Abstract
Protein folding is often hampered by protein aggregation, which can be prevented by a variety of chaperones in the cell. In this review, I summarize recent topics on in vitro and in vivo approaches to understand the role of Escherichia coli chaperones to prevent protein aggregations.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Protein folding
- Protein aggregation
- GroEL
- Chaperonin
- DnaK
- Trigger factor
- Reconstituted cell-free translation system
- Proteomics
1 Introduction
A protein must fold into the correct tertiary structure after emerging from the ribosome. The unique native structure of a protein is encoded in its amino acid sequence [1]. However, protein folding is often hampered by protein aggregation , which is generally prevented by a variety of chaperone proteins in the cell [2]. Chaperones are also involved in other multiple cellular processes associated with the conformational changes of proteins, such as stress responses [2].
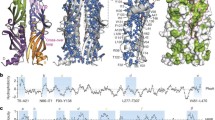
Many efforts over the past two decades have been dedicated to elucidate the mechanism of molecular chaperones. The best-characterized chaperones are those in Escherichia coli. In E. coli, three major chaperone systems are involved in the folding of newly synthesized proteins in the cytoplasm [2]. The first is trigger factor (TF) , which directly associates with ribosome and interacts with nascent chains cotranslationally [3]. TF associates with L23 ribosomal protein in a monomer with the dragon-shaped structure [3]. The second is DnaK , which is one of the Hsp70 family conserved widely in all kingdoms of life. DnaK, as other Hsp70 proteins, consists of two domains, an adenosine triphosphatase (ATPase) domain and a polypeptide-binding domain [2, 4]. The interaction of DnaK with the substrate polypeptide is ATP dependent. DnaK binds substrate proteins with high affinity in the adenosine diphosphate (ADP) state, and with low affinity in the ATP state. The function of DnaK is regulated by cofactors, DnaJ and GrpE. DnaJ binds denatured proteins in an ATP-independent manner to be targeted to DnaK. GrpE is a nucleotide exchange factor to regulate the ATPase cycle of DnaK. The third is GroEL , which belongs to conserved chaperonin family. GroEL forms a large cylindrical complex with cochaperonin GroES in the presence of ATP, which encapsulates substrate proteins into its cavity to assist the folding [2, 4, 5]. The bacterial GroEL and GroES are the only indispensable chaperones for the viability of E. coli [6, 7]. GroEL consists of two heptameric rings of 57 kDa subunits, and provides binding sites for non-native substrate proteins. GroES, a dome-shaped heptameric ring of ~ 10 kDa subunits, caps GroEL in the presence of adenine nucleotides, forming a central cavity that can accommodate substrate proteins up to ~ 60 kDa in size [8, 9].
It has been known that these three chaperone systems act cooperatively; TF and DnaK are known to exhibit overlapping co-translational roles in vivo [10–12], whereas GroEL is believed to be implicated in folding after the polypeptides are released from the ribosome, albeit that possible cotranslational involvement of GroEL has also been reported [13–16].
In addition to the “classical” functions in the protein folding , chaperones are proposed to promote protein evolution by buffering the destabilization of proteins caused by harmful genetic mutations [17–21]. The absolute requirement of chaperones for cellular functions and for protein evolution might account for the fact that an organism lacking chaperones has not been identified.
To understand protein folding in the cell, we at first need to know which proteins are aggregation-prone. Then, we have to tackle the question of which aggregation-prone proteins are rescued by chaperones. In this review, I summarize recent topics on in vitro and in vivo approaches to understand the role of E. coli chaperones to assist folding by preventing aggregates formation. Main topics include global in vitro analysis of protein aggregation and chaperone effects on aggregation-prone proteins, and in vivo substrates of chaperonin GroEL and chaperone DnaK .
2 In Vitro Analyses Using a Reconstituted Cell-Free Translation System
Chaperones are required to prevent protein aggregations [2] . Although we empirically know some proteins are aggregation-prone, there is no systematic global analysis on which proteins are actually aggregation-prone or not. In general, it is very difficult to evaluate the aggregation propensity of a protein of interest after the expression of the protein in cells, since cells contain a variety of chaperone proteins. Similar limitation would be applicable for conventional cell-free analysis since cell lysates contain endogenous chaperones. In this context, chaperone-free translation system would be ideal to evaluate the aggregation-prone propensity of proteins of interest. A highly controllable cell-free translation system called the Protein synthesizing Using Recombinant Elements (PURE) system was successfully reconstituted by using only the essential purified factors and enzymes that are responsible for gene expression in E. coli[22, 23]. The protein synthesis in this system has been proven to be highly manipulatable. Importantly, PURE system does not contain any chaperone [22, 24]. Therefore, we can evaluate the inherent aggregation propensities of proteins of interest in a translation-coupled manner under the chaperone-free conditions. In addition, this reconstituted translation system represents a straightforward approach for studying the chaperone-assisted folding of newly synthesized polypeptides, as the roles these chaperones play can be determined by simply adding them to the translation system .
2.1 Global Aggregation Analysis of the Entire Ensemble of E. coli Proteins
Understanding the mechanism underlying aggregate formation is required for the development of a wide variety of protein sciences. However, the relationship between the protein aggregation propensities and the primary sequences remains poorly understood. Since it is empirically known that some proteins tend to aggregate, several groups systematically studied the effects of mutations in proteins of interest that caused the formation of insoluble aggregates [25–28]. Aggregation-prone properties of thousands of E. coli proteins were evaluated using PURE system [29].
2.1.1 Experiment
The scheme of the global analysis is shown in Fig. 15.1: the one-by-one synthesis of individual E. coli proteins, the quantification of solubility by a centrifugation-based assay, and the statistical analyses of the collected data. This is an “in vitro (reconstituted) proteome” approach, in which the properties of thousands of proteins, including proteins with extremely low abundance in cells, are investigated individually after cell-free translation.
An in vitro expressed proteome approach for global aggregation analysis. Schematic illustration of the experiment. Escherichia coli proteins were separately expressed with a reconstituted cell-free translation system, the PURE system, in the absence and the presence of the major E. coli chaperones (trigger factor, TF; GroEL/GroES, GroE; DnaK/DnaJ/GrpK, KJE). Each translation product was labeled with [35S]methionine. After translation, the uncentrifuged total fraction (Total) and the supernatant fraction after centrifugation (Sup) were electrophoresed and quantified by autoradiography. The ratio of the translation products in the Total and Sup fractions was defined as the solubility, which represented the aggregation propensity of the protein. Reprinted from ref [44]
A comprehensive analysis, in which the complete E. coli open reading frame (ORF) library (ASKA library) [30] was translated in the PURE system under the same conditions, was conducted. The ASKA library consists of all predicted ORFs of the E. coli genome, including membrane proteins [30]. A total of 4132 ORFs were individually amplified by polymerase chain reaction (PCR) using a common primer set, and then were used for protein synthesis in the PURE system at 37 °C for 60 min.
The 35S-methionine-labeled proteins were quantified after electrophoresis of the translation products. Approximately 70 % of the E. coli ORFs (3173 proteins out of 4132) were successfully quantified. The remainder was not quantified, due to insufficient translation and trouble during the electrophoresis. The unquantifiable group contained ~ 60 % of the inner membrane proteins (435/754), whereas more than 80 % of the cytoplasmic proteins (2277/2688) were quantified.
The propensity for protein aggregation was examined by a centrifugation assay [15, 24]. An aliquot of the translation mixture was centrifuged. The proportion of the supernatant fraction, which was obtained after the centrifugation of the translation mixture, to the uncentrifuged total protein was defined as the solubility, the index of the aggregation propensity.
2.1.2 Bimodal Solubility Distribution
A histogram of the individual solubilities, based on data from 3173 translated proteins, showed a clear bimodal, rather than normal Gaussian, distribution (Fig. 15.2), indicating that the aggregation propensities are not evenly distributed across a continuum. Subtraction of the predicted integral membrane proteins from the data did not change the bimodal distribution, suggesting that the cytoplasmic proteins can be categorized into an aggregation-prone group and a highly soluble one. To elucidate which characteristics of the protein influence this bimodality, we compared a variety of protein properties in the aggregation-prone (Agg: defined as less than 30 %) and highly soluble (Sol: defined as more than 70 %) groups. One might expect that the bimodal distribution in the histogram is simply due to the difference in the synthesized yield of proteins, since it has been generally believed that higher protein concentrations generate more protein aggregates. However, this is not the case, since there is no apparent correlation between the solubilities and the yields.
Histogram of solubility for the 3173 quantified proteins. The proteins with solubilities below 30 % and above 70 % were defined as the aggregation-prone (Agg) and soluble (Sol) groups, respectively. Adapted from ref [29]
The essential proteins tended to be enriched in the high solubility group, suggesting that the essential proteins might have evolved to be soluble for their irreplaceable properties. In addition to the essentiality, the solubilities are strongly dependent on the functions. For example, structural component group, which is mainly composed of ribosomal proteins, and factor group, which includes transcription or translation factors, chaperones, and proteases, showed a strong bias to the high solubility group. In contrast, the proteins in the transporter group tended to be aggregation-prone. Regarding the oligomeric states of the proteins, preliminary analysis shows that hetero-oligomers seem to be aggregation-prone although we cannot say the tendency is statistically significant due to the incomplete database on the oligomeric states.
2.1.3 Physicochemical Properties
The physicochemical properties of the proteins, such as the molecular weights, the deduced isoelectric points (pI) and the amino acid residue content were compared to address the relationship between solubility and amino acid sequence. The distribution of molecular weights in the soluble group was shifted to smaller sizes as compared with the total histogram. Regarding the pI, an enrichment of low pI (5–7) proteins in the high solubility distribution was observed, whereas the aggregation-prone proteins showed a somewhat broader pI distribution (ranging from 5 to 10). Analysis to test whether the amino acid residue content affected the solubility revealed that higher contents of negatively charged residues (Asp and Glu) tended to be soluble. Higher contents of aromatic residues (Phe, Tyr, and Trp) were slightly biased to be aggregation-prone. In contrast, no significant difference was observed in the contents of hydrophobic residues (Val, Leu, and Ile) and positively charged residues (Lys, Arg, and His). Since it has been believed that the hydrophobic interaction is a critical driving force in aggregate formation, the lack of an apparent correlation in the hydrophobic residue content was unexpected. Other attempts to detect a bias between the solubility and the hydrophobicity, including a well-known hydropathy plot analysis [31, 32], which shows clusters of hydrophobic residues in the primary amino acid sequences, or several hydrophobic-polar alternates analyses also failed. We note that Gln/Asn-rich sequences including polyglutamine repeats, which tend to form amyloid fibrils, are very rare in the E. coli ORFs [33].
Several analyses related to the secondary structures were subsequently conducted. The secondary structure contents were predicted by using popular prediction methods, such as Chou-Fasman [34] and Psi-blast based secondary structure prediction (PSIPRED) [35, 36]. However, a notable correlation between the predicted secondary structure content and the solubility was not detected.
To address the correlation between the solubilities and the tertiary structures, the solubilities with the Structural Classification of Proteins (SCOP) database, which is a comprehensive ordering of all proteins with known structures, according to their evolutionary and structural relationships [37] were compared. The classification is based on hierarchical levels: class, fold, superfamily and family. Superfamilies and families are defined as having a common fold if their proteins have the same major secondary structures in the same arrangement and with the same topological connections. Roughly, the secondary structures did not correlate with the aggregation propensities. Regarding the SCOP folds some of the SCOP folds were extremely biased toward their solubilities. For example, in the periplasmic-binding protein-like II fold (SCOP fold: c94) group, which is largely dominated by deoxyribonucleic acid (DNA)-binding transcriptional regulator proteins, 83 % of the members were low solubility proteins (35 out of 42 assigned proteins), whereas only one protein was in a soluble group. Other low soluble folds included Pyridoxal-phosphate (PLP)-dependent transferases fold (c67), DNA/ribonucleic acid (RNA)-binding 3-helical bundle fold (a4), triose phosphate isomerase (TIM) β/α-barrel fold (c1) and P-loop containing nucleoside triphosphate hydrolases (c37). For the highly soluble folds, we assigned Flavodoxin-like fold (c23), oligonucleotide/oligosaccharide-binding (OB)-fold (b40), and Thioredoxin fold (c47).
2.1.4 Aggregation Propensity Prediction
These data can be applicable to several recently developed web tools to predict protein aggregation . However, none of the tools including TANGO [38], AGGRESCAN [39], and PASTA programs [40] extracted a notable positive correlation between our datasets and the predicted results, probably due to the fact that the algorithms used in those programs basically relied on data from amyloid aggregates in eukaryotes. Other attempts to predict the solubilities based on this dataset have been developed [41–43].
2.2 Global Analyses of Chaperone Effects on Aggregation-Prone Proteins
Global aggregation analysis under chaperone-free conditions has been extended to that in the presence of the major E. coli chaperones, TF , DnaK/DnaJ/GrpE (DnaKJE), and GroEL/GroES (GroE) [44].
In previous global aggregation analysis, aggregation-prone group was defined as the proteins with less than 30 % solubility [29]. Effects of chaperones were examined for the aggregation-prone E. coli proteins that are predicted to reside in the cytosol (792 proteins).
2.2.1 Experiment
All of the cytosolic aggregation-prone proteins were synthesized by the PURE system at 37 °C for 60 min in the absence or presence of each chaperone. Each chaperone was added at the approximate physiological concentration based on previous assessments of chaperone activities under cell-free conditions [15, 45, 46]. The 35S-methionine-labeled proteins were electrophoresed on Sodium dodecyl sulfate (SDS) gels and quantified. The aggregation propensity was examined by the centrifugation assay [29]. Typical results are shown in Fig. 15.3a. Almost all of the proteins (788/792) were quantified for their solubilities under each condition .
Global analysis of chaperone effects on the prevention of aggregate formation. a Typical examples. SDS-gels of four aggregation-prone E. coli cytosolic proteins (asd, hemB, yedS, and yajB) in the absence and the presence of the chaperones are shown. The numbers below the electrophoretic pattern indicate the solubility values, calculated by the ratio of the amount of translation products in the Sup (S) and Total (T) fractions. b Histograms of solubilities in the absence or the presence of three E. coli chaperones. c Two-dimensional distribution plot of ∆solubilities for DnaKJE and GroE. Dashed lines represent the boundaries of the lower and upper quartiles (34, 67 % and 26, 58 % solubility values for DnaKJE and GroE, respectively). Reprinted from ref [44]
In total, more than 3000 assays (788 proteins × 4 conditions = 3152) were conducted. Overall, the chaperones tested effectively increased the solubility of the aggregation-prone proteins (Fig. 15.3b). Overall, the solubilities of two-thirds of the proteins (526/788) were drastically increased, defined as more than a 50 % increase, in the presence of any one of the chaperones. The proteins that were not rescued by any one of the chaperones, defined as less than a 20 % increase in the solubility, represented only 3 % of the total (24 out of 788). Taken together, this comprehensive analysis has explicitly confirmed the global role of chaperones in preventing the aggregation of hundreds of proteins.
2.2.2 Each Chaperone Effects
The effects of each chaperone were compared. It is noteworthy that TF had only a marginal effect. On the other hand, DnaKJE and GroE increased the solubilities of many proteins. The solubilities of 409 and 287 proteins with DnaKJE and GroE, respectively, were drastically (> 50 ;%) increased. Approximately 30 % of the proteins with > 50 % increase in solubility (175 proteins) were common between DnaKJE and GroE, indicating that these overlapping proteins were rescued well by either DnaKJE or GroE. Taken together, the data clearly show the global effects of DnaKJE and GroE in preventing aggregation.
2.2.3 Physicochemical Properties
To further investigate the effects of DnaKJE and GroE, the data on DnaKJE and GroE were plotted in two dimensions. A substantial fraction of the proteins was biased toward DnaKJE or GroE. These biases suggest that DnaKJE and GroE could have different recognition modes for substrates. To extract the possible preferences of DnaKJE and GroE, physicochemical properties of the proteins were analyzed. Some biases were observed in molecular weights when the analysis was limited in the proteins that were well-solubilized by either DnaK or GroE, defined as the upper quartile (≥ 75th percentile) in the distribution. GroE is biased toward lower molecular weight proteins (20 ~ 50 kDa), whereas DnaKJE is effective for larger ones (> 60 kDa).
To address the correlation between the chaperone effects and the tertiary or quaternary structures, the SCOP database (class and fold) [37] and the oligomeric states of proteins were compared, although only a small number of proteins was analyzed, due to the limited database size. When classified by the SCOP classes (all-α, all-β, α/β, and α+β), DnaKJE was effective for the α+β class, whereas GroE was not effective for the all-α class. Furthermore, there are some biases for DnaKJE and GroE in several SCOP folds. GroE was biased toward the c1 (TIM barrel) fold, which is plausible since the most abundant fold in the in vivo obligate GroE substrates is the TIM barrel fold [47, 48]. Neither DnaKJE nor GroE was effective for the a4 (DNA/RNA-binding 3-helical bundle fold) and c94 (periplasmic-binding protein-like II) folds.
2.2.4 Cooperative Effects of Chaperones
Neither DnaKJE nor GroE could rescue a subset of proteins mapped around the lower left area in the plot (Fig. 15.3c) , termed recalcitrant proteins. Various combinations of chaperones were examined to investigate whether these recalcitrant proteins could be solubilized. All of the recalcitrant proteins, which were defined as the proteins categorized in the lower quartiles in DnaKJE and GroE (53 proteins), were translated in the presence of chaperone combinations: TF+DnaKJE, TF+GroE, DnaKJE+GroE, and TF+DnaKJE+GroE. The solubilities under the TF+DnaKJE and TF+GroE conditions were slightly increased, whereas the combination of DnaKJE and GroE was more effective for some of these recalcitrant proteins, reflecting the consensus that GroE and DnaKJE synergistically assist with the folding of nascent polypeptides [45, 49]. Strikingly, the addition of all three chaperones to the recalcitrant proteins drastically changed the solubility distribution: More than 70 % of the recalcitrant proteins (38/53) showed significantly improved solubilities in the presence of all chaperones. These results suggest that TF also has the potential to act cooperatively with DnaKJE and GroE, although TF itself was not very effective in preventing aggregation.
2.3 Solubility Database of all E. coli Proteins (eSOL)
The dataset of aggregation analyses including chaperone effects are freely accessible at online database (eSOL database: http://www.tanpaku.org/tp-esol).
3 In Vivo Substrates of Chaperonin GroEL
An important goal in chaperonin biology is to identify a subset of obligate GroEL/GroES (GroE) substrates that absolutely require GroE for folding in cells . Precise identification of the obligate GroE substrates should contribute to the identification of a distinctive role for GroE among chaperones, reveal the structural features of the obligate substrates, and shed light on the role of GroE in protein evolution.
3.1 Phenotype Analysis Using GroE-Knockdown Strain
One approach to identify obligate GroE substrates is a detailed analysis of the phenotypes of GroE-depleted cells. Since chaperonin GroE is the only indispensable chaperone for the viability of E. coli[6, 7], GroE-deletion strain is not available. However, a conditional GroE expression strain, MGM100, in which the native groE chromosomal promoter region has been replaced with the araC gene and the araBAD promoter [50]. When the sugar in the growth medium is changed from arabinose to glucose, the GroE levels decrease by 90 % within 2 h in this strain [50]. Investigations of E. coli phenotype after the GroE-depletion have identified DapA and FtsE as obligate GroE substrates in the cell lysis and filamentous morphology phenotypes, respectively [50, 51]. Although a detailed phenotypic analysis can precisely identify obligate GroE substrates, this approach is limited, in that the substrates can only be identified one by one, and only in the cells with experimentally tractable phenotypes.
3.2 Mass Spectrometry-based Proteomics of GroEL Interacting Proteins
Another approach to substrate identification is a proteome-wide analysis. Hundreds of GroEL substrates have been identified using mass spectrometry (MS) [47, 52] . In particular, Kerner et al. have identified ~ 250 substrates that interact with GroE in E. coli and categorized them into three classes depending on their enrichment in the GroE complex: Class I substrates as spontaneous folders, Class II as partial GroEL-dependent substrates, and Class III as the potential obligate GroE substrates [47]. Notably, ~ 84 Class III substrates are estimated to occupy ~ 80 % of the available GroEL capacity in the cell. This classification was primarily based on the proteomics of the GroE interactors. However, except for DapA, GatY, MetK, ADD, and YajO, which were verified as requiring GroE for folding, the in vivo GroE dependency of other Class III substrates has not been tested [47].
Impairment of GroE function in E. coli results in the accumulation or degradation of newly translated polypeptides due to misfolding. Wholesale accumulation of aggregates was observed in the severe temperature-sensitive GroE strain, which harbors GroEL (E461K) mutant instead of wild-type GroEL [53]. The proteomic analysis using MS of the aggregated proteins identified ~ 300 proteins [53]. Most of the identified proteins were cytoplasmic proteins, many known to be highly abundant [53] .
Note that similar approach to identify in vivo GroEL interactors has been applied to other bacteria besides E. coli. 24 ~ 28 GroEL interacting proteins were identified by MS-based proteomics in Thermus thermophilus and Bacillus subtils [54, 55] .
3.3 Combined Approach to Identify Obligate GroE-Dependent Substrates
Previous proteome-wide analysis of E. coli chaperonin GroEL interactors predicted obligate chaperonin substrates, which were termed Class III substrates. However, the requirement of chaperonins for in vivo folding has not been fully examined. In fact, one of the Class III proteins, ParC, was functional even under GroE-depleted conditions [51], raising the possibility that the predicted Class III proteins are not necessarily obligate substrates of GroE [48].
3.3.1 Proteomics of the Soluble Fraction in GroE-Depleted E. coli
In GroE-depleted cells, the known obligate GroE substrates either aggregate (e.g., MetK) or are degraded (e.g., DapA, FtsE, and GatY) [47, 50, 51] . Thus, the abundance of other potential GroE obligate substrates would also be reduced in the soluble fraction of GroE-depleted cells. A proteome-wide analysis of the soluble fraction of GroE-depleted cells, using a conditional GroE expression strain MGM100 [50], was therefore conducted to find candidate in vivo obligate GroE substrates. For proteomics, cells were subjected to a 2 h depletion of GroE in LB medium, during which the level of GroEL was reduced to less than 10 % of that in undepleted cells [50], and, as a control, cells with a normal level of GroE were also prepared. Note that diaminopimelic acid (DAP) was added to the medium to prevent cell lysis, a known consequence of DapA (a Class III substrate) deficiency in GroE-depleted cells [50]. The abundance of each protein was quantified by the exponentially modified Protein Abundance Index (emPAI), which provides an estimate of protein abundance by quantitating non-redundant peptides identified by MS [56, 57]. As a control for the sugar-associated changes in the proteome, MG1655, the wild-type parent strain of MGM100, was also examined by the same procedures.
The proteomics quantified a total of 986 proteins in MGM100 cells under glucose and arabinose conditions. The relative abundances of proteins defined as emPAIglucose/emPAIarabinose shows that ~ 33 % of proteins were reduced by more than 50 % in MGM100 cells. The drastic reduction of many proteins during glucose growth in MGM100 cells was caused by the GroE-depletion. We note that a significant number of proteins were increased in the GroE-depleted cells, including methionine biosynthetic enzymes, such as MetE [7, 53, 58, 59], and certain chaperones, such as DnaK and SecB .
The proteome data were used to roughly choose candidate GroE substrates by the following criteria. First, the proteins with soluble abundance that was reduced during depletion to less than 50 % of that found during arabinose growth in MGM100 (as a genetic control) were chosen. The cutoff value of 50 % was set to minimize false negatives, as the highest solubility of known in vivo obligate GroE substrates was 46 %, as found with MetK. The 347 proteins chosen by the first criterion contained many false positives due to a sugar-associated reduction in their levels, and thus were filtered by a second criterion, in which the proteins with expression in MGM100 during glucose growth reduced to less than 50 % of that found during glucose growth in MG1655 were chosen. Using the genetic and sugar controls, 252 proteins among the detected 986 proteins met both criteria for rough candidate GroE substrates. The candidates included all of the in vivo tested obligate GroE substrates (MetK, GatY, and DapA), except for FtsE, which was not quantified in the proteomics, confirming the reasonableness of the selected threshold for the growth conditions used here. Then, the percentages of the candidates showing protein reductions in each of the GroEL substrate classes defined by Kerner et al. [47] were calculated. Eight percent of Class I, 32 % of Class II, and 56 % of Class III substrates were reduced in the GroE-depleted cells. The fraction of class members showing reduced protein amounts increased with the degree of GroEL dependence. It is also noteworthy that about 44 % of the Class III substrates (24 out of the 43 quantified proteins) did not meet the criteria for GroE obligate substrates. This again suggests that a significant fraction of Class III members, in addition to ParC, are not obligate substrates in vivo .
3.3.2 About 40 % of Class III Substrates Do Not Require GroE for Solubility
To assess whether ~ 40 % of the Class III substrates are not actually obligate GroE substrates, we developed methods, independent of proteomics , to verify their GroE requirement for solubility. The methods also aimed to comprehensively cover all of the Class III proteins suggested by Kerner et al. [47], since the proteomics of MGM100 detected only about half of the Class III substrates (43 of 84). The expression of individual target proteins was induced from a tac promoter in MGM100 cells after a 2 h depletion of GroE, and their total amounts and the proportion in the soluble fraction were measured. The obligate GroE substrates would be expected to become insoluble or be degraded. To validate the strategy, proteins for which the status of GroE-dependence had already been verified were examined: Enolase (spontaneously folding in vitro, Class I), GatD (partial GroE dependent-folding in vitro, Class II), MetK, FtsE, DapA (the in vivo obligate GroE substrates, Class III), and ParC (assigned as Class III, but functional in the GroE-depleted cells) [47, 50, 51]. Enolase, GatD, and ParC were soluble irrespective of the GroE level, whereas MetF, MetK, FtsE, and DapA aggregated in the GroE-depleted cells. The disappearance or persistence of the bands under GroE-depleted conditions was almost complete, enabling easy and clear discrimination.
Next, the method was extended to all of the essential genes in the three GroEL substrate classes, to test the GroE requirement for solubility (G.R.) in cells. The solubility of the essential Class I and Class II proteins (proteins with low expression levels were not measured) was independent of the GroE levels, confirming that Classes I and II were not dependent on GroE for folding. The GroE-independence of Ppa (Class I), GatD, LpxA, HemL, and FabG (Class II), which were candidates of GroE substrates identified by the proteomics , indicated that not all of the candidates predicted by the proteomics are in vivo obligate GroE substrates. The results showed that the in vivo obligate GroE substrates were enriched in Class III, but not in Class I and II proteins. More importantly, the results also indicated that approximately half of the Class III proteins did not require GroE for solubility, as already suggested by the above data. Depending on the in vivo GroE requirement, the Kerner’s Class III substrates were divided into Class III+ (plus; GroE dependent for solubility in vivo) and Class III− (minus; not GroE dependent for solubility in vivo). Finally, all of the remaining Class III substrates were tested for an in vivo GroE requirement. When all of the solubility assays for the Class III substrates were combined, Class III was divided into 49 Class III+ and 34 Class III− substrates.
GroE-independent folding of Class III− proteins under GroE-depleted conditions was demonstrated for several representative Class III− proteins. First, the intracellular thymidine concentration was not decreased in the GroE-depleted cells, implying that FolE, one of the essential Class III− proteins, is functional in GroE-depleted cells, since FolE-defective cells only grow in thymidine-supplemented rich medium [60]. Second, the enzymatic activities of the Class III− proteins were directly assayed in the E. coli lysates. The activities of two essential Class III− proteins, an inositol monophosphatase, SuhB, and a tRNA methylase, TrmD, were measured in the lysates after the overexpression of Class III− proteins. The enzymes were active in both the GroE-depleted and -normal cells, indicating that the enzymes are both soluble and functional in the GroE-depleted cells. Among the Class III− substrates, only 4 proteins (ParC, FolE, SuhB, and TrmD) are essential. Although the functionality of the remaining Class III− proteins was not tested, at least all of the essential Class III− proteins were physiologically functional even in the GroE-depleted cells, further supporting the validity of the Class III+ and III− grouping.
3.3.3 Identification of Other In Vivo Obligate GroE Substrates that Were Not Previously Assigned as Class III Substrates
After the complete survey of the Class III substrates, other novel GroE obligate substrates besides the identified Class III proteins were searched. The metabolomics data showed that the level of O-phosphoserine, the product of a Class II substrate, SerC, was reduced in the GroE-depleted cells, suggesting that SerC was not active in the cells. In addition, the proteomics data also suggested that SerC was reduced in the supernatant of the GroE-depleted cells. SerC was aggregated in the GroE-depleted cells, strongly suggesting that the in vivo obligate GroE substrates are not confined to the identified Class III substrates.
Other putative GroE substrates were also identified. First, candidate proteins were selected based on the proteomics data under GroE-depleted conditions. The GroE requirement for a dozen drastically reduced proteins in the GroE-depleted cells was verified. These included 3 Class II proteins (KdsA, PyrC, and NuoC) and 6 proteins that had not appeared among the GroEL interactors (GuaC, ThiL, SdaB, PyrD, NemA, GdhA). The solubility assays of these candidate proteins revealed that all of the Class II candidates and 2 of the other 6 candidates (PyrD and GdhA) behaved as in vivo GroE obligate substrates.
In addition, the homologs of Class III+ substrates were searched on a database and were evaluated for the GroE requirement in GroE-depleted cells. From this strategy, TatD and YjhH were identified as in vivo GroE obligate substrates.
3.3.4 Class IV Substrates as In Vivo Obligate GroE Substrates
Eight new proteins were added to the list of in vivo GroE obligate substrates. We have combined these 8 in vivo substrates with the 49 Class III+ members and now suggest their classification together to form a new group, the Class IV substrates (Table 15.1).
Taken together, a comprehensive assessment of the GroE requirement using a conditional GroE expression strain (MGM100) revealed that only ~ 60 % of Class III substrates are bona fide obligate GroE substrates in vivo, and renamed as Class III+ (49 proteins). The in vivo obligate substrates, combined with newly identified 8 GroE obligate substrates, were termed Class IV substrates, for which folding is obligatorily dependent on GroE in vivo (Table 15.1) [48].
Class IV substrates are restricted to proteins with molecular weights that could be encapsulated in the chaperonin cavity, are enriched in alanine/glycine residues, and have a strong structural preference for aggregation-prone folds. Notably, ~ 70 % of the Class IV substrates appear to be metabolic enzymes, supporting a hypothetical role of GroE in enzyme evolution.
3.3.5 GroE-Dependency of In Vitro Translated Class III− and Newly Identified Class IV Proteins.
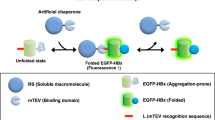
To elucidate the in vitro GroE-dependency of the newly identified substrates, a Class III− protein (FolE) and several Class IV proteins (DapA as a known obligate substrate, and SerC and KdsA as newly identified Class IV substrates) were translated by PURE system [15, 22, 29]. The requirements of the DnaK (DnaK, DnaJ, and GrpE) and GroE (GroEL and GroES) chaperone systems on the folding were monitored by the solubility and the appearance of folded structures, defined as a sharp band in native polyacrylamide gel electrophoresis (PAGE). FolE (Class III−) was soluble and formed a folded structure even in the absence of chaperones. In contrast, all of the Class IV proteins tested (DapA, SerC, and KdsA) were aggregation-prone without chaperones. The addition of the DnaK system increased the solubilities of the Class IV proteins to a greater or lesser extent, but the folded structures were not detected in native PAGE, implying that the soluble but unfolded structures in the presence of DnaK might be easily degraded in vivo. The Class IV proteins were soluble and formed folded structures only in the presence of GroE. The in vitro folding assay further confirmed our conclusion, in which the Class IV substrates, including the substrates that were not originally assigned as Class III (SerC and KdsA), stringently require GroE for correct folding.
3.3.6 Amino Acid Sequence Features of Class IV and III− Proteins
To define the features that are correlated with in vivo GroE dependency, physicochemical properties of the Class IV and III− proteins were compared. First, the molecular weights of the Class IV substrates were distributed normally, with a peak around 40 kDa, and ranging from 21 to 68 kDa, whereas the molecular weights of the 34 Class III− proteins ranged broadly, including 5 proteins smaller than 20 kDa and 4 proteins larger than 70 kDa, including ParC (84 kDa).
Second, the pI distribution and the hydrophobicity of Classes IV and III− were compared. The pI values of the Class IV substrates were distributed with a single peak around pI 5.8, whereas the pI distribution of the Class III− members was bimodal and similar to that of all cytosolic proteins of E. coli. The hydrophobicity distribution of the Class IV substrates was similar to that of all cytosolic proteins, whereas the Class III− proteins had lower hydrophobicity than either Class IV or all cytosolic proteins.
Third, the amino acid compositions of the Class IV and III− proteins were analyzed. As expected from the differences in the pI distributions and hydrophobicity, the amino acid compositions also differed between Classes IV and III−. Specifically, positively charged amino acids (arginine, lysine, and histidine) were enriched among the Class III− members. Neither hydrophobic amino acids (phenylalanine, tyrosine, tryptophan, isoleucine, leucine, and valine) nor other amino acids (negative, polar, neutral, and sulfur-containing amino acids) were enriched in either Class IV or III−.
3.3.7 Class IV Substrates Are Inherently Aggregation-Prone
Inherent solubility of Class IV substrates under chaperone-free conditions were already evaluated by global aggregation analysis using PURE system [29]. The histograms of the inherent solubilities of Classes IV and III− indicated a striking difference. The Class IV substrates were inherently highly aggregation-prone, whereas the Class III− proteins were broadly distributed from soluble to aggregation-prone.
3.3.8 Structural Features of Class IV Substrates
Previous studies on GroE substrates revealed that TIM barrel folds were substantially enriched in Class III proteins [47]. Surprisingly, all of the TIM barrel folds identified in the Class III proteins, except one (GatZ), were within the Class IV substrates (Table 15.1). As a result, the TIM barrel folds were further enriched in the Class IV substrates with 25 out of 57 Class IV substrates identified in this study possessing one (Table 15.1). This enrichment further supports the notion that the TIM barrel fold is correlated with GroE dependency [47, 58]. Not only TIM barrel folds (c.1 in SCOP database terminology [37]), but also FAD/NAD(P)-binding domains (c.3), PLP-dependent transferase like folds (c.67), and thiolase folds (c.95) were highly enriched in the class IV substrates, as compared with the frequency of their appearance in all cytosolic proteins. As Masters et al. pointed out in the review [58], the aldolase superfamily (c.1.10) subclass of TIM barrel folds is preferred among Class III proteins. In addition, metallo-dependent hydrolase (c.1.9) superfamily folds is also enriched within the Class IV members.
3.3.9 Class IV Homologs in a GroE-Lacking Organism
The genome of Ureaplasma urealyticum lacks the groELS gene [61]. Therefore, it is worth testing whether the Class IV homologs in Ureaplasma are GroE dependent. Five homologs of Class IV members (UuDeoA, UuCsdB, UuGatY, UuMetK, and UuYcfH) were found in Ureaplasma, by BLAST search. The genes encoding UuDeoA, UuMetK, and UuYcfH were cloned and overexpressed in MGM100, and their solubilities were assessed under GroE-depleted conditions. Strikingly, all of the Class IV homologs tested were soluble, even in the GroE-depleted cells, indicating that the GroE dependency was not conserved among the homologs. In addition, the S-adenosylmethionine synthase activity of UuMetK in the lysate of the GroE-depleted cells was comparable to that of the GroE-normal cells. Moreover, the leaky UuMetK expression suppressed the overexpression of MetE, which is one of the hallmarks of GroE-depleted cells [53, 58]. Collectively, UuMetK is active in the GroE-depleted cells. Intriguingly, the amino acid compositions of the Ureaplasma Class IV homologs revealed that the Ala/Gly fractions in all of the homologs were lower than those in their E. coli counterparts, whereas the contents of other amino acid groups, including aromatic, hydrophobic and positive amino acids, were indistinguishable from those in the E. coli counterparts. The amino acid content analysis again raises the possibility that a high Ala/Gly content might be involved in GroE dependency.
3.3.10 Possibility to Create GroE-Lacking E. coli
Verified obligate GroE substrates included only 6 genes (DapA, ASD, MetK, FtsE, HemB, and KdsA) essential for the viability of E. coli in rich medium. Although unidentified essential Class IV substrates may exist among the proteins that were not tested, we can predict the possible phenotypic defects caused by their inactivation in the GroE-depleted cells. If there are no further essential GroE-dependent proteins, then the complementation of these 6 essential genes by some means should generate an E. coli strain that can grow without groEL/ES. Such viable groEL/ES-knockout E. coli would provide the answer to the long-standing question of why GroE is essential for cell viability. Alternatively, the complementation of the 6 essential genes in E. coli lacking groEL/ES could be still lethal, due to the presence of unidentified essential Class IV substrates. In such a case, we can extend the phenotypic analysis to find the unidentified Class IV members, using the engineered E. coli.
3.3.11 Implication to Protein Evolution
Chaperones are known to provide a buffering system for genetic mutations, and thus promote genetic diversity [19, 62]. A recent quantitative assessment clearly showed that GroE promotes enzyme evolution by buffering the destabilizing mutations that confer improved enzymatic activities [21]. Since the destabilization of proteins generally results in their intracellular aggregation, aggregation-prone proteins, such as the Class IV substrates, could survive mutations if their aggregation is prevented by GroE, leading to the acquisition of diversity and/or the potential improvement of the enzymes in the Class IV substrates.
As already mentioned, many chaperonin GroE substrates are metabolic enzymes. However, the relationship between chaperonins and metabolism is still unclear. The distribution of GroE substrate enzymes in the metabolic network was investigated using network analysis techniques. Bioinformatics analysis revealed that as GroE requirement increases, substrate enzymes are more laterally distributed in the metabolic pathways [63]. In addition, comparative genome analysis showed that the GroE-dependent substrates were less conserved, suggesting that these substrates were acquired later on in evolutionary history [63]. This result implies the expansion of metabolic networks due to this chaperonin, and it supports the existing hypothesis of acceleration of evolution by chaperonins [64].
4 In Vivo Substrates of E. coli DnaK and Trigger Factor
DnaK , the major bacterial Hsp70 family is one of most abundant chaperones in the cytosol of E. coli. After an initial attempt to identify DnaK substrates in E. coli lacking trigger factor upon DnaK depletion [12], the direct isolation of DnaK-substrate complexes was conducted [65]. Quantitative MS-based proteomics revealed that DnaK interacts with at least ~ 700 proteins, including ~ 180 relatively aggregation-prone proteins. The study also focused on the role of DnaK in the E. coli chaperone network. Individual deletion of TF or depletion of GroE leads to specific changes in the DnaK interactome, suggesting that DnaK and other chaperones cooperates each other in E. coli cells. Taken together, Calloni et al. concluded that DnaK is a central organized of the chaperone network [65].
Regarding TF, modified ribosome profiling, which sequences messenger ribonucleic acid (mRNA) fragments covered by translating ribosomes, combined with a procedure to affinity purify ribosomes whose nascent polypeptides are associated with TF, called selective ribosome profiling, was used to reveal in vivo target proteins of TF [66]. Although TF can interact with many polypeptides, β-barrel outer membrane proteins are the most prominent substrates of TF [66].
5 Simulation of E. coli Chaperone Network
Proteins must fold into the unique native structures in a crowded environment in the cell. To understand the role of chaperones such as GroE or DnaK in the cell, we have to consider a balance between folding assisted by chaperones and degradation by proteases to maintain protein homeostasis, called proteostasis [2]. To gain insight into the interplay of processes and chaperones that maintain a functional proteome, a computational model called FoldEco was developed by Powers et al. [67].
FoldEco (http://foldeco.scripps.edu.) is the result of a joint effort to model how the proteostasis network affects protein folding in E. coli. E. coli was chosen as a model organism because its proteostasis networks are better characterized biochemically than those of any other organism. FoldEco simulates what happens to soluble proteins of interest as they are produced in the E. coli cytosol with five systems: protein synthesis and folding, DnaK system, GroE system, ClpB/DnaK disaggregation system, and protein degradation system. We can easily change parameter such as concentrations of chaperones in FoldEco to predict the fate of proteins of interest.
6 Concluding Remarks
After the emergence of chaperone concept in the late 1980s [68], extensive efforts have been dedicated to elucidate the molecular mechanism of chaperone proteins such as GroE or DnaK [2]. Compared to the understanding of chaperones as molecular machine, the role of chaperones in the cell still remains to be elucidated. Recent proteome-wide approaches from cell-free to in vivo would open the door to understand cellular functions of chaperones.
References
Anfinsen CB (1973) Principles that govern the folding of protein chains. Science 181:223–230
Hartl FU, Bracher A, Hayer-Hartl M (2011) Molecular chaperones in protein folding and proteostasis. Nature 475:324–332
Hoffmann A, Bukau B, Kramer G (2010) Structure and function of the molecular chaperone trigger factor. Biochim Biophys Acta 1803:650–661
Richter K, Haslbeck M, Buchner J (2010) The heat shock response: life on the verge of death. Mol Cell 40:253–266
Taguchi H (2005) Chaperonin GroEL meets the substrate protein as a “load” of the rings. J Biochem 137:543–549
Fayet O, Ziegelhoffer T, Georgopoulos C (1989) The groES and groEL heat shock gene products of Escherichia coli are essential for bacterial growth at all temperatures. J Bacteriol 171:1379–1385
Horwich AL, Low KB, Fenton WA, Hirshfield IN, Furtak K (1993) Folding in vivo of bacterial cytoplasmic proteins: role of GroEL. Cell 74:909–917
Xu Z, Horwich AL, Sigler PB (1997) The crystal structure of the asymmetric GroEL-GroES-(ADP)7 chaperonin complex. Nature 388:741–750
Sakikawa C, Taguchi H, Makino Y, Yoshida M (1999) On the maximum size of proteins to stay and fold in the cavity of GroEL underneath GroES. J Biol Chem 274:21251–21256
Deuerling E, Schulze-Specking A, Tomoyasu T, Mogk A, Bukau B (1999) Trigger factor and DnaK cooperate in folding of newly synthesized proteins. Nature 400:693–696
Teter SA, Houry WA, Ang D, Tradler T, Rockabrand D, Fischer G, Blum P, Georgopoulos C, Hartl FU (1999) Polypeptide flux through bacterial Hsp70: DnaK cooperates with trigger factor in chaperoning nascent chains. Cell 97:755–765
Deuerling E, Patzelt H, Vorderwulbecke S, Rauch T, Kramer G, Schaffitzel E, Mogk A, Schulze-Specking A, Langen H, Bukau B (2003) Trigger factor and DnaK possess overlapping substrate pools and binding specificities. Mol Microbiol 47:1317–1328
Vorderwulbecke S, Kramer G, Merz F, Kurz TA, Rauch T, Zachmann-Brand B, Bukau B, Deuerling E (2004) Low temperature or GroEL/ES overproduction permits growth of Escherichia coli cells lacking trigger factor and DnaK. FEBS Lett 559:181–187
Genevaux P, Keppel F, Schwager F, Langendijk-Genevaux PS, Hartl FU, Georgopoulos C (2004) In vivo analysis of the overlapping functions of DnaK and trigger factor. EMBO Rep 5:195–200
Ying BW, Taguchi H, Kondo M, Ueda T (2005) Co-translational involvement of the chaperonin GroEL in the folding of newly translated polypeptides. J Biol Chem 280:12035–12040
Ying BW, Taguchi H, Ueda T (2006) Co-translational binding of GroEL to nascent polypeptides is followed by post-translational encapsulation by GroES to mediate protein folding. J Biol Chem 281:21813–21819
Jenkins AJ, March JB, Oliver IR, Masters M (1986) A DNA fragment containing the groE genes can suppress mutations in the Escherichia coli dnaA gene. Mol Gen Genet 202(3):446–454
Van Dyk TK, Gatenby AA, LaRossa RA (1989) Demonstration by genetic suppression of interaction of GroE products with many proteins. Nature 342:451–453
Rutherford SL, Lindquist S (1998) Hsp90 as a capacitor for morphological evolution. Nature 396:336–342
Fares MA, Ruiz-Gonzalez MX, Moya A, Elena SF, Barrio E (2002) Endosymbiotic bacteria: groEL buffers against deleterious mutations. Nature 417:398
Tokuriki N, Oldfield CJ, Uversky VN, Berezovsky IN, Tawfik DS (2009) Do viral proteins possess unique biophysical features? Trends Biochem Sci 34:53–59
Shimizu Y, Inoue A, Tomari Y, Suzuki T, Yokogawa T, Nishikawa K, Ueda T (2001) Cell-free translation reconstituted with purified components. Nat Biotechnol 19:751–755
Shimizu Y, Kanamori T, Ueda T (2005) Protein synthesis by pure translation systems. Methods 36:299–304
Ying BW, Taguchi H, Ueda H, Ueda T (2004) Chaperone-assisted folding of a single-chain antibody in a reconstituted translation system. Biochem Biophys Res Commun 320:1359–1364
Chiti F, Taddei N, Baroni F, Capanni C, Stefani M, Ramponi G, Dobson CM (2002) Kinetic partitioning of protein folding and aggregation. Nat Struct Biol 9:137–143
Chiti F, Stefani M, Taddei N, Ramponi G, Dobson CM (2003) Rationalization of the effects of mutations on peptide and protein aggregation rates. Nature 424:805–808
Williams AD, Portelius E, Kheterpal I, Guo JT, Cook KD, Xu Y, Wetzel R (2004) Mapping abeta amyloid fibril secondary structure using scanning proline mutagenesis. J Mol Biol 335:833–842
de Groot NS, Aviles FX, Vendrell J, Ventura S (2006) Mutagenesis of the central hydrophobic cluster in Abeta42 Alzheimer’s peptide. Side-chain properties correlate with aggregation propensities. FEBS J 273:658–668
Niwa T, Ying BW, Saito K, Jin W, Takada S, Ueda T, Taguchi H (2009) Bimodal protein solubility distribution revealed by an aggregation analysis of the entire ensemble of Escherichia coli proteins. Proc Natl Acad Sci U S A 106:4201–4206
Kitagawa M, Ara T, Arifuzzaman M, Ioka-Nakamichi T, Inamoto E, Toyonaga H, Mori H (2005) Complete set of ORF clones of Escherichia coli ASKA library (a complete set of E. coli K-12 ORF archive): unique resources for biological research. DNA Res 12:291–299
Hopp TP, Woods KR (1981) Prediction of protein antigenic determinants from amino acid sequences. Proc Natl Acad Sci U S A 78:3824–3828
Kyte J, Doolittle RF (1982) A simple method for displaying the hydropathic character of a protein. J Mol Biol 157:105–132
Michelitsch MD, Weissman JS (2000) A census of glutamine/asparagine-rich regions: implications for their conserved function and the prediction of novel prions. Proc Natl Acad Sci U S A 97:11910–11915
Chou PY, Fasman GD (1974) Conformational parameters for amino acids in helical, beta-sheet, and random coil regions calculated from proteins. BioChemistry 13:211–222
Jones DT (1999) Protein secondary structure prediction based on position-specific scoring matrices. J Mol Biol 292:195–202
Bryson K, McGuffin LJ, Marsden RL, Ward JJ, Sodhi JS, Jones DT (2005) Protein structure prediction servers at University College London. Nucleic Acids Res 33:W36–W38
Murzin AG, Brenner SE, Hubbard T, Chothia C (1995) SCOP: a structural classification of proteins database for the investigation of sequences and structures. J Mol Biol 247:536–540
Fernandez-Escamilla AM, Rousseau F, Schymkowitz J, Serrano L (2004) Prediction of sequence-dependent and mutational effects on the aggregation of peptides and proteins. Nat Biotechnol 22:1302–1306
Conchillo-Sole O, de Groot NS, Aviles FX, Vendrell J, Daura X, Ventura S (2007) AGGRESCAN: a server for the prediction and evaluation of “hot spots” of aggregation in polypeptides. BMC Bioinformatics 8:65
Trovato A, Seno F, Tosatto SC (2007) The PASTA server for protein aggregation prediction. Protein Eng Des Sel 20:521–523
Agostini F, Vendruscolo M, Tartaglia GG (2012) Sequence-based prediction of protein solubility. J Mol Biol 421:237–241
Stiglic G, Kocbek S, Pernek I, Kokol P (2012) Comprehensive decision tree models in bioinformatics. PLoS ONE 7:e33812
Fang Y, Fang J (2013) Discrimination of soluble and aggregation-prone proteins based on sequence information. Mol Biosyst 9:806–811
Niwa T, Kanamori T, Ueda T, Taguchi H (2012) Global analysis of chaperone effects using a reconstituted cell-free translation system. Proc Natl Acad Sci U S A 109:8937–8942
Agashe VR, Guha S, Chang HC, Genevaux P, Hayer-Hartl M, Stemp M, Georgopoulos C, Hartl FU, Barral JM (2004) Function of trigger factor and DnaK in multidomain protein folding: increase in yield at the expense of folding speed. Cell 117:199–209
Hoffmann A, Merz F, Rutkowska A, Zachmann-Brand B, Deuerling E, Bukau B (2006) Trigger factor forms a protective shield for nascent polypeptides at the ribosome. J Biol Chem 281:6539–6545
Kerner MJ, Naylor DJ, Ishihama Y, Maier T, Chang HC, Stines AP, Georgopoulos C, Frishman D, Hayer-Hartl M, Mann M, Hartl FU (2005) Proteome-wide analysis of chaperonin-dependent protein folding in Escherichia coli. Cell 122:209–220
Fujiwara K, Ishihama Y, Nakahigashi K, Soga T, Taguchi H (2010) A systematic survey of in vivo obligate chaperonin-dependent substrates. EMBO J 29:1552–1564
Langer T, Lu C, Echols H, Flanagan J, Hayer MK, Hartl FU (1992) Successive action of DnaK, DnaJ and GroEL along the pathway of chaperone-mediated protein folding. Nature 356:683–689
McLennan N, Masters M (1998) GroE is vital for cell-wall synthesis. Nature 392:139
Fujiwara K, Taguchi H (2007) Filamentous morphology in GroE-depleted Escherichia coli induced by impaired folding of FtsE. J Bacteriol 189:5860–5866
Houry WA, Frishman D, Eckerskorn C, Lottspeich F, Hartl FU (1999) Identification of in vivo substrates of the chaperonin GroEL. Nature 402:147–154
Chapman E, Farr GW, Usaite R, Furtak K, Fenton WA, Chaudhuri TK, Hondorp ER, Matthews RG, Wolf SG, Yates JR, Pypaert M, Horwich AL (2006) Global aggregation of newly translated proteins in an Escherichia coli strain deficient of the chaperonin GroEL. Proc Natl Acad Sci U S A 103:15800–15805
Shimamura T, Koike-Takeshita A, Yokoyama K, Masui R, Murai N, Yoshida M, Taguchi H, Iwata S (2004) Crystal structure of the native chaperonin complex from Thermus thermophilus revealed unexpected asymmetry at the cis-cavity. Structure 12:1471–1480
Endo A, Kurusu Y (2007) Identification of in vivo substrates of the chaperonin GroEL from Bacillus subtilis. Biosci Biotechnol Biochem 71:1073–1077
Ishihama Y, Oda Y, Tabata T, Sato T, Nagasu T, Rappsilber J, Mann M (2005) Exponentially modified protein abundance index (emPAI) for estimation of absolute protein amount in proteomics by the number of sequenced peptides per protein. Mol Cell Proteomics 4:1265–1272
Ishihama Y, Schmidt T, Rappsilber J, Mann M, Hartl FU, Kerner MJ, Frishman D (2008) Protein abundance profiling of the Escherichia coli cytosol. BMC Genomics 9:102
Masters M, Blakely G, Coulson A, McLennan N, Yerko V, Acord J (2009) Protein folding in Escherichia coli: the chaperonin GroE and its substrates. Res Microbiol 160:267–277
Fujiwara K, Taguchi H (2012) Mechanism of methionine synthase overexpression in chaperonin-depleted Escherichia coli. Microbiology 158:917–924
El Yacoubi B, Bonnett S, Anderson JN, Swairjo MA, Iwata-Reuyl D, de Crecy-LagardV (2006) Discovery of a new prokaryotic type I GTP cyclohydrolase family. J Biol Chem 281:37586–37593
Glass JI, Lefkowitz EJ, Glass JS, Heiner CR, Chen EY, Cassell GH (2000) The complete sequence of the mucosal pathogen Ureaplasma urealyticum. Nature 407:757–762
Queitsch C, Sangster TA, Lindquist S (2002) Hsp90 as a capacitor of phenotypic variation. Nature 417:618–624
Takemoto K, Niwa T, Taguchi H (2011) Difference in the distribution pattern of substrate enzymes in the metabolic network of Escherichia coli, according to chaperonin requirement. BMC Syst Biol 5:98
Bogumil D, Dagan T (2012) Cumulative impact of chaperone-mediated folding on genome evolution. BioChemistry 51:9941–9953
Calloni G, Chen T, Schermann SM, Chang HC, Genevaux P, Agostini F, Tartaglia GG, Hayer-Hartl M, Hartl FU (2012) DnaK functions as a central hub in the E. coli chaperone network. Cell Rep 1:251–264
Oh E, Becker AH, Sandikci A, Huber D, Chaba R, Gloge F, Nichols RJ, Typas A, Gross CA, Kramer G, Weissman JS, Bukau B (2011) Selective ribosome profiling reveals the cotranslational chaperone action of trigger factor in vivo. Cell 147:1295–1308
Powers ET, Powers DL, Gierasch LM (2012) FoldEco: a model for proteostasis in E. coli. Cell Rep 1:265–276
Ellis J (1987) Proteins as molecular chaperones. Nature 328:378–379
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer Science+Business Media New York
About this chapter
Cite this chapter
Taguchi, H. (2014). The Interaction Networks of E. coli Chaperones. In: Houry, W. (eds) The Molecular Chaperones Interaction Networks in Protein Folding and Degradation. Interactomics and Systems Biology, vol 1. Springer, New York, NY. https://doi.org/10.1007/978-1-4939-1130-1_15
Download citation
DOI: https://doi.org/10.1007/978-1-4939-1130-1_15
Published:
Publisher Name: Springer, New York, NY
Print ISBN: 978-1-4939-1129-5
Online ISBN: 978-1-4939-1130-1
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)