E promotes tumor formation and cooperates with c-Myc in lymphomagenesis. Nat Med 10:484–486
Wendel HG, De Stanchina E, Fridman JS et al (2004) Survival signalling by Akt and eIF4E in oncogenesis and cancer therapy. Nature 428:332–337
Clemens MJ, Bommer UA (1999) Translational control: the cancer connection. Int J Biochem Cell Biol 31:1–23
Mamane Y, Petroulakis E, Rong L, Yoshida K, Ler LW, Sonenberg N (2004) eIF4E – from translation to transformation. Oncogene 23:3172–3179
De Benedetti A, Harris AL (1999) eIF4E expression in tumors: its possible role in progression of malignancies. Int J Biochem Cell Biol 31:59–72
Graff JR, Konicek BW, Carter JH, Marcusson EG (2008) Targeting the eukaryotic translation initiation factor 4E for cancer therapy. Cancer Res 68:631–634
Graff JR, Konicek BW, Vincent TM et al (2007) Therapeutic suppression of translation initiation factor eIF4E expression reduces tumor growth without toxicity. J Clin Invest 117:2638–2648
Zhou HY, Wong AS (2006) Activation of p70S6K induces expression of matrix metalloproteinase 9 associated with hepatocyte growth factor-mediated invasion in human ovarian cancer cells. Endocrinology 147:2557–2566
Fingar DC, Salama S, Tsou C, Harlow E, Blenis J (2002) Mammalian cell size is controlled by mTOR and its downstream targets S6K1 and 4EBP1/eIF4E. Genes Dev 16:1472–1487
Pon YL, Zhou HY, Cheung AN, Ngan HY, Wong AS (2008) p70 S6 kinase promotes epithelial to mesenchymal transition through snail induction in ovarian cancer cells. Cancer Res 68:6524–6532
Kwon HK, Bae GU, Yoon JW et al (2002) Constitutive activation of p70S6k in cancer cells. Arch Pharm Res 25:685–690
Couch FJ, Wang XY, Wu GJ, Qian J, Jenkins RB, James CD (1999) Localization of PS6K to chromosomal region 17q23 and determination of its amplification in breast cancer. Cancer Res 59:1408–1411
Marte BM, Graus-Porta D, Jeschke M, Fabbro D, Hynes NE, Taverna D (1995) NDF/heregulin activates MAP kinase and p70/p85 S6 kinase during proliferation or differentiation of mammary epithelial cells. Oncogene 10:167–175
Jastrzebski K, Hannan KM, Tchoubrieva EB, Hannan RD, Pearson RB (2007) Coordinate regulation of ribosome biogenesis and function by the ribosomal protein S6 kinase, a key mediator of mTOR function. Growth Factors 25:209–226
Nakamura JL, Garcia E, Pieper RO (2008) S6K1 plays a key role in glial transformation. Cancer Res 68:6516–6523
Barlund M, Forozan F, Kononen J et al (2000) Detecting activation of ribosomal protein S6 kinase by complementary DNA and tissue microarray analysis. J Natl Cancer Inst 92: 1252–1259
Barlund M, Monni O, Kononen J et al (2000) Multiple genes at 17q23 undergo amplification and overexpression in breast cancer. Cancer Res 60:5340–5344
Heinonen H, Nieminen A, Saarela M et al (2008) Deciphering downstream gene targets of PI3K/mTOR/p70S6K pathway in breast cancer. BMC Genomics 9:348
Fukuchi-Shimogori T, Ishii I, Kashiwagi K, Mashiba H, Ekimoto H, Igarashi K (1997) Malignant transformation by overproduction of translation initiation factor eIF4G. Cancer Res 57:5041–5044
Braunstein S, Karpisheva K, Pola C et al (2007) A hypoxia-controlled cap-dependent to cap-independent translation switch in breast cancer. Mol Cell 28:501–512
Bauer C, Brass N, Diesinger I, Kayser K, Grasser FA, Meese E (2002) Overexpression of the eukaryotic translation initiation factor 4G (eIF4G-1) in squamous cell lung carcinoma. Int J Cancer 98:181–185
Bauer C, Diesinger I, Brass N, Steinhart H, Iro H, Meese EU (2001) Translation initiation factor eIF-4G is immunogenic, overexpressed, and amplified in patients with squamous cell lung carcinoma. Cancer 92:822–829
Sarbassov DD, Ali SM, Sengupta S et al (2006) Prolonged rapamycin treatment inhibits mTORC2 assembly and Akt/PKB. Mol Cell 22:159–168
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2009 Springer Science+Business Media, LLC
About this chapter
Cite this chapter
Dowling, R.J.O., Sonenberg, N. (2009). Downstream of mTOR: Translational Control of Cancer. In: Polunovsky, V., Houghton, P. (eds) mTOR Pathway and mTOR Inhibitors in Cancer Therapy. Cancer Drug Discovery and Development. Humana Press, Totowa, NJ. https://doi.org/10.1007/978-1-60327-271-1_10
Download citation
DOI: https://doi.org/10.1007/978-1-60327-271-1_10
Published:
Publisher Name: Humana Press, Totowa, NJ
Print ISBN: 978-1-60327-270-4
Online ISBN: 978-1-60327-271-1
eBook Packages: MedicineMedicine (R0)