Abstract
Chronic infection with hepatitis B virus (HBV) occurs in approximately 5 % of the world’s human population and persistence of the virus is associated with serious complications of cirrhosis and liver cancer. Currently available treatments are modestly effective and advancing novel therapeutic strategies is a medical priority. Stability of the viral cccDNA replication intermediate is a major factor that has impeded the development of therapies that are capable of eliminating chronic infection. Recent advances that employ gene therapy strategies offer useful advantages over current therapeutics. Silencing of HBV gene expression by harnessing the RNA interference pathway has been shown to be highly effective in cell culture and in vivo. However, a potential limitation of this approach is that the post-transcriptional mechanism of gene silencing does not disable cccDNA. Early results using designer transcription activator-like effector nucleases (TALENs) and repressor TALEs (rTALEs) have shown potential as a mode of inactivating cccDNA. In this article, we review the recent advances that have been made in HBV gene therapy, with a particular emphasis on the potential anti-HBV therapeutic utility of designed sequence-specific DNA binding proteins and their derivatives.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- HBV
- cccDNA
- RNAi
- Designer nucleases
- TALENs
- Zinc finger nucleases
- Repressor TALEs
- Targeted disruption
- Site-directed mutagenesis
Hepatitis B Virus Epidemiology
Conservative estimates of the global prevalence of chronic hepatitis B virus (HBV) infection place the number of affected individuals in excess of 240 million worldwide [1, 2]. Although a third of the world’s population has been exposed to the virus [1, 2], most acute infections are cleared spontaneously. Infections that are not cleared progress to chronicity and it is this persistent HBV infection that is associated with serious sequelae, such as cirrhosis and liver cancer (hepatocellular carcinoma or HCC). Annually 600,000 deaths are attributable to complications that arise from chronic HBV infection. HBV alone accounts for 53 % of new HCC cases worldwide and the hepatitis C virus (HCV) is estimated to account for 25 % of new cases [3]. HCC is a particularly aggressive cancer that has a high mortality. In 2008, 92 % of new HCC cases were fatal [4]. Although vaccination effectively prevents HBV infection, the incidence of liver cancer has not changed significantly in the past 10 years. Moreover, modest curative efficacy of currently available treatment regimens is unlikely to prevent complications arising in those already infected with the virus. Liver cancer remains the sixth most common cancer worldwide and is still ranked as the third most common cause of cancer-related deaths [4, 5]. HBV is hyperendemic to sub-Saharan Africa, East and South-East Asia as well as the western Pacific islands. These regions, largely comprising developing countries, are also the most severely affected by HCC. HBV itself is a non-cytopathic virus; however, the increased risks of cirrhosis and HCC associated with chronic viral infection makes the disease a global priority.
Evidence from early intervention programmes have shown that decreasing the incidence of HBV has a positive impact on the incidence of cirrhosis and liver cancer [6, 7]. In 1984 Taiwan implemented the first anti-HBV vaccination programme and the most recent data from 2004 demonstrate that, with a 97 % vaccination coverage rate, HBV seroprevalence in children has decreased from 9.8 % to 0.6 % [7]. The decrease in HBV prevalence has been accompanied by a decrease in the incidence of HCC [7]. However, vaccination failure may occur and has been attributed to emergence of viral vaccine escape mutants. Nevertheless, these cases represent just a small number of individuals, only 33 in total. A second limitation of vaccination is that it is prophylactic and not therapeutic. As a consequence vaccination has little therapeutic benefit in cases where chronic HBV infection has already been established. Seven drugs are currently licensed for treatment of chronic HBV infection (reviewed in ref. [8]). These are broadly divided into two groups: (1) immunomodulators (interferon alpha (IFN-α) and pegylated IFN-α) and (2) nucleoside and nucleotide analogues (lamivudine, telbivudine, adefovir dipivoxil, tenofovir disoproxil fumarate and entecavir), which act by inhibiting viral reverse transcriptase. The efficacy of immunomodulators is limited as side effects are common, there are several contraindications and cure occurs in only a small subset of chronic HBV carriers. Nucleoside and nucleotide analogues exhibit a number of advantages over immunomodulators, which include ease of use (oral administration route) and better patient tolerance to the drugs. However, the first generation drugs exhibit a low barrier to resistance and viral escape rates have been reported to range from 29 % to 80 % [9–11]. Treatment with lamivudine in particular is complicated by high rates of viral escape by mutation [9, 10]. Newer drugs exhibit improved viral suppression and also limit development of viral escape (low in the case of entecavir and none reported for tenofovir (for review see ref. [8])). Although these data are promising, nucleoside and nucleotide analogues rarely eliminate HBV completely from infected hepatocytes. Stability of the covalently closed circular DNA (cccDNA) replication intermediate of HBV, and inability of available therapies to disable this viral transcription template, are mainly responsible for the limitations of available HBV treatments.
HBV Biology
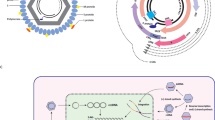
HBV is an enveloped DNA virus belonging to the Hepadnaviridae family of viruses (reviewed in ref. [12]). The DNA genome is contained within an icosahedral capsid which in turn is enveloped in a lipid bilayer. The genome of HBV exists in a partly double-stranded, circular conformation called the relaxed circular DNA or rcDNA [13]. The minus strand encompasses the entire genome and comprises approximately 3,200 bases. Until recently the receptor used by HBV to enter liver cells has remained elusive. Yan and colleagues identified the sodium taurocholate cotransporting peptide (NTCP) as the receptor employed by the virus to infect hepatocytes [14]. A basic overview of hepatocyte infection and viral replication is illustrated in Fig. 1. Upon infection the rcDNA, still contained within the capsid, is transported to the nucleus where it is released and “repaired” by cellular enzymes to form cccDNA [15]. The cccDNA serves as template for transcription of viral RNAs. These include the greater-than-genome length pregenomic RNA (pgRNA) which is reverse transcribed to form the viral rcDNA genome. The cccDNA exists episomally as a stable minichromosome [15, 16]. Since viral replication may be initiated from this stable minichromosome, an effective cure for chronic HBV infection requires that cccDNA be inactivated or eliminated.
Hepatitis B virus replication cycle. Following attachment to the hepatocyte cell membrane, the capsid is released into the cytoplasm and is then translocated to the nucleus. Here the rcDNA viral genome is released and repaired to form cccDNA. The cccDNA acts as the primary replication intermediate from which the pgRNA and viral subgenomic mRNAs are transcribed. These viral RNAs are transported to the cytoplasm where the subgenomic mRNAs are translated into proteins required for virion packaging and assembly. The pgRNA associates with polymerase and new core proteins to assemble the new capsid. Within the capsid, the pgRNA is reverse transcribed to form rcDNA. Within the endoplasmic reticulum, surface proteins surround the capsid to form new virions before secretion from the infected hepatocyte. Current gene therapy targets include the episomal cccDNA (A) for targeted gene disruption and transcriptional repression, and the cytoplasmic viral mRNAs (B) for a post-transcriptional gene silencing approach
HBV produces new virion DNA through error-prone reverse transcription of the pgRNA template. Although numerous mutations may be introduced during this reverse transcription step, the highly compact genome limits sequence plasticity and emergence of mutant strains. The four viral open reading frames (ORFs), which code for seven viral proteins, overlap with one another. Furthermore, the viral ORFs cover the entire genome of HBV and all viral regulatory elements are contained within protein coding regions (reviewed in ref. [17]). Since most regions of the HBV genome have dual use, mutations at one site commonly affect more than one genetic function to compromise viral fitness severely. The HBV genome is therefore more stable than genomes of other viruses that employ a reverse transcriptase during replication. This feature makes HBV an ideal target for gene therapy based on sequence-specific DNA recognition.
Until recently the technology required for efficient targeted disruption of specific DNA sequences was not readily available. Gene therapy using cccDNA-targeting engineered proteins has shown promise as a means of disabling HBV replication. Discovery of gene silencing by RNA interference (RNAi) was a major development in gene therapy. Several studies have reported that harnessing this pathway can be used successfully to inhibit HBV replication. In this chapter we summarise recent advances in HBV gene therapy, with particular emphasis on progress in adapting sequence-specific DNA binding proteins to counter the viral infection.
RNAi Against HBV
Activation of RNAi by microRNAs (miRNAs) is the prototypic mechanism by which endogenous post-transcriptional silencing of target genes is achieved [18]. The pathway occurs in metazoan cells and naturally involves stepwise processing of RNAs containing hairpin motifs. This leads to production of mature miRNAs, comprising hairpin-derived duplexes of approximately 23 bp. Post-transcriptional gene silencing by the guide strand from a mature miRNA is effected by the RNA induced silencing complex (RISC) and involves hybridisation of the guide to complementary sequences in the 3′ untranslated region of target genes. Efficient gene silencing may be achieved by introducing artificial RNA mimics of intermediates of the RNAi pathway into cells. Both synthetic [19–23] and expressed [24–31] exogenous activators of the RNAi pathway have been used to silence HBV replication in vitro and in vivo. Studies using synthetic RNAi activators against HBV were amongst the first to demonstrate the utility of harnessing RNAi in vivo [19, 21]. An important finding into the mechanistic aspect of these therapeutic sequences was that RNAi-mediated silencing did not require viral replication [19]. This is in contrast to the mechanism of action of nucleotide and nucleoside analogues, which need to be incorporated into a growing DNA chain to exert their inhibitory effects. RNAi-based activators interfere with gene expression at a post-transcriptional level and therefore act at a later stage of HBV replication than do nucleoside and nucleotide analogues. Since these first reports demonstrating potential utility of RNAi inducers, significant progress has been made using synthetic as wells as expressed anti-HBV effectors. Various chemical modifications have been introduced into synthetic anti-HBV sequences [32–35]. These have been formulated in non-viral vectors and shown to be effective inhibitors of HBV replication. Anti-HBV expression cassettes, which generate precursor miRNA (pre-miRNA) and primary miRNA (pri-miRNA) sequences transcribed from RNA polymerase (Pol) II and Pol III transcription regulatory elements, have also been found to be effective against the virus [36–42]. Some of these cassettes have been incorporated into recombinant adeno- and adeno-associated viral vectors, which have been effective against the virus in murine models of HBV replication. However, as with nucleoside and nucleotide analogues, RNAi-based therapy does not completely eliminate the stable pool of cccDNA. Consequently RNAi-based approaches are unlikely to cure HBV infection. This was demonstrated by Starkey et al. [43], who showed that expressed anti-HBV RNAi activators inhibited new formation of cccDNA in cultured hepatocyte-derived cells. However, concentrations of established cccDNA were unaffected by the HBV-targeting short hairpin RNAs (shRNAs). These results were not entirely unexpected and confirmed that RNAi-based anti-HBV agents function by a post-transcriptional mechanism to inhibit gene expression from the cccDNA template. Viral RNA, viral protein and new virion formation are subsequently suppressed but cccDNA levels remain unaffected. To have therapeutic utility against HBV, RNAi activators would need to be administered repeatedly or produced in a sustained manner from stable intrahepatic DNA expression cassettes. Both strategies pose challenges and highlight the need to develop strategies for directly disabling nuclear cccDNA.
Targeting cccDNA Using Engineered Sequence-Specific DNA Binding Proteins
During the past decade, engineered Zinc Finger Proteins (ZFPs) have been widely used to regulate expression of genes for therapeutic gain [44]. These DNA binding proteins, which occur naturally as eukaryotic transcription factors, may be designed to target specific DNA sequences. Since each zinc finger is capable of binding a triplet of nucleotides, the sequential arrangement of an array of six “fingers” enables site-specific targeting of up to 18 nucleotides by ZFPs (Fig. 2a) [45]. Although these DNA binding proteins have been engineered primarily for endogenous gene regulation, the chromosome-like structure of HBV cccDNA is likely to be amenable to similar modes of transcriptional manipulation. This was initially investigated by Zimmerman and colleagues who generated polydactyl ZFPs to manipulate duck hepatitis B virus (DHBV) gene expression [46]. They demonstrated that ZFPs designed to bind either 9 or 18 base pair sequences within the enhancer region of DHBV cccDNA reduced pgRNA expression by 41.6 %. Decreases in the DHBV surface and core proteins, as well as viral particle equivalents, were also observed. As the enhancer region includes multiple transcription factor binding sites, it is likely that these ZFPs functioned by competitively obstructing binding of transcription factors to cis elements of the cccDNA. However, as these ZFPs do not exert a permanent effect on their HBV targets, effectiveness is likely to be transient and dependent on the duration of their expression. Nevertheless, demonstration of an effect on cccDNA was a significant observation. This paved the way for investigating the utility of functionally enhanced DNA binding proteins that target cccDNA in human liver cells. Coupling transcriptional repressor domains such as the Krüppel associated box (KRAB) domain to DNA sequence-specific binding domains of Transcription Activator-Like Effectors (TALEs) to achieve more durable silencing may be preferable (Fig. 2b) (discussed later).
Proteins that bind specific DNA sequences and which have been used to inhibit HBV gene transcription. (a) Polydactyl ZFPs include multiple zinc fingers, each targeting a predefined nucleotide triplet. The length of the DNA binding region can be adjusted to target an 18 base pair sequence by adding up to six consecutive fingers within an array. (b) Repressor TALEs (rTALEs) are generated by fusing KRAB repressor domains to the N-terminal of the DNA binding TALE proteins. The repeat domain comprises 19 TALE modules, each with a nucleotide specific RVD to enable sequence-specific DNA binding. The nuclear localisation signal (NLS) facilitates efficient trafficking of the protein into the cell nucleus, whilst the haemagglutinin epitope (HA) sequence acts as a tag for convenient protein detection
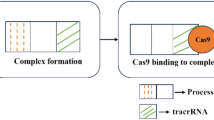
Designer nucleases have been developed by engineering DNA binding proteins to enable introduction of double stranded breaks (DSBs) at specific target sites (Fig. 3). DSBs are typically repaired by non-homologous end-joining (NHEJ) or homology directed repair (HDR) (Fig. 4) (reviewed in refs. [47, 48]). The NHEJ pathway is intrinsically error prone and may cause combinations of insertions, deletions and substitutions at sites of DSBs. By introducing DSBs designer nucleases may therefore be used to introduce disabling mutations at an intended target. When homologous sequences are introduced into target cells together with DSB-inducing nucleases, HDR may be used to restore gene function, as has been applied to the treatment of monogenic inherited diseases [49]. Gene correction, particularly for the treatment of monogenic disorders, has been a major focus of designer nuclease research [50]. However, the intentional disruption of pathology-causing viral DNA has generated considerable enthusiasm for this approach to viral gene therapy (reviewed in ref. [51]). Viruses that are capable of establishing latent infections, such as herpes simplex virus (HSV), human immunodeficiency virus (HIV) and HBV, are prime candidates for nuclease-mediated targeted disruption. Since HBV has a small compact genome arrangement with restricted sequence plasticity, it is suited to this gene therapy approach. Currently there are three different types of designer nucleases that have been used to target HBV cccDNA: homing endonucleases, otherwise known as meganucleases (Fig. 3a); zinc finger nucleases (ZFNs) (Fig. 3b) and TALE nucleases (TALENs) (Fig. 3c). Recent research on protein-RNA-based CRISPR (Clustered Regulatory Interspaced Short Palindromic Repeats) and CRISPR-associated (Cas) nucleases [52, 53] to modify genomic targets has been particularly vigorous. CRISPR/Cas nucleases have, however, not yet been used against HBV, but this topic is no doubt under investigation.
Designer nucleases used to disable HBV sequences. The three different engineered nucleases currently being investigated as potential anti-HBV therapies are illustrated schematically. (a) Homing endonucleases or meganucleases are found naturally as endonucleases which target DNA sequences between 12 and 40 bp in length. (b) ZFNs are engineered as pairs with each ZF in the array binding to a specific nucleotide triplet. When a polydactyl ZFP array consists of four ZFs, each left (L) or right (R) subunit targets a 12 bp sequence. (c) TALENs are engineered as pairs with each left (L) or right (R) subunit DNA-binding domain targeting a 19 bp sequence. Each subunit is assembled from single TALE repeats which confer single nucleotide specificity. Both ZFNs and TALENs cleave target DNA with FokI endonuclease domains to introduce double strand breaks (DSBs) indicated by the asterisks
Cellular repair pathways used during nuclease-mediated mutagenesis or gene correction. (a) TALEN dimerisation, depicted here, leads to cleavage at the DNA target sites and formation of DSBs, which are typically repaired by non-homologous end joining (NHEJ) (b) or homology-directed repair (HDR) (c). Meganucleases or ZFNs may also cleave double stranded DNA to initiate NHEJ or HDR. When NHEJ is triggered, DSB repair factors are recruited to the site. Repair may lead to restoration of the wild-type sequence or introduction of mutations. Abbreviations are spelled out in the legend to the figure to maintain consistency with the other figures. (insertions, deletions or substitutions), which occurs commonly when repeated cleavage occurs at a target site. If donor sequences with homologous regions flanking the DSB site are introduced, the HDR pathway may be triggered to repair or insert a sequence at the DSB
Homing endonucleases, also known as meganucleases, are DNA-specific cleavage proteins that were identified within mobile genetic elements of yeast (Fig. 3a) [54, 55]. They target DNA sequences ranging from 12 to 40 nucleotides in length, and have successfully been used to trigger HDR in eukaryotic cells [56]. Although homing endonucleases may be engineered to cleave defined DNA targets, the number of possible binding sites is limited by constraints of the effectiveness of motifs targeting these predefined targets. Despite this, homing endonucleases have recently been shown by Cellectis Bioresearch (Paris, France) to have therapeutic potential for treatment of HSV infection [57]. This group has also patented anti-HBV meganucleases (WO/2010/136981 and US2012/0171191A1) but published results of these studies are not yet available. Whilst meganucleases have natural endonuclease activity, ZFNs and TALENs are generated by fusing a FokI nuclease domain to the C-terminus of a ZFP or a TALE array (Fig. 3b, c) [58, 59]. The FokI enzyme exists as a monomer but requires interactions between the catalytic domains of two monomers to enable cleavage of double stranded DNA [60, 61]. For this reason, ZFNs and TALENs are designed in pairs comprising so-called left and right subunits. Although both subunits are required for efficient cleavage, the catalytic activity of the FokI nuclease domain itself may function in a potentially genotoxic sequence-independent fashion.
ZFN DNA-binding domains typically include ZFPs, with left and right subunits that each target sequences of between 9 and 12 base pairs [45]. The DNA binding sites are separated by a six base pair spacer region to enable FokI dimerisation and induction of DSBs at intended target sites. This configuration has been shown to cleave endogenous human genes efficiently [62, 63]. In an assessment of antiviral efficacy, Cradick and colleagues investigated the ability of engineered ZFNs to cleave HBV DNA targets in vitro [64]. The ZFNs were designed to bind to two 9 base pair target sequences within the core ORF that overlapped with the common polyadenylation site. Disruption of viral DNA targets was verified following co-transfection of Huh7 cells with plasmids encoding ZFN pairs and an HBV replication competent plasmid. In addition to inhibiting pgRNA transcription, the anti-HBV ZFNs cleaved 36 % of DNA targets. Sequencing indicated that site-directed mutations occurred in 6 % of HBV targets, of which 81 % produced frameshift mutations. Although the activity of these ZFNs on episomal cccDNA was not determined, this important study was the first to describe targeted disruption of HBV DNA as a potential gene therapy.
Recently designer nucleases have been re-engineered by substituting ZFPs with the plant bacterial TALEs as an alternative DNA binding domain (Fig. 3c). TALEs, which are transcription activators of host avirulence genes, are natural pro-survival proteins produced by the Xanthomonas plant pathogens [65]. Unlike ZFPs, which require specific context-dependent arrangement of each protein subunit for effective DNA binding [66, 67], a single monomer within the TALE DNA binding domain confers individual nucleotide specificity with little effect of neighbouring monomers [68, 69]. As a result, TALE monomers can be conveniently concatamerised to form sequence-specific DNA-binding domains that may be fused to the FokI nuclease domain [70]. Individual TALE monomers comprise approximately 34 amino acid polypeptide chains and TALE arrays are made up of tandem repeats of these monomers. Each monomer varies at amino acid positions 12 and 13, and these repeat variable diresidues (RVDs) confer specificity of binding by the TALE sequences to their DNA targets [68, 69, 71]. TALEN subunits are typically designed to target 19 base pair sequences. This confers higher sequence specificity than ZFNs, and may also account for their improved activity in vitro [72]. There are several commercial and publically available techniques currently used to assemble designer TALE (dTALE) arrays. Two commercial companies, Cellectis Bioresearch (Paris, France) and Life Technologies (New York, USA), offer design and synthesis services whilst several publically available methods have successfully been used to generate TALE arrays and TALENs [70]. As with the ZFNs, TALENs have been used to target multiple endogenous genes [73–75; however, the antiviral efficacy of TALENs has only recently been described [76, 77].
Disrupting the HBV cccDNA Minichromosome
Building on the success of the anti-HBV ZFNs [64], HBV-specific TALENs were investigated for their efficacy against viral DNA [76]. TALENs were engineered to target multifunctional sites within the surface (S), core (C), and polymerase (P1 and P2) ORFs. Co-transfection experiments, conducted using cultured liver-derived Huh7 cells, showed that the S-TALEN inhibited HBsAg secretion by 80 %. This result was corroborated by studies on the HepG2.2.15 cell line, which stably and constitutively produces HBV. In these cells, the S-TALEN-encoding sequences mediated inhibition of HBsAg secretion by approximately 60 % and caused targeted disruption of 31–35 % of cccDNA copies. The C-TALEN caused 12 % targeted disruption of cccDNA and predictably did not inhibit HBsAg secretion. Although TALENs provide an efficient means of disrupting HBV DNA sequences, cytotoxicity resulting from non-specific targeting is possible and may cause off target mutagenesis and cell death [78, 79]. A genome wide analysis confirmed that no potential off-target binding sites for P1-, P2-, S- and C-TALENs were found in either the mouse (Mus musculus) or human genomes. This finding is in line with the lack of cytotoxicity that was observed when treating cells with the panel of HBV-directed TALENs [76]. Moreover, multiple alignments of 26 different HBV genotypes and sub-genotypes showed high sequence homology at the selected TALEN target sites. This is particularly true for the S TALEN, and suggests that the HBV-targeting nucleases may be effective across HBV genotypes.
To establish whether TALENs were capable of inactivating HBV replication in vivo, the anti-HBV efficacy of the S and C TALENs were investigated in a murine model of HBV replication [76]. This was achieved by hydrodynamic tail vein injection of both TALEN and HBV-encoding plasmids. Importantly, murine hepatocytes do not support cccDNA formation [80] and the TALEN effects were a result of their action on the co-injected HBV replication-competent plasmid. In vivo, the S-TALEN inhibited HBsAg secretion by 95 % and induced disruption in 58–68 % of intrahepatic HBV DNA targets [76]. The C-TALEN inhibited HBcAg expression and induced disruption in 62–87 % of intrahepatic HBV DNA targets. Serological analysis showed a reduction in circulating virions and no apparent liver toxicity. Deep sequencing at the S- and C-TALEN binding sites showed targeted mutation of HBV that was specific to mice that had been treated with anti-HBV TALENs. As expected, deletions were predominantly detected at both the S- and C-TALEN target sites [81]. This effect is distinct from that of ZFNs, which may give rise to a combination of insertions, deletions and substitutions.
The therapeutic potential of anti-HBV TALENs has recently been corroborated by Chen and colleagues [77]. By engineering TALENs to bind to the core ORF and the RNaseH region of the polymerase ORF, significant knockdown of HBeAg, HBsAg and pgRNA levels in Huh7 cells could be achieved. This antiviral effect was observed for genotype B, C and D isolates, supporting the notion that TALENs designed to target conserved regions of HBV may be effective against several genotypes. Efficacy was also confirmed in vivo when using murine hydrodynamic injection. Importantly, this study also showed a synergistic antiviral effect when combining TALENs with INF-α in Huh7 cell cultures. As INF-α is a licensed HBV therapeutic, using TALENs in combination with other drugs is an interesting approach to improving treatment efficacy.
Transcriptional Gene Silencing with rTALEs
The endonuclease domains of TALENs and ZFNs may also be substituted with other effector proteins to enable gene-specific transcriptional regulation. Naturally, Xanthomonas TALEs contain activation domains, which increases transcription from specific plant gene promoters through association with the DNA-targeting repeat domains. After secretion from the bacteria, the TALEs enable survival of the bacteria through transcriptional activation of otherwise silent genes [65, 82–84]. As the TALE array can be engineered to bind to any target sequence, these transcriptional activators have been used in mammalian cells to trigger human gene expression [85–88]. As an alternative, TALE DNA binding arrays may be fused to gene repressors. These repressor TALEs (rTALEs) have recently been shown to be potent inhibitors of endogenous mammalian gene transcription [86, 87]. Different repressor domains have been fused to DNA-binding TALEs. Amongst the inhibitors of gene expression, the mSin interaction domain (SID) and KRAB domain silenced gene activity most efficiently [86, 87]. KRAB repressors occur naturally as zinc finger fusions [89], and are the largest group of mammalian transcriptional regulators (Fig. 2b). Although the exact mechanism of KRAB transcriptional repression has not been completely elucidated, KRAB-ZF protein binding results in the recruitment of heterochromatin forming complexes, which lead to gene silencing [90].
When studying anti-HBV efficacy of TALENs, it was found that one of the nucleases under investigation, the P1-TALEN, inhibited markers of viral replication without causing detectable mutation at the target site [76]. Since the HBV enhancer I sequence overlaps with the P1-TALEN target, it is likely that this TALEN inhibits viral replication through transcription inhibition rather than by mutating protein-coding sequences. In the same way that ZFPs were shown to inhibit DHBV transcription [46], transient transcriptional repression with TALENs is likely to involve competitive binding at the target site without any endonuclease cleavage. To investigate this further, we generated KRAB-rTALEs from the P1 and P2 left and right TALE DNA binding arrays (unpublished data). As with naturally occurring repressor proteins containing the KRAB domain, these rTALEs were designed with the repressor domain at the N-terminus (Fig. 2b). Individually, the left and right P1 rTALEs inhibited HBsAg expression in Huh7 cells following co-transfections with an HBV replication competent plasmid. Furthermore, surface and core mRNA concentrations were reduced by up to 80 %, suggesting the anti-HBV effect is a result of transcriptional inhibition. The HBV Enhancer I sequence regulates viral protein expression through binding of key hepatocyte transcription factors (reviewed in [17, 91, 92]). The P1 left rTALE is likely to inhibit the binding of retinoic acid response element (RARE) or regulatory factor X1 (RFX1), which in turn may prevent the cooperative binding of hepatocyte nuclear factor 4 (HNF-4) and retinoid X receptor alpha/peroxisome proliferator-activated receptor (RXRα/PPAR) heterodimers. The P1 right rTALE subunit, however, targets a region of the LSR element of Enhancer I motif that is essential for HBV gene expression. This sequence contains cis-elements that bind several hepatocyte transcription factors and is also involved in activation of HBx gene expression [93]. The Enhancer I motif operates in conjunction with Enhancer II, to increase surface protein expression [94]. As both the P1 left and P1 right rTALEs inhibit HBsAg expression, it is likely that these proteins are obstructing RARE and RFX1 binding or LSR regulation, and consequently inactivating the Enhancer I motif. Moreover, rTALE-induced formation of heterochromatin on the cccDNA may disable viral transcription permanently. Although TALENs are a promising therapeutic for the targeted inactivation of cccDNA, unwanted mutations and chromosome translocations are a concern [95]. HBV DNA is frequently integrated into the genomes of carriers of the virus, and TALENs acting at these sites may cause unwanted genotoxicity. By using transcriptional repression with rTALEs, instead of introducing targeted DSBs with TALENs, the likelihood of introducing undesirable mutagenic events may be diminished.
Conclusions
To date treatment with IFN-α or its pegylated derivatives has been the only intervention capable of eradicating HBV from infected cells [96]. The treatment is, however, only effective in a small subset of HBV carriers. Moreover use of IFN-α may be complicated by side effects, is contraindicated in several cases, and is expensive. Nucleoside and nucleotide analogues may effectively suppress viral replication but do not eliminate cccDNA and treatment withdrawal is associated with reactivation of HBV replication. With the advent of technology enabling the engineering of designer nucleases and transcriptional repressors, methods of efficiently silencing HBV replication by inactivating cccDNA have been added to the arsenal of potential anti-HBV therapies. Currently, the use of TALENs against HBV appears to be the most efficient. Although not yet reported, information on the utility of HBV-targeting CRISPR-Cas derivatives will be an interesting development.
Studies on viral resistance in cultured cell have been hampered by the lack of convenient models that simulate all steps of viral replication. Most reports describing anti-HBV therapy have employed replication competent plasmids or cell lines with an integrated copy of the viral genome. These strategies rely on assessing silencing of a single viral sequence and consequently provide little information on viral resistance. The discovery by Yan and colleagues that HBV uses the NTCP to infect hepatocytes may address this problem [14]. Cells ectopically expressing NTCP are permissive for HBV infection and should allow for analysis of emergence of resistance in response to treatment. Furthermore, as NTCP-expressing human cultured cells support cccDNA formation they will provide a useful model to assess the effect of anti-HBV therapeutics on cccDNA. Unfortunately, since mice do not synthesise cccDNA [80], NTCP-transgenic murine models will not recapitulate the entire HBV replication cycle.
Although results from testing anti-HBV nucleases and repressors have generated significant enthusiasm, significant hurdles need to be overcome before they are used in a therapeutic context. Safe and efficient delivery of sequences encoding anti-HBV nucleases is particularly challenging. The size of sequences encoding individual TALEN subunits (approximately 4 kb) is at the limit of the transgene capacity of single stranded adeno-associated viral (AAV) vectors. Delivery of complete TALENs with these favoured vectors would therefore require simultaneous administration of two recombinant viruses. Although constrained by the compact nature of its genome, selection of HBV escape mutants may be possible. It remains to be determined whether mutations within the HBV genome can confer resistance to TALEN- or rTALE-mediated silencing and if so, whether resistance can be prevented by simultaneously targeting the virus with multiple engineered DNA-binding proteins. The duration of expression of anti-HBV TALENs or rTALEs that is required to disable cccDNA completely is not yet known. Also, off target and long-term effects of expressing TALENs or rTALEs on hepatocytes need to be characterised. Thorough analysis of these topics will be required before this powerful technology is implemented as a therapeutic modality. Despite the unanswered questions, use of derivatives of engineered sequence-specific DNA binding proteins is an interesting novel strategy. Used alone or in combination with other antiviral approaches, such as RNAi and existing licensed therapies, they have the potential to improve the management of chronic carriers significantly.
References
Ott JJ, Stevens GA, Groeger J, Wiersma ST. Global epidemiology of hepatitis B virus infection: new estimates of age-specific HBsAg seroprevalence and endemicity. Vaccine. 2012;30(12):2212–9. PubMed PMID: 22273662. Epub 2012/01/26. eng.
WHO. Hepatitis B virus fact sheet no. 204 (Updated July 2013). 2013. http://www.who.int/mediacentre/factsheets/fs204/en/index.html.
Perz JF, Armstrong GL, Farrington LA, Hutin YJ, Bell BP. The contributions of hepatitis B virus and hepatitis C virus infections to cirrhosis and primary liver cancer worldwide. J Hepatol. 2006;45(4):529–38. PubMed PMID: 16879891. Epub 2006/08/02. eng.
Bray F, Jemal A, Grey N, Ferlay J, Forman D. Global cancer transitions according to the Human Development Index (2008-2030): a population-based study. Lancet Oncol. 2012;13(8):790–801. PubMed PMID: 22658655. Epub 2012/06/05. eng.
Parkin DM, Bray F, Ferlay J, Pisani P. Global cancer statistics, 2002. CA Cancer J Clin. 2005;55(2):74–108. PubMed PMID: 15761078. Epub 2005/03/12. eng.
Chien YC, Jan CF, Kuo HS, Chen CJ. Nationwide hepatitis B vaccination program in Taiwan: effectiveness in the 20 years after it was launched. Epidemiol Rev. 2006;28:126–35. PubMed PMID: 16782778. Epub 2006/06/20. eng.
Ni YH, Chen DS. Hepatitis B vaccination in children: the Taiwan experience. Pathol Biol (Paris). 2010;58(4):296–300. PubMed PMID: 20116181. Epub 2010/02/02. eng.
Zoulim F. Hepatitis B virus resistance to antiviral drugs: where are we going? Liver Int. 2011; 31 Suppl 1:111–6. PubMed PMID: 21205147. Pubmed Central PMCID: 3096621. Epub 2011/01/19. eng.
Lai CL, Dienstag J, Schiff E, Leung NW, Atkins M, Hunt C, et al. Prevalence and clinical correlates of YMDD variants during lamivudine therapy for patients with chronic hepatitis B. Clin Infect Dis. 2003;36(6):687–96. PubMed PMID: 12627352. Epub 2003/03/11. eng.
Lok AS, Lai CL, Leung N, Yao GB, Cui ZY, Schiff ER, et al. Long-term safety of lamivudine treatment in patients with chronic hepatitis B. Gastroenterology. 2003;125(6):1714–22. PubMed PMID: 14724824. Epub 2004/01/16. eng.
Marcellin P, Chang TT, Lim SG, Sievert W, Tong M, Arterburn S, et al. Long-term efficacy and safety of adefovir dipivoxil for the treatment of hepatitis B e antigen-positive chronic hepatitis B. Hepatology. 2008;48(3):750–8. PubMed PMID: 18752330. Epub 2008/08/30. eng.
Seeger C, Mason WS. Hepatitis B virus biology. Microbiol Mol Biol Rev. 2000;64(1):51–68. Pubmed Central PMCID: 98986. Epub 2000/03/08. eng.
Cummings IW, Browne JK, Salser WA, Tyler GV, Snyder RL, Smolec JM, et al. Isolation, characterization, and comparison of recombinant DNAs derived from genomes of human hepatitis B virus and woodchuck hepatitis virus. Proc Natl Acad Sci U S A. 1980;77(4):1842–6. PubMed PMID: 6246507. Pubmed Central PMCID: 348604. Epub 1980/04/01. eng.
Yan H, Zhong G, Xu G, He W, Jing Z, Gao Z, et al. Sodium taurocholate cotransporting polypeptide is a functional receptor for human hepatitis B and D virus. eLife. 2012;1:e00049.
Tuttleman JS, Pourcel C, Summers J. Formation of the pool of covalently closed circular viral DNA in hepadnavirus-infected cells. Cell. 1986;47(3):451–60. PubMed PMID: 3768961. Epub 1986/11/07. eng.
Newbold JE, Xin H, Tencza M, Sherman G, Dean J, Bowden S, et al. The covalently closed duplex form of the hepadnavirus genome exists in situ as a heterogeneous population of viral minichromosomes. J Virol. 1995;69(6):3350–7. PubMed PMID: 7745682. Pubmed Central PMCID: 189047. Epub 1995/06/01. eng.
Moolla N, Kew M, Arbuthnot P. Regulatory elements of hepatitis B virus transcription. J Viral Hepat. 2002;9(5):323–31. PubMed PMID: 12225325. Epub 2002/09/13. eng.
Kim VN, Han J, Siomi MC. Biogenesis of small RNAs in animals. Nat Rev Mol Cell Biol. 2009;10(2):126–39. PubMed PMID: 19165215. Epub 2009/01/24. eng.
Giladi H, Ketzinel-Gilad M, Rivkin L, Felig Y, Nussbaum O, Galun E. Small interfering RNA inhibits hepatitis B virus replication in mice. Mol Ther. 2003;8(5):769–76.
Hamasaki K, Nakao K, Matsumoto K, Ichikawa T, Ishikawa H, Eguchi K. Short interfering RNA-directed inhibition of hepatitis B virus replication. FEBS Lett. 2003;543(1–3):51–4. PubMed PMID: 12753904.
Klein C, Bock CT, Wedemeyer H, Wustefeld T, Locarnini S, Dienes HP, et al. Inhibition of hepatitis B virus replication in vivo by nucleoside analogues and siRNA. Gastroenterology. 2003;125(1):9–18. PubMed PMID: 12851866.
Konishi M, Wu CH, Wu GY. Inhibition of HBV replication by siRNA in a stable HBV-producing cell line. Hepatology. 2003;38(4):842–50. PubMed PMID: 14512871. Epub 2003/09/27. eng.
Qian ZK, Xuan BQ, Min TS, Xu JF, Li L, Huang WD. Cost-effective method of siRNA preparation and its application to inhibit hepatitis B virus replication in HepG2 cells. World J Gastroenterol. 2005;11(9):1297–302. PubMed PMID: 15761967. Epub 2005/03/12. eng.
Cheng TL, Chang WW, Su IJ, Lai MD, Huang W, Lei HY, et al. Therapeutic inhibition of hepatitis B virus surface antigen expression by RNA interference. Biochem Biophys Res Commun. 2005;336(3):820–30. PubMed PMID: 16153600. Epub 2005/09/13. eng.
McCaffrey AP, Nakai H, Pandey K, Huang Z, Salazar FH, Xu H, et al. Inhibition of hepatitis B virus in mice by RNA interference. Nat Biotechnol. 2003;21(6):639–44. PubMed PMID: 12740585.
Ren X, Luo G, Xie Z, Zhou L, Kong X, Xu A. Inhibition of multiple gene expression and virus replication of HBV by stable RNA interference in 2.2.15 cells. J Hepatol. 2006;44(4):663–70. PubMed PMID: 16466826.
Ren XR, Zhou LJ, Luo GB, Lin B, Xu A. Inhibition of hepatitis B virus replication in 2.2.15 cells by expressed shRNA. J Viral Hepat. 2005;12(3):236–42. PubMed PMID: 15850463.
Shin D, Kim SI, Kim M, Park M. Efficient inhibition of hepatitis B virus replication by small interfering RNAs targeted to the viral X gene in mice. Virus Res. 2006;119(2):146–53. PubMed PMID: 16443303. Epub 2006/01/31. eng.
Shlomai A, Shaul Y. Inhibition of hepatitis B virus expression and replication by RNA interference. Hepatology. 2003;37(4):764–70. PubMed PMID: 12668968.
Ying RS, Zhu C, Fan XG, Li N, Tian XF, Liu HB, et al. Hepatitis B virus is inhibited by RNA interference in cell culture and in mice. Antiviral Res. 2007;73(1):24–30. PubMed PMID: 16844238. Epub 2006/07/18. eng.
Zhang XN, Xiong W, Wang JD, Hu YW, Xiang L, Yuan ZH. siRNA-mediated inhibition of HBV replication and expression. World J Gastroenterol. 2004;10(20):2967–71. PubMed PMID: 15378775. Epub 2004/09/21. eng.
Hean J, Crowther C, Ely A, Ul Islam R, Barichievy S, Bloom K, et al. Inhibition of hepatitis B virus replication in vivo using lipoplexes containing altritol-modified antiviral siRNAs. Artifi DNA PNA XNA. 2010;1(1):17–26. PubMed PMID: 21687523. Pubmed Central PMCID: 3109439. Epub 2011/06/21. Eng.
Morrissey DV, Lockridge JA, Shaw L, Blanchard K, Jensen K, Breen W, et al. Potent and persistent in vivo anti-HBV activity of chemically modified siRNAs. Nat Biotechnol. 2005;23(8):1002–7. PubMed PMID: 16041363. Epub 2005/07/26. eng.
Marimani MD, Ely A, Buff MC, Bernhardt S, Engels JW, Arbuthnot P. Inhibition of hepatitis B virus replication in cultured cells and in vivo using 2′-O-guanidinopropyl modified siRNAs. Bioorg Med Chem. 2013;21(20):6145–55. PubMed PMID: 23743442.
Wooddell CI, Rozema DB, Hossbach M, John M, Hamilton HL, Chu Q, et al. Hepatocyte-targeted RNAi therapeutics for the treatment of chronic hepatitis B virus infection. Mol Ther. 2013;21(5):973–85. PubMed PMID: 23439496.
Carmona S, Ely A, Crowther C, Moolla N, Salazar FH, Marion PL, et al. Effective inhibition of HBV replication in vivo by anti-HBx short hairpin RNAs. Mol Ther. 2006;13(2):411–21. PubMed PMID: 16337206.
Ely A, Naidoo T, Arbuthnot P. Efficient silencing of gene expression with modular trimeric Pol II expression cassettes comprising microRNA shuttles. Nucleic Acids Res. 2009;37(13):e91. PubMed PMID: 19474340. Pubmed Central PMCID: 2715259. Epub 2009/05/29. eng.
Ely A, Naidoo T, Mufamadi S, Crowther C, Arbuthnot P. Expressed anti-HBV primary microRNA shuttles inhibit viral replication efficiently in vitro and in vivo. Mol Ther. 2008;16(6):1105–12. PubMed PMID: 18431360. Epub 2008/04/24. eng.
Grimm D, Streetz KL, Jopling CL, Storm TA, Pandey K, Davis CR, et al. Fatality in mice due to oversaturation of cellular microRNA/short hairpin RNA pathways. Nature. 2006;441(7092):537–41. PubMed PMID: 16724069.
Chen CC, Ko TM, Ma HI, Wu HL, Xiao X, Li J, et al. Long-term inhibition of hepatitis B virus in transgenic mice by double-stranded adeno-associated virus 8-delivered short hairpin RNA. Gene Ther. 2007;14(1):11–9. PubMed PMID: 16929350. Epub 2006/08/25. eng.
Chen CC, Sun CP, Ma HI, Fang CC, Wu PY, Xiao X, et al. Comparative study of anti-hepatitis B virus RNA interference by double-stranded adeno-associated virus serotypes 7, 8, and 9. Mol Ther. 2009;17(2):352–9. PubMed PMID: 19066602. Pubmed Central PMCID: 2835062. Epub 2008/12/11. eng.
Crowther C, Mowa MB, Ely A, Arbuthnot PB. Inhibition of hepatitis B virus replication in vivo using helper-dependent adenovirus vectors to deliver antiviral RNAi expression cassettes. Antivir Ther. 2014;19(4):363–73. PubMed PMID: 24296696.
Starkey JL, Chiari EF, Isom HC. Hepatitis B virus (HBV)-specific short hairpin RNA is capable of reducing the formation of HBV covalently closed circular (CCC) DNA but has no effect on established CCC DNA in vitro. J Gen Virol. 2009;90(Pt 1):115–26. PubMed PMID: 19088280. Pubmed Central PMCID: 2659548. Epub 2008/12/18. eng.
Jamieson AC, Miller JC, Pabo CO. Drug discovery with engineered zinc-finger proteins. Nat Rev Drug Discov. 2003;2(5):361–8. PubMed PMID: 12750739. Epub 2003/05/17. eng.
Klug A. The discovery of zinc fingers and their applications in gene regulation and genome manipulation. Annu Rev Biochem. 2010;79:213–31. PubMed PMID: 20192761.
Zimmerman KA, Fischer KP, Joyce MA, Tyrrell DL. Zinc finger proteins designed to specifically target duck hepatitis B virus covalently closed circular DNA inhibit viral transcription in tissue culture. J Virol. 2008;82(16):8013–21. PubMed PMID: 18524822. Pubmed Central PMCID: 2519588. Epub 2008/06/06. eng.
Bernstein KA, Rothstein R. At loose ends: resecting a double-strand break. Cell. 2009;137(5):807–10. PubMed PMID: 19490890. Epub 2009/06/06. eng.
Lieber MR. The mechanism of double-strand DNA break repair by the nonhomologous DNA end-joining pathway. Annu Rev Biochem. 2010;79:181–211. PubMed PMID: 20192759. Pubmed Central PMCID: 3079308. Epub 2010/03/03. eng.
Barker J, Voit R, Porteus M. Nuclease mediated targeted genome modification in mammalian cells. In: Renault S, Duchateau P, editors. Site-directed insertion of transgenes, Topics in current genetics, vol. 23. Netherlands: Springer; 2013. p. 327–52.
Wirt SE, Porteus MH. Development of nuclease-mediated site-specific genome modification. Curr Opin Immunol. 2012;24(5):609–16. PubMed PMID: 22981684.
Schiffer JT, Aubert M, Weber ND, Mintzer E, Stone D, Jerome KR. Targeted DNA mutagenesis for the cure of chronic viral infections. J Virol. 2012;86(17):8920–36. PubMed PMID: 22718830. Pubmed Central PMCID: 3416169. Epub 2012/06/22. eng.
Cong L, Ran FA, Cox D, Lin S, Barretto R, Habib N, et al. Multiplex genome engineering using CRISPR/Cas systems. Science. 2013;339(6121):819–23. PubMed PMID: 23287718. Epub 2013/01/05. eng.
Hwang WY, Fu Y, Reyon D, Maeder ML, Tsai SQ, Sander JD, et al. Efficient genome editing in zebrafish using a CRISPR-Cas system. Nat Biotechnol. 2013;31(3):227–9. PubMed PMID: 23360964. Pubmed Central PMCID: 3686313.
Jacquier A, Dujon B. An intron-encoded protein is active in a gene conversion process that spreads an intron into a mitochondrial gene. Cell. 1985;41(2):383–94. PubMed PMID: 3886163. Epub 1985/06/01. eng.
Kostriken R, Strathern JN, Klar AJ, Hicks JB, Heffron F. A site-specific endonuclease essential for mating-type switching in Saccharomyces cerevisiae. Cell. 1983;35(1):167–74. PubMed PMID: 6313222. Epub 1983/11/01. eng.
Thierry A, Dujon B. Nested chromosomal fragmentation in yeast using the meganuclease I-Sce I: a new method for physical mapping of eukaryotic genomes. Nucleic Acids Res. 1992;20(21):5625–31. PubMed PMID: 1333585. Pubmed Central PMCID: 334395. Epub 1992/11/11. eng.
Grosse S, Huot N, Mahiet C, Arnould S, Barradeau S, Clerre DL, et al. Meganuclease-mediated inhibition of HSV1 infection in cultured cells. Mol Ther. 2011;19(4):694–702. PubMed PMID: 21224832. Pubmed Central PMCID: 3070101. Epub 2011/01/13. eng.
Kim YG, Cha J, Chandrasegaran S. Hybrid restriction enzymes: zinc finger fusions to Fok I cleavage domain. Proc Natl Acad Sci U S A. 1996;93(3):1156–60. PubMed PMID: 8577732. Pubmed Central PMCID: 40048. Epub 1996/02/06. eng.
Christian M, Cermak T, Doyle EL, Schmidt C, Zhang F, Hummel A, et al. Targeting DNA double-strand breaks with TAL effector nucleases. Genetics. 2010;186(2):757–61. PubMed PMID: 20660643. Pubmed Central PMCID: 2942870Pubmed Central PMCID: 2942870. Epub 2010/07/28. eng.
Bitinaite J, Wah DA, Aggarwal AK, Schildkraut I. FokI dimerization is required for DNA cleavage. Proc Natl Acad Sci U S A. 1998;95(18):10570–5. PubMed PMID: 9724744. Pubmed Central PMCID: 27935. Epub 1998/09/02. eng.
Wah DA, Hirsch JA, Dorner LF, Schildkraut I, Aggarwal AK. Structure of the multimodular endonuclease FokI bound to DNA. Nature. 1997;388(6637):97–100. PubMed PMID: 92145100. Epub 1997/07/03. eng.
Porteus MH. Mammalian gene targeting with designed zinc finger nucleases. Mol Ther. 2006;13(2):438–46. PubMed PMID: 16169774. Epub 2005/09/20. eng.
Rahman SH, Maeder ML, Joung JK, Cathomen T. Zinc-finger nucleases for somatic gene therapy: the next frontier. Hum Gene Ther. 2011;22(8):925–33. PubMed PMID: 21631241. Pubmed Central PMCID: 3159524. Epub 2011/06/03. eng.
Cradick TJ, Keck K, Bradshaw S, Jamieson AC, McCaffrey AP. Zinc-finger nucleases as a novel therapeutic strategy for targeting hepatitis B virus DNAs. Mol Ther. 2010;18(5):947–54. PubMed PMID: 20160705. Pubmed Central PMCID: 2890117. Epub 2010/02/18. eng.
Boch J, Bonas U. Xanthomonas AvrBs3 family-type III effectors: discovery and function. Annu Rev Phytopathol. 2009;48:419–36. PubMed PMID: 19400638. Epub 2009/04/30. eng.
Ramirez CL, Foley JE, Wright DA, Muller-Lerch F, Rahman SH, Cornu TI, et al. Unexpected failure rates for modular assembly of engineered zinc fingers. Nat Methods. 2008;5(5):374–5. PubMed PMID: 18446154. Epub 2008/05/01. eng.
Sander JD, Dahlborg EJ, Goodwin MJ, Cade L, Zhang F, Cifuentes D, et al. Selection-free zinc-finger-nuclease engineering by context-dependent assembly (CoDA). Nat Methods. 2010;8(1):67–9. PubMed PMID: 21151135. Pubmed Central PMCID: 3018472. Epub 2010/12/15. eng.
Boch J, Scholze H, Schornack S, Landgraf A, Hahn S, Kay S, et al. Breaking the code of DNA binding specificity of TAL-type III effectors. Science. 2009;326(5959):1509–12. PubMed PMID: 19933107. Epub 2009/11/26. eng.
Moscou MJ, Bogdanove AJ. A simple cipher governs DNA recognition by TAL effectors. Science. 2009;326(5959):1501. PubMed PMID: 19933106. Epub 2009/11/26. eng.
Joung JK, Sander JD. TALENs: a widely applicable technology for targeted genome editing. Nat Rev Mol Cell Biol. 2013;14(1):49–55. PubMed PMID: 23169466. Pubmed Central PMCID: 3547402.
Mak AN, Bradley P, Cernadas RA, Bogdanove AJ, Stoddard BL. The crystal structure of TAL effector PthXo1 bound to its DNA target. Science. 2012;335(6069):716–9. PubMed PMID: 22223736. Pubmed Central PMCID: 3427646. Epub 2012/01/10. eng.
Mussolino C, Morbitzer R, Lutge F, Dannemann N, Lahaye T, Cathomen T. A novel TALE nuclease scaffold enables high genome editing activity in combination with low toxicity. Nucleic Acids Res. 2011;39(21):9283–93. PubMed PMID: 21813459. Pubmed Central PMCID: 3241638. Epub 2011/08/05. eng.
Hockemeyer D, Wang H, Kiani S, Lai CS, Gao Q, Cassady JP, et al. Genetic engineering of human pluripotent cells using TALE nucleases. Nat Biotechnol. 2011;29(8):731–4. PubMed PMID: 21738127. Pubmed Central PMCID: 3152587. Epub 2011/07/09. eng.
Li T, Huang S, Zhao X, Wright DA, Carpenter S, Spalding MH, et al. Modularly assembled designer TAL effector nucleases for targeted gene knockout and gene replacement in eukaryotes. Nucleic Acids Res. 2011;39(14):6315–25. PubMed PMID: 21459844. Pubmed Central PMCID: 3152341. Epub 2011/04/05. eng.
Ding Q, Lee YK, Schaefer EA, Peters DT, Veres A, Kim K, et al. A TALEN genome-editing system for generating human stem cell-based disease models. Cell Stem Cell. 2013;12(2):238–51. PubMed PMID: 23246482. Pubmed Central PMCID: 3570604. Epub 2012/12/19. eng.
Bloom K, Ely A, Mussolino C, Cathomen T, Arbuthnot P. Inactivation of hepatitis B virus replication in cultured cells and in vivo with engineered transcription activator-like effector nucleases. Mol Ther. 2013;21(10):1889–97. PubMed PMID: 23883864. Pubmed Central PMCID: 3808145. Epub 2013/07/26. eng.
Chen J, Zhang W, Lin J, Wang F, Wu M, Chen C, et al. An efficient antiviral strategy for targeting hepatitis B virus genome using transcription activator-like effector nucleases. Mol Ther. 2013;12. PubMed PMID: 24025750. Epub 2013/09/13. Eng.
Roos WP, Kaina B. DNA damage-induced cell death by apoptosis. Trends Mol Med. 2006;12(9):440–50. PubMed PMID: 16899408. Epub 2006/08/11. eng.
Bree RT, Neary C, Samali A, Lowndes NF. The switch from survival responses to apoptosis after chromosomal breaks. DNA Repair. 2004;3(8–9):989–95. PubMed PMID: 15279785. Epub 2004/07/29. eng.
Guidotti LG, Matzke B, Schaller H, Chisari FV. High-level hepatitis B virus replication in transgenic mice. J Virol. 1995;69(10):6158–69. PubMed PMID: 7666518. Pubmed Central PMCID: 189513. Epub 1995/10/01. eng.
Kim Y, Kweon J, Kim JS. TALENs and ZFNs are associated with different mutation signatures. Nat Methods. 2013;10(3):185. PubMed PMID: 23396284. Epub 2013/02/12. eng.
Buttner D, Bonas U. Regulation and secretion of Xanthomonas virulence factors. FEMS Microbiol Rev. 2010;34(2):107–33. PubMed PMID: 19925633. Epub 2009/11/21. eng.
Kay S, Hahn S, Marois E, Hause G, Bonas U. A bacterial effector acts as a plant transcription factor and induces a cell size regulator. Science. 2007;318(5850):648–51. PubMed PMID: 17962565. Epub 2007/10/27. eng.
Romer P, Hahn S, Jordan T, Strauss T, Bonas U, Lahaye T. Plant pathogen recognition mediated by promoter activation of the pepper Bs3 resistance gene. Science. 2007;318(5850):645–8. PubMed PMID: 17962564. Epub 2007/10/27. eng.
Zhang F, Cong L, Lodato S, Kosuri S, Church GM, Arlotta P. Efficient construction of sequence-specific TAL effectors for modulating mammalian transcription. Nat Biotechnol. 2011;29(2):149–53. PubMed PMID: 21248753. Pubmed Central PMCID: 3084533. Epub 2011/01/21. eng.
Cong L, Zhou R, Kuo YC, Cunniff M, Zhang F. Comprehensive interrogation of natural TALE DNA-binding modules and transcriptional repressor domains. Nat Commun. 2012;3:968. PubMed PMID: 22828628. Pubmed Central PMCID: 3556390. Epub 2012/07/26. eng.
Garg A, Lohmueller JJ, Silver PA, Armel TZ. Engineering synthetic TAL effectors with orthogonal target sites. Nucleic Acids Res. 2012;40(15):7584–95. PubMed PMID: 22581776. Pubmed Central PMCID: 3424557. Epub 2012/05/15. eng.
Geissler R, Scholze H, Hahn S, Streubel J, Bonas U, Behrens SE, et al. Transcriptional activators of human genes with programmable DNA-specificity. PLoS One. 2011;6(5):e19509. PubMed PMID: 21625585. Pubmed Central PMCID: 3098229. Epub 2011/06/01. eng.
Bellefroid EJ, Poncelet DA, Lecocq PJ, Revelant O, Martial JA. The evolutionarily conserved Kruppel-associated box domain defines a subfamily of eukaryotic multifingered proteins. Proc Natl Acad Sci U S A. 1991;88(9):3608–12. Pubmed Central PMCID: 51501. Epub 1991/05/01. eng.
Schultz DC, Ayyanathan K, Negorev D, Maul GG, Rauscher 3rd FJ. SETDB1: a novel KAP-1-associated histone H3, lysine 9-specific methyltransferase that contributes to HP1-mediated silencing of euchromatic genes by KRAB zinc-finger proteins. Genes Dev. 2002;16(8):919–32. PubMed PMID: 11959841. Pubmed Central PMCID: 152359. Epub 2002/04/18. eng.
Quasdorff M, Protzer U. Control of hepatitis B virus at the level of transcription. J Viral Hepat. 2010;17(8):527–36. PubMed PMID: 20546497. Epub 2010/06/16. eng.
Bar-Yishay I, Shaul Y, Shlomai A. Hepatocyte metabolic signalling pathways and regulation of hepatitis B virus expression. Liver Int. 2011;31(3):282–90. PubMed PMID: 21281428. Epub 2011/02/02. eng.
Fukai K, Takada S, Yokosuka O, Saisho H, Omata M, Koike K. Characterization of a specific region in the hepatitis B virus enhancer I for the efficient expression of X gene in the hepatic cell. Virology. 1997;236(2):279–87. PubMed PMID: 9325235. Epub 1997/11/05. eng.
Su H, Yee JK. Regulation of hepatitis B virus gene expression by its two enhancers. Proc Natl Acad Sci U S A. 1992;89(7):2708–12. PubMed PMID: 1313564. Pubmed Central PMCID: 48731. Epub 1992/04/01. eng.
Richardson C, Jasin M. Frequent chromosomal translocations induced by DNA double-strand breaks. Nature. 2000;405(6787):697–700. PubMed PMID: 10864328. Epub 2000/06/23. eng.
Perrillo R. Benefits and risks of interferon therapy for hepatitis B. Hepatology. 2009;49(5 Suppl):S103–11. PubMed PMID: 19399806.
Acknowledgements
Work in the authors’ laboratory is generously supported by funding from the National Research Foundation (NRF, GUNs 81768, 81692, 68339, 85981 & 77954) of South Africa, South African Medical Research Council (MRC), Poliomyelitis Research Foundation (PRF) and from the German Research Foundation (DFG).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 American Society of Gene and Cell Therapy
About this chapter
Cite this chapter
Bloom, K., Ely, A., Arbuthnot, P. (2015). Recent Advances in Use of Gene Therapy to Treat Hepatitis B Virus Infection. In: Berkhout, B., Ertl, H., Weinberg, M. (eds) Gene Therapy for HIV and Chronic Infections. Advances in Experimental Medicine and Biology(), vol 848. Springer, New York, NY. https://doi.org/10.1007/978-1-4939-2432-5_2
Download citation
DOI: https://doi.org/10.1007/978-1-4939-2432-5_2
Published:
Publisher Name: Springer, New York, NY
Print ISBN: 978-1-4939-2431-8
Online ISBN: 978-1-4939-2432-5
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)