Abstract
In this book, the role of many different proteins, pathways, or genetic events in affecting the function of leukemic cells, as well as their suitability for therapeutic modulation, is discussed. However, since acute myeloid leukemia (AML) is a very heterogeneous disease, it will be necessary to match the characteristics of individual patients to the selection of the right targeted agent in order to maximize efficacy. In this chapter, we discuss how the application of the numerous molecular profiling arrays that are now available can be utilized to provide the answer to this dilemma. The development of methods to combine data from multiple arrays, termed “integromics” will enable a more sophisticated application of “omic” data that will facilitate the selection of combinations of targeted agents on an individualized basis.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Omics
- Integromics
- Gene expression profiling (GEP)
- Whole genome sequencing
- Single nucleotide polymorphism (SNP)
- Reverse phase protein array (RPPA)
- Proteomics
2.1 Introduction
A central theme developed in the prior chapter is that acute myeloid leukemia (AML) is a heterogeneous disease with myriad changes occurring at multiple different levels that combine to determine the biology of the leukemic cell. In the chapters that follow, each author will discuss how a single gene (e.g., AKT, P53), or pathway (e.g., Ras/RAF/MEK/ERK), or genetic modification (e.g., methylation) affects AML cell biology and how that individual event might be targeted to improve therapy. The heterogeneous nature of AML suggests that each of these potential targets will be therapeutically efficacious, but in only a fraction of cases of AML. If we as a scientific community are successful at discovering the key points of attack and actually develop functional therapies for each of these, how will we determine when to use which? Furthermore, redundancy in pathways and mechanisms within a cell may require that targeted therapies be used in combination to achieve efficacy. Given the complexity of AML, how will we determine which agents to use in combination? In which cases? By what schedule and in what order? There is a very high risk that promising agents will be discarded as therapeutic failures as a result of faulty study design and the barriers erected by regulatory and proprietary interests. A drug may lack efficacy as single agent, but be of great benefit when used in combination with other chemotherapeutic agents. Current regulation makes it difficult to combine two non-approved agents that are currently in phase I–II evaluation into a single trial, and pharmaceutical companies are highly reluctant to pair a developmental molecule with one from another company. Consequently, combination trials of unapproved agents are nearly impossible to conduct. A second problem is that the typical phase I–II trial typically treats about 15 patients. If the frequency of the population that is susceptible to a given agent is low, then there is a high chance that no patients in a trial of that small size will have the characteristic (for event frequencies of 3, 5, 8, and 10 %, then 63, 46, 29, and 21 % of trials of n = 15 would have no patients with that characteristic). The activity of a highly efficacious agent can easily be missed if the correct patients do not make it onto the right trial. The successful translation of the discoveries discussed in the rest of this book into effective therapy that improves the outcome of AML is therefore critically dependent on having the ability to match patients on an individualized basis to the correct therapy.
In this chapter, we discuss how the various “omics” (Weinstein 2002) might be used to provide this missing information, to enable us to match patients to therapy, and to rationally combine different targeted therapies (Barretina et al. 2012) from a broad menu into a restricted list for an individual patient. High-throughput array-based technologies have been developed that enable the global characterization of multiple characteristics of a cell, enabling the collection of historically unprecedented amount of information. For virtually every aspect of a cell in which abnormalities could arise, an array now exists for its evaluation. The basic dogma of biology formerly was DNA → RNA → protein, with mutations leading to changes in the sequence or expression level of mRNA, resulting in quantitative or qualitative changes in protein function. But in light of the effects of epigenetics and noncoding RNAs on gene expression, this view is now overly simplistic.
2.2 Types of Alterations That Can Affect Leukemia Biology and Methods Available to Evaluate Them (Table 2.1)
2.2.1 Nucleic Acids
Mutations are theoretically the most stable form of change. A gene is mutated or not, and once a mutation develops, it should be passed on to all progeny. The location in a regulatory region can lead to over- or under-expression of the mRNA. Likewise, copy number abnormalities can lead to overexpression of an unmutated, oncogenic gene. A mutation in the coding region can result in a stop codon, or frameshift mutation, resulting in loss or gain of function, or result in an amino acid change that changes the function and regulation of the protein. Translocations can result in novel proteins with oncogenic function. Additional complexity can result from mutations in epigenetic regulators or affecting transcription factors, and by changes in splicing, for which the causative changes may not be known. Each of these can be determined using the currently available technologies, with each base pair analyzed in whole genome sequencing , or just known coding regions assayed in whole exome sequencing, while sequencing of those genes that are actually transcribed can be performed by RNA-seq (also called whole transcriptome shotgun sequencing). In addition to detecting mutations, the relative frequency of the detected fragments provides copy number data as well. With prices rapidly decreasing, it may soon be feasible, from a technical and economic standpoint, to sequence each individual patient’s tumor as the standard of care. However, among the important caveats is that the interpretation of such massive sequencing data remains challenging. For every “actionable” or “driver” mutation with a currently known prognostic and potentially therapeutic implication in AML, such as ones affecting FLT3, there may be many more non-synonymous differences from the reference genome or patient’s germ line sequence that could be oncogenic in unknown ways, “passenger” mutations due to genomic instability frequently present in tumors, or single-nucleotide polymorphisms (SNPs). Furthermore, sequencing is subject to technical and sampling variation. Two recent studies demonstrated that there is clonal heterogeneity within both solid tumors (Gerlinger et al. 2012) and myelodysplastic syndrome (MDS) and AML (Walter et al. 2012), suggesting that deep sequencing of multiple sites or samples may be necessary to fully recognize the spectrum of mutations within a patient.
The genome can be altered in ways that affect gene transcription without mutation. SNPs can regulate the level of expression of a gene, or the function of the resulting protein (e.g., SNP309 in MDM2, Pichiorri et al. 2010; Phillips et al. 2010). Arrays exist that can measure 250,000–600,000 to over a million SNPs. Likewise, DNA can be methylated, and methylation arrays have been developed to assess the status of methylation across the genome. The interaction of DNA with histones can also regulate the availability of the gene to transcription through histone methylation (Butler et al. 2012), acetylation, phosphorylation, and sumoylation. The first two can be assessed by genome-wide location analyses using chromatin immunoprecipitation linked to cDNA arrays or to high-throughput sequencing (Miao and Natarajan 2005) although there are not commercial kits for this readily available. Noncoding RNAs, also known as microRNAs (miRNAs), are small strands of RNAs that can bind to DNA to regulate transcription or to mRNA to regulate translation and mRNA longevity. They are subject to mutation, and variation in their levels has been associated with many malignancies (Garzon et al. 2009; Calin et al. 2004; Calin et al. 2005; Calin et al. 2007; Lee et al. 2012; Taby et al. 2009; Valastyan et al. 2011). They might also function in complex networks with pseudogenes to regulate protein levels (Salmena et al. 2011). Mutations in ncRNA can be detected by whole genome sequencing, and expression levels can be readily ascertained via the many commercially available miRNA arrays. This technology is in rapid flux as the number of ncRNA covered is rapidly increasing. Measurement of global gene expression at the mRNA level has become commonplace with two platforms, Affymetrix and Illumina, currently used widely. Gene expression profiling (GEP) is often thought of as a “detector” of causal changes, in which the changes in gene expression are themselves causative of a phenotype. However, the “unbiased,” genome-wide nature of GEP makes it good as a “reflector” of causal changes, in which the changes in gene expression are a downstream readout of changes, such as the activity of a signaling pathway, that are ultimately causative of a phenotype. The changes in gene expression may contribute to the phenotype, but that is not necessarily understood, nor does it need to be, when GEP is used as a detector. This “molecular pattern recognition” has a lot of clinical potential, and is in fact the way in which GEP is finally being Food and Drug Administration (FDA)-approved for clinical use, in breast cancer and myeloma. While many insights into cancer biology have emerged from GEP studies, these results are somewhat hampered by a lack of reproducibility from one dataset to the next (Xu et al. 2008; Michiels et al. 2005) and by the lack of correlation between mRNA expression and protein expression (Gygi et al. 1999; Griffin et al. 2002; Washburn et al. 2003; Greenbaum et al. 2003; Jansen et al. 2002; Kern et al. 2003).
2.2.2 Stroma, Chemokines, and Cytokines
External influences from chemokines and cytokines as well as from interaction with stroma can influence gene expression and protein activation. Multiplex kits are available to measure from 20 to 75 different chemokines and cytokines at once from very small quantities of plasma. In an analysis of the expression of 27 cytokines and chemokines in the serum from AML and MDS patients, we demonstrated that both of these diseases are characterized by recurrent patterns of expression, and surprisingly, these carried prognostic significance (Kornblau et al. 2010a). We have recently generated a reverse phase protein array (RPPA; see below) using mesenchymal stromal cells (MSC) derived from AML patients to define the characteristics of the stroma and determine how this correlates with characteristics of the AML blast.
2.2.3 Protein
Proteins are the predominant effectors of most genes (ribosomes, tRNA, etc., aside), and the net effect of mutations, DNA methylation, histone modifications, ncRNA and external influences, determines the level of expression of individual proteins. But since protein activity is regulated by post-translational modifications, it is not only sufficient to ascertain the protein expression level, but also the activation state must be known as well. The potential for splice variants and posttranslational modifications, including phosphorylation, glycosylation, ubiquitination, cleavage, etc., raise the number of different members of the proteome from the 20,000–40,000 known genes into more than a million potential variants. This complexity has made proteomics the least developed of the array technologies even though it is likely the most critical. Traditional proteomics involves two-dimensional (2-D) gel electrophoresis based on charge and mass, followed by tandem mass spectrometry (so-called 2D-MALDI-TOF). While the entire proteome can be analyzed, this technology is currently expensive, and impractical for individualized patient analysis as the amount of starting material required is high and the throughput low. There are alternatives to mass spectrometry-based proteomics.
2.2.4 Proteomics Methodologies
One approach, in commercial development (SOMA, Colorado), uses DNA-based aptamers, selected for their ability to specifically bind to conformed protein, with a tag that allows for quantitative detection (Gold et al. 2010). While phospho-specific aptamers could theoretically be developed, thus far the company has only developed aptamers to measure total protein levels. A second related approach, known as proximity ligation, uses two antibodies with overlapping tags that bind to each other enabling amplification to permit detection, in an enzyme-linked immunosorbent assay (ELISA)-like approach (Fredriksson et al. 2002). The other methodology uses an antibody -based proteomic approach. Two types of these exist. In forward phase protein arrays (FPPA), “capture” antibodies can be printed on an array in known positions, and these then bind proteins of interest which are then sandwiched by the binding of a second antibody with some form of detection capability. The difficulty of finding two structurally nonoverlapping, highly specific antibodies for each target have prevented this from generalized use. In the second type of antibody array, called RPPA, the protein from hundreds of patients are printed on the array and each slide is probed with a single highly validated antibody. Since only one antibody is utilized at a time, with proper validation, nonspecific binding can be minimized. The drawback of this technology is that samples must be batched to be printed at once, making this methodology impractical for real-time individual patient analysis. A second shortcoming is that it does not give data on all proteins and isoforms (e.g., it is biased as opposed to GEP that are unbiased for all potential open reading frames), and furthermore, only those targets with a validated antibody can be probed. On the plus side, the methodology is sample sparing, requiring only 50,000 cells to test about 150 different antibodies. Our laboratory has optimized the technology to do this in hematological malignancies and has generated arrays for AML, MDS, acute lymphoblastic leukemia (ALL) and with AML and MDS CD34+ and CD34+CD38− stem cell-enriched fractions, and recently with AML patient-derived MSC (Kornblau et al. 2009c; Tibes et al. 2006; Neeley et al. 2012; Quintas-Cardama et al. 2012; Kornblau et al. 2011a; Ruvolo et al. 2011a; Ruvolo et al. 2011b; Starkova et al. 2011; Kornblau et al. 2011b; Ravandi et al. 2011; Levin et al. 2010; Kornblau et al. 2010b; Kornblau et al. 2010c; Tsao et al. 2010; Ma et al. 2010a; Carter et al. 2010; Neeley et al. 2009; Poland et al. 2009; Pemmaraju et al. 2009; Lee et al. 2012; Minden et al. 2009; Kadia et al. 2009; Carter et al. 2009; Kornblau et al. 2009a; Kornblau et al. 2009b; Kornblau et al. 2013). Newer technologies like CyTOF (Bendall et al. 2011) offer higher throughput and individual cell analysis with a flow cytometry-like output, but the number of validated antibodies is currently limited. This technology will likely eventually be able to provide real-time throughput that RPPA cannot provide .
2.2.5 Metabolomics
Additional technologies to enable detailed profiling of the relative concentration of metabolic pathway components, including lipids, are also emerging . Ascertaining the levels of these would provide another level of validation that the changes observed at different levels are culminating in the expected changes. The unanticipated discovery that the excess 2-OH-glutarate resulting from isocitrate dehydrogenase (IDH) mutations (Lu et al. 2012) results in alteration of methylation highlights that the cell-wide consequences of a mutation extend beyond the canonical pathway that a gene is “contained” in .
2.2.6 Assessing Dynamic Change
A final consideration is expression in the setting of perturbation. The dynamic response of the cells to stress, such as chemotherapy, or apoptotic inducers, etc., can be a key to whether a cell is responsive or resistant. The dynamic adaptation of leukemic cells has been shown to relate to sensitivity. For example, changes in cholesterol synthesis and uptake correlate with sensitivity to ara-c and anthracyclines, (Banker et al. 2004) and blockade of this adaptation was shown to improve chemosensitivity in a clinical trial (Kornblau et al. 2007). Methods to assess adaptability are complex and still in development. The effect of perturbation with exogenous ligands to cell surface receptors, apoptosis inducers, chemotherapeutic agents on the expression and activation of various pathways, and the effect of this on cell viability has been measured using multiparameter flow cytometry as developed by The Nolan Lab (Krutzik et al. 2004; Irish et al. 2004). Identifying the key parameters to perturb and the critical outcomes to measure remains a daunting task. The development of this technology for common use is currently being pursued by Nodality Inc. In collaboration with Nodality, our group has conducted research aimed at the identification of certain stimulation-readout nodes (Kornblau et al. 2011a; Rosen et al. 2010; Cesano et al. 2009; Kornblau et al. 2009b). We also observed that the expression of cells with responsiveness in the Stat6 pathway in subfractions at diagnosis is predictive of relapse (Kornblau et al. 2011a). With further development, omics approaches for dynamic adaptability hold additional promise for improved identification of key targets on an individualized basis.
2.3 Integromics
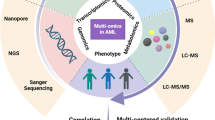
The goal, once the information from each of these arrays has been performed, is to integrate information from many levels into a single dataset reflecting what is happening in a particular patient’s leukemic cells . This process has been named “integromics” (Fig. 2.1). Once the integrated dataset is generated, then the effects of the different changes on a particular pathway can be determined. Expression of components of a pathway might be affected by different processes that combine to result in a “net” effect on a downstream effector that determines activity. For example, a cell might have an activating rat sarcoma (RAS) mutation, which would lead to expectation that the downstream pathway would be active, but lack mitogen extracellular signal-regulated kinase (MEK) and extracellular signal-regulated kinase (ERK) activity due to the combined action of histone acetylation preventing MEK expression, gene methylation preventing ERK expression, and the overexpression of ncRNA resulting in negligible expression of an ERK target. Conversely, small effects at many levels might combine to lead to highly significant overexpression of ERK protein and cross-activation by a kinase with ERK phosphorylation activity other than MEK. In the first case, detecting a RAS mutation might lead us to consider use of a RAS, MEK, or ERK inhibitor, but such a patient would be unlikely to benefit as all the components downstream of the RAS mutation are already under-expressed. Conversely, in the second example, an ERK inhibitor might be therapeutically highly effective, but simply searching for RAS mutations would not have selected this patient for such a trial. So the multilevel information that can be gained from an integromic approach will be key to applying omics to the direction of targeted therapy . The integration of datasets is not a trivial computational matter, and there are many technical considerations that must be addressed if this is to be successfully accomplished .
Integromics. The goal is to understand how much function there is of a given protein (gene1) and what factors regulate that. The assays that can currently go into an integromics analysis and the level they measure are shown at the left side. The combined effects of DNA methylation, Histone modifications, SNPs and non-coding RNA can have direct effects on the transcription of the gene in question. Likewise they can affect the level of expression of other gene(s) that modulate the expression of gene 1 and thereby have an indirect effect. This sets the rheostat for how much gene 1 mRNA is produced. Non-coding RNAs can also modulate gene 1 mRNA fate, affecting translation along with other cell characteristics that affect the rate of translation to set the rheostat for how much total protein exists. Post translational modification, which is regulated by direct or indirect effects of internal and external factors can affect the activation state of protein 1 thereby setting the rheostat for protein 1 function
2.3.1 Binary Outcomes Versus Quantitative Measurements
Regardless of the material that is being measured (DNA, RNA, protein miRNA, etc.), there are certain statistical analytic challenges that must be met. The first is a determination of whether detection is adequate or whether relative quantity is needed. In the case of mutations, simply detecting the presence or absence of an event provides one level of answer, but given the heterogeneity of tumor tissues, it may be important to also quantify what percentage of a tumor carries the mutation. For most other analytes, such as mRNA, miRNA, methylation events, protein quantity, etc., relative quantification of the level of expression is required. While the complete absence of expression of a given analyte would provide important information, for many of them it is the relative change in expression level that will determine the functional consequences, and it must be remembered that small coordinated changes in a pathway can be cumulatively important, even if the changes in individual genes do not rise to the level of statistical significance. GEP typically uses a cutoff of threefold expression differences between two samples to define a significant difference. While such cutoffs have the potential to provide higher signal-to-noise ratios, they introduce the potential to miss important changes. Consider the interaction of the pro- and antiapoptotic proteins Bcl2 and Bax, where the relative concentration of each is thought to regulate the apoptosis rheostat. A twofold increase in one accompanied by a twofold decrease in the other would have both being listed as not significantly different, although the relative ratio change would be 4:1 and would likely have a functional consequence. Similarly, if both increased by fourfold, they would be included in the list of significantly different proteins, yet the ratio of each to the other would remain unchanged, and there might be no functional consequences despite a large change in expression.
One approach to generating a more systematic network-based approach to the analysis of GEP data is to evaluate multiple genes simultaneously in sets or signatures linked to some biological feature, such as a signaling pathway. Many gene sets have been prepared in collections that are available online for application to gene expression data, such as Gene Ontology (GO), (Ma et al. 2009) the Molecular Signatures Database (MSigDB), (Subramanian et al. 2005) and the Kyoto Encyclopedia of Genes and Genomes (KEGG) (Ogata et al. 1999). One frequently used approach, so-called gene set enrichment analysis (GSEA), (Gerlinger et al. 2012) has the advantage of not being dependent on the identification of differentially expressed genes (DEGs), according to some arbitrary threshold (fold change, statistical test, etc.). GSEA determines whether the overall expression of a set of genes associated with a particular biological feature differs significantly between two groups of samples, i.e., whether the feature is enriched, even if few or even none of the differences in individual genes meets the threshold of a DEG. Gene set analysis can also be based on DEGs, using the hypergeometric distribution test to determine the significance of enrichment (Subramanian et al. 2005), as in the popular proprietary application Ingenuity Pathway Analysis. While some caveats exist, gene set analysis has been a major advance in the analysis of gene expression data; investigators can now do more than stare at a heat map or list of DEGs and hope for inspiration. However, experience shows that gene set analysis typically only “explains” a small proportion of the differences in gene expression between two groups of samples. Gene set analysis does not eliminate the need for thoughtful interpretation and validation of gene expression data. GSEA serves as a paradigm for how a more integrated analysis can improve the strength of the data emerging from array approaches.
2.3.2 Defining the “Normal” Comparator
For each analyte with a quantitative expression measurement, the first question is whether expression is changed from normal. However, defining what is normal is complicated. What is the “normal” comparator for an AML blast, is this a CD33+ cell? A CD34+ cell? A CD34+ CD38− cell? Normal hematopoietic stem cell (HSC) and leukemia stem cell (LSC) tend to be CD90−/CD123+/CD117−/HLA-DR− and also express CLL-1, CD44, CD47, and CD96 (Chan and Huntley 2008; Roboz and Guzman 2009; Blair et al. 1998; Blair and Sutherland 2000; Jordan et al. 2000). The restriction of LSC to this phenotype is controversial as some studies have demonstrated LIC with specific molecular abnormalities (del 5q) (Tehranchi et al. 2010) in this pool while other have found persistence of cells harboring specific mutations (NPM1) in the CD34− fractions (Taussig et al. 2010). Ideally, a study of normal HSC might use CD34+ CD38− cells, but these cannot be obtained in sufficient numbers even for RPPA . Normal HSC are much rarer than LSC, and it was not possible to purchase sufficient normal CD34+ cells to isolate 2 × 105 normal HSC from available. If normal CD34+ cells are 5 % of normal bone marrow and normal HSC comprise 1:10,000 marrow cells, then there would be 1 HSC per 500 CD34+. Assuming a 50 % recovery, this would require 1000 CD34+ cells to recover a single HSC or that we start with 2 × 108 CD34+ cells (= 20 vials × $750/vial = $15,000/sample), enough to engraft a 50 kg patient to recover 2 × 105 cells. If the normal HSC frequency is 1:100,000, then multiply this by 10. For our protein work, we have therefore used a CD34+ cell from a normal bone marrow as our standard comparator (we have successfully used CD34+ cells from AllCells for our RPPA work). We also evaluated granulocyte colony-stimulating factor (G-CSF)-primed peripheral blood derived CD34+ cells by RPPA. As expected, the expression of many proteins was markedly different upon G-CSF stimulation. We have demonstrated that protein expression is markedly different between bulk leukemia cells, CD34+ cells, or cells from the stem cell-enriched CD34+ CD38− fraction (Kornblau et al. 2013). If the cell that gives rise to resistance comes predominantly from one compartment, then the biology of that compartment needs to be targeted, and it is important to know what is normal for the non-leukemic counterpart from that compartment.
2.3.3 Effect of Sample Source
AML samples can come from the blood or the marrow. Does expression change by the origin of the sample? For DNA and epigenetic events, location should not affect the readout, but for quantitative analytes, this could be changed. In our RPPA analyses, we did not observe a statistically significant difference in expression between blood and marrow samples for 85 % of proteins measured, and for most of these, the fold change was less than 20 %, so, therefore, of uncertain functional consequence. However, real differences could have been missed due to dilution of “marrow” samples with blood as our research collection sample is the last aspirate added on at the end of a diagnostic study. If resistance arises from cells in particular environments, such as adjacent to stroma , or in areas of hypoxia, then the key feature may be present in only a small fraction of cells, and it may be important to isolate specific subsets of cells and to maintain those cells in the native condition until the labile fraction can be isolated and stabilized.
2.3.4 Use of Statistical Versus Biologically Functional Thresholds
Another crucial question, that is very difficult to answer, is what is the correlation between the difference in expression and a biologically functional difference? What level correlates to a functional difference? Is this a level that reaches statistical significance by standard methods, or is it a level that is much greater than the threshold for statistical significance? Rigorous testing of each change that appears significantly different by an array to determine the actual functional difference, using other methods for altering level (e.g., siRNA, knock in, knock out), would be costly and time consuming. The level that gives a survival difference on a Kaplan–Meier survival plot may be unrelated to the level that can be pharmacologically modified.
2.4 Issues in Applying Integromics to Patient Care
Despite these obvious potentially confounding issues (and in ignorance of the nonobvious ones), the integromic dataset has the potential to guide the intelligent selection of targeted therapies. Let us consider how having this knowledge might be applied in different circumstances .
2.4.1 Data for Individual Proteins
For an agent targeting one molecule, the first question is whether that molecule is expressed in that patient’s leukemic cells . As mutations are recognized and studied, we collectively develop a catalog of the functional consequences of that mutation. Is it activating? Does it eliminate kinase activity, etc.? For non-mutational events, quantity is relevant to therapeutic targeting. Intuitively, a target that is not expressed would not be expected to respond to an agent that inhibits its function. However, if dynamic studies show that the target is significantly upregulated as a defensive adaptation in response to exposure to a therapeutic agent, then even a target with low expression at baseline may be targetable. Both the individual arrays and the integromic approach, especially if dynamic measures are included, can therefore suggest if an individual marker is a candidate.
2.4.2 Data from Multiple Proteins
A stronger level of evidence that a pathway is a good target would come from evidence that components up and down a pathway are altered in the same direction . Observing high levels of phospho-RAF, phospho-MEK, phospho-ERK, and phosphorylated ERK targets in a case, with or without RAS mutation, would suggest that this pathway was relevant to the biology of that leukemic cell and a good target. We can envision having heat maps of pathway activation as opposed to heat maps of individual components, showing which pathways are most heavily utilized. Use of available databases such as Ingenuity or Cytoscape can aid in development of lists of genes or proteins that have known canonical associations. However, neither is currently very useful for the incorporation of data on protein activation states. It is important to recognize the novel consequences of oncogenic changes as well, such as the noncanonical activation of one pathway by mutated kinases in another pathway, or the previously mentioned effect of IDH mutations on methylation. The ability to build networks from integromic data will permit the recognition of these novel consequences by differential mapping of actual networks onto canonical pathways. We are attempting to build the technology to do this with RPPA data and have used this to recognize new connections in favorable prognosis core binding factor leukemias (York et al. 2012) . Novel connections are likely to be prime targets for therapeutic intervention as the consequences of their blockade are likely to only occur in neoplastic cells and not in the normal cells that lack these connections .
2.4.3 Data from Multiple Different Analytes
It will also be important to be able to recognize when more subtle changes at different levels, or changes in multiple components of one level, combine to result in biologically relevant changes in a target . Understanding the consequences of ncRNA expression can be mind-boggling as individual ncRNAs affect many genes and many ncRNA can affect the same gene. Recently, a “ceRNA hypothesis” has been proposed as a major regulator of gene expression and hence protein levels (Salmena et al. 2011). Furthermore, a given target is subject to direct modulation by one ncRNA and indirect modulation by others that act on the other modulators of that gene. Sensing the integrated effect of all the ncRNA on a given target may therefore be more important than measurement of an individual ncRNA. Partial methylation or histone acetylation in combination with the effects of ncRNAs adds another layer of complexity. Detecting when the regulation of a target is arising solely from one layer, or as the integrated effect of multiple levels, can suggest which modulator to target, or whether a combinatorial approach is required. Protein-based analysis can provide the answer on where the rheostat is “set,” but each layer must be assessed to determine which hand is controlling the knob.
2.4.4 How to Test Rationally
Testing this will require the development of catalogs of ncRNA that can affect a gene and the development of computational methods that can correlate data from the actual levels of all the ncRNA relevant to a given gene with actual mRNA expression and protein level. Likewise, having highly accurate and detailed maps of network interactions of all the proteins that regulate expression of a target must exist before a computational prediction of how epigenetic changes at all the relevant sites combine to affect mRNA and protein levels. Ideally, these predictions would then be validated in controlled systems and the results from those used as feedback on the computational predictions. This can practically be done in cell lines for some interactomes using high-throughput testing systems capable of measuring the consequences of individual and combinatorial multigene modulations. However, in the forthcoming era of individualized medicine, it will not be possible to validate each individual prediction for each patient. It will therefore be necessary to extrapolate from generalized knowledge to individual patients.
2.4.5 If It Is Low, Can You Elevate?
The ability to restore expression of a missing gene depends on what mechanism is decreasing expression. Since repression could arise from histone acetylation, histone methylation, gene methylation, or blockade by ncRNA, or by protein-mediated repression, identifying which is the responsible event and the in vitro testing of whether use of an agent to reverse this has the desired result. Posttranslational modulation of wild-type p53 by the MDM2 modulating agent Nutlin and KST185 (discussed later in this book) is a good example of this.
2.4.6 If It Is High, Can You Successfully Suppress?
Similarly, omics could identify the mechanism by which a protein is overexpressed or activated and suggest where to target. Additionally, specific ncRNA could be used therapeutically to repress a gene directly, or through the repression on proteins than enhance expression (Ma et al. 2010b). This could be tested in vitro to verify that the modulation produces the desired effect or to identify unexpected consequences in other pathways.
Conclusion
In summary, the ever-growing list of available arrays has the potential to provide an overwhelming amount of data for each patient who can reveal the workings of their leukemia cells at a level and degree of detail that was unimaginable until recently. Combined with the rapidly expanding list of targeted agents designed to interfere with the expression or function of these genes and proteins, this presents the theoretical opportunity to match targets to patients in real time and in a manner that dramatically improves response and outcome. Achieving this requires the development of integromic computational capabilities to enable us as physicians to see the relevant from among a torrential flood of data. Or, as once stated by Mao Zedong, “There is great chaos under heaven—the situation is excellent.”
References
Banker DE, Mayer SJ, Li HY et al (2004) Cholesterol synthesis and import contribute to protective cholesterol increments in acute myeloid leukemia cells. Blood 104:1816–1824
Barretina J, Caponigro G, Stransky N et al (2012) The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature 483:603–607
Bendall SC, Simonds EF, Qiu P et al (2011) Single-cell mass cytometry of differential immune and drug responses across a human hematopoietic continuum. Science 332:687–696
Blair A, Sutherland HJ (2000) Primitive acute myeloid leukemia cells with long-term proliferative ability in vitro and in vivo lack surface expression of c-kit (CD117). Exp Hematol 28:660–671
Blair A, Hogge DE, Sutherland HJ (1998) Most acute myeloid leukemia progenitor cells with long-term proliferative ability in vitro and in vivo have the phenotype CD34+/CD71–/HLA-DR−. Blood 92:4325–4335
Butler JS, Koutelou E, Schibler AC, Dent SY (2012) Histone-modifying enzymes: regulators of developmental decisions and drivers of human disease. Epigenomics 4:163–177
Calin GA, Liu CG, Sevignani C et al (2004) MicroRNA profiling reveals distinct signatures in B cell chronic lymphocytic leukemias. Proc Natl Acad Sci U S A 101:11755–11760
Calin GA, Ferracin M, Cimmino A et al (2005) A MicroRNA signature associated with prognosis and progression in chronic lymphocytic leukemia. N Engl J Med 353:1793–1801
Calin GA, Liu CG, Ferracin M et al (2007) Ultraconserved regions encoding ncRNAs are altered in human leukemias and carcinomas. Cancer Cell 12:215–229
Carter BZ, Zhang N, Coombes KR et al (2009) Expression of apoptosis repressor with caspase recruitment domain (ARC), an antiapoptotic protein, is strongly prognostic in acute myeloid leukemia (AML) [abstract]. Blood 114:166
Carter BZ, Qiu YH, Zhang N et al (2010) Expression of ARC (apoptosis repressor with caspase recruitment domain), an antiapoptotic protein, is strongly prognostic in AML. Blood 117(3):780–787
Cesano A, Woronicz J, Cohen AC et al (2009) Single cell network profiling as a tool to identify AML chemotherapy resistant cell phenotypes under in vivo therapeutic pressure [abstract]. Blood 114:165
Chan WI, Huntly BJ (2008) Leukemia stem cells in acute myeloid leukemia. Semin Oncol 35:326–335
Fredriksson S, Gullberg M, Jarvius J et al (2002) Protein detection using proximity-dependent DNA ligation assays. Nat Biotechnol 20:473–477
Garzon R, Heaphy CE, Havelange V et al (2009) MicroRNA 29b functions in acute myeloid leukemia. Blood 114:5331–5341
Gerlinger M, Rowan AJ, Horswell S et al (2012) Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N Engl J Med 366:883–892
Gold L, Ayers D, Bertino J et al (2010) Aptamer-based multiplexed proteomic technology for biomarker discovery. PLoS ONE 5:e15004
Greenbaum D, Colangelo C, Williams K, Gerstein M (2003) Comparing protein abundance and mRNA expression levels on a genomic scale. Genome Biol 4:117
Griffin TJ, Gygi SP, Ideker T et al (2002) Complementary profiling of gene expression at the transcriptome and proteome levels in Saccharomyces cerevisiae. Mol Cell Proteomics 1:323–333
Gygi SP, Rochon Y, Franza BR, Aebersold R (1999) Correlation between protein and mRNA abundance in yeast. Mol Cell Biol 19:1720–1730
Irish JM, Hovland R, Krutzik PO et al (2004) Single cell profiling of potentiated phospho-protein networks in cancer cells. Cell 118:217–228
Jansen R, Greenbaum D, Gerstein M (2002) Relating whole-genome expression data with protein-protein interactions. Genome Res 12:37–46
Jordan CT, Upchurch D, Szilvassy SJ et al (2000) The interleukin-3 receptor alpha chain is a unique marker for human acute myelogenous leukemia stem cells. Leukemia 14:1777–1784
Kadia T, Kornblau SM, Kantarjian H et al (2009) Clinical characterization and proteomic consequences of mutated Ras in acute myeloid leukemia [abstract]. Blood 114:139–140
Kern W, Kohlmann A, Wuchter C et al (2003) Correlation of protein expression and gene expression in acute leukemia. Cytometry B Clin Cytom 55:29–36
Kornblau SM, Banker DE, Stirewalt D et al (2007) Blockade of adaptive defensive changes in cholesterol uptake and synthesis in AML by the addition of pravastatin to idarubicin + high dose Ara-C: a phase I study. Blood 109(7):2999–3006
Kornblau SM, Singh N, Qiu YH et al (2009a) Highly phosphorylated FOXO3 A is an adverse prognostic factor in acute myeloid leukemia [abstract]. Blood 114:166–167
Kornblau SM, Minden MD, Rosen DB et al (2009b) Single cell network profiles in non-M3 AML associated with patient response to standard induction therapy [abstract]. Blood 114:633
Kornblau SM, Tibes R, Qiu YH et al (2009c) Functional proteomic profiling of AML predicts response and survival. Blood 113:154–164
Kornblau SM, McCue D, Singh N et al (2010a) Recurrent expression signatures of cytokines and chemokines are present and are independently prognostic in acute myelogenous leukemia and myelodysplasia. Blood 116(20):4251–4261
Kornblau SM, Minden A, Hogge D, Chohen A, Cesano A (2010b) Insights into acute myeloid leukemia via single cell network profiling. Clin Lab Int 1:12–15
Kornblau SM, Minden M, Rosen D et al (2010c) Dynamic single cell network profiles in AML are associated with patient response to standard induction therapy. Clin Can Res 16(14):3721–3733
Kornblau SM, Covey T, Putta S et al (2011a) Signaling changes in the stem cell factor-AKT-S6 pathway in diagnostic AML samples are associated with disease relapse. Blood Cell J 1(2):e3
Kornblau SM, Qiu YH, Zhang N et al (2011b) Abnormal expression of FLI1 protein is an adverse prognostic factor in acute myeloid leukemia. Blood 118:5604–5612
Kornblau SM, Qutub A, Yao H et al (2013) Proteomic profiling identifies distinct protein patterns in acute myelogenous leukemia CD34+CD38- stem-like cells. PLoS One 8(10):e78453
Krutzik PO, Irish JM, Nolan GP, Perez OD (2004) Analysis of protein phosphorylation and cellular signaling events by flow cytometry: techniques and clinical applications. Clin Immunol 110:206–221
Lee DW, Futami M, Carroll M et al (2012) Loss of SHIP-1 protein expression in high-risk myelodysplastic syndromes is associated with miR-210 and miR-155. Oncogene 31:4085–4094 (Epub)
Levin VA, Panchabhai SC, Shen L et al (2010) Different changes in protein and phosphoprotein levels result from serum starvation of high-grade glioma and adenocarcinoma cell lines. J Proteome Res 9:179–191
Lu C, Ward PS, Kapoor GS et al (2012) IDH mutation impairs histone demethylation and results in a block to cell differentiation. Nature 483:474–478
Ma W, Yang D, Gu Y et al (2009) Finding disease-specific coordinated functions by multi-function genes: insight into the coordination mechanisms in diseases. Genomics 94:94–100
Ma W, Wang M, Wang ZQ et al (2010a) Effect of long-term storage in TRIzol on microarray-based gene expression profiling. Cancer Epidemiol Biomarkers Prev 19:2445–2452
Ma L, Reinhardt F, Pan E et al (2010b) Therapeutic silencing of miR-10b inhibits metastasis in a mouse mammary tumor model. Nat Biotechnol 28:341–347
Miao F, Natarajan R (2005) Mapping global histone methylation patterns in the coding regions of human genes. Mol Cell Biol 25:4650–4661
Michiels S, Koscielny S, Hill C (2005) Prediction of cancer outcome with microarrays: a multiple random validation strategy. Lancet 365:488–492
Minden MD, Kornblau SM, Rosen DB et al (2009) FLT3 ITD signaling profiles in AML samples harboring mutations [abstract]. Blood 114:635
Neeley ES, Kornblau SM, Coombes KR, Baggerly KA (2009) Variable slope normalization of reverse phase protein arrays. Bioinformatics 25:1384–1389
Neeley ES, Baggerly KA., Kornblau SM (2012) Surface adjustment of reverse phase protein arrays using positive controls spots. Cancer Inform 2012:77–86
Ogata H, Goto S, Sato K et al (1999) KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res 27:29–34
Pemmaraju N, Kantarjian HM, Ravandi F et al (2009) FLT3 inhibitor therapy for patients with myelodysplastic syndromes (MDS) and acute myeloid leukemia (AML): impact on survival according to FLT3 status [abstract]. Blood 114:424–425
Phillips CL, Gerbing R, Alonzo T et al (2010) MDM2 polymorphism increases susceptibility to childhood acute myeloid leukemia: a report from the Children’s Oncology Group. Pediatr Blood Cancer 55:248–253
Pichiorri F, Suh SS, Rocci A et al (2010) Downregulation of p53-inducible microRNAs 192, 194, and 215 impairs the p53/MDM2 autoregulatory loop in multiple myeloma development. Cancer Cell 18:367–381
Poland KS, Shardy DL, Azim M et al (2009) Overexpression of ZNF342 by juxtaposition with MPO promoter/enhancer in the novel translocation t(17;19)(q23;q13.32) in pediatric acute myeloid leukemia and analysis of ZNF342 expression in leukemia. Genes Chromosomes Cancer 48:480–489
Quintas-Cardama A, Qiu YH, Post SM et al (2012) Reverse phase protein array profiling reveals distinct proteomic signatures associated with chronic myeloid leukemia progression and with chronic phase in the CD34-positive compartment. Cancer 118:5283–5292 (Epub)
Ravandi F, Patel K, Luthra R et al (2011) Prognostic significance of alterations in IDH enzyme isoforms in patients with AML treated with high-dose cytarabine and idarubicin. Cancer 118:(10):2665–2673
Roboz GJ, Guzman M (2009) Acute myeloid leukemia stem cells: seek and destroy. Expert Rev Hematol 2:663–672
Rosen DB, Minden MD, Kornblau SM et al (2010) Functional characterization of FLT3 receptor signaling deregulation in acute myeloid leukemia by single cell network profiling (SCNP). PLoS ONE 5:e13543
Ruvolo PP, Qui YH, Coombes KR et al (2011a) Low expression of PP2 A regulatory subunit B55alpha is associated with T308 phosphorylation of AKT and shorter complete remission duration in acute myeloid leukemia patients. Leukemia 25:1711–1717
Ruvolo PP, Zhou L, Watt JC et al (2011b) Targeting PKC-mediated signal transduction pathways using enzastaurin to promote apoptosis in acute myeloid leukemia-derived cell lines and blast cells. J Cell Biochem 112:1696–1707
Salmena L, Poliseno L, Tay Y, Kats L, Pandolfi PP (2011) A ceRNA hypothesis: the Rosetta Stone of a hidden RNA language? Cell 146:353–358
Starkova J, Gadgil S, Qiu YH et al (2011) Up-regulation of homeodomain genes, DLX1 and DLX2, by FLT3 signaling. Haematologica 96:820–828
Subramanian A, Tamayo P, Mootha VK et al (2005) Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles [abstract]. Proc Natl Acad Sci U S A 102:15545–15550
Taby RF, Yellapragada SV, Kroeger H et al (2009) Validation of a DNA methylation signature of favorable prognosis in newly diagnosed acute myeloid leukemia [abstract]. Blood 114:645
Taussig DC, Vargaftig J, Miraki-Moud F et al (2010) Leukemia-initiating cells from some acute myeloid leukemia patients with mutated nucleophosmin reside in the CD34– fraction. Blood 115:1976–1984
Tehranchi R, Woll PS, Anderson K et al (2010) Persistent malignant stem cells in del(5q) myelodysplasia in remission. N Engl J Med 363:1025–1037
Tibes R, Qiu YH, Lu Y et al (2006) Reverse phase protein array (RPPA): validation of a novel proteomic technology and utility for analysis of primary leukemia specimens and hematopoietic stem cells (HSC). Mol Cancer Ther 5:2512–2521
Tsao T, Kornblau S, Safe S et al. (2010) Role of peroxisome proliferator-activated receptor-gamma and its coactivator DRIP205 in cellular responses to CDDO (RTA –401) in acute myelogenous leukemia. Cancer Res 70:4949–4960
Valastyan S, Chang A, Benaich N, Reinhardt F, Weinberg RA (2011) Activation of miR-31 function in already-established metastases elicits metastatic regression. Genes Dev 25:646–659
Walter MJ, Shen D, Ding L et al (2012) Clonal architecture of secondary acute myeloid leukemia. N Engl J Med 366:1090–1098
Washburn MP, Koller A, Oshiro G et al (2003) Protein pathway and complex clustering of correlated mRNA and protein expression analyses in Saccharomyces cerevisiae. Proc Natl Acad Sci U S A 100:3107–3112
Weinstein JN (2002) ‘Omic’ and hypothesis-driven research in the molecular pharmacology of cancer. Curr Opin Pharmacol 2:361–365
Xu L, Tan AC, Winslow RL, Geman D (2008) Merging microarray data from separate breast cancer studies provides a robust prognostic test. BMC Bioinformatics 9:125
York H, Kornblau SM, Qutub AA (2012) Network analysis of reverse phase protein expression data: Characterizing protein signatures in acute myeloid leukemia cytogenetic categories t(8;21) and inv(16). Proteomics 12:2084–2093
Acknowledgments
Thanks to Dr. Richard Eric Davis for critical reading and suggestions.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer-Verlag New York
About this chapter
Cite this chapter
Kornblau, S. (2015). The Use of “Omics” to Guide the Selection of Targeted Therapy. In: Andreeff, M. (eds) Targeted Therapy of Acute Myeloid Leukemia. Current Cancer Research. Springer, New York, NY. https://doi.org/10.1007/978-1-4939-1393-0_2
Download citation
DOI: https://doi.org/10.1007/978-1-4939-1393-0_2
Published:
Publisher Name: Springer, New York, NY
Print ISBN: 978-1-4939-1392-3
Online ISBN: 978-1-4939-1393-0
eBook Packages: MedicineMedicine (R0)